Please be patient as the page loads
|
EPAS1_MOUSE
|
||||||
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
EPAS1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.994197 (rank : 2) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
EPAS1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 94 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
HIF1A_HUMAN
|
||||||
| θ value | 2.04462e-161 (rank : 3) | NC score | 0.975288 (rank : 4) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
HIF1A_MOUSE
|
||||||
| θ value | 2.04462e-161 (rank : 4) | NC score | 0.977646 (rank : 3) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
SIM2_MOUSE
|
||||||
| θ value | 2.32439e-64 (rank : 5) | NC score | 0.923554 (rank : 5) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
SIM1_HUMAN
|
||||||
| θ value | 3.03575e-64 (rank : 6) | NC score | 0.921705 (rank : 7) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM1_MOUSE
|
||||||
| θ value | 3.35642e-63 (rank : 7) | NC score | 0.920668 (rank : 8) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
SIM2_HUMAN
|
||||||
| θ value | 4.10295e-61 (rank : 8) | NC score | 0.923527 (rank : 6) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
NPAS3_HUMAN
|
||||||
| θ value | 1.36585e-56 (rank : 9) | NC score | 0.899765 (rank : 11) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_MOUSE
|
||||||
| θ value | 1.78386e-56 (rank : 10) | NC score | 0.899814 (rank : 10) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS1_MOUSE
|
||||||
| θ value | 1.23823e-49 (rank : 11) | NC score | 0.900850 (rank : 9) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
NPAS1_HUMAN
|
||||||
| θ value | 6.7944e-48 (rank : 12) | NC score | 0.899572 (rank : 12) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
ARNT2_MOUSE
|
||||||
| θ value | 6.82597e-32 (rank : 13) | NC score | 0.793341 (rank : 21) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_HUMAN
|
||||||
| θ value | 8.91499e-32 (rank : 14) | NC score | 0.794139 (rank : 20) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | 1.98606e-31 (rank : 15) | NC score | 0.829322 (rank : 13) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | 3.3877e-31 (rank : 16) | NC score | 0.826994 (rank : 14) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
ARNT_HUMAN
|
||||||
| θ value | 2.86786e-30 (rank : 17) | NC score | 0.788681 (rank : 23) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
NPAS2_MOUSE
|
||||||
| θ value | 2.86786e-30 (rank : 18) | NC score | 0.796372 (rank : 19) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
CLOCK_HUMAN
|
||||||
| θ value | 4.89182e-30 (rank : 19) | NC score | 0.807441 (rank : 17) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | 8.3442e-30 (rank : 20) | NC score | 0.775106 (rank : 24) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
NPAS2_HUMAN
|
||||||
| θ value | 1.4233e-29 (rank : 21) | NC score | 0.789720 (rank : 22) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
CLOCK_MOUSE
|
||||||
| θ value | 4.14116e-29 (rank : 22) | NC score | 0.803980 (rank : 18) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
AHR_MOUSE
|
||||||
| θ value | 4.43474e-23 (rank : 23) | NC score | 0.812231 (rank : 16) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |
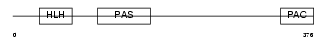
|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
AHR_HUMAN
|
||||||
| θ value | 9.87957e-23 (rank : 24) | NC score | 0.815426 (rank : 15) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
NPAS4_HUMAN
|
||||||
| θ value | 1.13037e-18 (rank : 25) | NC score | 0.678107 (rank : 25) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
NPAS4_MOUSE
|
||||||
| θ value | 1.24977e-17 (rank : 26) | NC score | 0.661451 (rank : 26) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
NCOA2_MOUSE
|
||||||
| θ value | 6.41864e-14 (rank : 27) | NC score | 0.567186 (rank : 30) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
NCOA2_HUMAN
|
||||||
| θ value | 1.09485e-13 (rank : 28) | NC score | 0.566473 (rank : 31) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NCOA3_HUMAN
|
||||||
| θ value | 7.84624e-12 (rank : 29) | NC score | 0.569862 (rank : 29) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NCOA1_HUMAN
|
||||||
| θ value | 1.74796e-11 (rank : 30) | NC score | 0.617665 (rank : 28) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NCOA1_MOUSE
|
||||||
| θ value | 3.89403e-11 (rank : 31) | NC score | 0.620193 (rank : 27) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA3_MOUSE
|
||||||
| θ value | 3.29651e-10 (rank : 32) | NC score | 0.533502 (rank : 32) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
PER1_MOUSE
|
||||||
| θ value | 3.77169e-06 (rank : 33) | NC score | 0.325698 (rank : 35) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
PER3_MOUSE
|
||||||
| θ value | 6.43352e-06 (rank : 34) | NC score | 0.348456 (rank : 33) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
PER1_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 35) | NC score | 0.326997 (rank : 34) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 36) | NC score | 0.313189 (rank : 37) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
PER2_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 37) | NC score | 0.319170 (rank : 36) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER2_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 38) | NC score | 0.295480 (rank : 38) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
DDEF2_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 39) | NC score | 0.021747 (rank : 64) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 599 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43150 | Gene names | DDEF2, KIAA0400 | |||
|
Domain Architecture |

