Please be patient as the page loads
|
KCNKD_MOUSE
|
||||||
| SwissProt Accessions | Q8R1P5 | Gene names | Kcnk13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
KCNKD_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.997528 (rank : 2) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9HB14, Q96E79 | Gene names | KCNK13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNKD_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8R1P5 | Gene names | Kcnk13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNKC_HUMAN
|
||||||
| θ value | 7.57797e-132 (rank : 3) | NC score | 0.990879 (rank : 3) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9HB15 | Gene names | KCNK12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 12 (Tandem pore domain halothane- inhibited potassium channel 2) (THIK-2). | |||||
|
KCNK3_MOUSE
|
||||||
| θ value | 3.17079e-29 (rank : 4) | NC score | 0.845538 (rank : 7) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O35111, O35163 | Gene names | Kcnk3, Ctbak, Task, Task1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Cardiac two-pore background K(+) channel) (cTBAK-1) (Two pore potassium channel KT3.1). | |||||
|
KCNK3_HUMAN
|
||||||
| θ value | 4.14116e-29 (rank : 5) | NC score | 0.845227 (rank : 8) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O14649 | Gene names | KCNK3, TASK, TASK1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Two pore potassium channel KT3.1). | |||||
|
KCNK9_HUMAN
|
||||||
| θ value | 1.92365e-26 (rank : 6) | NC score | 0.848616 (rank : 5) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9NPC2, Q2M290, Q540F2 | Gene names | KCNK9, TASK3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3) (Two pore potassium channel KT3.2). | |||||
|
KCNK9_MOUSE
|
||||||
| θ value | 9.54697e-26 (rank : 7) | NC score | 0.849190 (rank : 4) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q3LS21 | Gene names | Kcnk9, Task3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3). | |||||
|
KCNKF_HUMAN
|
||||||
| θ value | 1.5242e-23 (rank : 8) | NC score | 0.839136 (rank : 9) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9H427, Q9HBC8 | Gene names | KCNK15, TASK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 15 (Acid-sensitive potassium channel protein TASK-5) (TWIK-related acid-sensitive K(+) channel 5) (Two pore potassium channel KT3.3). | |||||
|
KCNK1_HUMAN
|
||||||
| θ value | 2.20094e-22 (rank : 9) | NC score | 0.830514 (rank : 10) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O00180, Q13307 | Gene names | KCNK1, HOHO1, KCNO1, TWIK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1) (Potassium channel KCNO1). | |||||
|
KCNKG_HUMAN
|
||||||
| θ value | 3.75424e-22 (rank : 10) | NC score | 0.845758 (rank : 6) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96T55, Q5TCF3, Q9H591 | Gene names | KCNK16, TALK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 16 (TWIK-related alkaline pH- activated K(+) channel 1) (2P domain potassium channel Talk-1). | |||||
|
KCNK1_MOUSE
|
||||||
| θ value | 1.42661e-21 (rank : 11) | NC score | 0.829987 (rank : 11) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08581 | Gene names | Kcnk1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1). | |||||
|
KCNKA_HUMAN
|
||||||
| θ value | 5.42112e-21 (rank : 12) | NC score | 0.794424 (rank : 18) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P57789, Q8TDK7, Q8TDK8, Q9HB59 | Gene names | KCNK10, TREK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 10 (Outward rectifying potassium channel protein TREK-2) (TREK-2 K(+) channel subunit). | |||||
|
KCNK2_MOUSE
|
||||||
| θ value | 7.82807e-20 (rank : 13) | NC score | 0.800881 (rank : 15) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97438 | Gene names | Kcnk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (Two-pore potassium channel TPKC1) (TREK-1 K(+) channel subunit). | |||||
|
KCNK2_HUMAN
|
||||||
| θ value | 1.02238e-19 (rank : 14) | NC score | 0.803714 (rank : 14) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O95069, Q9UNE3 | Gene names | KCNK2, TREK, TREK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (TREK-1 K(+) channel subunit) (Two-pore potassium channel TPKC1). | |||||
|
KCNK5_HUMAN
|
||||||
| θ value | 1.92812e-18 (rank : 15) | NC score | 0.821524 (rank : 12) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
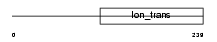
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
KCNK4_HUMAN
|
||||||
