Please be patient as the page loads
|
ARNT2_MOUSE
|
||||||
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ARNT2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.997952 (rank : 2) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 115 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.987210 (rank : 3) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.972909 (rank : 4) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | 1.95407e-87 (rank : 5) | NC score | 0.941129 (rank : 5) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | 6.28603e-86 (rank : 6) | NC score | 0.939326 (rank : 6) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
EPAS1_MOUSE
|
||||||
| θ value | 6.82597e-32 (rank : 7) | NC score | 0.793341 (rank : 10) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
EPAS1_HUMAN
|
||||||
| θ value | 9.8567e-31 (rank : 8) | NC score | 0.793388 (rank : 9) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
HIF1A_HUMAN
|
||||||
| θ value | 5.59698e-26 (rank : 9) | NC score | 0.779499 (rank : 14) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
HIF1A_MOUSE
|
||||||
| θ value | 1.24688e-25 (rank : 10) | NC score | 0.780485 (rank : 13) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
SIM2_HUMAN
|
||||||
| θ value | 2.77775e-25 (rank : 11) | NC score | 0.747171 (rank : 17) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
NPAS2_HUMAN
|
||||||
| θ value | 3.62785e-25 (rank : 12) | NC score | 0.793339 (rank : 11) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
NPAS2_MOUSE
|
||||||
| θ value | 3.62785e-25 (rank : 13) | NC score | 0.792977 (rank : 12) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
CLOCK_MOUSE
|
||||||
| θ value | 1.37858e-24 (rank : 14) | NC score | 0.796400 (rank : 7) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
SIM1_HUMAN
|
||||||
| θ value | 1.80048e-24 (rank : 15) | NC score | 0.751988 (rank : 15) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM1_MOUSE
|
||||||
| θ value | 5.23862e-24 (rank : 16) | NC score | 0.750912 (rank : 16) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
CLOCK_HUMAN
|
||||||
| θ value | 1.16704e-23 (rank : 17) | NC score | 0.795797 (rank : 8) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
SIM2_MOUSE
|
||||||
| θ value | 1.16704e-23 (rank : 18) | NC score | 0.746194 (rank : 18) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
NPAS3_MOUSE
|
||||||
| θ value | 4.29542e-18 (rank : 19) | NC score | 0.707527 (rank : 19) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_HUMAN
|
||||||
| θ value | 8.95645e-16 (rank : 20) | NC score | 0.700658 (rank : 20) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NCOA3_HUMAN
|
||||||
| θ value | 4.44505e-15 (rank : 21) | NC score | 0.575044 (rank : 27) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NCOA1_MOUSE
|
||||||
| θ value | 7.58209e-15 (rank : 22) | NC score | 0.610391 (rank : 25) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA1_HUMAN
|
||||||
| θ value | 1.29331e-14 (rank : 23) | NC score | 0.606337 (rank : 26) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NPAS1_MOUSE
|
||||||
| θ value | 4.16044e-13 (rank : 24) | NC score | 0.685983 (rank : 21) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
PER2_HUMAN
|
||||||
| θ value | 1.2105e-12 (rank : 25) | NC score | 0.481591 (rank : 34) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER2_MOUSE
|
||||||
| θ value | 1.58096e-12 (rank : 26) | NC score | 0.451576 (rank : 38) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
NPAS1_HUMAN
|
||||||
| θ value | 4.59992e-12 (rank : 27) | NC score | 0.681275 (rank : 22) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
NCOA3_MOUSE
|
||||||
| θ value | 7.84624e-12 (rank : 28) | NC score | 0.535772 (rank : 28) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
PER1_HUMAN
|
||||||
| θ value | 7.84624e-12 (rank : 29) | NC score | 0.473489 (rank : 35) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER1_MOUSE
|
||||||
| θ value | 7.84624e-12 (rank : 30) | NC score | 0.470832 (rank : 36) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 1.02475e-11 (rank : 31) | NC score | 0.454827 (rank : 37) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
PER3_MOUSE
|
||||||
| θ value | 4.30538e-10 (rank : 32) | NC score | 0.488945 (rank : 32) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
AHR_HUMAN
|
||||||
| θ value | 5.62301e-10 (rank : 33) | NC score | 0.663710 (rank : 23) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
NCOA2_MOUSE
|
||||||
| θ value | 1.38499e-08 (rank : 34) | NC score | 0.525450 (rank : 29) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
AHR_MOUSE
|
||||||
| θ value | 2.36244e-08 (rank : 35) | NC score | 0.648843 (rank : 24) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |
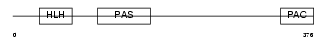
|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
NCOA2_HUMAN
|
||||||
| θ value | 2.36244e-08 (rank : 36) | NC score | 0.524259 (rank : 30) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NPAS4_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 37) | NC score | 0.506395 (rank : 31) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
USF2_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 38) | NC score | 0.258678 (rank : 41) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