|
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
ROBO3_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 40) | NC score | 0.007821 (rank : 89) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96MS0 | Gene names | ROBO3 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Roundabout-like protein 3). | |||||
|
SETB1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 41) | NC score | 0.022215 (rank : 63) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88974, Q922K1 | Gene names | Setdb1, Eset | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.365318 (rank : 42) | NC score | 0.031434 (rank : 58) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
NF2L2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 43) | NC score | 0.022972 (rank : 62) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q16236, Q96F71 | Gene names | NFE2L2, NRF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor erythroid 2-related factor 2 (NF-E2-related factor 2) (NFE2-related factor 2) (Nuclear factor, erythroid derived 2, like 2) (HEBP1). | |||||
|
ERF_HUMAN
|
||||||
| θ value | 0.62314 (rank : 44) | NC score | 0.017675 (rank : 72) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P50548, Q9UPI7 | Gene names | ERF | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF (Ets2 repressor factor). | |||||
|
ERF_MOUSE
|
||||||
| θ value | 0.62314 (rank : 45) | NC score | 0.017856 (rank : 70) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
HLF_MOUSE
|
||||||
| θ value | 0.62314 (rank : 46) | NC score | 0.026412 (rank : 59) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BW74, Q6PF83 | Gene names | Hlf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatic leukemia factor. | |||||
|
ENAH_MOUSE
|
||||||
| θ value | 0.813845 (rank : 47) | NC score | 0.017381 (rank : 73) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
HLF_HUMAN
|
||||||
| θ value | 0.813845 (rank : 48) | NC score | 0.025818 (rank : 60) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q16534 | Gene names | HLF | |||
|
Domain Architecture |

|
|||||
| Description | Hepatic leukemia factor. | |||||
|
JPH4_HUMAN
|
||||||
| θ value | 0.813845 (rank : 49) | NC score | 0.012692 (rank : 80) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96JJ6, Q8ND44, Q96DQ0 | Gene names | JPH4, JPHL1, KIAA1831 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-4 (Junctophilin-like 1 protein). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.021469 (rank : 66) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
DRD3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 51) | NC score | 0.000490 (rank : 109) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P30728 | Gene names | Drd3 | |||
|
Domain Architecture |

|
|||||
| Description | D(3) dopamine receptor. | |||||
|
GPS2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 52) | NC score | 0.019917 (rank : 68) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13227 | Gene names | GPS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein pathway suppressor 2 (Protein GPS2). | |||||
|
PASK_HUMAN
|
||||||
| θ value | 1.38821 (rank : 53) | NC score | 0.005154 (rank : 97) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 857 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96RG2, Q86XH6, Q99763, Q9UFR7 | Gene names | PASK, KIAA0135 | |||
|
Domain Architecture |

|
|||||
| Description | PAS domain-containing serine/threonine-protein kinase (EC 2.7.11.1) (PAS-kinase) (PASKIN) (hPASK). | |||||
|
SYN3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.014985 (rank : 75) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8JZP2 | Gene names | Syn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
CDC20_MOUSE
|
||||||
| θ value | 2.36792 (rank : 55) | NC score | 0.005886 (rank : 94) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JJ66, Q3TGP1, Q8BPG4, Q99LK3 | Gene names | Cdc20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC) (mmCdc20). | |||||
|
CSF1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 56) | NC score | 0.016649 (rank : 74) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P07141, Q8R3C8 | Gene names | Csf1, Csfm | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage colony-stimulating factor 1 precursor (CSF-1) (MCSF). | |||||
|
NF2L2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 57) | NC score | 0.017720 (rank : 71) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60795 | Gene names | Nfe2l2, Nrf-2, Nrf2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor erythroid 2-related factor 2 (NF-E2-related factor 2) (NFE2-related factor 2) (Nuclear factor, erythroid derived 2, like 2). | |||||
|
ZEP2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 58) | NC score | 0.001046 (rank : 106) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3UHF7, O55140, Q3UVD4, Q3UVH5 | Gene names | Hivep2, Mibp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 homolog (Myc intron-binding protein 1) (MIBP-1). | |||||
|
CNOT3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.012095 (rank : 82) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75175, Q9NZN7, Q9UF76 | Gene names | CNOT3, KIAA0691, NOT3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
DDEF1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.014078 (rank : 78) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QWY8, O08612, Q80TG8, Q80UV6, Q99LV8, Q9Z2B6 | Gene names | Ddef1, Asap1, Kiaa1249, Shag1 | |||
|
Domain Architecture |