| θ value | 6.20254e-17 (rank : 16) | NC score | 0.794823 (rank : 17) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9NYG8, Q96T94 | Gene names | KCNK4, TRAAK | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK) (Two pore K(+) channel KT4.1). | |||||
|
KCNK6_HUMAN
|
||||||
| θ value | 8.10077e-17 (rank : 17) | NC score | 0.819788 (rank : 13) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9Y257, Q9HB47 | Gene names | KCNK6, TOSS, TWIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 6 (Inward rectifying potassium channel protein TWIK-2) (TWIK-originated similarity sequence). | |||||
|
KCNK4_MOUSE
|
||||||
| θ value | 6.85773e-16 (rank : 18) | NC score | 0.790999 (rank : 19) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
KCNKH_HUMAN
|
||||||
| θ value | 1.16975e-15 (rank : 19) | NC score | 0.797222 (rank : 16) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96T54, Q5TCF4, Q8TAW4, Q9BXD1, Q9H592 | Gene names | KCNK17, TALK2, TASK4 | |||
|
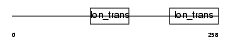
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 17 (TWIK-related alkaline pH- activated K(+) channel 2) (2P domain potassium channel Talk-2) (TWIK- related acid-sensitive K(+) channel 4) (TASK-4). | |||||
|
KCNK7_MOUSE
|
||||||
| θ value | 1.25267e-09 (rank : 20) | NC score | 0.751682 (rank : 20) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Z2T1, Q9QXY0, Q9QYE8, Q9R1V1, Q9R242 | Gene names | Kcnk7, Dpkch3, Kcnk6, Kcnk8, Knot1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7 (Putative potassium channel DP3) (Double-pore K(+) channel 3) (Neuromuscular two p domain potassium channel). | |||||
|
KCNK7_HUMAN
|
||||||
| θ value | 5.26297e-08 (rank : 21) | NC score | 0.731163 (rank : 21) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y2U2, Q9Y2U3, Q9Y2U4 | Gene names | KCNK7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7. | |||||
|
KCNT1_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 22) | NC score | 0.200747 (rank : 22) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5JUK3, Q9P2C5 | Gene names | KCNT1, KIAA1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1 (KCa4.1). | |||||
|
KCNT1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 23) | NC score | 0.188943 (rank : 23) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6ZPR4, Q8C3E7 | Gene names | Kcnt1, Kiaa1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1. | |||||
|
KCNH5_HUMAN
|
||||||
| θ value | 0.47712 (rank : 24) | NC score | 0.128247 (rank : 25) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8NCM2 | Gene names | KCNH5, EAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (hEAG2). | |||||
|
KCNH5_MOUSE
|
||||||
| θ value | 0.47712 (rank : 25) | NC score | 0.128288 (rank : 24) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q920E3, Q8C035 | Gene names | Kcnh5, Eag2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (Eag2). | |||||
|
VIPR2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 26) | NC score | 0.021655 (rank : 57) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P41588, P97750 | Gene names | Vipr2 | |||
|
Domain Architecture |

|
|||||
| Description | Vasoactive intestinal polypeptide receptor 2 precursor (VIP-R-2) (Pituitary adenylate cyclase-activating polypeptide type III receptor) (PACAP type III receptor) (PACAP-R-3). | |||||
|
KCNH2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 27) | NC score | 0.101172 (rank : 33) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q12809, O75418, O75680, Q9BT72, Q9BUT7, Q9H3P0 | Gene names | KCNH2, ERG, ERG1, HERG, HERG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (H-ERG) (Erg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1) (eag homolog). | |||||
|
KCNH2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 28) | NC score | 0.101106 (rank : 34) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35219, O35220, O35221, O35989 | Gene names | Kcnh2, Erg, Merg1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (ERG1) (MERG) (Merg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1). | |||||
|
KCMA1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 29) | NC score | 0.109733 (rank : 29) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
KCMA1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 30) | NC score | 0.109936 (rank : 28) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCNH7_HUMAN
|
||||||