USF1_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 39) | NC score | 0.266737 (rank : 40) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 40) | NC score | 0.266865 (rank : 39) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 41) | NC score | 0.249722 (rank : 42) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
MYCL1_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 42) | NC score | 0.188080 (rank : 52) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
HEY2_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 43) | NC score | 0.232755 (rank : 45) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 44) | NC score | 0.229817 (rank : 48) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
MYCL1_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 45) | NC score | 0.182085 (rank : 54) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
TFE3_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 46) | NC score | 0.239484 (rank : 43) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
NPAS4_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 47) | NC score | 0.485968 (rank : 33) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
TFEB_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 48) | NC score | 0.231864 (rank : 47) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 49) | NC score | 0.232153 (rank : 46) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
MITF_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 50) | NC score | 0.233970 (rank : 44) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 51) | NC score | 0.227230 (rank : 49) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
HEY1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 52) | NC score | 0.201094 (rank : 51) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HEY1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 53) | NC score | 0.208249 (rank : 50) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HAND1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 54) | NC score | 0.040197 (rank : 89) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q64279, Q61099 | Gene names | Hand1, Ehand, Hxt, Thing1 | |||
|
Domain Architecture |
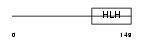
|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND) (Helix-loop-helix transcription factor expressed in extraembryonic mesoderm and trophoblast) (Thing-1) (Th1). | |||||
|
MAX_MOUSE
|
||||||
| θ value | 0.21417 (rank : 55) | NC score | 0.175834 (rank : 55) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
MYCN_HUMAN
|
||||||
| θ value | 0.21417 (rank : 56) | NC score | 0.128427 (rank : 63) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCN_MOUSE
|
||||||
| θ value | 0.21417 (rank : 57) | NC score | 0.124979 (rank : 67) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYC_MOUSE
|
||||||
| θ value | 0.21417 (rank : 58) | NC score | 0.122717 (rank : 71) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |
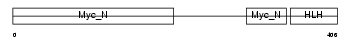
|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
BACH_MOUSE
|
||||||
| θ value | 0.279714 (rank : 59) | NC score | 0.041229 (rank : 88) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91V12, Q59HQ2 | Gene names | Acot7, Bach | |||
|
Domain Architecture |

|
|||||
| Description | Cytosolic acyl coenzyme A thioester hydrolase (EC 3.1.2.2) (Long chain acyl-CoA thioester hydrolase) (CTE-II) (CTE-IIa) (Brain acyl-CoA hydrolase) (Acyl-CoA thioesterase 7). | |||||
|
BHLH2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 60) | NC score | 0.126585 (rank : 65) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
MAD4_HUMAN
|
||||||
| θ value | 0.279714 (rank : 61) | NC score | 0.123879 (rank : 69) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
SRBP1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 62) | NC score | 0.153213 (rank : 59) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
ATF7_HUMAN
|
||||||
| θ value | 0.47712 (rank : 63) | NC score | 0.032898 (rank : 93) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P17544, Q13814 | Gene names | ATF7, ATFA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
ATF7_MOUSE
|
||||||
| θ value | 0.47712 (rank : 64) | NC score | 0.035620 (rank : 91) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R0S1 | Gene names | Atf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
BHLH2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 65) | NC score | 0.126702 (rank : 64) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
K0310_HUMAN
|
||||||
| θ value | 0.47712 (rank : 66) | NC score | 0.038807 (rank : 90) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
MAX_HUMAN
|
||||||
| θ value | 0.47712 (rank : 67) | NC score | 0.165157 (rank : 56) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
MYC_HUMAN
|
||||||
| θ value | 0.47712 (rank : 68) | NC score | 0.122811 (rank : 70) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
PELP1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 69) | NC score | 0.034914 (rank : 92) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
TCFL5_HUMAN
|
||||||
| θ value | 0.47712 (rank : 70) | NC score | 0.184628 (rank : 53) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
ZN485_HUMAN
|
||||||
| θ value | 0.47712 (rank : 71) | NC score | -0.001215 (rank : 123) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NCK3, Q96CL0 | Gene names | ZNF485 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 485. | |||||
|
TBX21_HUMAN
|
||||||
| θ value | 0.62314 (rank : 72) | NC score | 0.009348 (rank : 109) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UL17 | Gene names | TBX21, TBET, TBLYM | |||
