Please be patient as the page loads
|
KCNH5_HUMAN
|
||||||
| SwissProt Accessions | Q8NCM2 | Gene names | KCNH5, EAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (hEAG2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
KCNH1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.996375 (rank : 4) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O95259, O76035 | Gene names | KCNH1, EAG, EAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (hEAG1) (h-eag). | |||||
|
KCNH1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.996566 (rank : 3) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q60603 | Gene names | Kcnh1, Eag | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (EAG1) (m-eag). | |||||
|
KCNH5_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 99 | |
| SwissProt Accessions | Q8NCM2 | Gene names | KCNH5, EAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (hEAG2). | |||||
|
KCNH5_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.998931 (rank : 2) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q920E3, Q8C035 | Gene names | Kcnh5, Eag2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (Eag2). | |||||
|
KCNH8_HUMAN
|
||||||
| θ value | 2.19974e-139 (rank : 5) | NC score | 0.968428 (rank : 5) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q96L42 | Gene names | KCNH8 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 8 (Voltage-gated potassium channel subunit Kv12.1) (Ether-a-go-go-like potassium channel 3) (ELK channel 3) (ELK3) (ELK1) (hElk1). | |||||
|
KCNH6_HUMAN
|
||||||
| θ value | 1.74296e-136 (rank : 6) | NC score | 0.960872 (rank : 9) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9H252, Q9BRD7 | Gene names | KCNH6, ERG2, HERG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 6 (Voltage-gated potassium channel subunit Kv11.2) (Ether-a-go-go-related gene potassium channel 2) (Ether-a-go-go-related protein 2) (Eag-related protein 2). | |||||
|
KCNH3_HUMAN
|
||||||
| θ value | 4.01817e-133 (rank : 7) | NC score | 0.961603 (rank : 7) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9ULD8, Q9UQ06 | Gene names | KCNH3, KIAA1282 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (ELK2) (Brain-specific eag-like channel 1) (BEC1). | |||||
|
KCNH3_MOUSE
|
||||||
| θ value | 8.9516e-133 (rank : 8) | NC score | 0.961425 (rank : 8) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9WVJ0 | Gene names | Kcnh3, Elk2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (mElk2). | |||||
|
KCNH4_HUMAN
|
||||||
| θ value | 3.1838e-130 (rank : 9) | NC score | 0.964158 (rank : 6) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9UQ05 | Gene names | KCNH4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 4 (Voltage-gated potassium channel subunit Kv12.3) (Ether-a-go-go-like potassium channel 1) (ELK channel 1) (ELK1) (Brain-specific eag-like channel 2) (BEC2). | |||||
|
KCNH2_HUMAN
|
||||||
| θ value | 3.19119e-122 (rank : 10) | NC score | 0.954467 (rank : 10) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q12809, O75418, O75680, Q9BT72, Q9BUT7, Q9H3P0 | Gene names | KCNH2, ERG, ERG1, HERG, HERG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (H-ERG) (Erg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1) (eag homolog). | |||||
|
KCNH2_MOUSE
|
||||||
| θ value | 3.19119e-122 (rank : 11) | NC score | 0.954007 (rank : 11) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | O35219, O35220, O35221, O35989 | Gene names | Kcnh2, Erg, Merg1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (ERG1) (MERG) (Merg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1). | |||||
|
KCNH7_MOUSE
|
||||||
| θ value | 8.99321e-117 (rank : 12) | NC score | 0.953148 (rank : 12) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9ER47 | Gene names | Kcnh7, Erg3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (Ether-a-go-go-related protein 3) (Eag-related protein 3). | |||||
|
KCNH7_HUMAN
|
||||||
| θ value | 1.21547e-113 (rank : 13) | NC score | 0.951974 (rank : 13) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9NS40 | Gene names | KCNH7, ERG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (HERG-3) (Ether-a-go-go-related protein 3) (Eag- related protein 3). | |||||
|
HCN1_MOUSE
|
||||||
| θ value | 2.19075e-38 (rank : 14) | NC score | 0.795328 (rank : 19) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O88704, O54899, Q9D613 | Gene names | Hcn1, Bcng1, Hac2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1) (Hyperpolarization-activated cation channel 2) (HAC-2). | |||||
|
HCN1_HUMAN
|
||||||
| θ value | 4.88048e-38 (rank : 15) | NC score | 0.798895 (rank : 16) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O60741 | Gene names | HCN1, BCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1). | |||||
|
HCN2_MOUSE
|
||||||
| θ value | 1.57e-36 (rank : 16) | NC score | 0.780277 (rank : 23) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O88703, O70506 | Gene names | Hcn2, Bcng2, Hac1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2) (Hyperpolarization-activated cation channel 1) (HAC-1). | |||||
|
HCN4_HUMAN
|
||||||
| θ value | 4.568e-36 (rank : 17) | NC score | 0.777496 (rank : 24) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Y3Q4, Q9UMQ7 | Gene names | HCN4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4. | |||||
|
HCN4_MOUSE
|
||||||
| θ value | 1.01765e-35 (rank : 18) | NC score | 0.775728 (rank : 25) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O70507 | Gene names | Hcn4, Bcng3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4 (Brain cyclic nucleotide-gated channel 3) (BCNG-3). | |||||
|
HCN2_HUMAN
|
||||||
| θ value | 1.32909e-35 (rank : 19) | NC score | 0.771816 (rank : 26) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UL51, O60742, O60743, O75267, Q9UBS2 | Gene names | HCN2, BCNG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2). | |||||
|
CNGA2_MOUSE
|
||||||
| θ value | 2.96089e-35 (rank : 20) | NC score | 0.812577 (rank : 14) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q62398 | Gene names | Cnga2, Cncg2, Cncg4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated olfactory channel (Cyclic nucleotide-gated cation channel 2) (CNG channel 2) (CNG-2) (CNG2). | |||||
|
HCN3_HUMAN
|
||||||
| θ value | 5.584e-34 (rank : 21) | NC score | 0.792875 (rank : 22) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9P1Z3, Q4VX12, Q8N6W6, Q9BWQ2 | Gene names | HCN3, KIAA1535 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3. | |||||
|
HCN3_MOUSE
|
||||||
| θ value | 1.24399e-33 (rank : 22) | NC score | 0.792903 (rank : 20) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88705 | Gene names | Hcn3, Hac3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3 (Hyperpolarization-activated cation channel 3) (HAC-3). | |||||
|
CNGA1_HUMAN
|
||||||
| θ value | 4.00176e-32 (rank : 23) | NC score | 0.800556 (rank : 15) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P29973, Q16279, Q16485 | Gene names | CNGA1, CNCG, CNCG1 | |||
|
Domain Architecture |
|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
CNGA1_MOUSE
|
||||||
| θ value | 2.59387e-31 (rank : 24) | NC score | 0.796239 (rank : 18) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P29974, Q60776 | Gene names | Cnga1, Cncg, Cncg1 | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
CNGA3_MOUSE
|
||||||
| θ value | 2.42779e-29 (rank : 25) | NC score | 0.798088 (rank : 17) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9JJZ8, Q9WV01 | Gene names | Cnga3, Cng3 | |||