|
|||||
| Description | 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase- activating protein (PIP2-dependent ARF1 GAP) (ADP-ribosylation factor- directed GTPase-activating protein 1) (ARF GTPase-activating protein 1) (Development and differentiation-enhancing factor 1) (Differentiation-enhancing factor 1) (DEF-1). | |||||
|
K1802_HUMAN
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.013125 (rank : 79) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
K1802_MOUSE
|
||||||
| θ value | 3.0926 (rank : 62) | NC score | 0.008445 (rank : 87) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
HES5_MOUSE
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.065676 (rank : 54) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
KCNH3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.004394 (rank : 99) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULD8, Q9UQ06 | Gene names | KCNH3, KIAA1282 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (ELK2) (Brain-specific eag-like channel 1) (BEC1). | |||||
|
MAGC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.032762 (rank : 57) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O60732, O75451, Q8TCV4 | Gene names | MAGEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen C1 (MAGE-C1 antigen) (Cancer-testis antigen CT7). | |||||
|
NEST_HUMAN
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.001012 (rank : 107) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
PASK_MOUSE
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.003045 (rank : 102) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 867 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CEE6, Q91YD8 | Gene names | Pask, Kiaa0135 | |||
|
Domain Architecture |

|
|||||
| Description | PAS domain-containing serine/threonine-protein kinase (EC 2.7.11.1) (PAS-kinase) (PASKIN). | |||||
|
R3HD1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.014469 (rank : 76) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15032, Q8IW32 | Gene names | R3HDM1, KIAA0029, R3HDM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R3H domain-containing protein 1. | |||||
|
SYP2L_HUMAN
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.008360 (rank : 88) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H987 | Gene names | SYNPO2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin 2-like protein. | |||||
|
AMELX_MOUSE
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.012294 (rank : 81) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P63277, P45559, P70592, Q61293 | Gene names | Amelx, Amel | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amelogenin, X isoform precursor (Leucine-rich amelogenin peptide) (LRAP). | |||||
|
CENG1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.007748 (rank : 90) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3UHD9, Q5DU45 | Gene names | Centg1, Agap2, Kiaa0167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-gamma 1 (ARF-GAP with GTP-binding protein-like, ankyrin repeat and pleckstrin homology domains 2) (AGAP-2) (Phosphatidylinositol-3-kinase enhancer) (PIKE). | |||||
|
SPG16_MOUSE
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.003314 (rank : 101) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8K450, Q8C450, Q9CPU8, Q9D522 | Gene names | Spag16, Pf20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 16 protein (PF20 protein homolog). | |||||
|
SSPO_MOUSE
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.000847 (rank : 108) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
TENX_HUMAN
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.000077 (rank : 110) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P22105, P78530, P78531, Q08424, Q9UMG7 | Gene names | TNXB, HXBL, TNX, XB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-X precursor (TN-X) (Hexabrachion-like protein). | |||||
|
TP53B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.010048 (rank : 83) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
3BP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.004960 (rank : 98) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y3L3, Q6IBZ2, Q6ZVL9, Q96HQ5, Q9NSQ9 | Gene names | SH3BP1 | |||
|
Domain Architecture |
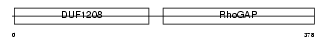
|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.021030 (rank : 67) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
BCL9_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.021620 (rank : 65) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
CENG1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.008602 (rank : 85) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 544 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99490, O00578, Q548E0, Q8IWU3 | Gene names | CENTG1, AGAP2, KIAA0167 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-gamma 1 (ARF-GAP with GTP-binding protein-like, ankyrin repeat and pleckstrin homology domains 2) (AGAP-2) (Phosphatidylinositol-3-kinase enhancer) (PIKE) (GTP-binding and GTPase-activating protein 2) (GGAP2). | |||||
|
ENV2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.008473 (rank : 86) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P11370, Q61529 | Gene names | Fv4, Env | |||
|
Domain Architecture |