|
Domain Architecture |

|
|||||
| Description | T-box transcription factor TBX21 (T-box protein 21) (Transcription factor TBLYM) (T-cell-specific T-box transcription factor T-bet). | |||||
|
DDX24_HUMAN
|
||||||
| θ value | 0.813845 (rank : 73) | NC score | 0.012441 (rank : 106) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9GZR7 | Gene names | DDX24 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX24 (EC 3.6.1.-) (DEAD box protein 24). | |||||
|
MYCS_MOUSE
|
||||||
| θ value | 0.813845 (rank : 74) | NC score | 0.124628 (rank : 68) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |
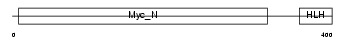
|
|||||
| Description | Protein S-myc. | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 0.813845 (rank : 75) | NC score | 0.023790 (rank : 96) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
K0240_HUMAN
|
||||||
| θ value | 1.06291 (rank : 76) | NC score | 0.026748 (rank : 95) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6AI39, Q5TFZ3, Q92514 | Gene names | KIAA0240 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0240. | |||||
|
PLEC1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 77) | NC score | 0.003254 (rank : 121) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
SRBP1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 78) | NC score | 0.135564 (rank : 60) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
ST2B1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 79) | NC score | 0.009110 (rank : 110) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35400, Q8K472, Q91V03 | Gene names | Sult2b1, Sult2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sulfotransferase family cytosolic 2B member 1 (EC 2.8.2.2) (Sulfotransferase 2B1) (Sulfotransferase 2B) (Alcohol sulfotransferase) (Hydroxysteroid sulfotransferase 2). | |||||
|
PELP1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 80) | NC score | 0.030156 (rank : 94) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
SRBP2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 81) | NC score | 0.125371 (rank : 66) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
PLCD1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 82) | NC score | 0.015514 (rank : 101) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R3B1 | Gene names | Plcd1, Plcd | |||
|
Domain Architecture |

|
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase delta 1 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (PLC-delta-1) (Phospholipase C-delta-1) (PLC-III). | |||||
|
SYNJ1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 83) | NC score | 0.013419 (rank : 105) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
BHLH3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 84) | NC score | 0.130895 (rank : 61) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BHLH3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 85) | NC score | 0.129075 (rank : 62) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
CASL_HUMAN
|
||||||
| θ value | 2.36792 (rank : 86) | NC score | 0.014331 (rank : 104) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14511 | Gene names | NEDD9, CASL | |||
|
Domain Architecture |
|
|||||
| Description | Enhancer of filamentation 1 (HEF1) (CRK-associated substrate-related protein) (CAS-L) (CasL) (p105) (Protein NEDD9) (NY-REN-12 antigen). | |||||
|
COKA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 87) | NC score | 0.004287 (rank : 118) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9P218, Q4VXQ4, Q8WUT2, Q96CY9, Q9BQU6, Q9BQU7 | Gene names | COL20A1, KIAA1510 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XX) chain precursor. | |||||
|
HAIR_MOUSE
|
||||||
| θ value | 3.0926 (rank : 88) | NC score | 0.009825 (rank : 108) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61645, Q80Y47 | Gene names | Hr | |||
|
Domain Architecture |

|
|||||
| Description | Protein hairless. | |||||
|
PDLI1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 89) | NC score | 0.006191 (rank : 115) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70400, Q99K93 | Gene names | Pdlim1, Clim1 | |||
|
Domain Architecture |
|
|||||
| Description | PDZ and LIM domain protein 1 (Elfin) (LIM domain protein CLP-36) (C- terminal LIM domain protein 1). | |||||
|
PGCA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 90) | NC score | 0.003938 (rank : 119) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
ZN228_HUMAN
|
||||||
| θ value | 3.0926 (rank : 91) | NC score | -0.002739 (rank : 124) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UJU3, Q9HCA7 | Gene names | ZNF228 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 228. | |||||
|
ASPP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.009023 (rank : 111) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q62415 | Gene names | Ppp1r13b, Aspp1 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 1 (Protein phosphatase 1 regulatory subunit 13B). | |||||
|
MAD3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.073191 (rank : 77) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 94) | NC score | 0.023741 (rank : 97) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
PSL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 95) | NC score | 0.011426 (rank : 107) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