| θ value | 1.06291 (rank : 31) | NC score | 0.105634 (rank : 30) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NS40 | Gene names | KCNH7, ERG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (HERG-3) (Ether-a-go-go-related protein 3) (Eag- related protein 3). | |||||
|
KCNH7_MOUSE
|
||||||
| θ value | 1.06291 (rank : 32) | NC score | 0.104602 (rank : 31) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9ER47 | Gene names | Kcnh7, Erg3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (Ether-a-go-go-related protein 3) (Eag-related protein 3). | |||||
|
KCNH6_HUMAN
|
||||||
| θ value | 1.38821 (rank : 33) | NC score | 0.104547 (rank : 32) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H252, Q9BRD7 | Gene names | KCNH6, ERG2, HERG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 6 (Voltage-gated potassium channel subunit Kv11.2) (Ether-a-go-go-related gene potassium channel 2) (Ether-a-go-go-related protein 2) (Eag-related protein 2). | |||||
|
VIPR2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 34) | NC score | 0.020723 (rank : 58) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P41587, Q13053, Q15870 | Gene names | VIPR2, VIP2R | |||
|
Domain Architecture |

|
|||||
| Description | Vasoactive intestinal polypeptide receptor 2 precursor (VIP-R-2) (Pituitary adenylate cyclase-activating polypeptide type III receptor) (PACAP type III receptor) (PACAP-R-3) (Helodermin-preferring VIP receptor). | |||||
|
KCNA5_MOUSE
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.049412 (rank : 56) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61762 | Gene names | Kcna5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (KV1-5). | |||||
|
KCNH1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.117731 (rank : 26) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O95259, O76035 | Gene names | KCNH1, EAG, EAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (hEAG1) (h-eag). | |||||
|
KCNH1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.117702 (rank : 27) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q60603 | Gene names | Kcnh1, Eag | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (EAG1) (m-eag). | |||||
|
TRPV3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.010332 (rank : 60) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NET8, Q8NDW7, Q8NET9, Q8NFH2 | Gene names | TRPV3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3) (Vanilloid receptor-like 3) (VRL-3). | |||||
|
KCNA4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.055701 (rank : 53) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
KCNA5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.050021 (rank : 55) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P22460 | Gene names | KCNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (HK2) (HPCN1). | |||||
|
KCNN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.060270 (rank : 44) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q92952 | Gene names | KCNN1, SK | |||
|
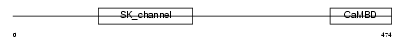
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCNN1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.061911 (rank : 42) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9EQR3, Q99JA9 | Gene names | Kcnn1, Sk1 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCNN2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.059204 (rank : 45) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9H2S1 | Gene names | KCNN2 | |||
|
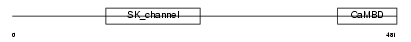
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
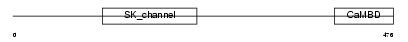
KCNN2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.059184 (rank : 46) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P58390 | Gene names | Kcnn2, Sk2 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.056625 (rank : 51) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UGI6, O43517 | Gene names | KCNN3, K3 | |||
|
Domain Architecture |
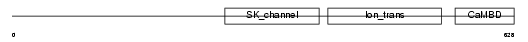
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3) (SKCa3). | |||||
|
KCNN3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.056743 (rank : 50) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P58391 | Gene names | Kcnn3, Sk3 | |||
|
Domain Architecture |
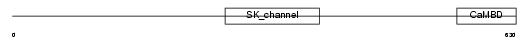
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3). | |||||
|
KCNN4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.085383 (rank : 38) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O89109, Q3U1J8, Q9CY39 | Gene names | Kcnn4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1). | |||||