|
|||||
| Description | Retrovirus-related Env polyprotein from Fv-4 locus. | |||||
|
NEB1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.001290 (rank : 104) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULJ8, O76059, Q9NXT2 | Gene names | PPP1R9A, KIAA1222 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-1 (Neurabin-I) (Neural tissue-specific F-actin-binding protein I) (Protein phosphatase 1 regulatory subunit 9A). | |||||
|
PRR12_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.018821 (rank : 69) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
TRI47_MOUSE
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.005431 (rank : 96) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C0E3, Q6P249, Q811J7, Q8BVZ8, Q8R1K0, Q8R3Y1 | Gene names | Trim47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 47. | |||||
|
TROAP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.022999 (rank : 61) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
ABRA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.014126 (rank : 77) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N0Z2 | Gene names | ABRA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding Rho-activating protein (Striated muscle activator of Rho-dependent signaling) (STARS). | |||||
|
CCD1A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.007106 (rank : 91) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K1A6, Q2MJB5, Q8R3Z4 | Gene names | Cc2d1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil and C2 domain-containing protein 1A (Five repressor element under dual repression-binding protein 1) (FRE under dual repression-binding protein 1) (Freud-1). | |||||
|
CDC20_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.003765 (rank : 100) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12834, Q9BW56, Q9UQI9 | Gene names | CDC20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC). | |||||
|
CHD1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.001213 (rank : 105) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
EDRF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.006590 (rank : 93) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3B7T1, Q4G190, Q8IZ74, Q9Y3W4 | Gene names | EDRF1, C10orf137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Erythroid differentiation-related factor 1. | |||||
|
MKL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.009544 (rank : 84) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ULH7, Q68CT1, Q6UB16, Q86WW2, Q8N226 | Gene names | MKL2, KIAA1243, MRTFB | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 2 (Myocardin-related transcription factor B) (MRTF-B) (Megakaryoblastic leukemia 2). | |||||
|
NEO1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.001494 (rank : 103) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92859, O00340 | Gene names | NEO1, NGN | |||
|
Domain Architecture |

|
|||||
| Description | Neogenin precursor. | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | -0.002570 (rank : 111) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
PRDM8_MOUSE
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.005559 (rank : 95) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BZ97 | Gene names | Prdm8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PR domain zinc finger protein 8 (PR domain-containing protein 8). | |||||
|
ZYX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.006643 (rank : 92) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15942 | Gene names | ZYX | |||
|
Domain Architecture |

