Please be patient as the page loads
|
KCNH8_HUMAN
|
||||||
| SwissProt Accessions | Q96L42 | Gene names | KCNH8 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 8 (Voltage-gated potassium channel subunit Kv12.1) (Ether-a-go-go-like potassium channel 3) (ELK channel 3) (ELK3) (ELK1) (hElk1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
KCNH3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.984717 (rank : 4) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9ULD8, Q9UQ06 | Gene names | KCNH3, KIAA1282 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (ELK2) (Brain-specific eag-like channel 1) (BEC1). | |||||
|
KCNH3_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.984758 (rank : 3) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9WVJ0 | Gene names | Kcnh3, Elk2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (mElk2). | |||||
|
KCNH4_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.988115 (rank : 2) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9UQ05 | Gene names | KCNH4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 4 (Voltage-gated potassium channel subunit Kv12.3) (Ether-a-go-go-like potassium channel 1) (ELK channel 1) (ELK1) (Brain-specific eag-like channel 2) (BEC2). | |||||
|
KCNH8_HUMAN
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 104 | |
| SwissProt Accessions | Q96L42 | Gene names | KCNH8 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 8 (Voltage-gated potassium channel subunit Kv12.1) (Ether-a-go-go-like potassium channel 3) (ELK channel 3) (ELK3) (ELK1) (hElk1). | |||||
|
KCNH6_HUMAN
|
||||||
| θ value | 2.95243e-160 (rank : 5) | NC score | 0.968133 (rank : 9) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9H252, Q9BRD7 | Gene names | KCNH6, ERG2, HERG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 6 (Voltage-gated potassium channel subunit Kv11.2) (Ether-a-go-go-related gene potassium channel 2) (Ether-a-go-go-related protein 2) (Eag-related protein 2). | |||||
|
KCNH1_HUMAN
|
||||||
| θ value | 1.28961e-139 (rank : 6) | NC score | 0.968590 (rank : 6) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O95259, O76035 | Gene names | KCNH1, EAG, EAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (hEAG1) (h-eag). | |||||
|
KCNH5_MOUSE
|
||||||
| θ value | 1.28961e-139 (rank : 7) | NC score | 0.968588 (rank : 7) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q920E3, Q8C035 | Gene names | Kcnh5, Eag2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (Eag2). | |||||
|
KCNH1_MOUSE
|
||||||
| θ value | 1.68428e-139 (rank : 8) | NC score | 0.968716 (rank : 5) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q60603 | Gene names | Kcnh1, Eag | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (EAG1) (m-eag). | |||||
|
KCNH5_HUMAN
|
||||||
| θ value | 2.19974e-139 (rank : 9) | NC score | 0.968428 (rank : 8) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8NCM2 | Gene names | KCNH5, EAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (hEAG2). | |||||
|
KCNH7_MOUSE
|
||||||
| θ value | 5.61992e-127 (rank : 10) | NC score | 0.960306 (rank : 11) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9ER47 | Gene names | Kcnh7, Erg3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (Ether-a-go-go-related protein 3) (Eag-related protein 3). | |||||
|
KCNH2_HUMAN
|
||||||
| θ value | 4.75757e-126 (rank : 11) | NC score | 0.960370 (rank : 10) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q12809, O75418, O75680, Q9BT72, Q9BUT7, Q9H3P0 | Gene names | KCNH2, ERG, ERG1, HERG, HERG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (H-ERG) (Erg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1) (eag homolog). | |||||
|
KCNH2_MOUSE
|
||||||
| θ value | 6.21357e-126 (rank : 12) | NC score | 0.960275 (rank : 12) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O35219, O35220, O35221, O35989 | Gene names | Kcnh2, Erg, Merg1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (ERG1) (MERG) (Merg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1). | |||||
|
KCNH7_HUMAN
|
||||||
| θ value | 1.99884e-124 (rank : 13) | NC score | 0.959333 (rank : 13) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9NS40 | Gene names | KCNH7, ERG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (HERG-3) (Ether-a-go-go-related protein 3) (Eag- related protein 3). | |||||
|
CNGA1_HUMAN
|
||||||
| θ value | 8.61488e-35 (rank : 14) | NC score | 0.808073 (rank : 15) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P29973, Q16279, Q16485 | Gene names | CNGA1, CNCG, CNCG1 | |||
|
Domain Architecture |
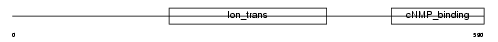
|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
HCN4_MOUSE
|
||||||
| θ value | 1.9192e-34 (rank : 15) | NC score | 0.772171 (rank : 25) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O70507 | Gene names | Hcn4, Bcng3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4 (Brain cyclic nucleotide-gated channel 3) (BCNG-3). | |||||
|
CNGA1_MOUSE
|
||||||
| θ value | 9.52487e-34 (rank : 16) | NC score | 0.803774 (rank : 17) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P29974, Q60776 | Gene names | Cnga1, Cncg, Cncg1 | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
HCN4_HUMAN
|
||||||
| θ value | 3.61944e-33 (rank : 17) | NC score | 0.773961 (rank : 24) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9Y3Q4, Q9UMQ7 | Gene names | HCN4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4. | |||||
|
CNGA2_MOUSE
|
||||||
| θ value | 4.72714e-33 (rank : 18) | NC score | 0.817702 (rank : 14) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q62398 | Gene names | Cnga2, Cncg2, Cncg4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated olfactory channel (Cyclic nucleotide-gated cation channel 2) (CNG channel 2) (CNG-2) (CNG2). | |||||
|
HCN2_MOUSE
|
||||||
| θ value | 4.72714e-33 (rank : 19) | NC score | 0.776791 (rank : 23) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O88703, O70506 | Gene names | Hcn2, Bcng2, Hac1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2) (Hyperpolarization-activated cation channel 1) (HAC-1). | |||||
|
CNGA3_HUMAN
|
||||||
| θ value | 8.06329e-33 (rank : 20) | NC score | 0.802989 (rank : 18) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q16281, Q9UP64 | Gene names | CNGA3, CNCG3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel alpha 3 (CNG channel alpha 3) (CNG-3) (CNG3) (Cyclic nucleotide-gated channel alpha 3) (Cone photoreceptor cGMP-gated channel subunit alpha). | |||||
|
HCN2_HUMAN
|
||||||
| θ value | 1.0531e-32 (rank : 21) | NC score | 0.767470 (rank : 26) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UL51, O60742, O60743, O75267, Q9UBS2 | Gene names | HCN2, BCNG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2). | |||||
|
HCN1_MOUSE
|
||||||
| θ value | 5.22648e-32 (rank : 22) | NC score | 0.789504 (rank : 20) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O88704, O54899, Q9D613 | Gene names | Hcn1, Bcng1, Hac2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1) (Hyperpolarization-activated cation channel 2) (HAC-2). | |||||
|
HCN1_HUMAN
|
||||||
| θ value | 6.82597e-32 (rank : 23) | NC score | 0.793221 (rank : 19) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O60741 | Gene names | HCN1, BCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1). | |||||
|
CNGA3_MOUSE
|
||||||
| θ value | 8.91499e-32 (rank : 24) | NC score | 0.806520 (rank : 16) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JJZ8, Q9WV01 | Gene names | Cnga3, Cng3 | |||
|
Domain Architecture |
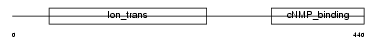
|
|||||
| Description | Cyclic nucleotide-gated cation channel alpha 3 (CNG channel alpha 3) (CNG-3) (CNG3) (Cyclic nucleotide-gated channel alpha 3) (Cone photoreceptor cGMP-gated channel subunit alpha). | |||||
|
HCN3_HUMAN
|
||||||
| θ value | 4.57862e-28 (rank : 25) | NC score | 0.787354 (rank : 21) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9P1Z3, Q4VX12, Q8N6W6, Q9BWQ2 | Gene names | HCN3, KIAA1535 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3. | |||||
|
HCN3_MOUSE
|
||||||
| θ value | 1.33217e-27 (rank : 26) | NC score | 0.787246 (rank : 22) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O88705 | Gene names | Hcn3, Hac3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3 (Hyperpolarization-activated cation channel 3) (HAC-3). | |||||
|
CNGB3_MOUSE
|
||||||
| θ value | 9.56915e-18 (rank : 27) | NC score | 0.741179 (rank : 27) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9JJZ9 | Gene names | Cngb3, Cng6 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel beta 3 (CNG channel beta 3) (Cyclic nucleotide-gated channel beta 3) (Cone photoreceptor cGMP- gated channel subunit beta) (Cyclic nucleotide-gated cation channel modulatory subunit) (Cyclic nucleotide-gated channel subunit CNG6). | |||||
|
CNGB3_HUMAN
|
||||||
| θ value | 3.63628e-17 (rank : 28) | NC score | 0.728519 (rank : 28) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9NQW8, Q9NRE9 | Gene names | CNGB3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel beta 3 (CNG channel beta 3) (Cyclic nucleotide-gated channel beta 3) (Cone photoreceptor cGMP- gated channel subunit beta) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
CNGB1_HUMAN
|
||||||
| θ value | 1.16975e-15 (rank : 29) | NC score | 0.724180 (rank : 29) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q14028, Q14029 | Gene names | CNGB1, CNCG4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel 4 (CNG channel 4) (CNG-4) (CNG4) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
KCNKH_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 30) | NC score | 0.258199 (rank : 30) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96T54, Q5TCF4, Q8TAW4, Q9BXD1, Q9H592 | Gene names | KCNK17, TALK2, TASK4 | |||
|
Domain Architecture |
