Please be patient as the page loads
|
NPAS2_HUMAN
|
||||||
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CLOCK_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.968979 (rank : 3) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
CLOCK_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.967165 (rank : 4) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
NPAS2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 150 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
NPAS2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.988447 (rank : 2) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | 1.20211e-36 (rank : 5) | NC score | 0.844131 (rank : 5) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | 2.26708e-35 (rank : 6) | NC score | 0.840258 (rank : 6) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
ARNT_HUMAN
|
||||||
| θ value | 1.0531e-32 (rank : 7) | NC score | 0.807856 (rank : 7) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | 8.91499e-32 (rank : 8) | NC score | 0.794288 (rank : 8) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
EPAS1_HUMAN
|
||||||
| θ value | 6.38894e-30 (rank : 9) | NC score | 0.792996 (rank : 10) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
EPAS1_MOUSE
|
||||||
| θ value | 1.4233e-29 (rank : 10) | NC score | 0.789720 (rank : 12) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
HIF1A_HUMAN
|
||||||
| θ value | 7.06379e-29 (rank : 11) | NC score | 0.783999 (rank : 13) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
HIF1A_MOUSE
|
||||||
| θ value | 1.47289e-26 (rank : 12) | NC score | 0.781677 (rank : 14) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
ARNT2_MOUSE
|
||||||
| θ value | 3.62785e-25 (rank : 13) | NC score | 0.793339 (rank : 9) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_HUMAN
|
||||||
| θ value | 4.73814e-25 (rank : 14) | NC score | 0.792793 (rank : 11) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
SIM1_HUMAN
|
||||||
| θ value | 1.13037e-18 (rank : 15) | NC score | 0.731785 (rank : 16) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM1_MOUSE
|
||||||
| θ value | 1.13037e-18 (rank : 16) | NC score | 0.731920 (rank : 15) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
SIM2_HUMAN
|
||||||
| θ value | 5.60996e-18 (rank : 17) | NC score | 0.727511 (rank : 17) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
SIM2_MOUSE
|
||||||
| θ value | 3.63628e-17 (rank : 18) | NC score | 0.726641 (rank : 18) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
AHR_HUMAN
|
||||||
| θ value | 1.38178e-16 (rank : 19) | NC score | 0.721429 (rank : 19) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
AHR_MOUSE
|
||||||
| θ value | 8.95645e-16 (rank : 20) | NC score | 0.712314 (rank : 20) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |
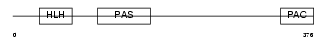
|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
NCOA1_MOUSE
|
||||||
| θ value | 8.95645e-16 (rank : 21) | NC score | 0.628594 (rank : 25) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA1_HUMAN
|
||||||
| θ value | 1.99529e-15 (rank : 22) | NC score | 0.623987 (rank : 26) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NPAS3_MOUSE
|
||||||
| θ value | 6.41864e-14 (rank : 23) | NC score | 0.686872 (rank : 21) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NCOA2_HUMAN
|
||||||
| θ value | 1.86753e-13 (rank : 24) | NC score | 0.569581 (rank : 27) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NCOA2_MOUSE
|
||||||
| θ value | 3.18553e-13 (rank : 25) | NC score | 0.568298 (rank : 29) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
NPAS3_HUMAN
|
||||||
| θ value | 5.43371e-13 (rank : 26) | NC score | 0.682649 (rank : 22) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
PER2_MOUSE
|
||||||
| θ value | 7.34386e-10 (rank : 27) | NC score | 0.414964 (rank : 38) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
PER2_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 28) | NC score | 0.434540 (rank : 34) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER3_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 29) | NC score | 0.456176 (rank : 33) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
NCOA3_HUMAN
|
||||||
| θ value | 3.64472e-09 (rank : 30) | NC score | 0.569420 (rank : 28) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 6.21693e-09 (rank : 31) | NC score | 0.419036 (rank : 37) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
NPAS1_MOUSE
|
||||||
| θ value | 1.38499e-08 (rank : 32) | NC score | 0.664367 (rank : 23) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
NPAS1_HUMAN
|
||||||
| θ value | 2.36244e-08 (rank : 33) | NC score | 0.661296 (rank : 24) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
PER1_HUMAN
|
||||||
| θ value | 4.0297e-08 (rank : 34) | NC score | 0.427623 (rank : 35) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER1_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 35) | NC score | 0.423123 (rank : 36) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
NCOA3_MOUSE
|
||||||
| θ value | 2.61198e-07 (rank : 36) | NC score | 0.531060 (rank : 30) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
NPAS4_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 37) | NC score | 0.507368 (rank : 31) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
NPAS4_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 38) | NC score | 0.491599 (rank : 32) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
HNF1A_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 39) | NC score | 0.041654 (rank : 83) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
EMSY_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 40) | NC score | 0.087680 (rank : 63) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |
|
|||||
| Description | Protein EMSY. | |||||
|
RUNX1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 41) | NC score | 0.043345 (rank : 81) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q01196, O60472, O60473, O76047, O76089, Q13081, Q13755, Q13756, Q13757, Q13758, Q13759, Q15341, Q15343, Q16122, Q16284, Q16285, Q16286, Q16346, Q16347, Q92479 | Gene names | RUNX1, AML1, CBFA2 | |||
|
Domain Architecture |