|
Domain Architecture |
|
|||||
| Description | Cyclic nucleotide-gated cation channel alpha 3 (CNG channel alpha 3) (CNG-3) (CNG3) (Cyclic nucleotide-gated channel alpha 3) (Cone photoreceptor cGMP-gated channel subunit alpha). | |||||
|
CNGA3_HUMAN
|
||||||
| θ value | 3.28125e-26 (rank : 26) | NC score | 0.792898 (rank : 21) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q16281, Q9UP64 | Gene names | CNGA3, CNCG3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel alpha 3 (CNG channel alpha 3) (CNG-3) (CNG3) (Cyclic nucleotide-gated channel alpha 3) (Cone photoreceptor cGMP-gated channel subunit alpha). | |||||
|
CNGB3_HUMAN
|
||||||
| θ value | 2.20094e-22 (rank : 27) | NC score | 0.735686 (rank : 28) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9NQW8, Q9NRE9 | Gene names | CNGB3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel beta 3 (CNG channel beta 3) (Cyclic nucleotide-gated channel beta 3) (Cone photoreceptor cGMP- gated channel subunit beta) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
CNGB3_MOUSE
|
||||||
| θ value | 3.51386e-20 (rank : 28) | NC score | 0.745273 (rank : 27) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9JJZ9 | Gene names | Cngb3, Cng6 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel beta 3 (CNG channel beta 3) (Cyclic nucleotide-gated channel beta 3) (Cone photoreceptor cGMP- gated channel subunit beta) (Cyclic nucleotide-gated cation channel modulatory subunit) (Cyclic nucleotide-gated channel subunit CNG6). | |||||
|
CNGB1_HUMAN
|
||||||
| θ value | 6.62687e-19 (rank : 29) | NC score | 0.725681 (rank : 29) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14028, Q14029 | Gene names | CNGB1, CNCG4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel 4 (CNG channel 4) (CNG-4) (CNG4) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
KCNKH_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 30) | NC score | 0.288939 (rank : 30) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q96T54, Q5TCF4, Q8TAW4, Q9BXD1, Q9H592 | Gene names | KCNK17, TALK2, TASK4 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 17 (TWIK-related alkaline pH- activated K(+) channel 2) (2P domain potassium channel Talk-2) (TWIK- related acid-sensitive K(+) channel 4) (TASK-4). | |||||
|
KCNK2_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 31) | NC score | 0.225128 (rank : 31) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O95069, Q9UNE3 | Gene names | KCNK2, TREK, TREK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (TREK-1 K(+) channel subunit) (Two-pore potassium channel TPKC1). | |||||
|
KCNK2_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 32) | NC score | 0.224145 (rank : 32) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P97438 | Gene names | Kcnk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (Two-pore potassium channel TPKC1) (TREK-1 K(+) channel subunit). | |||||
|
KCNK3_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 33) | NC score | 0.219668 (rank : 34) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O14649 | Gene names | KCNK3, TASK, TASK1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Two pore potassium channel KT3.1). | |||||
|
KCNK3_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 34) | NC score | 0.215138 (rank : 36) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O35111, O35163 | Gene names | Kcnk3, Ctbak, Task, Task1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Cardiac two-pore background K(+) channel) (cTBAK-1) (Two pore potassium channel KT3.1). | |||||
|
KCNK4_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 35) | NC score | 0.206307 (rank : 39) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9NYG8, Q96T94 | Gene names | KCNK4, TRAAK | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK) (Two pore K(+) channel KT4.1). | |||||
|
KCNK6_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 36) | NC score | 0.214788 (rank : 37) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Y257, Q9HB47 | Gene names | KCNK6, TOSS, TWIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 6 (Inward rectifying potassium channel protein TWIK-2) (TWIK-originated similarity sequence). | |||||
|
KCNKA_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 37) | NC score | 0.217676 (rank : 35) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P57789, Q8TDK7, Q8TDK8, Q9HB59 | Gene names | KCNK10, TREK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 10 (Outward rectifying potassium channel protein TREK-2) (TREK-2 K(+) channel subunit). | |||||
|
KCNKG_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 38) | NC score | 0.209863 (rank : 38) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q96T55, Q5TCF3, Q9H591 | Gene names | KCNK16, TALK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 16 (TWIK-related alkaline pH- activated K(+) channel 1) (2P domain potassium channel Talk-1). | |||||
|
KCNK9_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 39) | NC score | 0.191570 (rank : 42) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9NPC2, Q2M290, Q540F2 | Gene names | KCNK9, TASK3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3) (Two pore potassium channel KT3.2). | |||||
|
KCNK9_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 40) | NC score | 0.191984 (rank : 41) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q3LS21 | Gene names | Kcnk9, Task3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3). | |||||
|
KCND3_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 41) | NC score | 0.080990 (rank : 57) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND3_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 42) | NC score | 0.080981 (rank : 58) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCNK4_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 43) | NC score | 0.193867 (rank : 40) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
KCNKF_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 44) | NC score | 0.224081 (rank : 33) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9H427, Q9HBC8 | Gene names | KCNK15, TASK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 15 (Acid-sensitive potassium channel protein TASK-5) (TWIK-related acid-sensitive K(+) channel 5) (Two pore potassium channel KT3.3). | |||||
|
KCND1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 45) | NC score | 0.087230 (rank : 54) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KCND1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 46) | NC score | 0.082387 (rank : 56) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
KCND2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 47) | NC score | 0.080908 (rank : 59) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_MOUSE
|
||||||
| θ value | 0.125558 (rank : 48) | NC score | 0.080907 (rank : 60) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.21417 (rank : 49) | NC score | 0.017233 (rank : 90) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
KCNK1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 50) | NC score | 0.164758 (rank : 43) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O00180, Q13307 | Gene names | KCNK1, HOHO1, KCNO1, TWIK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1) (Potassium channel KCNO1). | |||||
|
KCNK5_HUMAN
|
||||||
| θ value | 0.279714 (rank : 51) | NC score | 0.163634 (rank : 44) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