|
|||||
| Description | Zyxin (Zyxin-2). | |||||
|
HEY1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.081410 (rank : 42) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HEY1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.085320 (rank : 41) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HEY2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.092271 (rank : 39) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.090280 (rank : 40) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
MITF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.074516 (rank : 46) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.076552 (rank : 45) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MLX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.066027 (rank : 53) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.066095 (rank : 52) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MYCL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.050907 (rank : 56) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
TCFL5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.078190 (rank : 43) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
TFE3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.077068 (rank : 44) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
TFEB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.071448 (rank : 47) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.071256 (rank : 48) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
USF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.070965 (rank : 49) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.070933 (rank : 50) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.067952 (rank : 51) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
USF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.063357 (rank : 55) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
EPAS1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 94 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
EPAS1_HUMAN
|
||||||
| NC score | 0.994197 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
HIF1A_MOUSE
|
||||||
| NC score | 0.977646 (rank : 3) | θ value | 2.04462e-161 (rank : 4) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
HIF1A_HUMAN
|
||||||
| NC score | 0.975288 (rank : 4) | θ value | 2.04462e-161 (rank : 3) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
SIM2_MOUSE
|
||||||
| NC score | 0.923554 (rank : 5) | θ value | 2.32439e-64 (rank : 5) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
SIM2_HUMAN
|
||||||
| NC score | 0.923527 (rank : 6) | θ value | 4.10295e-61 (rank : 8) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
SIM1_HUMAN
|
||||||
| NC score | 0.921705 (rank : 7) | θ value | 3.03575e-64 (rank : 6) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM1_MOUSE
|
||||||
| NC score | 0.920668 (rank : 8) | θ value | 3.35642e-63 (rank : 7) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
NPAS1_MOUSE
|
||||||
| NC score | 0.900850 (rank : 9) | θ value | 1.23823e-49 (rank : 11) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
NPAS3_MOUSE
|
||||||
| NC score | 0.899814 (rank : 10) | θ value | 1.78386e-56 (rank : 10) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_HUMAN
|
||||||
| NC score | 0.899765 (rank : 11) | θ value | 1.36585e-56 (rank : 9) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS1_HUMAN
|
||||||
| NC score | 0.899572 (rank : 12) | θ value | 6.7944e-48 (rank : 12) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.829322 (rank : 13) | θ value | 1.98606e-31 (rank : 15) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.826994 (rank : 14) | θ value | 3.3877e-31 (rank : 16) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
AHR_HUMAN
|
||||||
| NC score | 0.815426 (rank : 15) | θ value | 9.87957e-23 (rank : 24) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
AHR_MOUSE
|
||||||
| NC score | 0.812231 (rank : 16) | θ value | 4.43474e-23 (rank : 23) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
CLOCK_HUMAN
|
||||||
| NC score | 0.807441 (rank : 17) | θ value | 4.89182e-30 (rank : 19) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
CLOCK_MOUSE
|
||||||
| NC score | 0.803980 (rank : 18) | θ value | 4.14116e-29 (rank : 22) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
NPAS2_MOUSE
|
||||||
| NC score | 0.796372 (rank : 19) | θ value | 2.86786e-30 (rank : 18) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
ARNT2_HUMAN
|
||||||
| NC score | 0.794139 (rank : 20) | θ value | 8.91499e-32 (rank : 14) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_MOUSE
|
||||||
| NC score | 0.793341 (rank : 21) | θ value | 6.82597e-32 (rank : 13) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
NPAS2_HUMAN
|
||||||
| NC score | 0.789720 (rank : 22) | θ value | 1.4233e-29 (rank : 21) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
ARNT_HUMAN
|
||||||
| NC score | 0.788681 (rank : 23) | θ value | 2.86786e-30 (rank : 17) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.775106 (rank : 24) | θ value | 8.3442e-30 (rank : 20) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
NPAS4_HUMAN
|
||||||
| NC score | 0.678107 (rank : 25) | θ value | 1.13037e-18 (rank : 25) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
NPAS4_MOUSE
|
||||||
| NC score | 0.661451 (rank : 26) | θ value | 1.24977e-17 (rank : 26) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
NCOA1_MOUSE
|
||||||
| NC score | 0.620193 (rank : 27) | θ value | 3.89403e-11 (rank : 31) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA1_HUMAN
|
||||||
| NC score | 0.617665 (rank : 28) | θ value | 1.74796e-11 (rank : 30) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NCOA3_HUMAN
|
||||||
| NC score | 0.569862 (rank : 29) | θ value | 7.84624e-12 (rank : 29) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NCOA2_MOUSE
|
||||||
| NC score | 0.567186 (rank : 30) | θ value | 6.41864e-14 (rank : 27) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
NCOA2_HUMAN
|
||||||
| NC score | 0.566473 (rank : 31) | θ value | 1.09485e-13 (rank : 28) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NCOA3_MOUSE
|
||||||
| NC score | 0.533502 (rank : 32) | θ value | 3.29651e-10 (rank : 32) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
PER3_MOUSE
|
||||||
| NC score | 0.348456 (rank : 33) | θ value | 6.43352e-06 (rank : 34) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
PER1_HUMAN
|
||||||
| NC score | 0.326997 (rank : 34) | θ value | 8.40245e-06 (rank : 35) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER1_MOUSE
|
||||||
| NC score | 0.325698 (rank : 35) | θ value | 3.77169e-06 (rank : 33) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
PER2_HUMAN
|
||||||
| NC score | 0.319170 (rank : 36) | θ value | 0.00102713 (rank : 37) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.313189 (rank : 37) | θ value | 0.000158464 (rank : 36) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
PER2_MOUSE
|
||||||
| NC score | 0.295480 (rank : 38) | θ value | 0.00298849 (rank : 38) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
HEY2_HUMAN
|
||||||
| NC score | 0.092271 (rank : 39) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.090280 (rank : 40) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HEY1_MOUSE
|
||||||
| NC score | 0.085320 (rank : 41) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HEY1_HUMAN
|
||||||
| NC score | 0.081410 (rank : 42) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
TCFL5_HUMAN
|
||||||
| NC score | 0.078190 (rank : 43) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.077068 (rank : 44) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.076552 (rank : 45) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.074516 (rank : 46) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.071448 (rank : 47) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.071256 (rank : 48) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.070965 (rank : 49) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.070933 (rank : 50) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.067952 (rank : 51) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.066095 (rank : 52) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.066027 (rank : 53) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
HES5_MOUSE
|
||||||
| NC score | 0.065676 (rank : 54) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.063357 (rank : 55) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
MYCL1_MOUSE
|
||||||
| NC score | 0.050907 (rank : 56) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MAGC1_HUMAN
|
||||||
| NC score | 0.032762 (rank : 57) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O60732, O75451, Q8TCV4 | Gene names | MAGEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen C1 (MAGE-C1 antigen) (Cancer-testis antigen CT7). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.031434 (rank : 58) | θ value | 0.365318 (rank : 42) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
HLF_MOUSE
|
||||||
| NC score | 0.026412 (rank : 59) | θ value | 0.62314 (rank : 46) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BW74, Q6PF83 | Gene names | Hlf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatic leukemia factor. | |||||
|
HLF_HUMAN
|
||||||
| NC score | 0.025818 (rank : 60) | θ value | 0.813845 (rank : 48) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q16534 | Gene names | HLF | |||
|
Domain Architecture |