ULK2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 96) | NC score | -0.004731 (rank : 128) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 979 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IYT8, O75119 | Gene names | ULK2, KIAA0623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK2 (EC 2.7.11.1) (Unc-51-like kinase 2). | |||||
|
ALEX_HUMAN
|
||||||
| θ value | 5.27518 (rank : 97) | NC score | 0.016391 (rank : 100) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P84996 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.021734 (rank : 99) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
MLX_MOUSE
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.161243 (rank : 57) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
OSBP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.006928 (rank : 114) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5QNQ6, Q8CF21 | Gene names | Osbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxysterol-binding protein 2. | |||||
|
R3HD2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.023057 (rank : 98) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
RHES_MOUSE
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | 0.003529 (rank : 120) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P63032, Q9WVD3 | Gene names | Rasd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTP-binding protein Rhes (Ras homolog enriched in striatum). | |||||
|
EPHA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | -0.002770 (rank : 125) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P21709, Q15405 | Gene names | EPHA1, EPH, EPHT, EPHT1 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-A receptor 1 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor EPH). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.004572 (rank : 117) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.007650 (rank : 112) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MLX_HUMAN
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.155944 (rank : 58) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
PDE8A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.006955 (rank : 113) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88502 | Gene names | Pde8a, Pde8 | |||
|
Domain Architecture |

|
|||||
| Description | High-affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8A (EC 3.1.4.17) (MMPDE8). | |||||
|
RHES_HUMAN
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.002790 (rank : 122) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96D21, O95520, Q5THY8 | Gene names | RASD2, TEM2 | |||
|
Domain Architecture |
|
|||||
| Description | GTP-binding protein Rhes (Ras homolog enriched in striatum) (Tumor endothelial marker 2). | |||||
|
FHOD1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.015346 (rank : 102) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
FHOD1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.015096 (rank : 103) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6P9Q4, Q8BMK2 | Gene names | Fhod1, Fhos1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
HES4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.096194 (rank : 72) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
MAD1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.053340 (rank : 83) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
PDE8A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.006147 (rank : 116) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60658, Q969I1, Q96PC9, Q96PD0, Q96PD1, Q9UMB7 | Gene names | PDE8A | |||
|
Domain Architecture |

|
|||||
| Description | High-affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8A (EC 3.1.4.17). | |||||
|
ZN473_MOUSE
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | -0.003593 (rank : 127) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 824 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BI67, Q8BI98, Q8BIB7 | Gene names | Znf473, Zfp100, Zfp473 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 473 homolog (Zinc finger protein 100) (Zfp-100). | |||||
|
ZNF7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | -0.003340 (rank : 126) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P17097, P17015 | Gene names | ZNF7, KOX4 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 7 (Zinc finger protein KOX4) (Zinc finger protein HF.16). | |||||
|
HES1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.074972 (rank : 76) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
HES1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.075318 (rank : 75) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.066273 (rank : 78) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
HES5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.095276 (rank : 73) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
HES7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.062514 (rank : 80) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.060670 (rank : 81) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
MAD3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.050258 (rank : 86) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80US8, Q60947 | Gene names | Mxd3, Mad3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3). | |||||
|
MAD4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.050680 (rank : 85) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q60948 | Gene names | Mxd4, Mad4 | |||
|
Domain Architecture |