KCNK1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 52) | NC score | 0.161118 (rank : 45) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08581 | Gene names | Kcnk1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1). | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 53) | NC score | 0.005766 (rank : 101) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
KCNKD_MOUSE
|
||||||
| θ value | 0.47712 (rank : 54) | NC score | 0.128247 (rank : 47) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8R1P5 | Gene names | Kcnk13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
PHC1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 55) | NC score | 0.014639 (rank : 92) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P78364, Q8WVM3, Q9BU63 | Gene names | PHC1, EDR1, PH1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (hPH1) (Early development regulatory protein 1). | |||||
|
KGP1B_MOUSE
|
||||||
| θ value | 0.62314 (rank : 56) | NC score | 0.039341 (rank : 79) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Z0Z0 | Gene names | Prkg1, Prkg1b, Prkgr1b | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, beta isozyme (EC 2.7.11.12) (cGK 1 beta) (cGKI-beta). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 0.62314 (rank : 57) | NC score | 0.006417 (rank : 99) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
KCNG4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 58) | NC score | 0.044228 (rank : 73) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8TDN1, Q96H24 | Gene names | KCNG4, KCNG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 4 (Voltage-gated potassium channel subunit Kv6.4). | |||||
|
DTX3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.020546 (rank : 88) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80V91, Q9ER06 | Gene names | Dtx3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein deltex-3 (Deltex-3) (Deltex3) (mDTX3). | |||||
|
KGP1A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.039166 (rank : 81) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13976 | Gene names | PRKG1, PRKGR1A | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, alpha isozyme (EC 2.7.11.12) (CGK 1 alpha) (cGKI-alpha). | |||||
|
KGP1B_HUMAN
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.039195 (rank : 80) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 921 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P14619 | Gene names | PRKG1, PRKG1B, PRKGR1B | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, beta isozyme (EC 2.7.11.12) (cGK 1 beta) (cGKI-beta). | |||||
|
RAI14_MOUSE
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | -0.000050 (rank : 112) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1505 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9EP71, Q3URT3, Q6ZPT6 | Gene names | Rai14, Kiaa1334, Norpeg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein) (p125). | |||||
|
DTX3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.019476 (rank : 89) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N9I9, Q8NAU6, Q8NDS8 | Gene names | DTX3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein deltex-3 (Deltex-3) (Deltex3). | |||||
|
KCNQ4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.049993 (rank : 72) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P56696, O96025 | Gene names | KCNQ4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 4 (Voltage-gated potassium channel subunit Kv7.4) (Potassium channel subunit alpha KvLQT4) (KQT-like 4). | |||||
|
KCNQ5_HUMAN
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.037156 (rank : 84) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NR82, Q9NRN0, Q9NYA6 | Gene names | KCNQ5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 5 (Voltage-gated potassium channel subunit Kv7.5) (Potassium channel subunit alpha KvLQT5) (KQT-like 5). | |||||
|
KCNK7_MOUSE
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.109768 (rank : 49) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9Z2T1, Q9QXY0, Q9QYE8, Q9R1V1, Q9R242 | Gene names | Kcnk7, Dpkch3, Kcnk6, Kcnk8, Knot1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7 (Putative potassium channel DP3) (Double-pore K(+) channel 3) (Neuromuscular two p domain potassium channel). | |||||
|
KCNKD_HUMAN
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.109156 (rank : 50) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HB14, Q96E79 | Gene names | KCNK13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNS1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.036465 (rank : 86) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96KK3, O43652, Q6DJU6 | Gene names | KCNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
FMN1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.011210 (rank : 94) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q05859 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoform IV (Limb deformity protein). | |||||
|
ARMX5_MOUSE
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.014736 (rank : 91) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UZB0 | Gene names | Armcx5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 5. | |||||
|
DMP1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.010526 (rank : 95) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
KCNA2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.038574 (rank : 82) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P63141, P15386, Q02010, Q8C8W4 | Gene names | Kcna2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (MK2). | |||||
|
KCNB1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.042453 (rank : 76) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q14721, Q14193 | Gene names | KCNB1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (h-DRK1). | |||||
|
KCNB1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.042471 (rank : 75) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q03717 | Gene names | Kcnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (mShab). | |||||
|
KCNB2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.043655 (rank : 74) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
KCNS2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.051801 (rank : 70) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9ULS6 | Gene names | KCNS2, KIAA1144 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.001618 (rank : 109) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
VINEX_MOUSE
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.003450 (rank : 104) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9R1Z8, Q62423 | Gene names | Sorbs3, Scam1, Sh3d4 | |||
|
Domain Architecture |

|
|||||
| Description | Vinexin (Sorbin and SH3 domain-containing protein 3) (SH3-containing adapter molecule 1) (SCAM-1) (SH3 domain-containing protein SH3P3). | |||||
|
CAYP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.009115 (rank : 97) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BUG5, Q8C1J6 | Gene names | Caps2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcyphosin-2 (Calcyphosine-2). | |||||
|
GON4L_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.010470 (rank : 96) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
KCNS2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.051196 (rank : 71) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O35174, Q543P3 | Gene names | Kcns2 | |||
|
Domain Architecture |
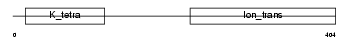