|
|||||
| Description | Hepatic leukemia factor. | |||||
|
TROAP_HUMAN
|
||||||
| NC score | 0.022999 (rank : 61) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
NF2L2_HUMAN
|
||||||
| NC score | 0.022972 (rank : 62) | θ value | 0.47712 (rank : 43) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q16236, Q96F71 | Gene names | NFE2L2, NRF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor erythroid 2-related factor 2 (NF-E2-related factor 2) (NFE2-related factor 2) (Nuclear factor, erythroid derived 2, like 2) (HEBP1). | |||||
|
SETB1_MOUSE
|
||||||
| NC score | 0.022215 (rank : 63) | θ value | 0.21417 (rank : 41) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88974, Q922K1 | Gene names | Setdb1, Eset | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
DDEF2_HUMAN
|
||||||
| NC score | 0.021747 (rank : 64) | θ value | 0.0148317 (rank : 39) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 599 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43150 | Gene names | DDEF2, KIAA0400 | |||
|
Domain Architecture |

|
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
BCL9_MOUSE
|
||||||
| NC score | 0.021620 (rank : 65) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.021469 (rank : 66) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.021030 (rank : 67) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
GPS2_HUMAN
|
||||||
| NC score | 0.019917 (rank : 68) | θ value | 1.38821 (rank : 52) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13227 | Gene names | GPS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein pathway suppressor 2 (Protein GPS2). | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.018821 (rank : 69) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
ERF_MOUSE
|
||||||
| NC score | 0.017856 (rank : 70) | θ value | 0.62314 (rank : 45) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
NF2L2_MOUSE
|
||||||
| NC score | 0.017720 (rank : 71) | θ value | 2.36792 (rank : 57) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60795 | Gene names | Nfe2l2, Nrf-2, Nrf2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor erythroid 2-related factor 2 (NF-E2-related factor 2) (NFE2-related factor 2) (Nuclear factor, erythroid derived 2, like 2). | |||||
|
ERF_HUMAN
|
||||||
| NC score | 0.017675 (rank : 72) | θ value | 0.62314 (rank : 44) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P50548, Q9UPI7 | Gene names | ERF | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF (Ets2 repressor factor). | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.017381 (rank : 73) | θ value | 0.813845 (rank : 47) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
CSF1_MOUSE
|
||||||
| NC score | 0.016649 (rank : 74) | θ value | 2.36792 (rank : 56) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P07141, Q8R3C8 | Gene names | Csf1, Csfm | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage colony-stimulating factor 1 precursor (CSF-1) (MCSF). | |||||
|
SYN3_MOUSE
|
||||||
| NC score | 0.014985 (rank : 75) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8JZP2 | Gene names | Syn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
R3HD1_HUMAN
|
||||||
| NC score | 0.014469 (rank : 76) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15032, Q8IW32 | Gene names | R3HDM1, KIAA0029, R3HDM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R3H domain-containing protein 1. | |||||
|
ABRA_HUMAN
|
||||||
| NC score | 0.014126 (rank : 77) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N0Z2 | Gene names | ABRA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding Rho-activating protein (Striated muscle activator of Rho-dependent signaling) (STARS). | |||||
|
DDEF1_MOUSE
|
||||||
| NC score | 0.014078 (rank : 78) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QWY8, O08612, Q80TG8, Q80UV6, Q99LV8, Q9Z2B6 | Gene names | Ddef1, Asap1, Kiaa1249, Shag1 | |||
|
Domain Architecture |