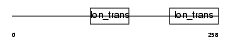
|
|||||
| Description | Potassium channel subfamily K member 17 (TWIK-related alkaline pH- activated K(+) channel 2) (2P domain potassium channel Talk-2) (TWIK- related acid-sensitive K(+) channel 4) (TASK-4). | |||||
|
KCNK2_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 31) | NC score | 0.188297 (rank : 31) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O95069, Q9UNE3 | Gene names | KCNK2, TREK, TREK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (TREK-1 K(+) channel subunit) (Two-pore potassium channel TPKC1). | |||||
|
KCNK2_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 32) | NC score | 0.187561 (rank : 33) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P97438 | Gene names | Kcnk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (Two-pore potassium channel TPKC1) (TREK-1 K(+) channel subunit). | |||||
|
ANR21_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 33) | NC score | 0.013257 (rank : 89) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86YR6 | Gene names | ANKRD21, POTE | |||
|
Domain Architecture |
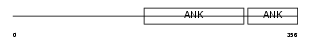
|
|||||
| Description | Ankyrin repeat domain-containing protein 21 (Protein POTE) (Prostate, ovary, testis-expressed protein). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 34) | NC score | 0.042796 (rank : 70) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
KCNS2_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 35) | NC score | 0.052243 (rank : 68) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9ULS6 | Gene names | KCNS2, KIAA1144 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNS2_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 36) | NC score | 0.051807 (rank : 69) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35174, Q543P3 | Gene names | Kcns2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNK4_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 37) | NC score | 0.172892 (rank : 38) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NYG8, Q96T94 | Gene names | KCNK4, TRAAK | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK) (Two pore K(+) channel KT4.1). | |||||
|
KCMA1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 38) | NC score | 0.081971 (rank : 52) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
KCMA1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 39) | NC score | 0.082080 (rank : 51) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCND1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 40) | NC score | 0.075400 (rank : 59) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
KCNK1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 41) | NC score | 0.133663 (rank : 43) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O00180, Q13307 | Gene names | KCNK1, HOHO1, KCNO1, TWIK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1) (Potassium channel KCNO1). | |||||
|
KCNK1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 42) | NC score | 0.129494 (rank : 45) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O08581 | Gene names | Kcnk1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1). | |||||
|
KCNK6_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 43) | NC score | 0.180145 (rank : 35) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Y257, Q9HB47 | Gene names | KCNK6, TOSS, TWIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 6 (Inward rectifying potassium channel protein TWIK-2) (TWIK-originated similarity sequence). | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 44) | NC score | 0.031658 (rank : 77) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
KCND1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 45) | NC score | 0.081172 (rank : 53) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KCNK4_MOUSE
|
||||||
| θ value | 0.125558 (rank : 46) | NC score | 0.160256 (rank : 40) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
KCND2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 47) | NC score | 0.073591 (rank : 61) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 48) | NC score | 0.073620 (rank : 60) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
POT15_HUMAN
|
||||||
| θ value | 0.163984 (rank : 49) | NC score | 0.010994 (rank : 93) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6S5H4, Q6NXN7, Q6S5H7 | Gene names | POTE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 15. | |||||
|
IL7RA_MOUSE
|
||||||
| θ value | 0.21417 (rank : 50) | NC score | 0.027233 (rank : 83) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P16872 | Gene names | Il7r | |||
|
Domain Architecture |
|
|||||
| Description | Interleukin-7 receptor alpha chain precursor (IL-7R-alpha) (CD127 antigen). | |||||
|
KCNKA_HUMAN
|
||||||
| θ value | 0.21417 (rank : 51) | NC score | 0.181147 (rank : 34) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P57789, Q8TDK7, Q8TDK8, Q9HB59 | Gene names | KCNK10, TREK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 10 (Outward rectifying potassium channel protein TREK-2) (TREK-2 K(+) channel subunit). | |||||
|
KCNK3_HUMAN
|
||||||
| θ value | 0.279714 (rank : 52) | NC score | 0.179237 (rank : 36) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O14649 | Gene names | KCNK3, TASK, TASK1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Two pore potassium channel KT3.1). | |||||
|
KCNK3_MOUSE
|
||||||
| θ value | 0.279714 (rank : 53) | NC score | 0.175758 (rank : 37) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O35111, O35163 | Gene names | Kcnk3, Ctbak, Task, Task1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Cardiac two-pore background K(+) channel) (cTBAK-1) (Two pore potassium channel KT3.1). | |||||
|
KCNKF_HUMAN
|
||||||
| θ value | 0.279714 (rank : 54) | NC score | 0.188275 (rank : 32) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9H427, Q9HBC8 | Gene names | KCNK15, TASK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 15 (Acid-sensitive potassium channel protein TASK-5) (TWIK-related acid-sensitive K(+) channel 5) (Two pore potassium channel KT3.3). | |||||
|
IRK13_HUMAN
|
||||||
| θ value | 0.365318 (rank : 55) | NC score | 0.014596 (rank : 86) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60928, O76023, Q8N3Y4 | Gene names | KCNJ13 | |||
|
Domain Architecture |
|
|||||
| Description | Inward rectifier potassium channel 13 (Potassium channel, inwardly rectifying subfamily J member 13) (Inward rectifier K(+) channel Kir7.1). | |||||
|
KCND3_HUMAN
|
||||||
| θ value | 0.365318 (rank : 56) | NC score | 0.071750 (rank : 63) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND3_MOUSE
|
||||||
| θ value | 0.365318 (rank : 57) | NC score | 0.071754 (rank : 62) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
AF17_HUMAN
|
||||||
| θ value | 0.813845 (rank : 58) | NC score | 0.013402 (rank : 88) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P55198, Q9UF49 | Gene names | MLLT6, AF17 | |||
|
Domain Architecture |
|
|||||
| Description | Protein AF-17. | |||||
|
KCNK9_HUMAN
|
||||||
| θ value | 0.813845 (rank : 59) | NC score | 0.151495 (rank : 42) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9NPC2, Q2M290, Q540F2 | Gene names | KCNK9, TASK3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3) (Two pore potassium channel KT3.2). | |||||
|
KCNK9_MOUSE
|
||||||
| θ value | 0.813845 (rank : 60) | NC score | 0.151517 (rank : 41) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q3LS21 | Gene names | Kcnk9, Task3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3). | |||||
|
LRCH2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 61) | NC score | 0.008148 (rank : 101) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UMG5, Q7TNP9, Q80TC9 | Gene names | Lrch2, Kiaa1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
KCNA4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.034769 (rank : 76) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
GP108_MOUSE
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.011742 (rank : 91) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91WD0, Q8BXE9, Q925B1 | Gene names | Gpr108, Lustr2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein GPR108 precursor (Lung seven transmembrane receptor 2). | |||||
|
KGP1A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.040156 (rank : 72) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q13976 | Gene names | PRKG1, PRKGR1A | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, alpha isozyme (EC 2.7.11.12) (CGK 1 alpha) (cGKI-alpha). | |||||
|
KGP1B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.040161 (rank : 71) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 921 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P14619 | Gene names | PRKG1, PRKG1B, PRKGR1B | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, beta isozyme (EC 2.7.11.12) (cGK 1 beta) (cGKI-beta). | |||||
|
NDE1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.004398 (rank : 108) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 823 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NXR1, Q49AQ2 | Gene names | NDE1, NUDE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear distribution protein nudE homolog 1 (NudE). | |||||
|
AP4E1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.009701 (rank : 98) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80V94 | Gene names | Ap4e1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-4 complex subunit epsilon-1 (Adapter-related protein complex 4 epsilon-1 subunit) (Epsilon subunit of AP-4) (AP-4 adapter complex epsilon subunit). | |||||
|
CKLF_HUMAN
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.026164 (rank : 84) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UBR5, Q9UHM7, Q9UHN8, Q9UI41 | Gene names | CKLF | |||
|
Domain Architecture |