|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNT1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.040324 (rank : 78) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5JUK3, Q9P2C5 | Gene names | KCNT1, KIAA1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1 (KCa4.1). | |||||
|
MYH13_HUMAN
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | -0.000339 (rank : 113) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
RPGF2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.013812 (rank : 93) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y4G8 | Gene names | RAPGEF2, KIAA0313, NRAPGEP, PDZGEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rap guanine nucleotide exchange factor 2 (Neural RAP guanine nucleotide exchange protein) (nRap GEP) (PDZ domain-containing guanine nucleotide exchange factor 1) (PDZ-GEF1) (RA-GEF). | |||||
|
TSH2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.007414 (rank : 98) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q68FE9, Q5DTF1, Q6AXH5, Q9JL71 | Gene names | Tshz2, Kiaa4248, Sdccag33l, Tsh2, Zfp218, Znf218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 2 (Zinc finger protein 218) (SDCCAG33-like protein). | |||||
|
CORO6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.003159 (rank : 105) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q920M5, Q920M3, Q920M4 | Gene names | Coro6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coronin-6 (Clipin E). | |||||
|
K1H5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.001943 (rank : 106) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92764, O76012, Q92651 | Gene names | KRTHA5, HHA5, HKA5 | |||
|
Domain Architecture |
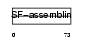
|
|||||
| Description | Keratin, type I cuticular Ha5 (Hair keratin, type I Ha5). | |||||
|
KCNG2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.041688 (rank : 77) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UJ96 | Gene names | KCNG2, KCNF2 | |||
|
Domain Architecture |
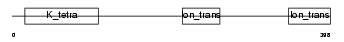
|
|||||
| Description | Potassium voltage-gated channel subfamily G member 2 (Voltage-gated potassium channel subunit Kv6.2) (Cardiac potassium channel subunit). | |||||
|
KCNKC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.104002 (rank : 52) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HB15 | Gene names | KCNK12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 12 (Tandem pore domain halothane- inhibited potassium channel 2) (THIK-2). | |||||
|
KCNT1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.035147 (rank : 87) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6ZPR4, Q8C3E7 | Gene names | Kcnt1, Kiaa1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1. | |||||
|
KIF22_HUMAN
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.001176 (rank : 110) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14807, O94814, Q9BT46 | Gene names | KIF22, KID, KNSL4 | |||
|
Domain Architecture |
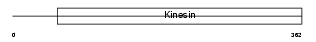
|
|||||
| Description | Kinesin-like protein KIF22 (Kinesin-like DNA-binding protein) (Kinesin-like protein 4). | |||||
|
NEST_HUMAN
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.001648 (rank : 108) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
SEPT8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.001687 (rank : 107) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CHH9, Q80YC7, Q9ESF7 | Gene names | Sept8, Kiaa0202 | |||
|
Domain Architecture |
|
|||||
| Description | Septin-8. | |||||
|
CCP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.004617 (rank : 102) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHG6, Q91WQ4, Q9JK50, Q9JK51, Q9JKK2, Q9JKK3 | Gene names | Dscr1 | |||
|
Domain Architecture |
|
|||||
| Description | Calcipressin-1 (Down syndrome critical region protein 1 homolog) (Myocyte-enriched calcineurin-interacting protein 1) (MCIP1). | |||||
|
CD276_HUMAN
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.000668 (rank : 111) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5ZPR3, Q6P5Y4, Q6UXI2, Q8NBI8, Q8NC34, Q8NCB6, Q9BXR1 | Gene names | CD276, B7H3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CD276 antigen precursor (Costimulatory molecule) (B7 homolog 3) (B7- H3) (4Ig-B7-H3). | |||||
|
HPS4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.006333 (rank : 100) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99KG7 | Gene names | Hps4, Le | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 4 protein homolog (Light-ear protein) (Le protein). | |||||
|
KCNA2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.037051 (rank : 85) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P16389 | Gene names | KCNA2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (HBK5) (NGK1) (HUKIV). | |||||
|
KCNA4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.038270 (rank : 83) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
SMCE1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.004090 (rank : 103) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q969G3, O43539 | Gene names | SMARCE1, BAF57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
CT152_MOUSE
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.060447 (rank : 65) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9D5U8, Q8R083 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | Uncharacterized protein C20orf152 homolog. | |||||
|
KAP0_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.130090 (rank : 46) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P10644, Q567S7 | Gene names | PRKAR1A, PKR1, PRKAR1 | |||
|
Domain Architecture |
|
|||||
| Description | cAMP-dependent protein kinase type I-alpha regulatory subunit (Tissue- specific extinguisher 1) (TSE1). | |||||
|
KAP0_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.117989 (rank : 48) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9DBC7, Q3UKU7, Q9JHR5, Q9JHR6 | Gene names | Prkar1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | cAMP-dependent protein kinase type I-alpha regulatory subunit. | |||||
|
KAP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.104611 (rank : 51) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P31321 | Gene names | PRKAR1B | |||
|
Domain Architecture |
|
|||||
| Description | cAMP-dependent protein kinase type I-beta regulatory subunit. | |||||
|
KAP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.102273 (rank : 53) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P12849 | Gene names | Prkar1b | |||
|
Domain Architecture |
|
|||||
| Description | cAMP-dependent protein kinase type I-beta regulatory subunit. | |||||
|
KAP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.075706 (rank : 63) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P13861, Q16823 | Gene names | PRKAR2A, PKR2, PRKAR2 | |||
|
Domain Architecture |
|
|||||
| Description | cAMP-dependent protein kinase type II-alpha regulatory subunit. | |||||
|
KAP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.077896 (rank : 61) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P12367 | Gene names | Prkar2a | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type II-alpha regulatory subunit. | |||||
|
KAP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.074483 (rank : 64) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P31323 | Gene names | PRKAR2B | |||
|
Domain Architecture |
|
|||||
| Description | cAMP-dependent protein kinase type II-beta regulatory subunit. | |||||
|