|
TRPV3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.009380 (rank : 61) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K424, Q5SV08 | Gene names | Trpv3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3). | |||||
|
KCND1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.062502 (rank : 41) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
KCND1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.061081 (rank : 43) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KCNS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.020655 (rank : 59) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96KK3, O43652, Q6DJU6 | Gene names | KCNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
KCND2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.058740 (rank : 48) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.058730 (rank : 49) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.055700 (rank : 54) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.055772 (rank : 52) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCNH3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.088200 (rank : 37) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9ULD8, Q9UQ06 | Gene names | KCNH3, KIAA1282 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (ELK2) (Brain-specific eag-like channel 1) (BEC1). | |||||
|
KCNH3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.088946 (rank : 36) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WVJ0 | Gene names | Kcnh3, Elk2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (mElk2). | |||||
|
KCNH4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.081180 (rank : 39) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UQ05 | Gene names | KCNH4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 4 (Voltage-gated potassium channel subunit Kv12.3) (Ether-a-go-go-like potassium channel 1) (ELK channel 1) (ELK1) (Brain-specific eag-like channel 2) (BEC2). | |||||
|
KCNH8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.089190 (rank : 35) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96L42 | Gene names | KCNH8 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 8 (Voltage-gated potassium channel subunit Kv12.1) (Ether-a-go-go-like potassium channel 3) (ELK channel 3) (ELK3) (ELK1) (hElk1). | |||||
|
KCNN4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.078191 (rank : 40) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O15554 | Gene names | KCNN4, IK1, IKCA1, KCA4, SK4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1) (IKCa1) (Putative Gardos channel). | |||||
|
KCNQ4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.059001 (rank : 47) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P56696, O96025 | Gene names | KCNQ4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 4 (Voltage-gated potassium channel subunit Kv7.4) (Potassium channel subunit alpha KvLQT4) (KQT-like 4). | |||||
|
KCNKD_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8R1P5 | Gene names | Kcnk13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNKD_HUMAN
|
||||||
| NC score | 0.997528 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9HB14, Q96E79 | Gene names | KCNK13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNKC_HUMAN
|
||||||
| NC score | 0.990879 (rank : 3) | θ value | 7.57797e-132 (rank : 3) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9HB15 | Gene names | KCNK12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 12 (Tandem pore domain halothane- inhibited potassium channel 2) (THIK-2). | |||||
|
KCNK9_MOUSE
|
||||||
| NC score | 0.849190 (rank : 4) | θ value | 9.54697e-26 (rank : 7) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q3LS21 | Gene names | Kcnk9, Task3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3). | |||||
|
KCNK9_HUMAN
|
||||||
| NC score | 0.848616 (rank : 5) | θ value | 1.92365e-26 (rank : 6) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9NPC2, Q2M290, Q540F2 | Gene names | KCNK9, TASK3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3) (Two pore potassium channel KT3.2). | |||||
|
KCNKG_HUMAN
|
||||||
| NC score | 0.845758 (rank : 6) | θ value | 3.75424e-22 (rank : 10) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96T55, Q5TCF3, Q9H591 | Gene names | KCNK16, TALK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 16 (TWIK-related alkaline pH- activated K(+) channel 1) (2P domain potassium channel Talk-1). | |||||
|