|
|||||
| Description | Chemokine-like factor (C32). | |||||
|
HD_MOUSE
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.011008 (rank : 92) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42859 | Gene names | Hd, Hdh | |||
|
Domain Architecture |
|
|||||
| Description | Huntingtin (Huntington disease protein homolog) (HD protein). | |||||
|
KGP1B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.039755 (rank : 73) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Z0Z0 | Gene names | Prkg1, Prkg1b, Prkgr1b | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, beta isozyme (EC 2.7.11.12) (cGK 1 beta) (cGKI-beta). | |||||
|
KGP2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.028954 (rank : 79) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1023 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13237, O00125, O60916 | Gene names | PRKG2, PRKGR2 | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 2 (EC 2.7.11.12) (CGK 2) (cGKII) (Type II cGMP-dependent protein kinase). | |||||
|
LRCH2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.008797 (rank : 100) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5VUJ6, Q9HA88, Q9P233 | Gene names | LRCH2, KIAA1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
RAI14_HUMAN
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.000816 (rank : 113) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1367 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9P0K7, Q6V1W9, Q7Z5I4, Q7Z733, Q9P2L2, Q9Y3T5 | Gene names | RAI14, KIAA1334, NORPEG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein). | |||||
|
KCNKG_HUMAN
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.164679 (rank : 39) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q96T55, Q5TCF3, Q9H591 | Gene names | KCNK16, TALK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 16 (TWIK-related alkaline pH- activated K(+) channel 1) (2P domain potassium channel Talk-1). | |||||
|
NAB2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 75) | NC score | 0.010081 (rank : 97) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61127 | Gene names | Nab2 | |||
|
Domain Architecture |