KAP3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.077748 (rank : 62) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P31324, Q3UTZ1, Q80ZM4, Q8BRZ7 | Gene names | Prkar2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | cAMP-dependent protein kinase type II-beta regulatory subunit. | |||||
|
KCMA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.058338 (rank : 67) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
KCMA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.058476 (rank : 66) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCNK7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.086340 (rank : 55) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y2U2, Q9Y2U3, Q9Y2U4 | Gene names | KCNK7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7. | |||||
|
KCNQ3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.053020 (rank : 69) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O43525 | Gene names | KCNQ3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
KCNQ3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.054325 (rank : 68) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8K3F6 | Gene names | Kcnq3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
KCNH5_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 99 | |
| SwissProt Accessions | Q8NCM2 | Gene names | KCNH5, EAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (hEAG2). | |||||
|
KCNH5_MOUSE
|
||||||
| NC score | 0.998931 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q920E3, Q8C035 | Gene names | Kcnh5, Eag2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (Eag2). | |||||
|
KCNH1_MOUSE
|
||||||
| NC score | 0.996566 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q60603 | Gene names | Kcnh1, Eag | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (EAG1) (m-eag). | |||||
|
KCNH1_HUMAN
|
||||||
| NC score | 0.996375 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O95259, O76035 | Gene names | KCNH1, EAG, EAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (hEAG1) (h-eag). | |||||
|
KCNH8_HUMAN
|
||||||
| NC score | 0.968428 (rank : 5) | θ value | 2.19974e-139 (rank : 5) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q96L42 | Gene names | KCNH8 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 8 (Voltage-gated potassium channel subunit Kv12.1) (Ether-a-go-go-like potassium channel 3) (ELK channel 3) (ELK3) (ELK1) (hElk1). | |||||
|
KCNH4_HUMAN
|
||||||
| NC score | 0.964158 (rank : 6) | θ value | 3.1838e-130 (rank : 9) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9UQ05 | Gene names | KCNH4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 4 (Voltage-gated potassium channel subunit Kv12.3) (Ether-a-go-go-like potassium channel 1) (ELK channel 1) (ELK1) (Brain-specific eag-like channel 2) (BEC2). | |||||
|
KCNH3_HUMAN
|
||||||
| NC score | 0.961603 (rank : 7) | θ value | 4.01817e-133 (rank : 7) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9ULD8, Q9UQ06 | Gene names | KCNH3, KIAA1282 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (ELK2) (Brain-specific eag-like channel 1) (BEC1). | |||||
|
KCNH3_MOUSE
|
||||||
| NC score | 0.961425 (rank : 8) | θ value | 8.9516e-133 (rank : 8) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9WVJ0 | Gene names | Kcnh3, Elk2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (mElk2). | |||||
|
KCNH6_HUMAN
|
||||||
| NC score | 0.960872 (rank : 9) | θ value | 1.74296e-136 (rank : 6) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9H252, Q9BRD7 | Gene names | KCNH6, ERG2, HERG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 6 (Voltage-gated potassium channel subunit Kv11.2) (Ether-a-go-go-related gene potassium channel 2) (Ether-a-go-go-related protein 2) (Eag-related protein 2). | |||||
|
KCNH2_HUMAN
|
||||||
| NC score | 0.954467 (rank : 10) | θ value | 3.19119e-122 (rank : 10) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q12809, O75418, O75680, Q9BT72, Q9BUT7, Q9H3P0 | Gene names | KCNH2, ERG, ERG1, HERG, HERG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (H-ERG) (Erg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1) (eag homolog). | |||||
|
KCNH2_MOUSE
|
||||||
| NC score | 0.954007 (rank : 11) | θ value | 3.19119e-122 (rank : 11) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | O35219, O35220, O35221, O35989 | Gene names | Kcnh2, Erg, Merg1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (ERG1) (MERG) (Merg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1). | |||||
|
KCNH7_MOUSE
|
||||||
| NC score | 0.953148 (rank : 12) | θ value | 8.99321e-117 (rank : 12) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9ER47 | Gene names | Kcnh7, Erg3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (Ether-a-go-go-related protein 3) (Eag-related protein 3). | |||||
|
KCNH7_HUMAN
|
||||||
| NC score | 0.951974 (rank : 13) | θ value | 1.21547e-113 (rank : 13) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9NS40 | Gene names | KCNH7, ERG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (HERG-3) (Ether-a-go-go-related protein 3) (Eag- related protein 3). | |||||
|
CNGA2_MOUSE
|
||||||
| NC score | 0.812577 (rank : 14) | θ value | 2.96089e-35 (rank : 20) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q62398 | Gene names | Cnga2, Cncg2, Cncg4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated olfactory channel (Cyclic nucleotide-gated cation channel 2) (CNG channel 2) (CNG-2) (CNG2). | |||||
|
CNGA1_HUMAN
|
||||||
| NC score | 0.800556 (rank : 15) | θ value | 4.00176e-32 (rank : 23) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P29973, Q16279, Q16485 | Gene names | CNGA1, CNCG, CNCG1 | |||
|
Domain Architecture |
|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
HCN1_HUMAN
|
||||||
| NC score | 0.798895 (rank : 16) | θ value | 4.88048e-38 (rank : 15) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O60741 | Gene names | HCN1, BCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1). | |||||
|
CNGA3_MOUSE
|
||||||
| NC score | 0.798088 (rank : 17) | θ value | 2.42779e-29 (rank : 25) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9JJZ8, Q9WV01 | Gene names | Cnga3, Cng3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel alpha 3 (CNG channel alpha 3) (CNG-3) (CNG3) (Cyclic nucleotide-gated channel alpha 3) (Cone photoreceptor cGMP-gated channel subunit alpha). | |||||
|
CNGA1_MOUSE
|
||||||
| NC score | 0.796239 (rank : 18) | θ value | 2.59387e-31 (rank : 24) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P29974, Q60776 | Gene names | Cnga1, Cncg, Cncg1 | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
HCN1_MOUSE
|
||||||
| NC score | 0.795328 (rank : 19) | θ value | 2.19075e-38 (rank : 14) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O88704, O54899, Q9D613 | Gene names | Hcn1, Bcng1, Hac2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1) (Hyperpolarization-activated cation channel 2) (HAC-2). | |||||
|
HCN3_MOUSE
|
||||||
| NC score | 0.792903 (rank : 20) | θ value | 1.24399e-33 (rank : 22) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88705 | Gene names | Hcn3, Hac3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3 (Hyperpolarization-activated cation channel 3) (HAC-3). | |||||
|
CNGA3_HUMAN
|
||||||
| NC score | 0.792898 (rank : 21) | θ value | 3.28125e-26 (rank : 26) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q16281, Q9UP64 | Gene names | CNGA3, CNCG3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel alpha 3 (CNG channel alpha 3) (CNG-3) (CNG3) (Cyclic nucleotide-gated channel alpha 3) (Cone photoreceptor cGMP-gated channel subunit alpha). | |||||