|
|||||
| Description | Runt-related transcription factor 1 (Core-binding factor, alpha 2 subunit) (CBF-alpha 2) (Acute myeloid leukemia 1 protein) (Oncogene AML-1) (Polyomavirus enhancer-binding protein 2 alpha B subunit) (PEBP2-alpha B) (PEA2-alpha B) (SL3-3 enhancer factor 1 alpha B subunit) (SL3/AKV core-binding factor alpha B subunit). | |||||
|
ZIMP7_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 42) | NC score | 0.039672 (rank : 84) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CIE2, Q6P558, Q6P9Q5, Q8BJ00 | Gene names | Zimp7, D11Bwg0280e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
RUNX1_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 43) | NC score | 0.041745 (rank : 82) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q03347, O08598, Q62049, Q9ESB9, Q9ET65 | Gene names | Runx1, Aml1, Cbfa2, Pebp2ab | |||
|
Domain Architecture |
|
|||||
| Description | Runt-related transcription factor 1 (Core-binding factor, alpha 2 subunit) (CBF-alpha 2) (Acute myeloid leukemia 1 protein) (Oncogene AML-1) (Polyomavirus enhancer-binding protein 2 alpha B subunit) (PEBP2-alpha B) (PEA2-alpha B) (SL3-3 enhancer factor 1 alpha B subunit) (SL3/AKV core-binding factor alpha B subunit). | |||||
|
MYH9_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 44) | NC score | 0.024163 (rank : 96) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH9_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 45) | NC score | 0.023462 (rank : 97) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
EMSY_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 46) | NC score | 0.080939 (rank : 68) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BMB0, Q5FWK5, Q80XU1, Q8VDW9 | Gene names | Emsy | |||
|
Domain Architecture |
|
|||||
| Description | Protein EMSY. | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | 0.125558 (rank : 47) | NC score | 0.072939 (rank : 71) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
NCOA6_MOUSE
|
||||||
| θ value | 0.125558 (rank : 48) | NC score | 0.082141 (rank : 67) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.163984 (rank : 49) | NC score | 0.052450 (rank : 78) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
US6NL_HUMAN
|
||||||
| θ value | 0.163984 (rank : 50) | NC score | 0.028367 (rank : 88) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92738, Q15400, Q7L0K9 | Gene names | USP6NL, KIAA0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein (Related to the N-terminus of tre) (RN- tre). | |||||
|
CD248_HUMAN
|
||||||
| θ value | 0.21417 (rank : 51) | NC score | 0.022363 (rank : 99) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9HCU0, Q3SX55, Q96KB6 | Gene names | CD248, CD164L1, TEM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
MAML3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 52) | NC score | 0.065583 (rank : 74) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96JK9 | Gene names | MAML3, KIAA1816 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 3 (Mam-3). | |||||
|
TFE3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 53) | NC score | 0.177910 (rank : 39) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
HES7_HUMAN
|
||||||
| θ value | 0.279714 (rank : 54) | NC score | 0.096405 (rank : 61) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
MITF_HUMAN
|
||||||
| θ value | 0.279714 (rank : 55) | NC score | 0.170197 (rank : 41) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| θ value | 0.279714 (rank : 56) | NC score | 0.171370 (rank : 40) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
BHLH2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 57) | NC score | 0.111474 (rank : 55) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
BHLH2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 58) | NC score | 0.110609 (rank : 56) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
HES7_MOUSE
|
||||||
| θ value | 0.365318 (rank : 59) | NC score | 0.092084 (rank : 62) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
MLX_HUMAN
|
||||||
| θ value | 0.365318 (rank : 60) | NC score | 0.160216 (rank : 42) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| θ value | 0.365318 (rank : 61) | NC score | 0.159794 (rank : 43) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
TTLL5_MOUSE
|
||||||
| θ value | 0.365318 (rank : 62) | NC score | 0.021006 (rank : 101) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CHB8 | Gene names | Ttll5, Kiaa0998 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 5. | |||||
|
WNK1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 63) | NC score | 0.008020 (rank : 142) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1179 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P83741 | Gene names | Wnk1, Prkwnk1 | |||
|
Domain Architecture |
|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1). | |||||
|
PO2F1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 64) | NC score | 0.012660 (rank : 126) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P25425, Q61994, Q63891, Q6Y681, Q7TSD0, Q8BT04, Q8K570, Q99JH0, Q9WTZ4, Q9WTZ5, Q9WTZ6 | Gene names | Pou2f1, Oct-1, Otf-1, Otf1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 2, transcription factor 1 (Octamer-binding transcription factor 1) (Oct-1) (OTF-1) (NF-A1). | |||||
|
SDC3_MOUSE
|
||||||
| θ value | 0.47712 (rank : 65) | NC score | 0.039241 (rank : 85) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64519, Q3UDD9, Q6ZQA4, Q7TQD4 | Gene names | Sdc3, Kiaa0468 | |||
|
Domain Architecture |
|
|||||
| Description | Syndecan-3 precursor (SYND3). | |||||
|
USF1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 66) | NC score | 0.142897 (rank : 46) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 67) | NC score | 0.142819 (rank : 47) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
HCN1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 68) | NC score | 0.014261 (rank : 119) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O88704, O54899, Q9D613 | Gene names | Hcn1, Bcng1, Hac2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1) (Hyperpolarization-activated cation channel 2) (HAC-2). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 69) | NC score | 0.027569 (rank : 89) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
ULK1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 70) | NC score | -0.000603 (rank : 158) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1121 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O75385 | Gene names | ULK1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase ULK1 (EC 2.7.11.1) (Unc-51-like kinase 1). | |||||
|
USF2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 71) | NC score | 0.135239 (rank : 48) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
DACH1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 72) | NC score | 0.022404 (rank : 98) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UI36, O75523, O75687, Q5VYY3, Q5VYY4, Q96SG3, Q96SG4, Q9H524, Q9UMH4 | Gene names | DACH1, DACH | |||
|
Domain Architecture |

|
|||||
| Description | Dachshund homolog 1 (Dach1). | |||||
|
DCC_MOUSE
|
||||||
| θ value | 1.06291 (rank : 73) | NC score | 0.005681 (rank : 150) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
MYH10_HUMAN
|
||||||
| θ value | 1.06291 (rank : 74) | NC score | 0.018510 (rank : 106) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
RN165_HUMAN
|
||||||
| θ value | 1.06291 (rank : 75) | NC score | 0.018460 (rank : 108) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZSG1 | Gene names | RNF165 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 165. | |||||
|
TULP4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 76) | NC score | 0.026480 (rank : 91) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NRJ4, Q9HD22, Q9P2F0 | Gene names | TULP4, KIAA1397, TUSP | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-like protein 4 (Tubby superfamily protein). | |||||
|
DCP1A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 77) | NC score | 0.046854 (rank : 80) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NPI6 | Gene names | DCP1A, SMIF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (Smad4-interacting transcriptional co-activator). | |||||
|
GAK2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 78) | NC score | 0.015991 (rank : 116) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
KIF17_HUMAN
|
||||||
| θ value | 1.38821 (rank : 79) | NC score | 0.008592 (rank : 139) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P2E2, O95077, Q53YS6, Q5VWA9, Q6GSA8, Q8N411 | Gene names | KIF17, KIAA1405, KIF3X | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF17 (KIF3-related motor protein). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 80) | NC score | 0.033331 (rank : 86) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MYH10_MOUSE
|
||||||
| θ value | 1.38821 (rank : 81) | NC score | 0.018268 (rank : 109) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
USF2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 82) | NC score | 0.127512 (rank : 54) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
ADA12_MOUSE
|
||||||
| θ value | 1.81305 (rank : 83) | NC score | 0.004716 (rank : 151) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61824 | Gene names | Adam12, Mltna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
ATOH1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 84) | NC score | 0.016670 (rank : 114) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92858 | Gene names | ATOH1, ATH1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein hATH-1). | |||||
|
BHLH3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 85) | NC score | 0.110389 (rank : 57) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BHLH3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 86) | NC score | 0.108672 (rank : 58) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
BRCA1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 87) | NC score | 0.010432 (rank : 134) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48754, Q60957, Q60983 | Gene names | Brca1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein homolog. | |||||
|
MUC5B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 88) | NC score | 0.026973 (rank : 90) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
RANB3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 89) | NC score | 0.018471 (rank : 107) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H6Z4, O60405, O75759, O75760, Q9BT47, Q9UG74 | Gene names | RANBP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 3 (RanBP3). | |||||
|
TAF3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 90) | NC score | 0.018600 (rank : 104) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
DLX5_MOUSE
|
||||||
| θ value | 2.36792 (rank : 91) | NC score | 0.003100 (rank : 154) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70396, O54876, O54877, O54878, Q9JJ45 | Gene names | Dlx5 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein DLX-5. | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 2.36792 (rank : 92) | NC score | 0.017515 (rank : 112) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 2.36792 (rank : 93) | NC score | 0.017678 (rank : 111) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
KIF17_MOUSE
|
||||||
| θ value | 3.0926 (rank : 94) | NC score | 0.009807 (rank : 137) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99PW8 | Gene names | Kif17 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF17 (MmKIF17). | |||||
|
PI51A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 95) | NC score | 0.008476 (rank : 140) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99755, Q99754, Q99756 | Gene names | PIP5K1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 5-kinase type-1 alpha (EC 2.7.1.68) (Phosphatidylinositol-4-phosphate 5-kinase type I alpha) (PtdIns(4)P- 5-kinase alpha) (PIP5KIalpha) (68 kDa type I phosphatidylinositol-4- phosphate 5-kinase alpha). | |||||
|
RFFL_HUMAN
|
||||||
| θ value | 3.0926 (rank : 96) | NC score | 0.011680 (rank : 131) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WZ73, Q8NHW0, Q8TBY7, Q96BE6 | Gene names | RFFL, RNF189, RNF34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (FYVE-RING finger protein Sakura) (Fring) (Caspases-8 and -10- associated RING finger protein 2) (CARP-2) (Caspase regulator CARP2) (RING finger protein 189) (RING finger protein 34-like). | |||||
|
TARA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 97) | NC score | 0.024266 (rank : 95) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TFEB_HUMAN
|
||||||
| θ value | 3.0926 (rank : 98) | NC score | 0.150564 (rank : 45) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| θ value | 3.0926 (rank : 99) | NC score | 0.151452 (rank : 44) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TIMD1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 100) | NC score | 0.021785 (rank : 100) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96D42, O43656 | Gene names | HAVCR1, TIM1, TIMD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatitis A virus cellular receptor 1 precursor (HAVcr-1) (T cell immunoglobulin and mucin domain-containing protein 1) (TIMD-1) (T cell membrane protein 1) (TIM-1) (TIM). | |||||
|
AMOT_HUMAN
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.016391 (rank : 115) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
APOB_HUMAN
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.020144 (rank : 102) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P04114, O00502, P78479, P78480, P78481, Q13779, Q13785, Q13786, Q13787, Q13788, Q7Z600, Q9UMN0 | Gene names | APOB | |||
|
Domain Architecture |