|
|||||
| Description | NGFI-A-binding protein 2 (EGR-1-binding protein 2). | |||||
|
TRI73_HUMAN
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.005835 (rank : 106) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86UV7, Q8N0S3 | Gene names | TRIM73, TRIM50B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 73 (Tripartite motif-containing protein 50B). | |||||
|
ZHX3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.006807 (rank : 105) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H4I2, O43145 | Gene names | ZHX3, KIAA0395, TIX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and homeodomain protein 3 (Triple homeobox protein 1). | |||||
|
BRSK2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | -0.002672 (rank : 116) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 897 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IWQ3, O60843, O95099, Q5J5B4, Q8TB60 | Gene names | BRSK2, C11orf7, PEN11B, STK29 | |||
|
Domain Architecture |

|
|||||
| Description | BR serine/threonine-protein kinase 2 (EC 2.7.11.1) (Serine/threonine- protein kinase 29) (SAD1B). | |||||
|
KCNN4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 79) | NC score | 0.038761 (rank : 75) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O89109, Q3U1J8, Q9CY39 | Gene names | Kcnn4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1). | |||||
|
MTR1L_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | -0.001526 (rank : 115) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1180 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13585 | Gene names | GPR50 | |||
|
Domain Architecture |

|
|||||
| Description | Melatonin-related receptor (G protein-coupled receptor 50) (H9). | |||||
|
OR2V2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | -0.004130 (rank : 118) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96R30, Q6IFL6, Q8NGV1 | Gene names | OR2V2, OR2V3 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 2V2 (Olfactory receptor OR5-3). | |||||
|
PROM1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.012987 (rank : 90) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54990, O35408 | Gene names | Prom1, Prom, Proml1 | |||
|
Domain Architecture |

|
|||||
| Description | Prominin-1 precursor (Prominin-like protein 1) (Antigen AC133 homolog). | |||||
|
APOA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | -0.000148 (rank : 114) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P08519 | Gene names | LPA | |||
|
Domain Architecture |