|
HCN3_HUMAN
|
||||||
| NC score | 0.792875 (rank : 22) | θ value | 5.584e-34 (rank : 21) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9P1Z3, Q4VX12, Q8N6W6, Q9BWQ2 | Gene names | HCN3, KIAA1535 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3. | |||||
|
HCN2_MOUSE
|
||||||
| NC score | 0.780277 (rank : 23) | θ value | 1.57e-36 (rank : 16) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O88703, O70506 | Gene names | Hcn2, Bcng2, Hac1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2) (Hyperpolarization-activated cation channel 1) (HAC-1). | |||||
|
HCN4_HUMAN
|
||||||
| NC score | 0.777496 (rank : 24) | θ value | 4.568e-36 (rank : 17) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Y3Q4, Q9UMQ7 | Gene names | HCN4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4. | |||||
|
HCN4_MOUSE
|
||||||
| NC score | 0.775728 (rank : 25) | θ value | 1.01765e-35 (rank : 18) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O70507 | Gene names | Hcn4, Bcng3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4 (Brain cyclic nucleotide-gated channel 3) (BCNG-3). | |||||
|
HCN2_HUMAN
|
||||||
| NC score | 0.771816 (rank : 26) | θ value | 1.32909e-35 (rank : 19) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UL51, O60742, O60743, O75267, Q9UBS2 | Gene names | HCN2, BCNG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2). | |||||
|
CNGB3_MOUSE
|
||||||
| NC score | 0.745273 (rank : 27) | θ value | 3.51386e-20 (rank : 28) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9JJZ9 | Gene names | Cngb3, Cng6 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel beta 3 (CNG channel beta 3) (Cyclic nucleotide-gated channel beta 3) (Cone photoreceptor cGMP- gated channel subunit beta) (Cyclic nucleotide-gated cation channel modulatory subunit) (Cyclic nucleotide-gated channel subunit CNG6). | |||||
|
CNGB3_HUMAN
|
||||||
| NC score | 0.735686 (rank : 28) | θ value | 2.20094e-22 (rank : 27) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9NQW8, Q9NRE9 | Gene names | CNGB3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel beta 3 (CNG channel beta 3) (Cyclic nucleotide-gated channel beta 3) (Cone photoreceptor cGMP- gated channel subunit beta) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
CNGB1_HUMAN
|
||||||
| NC score | 0.725681 (rank : 29) | θ value | 6.62687e-19 (rank : 29) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14028, Q14029 | Gene names | CNGB1, CNCG4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel 4 (CNG channel 4) (CNG-4) (CNG4) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
KCNKH_HUMAN
|
||||||
| NC score | 0.288939 (rank : 30) | θ value | 0.00020696 (rank : 30) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q96T54, Q5TCF4, Q8TAW4, Q9BXD1, Q9H592 | Gene names | KCNK17, TALK2, TASK4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 17 (TWIK-related alkaline pH- activated K(+) channel 2) (2P domain potassium channel Talk-2) (TWIK- related acid-sensitive K(+) channel 4) (TASK-4). | |||||
|
KCNK2_HUMAN
|
||||||
| NC score | 0.225128 (rank : 31) | θ value | 0.00035302 (rank : 31) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O95069, Q9UNE3 | Gene names | KCNK2, TREK, TREK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (TREK-1 K(+) channel subunit) (Two-pore potassium channel TPKC1). | |||||
|
KCNK2_MOUSE
|
||||||
| NC score | 0.224145 (rank : 32) | θ value | 0.000602161 (rank : 32) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P97438 | Gene names | Kcnk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (Two-pore potassium channel TPKC1) (TREK-1 K(+) channel subunit). | |||||
|
KCNKF_HUMAN
|
||||||
| NC score | 0.224081 (rank : 33) | θ value | 0.0252991 (rank : 44) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9H427, Q9HBC8 | Gene names | KCNK15, TASK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 15 (Acid-sensitive potassium channel protein TASK-5) (TWIK-related acid-sensitive K(+) channel 5) (Two pore potassium channel KT3.3). | |||||
|
KCNK3_HUMAN
|
||||||
| NC score | 0.219668 (rank : 34) | θ value | 0.00175202 (rank : 33) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O14649 | Gene names | KCNK3, TASK, TASK1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Two pore potassium channel KT3.1). | |||||
|
KCNKA_HUMAN
|
||||||
| NC score | 0.217676 (rank : 35) | θ value | 0.00509761 (rank : 37) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P57789, Q8TDK7, Q8TDK8, Q9HB59 | Gene names | KCNK10, TREK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 10 (Outward rectifying potassium channel protein TREK-2) (TREK-2 K(+) channel subunit). | |||||
|
KCNK3_MOUSE
|
||||||
| NC score | 0.215138 (rank : 36) | θ value | 0.00298849 (rank : 34) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O35111, O35163 | Gene names | Kcnk3, Ctbak, Task, Task1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Cardiac two-pore background K(+) channel) (cTBAK-1) (Two pore potassium channel KT3.1). | |||||
|
KCNK6_HUMAN
|
||||||
| NC score | 0.214788 (rank : 37) | θ value | 0.00298849 (rank : 36) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Y257, Q9HB47 | Gene names | KCNK6, TOSS, TWIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 6 (Inward rectifying potassium channel protein TWIK-2) (TWIK-originated similarity sequence). | |||||
|
KCNKG_HUMAN
|
||||||
| NC score | 0.209863 (rank : 38) | θ value | 0.00509761 (rank : 38) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q96T55, Q5TCF3, Q9H591 | Gene names | KCNK16, TALK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 16 (TWIK-related alkaline pH- activated K(+) channel 1) (2P domain potassium channel Talk-1). | |||||
|
KCNK4_HUMAN
|
||||||
| NC score | 0.206307 (rank : 39) | θ value | 0.00298849 (rank : 35) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9NYG8, Q96T94 | Gene names | KCNK4, TRAAK | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK) (Two pore K(+) channel KT4.1). | |||||
|
KCNK4_MOUSE
|
||||||
| NC score | 0.193867 (rank : 40) | θ value | 0.0252991 (rank : 43) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
KCNK9_MOUSE
|
||||||
| NC score | 0.191984 (rank : 41) | θ value | 0.0113563 (rank : 40) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q3LS21 | Gene names | Kcnk9, Task3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3). | |||||
|
KCNK9_HUMAN
|
||||||
| NC score | 0.191570 (rank : 42) | θ value | 0.0113563 (rank : 39) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9NPC2, Q2M290, Q540F2 | Gene names | KCNK9, TASK3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3) (Two pore potassium channel KT3.2). | |||||
|
KCNK1_HUMAN
|
||||||
| NC score | 0.164758 (rank : 43) | θ value | 0.279714 (rank : 50) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O00180, Q13307 | Gene names | KCNK1, HOHO1, KCNO1, TWIK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1) (Potassium channel KCNO1). | |||||
|
KCNK5_HUMAN
|
||||||
| NC score | 0.163634 (rank : 44) | θ value | 0.279714 (rank : 51) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
KCNK1_MOUSE
|
||||||
| NC score | 0.161118 (rank : 45) | θ value | 0.365318 (rank : 52) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08581 | Gene names | Kcnk1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1). | |||||
|
KAP0_HUMAN
|
||||||
| NC score | 0.130090 (rank : 46) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P10644, Q567S7 | Gene names | PRKAR1A, PKR1, PRKAR1 | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type I-alpha regulatory subunit (Tissue- specific extinguisher 1) (TSE1). | |||||