KCNK3_MOUSE
|
||||||
| NC score | 0.845538 (rank : 7) | θ value | 3.17079e-29 (rank : 4) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O35111, O35163 | Gene names | Kcnk3, Ctbak, Task, Task1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Cardiac two-pore background K(+) channel) (cTBAK-1) (Two pore potassium channel KT3.1). | |||||
|
KCNK3_HUMAN
|
||||||
| NC score | 0.845227 (rank : 8) | θ value | 4.14116e-29 (rank : 5) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O14649 | Gene names | KCNK3, TASK, TASK1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Two pore potassium channel KT3.1). | |||||
|
KCNKF_HUMAN
|
||||||
| NC score | 0.839136 (rank : 9) | θ value | 1.5242e-23 (rank : 8) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9H427, Q9HBC8 | Gene names | KCNK15, TASK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 15 (Acid-sensitive potassium channel protein TASK-5) (TWIK-related acid-sensitive K(+) channel 5) (Two pore potassium channel KT3.3). | |||||
|
KCNK1_HUMAN
|
||||||
| NC score | 0.830514 (rank : 10) | θ value | 2.20094e-22 (rank : 9) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O00180, Q13307 | Gene names | KCNK1, HOHO1, KCNO1, TWIK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1) (Potassium channel KCNO1). | |||||
|
KCNK1_MOUSE
|
||||||
| NC score | 0.829987 (rank : 11) | θ value | 1.42661e-21 (rank : 11) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08581 | Gene names | Kcnk1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1). | |||||
|
KCNK5_HUMAN
|
||||||
| NC score | 0.821524 (rank : 12) | θ value | 1.92812e-18 (rank : 15) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
KCNK6_HUMAN
|
||||||
| NC score | 0.819788 (rank : 13) | θ value | 8.10077e-17 (rank : 17) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9Y257, Q9HB47 | Gene names | KCNK6, TOSS, TWIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 6 (Inward rectifying potassium channel protein TWIK-2) (TWIK-originated similarity sequence). | |||||
|
KCNK2_HUMAN
|
||||||
| NC score | 0.803714 (rank : 14) | θ value | 1.02238e-19 (rank : 14) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O95069, Q9UNE3 | Gene names | KCNK2, TREK, TREK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (TREK-1 K(+) channel subunit) (Two-pore potassium channel TPKC1). | |||||
|
KCNK2_MOUSE
|
||||||
| NC score | 0.800881 (rank : 15) | θ value | 7.82807e-20 (rank : 13) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97438 | Gene names | Kcnk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (Two-pore potassium channel TPKC1) (TREK-1 K(+) channel subunit). | |||||
|
KCNKH_HUMAN
|
||||||
| NC score | 0.797222 (rank : 16) | θ value | 1.16975e-15 (rank : 19) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96T54, Q5TCF4, Q8TAW4, Q9BXD1, Q9H592 | Gene names | KCNK17, TALK2, TASK4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 17 (TWIK-related alkaline pH- activated K(+) channel 2) (2P domain potassium channel Talk-2) (TWIK- related acid-sensitive K(+) channel 4) (TASK-4). | |||||
|
KCNK4_HUMAN
|
||||||
| NC score | 0.794823 (rank : 17) | θ value | 6.20254e-17 (rank : 16) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9NYG8, Q96T94 | Gene names | KCNK4, TRAAK | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK) (Two pore K(+) channel KT4.1). | |||||
|
KCNKA_HUMAN
|
||||||
| NC score | 0.794424 (rank : 18) | θ value | 5.42112e-21 (rank : 12) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P57789, Q8TDK7, Q8TDK8, Q9HB59 | Gene names | KCNK10, TREK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 10 (Outward rectifying potassium channel protein TREK-2) (TREK-2 K(+) channel subunit). | |||||
|
KCNK4_MOUSE
|
||||||
| NC score | 0.790999 (rank : 19) | θ value | 6.85773e-16 (rank : 18) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
KCNK7_MOUSE
|
||||||
| NC score | 0.751682 (rank : 20) | θ value | 1.25267e-09 (rank : 20) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Z2T1, Q9QXY0, Q9QYE8, Q9R1V1, Q9R242 | Gene names | Kcnk7, Dpkch3, Kcnk6, Kcnk8, Knot1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7 (Putative potassium channel DP3) (Double-pore K(+) channel 3) (Neuromuscular two p domain potassium channel). | |||||
|
KCNK7_HUMAN
|
||||||
| NC score | 0.731163 (rank : 21) | θ value | 5.26297e-08 (rank : 21) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y2U2, Q9Y2U3, Q9Y2U4 | Gene names | KCNK7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7. | |||||