|
|||||
| Description | Apolipoprotein(a) precursor (EC 3.4.21.-) (Apo(a)) (Lp(a)). | |||||
|
KCNA4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.030935 (rank : 78) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61423 | Gene names | Kcna4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4). | |||||
|
KCNN2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.028771 (rank : 80) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9H2S1 | Gene names | KCNN2 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | 0.028748 (rank : 81) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P58390 | Gene names | Kcnn2, Sk2 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KGP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 87) | NC score | 0.028199 (rank : 82) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1029 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61410 | Gene names | Prkg2, Prkgr2 | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 2 (EC 2.7.11.12) (CGK 2) (cGKII) (Type II cGMP-dependent protein kinase). | |||||
|
LARP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | 0.018263 (rank : 85) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6PKG0, O94836, Q8N4M2, Q8NB73, Q9UFD7 | Gene names | LARP1, KIAA0731, LARP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 1 (La ribonucleoprotein domain family member 1). | |||||
|
T2FA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.009622 (rank : 99) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
TINF2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 90) | NC score | 0.010810 (rank : 95) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BSI4, Q9H904, Q9UHC2 | Gene names | TINF2, TIN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TERF1-interacting nuclear factor 2 (TRF1-interacting nuclear protein 2). | |||||
|
TRI50_MOUSE
|
||||||
| θ value | 5.27518 (rank : 91) | NC score | 0.002536 (rank : 111) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q810I2 | Gene names | Trim50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 50. | |||||
|
BCL7A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.006859 (rank : 104) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CXE2, Q8C361, Q8C8M8, Q8VD15 | Gene names | Bcl7a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell CLL/lymphoma 7 protein family member A. | |||||
|
FA35A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.007902 (rank : 102) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3UEN2 | Gene names | Fam35a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
KCNB2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.038897 (rank : 74) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
KCNK5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.130061 (rank : 44) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.003582 (rank : 109) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
RFIP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.007494 (rank : 103) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D620, Q3UBC2, Q8BN24 | Gene names | Rab11fip1, Rcp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
RPGR_MOUSE
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.003169 (rank : 110) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
TRI74_HUMAN
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.004809 (rank : 107) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86UV6 | Gene names | TRIM74, TRIM50C | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 74 (Tripartite motif-containing protein 50C). | |||||
|
AGGF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.013791 (rank : 87) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N302, O00581, Q53YS3, Q9BU84, Q9NW66 | Gene names | AGGF1, VG5Q | |||
|
Domain Architecture |
|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (Vasculogenesis gene on 5q protein) (hVG5Q). | |||||
|
CEP35_HUMAN
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.002024 (rank : 112) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
MTR1L_MOUSE
|
||||||
| θ value | 8.99809 (rank : 102) | NC score | -0.003327 (rank : 117) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88495 | Gene names | Gpr50 | |||
|
Domain Architecture |
|
|||||
| Description | Melatonin-related receptor (G protein-coupled receptor 50) (H9). | |||||
|
PROM1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 103) | NC score | 0.010856 (rank : 94) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43490 | Gene names | PROM1, PROML1 | |||
|
Domain Architecture |