|
|||||
| Description | Apolipoprotein B-100 precursor (Apo B-100) [Contains: Apolipoprotein B-48 (Apo B-48)]. | |||||
|
CDK8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 103) | NC score | -0.001377 (rank : 161) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P49336 | Gene names | CDK8 | |||
|
Domain Architecture |
|
|||||
| Description | Cell division protein kinase 8 (EC 2.7.11.22) (EC 2.7.11.23) (Protein kinase K35). | |||||
|
GAK5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 104) | NC score | 0.014640 (rank : 118) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GMEB2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 105) | NC score | 0.011072 (rank : 133) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKD1, Q5TDS0, Q9H431, Q9H4X7, Q9H4X8, Q9UF78, Q9ULF1 | Gene names | GMEB2, KIAA1269 | |||
|
Domain Architecture |
|
|||||
| Description | Glucocorticoid modulatory element-binding protein 2 (GMEB-2) (Parvovirus initiation factor p79) (PIF p79) (DNA-binding protein p79PIF). | |||||
|
CBL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.007241 (rank : 146) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P22682, Q8CEA1 | Gene names | Cbl | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase CBL (EC 6.3.2.-) (Signal transduction protein CBL) (Proto-oncogene c-CBL) (Casitas B-lineage lymphoma proto- oncogene). | |||||
|
CO039_MOUSE
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.017184 (rank : 113) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
HAX1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.013808 (rank : 122) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35387 | Gene names | Hax1, Hs1bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HS1-associating protein X-1 (HAX-1) (HS1-binding protein). | |||||
|
HGS_HUMAN
|
||||||
| θ value | 5.27518 (rank : 109) | NC score | 0.025434 (rank : 94) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
K0240_MOUSE
|
||||||
| θ value | 5.27518 (rank : 110) | NC score | 0.014151 (rank : 120) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CHH5 | Gene names | Kiaa0240 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0240. | |||||
|
LIPA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 111) | NC score | 0.006245 (rank : 149) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
RFX1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 112) | NC score | 0.018048 (rank : 110) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P22670 | Gene names | RFX1 | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II regulatory factor RFX1 (RFX) (Enhancer factor C) (EF-C). | |||||
|
STK39_MOUSE
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | -0.002592 (rank : 162) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 915 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Z1W9, Q80W13 | Gene names | Stk39, Spak | |||
|
Domain Architecture |
|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39). | |||||
|
T22D2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 114) | NC score | 0.025495 (rank : 93) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
TAF1C_MOUSE
|
||||||
| θ value | 5.27518 (rank : 115) | NC score | 0.018729 (rank : 103) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
TCFL5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 116) | NC score | 0.129367 (rank : 52) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
XKR5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 117) | NC score | 0.009379 (rank : 138) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5GH66 | Gene names | Xkr5, Xrg5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | XK-related protein 5. | |||||
|
ZN645_HUMAN
|
||||||
| θ value | 5.27518 (rank : 118) | NC score | 0.029317 (rank : 87) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8N7E2, Q6DJY9 | Gene names | ZNF645 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 645. | |||||
|
GAK6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 119) | NC score | 0.013899 (rank : 121) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
HES5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 120) | NC score | 0.106422 (rank : 59) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
HEY1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 121) | NC score | 0.128273 (rank : 53) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HEY1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 122) | NC score | 0.133491 (rank : 49) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
MCM6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 123) | NC score | 0.007353 (rank : 144) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97311, Q80YQ7 | Gene names | Mcm6, Mcmd6, Mis5 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM6 (Mis5 homolog). | |||||
|
MK07_MOUSE
|
||||||
| θ value | 6.88961 (rank : 124) | NC score | -0.001179 (rank : 160) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1054 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9WVS8 | Gene names | Mapk7, Erk5 | |||
|
Domain Architecture |
|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (BMK1 kinase). | |||||
|
MYCL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 125) | NC score | 0.078269 (rank : 69) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCL1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 126) | NC score | 0.082784 (rank : 66) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
POGK_MOUSE
|
||||||
| θ value | 6.88961 (rank : 127) | NC score | 0.001875 (rank : 157) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80TC5, Q8BPJ3, Q8C887 | Gene names | Pogk, Kiaa1513 | |||
|
Domain Architecture |