|
KCNT1_HUMAN
|
||||||
| NC score | 0.200747 (rank : 22) | θ value | 0.0113563 (rank : 22) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5JUK3, Q9P2C5 | Gene names | KCNT1, KIAA1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1 (KCa4.1). | |||||
|
KCNT1_MOUSE
|
||||||
| NC score | 0.188943 (rank : 23) | θ value | 0.0736092 (rank : 23) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6ZPR4, Q8C3E7 | Gene names | Kcnt1, Kiaa1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1. | |||||
|
KCNH5_MOUSE
|
||||||
| NC score | 0.128288 (rank : 24) | θ value | 0.47712 (rank : 25) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q920E3, Q8C035 | Gene names | Kcnh5, Eag2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (Eag2). | |||||
|
KCNH5_HUMAN
|
||||||
| NC score | 0.128247 (rank : 25) | θ value | 0.47712 (rank : 24) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8NCM2 | Gene names | KCNH5, EAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 5 (Voltage-gated potassium channel subunit Kv10.2) (Ether-a-go-go potassium channel 2) (hEAG2). | |||||
|
KCNH1_HUMAN
|
||||||
| NC score | 0.117731 (rank : 26) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O95259, O76035 | Gene names | KCNH1, EAG, EAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (hEAG1) (h-eag). | |||||
|
KCNH1_MOUSE
|
||||||
| NC score | 0.117702 (rank : 27) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q60603 | Gene names | Kcnh1, Eag | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (EAG1) (m-eag). | |||||
|
KCMA1_MOUSE
|
||||||
| NC score | 0.109936 (rank : 28) | θ value | 1.06291 (rank : 30) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCMA1_HUMAN
|
||||||
| NC score | 0.109733 (rank : 29) | θ value | 1.06291 (rank : 29) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
KCNH7_HUMAN
|
||||||
| NC score | 0.105634 (rank : 30) | θ value | 1.06291 (rank : 31) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NS40 | Gene names | KCNH7, ERG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (HERG-3) (Ether-a-go-go-related protein 3) (Eag- related protein 3). | |||||
|
KCNH7_MOUSE
|
||||||
| NC score | 0.104602 (rank : 31) | θ value | 1.06291 (rank : 32) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9ER47 | Gene names | Kcnh7, Erg3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (Ether-a-go-go-related protein 3) (Eag-related protein 3). | |||||
|
KCNH6_HUMAN
|
||||||
| NC score | 0.104547 (rank : 32) | θ value | 1.38821 (rank : 33) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H252, Q9BRD7 | Gene names | KCNH6, ERG2, HERG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 6 (Voltage-gated potassium channel subunit Kv11.2) (Ether-a-go-go-related gene potassium channel 2) (Ether-a-go-go-related protein 2) (Eag-related protein 2). | |||||
|
KCNH2_HUMAN
|
||||||
| NC score | 0.101172 (rank : 33) | θ value | 0.813845 (rank : 27) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q12809, O75418, O75680, Q9BT72, Q9BUT7, Q9H3P0 | Gene names | KCNH2, ERG, ERG1, HERG, HERG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (H-ERG) (Erg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1) (eag homolog). | |||||
|
KCNH2_MOUSE
|
||||||
| NC score | 0.101106 (rank : 34) | θ value | 0.813845 (rank : 28) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35219, O35220, O35221, O35989 | Gene names | Kcnh2, Erg, Merg1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (ERG1) (MERG) (Merg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1). | |||||
|
KCNH8_HUMAN
|
||||||
| NC score | 0.089190 (rank : 35) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96L42 | Gene names | KCNH8 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 8 (Voltage-gated potassium channel subunit Kv12.1) (Ether-a-go-go-like potassium channel 3) (ELK channel 3) (ELK3) (ELK1) (hElk1). | |||||
|
KCNH3_MOUSE
|
||||||
| NC score | 0.088946 (rank : 36) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WVJ0 | Gene names | Kcnh3, Elk2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (mElk2). | |||||
|
KCNH3_HUMAN
|
||||||
| NC score | 0.088200 (rank : 37) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9ULD8, Q9UQ06 | Gene names | KCNH3, KIAA1282 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 3 (Voltage-gated potassium channel subunit Kv12.2) (Ether-a-go-go-like potassium channel 2) (ELK channel 2) (ELK2) (Brain-specific eag-like channel 1) (BEC1). | |||||
|
KCNN4_MOUSE