|
KCNKD_MOUSE
|
||||||
| NC score | 0.128247 (rank : 47) | θ value | 0.47712 (rank : 54) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8R1P5 | Gene names | Kcnk13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KAP0_MOUSE
|
||||||
| NC score | 0.117989 (rank : 48) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9DBC7, Q3UKU7, Q9JHR5, Q9JHR6 | Gene names | Prkar1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | cAMP-dependent protein kinase type I-alpha regulatory subunit. | |||||
|
KCNK7_MOUSE
|
||||||
| NC score | 0.109768 (rank : 49) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9Z2T1, Q9QXY0, Q9QYE8, Q9R1V1, Q9R242 | Gene names | Kcnk7, Dpkch3, Kcnk6, Kcnk8, Knot1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7 (Putative potassium channel DP3) (Double-pore K(+) channel 3) (Neuromuscular two p domain potassium channel). | |||||
|
KCNKD_HUMAN
|
||||||
| NC score | 0.109156 (rank : 50) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HB14, Q96E79 | Gene names | KCNK13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KAP1_HUMAN
|
||||||
| NC score | 0.104611 (rank : 51) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P31321 | Gene names | PRKAR1B | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type I-beta regulatory subunit. | |||||
|
KCNKC_HUMAN
|
||||||
| NC score | 0.104002 (rank : 52) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HB15 | Gene names | KCNK12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 12 (Tandem pore domain halothane- inhibited potassium channel 2) (THIK-2). | |||||
|
KAP1_MOUSE
|
||||||
| NC score | 0.102273 (rank : 53) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P12849 | Gene names | Prkar1b | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type I-beta regulatory subunit. | |||||
|
KCND1_MOUSE
|
||||||
| NC score | 0.087230 (rank : 54) | θ value | 0.0961366 (rank : 45) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KCNK7_HUMAN
|
||||||
| NC score | 0.086340 (rank : 55) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y2U2, Q9Y2U3, Q9Y2U4 | Gene names | KCNK7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7. | |||||
|
KCND1_HUMAN
|
||||||
| NC score | 0.082387 (rank : 56) | θ value | 0.125558 (rank : 46) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
KCND3_HUMAN
|
||||||
| NC score | 0.080990 (rank : 57) | θ value | 0.0193708 (rank : 41) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND3_MOUSE
|
||||||
| NC score | 0.080981 (rank : 58) | θ value | 0.0193708 (rank : 42) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND2_HUMAN
|
||||||
| NC score | 0.080908 (rank : 59) | θ value | 0.125558 (rank : 47) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_MOUSE
|
||||||
| NC score | 0.080907 (rank : 60) | θ value | 0.125558 (rank : 48) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KAP2_MOUSE
|
||||||
| NC score | 0.077896 (rank : 61) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P12367 | Gene names | Prkar2a | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type II-alpha regulatory subunit. | |||||
|
KAP3_MOUSE
|
||||||
| NC score | 0.077748 (rank : 62) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P31324, Q3UTZ1, Q80ZM4, Q8BRZ7 | Gene names | Prkar2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | cAMP-dependent protein kinase type II-beta regulatory subunit. | |||||
|
KAP2_HUMAN
|
||||||
| NC score | 0.075706 (rank : 63) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P13861, Q16823 | Gene names | PRKAR2A, PKR2, PRKAR2 | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type II-alpha regulatory subunit. | |||||
|
KAP3_HUMAN
|
||||||
| NC score | 0.074483 (rank : 64) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P31323 | Gene names | PRKAR2B | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type II-beta regulatory subunit. | |||||
|
CT152_MOUSE
|
||||||
| NC score | 0.060447 (rank : 65) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9D5U8, Q8R083 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf152 homolog. | |||||
|
KCMA1_MOUSE
|
||||||
| NC score | 0.058476 (rank : 66) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCMA1_HUMAN
|
||||||
| NC score | 0.058338 (rank : 67) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
KCNQ3_MOUSE
|
||||||
| NC score | 0.054325 (rank : 68) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8K3F6 | Gene names | Kcnq3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
KCNQ3_HUMAN
|
||||||
| NC score | 0.053020 (rank : 69) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O43525 | Gene names | KCNQ3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
KCNS2_HUMAN
|
||||||
| NC score | 0.051801 (rank : 70) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9ULS6 | Gene names | KCNS2, KIAA1144 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNS2_MOUSE
|
||||||
| NC score | 0.051196 (rank : 71) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O35174, Q543P3 | Gene names | Kcns2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNQ4_HUMAN
|
||||||
| NC score | 0.049993 (rank : 72) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P56696, O96025 | Gene names | KCNQ4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 4 (Voltage-gated potassium channel subunit Kv7.4) (Potassium channel subunit alpha KvLQT4) (KQT-like 4). | |||||
|
KCNG4_HUMAN
|
||||||
| NC score | 0.044228 (rank : 73) | θ value | 1.06291 (rank : 58) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8TDN1, Q96H24 | Gene names | KCNG4, KCNG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 4 (Voltage-gated potassium channel subunit Kv6.4). | |||||
|
KCNB2_HUMAN
|
||||||
| NC score | 0.043655 (rank : 74) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
KCNB1_MOUSE
|
||||||
| NC score | 0.042471 (rank : 75) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q03717 | Gene names | Kcnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (mShab). | |||||
|
KCNB1_HUMAN
|
||||||
| NC score | 0.042453 (rank : 76) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q14721, Q14193 | Gene names | KCNB1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (h-DRK1). | |||||
|
KCNG2_HUMAN
|
||||||
| NC score | 0.041688 (rank : 77) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UJ96 | Gene names | KCNG2, KCNF2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 2 (Voltage-gated potassium channel subunit Kv6.2) (Cardiac potassium channel subunit). | |||||
|
KCNT1_HUMAN
|
||||||
| NC score | 0.040324 (rank : 78) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5JUK3, Q9P2C5 | Gene names | KCNT1, KIAA1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1 (KCa4.1). | |||||
|
KGP1B_MOUSE
|
||||||
| NC score | 0.039341 (rank : 79) | θ value | 0.62314 (rank : 56) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Z0Z0 | Gene names | Prkg1, Prkg1b, Prkgr1b | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, beta isozyme (EC 2.7.11.12) (cGK 1 beta) (cGKI-beta). | |||||
|
KGP1B_HUMAN
|
||||||
| NC score | 0.039195 (rank : 80) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 921 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P14619 | Gene names | PRKG1, PRKG1B, PRKGR1B | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, beta isozyme (EC 2.7.11.12) (cGK 1 beta) (cGKI-beta). | |||||
|
KGP1A_HUMAN
|
||||||