|
|||||
| Description | Pogo transposable element with KRAB domain. | |||||
|
RUFY1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 128) | NC score | 0.012508 (rank : 128) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 848 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96T51, Q59FF3, Q71S93, Q9H6I3 | Gene names | RUFY1, RABIP4, ZFYVE12 | |||
|
Domain Architecture |

|
|||||
| Description | RUN and FYVE domain-containing protein 1 (FYVE-finger protein EIP1) (Zinc finger FYVE domain-containing protein 12) (La-binding protein 1) (Rab4-interacting protein). | |||||
|
SDC3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 129) | NC score | 0.025988 (rank : 92) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75056, Q5T1Z6, Q5T1Z7, Q96CT3, Q96PR8 | Gene names | SDC3, KIAA0468 | |||
|
Domain Architecture |
|
|||||
| Description | Syndecan-3 (SYND3). | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 6.88961 (rank : 130) | NC score | -0.000892 (rank : 159) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
TTLL4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 131) | NC score | 0.012040 (rank : 130) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14679, Q8WW29 | Gene names | TTLL4, KIAA0173 | |||
|
Domain Architecture |

|
|||||
| Description | Tubulin--tyrosine ligase-like protein 4. | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 132) | NC score | 0.007259 (rank : 145) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
ADA19_HUMAN
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.003874 (rank : 152) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
AKAP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | 0.010278 (rank : 135) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
AMOL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 135) | NC score | 0.011306 (rank : 132) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.018576 (rank : 105) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
CEP63_HUMAN
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.008023 (rank : 141) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1005 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96MT8, Q96CR0, Q9H8F5, Q9H8N0 | Gene names | CEP63 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 63 kDa (Cep63 protein). | |||||
|
CIC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.015117 (rank : 117) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
ELF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.006854 (rank : 147) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q60775 | Gene names | Elf1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
GAK11_HUMAN
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.013597 (rank : 124) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.013529 (rank : 125) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.013602 (rank : 123) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GOGA5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.007998 (rank : 143) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
IF35_HUMAN
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.009867 (rank : 136) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O00303, Q8N978 | Gene names | EIF3S5 | |||
|
Domain Architecture |
|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 5 (eIF-3 epsilon) (eIF3 p47 subunit) (eIF3f). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.003278 (rank : 153) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LTB1L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | 0.002560 (rank : 155) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14766 | Gene names | LTBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein, isoform 1L precursor (LTBP-1) (Transforming growth factor beta-1-binding protein 1) (TGF-beta1-BP-1). | |||||
|
LTB1S_HUMAN
|
||||||
| θ value | 8.99809 (rank : 147) | NC score | 0.002506 (rank : 156) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P22064, Q8TD95 | Gene names | LTBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein, isoform 1S precursor (LTBP-1) (Transforming growth factor beta-1-binding protein 1) (TGF-beta1-BP-1). | |||||
|
MUCEN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 148) | NC score | 0.012657 (rank : 127) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULC0, Q8NEY5, Q8WWE7, Q9NRM8 | Gene names | EMCN, EMCN2. MUC14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endomucin precursor (Endomucin-2) (Gastric cancer antigen Ga34). | |||||
|
NMNA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 149) | NC score | 0.006534 (rank : 148) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9EPA7, Q6B504 | Gene names | Nmnat1, D4Cole1e, Nmnat | |||
|
Domain Architecture |
|
|||||
| Description | Nicotinamide mononucleotide adenylyltransferase 1 (EC 2.7.7.1) (NMN adenylyltransferase 1). | |||||
|
TEKT2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 150) | NC score | 0.012407 (rank : 129) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UIF3, O60638 | Gene names | TEKT2 | |||
|
Domain Architecture |