|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MNT_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.063071 (rank : 79) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MNT_MOUSE
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.059998 (rank : 82) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MXI1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.050093 (rank : 87) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MXI1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.050982 (rank : 84) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MYCL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.076638 (rank : 74) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P12525 | Gene names | MYCL2 | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-2 protein. | |||||
|
ARNT2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 115 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_HUMAN
|
||||||
| NC score | 0.997952 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT_HUMAN
|
||||||
| NC score | 0.987210 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.972909 (rank : 4) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.941129 (rank : 5) | θ value | 1.95407e-87 (rank : 5) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.939326 (rank : 6) | θ value | 6.28603e-86 (rank : 6) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
CLOCK_MOUSE
|
||||||
| NC score | 0.796400 (rank : 7) | θ value | 1.37858e-24 (rank : 14) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
CLOCK_HUMAN
|
||||||
| NC score | 0.795797 (rank : 8) | θ value | 1.16704e-23 (rank : 17) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
EPAS1_HUMAN
|
||||||
| NC score | 0.793388 (rank : 9) | θ value | 9.8567e-31 (rank : 8) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
EPAS1_MOUSE
|
||||||
| NC score | 0.793341 (rank : 10) | θ value | 6.82597e-32 (rank : 7) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
NPAS2_HUMAN
|
||||||
| NC score | 0.793339 (rank : 11) | θ value | 3.62785e-25 (rank : 12) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
NPAS2_MOUSE
|
||||||
| NC score | 0.792977 (rank : 12) | θ value | 3.62785e-25 (rank : 13) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
HIF1A_MOUSE
|
||||||
| NC score | 0.780485 (rank : 13) | θ value | 1.24688e-25 (rank : 10) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
HIF1A_HUMAN
|
||||||
| NC score | 0.779499 (rank : 14) | θ value | 5.59698e-26 (rank : 9) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
SIM1_HUMAN
|
||||||
| NC score | 0.751988 (rank : 15) | θ value | 1.80048e-24 (rank : 15) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM1_MOUSE
|
||||||
| NC score | 0.750912 (rank : 16) | θ value | 5.23862e-24 (rank : 16) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
SIM2_HUMAN
|
||||||
| NC score | 0.747171 (rank : 17) | θ value | 2.77775e-25 (rank : 11) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
SIM2_MOUSE
|
||||||
| NC score | 0.746194 (rank : 18) | θ value | 1.16704e-23 (rank : 18) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
NPAS3_MOUSE
|
||||||
| NC score | 0.707527 (rank : 19) | θ value | 4.29542e-18 (rank : 19) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_HUMAN
|
||||||
| NC score | 0.700658 (rank : 20) | θ value | 8.95645e-16 (rank : 20) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS1_MOUSE
|
||||||
| NC score | 0.685983 (rank : 21) | θ value | 4.16044e-13 (rank : 24) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
NPAS1_HUMAN
|
||||||
| NC score | 0.681275 (rank : 22) | θ value | 4.59992e-12 (rank : 27) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
AHR_HUMAN
|
||||||
| NC score | 0.663710 (rank : 23) | θ value | 5.62301e-10 (rank : 33) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
AHR_MOUSE
|
||||||
| NC score | 0.648843 (rank : 24) | θ value | 2.36244e-08 (rank : 35) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
NCOA1_MOUSE
|
||||||
| NC score | 0.610391 (rank : 25) | θ value | 7.58209e-15 (rank : 22) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA1_HUMAN
|
||||||
| NC score | 0.606337 (rank : 26) | θ value | 1.29331e-14 (rank : 23) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NCOA3_HUMAN
|
||||||
| NC score | 0.575044 (rank : 27) | θ value | 4.44505e-15 (rank : 21) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NCOA3_MOUSE
|
||||||
| NC score | 0.535772 (rank : 28) | θ value | 7.84624e-12 (rank : 28) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
NCOA2_MOUSE
|
||||||
| NC score | 0.525450 (rank : 29) | θ value | 1.38499e-08 (rank : 34) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
NCOA2_HUMAN
|
||||||
| NC score | 0.524259 (rank : 30) | θ value | 2.36244e-08 (rank : 36) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NPAS4_HUMAN
|
||||||
| NC score | 0.506395 (rank : 31) | θ value | 3.19293e-05 (rank : 37) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
PER3_MOUSE
|
||||||
| NC score | 0.488945 (rank : 32) | θ value | 4.30538e-10 (rank : 32) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
NPAS4_MOUSE
|
||||||
| NC score | 0.485968 (rank : 33) | θ value | 0.00869519 (rank : 47) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
PER2_HUMAN
|
||||||
| NC score | 0.481591 (rank : 34) | θ value | 1.2105e-12 (rank : 25) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER1_HUMAN
|
||||||
| NC score | 0.473489 (rank : 35) | θ value | 7.84624e-12 (rank : 29) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER1_MOUSE