|
|||||
| Description | Prominin-1 precursor (Prominin-like protein 1) (Antigen AC133) (CD133 antigen). | |||||
|
TP53B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.010654 (rank : 96) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
CT152_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.060555 (rank : 66) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9D5U8, Q8R083 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | Uncharacterized protein C20orf152 homolog. | |||||
|
KAP0_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.128849 (rank : 46) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P10644, Q567S7 | Gene names | PRKAR1A, PKR1, PRKAR1 | |||
|
Domain Architecture |
|
|||||
| Description | cAMP-dependent protein kinase type I-alpha regulatory subunit (Tissue- specific extinguisher 1) (TSE1). | |||||
|
KAP0_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.117243 (rank : 47) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9DBC7, Q3UKU7, Q9JHR5, Q9JHR6 | Gene names | Prkar1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | cAMP-dependent protein kinase type I-alpha regulatory subunit. | |||||
|
KAP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.106325 (rank : 48) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P31321 | Gene names | PRKAR1B | |||
|
Domain Architecture |
|
|||||
| Description | cAMP-dependent protein kinase type I-beta regulatory subunit. | |||||
|
KAP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.103875 (rank : 49) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P12849 | Gene names | Prkar1b | |||
|
Domain Architecture |
|
|||||
| Description | cAMP-dependent protein kinase type I-beta regulatory subunit. | |||||
|
KAP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.075581 (rank : 58) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P13861, Q16823 | Gene names | PRKAR2A, PKR2, PRKAR2 | |||
|
Domain Architecture |
|
|||||
| Description | cAMP-dependent protein kinase type II-alpha regulatory subunit. | |||||
|
KAP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.077626 (rank : 55) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P12367 | Gene names | Prkar2a | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type II-alpha regulatory subunit. | |||||
|
KAP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.075624 (rank : 57) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P31323 | Gene names | PRKAR2B | |||
|
Domain Architecture |
|
|||||
| Description | cAMP-dependent protein kinase type II-beta regulatory subunit. | |||||
|
KAP3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.078896 (rank : 54) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P31324, Q3UTZ1, Q80ZM4, Q8BRZ7 | Gene names | Prkar2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | cAMP-dependent protein kinase type II-beta regulatory subunit. | |||||
|
KCNK7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.058607 (rank : 67) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y2U2, Q9Y2U3, Q9Y2U4 | Gene names | KCNK7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7. | |||||
|
KCNK7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.076437 (rank : 56) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9Z2T1, Q9QXY0, Q9QYE8, Q9R1V1, Q9R242 | Gene names | Kcnk7, Dpkch3, Kcnk6, Kcnk8, Knot1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7 (Putative potassium channel DP3) (Double-pore K(+) channel 3) (Neuromuscular two p domain potassium channel). | |||||
|
KCNKC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.068393 (rank : 65) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9HB15 | Gene names | KCNK12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 12 (Tandem pore domain halothane- inhibited potassium channel 2) (THIK-2). | |||||
|
KCNKD_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.071252 (rank : 64) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9HB14, Q96E79 | Gene names | KCNK13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNKD_MOUSE
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.089190 (rank : 50) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8R1P5 | Gene names | Kcnk13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNH8_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 104 | |
| SwissProt Accessions | Q96L42 | Gene names | KCNH8 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 8 (Voltage-gated potassium channel subunit Kv12.1) (Ether-a-go-go-like potassium channel 3) (ELK channel 3) (ELK3) (ELK1) (hElk1). | |||||
|
KCNH4_HUMAN
|
||||||
| NC score | 0.988115 (rank : 2) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9UQ05 | Gene names | KCNH4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 4 (Voltage-gated potassium channel subunit Kv12.3) (Ether-a-go-go-like potassium channel 1) (ELK channel 1) (ELK1) (Brain-specific eag-like channel 2) (BEC2). | |||||
|
KCNH3_MOUSE
|
||||||
| NC score | 0.984758 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9WVJ0 | Gene names | Kcnh3, Elk2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (mElk2). | |||||
|
KCNH3_HUMAN
|
||||||
| NC score | 0.984717 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9ULD8, Q9UQ06 | Gene names | KCNH3, KIAA1282 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (ELK2) (Brain-specific eag-like channel 1) (BEC1). | |||||
|
KCNH1_MOUSE
|
||||||
| NC score | 0.968716 (rank : 5) | θ value | 1.68428e-139 (rank : 8) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q60603 | Gene names | Kcnh1, Eag | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (EAG1) (m-eag). | |||||
|
KCNH1_HUMAN
|
||||||
| NC score | 0.968590 (rank : 6) | θ value | 1.28961e-139 (rank : 6) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O95259, O76035 | Gene names | KCNH1, EAG, EAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (hEAG1) (h-eag). | |||||
|
KCNH5_MOUSE
|
||||||
| NC score | 0.968588 (rank : 7) | θ value | 1.28961e-139 (rank : 7) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q920E3, Q8C035 | Gene names | Kcnh5, Eag2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (Eag2). | |||||
|
KCNH5_HUMAN
|
||||||
| NC score | 0.968428 (rank : 8) | θ value | 2.19974e-139 (rank : 9) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8NCM2 | Gene names | KCNH5, EAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (hEAG2). | |||||
|
KCNH6_HUMAN
|
||||||
| NC score | 0.968133 (rank : 9) | θ value | 2.95243e-160 (rank : 5) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9H252, Q9BRD7 | Gene names | KCNH6, ERG2, HERG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 6 (Voltage-gated potassium channel subunit Kv11.2) (Ether-a-go-go-related gene potassium channel 2) (Ether-a-go-go-related protein 2) (Eag-related protein 2). | |||||
|
KCNH2_HUMAN
|
||||||
| NC score | 0.960370 (rank : 10) | θ value | 4.75757e-126 (rank : 11) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q12809, O75418, O75680, Q9BT72, Q9BUT7, Q9H3P0 | Gene names | KCNH2, ERG, ERG1, HERG, HERG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (H-ERG) (Erg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1) (eag homolog). | |||||
|
KCNH7_MOUSE
|
||||||
| NC score | 0.960306 (rank : 11) | θ value | 5.61992e-127 (rank : 10) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9ER47 | Gene names | Kcnh7, Erg3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (Ether-a-go-go-related protein 3) (Eag-related protein 3). | |||||
|
KCNH2_MOUSE
|
||||||
| NC score | 0.960275 (rank : 12) | θ value | 6.21357e-126 (rank : 12) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O35219, O35220, O35221, O35989 | Gene names | Kcnh2, Erg, Merg1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (ERG1) (MERG) (Merg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1). | |||||
|
KCNH7_HUMAN
|
||||||
| NC score | 0.959333 (rank : 13) | θ value | 1.99884e-124 (rank : 13) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9NS40 | Gene names | KCNH7, ERG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (HERG-3) (Ether-a-go-go-related protein 3) (Eag- related protein 3). | |||||
|
CNGA2_MOUSE
|
||||||
| NC score | 0.817702 (rank : 14) | θ value | 4.72714e-33 (rank : 18) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q62398 | Gene names | Cnga2, Cncg2, Cncg4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated olfactory channel (Cyclic nucleotide-gated cation channel 2) (CNG channel 2) (CNG-2) (CNG2). | |||||
|
CNGA1_HUMAN
|
||||||
| NC score | 0.808073 (rank : 15) | θ value | 8.61488e-35 (rank : 14) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P29973, Q16279, Q16485 | Gene names | CNGA1, CNCG, CNCG1 | |||
|
Domain Architecture |
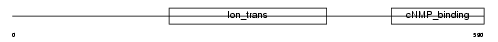
|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
CNGA3_MOUSE
|
||||||
| NC score | 0.806520 (rank : 16) | θ value | 8.91499e-32 (rank : 24) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JJZ8, Q9WV01 | Gene names | Cnga3, Cng3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel alpha 3 (CNG channel alpha 3) (CNG-3) (CNG3) (Cyclic nucleotide-gated channel alpha 3) (Cone photoreceptor cGMP-gated channel subunit alpha). | |||||
|
CNGA1_MOUSE
|
||||||
| NC score | 0.803774 (rank : 17) | θ value | 9.52487e-34 (rank : 16) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P29974, Q60776 | Gene names | Cnga1, Cncg, Cncg1 | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
CNGA3_HUMAN
|
||||||
| NC score | 0.802989 (rank : 18) | θ value | 8.06329e-33 (rank : 20) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q16281, Q9UP64 | Gene names | CNGA3, CNCG3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel alpha 3 (CNG channel alpha 3) (CNG-3) (CNG3) (Cyclic nucleotide-gated channel alpha 3) (Cone photoreceptor cGMP-gated channel subunit alpha). | |||||
|
HCN1_HUMAN
|
||||||
| NC score | 0.793221 (rank : 19) | θ value | 6.82597e-32 (rank : 23) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O60741 | Gene names | HCN1, BCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1). | |||||
|
HCN1_MOUSE
|
||||||
| NC score | 0.789504 (rank : 20) | θ value | 5.22648e-32 (rank : 22) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O88704, O54899, Q9D613 | Gene names | Hcn1, Bcng1, Hac2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1) (Hyperpolarization-activated cation channel 2) (HAC-2). | |||||
|
HCN3_HUMAN
|
||||||
| NC score | 0.787354 (rank : 21) | θ value | 4.57862e-28 (rank : 25) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9P1Z3, Q4VX12, Q8N6W6, Q9BWQ2 | Gene names | HCN3, KIAA1535 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3. | |||||
|
HCN3_MOUSE
|
||||||
| NC score | 0.787246 (rank : 22) | θ value | 1.33217e-27 (rank : 26) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O88705 | Gene names | Hcn3, Hac3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3 (Hyperpolarization-activated cation channel 3) (HAC-3). | |||||
|
HCN2_MOUSE
|
||||||
| NC score | 0.776791 (rank : 23) | θ value | 4.72714e-33 (rank : 19) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O88703, O70506 | Gene names | Hcn2, Bcng2, Hac1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2) (Hyperpolarization-activated cation channel 1) (HAC-1). | |||||
|
HCN4_HUMAN
|
||||||
| NC score | 0.773961 (rank : 24) | θ value | 3.61944e-33 (rank : 17) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9Y3Q4, Q9UMQ7 | Gene names | HCN4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4. | |||||
|
HCN4_MOUSE
|
||||||
| NC score | 0.772171 (rank : 25) | θ value | 1.9192e-34 (rank : 15) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O70507 | Gene names | Hcn4, Bcng3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4 (Brain cyclic nucleotide-gated channel 3) (BCNG-3). | |||||
|
HCN2_HUMAN
|
||||||
| NC score | 0.767470 (rank : 26) | θ value | 1.0531e-32 (rank : 21) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UL51, O60742, O60743, O75267, Q9UBS2 | Gene names | HCN2, BCNG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2). | |||||
|
CNGB3_MOUSE
|
||||||
| NC score | 0.741179 (rank : 27) | θ value | 9.56915e-18 (rank : 27) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9JJZ9 | Gene names | Cngb3, Cng6 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel beta 3 (CNG channel beta 3) (Cyclic nucleotide-gated channel beta 3) (Cone photoreceptor cGMP- gated channel subunit beta) (Cyclic nucleotide-gated cation channel modulatory subunit) (Cyclic nucleotide-gated channel subunit CNG6). | |||||
|
CNGB3_HUMAN
|
||||||
| NC score | 0.728519 (rank : 28) | θ value | 3.63628e-17 (rank : 28) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9NQW8, Q9NRE9 | Gene names | CNGB3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel beta 3 (CNG channel beta 3) (Cyclic nucleotide-gated channel beta 3) (Cone photoreceptor cGMP- gated channel subunit beta) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
CNGB1_HUMAN
|
||||||
| NC score | 0.724180 (rank : 29) | θ value | 1.16975e-15 (rank : 29) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q14028, Q14029 | Gene names | CNGB1, CNCG4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel 4 (CNG channel 4) (CNG-4) (CNG4) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