|
|||||
| Description | Tektin-2 (Testicular tektin) (Tektin-t) (Testicular tektin B1-like protein) (Tektin-B1) (TEKTB1). | |||||
|
HES3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.070983 (rank : 73) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
HES4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.055530 (rank : 75) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
HEY2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.132357 (rank : 50) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.130818 (rank : 51) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
MAD4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.052697 (rank : 76) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MAX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.072109 (rank : 72) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
MAX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.083968 (rank : 65) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.050342 (rank : 79) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
PCQAP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.052464 (rank : 77) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 549 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q96RN5, O15413, Q6IC31, Q8NF16, Q96CT0, Q96IH7, Q9P1T3 | Gene names | PCQAP, ARC105, CTG7A, TIG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Positive cofactor 2 glutamine/Q-rich-associated protein (PC2 glutamine/Q-rich-associated protein) (TPA-inducible gene 1 protein) (TIG-1) (Activator-recruited cofactor 105 kDa component) (ARC105) (CTG repeat protein 7a). | |||||
|
SRBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.097196 (rank : 60) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
SRBP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.087017 (rank : 64) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
SRBP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.077100 (rank : 70) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
NPAS2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 150 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
NPAS2_MOUSE
|
||||||
| NC score | 0.988447 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
CLOCK_HUMAN
|
||||||
| NC score | 0.968979 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
CLOCK_MOUSE
|
||||||
| NC score | 0.967165 (rank : 4) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.844131 (rank : 5) | θ value | 1.20211e-36 (rank : 5) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.840258 (rank : 6) | θ value | 2.26708e-35 (rank : 6) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
ARNT_HUMAN
|
||||||
| NC score | 0.807856 (rank : 7) | θ value | 1.0531e-32 (rank : 7) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.794288 (rank : 8) | θ value | 8.91499e-32 (rank : 8) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT2_MOUSE
|
||||||
| NC score | 0.793339 (rank : 9) | θ value | 3.62785e-25 (rank : 13) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
EPAS1_HUMAN
|
||||||
| NC score | 0.792996 (rank : 10) | θ value | 6.38894e-30 (rank : 9) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
ARNT2_HUMAN
|
||||||
| NC score | 0.792793 (rank : 11) | θ value | 4.73814e-25 (rank : 14) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
EPAS1_MOUSE
|
||||||
| NC score | 0.789720 (rank : 12) | θ value | 1.4233e-29 (rank : 10) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
HIF1A_HUMAN
|
||||||
| NC score | 0.783999 (rank : 13) | θ value | 7.06379e-29 (rank : 11) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
HIF1A_MOUSE
|
||||||
| NC score | 0.781677 (rank : 14) | θ value | 1.47289e-26 (rank : 12) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
SIM1_MOUSE
|
||||||
| NC score | 0.731920 (rank : 15) | θ value | 1.13037e-18 (rank : 16) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
SIM1_HUMAN
|
||||||
| NC score | 0.731785 (rank : 16) | θ value | 1.13037e-18 (rank : 15) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM2_HUMAN
|
||||||
| NC score | 0.727511 (rank : 17) | θ value | 5.60996e-18 (rank : 17) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
SIM2_MOUSE
|
||||||
| NC score | 0.726641 (rank : 18) | θ value | 3.63628e-17 (rank : 18) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
AHR_HUMAN
|
||||||
| NC score | 0.721429 (rank : 19) | θ value | 1.38178e-16 (rank : 19) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
AHR_MOUSE
|
||||||
| NC score | 0.712314 (rank : 20) | θ value | 8.95645e-16 (rank : 20) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
NPAS3_MOUSE
|
||||||
| NC score | 0.686872 (rank : 21) | θ value | 6.41864e-14 (rank : 23) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_HUMAN
|
||||||
| NC score | 0.682649 (rank : 22) | θ value | 5.43371e-13 (rank : 26) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS1_MOUSE
|
||||||
| NC score | 0.664367 (rank : 23) | θ value | 1.38499e-08 (rank : 32) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
NPAS1_HUMAN
|
||||||
| NC score | 0.661296 (rank : 24) | θ value | 2.36244e-08 (rank : 33) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
NCOA1_MOUSE
|
||||||
| NC score | 0.628594 (rank : 25) | θ value | 8.95645e-16 (rank : 21) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA1_HUMAN
|
||||||
| NC score | 0.623987 (rank : 26) | θ value | 1.99529e-15 (rank : 22) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NCOA2_HUMAN
|
||||||
| NC score | 0.569581 (rank : 27) | θ value | 1.86753e-13 (rank : 24) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NCOA3_HUMAN
|
||||||
| NC score | 0.569420 (rank : 28) | θ value | 3.64472e-09 (rank : 30) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NCOA2_MOUSE
|
||||||
| NC score | 0.568298 (rank : 29) | θ value | 3.18553e-13 (rank : 25) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
NCOA3_MOUSE
|
||||||
| NC score | 0.531060 (rank : 30) | θ value | 2.61198e-07 (rank : 36) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
NPAS4_HUMAN
|
||||||
| NC score | 0.507368 (rank : 31) | θ value | 0.00020696 (rank : 37) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
NPAS4_MOUSE
|
||||||
| NC score | 0.491599 (rank : 32) | θ value | 0.000270298 (rank : 38) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
PER3_MOUSE
|
||||||
| NC score | 0.456176 (rank : 33) | θ value | 2.79066e-09 (rank : 29) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
PER2_HUMAN
|
||||||
| NC score | 0.434540 (rank : 34) | θ value | 2.79066e-09 (rank : 28) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER1_HUMAN
|
||||||
| NC score | 0.427623 (rank : 35) | θ value | 4.0297e-08 (rank : 34) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER1_MOUSE
|
||||||
| NC score | 0.423123 (rank : 36) | θ value | 5.26297e-08 (rank : 35) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.419036 (rank : 37) | θ value | 6.21693e-09 (rank : 31) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
PER2_MOUSE
|
||||||
| NC score | 0.414964 (rank : 38) | θ value | 7.34386e-10 (rank : 27) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.177910 (rank : 39) | θ value | 0.21417 (rank : 53) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.171370 (rank : 40) | θ value | 0.279714 (rank : 56) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.170197 (rank : 41) | θ value | 0.279714 (rank : 55) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.160216 (rank : 42) | θ value | 0.365318 (rank : 60) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.159794 (rank : 43) | θ value | 0.365318 (rank : 61) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.151452 (rank : 44) | θ value | 3.0926 (rank : 99) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.150564 (rank : 45) | θ value | 3.0926 (rank : 98) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.142897 (rank : 46) | θ value | 0.47712 (rank : 66) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.142819 (rank : 47) | θ value | 0.47712 (rank : 67) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.135239 (rank : 48) | θ value | 0.813845 (rank : 71) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
HEY1_MOUSE
|
||||||
| NC score | 0.133491 (rank : 49) | θ value | 6.88961 (rank : 122) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HEY2_HUMAN
|
||||||
| NC score | 0.132357 (rank : 50) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.130818 (rank : 51) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
TCFL5_HUMAN
|
||||||
| NC score | 0.129367 (rank : 52) | θ value | 5.27518 (rank : 116) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
HEY1_HUMAN
|
||||||
| NC score | 0.128273 (rank : 53) | θ value | 6.88961 (rank : 121) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.127512 (rank : 54) | θ value | 1.38821 (rank : 82) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
BHLH2_HUMAN
|
||||||
| NC score | 0.111474 (rank : 55) | θ value | 0.365318 (rank : 57) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
BHLH2_MOUSE
|
||||||
| NC score | 0.110609 (rank : 56) | θ value | 0.365318 (rank : 58) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
BHLH3_HUMAN
|
||||||
| NC score | 0.110389 (rank : 57) | θ value | 1.81305 (rank : 85) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BHLH3_MOUSE
|
||||||
| NC score | 0.108672 (rank : 58) | θ value | 1.81305 (rank : 86) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
HES5_MOUSE
|
||||||
| NC score | 0.106422 (rank : 59) | θ value | 6.88961 (rank : 120) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
SRBP1_HUMAN
|
||||||
| NC score | 0.097196 (rank : 60) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
HES7_HUMAN
|
||||||
| NC score | 0.096405 (rank : 61) | θ value | 0.279714 (rank : 54) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES7_MOUSE
|
||||||
| NC score | 0.092084 (rank : 62) | θ value | 0.365318 (rank : 59) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
EMSY_HUMAN
|
||||||
| NC score | 0.087680 (rank : 63) | θ value | 0.0193708 (rank : 40) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |

|
|||||
| Description | Protein EMSY. | |||||
|
SRBP1_MOUSE
|
||||||
| NC score | 0.087017 (rank : 64) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
MAX_MOUSE
|
||||||
| NC score | 0.083968 (rank : 65) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
MYCL1_MOUSE
|
||||||
| NC score | 0.082784 (rank : 66) | θ value | 6.88961 (rank : 126) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
NCOA6_MOUSE
|
||||||
| NC score | 0.082141 (rank : 67) | θ value | 0.125558 (rank : 48) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
EMSY_MOUSE
|
||||||
| NC score | 0.080939 (rank : 68) | θ value | 0.0961366 (rank : 46) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BMB0, Q5FWK5, Q80XU1, Q8VDW9 | Gene names | Emsy | |||
|
Domain Architecture |