|
||||||
| NC score | 0.470832 (rank : 36) | θ value | 7.84624e-12 (rank : 30) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.454827 (rank : 37) | θ value | 1.02475e-11 (rank : 31) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
PER2_MOUSE
|
||||||
| NC score | 0.451576 (rank : 38) | θ value | 1.58096e-12 (rank : 26) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.266865 (rank : 39) | θ value | 0.000786445 (rank : 40) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.266737 (rank : 40) | θ value | 0.000786445 (rank : 39) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.258678 (rank : 41) | θ value | 0.000602161 (rank : 38) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.249722 (rank : 42) | θ value | 0.00102713 (rank : 41) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.239484 (rank : 43) | θ value | 0.00665767 (rank : 46) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.233970 (rank : 44) | θ value | 0.0113563 (rank : 50) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
HEY2_HUMAN
|
||||||
| NC score | 0.232755 (rank : 45) | θ value | 0.00298849 (rank : 43) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.232153 (rank : 46) | θ value | 0.00869519 (rank : 49) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.231864 (rank : 47) | θ value | 0.00869519 (rank : 48) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.229817 (rank : 48) | θ value | 0.00298849 (rank : 44) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.227230 (rank : 49) | θ value | 0.0431538 (rank : 51) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
HEY1_MOUSE
|
||||||
| NC score | 0.208249 (rank : 50) | θ value | 0.0736092 (rank : 53) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HEY1_HUMAN
|
||||||
| NC score | 0.201094 (rank : 51) | θ value | 0.0736092 (rank : 52) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
MYCL1_MOUSE
|
||||||
| NC score | 0.188080 (rank : 52) | θ value | 0.00175202 (rank : 42) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
TCFL5_HUMAN
|
||||||
| NC score | 0.184628 (rank : 53) | θ value | 0.47712 (rank : 70) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
MYCL1_HUMAN
|
||||||
| NC score | 0.182085 (rank : 54) | θ value | 0.00390308 (rank : 45) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MAX_MOUSE
|
||||||
| NC score | 0.175834 (rank : 55) | θ value | 0.21417 (rank : 55) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
MAX_HUMAN
|
||||||
| NC score | 0.165157 (rank : 56) | θ value | 0.47712 (rank : 67) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.161243 (rank : 57) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.155944 (rank : 58) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
SRBP1_HUMAN
|
||||||
| NC score | 0.153213 (rank : 59) | θ value | 0.279714 (rank : 62) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
SRBP1_MOUSE
|
||||||
| NC score | 0.135564 (rank : 60) | θ value | 1.06291 (rank : 78) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
BHLH3_HUMAN
|
||||||
| NC score | 0.130895 (rank : 61) | θ value | 2.36792 (rank : 84) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BHLH3_MOUSE
|
||||||
| NC score | 0.129075 (rank : 62) | θ value | 2.36792 (rank : 85) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
MYCN_HUMAN
|
||||||
| NC score | 0.128427 (rank : 63) | θ value | 0.21417 (rank : 56) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
BHLH2_HUMAN
|
||||||
| NC score | 0.126702 (rank : 64) | θ value | 0.47712 (rank : 65) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
BHLH2_MOUSE
|
||||||
| NC score | 0.126585 (rank : 65) | θ value | 0.279714 (rank : 60) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
SRBP2_HUMAN
|
||||||
| NC score | 0.125371 (rank : 66) | θ value | 1.38821 (rank : 81) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
MYCN_MOUSE
|
||||||
| NC score | 0.124979 (rank : 67) | θ value | 0.21417 (rank : 57) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCS_MOUSE
|
||||||
| NC score | 0.124628 (rank : 68) | θ value | 0.813845 (rank : 74) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |

|
|||||
| Description | Protein S-myc. | |||||
|
MAD4_HUMAN
|
||||||
| NC score | 0.123879 (rank : 69) | θ value | 0.279714 (rank : 61) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MYC_HUMAN
|
||||||
| NC score | 0.122811 (rank : 70) | θ value | 0.47712 (rank : 68) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYC_MOUSE
|
||||||
| NC score | 0.122717 (rank : 71) | θ value | 0.21417 (rank : 58) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
HES4_HUMAN
|
||||||
| NC score | 0.096194 (rank : 72) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
HES5_MOUSE
|
||||||
| NC score | 0.095276 (rank : 73) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
MYCL2_HUMAN
|
||||||
| NC score | 0.076638 (rank : 74) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P12525 | Gene names | MYCL2 | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-2 protein. | |||||
|
HES1_MOUSE
|
||||||
| NC score | 0.075318 (rank : 75) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES1_HUMAN
|
||||||
| NC score | 0.074972 (rank : 76) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
MAD3_HUMAN
|
||||||
| NC score | 0.073191 (rank : 77) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
HES3_MOUSE
|
||||||
| NC score | 0.066273 (rank : 78) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