|
|||||
| Description | 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase- activating protein (PIP2-dependent ARF1 GAP) (ADP-ribosylation factor- directed GTPase-activating protein 1) (ARF GTPase-activating protein 1) (Development and differentiation-enhancing factor 1) (Differentiation-enhancing factor 1) (DEF-1). | |||||
|
K1802_HUMAN
|
||||||
| NC score | 0.013125 (rank : 79) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
JPH4_HUMAN
|
||||||
| NC score | 0.012692 (rank : 80) | θ value | 0.813845 (rank : 49) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96JJ6, Q8ND44, Q96DQ0 | Gene names | JPH4, JPHL1, KIAA1831 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-4 (Junctophilin-like 1 protein). | |||||
|
AMELX_MOUSE
|
||||||
| NC score | 0.012294 (rank : 81) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P63277, P45559, P70592, Q61293 | Gene names | Amelx, Amel | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amelogenin, X isoform precursor (Leucine-rich amelogenin peptide) (LRAP). | |||||
|
CNOT3_HUMAN
|
||||||
| NC score | 0.012095 (rank : 82) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75175, Q9NZN7, Q9UF76 | Gene names | CNOT3, KIAA0691, NOT3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
TP53B_MOUSE
|
||||||
| NC score | 0.010048 (rank : 83) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
MKL2_HUMAN
|
||||||
| NC score | 0.009544 (rank : 84) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ULH7, Q68CT1, Q6UB16, Q86WW2, Q8N226 | Gene names | MKL2, KIAA1243, MRTFB | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 2 (Myocardin-related transcription factor B) (MRTF-B) (Megakaryoblastic leukemia 2). | |||||
|
CENG1_HUMAN
|
||||||
| NC score | 0.008602 (rank : 85) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 544 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99490, O00578, Q548E0, Q8IWU3 | Gene names | CENTG1, AGAP2, KIAA0167 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-gamma 1 (ARF-GAP with GTP-binding protein-like, ankyrin repeat and pleckstrin homology domains 2) (AGAP-2) (Phosphatidylinositol-3-kinase enhancer) (PIKE) (GTP-binding and GTPase-activating protein 2) (GGAP2). | |||||
|
ENV2_MOUSE
|
||||||
| NC score | 0.008473 (rank : 86) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P11370, Q61529 | Gene names | Fv4, Env | |||
|
Domain Architecture |

|
|||||
| Description | Retrovirus-related Env polyprotein from Fv-4 locus. | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.008445 (rank : 87) | θ value | 3.0926 (rank : 62) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
SYP2L_HUMAN
|
||||||
| NC score | 0.008360 (rank : 88) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H987 | Gene names | SYNPO2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin 2-like protein. | |||||
|
ROBO3_HUMAN
|
||||||
| NC score | 0.007821 (rank : 89) | θ value | 0.0431538 (rank : 40) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96MS0 | Gene names | ROBO3 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Roundabout-like protein 3). | |||||
|
CENG1_MOUSE
|
||||||
| NC score | 0.007748 (rank : 90) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3UHD9, Q5DU45 | Gene names | Centg1, Agap2, Kiaa0167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-gamma 1 (ARF-GAP with GTP-binding protein-like, ankyrin repeat and pleckstrin homology domains 2) (AGAP-2) (Phosphatidylinositol-3-kinase enhancer) (PIKE). | |||||
|
CCD1A_MOUSE
|
||||||
| NC score | 0.007106 (rank : 91) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K1A6, Q2MJB5, Q8R3Z4 | Gene names | Cc2d1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil and C2 domain-containing protein 1A (Five repressor element under dual repression-binding protein 1) (FRE under dual repression-binding protein 1) (Freud-1). | |||||
|
ZYX_HUMAN
|
||||||
| NC score | 0.006643 (rank : 92) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15942 | Gene names | ZYX | |||
|
Domain Architecture |