|
|||||
| Description | Protein EMSY. | |||||
|
MYCL1_HUMAN
|
||||||
| NC score | 0.078269 (rank : 69) | θ value | 6.88961 (rank : 125) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
SRBP2_HUMAN
|
||||||
| NC score | 0.077100 (rank : 70) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.072939 (rank : 71) | θ value | 0.125558 (rank : 47) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
MAX_HUMAN
|
||||||
| NC score | 0.072109 (rank : 72) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
HES3_MOUSE
|
||||||
| NC score | 0.070983 (rank : 73) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
MAML3_HUMAN
|
||||||
| NC score | 0.065583 (rank : 74) | θ value | 0.21417 (rank : 52) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96JK9 | Gene names | MAML3, KIAA1816 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 3 (Mam-3). | |||||
|
HES4_HUMAN
|
||||||
| NC score | 0.055530 (rank : 75) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
MAD4_HUMAN
|
||||||
| NC score | 0.052697 (rank : 76) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
PCQAP_HUMAN
|
||||||
| NC score | 0.052464 (rank : 77) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 549 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q96RN5, O15413, Q6IC31, Q8NF16, Q96CT0, Q96IH7, Q9P1T3 | Gene names | PCQAP, ARC105, CTG7A, TIG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Positive cofactor 2 glutamine/Q-rich-associated protein (PC2 glutamine/Q-rich-associated protein) (TPA-inducible gene 1 protein) (TIG-1) (Activator-recruited cofactor 105 kDa component) (ARC105) (CTG repeat protein 7a). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.052450 (rank : 78) | θ value | 0.163984 (rank : 49) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.050342 (rank : 79) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
DCP1A_HUMAN
|
||||||
| NC score | 0.046854 (rank : 80) | θ value | 1.38821 (rank : 77) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NPI6 | Gene names | DCP1A, SMIF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (Smad4-interacting transcriptional co-activator). | |||||
|
RUNX1_HUMAN
|
||||||
| NC score | 0.043345 (rank : 81) | θ value | 0.0252991 (rank : 41) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q01196, O60472, O60473, O76047, O76089, Q13081, Q13755, Q13756, Q13757, Q13758, Q13759, Q15341, Q15343, Q16122, Q16284, Q16285, Q16286, Q16346, Q16347, Q92479 | Gene names | RUNX1, AML1, CBFA2 | |||
|
Domain Architecture |

|
|||||
| Description | Runt-related transcription factor 1 (Core-binding factor, alpha 2 subunit) (CBF-alpha 2) (Acute myeloid leukemia 1 protein) (Oncogene AML-1) (Polyomavirus enhancer-binding protein 2 alpha B subunit) (PEBP2-alpha B) (PEA2-alpha B) (SL3-3 enhancer factor 1 alpha B subunit) (SL3/AKV core-binding factor alpha B subunit). | |||||
|
RUNX1_MOUSE
|
||||||
| NC score | 0.041745 (rank : 82) | θ value | 0.0431538 (rank : 43) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q03347, O08598, Q62049, Q9ESB9, Q9ET65 | Gene names | Runx1, Aml1, Cbfa2, Pebp2ab | |||
|
Domain Architecture |

|
|||||
| Description | Runt-related transcription factor 1 (Core-binding factor, alpha 2 subunit) (CBF-alpha 2) (Acute myeloid leukemia 1 protein) (Oncogene AML-1) (Polyomavirus enhancer-binding protein 2 alpha B subunit) (PEBP2-alpha B) (PEA2-alpha B) (SL3-3 enhancer factor 1 alpha B subunit) (SL3/AKV core-binding factor alpha B subunit). | |||||
|
HNF1A_MOUSE
|
||||||
| NC score | 0.041654 (rank : 83) | θ value | 0.0113563 (rank : 39) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
ZIMP7_MOUSE
|
||||||
| NC score | 0.039672 (rank : 84) | θ value | 0.0252991 (rank : 42) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CIE2, Q6P558, Q6P9Q5, Q8BJ00 | Gene names | Zimp7, D11Bwg0280e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
SDC3_MOUSE
|
||||||
| NC score | 0.039241 (rank : 85) | θ value | 0.47712 (rank : 65) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64519, Q3UDD9, Q6ZQA4, Q7TQD4 | Gene names | Sdc3, Kiaa0468 | |||
|
Domain Architecture |

|
|||||
| Description | Syndecan-3 precursor (SYND3). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.033331 (rank : 86) | θ value | 1.38821 (rank : 80) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
ZN645_HUMAN
|
||||||
| NC score | 0.029317 (rank : 87) | θ value | 5.27518 (rank : 118) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8N7E2, Q6DJY9 | Gene names | ZNF645 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 645. | |||||
|
US6NL_HUMAN
|
||||||
| NC score | 0.028367 (rank : 88) | θ value | 0.163984 (rank : 50) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92738, Q15400, Q7L0K9 | Gene names | USP6NL, KIAA0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein (Related to the N-terminus of tre) (RN- tre). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.027569 (rank : 89) | θ value | 0.813845 (rank : 69) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
MUC5B_HUMAN
|
||||||
| NC score | 0.026973 (rank : 90) | θ value | 1.81305 (rank : 88) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
TULP4_HUMAN
|
||||||
| NC score | 0.026480 (rank : 91) | θ value | 1.06291 (rank : 76) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NRJ4, Q9HD22, Q9P2F0 | Gene names | TULP4, KIAA1397, TUSP | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-like protein 4 (Tubby superfamily protein). | |||||
|
SDC3_HUMAN
|
||||||
| NC score | 0.025988 (rank : 92) | θ value | 6.88961 (rank : 129) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75056, Q5T1Z6, Q5T1Z7, Q96CT3, Q96PR8 | Gene names | SDC3, KIAA0468 | |||
|
Domain Architecture |

|
|||||
| Description | Syndecan-3 (SYND3). | |||||
|
T22D2_HUMAN
|
||||||
| NC score | 0.025495 (rank : 93) | θ value | 5.27518 (rank : 114) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
HGS_HUMAN
|
||||||
| NC score | 0.025434 (rank : 94) | θ value | 5.27518 (rank : 109) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.024266 (rank : 95) | θ value | 3.0926 (rank : 97) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.024163 (rank : 96) | θ value | 0.0563607 (rank : 44) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH9_MOUSE
|
||||||
| NC score | 0.023462 (rank : 97) | θ value | 0.0736092 (rank : 45) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
DACH1_HUMAN
|
||||||
| NC score | 0.022404 (rank : 98) | θ value | 1.06291 (rank : 72) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UI36, O75523, O75687, Q5VYY3, Q5VYY4, Q96SG3, Q96SG4, Q9H524, Q9UMH4 | Gene names | DACH1, DACH | |||
|
Domain Architecture |

|
|||||
| Description | Dachshund homolog 1 (Dach1). | |||||
|
CD248_HUMAN
|
||||||
| NC score | 0.022363 (rank : 99) | θ value | 0.21417 (rank : 51) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9HCU0, Q3SX55, Q96KB6 | Gene names | CD248, CD164L1, TEM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
TIMD1_HUMAN
|
||||||
| NC score | 0.021785 (rank : 100) | θ value | 3.0926 (rank : 100) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96D42, O43656 | Gene names | HAVCR1, TIM1, TIMD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatitis A virus cellular receptor 1 precursor (HAVcr-1) (T cell immunoglobulin and mucin domain-containing protein 1) (TIMD-1) (T cell membrane protein 1) (TIM-1) (TIM). | |||||
|
TTLL5_MOUSE
|
||||||
| NC score | 0.021006 (rank : 101) | θ value | 0.365318 (rank : 62) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CHB8 | Gene names | Ttll5, Kiaa0998 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 5. | |||||
|
APOB_HUMAN
|
||||||
| NC score | 0.020144 (rank : 102) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P04114, O00502, P78479, P78480, P78481, Q13779, Q13785, Q13786, Q13787, Q13788, Q7Z600, Q9UMN0 | Gene names | APOB | |||
|
Domain Architecture |