MNT_HUMAN
|
||||||
| NC score | 0.063071 (rank : 79) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
HES7_HUMAN
|
||||||
| NC score | 0.062514 (rank : 80) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES7_MOUSE
|
||||||
| NC score | 0.060670 (rank : 81) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
MNT_MOUSE
|
||||||
| NC score | 0.059998 (rank : 82) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MAD1_MOUSE
|
||||||
| NC score | 0.053340 (rank : 83) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MXI1_MOUSE
|
||||||
| NC score | 0.050982 (rank : 84) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MAD4_MOUSE
|
||||||
| NC score | 0.050680 (rank : 85) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q60948 | Gene names | Mxd4, Mad4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MAD3_MOUSE
|
||||||
| NC score | 0.050258 (rank : 86) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80US8, Q60947 | Gene names | Mxd3, Mad3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3). | |||||
|
MXI1_HUMAN
|
||||||
| NC score | 0.050093 (rank : 87) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
BACH_MOUSE
|
||||||
| NC score | 0.041229 (rank : 88) | θ value | 0.279714 (rank : 59) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91V12, Q59HQ2 | Gene names | Acot7, Bach | |||
|
Domain Architecture |

|
|||||
| Description | Cytosolic acyl coenzyme A thioester hydrolase (EC 3.1.2.2) (Long chain acyl-CoA thioester hydrolase) (CTE-II) (CTE-IIa) (Brain acyl-CoA hydrolase) (Acyl-CoA thioesterase 7). | |||||
|
HAND1_MOUSE
|
||||||
| NC score | 0.040197 (rank : 89) | θ value | 0.21417 (rank : 54) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q64279, Q61099 | Gene names | Hand1, Ehand, Hxt, Thing1 | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND) (Helix-loop-helix transcription factor expressed in extraembryonic mesoderm and trophoblast) (Thing-1) (Th1). | |||||
|
K0310_HUMAN
|
||||||
| NC score | 0.038807 (rank : 90) | θ value | 0.47712 (rank : 66) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
ATF7_MOUSE
|
||||||
| NC score | 0.035620 (rank : 91) | θ value | 0.47712 (rank : 64) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R0S1 | Gene names | Atf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
PELP1_HUMAN
|
||||||
| NC score | 0.034914 (rank : 92) | θ value | 0.47712 (rank : 69) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
ATF7_HUMAN
|
||||||
| NC score | 0.032898 (rank : 93) | θ value | 0.47712 (rank : 63) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P17544, Q13814 | Gene names | ATF7, ATFA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.030156 (rank : 94) | θ value | 1.38821 (rank : 80) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
K0240_HUMAN
|
||||||
| NC score | 0.026748 (rank : 95) | θ value | 1.06291 (rank : 76) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6AI39, Q5TFZ3, Q92514 | Gene names | KIAA0240 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0240. | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.023790 (rank : 96) | θ value | 0.813845 (rank : 75) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.023741 (rank : 97) | θ value | 4.03905 (rank : 94) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
R3HD2_MOUSE
|
||||||
| NC score | 0.023057 (rank : 98) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.021734 (rank : 99) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
ALEX_HUMAN
|
||||||
| NC score | 0.016391 (rank : 100) | θ value | 5.27518 (rank : 97) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P84996 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
PLCD1_MOUSE
|
||||||
| NC score | 0.015514 (rank : 101) | θ value | 1.81305 (rank : 82) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R3B1 | Gene names | Plcd1, Plcd | |||
|
Domain Architecture |

|
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase delta 1 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (PLC-delta-1) (Phospholipase C-delta-1) (PLC-III). | |||||
|
FHOD1_HUMAN
|
||||||
| NC score | 0.015346 (rank : 102) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
FHOD1_MOUSE
|
||||||
| NC score | 0.015096 (rank : 103) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6P9Q4, Q8BMK2 | Gene names | Fhod1, Fhos1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
CASL_HUMAN
|
||||||
| NC score | 0.014331 (rank : 104) | θ value | 2.36792 (rank : 86) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14511 | Gene names | NEDD9, CASL | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of filamentation 1 (HEF1) (CRK-associated substrate-related protein) (CAS-L) (CasL) (p105) (Protein NEDD9) (NY-REN-12 antigen). | |||||
|
SYNJ1_MOUSE
|
||||||
| NC score | 0.013419 (rank : 105) | θ value | 1.81305 (rank : 83) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
DDX24_HUMAN
|
||||||
| NC score | 0.012441 (rank : 106) | θ value | 0.813845 (rank : 73) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9GZR7 | Gene names | DDX24 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX24 (EC 3.6.1.-) (DEAD box protein 24). | |||||
|
PSL1_HUMAN
|
||||||
| NC score | 0.011426 (rank : 107) | θ value | 4.03905 (rank : 95) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
HAIR_MOUSE
|
||||||
| NC score | 0.009825 (rank : 108) | θ value | 3.0926 (rank : 88) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61645, Q80Y47 | Gene names | Hr | |||
|