|
|||||
| Description | Zyxin (Zyxin-2). | |||||
|
EDRF1_HUMAN
|
||||||
| NC score | 0.006590 (rank : 93) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3B7T1, Q4G190, Q8IZ74, Q9Y3W4 | Gene names | EDRF1, C10orf137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Erythroid differentiation-related factor 1. | |||||
|
CDC20_MOUSE
|
||||||
| NC score | 0.005886 (rank : 94) | θ value | 2.36792 (rank : 55) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JJ66, Q3TGP1, Q8BPG4, Q99LK3 | Gene names | Cdc20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC) (mmCdc20). | |||||
|
PRDM8_MOUSE
|
||||||
| NC score | 0.005559 (rank : 95) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BZ97 | Gene names | Prdm8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PR domain zinc finger protein 8 (PR domain-containing protein 8). | |||||
|
TRI47_MOUSE
|
||||||
| NC score | 0.005431 (rank : 96) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C0E3, Q6P249, Q811J7, Q8BVZ8, Q8R1K0, Q8R3Y1 | Gene names | Trim47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 47. | |||||
|
PASK_HUMAN
|
||||||
| NC score | 0.005154 (rank : 97) | θ value | 1.38821 (rank : 53) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 857 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96RG2, Q86XH6, Q99763, Q9UFR7 | Gene names | PASK, KIAA0135 | |||
|
Domain Architecture |

|
|||||
| Description | PAS domain-containing serine/threonine-protein kinase (EC 2.7.11.1) (PAS-kinase) (PASKIN) (hPASK). | |||||
|
3BP1_HUMAN
|
||||||
| NC score | 0.004960 (rank : 98) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y3L3, Q6IBZ2, Q6ZVL9, Q96HQ5, Q9NSQ9 | Gene names | SH3BP1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
KCNH3_HUMAN
|
||||||
| NC score | 0.004394 (rank : 99) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULD8, Q9UQ06 | Gene names | KCNH3, KIAA1282 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (ELK2) (Brain-specific eag-like channel 1) (BEC1). | |||||
|
CDC20_HUMAN
|
||||||
| NC score | 0.003765 (rank : 100) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12834, Q9BW56, Q9UQI9 | Gene names | CDC20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC). | |||||
|
SPG16_MOUSE
|
||||||
| NC score | 0.003314 (rank : 101) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8K450, Q8C450, Q9CPU8, Q9D522 | Gene names | Spag16, Pf20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 16 protein (PF20 protein homolog). | |||||
|
PASK_MOUSE
|
||||||
| NC score | 0.003045 (rank : 102) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 867 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CEE6, Q91YD8 | Gene names | Pask, Kiaa0135 | |||
|
Domain Architecture |

|
|||||
| Description | PAS domain-containing serine/threonine-protein kinase (EC 2.7.11.1) (PAS-kinase) (PASKIN). | |||||
|
NEO1_HUMAN
|
||||||
| NC score | 0.001494 (rank : 103) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92859, O00340 | Gene names | NEO1, NGN | |||
|
Domain Architecture |

|
|||||
| Description | Neogenin precursor. | |||||
|
NEB1_HUMAN
|
||||||
| NC score | 0.001290 (rank : 104) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULJ8, O76059, Q9NXT2 | Gene names | PPP1R9A, KIAA1222 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-1 (Neurabin-I) (Neural tissue-specific F-actin-binding protein I) (Protein phosphatase 1 regulatory subunit 9A). | |||||
|
CHD1_MOUSE
|
||||||
| NC score | 0.001213 (rank : 105) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
ZEP2_MOUSE
|
||||||
| NC score | 0.001046 (rank : 106) | θ value | 2.36792 (rank : 58) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3UHF7, O55140, Q3UVD4, Q3UVH5 | Gene names | Hivep2, Mibp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 homolog (Myc intron-binding protein 1) (MIBP-1). | |||||
|
NEST_HUMAN
|
||||||
| NC score | 0.001012 (rank : 107) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
SSPO_MOUSE
|
||||||
| NC score | 0.000847 (rank : 108) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
DRD3_MOUSE
|
||||||
| NC score | 0.000490 (rank : 109) | θ value | 1.38821 (rank : 51) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P30728 | Gene names | Drd3 | |||
|
Domain Architecture |

|
|||||
| Description | D(3) dopamine receptor. | |||||
|
TENX_HUMAN
|
||||||
| NC score | 0.000077 (rank : 110) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P22105, P78530, P78531, Q08424, Q9UMG7 | Gene names | TNXB, HXBL, TNX, XB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-X precursor (TN-X) (Hexabrachion-like protein). | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | -0.002570 (rank : 111) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||