KCNKH_HUMAN
|
||||||
| NC score | 0.258199 (rank : 30) | θ value | 0.000158464 (rank : 30) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96T54, Q5TCF4, Q8TAW4, Q9BXD1, Q9H592 | Gene names | KCNK17, TALK2, TASK4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 17 (TWIK-related alkaline pH- activated K(+) channel 2) (2P domain potassium channel Talk-2) (TWIK- related acid-sensitive K(+) channel 4) (TASK-4). | |||||
|
KCNK2_HUMAN
|
||||||
| NC score | 0.188297 (rank : 31) | θ value | 0.00869519 (rank : 31) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O95069, Q9UNE3 | Gene names | KCNK2, TREK, TREK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (TREK-1 K(+) channel subunit) (Two-pore potassium channel TPKC1). | |||||
|
KCNKF_HUMAN
|
||||||
| NC score | 0.188275 (rank : 32) | θ value | 0.279714 (rank : 54) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9H427, Q9HBC8 | Gene names | KCNK15, TASK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 15 (Acid-sensitive potassium channel protein TASK-5) (TWIK-related acid-sensitive K(+) channel 5) (Two pore potassium channel KT3.3). | |||||
|
KCNK2_MOUSE
|
||||||
| NC score | 0.187561 (rank : 33) | θ value | 0.00869519 (rank : 32) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P97438 | Gene names | Kcnk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (Two-pore potassium channel TPKC1) (TREK-1 K(+) channel subunit). | |||||
|
KCNKA_HUMAN
|
||||||
| NC score | 0.181147 (rank : 34) | θ value | 0.21417 (rank : 51) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P57789, Q8TDK7, Q8TDK8, Q9HB59 | Gene names | KCNK10, TREK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 10 (Outward rectifying potassium channel protein TREK-2) (TREK-2 K(+) channel subunit). | |||||
|
KCNK6_HUMAN
|
||||||
| NC score | 0.180145 (rank : 35) | θ value | 0.0961366 (rank : 43) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Y257, Q9HB47 | Gene names | KCNK6, TOSS, TWIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 6 (Inward rectifying potassium channel protein TWIK-2) (TWIK-originated similarity sequence). | |||||
|
KCNK3_HUMAN
|
||||||
| NC score | 0.179237 (rank : 36) | θ value | 0.279714 (rank : 52) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O14649 | Gene names | KCNK3, TASK, TASK1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Two pore potassium channel KT3.1). | |||||
|
KCNK3_MOUSE
|
||||||
| NC score | 0.175758 (rank : 37) | θ value | 0.279714 (rank : 53) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O35111, O35163 | Gene names | Kcnk3, Ctbak, Task, Task1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Cardiac two-pore background K(+) channel) (cTBAK-1) (Two pore potassium channel KT3.1). | |||||
|
KCNK4_HUMAN
|
||||||
| NC score | 0.172892 (rank : 38) | θ value | 0.0563607 (rank : 37) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NYG8, Q96T94 | Gene names | KCNK4, TRAAK | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK) (Two pore K(+) channel KT4.1). | |||||
|
KCNKG_HUMAN
|
||||||
| NC score | 0.164679 (rank : 39) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q96T55, Q5TCF3, Q9H591 | Gene names | KCNK16, TALK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 16 (TWIK-related alkaline pH- activated K(+) channel 1) (2P domain potassium channel Talk-1). | |||||
|
KCNK4_MOUSE
|
||||||
| NC score | 0.160256 (rank : 40) | θ value | 0.125558 (rank : 46) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
KCNK9_MOUSE
|
||||||
| NC score | 0.151517 (rank : 41) | θ value | 0.813845 (rank : 60) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q3LS21 | Gene names | Kcnk9, Task3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3). | |||||
|
KCNK9_HUMAN
|
||||||
| NC score | 0.151495 (rank : 42) | θ value | 0.813845 (rank : 59) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9NPC2, Q2M290, Q540F2 | Gene names | KCNK9, TASK3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3) (Two pore potassium channel KT3.2). | |||||
|
KCNK1_HUMAN
|
||||||
| NC score | 0.133663 (rank : 43) | θ value | 0.0736092 (rank : 41) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O00180, Q13307 | Gene names | KCNK1, HOHO1, KCNO1, TWIK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1) (Potassium channel KCNO1). | |||||
|
KCNK5_HUMAN
|
||||||
| NC score | 0.130061 (rank : 44) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
KCNK1_MOUSE
|
||||||
| NC score | 0.129494 (rank : 45) | θ value | 0.0961366 (rank : 42) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O08581 | Gene names | Kcnk1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1). | |||||
|
KAP0_HUMAN
|
||||||
| NC score | 0.128849 (rank : 46) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P10644, Q567S7 | Gene names | PRKAR1A, PKR1, PRKAR1 | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type I-alpha regulatory subunit (Tissue- specific extinguisher 1) (TSE1). | |||||
|
KAP0_MOUSE
|
||||||
| NC score | 0.117243 (rank : 47) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9DBC7, Q3UKU7, Q9JHR5, Q9JHR6 | Gene names | Prkar1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | cAMP-dependent protein kinase type I-alpha regulatory subunit. | |||||
|
KAP1_HUMAN
|
||||||
| NC score | 0.106325 (rank : 48) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P31321 | Gene names | PRKAR1B | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type I-beta regulatory subunit. | |||||
|
KAP1_MOUSE
|
||||||
| NC score | 0.103875 (rank : 49) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P12849 | Gene names | Prkar1b | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type I-beta regulatory subunit. | |||||
|
KCNKD_MOUSE
|
||||||
| NC score | 0.089190 (rank : 50) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8R1P5 | Gene names | Kcnk13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCMA1_MOUSE
|
||||||
| NC score | 0.082080 (rank : 51) | θ value | 0.0736092 (rank : 39) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCMA1_HUMAN
|
||||||
| NC score | 0.081971 (rank : 52) | θ value | 0.0736092 (rank : 38) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
KCND1_MOUSE
|
||||||
| NC score | 0.081172 (rank : 53) | θ value | 0.125558 (rank : 45) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KAP3_MOUSE
|
||||||
| NC score | 0.078896 (rank : 54) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P31324, Q3UTZ1, Q80ZM4, Q8BRZ7 | Gene names | Prkar2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | cAMP-dependent protein kinase type II-beta regulatory subunit. | |||||
|
KAP2_MOUSE
|
||||||
| NC score | 0.077626 (rank : 55) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P12367 | Gene names | Prkar2a | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type II-alpha regulatory subunit. | |||||
|
KCNK7_MOUSE
|
||||||
| NC score | 0.076437 (rank : 56) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9Z2T1, Q9QXY0, Q9QYE8, Q9R1V1, Q9R242 | Gene names | Kcnk7, Dpkch3, Kcnk6, Kcnk8, Knot1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7 (Putative potassium channel DP3) (Double-pore K(+) channel 3) (Neuromuscular two p domain potassium channel). | |||||
|
KAP3_HUMAN
|
||||||
| NC score | 0.075624 (rank : 57) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P31323 | Gene names | PRKAR2B | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type II-beta regulatory subunit. | |||||
|
KAP2_HUMAN
|
||||||
| NC score | 0.075581 (rank : 58) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P13861, Q16823 | Gene names | PRKAR2A, PKR2, PRKAR2 | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type II-alpha regulatory subunit. | |||||
|
KCND1_HUMAN
|
||||||
| NC score | 0.075400 (rank : 59) | θ value | 0.0736092 (rank : 40) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
KCND2_MOUSE
|
||||||
| NC score | 0.073620 (rank : 60) | θ value | 0.163984 (rank : 48) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_HUMAN
|
||||||
| NC score | 0.073591 (rank : 61) | θ value | 0.163984 (rank : 47) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND3_MOUSE
|
||||||
| NC score | 0.071754 (rank : 62) | θ value | 0.365318 (rank : 57) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND3_HUMAN
|
||||||
| NC score | 0.071750 (rank : 63) | θ value | 0.365318 (rank : 56) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCNKD_HUMAN
|
||||||
| NC score | 0.071252 (rank : 64) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9HB14, Q96E79 | Gene names | KCNK13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNKC_HUMAN
|
||||||
| NC score | 0.068393 (rank : 65) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9HB15 | Gene names | KCNK12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 12 (Tandem pore domain halothane- inhibited potassium channel 2) (THIK-2). | |||||
|
CT152_MOUSE
|
||||||
| NC score | 0.060555 (rank : 66) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9D5U8, Q8R083 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf152 homolog. | |||||
|
KCNK7_HUMAN
|
||||||
| NC score | 0.058607 (rank : 67) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y2U2, Q9Y2U3, Q9Y2U4 | Gene names | KCNK7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7. | |||||
|
KCNS2_HUMAN
|
||||||
| NC score | 0.052243 (rank : 68) | θ value | 0.0193708 (rank : 35) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9ULS6 | Gene names | KCNS2, KIAA1144 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNS2_MOUSE
|
||||||
| NC score | 0.051807 (rank : 69) | θ value | 0.0193708 (rank : 36) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35174, Q543P3 | Gene names | Kcns2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.042796 (rank : 70) | θ value | 0.0113563 (rank : 34) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
KGP1B_HUMAN
|
||||||
| NC score | 0.040161 (rank : 71) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 921 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P14619 | Gene names | PRKG1, PRKG1B, PRKGR1B | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, beta isozyme (EC 2.7.11.12) (cGK 1 beta) (cGKI-beta). | |||||
|
KGP1A_HUMAN
|
||||||
| NC score | 0.040156 (rank : 72) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q13976 | Gene names | PRKG1, PRKGR1A | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, alpha isozyme (EC 2.7.11.12) (CGK 1 alpha) (cGKI-alpha). | |||||
|
KGP1B_MOUSE
|
||||||
| NC score | 0.039755 (rank : 73) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Z0Z0 | Gene names | Prkg1, Prkg1b, Prkgr1b | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, beta isozyme (EC 2.7.11.12) (cGK 1 beta) (cGKI-beta). | |||||
|
KCNB2_HUMAN
|
||||||
| NC score | 0.038897 (rank : 74) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
KCNN4_MOUSE
|
||||||
| NC score | 0.038761 (rank : 75) | θ value | 4.03905 (rank : 79) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O89109, Q3U1J8, Q9CY39 | Gene names | Kcnn4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1). | |||||
|
KCNA4_HUMAN
|
||||||
| NC score | 0.034769 (rank : 76) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.031658 (rank : 77) | θ value | 0.0961366 (rank : 44) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
KCNA4_MOUSE
|
||||||
| NC score | 0.030935 (rank : 78) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61423 | Gene names | Kcna4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4). | |||||
|
KGP2_HUMAN
|
||||||
| NC score | 0.028954 (rank : 79) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1023 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13237, O00125, O60916 | Gene names | PRKG2, PRKGR2 | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 2 (EC 2.7.11.12) (CGK 2) (cGKII) (Type II cGMP-dependent protein kinase). | |||||
|
KCNN2_HUMAN
|
||||||
| NC score | 0.028771 (rank : 80) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9H2S1 | Gene names | KCNN2 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN2_MOUSE
|
||||||
| NC score | 0.028748 (rank : 81) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P58390 | Gene names | Kcnn2, Sk2 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KGP2_MOUSE
|
||||||
| NC score | 0.028199 (rank : 82) | θ value | 5.27518 (rank : 87) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1029 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61410 | Gene names | Prkg2, Prkgr2 | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 2 (EC 2.7.11.12) (CGK 2) (cGKII) (Type II cGMP-dependent protein kinase). | |||||
|
IL7RA_MOUSE
|
||||||
| NC score | 0.027233 (rank : 83) | θ value | 0.21417 (rank : 50) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P16872 | Gene names | Il7r | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-7 receptor alpha chain precursor (IL-7R-alpha) (CD127 antigen). | |||||
|
CKLF_HUMAN
|
||||||
| NC score | 0.026164 (rank : 84) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UBR5, Q9UHM7, Q9UHN8, Q9UI41 | Gene names | CKLF | |||
|
Domain Architecture |