|
|||||
| Description | Apolipoprotein B-100 precursor (Apo B-100) [Contains: Apolipoprotein B-48 (Apo B-48)]. | |||||
|
TAF1C_MOUSE
|
||||||
| NC score | 0.018729 (rank : 103) | θ value | 5.27518 (rank : 115) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
TAF3_HUMAN
|
||||||
| NC score | 0.018600 (rank : 104) | θ value | 1.81305 (rank : 90) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.018576 (rank : 105) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
MYH10_HUMAN
|
||||||
| NC score | 0.018510 (rank : 106) | θ value | 1.06291 (rank : 74) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
RANB3_HUMAN
|
||||||
| NC score | 0.018471 (rank : 107) | θ value | 1.81305 (rank : 89) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H6Z4, O60405, O75759, O75760, Q9BT47, Q9UG74 | Gene names | RANBP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 3 (RanBP3). | |||||
|
RN165_HUMAN
|
||||||
| NC score | 0.018460 (rank : 108) | θ value | 1.06291 (rank : 75) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZSG1 | Gene names | RNF165 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 165. | |||||
|
MYH10_MOUSE
|
||||||
| NC score | 0.018268 (rank : 109) | θ value | 1.38821 (rank : 81) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
RFX1_HUMAN
|
||||||
| NC score | 0.018048 (rank : 110) | θ value | 5.27518 (rank : 112) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P22670 | Gene names | RFX1 | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II regulatory factor RFX1 (RFX) (Enhancer factor C) (EF-C). | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.017678 (rank : 111) | θ value | 2.36792 (rank : 93) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | 0.017515 (rank : 112) | θ value | 2.36792 (rank : 92) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
CO039_MOUSE
|
||||||
| NC score | 0.017184 (rank : 113) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
ATOH1_HUMAN
|
||||||
| NC score | 0.016670 (rank : 114) | θ value | 1.81305 (rank : 84) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92858 | Gene names | ATOH1, ATH1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein hATH-1). | |||||
|
AMOT_HUMAN
|
||||||
| NC score | 0.016391 (rank : 115) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
GAK2_HUMAN
|
||||||
| NC score | 0.015991 (rank : 116) | θ value | 1.38821 (rank : 78) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.015117 (rank : 117) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
GAK5_HUMAN
|
||||||
| NC score | 0.014640 (rank : 118) | θ value | 4.03905 (rank : 104) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
HCN1_MOUSE
|
||||||
| NC score | 0.014261 (rank : 119) | θ value | 0.813845 (rank : 68) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O88704, O54899, Q9D613 | Gene names | Hcn1, Bcng1, Hac2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1) (Hyperpolarization-activated cation channel 2) (HAC-2). | |||||
|
K0240_MOUSE
|
||||||
| NC score | 0.014151 (rank : 120) | θ value | 5.27518 (rank : 110) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CHH5 | Gene names | Kiaa0240 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0240. | |||||
|
GAK6_HUMAN
|
||||||
| NC score | 0.013899 (rank : 121) | θ value | 6.88961 (rank : 119) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
HAX1_MOUSE
|
||||||
| NC score | 0.013808 (rank : 122) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35387 | Gene names | Hax1, Hs1bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HS1-associating protein X-1 (HAX-1) (HS1-binding protein). | |||||
|
GAK4_HUMAN
|
||||||
| NC score | 0.013602 (rank : 123) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK11_HUMAN
|
||||||
| NC score | 0.013597 (rank : 124) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK12_HUMAN
|
||||||
| NC score | 0.013529 (rank : 125) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
PO2F1_MOUSE
|
||||||
| NC score | 0.012660 (rank : 126) | θ value | 0.47712 (rank : 64) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P25425, Q61994, Q63891, Q6Y681, Q7TSD0, Q8BT04, Q8K570, Q99JH0, Q9WTZ4, Q9WTZ5, Q9WTZ6 | Gene names | Pou2f1, Oct-1, Otf-1, Otf1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 2, transcription factor 1 (Octamer-binding transcription factor 1) (Oct-1) (OTF-1) (NF-A1). | |||||
|
MUCEN_HUMAN
|
||||||
| NC score | 0.012657 (rank : 127) | θ value | 8.99809 (rank : 148) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULC0, Q8NEY5, Q8WWE7, Q9NRM8 | Gene names | EMCN, EMCN2. MUC14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endomucin precursor (Endomucin-2) (Gastric cancer antigen Ga34). | |||||
|
RUFY1_HUMAN
|
||||||
| NC score | 0.012508 (rank : 128) | θ value | 6.88961 (rank : 128) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 848 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96T51, Q59FF3, Q71S93, Q9H6I3 | Gene names | RUFY1, RABIP4, ZFYVE12 | |||
|
Domain Architecture |

|
|||||
| Description | RUN and FYVE domain-containing protein 1 (FYVE-finger protein EIP1) (Zinc finger FYVE domain-containing protein 12) (La-binding protein 1) (Rab4-interacting protein). | |||||
|
TEKT2_HUMAN
|
||||||
| NC score | 0.012407 (rank : 129) | θ value | 8.99809 (rank : 150) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UIF3, O60638 | Gene names | TEKT2 | |||
|
Domain Architecture |

|
|||||
| Description | Tektin-2 (Testicular tektin) (Tektin-t) (Testicular tektin B1-like protein) (Tektin-B1) (TEKTB1). | |||||
|
TTLL4_HUMAN
|
||||||
| NC score | 0.012040 (rank : 130) | θ value | 6.88961 (rank : 131) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14679, Q8WW29 | Gene names | TTLL4, KIAA0173 | |||
|
Domain Architecture |

|
|||||
| Description | Tubulin--tyrosine ligase-like protein 4. | |||||
|
RFFL_HUMAN
|
||||||
| NC score | 0.011680 (rank : 131) | θ value | 3.0926 (rank : 96) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WZ73, Q8NHW0, Q8TBY7, Q96BE6 | Gene names | RFFL, RNF189, RNF34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (FYVE-RING finger protein Sakura) (Fring) (Caspases-8 and -10- associated RING finger protein 2) (CARP-2) (Caspase regulator CARP2) (RING finger protein 189) (RING finger protein 34-like). | |||||
|
AMOL1_MOUSE
|
||||||
| NC score | 0.011306 (rank : 132) | θ value | 8.99809 (rank : 135) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
GMEB2_HUMAN
|
||||||
| NC score | 0.011072 (rank : 133) | θ value | 4.03905 (rank : 105) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKD1, Q5TDS0, Q9H431, Q9H4X7, Q9H4X8, Q9UF78, Q9ULF1 | Gene names | GMEB2, KIAA1269 | |||
|
Domain Architecture |