Domain Architecture |

|
|||||
| Description | Protein hairless. | |||||
|
TBX21_HUMAN
|
||||||
| NC score | 0.009348 (rank : 109) | θ value | 0.62314 (rank : 72) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UL17 | Gene names | TBX21, TBET, TBLYM | |||
|
Domain Architecture |

|
|||||
| Description | T-box transcription factor TBX21 (T-box protein 21) (Transcription factor TBLYM) (T-cell-specific T-box transcription factor T-bet). | |||||
|
ST2B1_MOUSE
|
||||||
| NC score | 0.009110 (rank : 110) | θ value | 1.06291 (rank : 79) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35400, Q8K472, Q91V03 | Gene names | Sult2b1, Sult2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sulfotransferase family cytosolic 2B member 1 (EC 2.8.2.2) (Sulfotransferase 2B1) (Sulfotransferase 2B) (Alcohol sulfotransferase) (Hydroxysteroid sulfotransferase 2). | |||||
|
ASPP1_MOUSE
|
||||||
| NC score | 0.009023 (rank : 111) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q62415 | Gene names | Ppp1r13b, Aspp1 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 1 (Protein phosphatase 1 regulatory subunit 13B). | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.007650 (rank : 112) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PDE8A_MOUSE
|
||||||
| NC score | 0.006955 (rank : 113) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88502 | Gene names | Pde8a, Pde8 | |||
|
Domain Architecture |

|
|||||
| Description | High-affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8A (EC 3.1.4.17) (MMPDE8). | |||||
|
OSBP2_MOUSE
|
||||||
| NC score | 0.006928 (rank : 114) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5QNQ6, Q8CF21 | Gene names | Osbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxysterol-binding protein 2. | |||||
|
PDLI1_MOUSE
|
||||||
| NC score | 0.006191 (rank : 115) | θ value | 3.0926 (rank : 89) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70400, Q99K93 | Gene names | Pdlim1, Clim1 | |||
|
Domain Architecture |

|
|||||
| Description | PDZ and LIM domain protein 1 (Elfin) (LIM domain protein CLP-36) (C- terminal LIM domain protein 1). | |||||
|
PDE8A_HUMAN
|
||||||
| NC score | 0.006147 (rank : 116) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60658, Q969I1, Q96PC9, Q96PD0, Q96PD1, Q9UMB7 | Gene names | PDE8A | |||
|
Domain Architecture |

|
|||||
| Description | High-affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8A (EC 3.1.4.17). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.004572 (rank : 117) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
COKA1_HUMAN
|
||||||
| NC score | 0.004287 (rank : 118) | θ value | 2.36792 (rank : 87) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9P218, Q4VXQ4, Q8WUT2, Q96CY9, Q9BQU6, Q9BQU7 | Gene names | COL20A1, KIAA1510 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XX) chain precursor. | |||||
|
PGCA_HUMAN
|
||||||
| NC score | 0.003938 (rank : 119) | θ value | 3.0926 (rank : 90) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
RHES_MOUSE
|
||||||
| NC score | 0.003529 (rank : 120) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P63032, Q9WVD3 | Gene names | Rasd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTP-binding protein Rhes (Ras homolog enriched in striatum). | |||||
|
PLEC1_HUMAN
|
||||||
| NC score | 0.003254 (rank : 121) | θ value | 1.06291 (rank : 77) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
RHES_HUMAN
|
||||||
| NC score | 0.002790 (rank : 122) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96D21, O95520, Q5THY8 | Gene names | RASD2, TEM2 | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein Rhes (Ras homolog enriched in striatum) (Tumor endothelial marker 2). | |||||
|
ZN485_HUMAN
|
||||||
| NC score | -0.001215 (rank : 123) | θ value | 0.47712 (rank : 71) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NCK3, Q96CL0 | Gene names | ZNF485 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 485. | |||||
|
ZN228_HUMAN
|
||||||
| NC score | -0.002739 (rank : 124) | θ value | 3.0926 (rank : 91) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UJU3, Q9HCA7 | Gene names | ZNF228 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 228. | |||||
|
EPHA1_HUMAN
|
||||||
| NC score | -0.002770 (rank : 125) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P21709, Q15405 | Gene names | EPHA1, EPH, EPHT, EPHT1 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-A receptor 1 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor EPH). | |||||
|
ZNF7_HUMAN
|
||||||
| NC score | -0.003340 (rank : 126) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P17097, P17015 | Gene names | ZNF7, KOX4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 7 (Zinc finger protein KOX4) (Zinc finger protein HF.16). | |||||
|
ZN473_MOUSE
|
||||||
| NC score | -0.003593 (rank : 127) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 824 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BI67, Q8BI98, Q8BIB7 | Gene names | Znf473, Zfp100, Zfp473 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 473 homolog (Zinc finger protein 100) (Zfp-100). | |||||
|
ULK2_HUMAN
|
||||||
| NC score | -0.004731 (rank : 128) | θ value | 4.03905 (rank : 96) | |||
| Query Neighborhood Hits | 115 | Target Neighborhood Hits | 979 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IYT8, O75119 | Gene names | ULK2, KIAA0623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK2 (EC 2.7.11.1) (Unc-51-like kinase 2). | |||||