|
|||||
| Description | Chemokine-like factor (C32). | |||||
|
LARP1_HUMAN
|
||||||
| NC score | 0.018263 (rank : 85) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6PKG0, O94836, Q8N4M2, Q8NB73, Q9UFD7 | Gene names | LARP1, KIAA0731, LARP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 1 (La ribonucleoprotein domain family member 1). | |||||
|
IRK13_HUMAN
|
||||||
| NC score | 0.014596 (rank : 86) | θ value | 0.365318 (rank : 55) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60928, O76023, Q8N3Y4 | Gene names | KCNJ13 | |||
|
Domain Architecture |

|
|||||
| Description | Inward rectifier potassium channel 13 (Potassium channel, inwardly rectifying subfamily J member 13) (Inward rectifier K(+) channel Kir7.1). | |||||
|
AGGF1_HUMAN
|
||||||
| NC score | 0.013791 (rank : 87) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N302, O00581, Q53YS3, Q9BU84, Q9NW66 | Gene names | AGGF1, VG5Q | |||
|
Domain Architecture |

|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (Vasculogenesis gene on 5q protein) (hVG5Q). | |||||
|
AF17_HUMAN
|
||||||
| NC score | 0.013402 (rank : 88) | θ value | 0.813845 (rank : 58) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P55198, Q9UF49 | Gene names | MLLT6, AF17 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-17. | |||||
|
ANR21_HUMAN
|
||||||
| NC score | 0.013257 (rank : 89) | θ value | 0.0113563 (rank : 33) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86YR6 | Gene names | ANKRD21, POTE | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat domain-containing protein 21 (Protein POTE) (Prostate, ovary, testis-expressed protein). | |||||
|
PROM1_MOUSE
|
||||||
| NC score | 0.012987 (rank : 90) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54990, O35408 | Gene names | Prom1, Prom, Proml1 | |||
|
Domain Architecture |

|
|||||
| Description | Prominin-1 precursor (Prominin-like protein 1) (Antigen AC133 homolog). | |||||
|
GP108_MOUSE
|
||||||
| NC score | 0.011742 (rank : 91) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91WD0, Q8BXE9, Q925B1 | Gene names | Gpr108, Lustr2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein GPR108 precursor (Lung seven transmembrane receptor 2). | |||||
|
HD_MOUSE
|
||||||
| NC score | 0.011008 (rank : 92) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42859 | Gene names | Hd, Hdh | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein homolog) (HD protein). | |||||
|
POT15_HUMAN
|
||||||
| NC score | 0.010994 (rank : 93) | θ value | 0.163984 (rank : 49) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6S5H4, Q6NXN7, Q6S5H7 | Gene names | POTE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 15. | |||||
|
PROM1_HUMAN
|
||||||
| NC score | 0.010856 (rank : 94) | θ value | 8.99809 (rank : 103) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43490 | Gene names | PROM1, PROML1 | |||
|
Domain Architecture |

|
|||||
| Description | Prominin-1 precursor (Prominin-like protein 1) (Antigen AC133) (CD133 antigen). | |||||
|
TINF2_HUMAN
|
||||||
| NC score | 0.010810 (rank : 95) | θ value | 5.27518 (rank : 90) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BSI4, Q9H904, Q9UHC2 | Gene names | TINF2, TIN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TERF1-interacting nuclear factor 2 (TRF1-interacting nuclear protein 2). | |||||
|
TP53B_MOUSE
|
||||||
| NC score | 0.010654 (rank : 96) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
NAB2_MOUSE
|
||||||
| NC score | 0.010081 (rank : 97) | θ value | 3.0926 (rank : 75) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61127 | Gene names | Nab2 | |||
|
Domain Architecture |

|
|||||
| Description | NGFI-A-binding protein 2 (EGR-1-binding protein 2). | |||||
|
AP4E1_MOUSE
|
||||||
| NC score | 0.009701 (rank : 98) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80V94 | Gene names | Ap4e1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-4 complex subunit epsilon-1 (Adapter-related protein complex 4 epsilon-1 subunit) (Epsilon subunit of AP-4) (AP-4 adapter complex epsilon subunit). | |||||
|
T2FA_HUMAN
|
||||||
| NC score | 0.009622 (rank : 99) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
LRCH2_HUMAN
|
||||||
| NC score | 0.008797 (rank : 100) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5VUJ6, Q9HA88, Q9P233 | Gene names | LRCH2, KIAA1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
LRCH2_MOUSE
|
||||||
| NC score | 0.008148 (rank : 101) | θ value | 1.06291 (rank : 61) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UMG5, Q7TNP9, Q80TC9 | Gene names | Lrch2, Kiaa1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
FA35A_MOUSE
|
||||||
| NC score | 0.007902 (rank : 102) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3UEN2 | Gene names | Fam35a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
RFIP1_MOUSE
|
||||||
| NC score | 0.007494 (rank : 103) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D620, Q3UBC2, Q8BN24 | Gene names | Rab11fip1, Rcp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
BCL7A_MOUSE
|
||||||
| NC score | 0.006859 (rank : 104) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CXE2, Q8C361, Q8C8M8, Q8VD15 | Gene names | Bcl7a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell CLL/lymphoma 7 protein family member A. | |||||
|
ZHX3_HUMAN
|
||||||
| NC score | 0.006807 (rank : 105) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H4I2, O43145 | Gene names | ZHX3, KIAA0395, TIX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and homeodomain protein 3 (Triple homeobox protein 1). | |||||
|
TRI73_HUMAN
|
||||||
| NC score | 0.005835 (rank : 106) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86UV7, Q8N0S3 | Gene names | TRIM73, TRIM50B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 73 (Tripartite motif-containing protein 50B). | |||||
|
TRI74_HUMAN
|
||||||
| NC score | 0.004809 (rank : 107) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86UV6 | Gene names | TRIM74, TRIM50C | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 74 (Tripartite motif-containing protein 50C). | |||||
|
NDE1_HUMAN
|
||||||
| NC score | 0.004398 (rank : 108) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 823 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NXR1, Q49AQ2 | Gene names | NDE1, NUDE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear distribution protein nudE homolog 1 (NudE). | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.003582 (rank : 109) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
RPGR_MOUSE
|
||||||
| NC score | 0.003169 (rank : 110) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
TRI50_MOUSE
|
||||||
| NC score | 0.002536 (rank : 111) | θ value | 5.27518 (rank : 91) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q810I2 | Gene names | Trim50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 50. | |||||
|
CEP35_HUMAN
|
||||||
| NC score | 0.002024 (rank : 112) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
RAI14_HUMAN
|
||||||
| NC score | 0.000816 (rank : 113) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1367 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9P0K7, Q6V1W9, Q7Z5I4, Q7Z733, Q9P2L2, Q9Y3T5 | Gene names | RAI14, KIAA1334, NORPEG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein). | |||||
|
APOA_HUMAN
|
||||||
| NC score | -0.000148 (rank : 114) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P08519 | Gene names | LPA | |||
|
Domain Architecture |

|
|||||
| Description | Apolipoprotein(a) precursor (EC 3.4.21.-) (Apo(a)) (Lp(a)). | |||||
|
MTR1L_HUMAN
|
||||||
| NC score | -0.001526 (rank : 115) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1180 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13585 | Gene names | GPR50 | |||
|
Domain Architecture |

|
|||||
| Description | Melatonin-related receptor (G protein-coupled receptor 50) (H9). | |||||
|
BRSK2_HUMAN
|
||||||
| NC score | -0.002672 (rank : 116) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 897 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IWQ3, O60843, O95099, Q5J5B4, Q8TB60 | Gene names | BRSK2, C11orf7, PEN11B, STK29 | |||
|
Domain Architecture |

|
|||||
| Description | BR serine/threonine-protein kinase 2 (EC 2.7.11.1) (Serine/threonine- protein kinase 29) (SAD1B). | |||||
|
MTR1L_MOUSE
|
||||||
| NC score | -0.003327 (rank : 117) | θ value | 8.99809 (rank : 102) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88495 | Gene names | Gpr50 | |||
|
Domain Architecture |

|
|||||
| Description | Melatonin-related receptor (G protein-coupled receptor 50) (H9). | |||||
|
OR2V2_HUMAN
|
||||||
| NC score | -0.004130 (rank : 118) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96R30, Q6IFL6, Q8NGV1 | Gene names | OR2V2, OR2V3 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 2V2 (Olfactory receptor OR5-3). | |||||