|
|||||
| Description | Glucocorticoid modulatory element-binding protein 2 (GMEB-2) (Parvovirus initiation factor p79) (PIF p79) (DNA-binding protein p79PIF). | |||||
|
BRCA1_MOUSE
|
||||||
| NC score | 0.010432 (rank : 134) | θ value | 1.81305 (rank : 87) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48754, Q60957, Q60983 | Gene names | Brca1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein homolog. | |||||
|
AKAP2_HUMAN
|
||||||
| NC score | 0.010278 (rank : 135) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
IF35_HUMAN
|
||||||
| NC score | 0.009867 (rank : 136) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O00303, Q8N978 | Gene names | EIF3S5 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 5 (eIF-3 epsilon) (eIF3 p47 subunit) (eIF3f). | |||||
|
KIF17_MOUSE
|
||||||
| NC score | 0.009807 (rank : 137) | θ value | 3.0926 (rank : 94) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99PW8 | Gene names | Kif17 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF17 (MmKIF17). | |||||
|
XKR5_MOUSE
|
||||||
| NC score | 0.009379 (rank : 138) | θ value | 5.27518 (rank : 117) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5GH66 | Gene names | Xkr5, Xrg5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | XK-related protein 5. | |||||
|
KIF17_HUMAN
|
||||||
| NC score | 0.008592 (rank : 139) | θ value | 1.38821 (rank : 79) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P2E2, O95077, Q53YS6, Q5VWA9, Q6GSA8, Q8N411 | Gene names | KIF17, KIAA1405, KIF3X | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF17 (KIF3-related motor protein). | |||||
|
PI51A_HUMAN
|
||||||
| NC score | 0.008476 (rank : 140) | θ value | 3.0926 (rank : 95) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99755, Q99754, Q99756 | Gene names | PIP5K1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 5-kinase type-1 alpha (EC 2.7.1.68) (Phosphatidylinositol-4-phosphate 5-kinase type I alpha) (PtdIns(4)P- 5-kinase alpha) (PIP5KIalpha) (68 kDa type I phosphatidylinositol-4- phosphate 5-kinase alpha). | |||||
|
CEP63_HUMAN
|
||||||
| NC score | 0.008023 (rank : 141) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1005 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96MT8, Q96CR0, Q9H8F5, Q9H8N0 | Gene names | CEP63 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 63 kDa (Cep63 protein). | |||||
|
WNK1_MOUSE
|
||||||
| NC score | 0.008020 (rank : 142) | θ value | 0.365318 (rank : 63) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1179 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P83741 | Gene names | Wnk1, Prkwnk1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1). | |||||
|
GOGA5_MOUSE
|
||||||
| NC score | 0.007998 (rank : 143) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
MCM6_MOUSE
|
||||||
| NC score | 0.007353 (rank : 144) | θ value | 6.88961 (rank : 123) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97311, Q80YQ7 | Gene names | Mcm6, Mcmd6, Mis5 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM6 (Mis5 homolog). | |||||
|
WNK2_HUMAN
|
||||||
| NC score | 0.007259 (rank : 145) | θ value | 6.88961 (rank : 132) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
CBL_MOUSE
|
||||||
| NC score | 0.007241 (rank : 146) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P22682, Q8CEA1 | Gene names | Cbl | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase CBL (EC 6.3.2.-) (Signal transduction protein CBL) (Proto-oncogene c-CBL) (Casitas B-lineage lymphoma proto- oncogene). | |||||
|
ELF1_MOUSE
|
||||||
| NC score | 0.006854 (rank : 147) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q60775 | Gene names | Elf1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
NMNA1_MOUSE
|
||||||
| NC score | 0.006534 (rank : 148) | θ value | 8.99809 (rank : 149) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9EPA7, Q6B504 | Gene names | Nmnat1, D4Cole1e, Nmnat | |||
|
Domain Architecture |

|
|||||
| Description | Nicotinamide mononucleotide adenylyltransferase 1 (EC 2.7.7.1) (NMN adenylyltransferase 1). | |||||
|
LIPA1_HUMAN
|
||||||
| NC score | 0.006245 (rank : 149) | θ value | 5.27518 (rank : 111) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.005681 (rank : 150) | θ value | 1.06291 (rank : 73) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
ADA12_MOUSE
|
||||||
| NC score | 0.004716 (rank : 151) | θ value | 1.81305 (rank : 83) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61824 | Gene names | Adam12, Mltna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
ADA19_HUMAN
|
||||||
| NC score | 0.003874 (rank : 152) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.003278 (rank : 153) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
DLX5_MOUSE
|
||||||
| NC score | 0.003100 (rank : 154) | θ value | 2.36792 (rank : 91) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70396, O54876, O54877, O54878, Q9JJ45 | Gene names | Dlx5 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein DLX-5. | |||||
|
LTB1L_HUMAN
|
||||||
| NC score | 0.002560 (rank : 155) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14766 | Gene names | LTBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein, isoform 1L precursor (LTBP-1) (Transforming growth factor beta-1-binding protein 1) (TGF-beta1-BP-1). | |||||
|
LTB1S_HUMAN
|
||||||
| NC score | 0.002506 (rank : 156) | θ value | 8.99809 (rank : 147) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P22064, Q8TD95 | Gene names | LTBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein, isoform 1S precursor (LTBP-1) (Transforming growth factor beta-1-binding protein 1) (TGF-beta1-BP-1). | |||||
|
POGK_MOUSE
|
||||||
| NC score | 0.001875 (rank : 157) | θ value | 6.88961 (rank : 127) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80TC5, Q8BPJ3, Q8C887 | Gene names | Pogk, Kiaa1513 | |||
|
Domain Architecture |

|
|||||
| Description | Pogo transposable element with KRAB domain. | |||||
|
ULK1_HUMAN
|
||||||
| NC score | -0.000603 (rank : 158) | θ value | 0.813845 (rank : 70) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1121 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O75385 | Gene names | ULK1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase ULK1 (EC 2.7.11.1) (Unc-51-like kinase 1). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | -0.000892 (rank : 159) | θ value | 6.88961 (rank : 130) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
MK07_MOUSE
|
||||||
| NC score | -0.001179 (rank : 160) | θ value | 6.88961 (rank : 124) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1054 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9WVS8 | Gene names | Mapk7, Erk5 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (BMK1 kinase). | |||||
|
CDK8_HUMAN
|
||||||
| NC score | -0.001377 (rank : 161) | θ value | 4.03905 (rank : 103) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P49336 | Gene names | CDK8 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division protein kinase 8 (EC 2.7.11.22) (EC 2.7.11.23) (Protein kinase K35). | |||||
|
STK39_MOUSE
|
||||||
| NC score | -0.002592 (rank : 162) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 915 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Z1W9, Q80W13 | Gene names | Stk39, Spak | |||
|
Domain Architecture |

|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39). | |||||