| NC score | 0.039166 (rank : 81) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13976 | Gene names | PRKG1, PRKGR1A | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-dependent protein kinase 1, alpha isozyme (EC 2.7.11.12) (CGK 1 alpha) (cGKI-alpha). | |||||
|
KCNA2_MOUSE
|
||||||
| NC score | 0.038574 (rank : 82) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P63141, P15386, Q02010, Q8C8W4 | Gene names | Kcna2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (MK2). | |||||
|
KCNA4_HUMAN
|
||||||
| NC score | 0.038270 (rank : 83) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
KCNQ5_HUMAN
|
||||||
| NC score | 0.037156 (rank : 84) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NR82, Q9NRN0, Q9NYA6 | Gene names | KCNQ5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 5 (Voltage-gated potassium channel subunit Kv7.5) (Potassium channel subunit alpha KvLQT5) (KQT-like 5). | |||||
|
KCNA2_HUMAN
|
||||||
| NC score | 0.037051 (rank : 85) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P16389 | Gene names | KCNA2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (HBK5) (NGK1) (HUKIV). | |||||
|
KCNS1_HUMAN
|
||||||
| NC score | 0.036465 (rank : 86) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96KK3, O43652, Q6DJU6 | Gene names | KCNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
KCNT1_MOUSE
|
||||||
| NC score | 0.035147 (rank : 87) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6ZPR4, Q8C3E7 | Gene names | Kcnt1, Kiaa1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1. | |||||
|
DTX3_MOUSE
|
||||||
| NC score | 0.020546 (rank : 88) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80V91, Q9ER06 | Gene names | Dtx3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein deltex-3 (Deltex-3) (Deltex3) (mDTX3). | |||||
|
DTX3_HUMAN
|
||||||
| NC score | 0.019476 (rank : 89) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N9I9, Q8NAU6, Q8NDS8 | Gene names | DTX3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein deltex-3 (Deltex-3) (Deltex3). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.017233 (rank : 90) | θ value | 0.21417 (rank : 49) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
ARMX5_MOUSE
|
||||||
| NC score | 0.014736 (rank : 91) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UZB0 | Gene names | Armcx5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 5. | |||||
|
PHC1_HUMAN
|
||||||
| NC score | 0.014639 (rank : 92) | θ value | 0.47712 (rank : 55) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P78364, Q8WVM3, Q9BU63 | Gene names | PHC1, EDR1, PH1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (hPH1) (Early development regulatory protein 1). | |||||
|
RPGF2_HUMAN
|
||||||
| NC score | 0.013812 (rank : 93) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y4G8 | Gene names | RAPGEF2, KIAA0313, NRAPGEP, PDZGEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rap guanine nucleotide exchange factor 2 (Neural RAP guanine nucleotide exchange protein) (nRap GEP) (PDZ domain-containing guanine nucleotide exchange factor 1) (PDZ-GEF1) (RA-GEF). | |||||
|
FMN1B_MOUSE
|
||||||
| NC score | 0.011210 (rank : 94) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q05859 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoform IV (Limb deformity protein). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.010526 (rank : 95) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
GON4L_HUMAN
|
||||||
| NC score | 0.010470 (rank : 96) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
CAYP2_MOUSE
|
||||||
| NC score | 0.009115 (rank : 97) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BUG5, Q8C1J6 | Gene names | Caps2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcyphosin-2 (Calcyphosine-2). | |||||
|
TSH2_MOUSE
|
||||||
| NC score | 0.007414 (rank : 98) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q68FE9, Q5DTF1, Q6AXH5, Q9JL71 | Gene names | Tshz2, Kiaa4248, Sdccag33l, Tsh2, Zfp218, Znf218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 2 (Zinc finger protein 218) (SDCCAG33-like protein). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.006417 (rank : 99) | θ value | 0.62314 (rank : 57) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
HPS4_MOUSE
|
||||||
| NC score | 0.006333 (rank : 100) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99KG7 | Gene names | Hps4, Le | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 4 protein homolog (Light-ear protein) (Le protein). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.005766 (rank : 101) | θ value | 0.47712 (rank : 53) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
CCP1_MOUSE
|
||||||
| NC score | 0.004617 (rank : 102) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHG6, Q91WQ4, Q9JK50, Q9JK51, Q9JKK2, Q9JKK3 | Gene names | Dscr1 | |||
|
Domain Architecture |

|
|||||
| Description | Calcipressin-1 (Down syndrome critical region protein 1 homolog) (Myocyte-enriched calcineurin-interacting protein 1) (MCIP1). | |||||
|
SMCE1_HUMAN
|
||||||
| NC score | 0.004090 (rank : 103) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q969G3, O43539 | Gene names | SMARCE1, BAF57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
VINEX_MOUSE
|
||||||
| NC score | 0.003450 (rank : 104) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9R1Z8, Q62423 | Gene names | Sorbs3, Scam1, Sh3d4 | |||
|
Domain Architecture |

|
|||||
| Description | Vinexin (Sorbin and SH3 domain-containing protein 3) (SH3-containing adapter molecule 1) (SCAM-1) (SH3 domain-containing protein SH3P3). | |||||
|
CORO6_MOUSE
|
||||||
| NC score | 0.003159 (rank : 105) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q920M5, Q920M3, Q920M4 | Gene names | Coro6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coronin-6 (Clipin E). | |||||
|
K1H5_HUMAN
|
||||||
| NC score | 0.001943 (rank : 106) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92764, O76012, Q92651 | Gene names | KRTHA5, HHA5, HKA5 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cuticular Ha5 (Hair keratin, type I Ha5). | |||||
|
SEPT8_MOUSE
|
||||||
| NC score | 0.001687 (rank : 107) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CHH9, Q80YC7, Q9ESF7 | Gene names | Sept8, Kiaa0202 | |||
|
Domain Architecture |

|
|||||
| Description | Septin-8. | |||||
|
NEST_HUMAN
|
||||||
| NC score | 0.001648 (rank : 108) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.001618 (rank : 109) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
KIF22_HUMAN
|
||||||
| NC score | 0.001176 (rank : 110) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14807, O94814, Q9BT46 | Gene names | KIF22, KID, KNSL4 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF22 (Kinesin-like DNA-binding protein) (Kinesin-like protein 4). | |||||
|
CD276_HUMAN
|
||||||
| NC score | 0.000668 (rank : 111) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5ZPR3, Q6P5Y4, Q6UXI2, Q8NBI8, Q8NC34, Q8NCB6, Q9BXR1 | Gene names | CD276, B7H3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CD276 antigen precursor (Costimulatory molecule) (B7 homolog 3) (B7- H3) (4Ig-B7-H3). | |||||
|
RAI14_MOUSE
|
||||||
| NC score | -0.000050 (rank : 112) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1505 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9EP71, Q3URT3, Q6ZPT6 | Gene names | Rai14, Kiaa1334, Norpeg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein) (p125). | |||||
|
MYH13_HUMAN
|
||||||
| NC score | -0.000339 (rank : 113) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||