|
||||||
| NC score | 0.085383 (rank : 38) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O89109, Q3U1J8, Q9CY39 | Gene names | Kcnn4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1). | |||||
|
KCNH4_HUMAN
|
||||||
| NC score | 0.081180 (rank : 39) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UQ05 | Gene names | KCNH4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 4 (Voltage-gated potassium channel subunit Kv12.3) (Ether-a-go-go-like potassium channel 1) (ELK channel 1) (ELK1) (Brain-specific eag-like channel 2) (BEC2). | |||||
|
KCNN4_HUMAN
|
||||||
| NC score | 0.078191 (rank : 40) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O15554 | Gene names | KCNN4, IK1, IKCA1, KCA4, SK4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1) (IKCa1) (Putative Gardos channel). | |||||
|
KCND1_HUMAN
|
||||||
| NC score | 0.062502 (rank : 41) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
KCNN1_MOUSE
|
||||||
| NC score | 0.061911 (rank : 42) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9EQR3, Q99JA9 | Gene names | Kcnn1, Sk1 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCND1_MOUSE
|
||||||
| NC score | 0.061081 (rank : 43) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KCNN1_HUMAN
|
||||||
| NC score | 0.060270 (rank : 44) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q92952 | Gene names | KCNN1, SK | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCNN2_HUMAN
|
||||||
| NC score | 0.059204 (rank : 45) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9H2S1 | Gene names | KCNN2 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN2_MOUSE
|
||||||
| NC score | 0.059184 (rank : 46) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P58390 | Gene names | Kcnn2, Sk2 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNQ4_HUMAN
|
||||||
| NC score | 0.059001 (rank : 47) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P56696, O96025 | Gene names | KCNQ4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 4 (Voltage-gated potassium channel subunit Kv7.4) (Potassium channel subunit alpha KvLQT4) (KQT-like 4). | |||||
|
KCND2_HUMAN
|
||||||
| NC score | 0.058740 (rank : 48) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_MOUSE
|
||||||
| NC score | 0.058730 (rank : 49) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCNN3_MOUSE
|
||||||
| NC score | 0.056743 (rank : 50) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P58391 | Gene names | Kcnn3, Sk3 | |||
|
Domain Architecture |
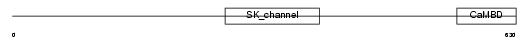
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3). | |||||
|
KCNN3_HUMAN
|
||||||
| NC score | 0.056625 (rank : 51) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UGI6, O43517 | Gene names | KCNN3, K3 | |||
|
Domain Architecture |
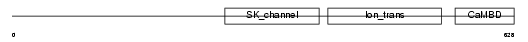
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3) (SKCa3). | |||||
|
KCND3_MOUSE
|
||||||
| NC score | 0.055772 (rank : 52) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCNA4_HUMAN
|
||||||
| NC score | 0.055701 (rank : 53) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
KCND3_HUMAN
|
||||||
| NC score | 0.055700 (rank : 54) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCNA5_HUMAN
|
||||||
| NC score | 0.050021 (rank : 55) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P22460 | Gene names | KCNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (HK2) (HPCN1). | |||||
|
KCNA5_MOUSE
|
||||||
| NC score | 0.049412 (rank : 56) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61762 | Gene names | Kcna5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (KV1-5). | |||||
|
VIPR2_MOUSE
|
||||||
| NC score | 0.021655 (rank : 57) | θ value | 0.47712 (rank : 26) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P41588, P97750 | Gene names | Vipr2 | |||
|
Domain Architecture |

|
|||||
| Description | Vasoactive intestinal polypeptide receptor 2 precursor (VIP-R-2) (Pituitary adenylate cyclase-activating polypeptide type III receptor) (PACAP type III receptor) (PACAP-R-3). | |||||
|
VIPR2_HUMAN
|
||||||
| NC score | 0.020723 (rank : 58) | θ value | 1.38821 (rank : 34) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P41587, Q13053, Q15870 | Gene names | VIPR2, VIP2R | |||
|
Domain Architecture |

|
|||||
| Description | Vasoactive intestinal polypeptide receptor 2 precursor (VIP-R-2) (Pituitary adenylate cyclase-activating polypeptide type III receptor) (PACAP type III receptor) (PACAP-R-3) (Helodermin-preferring VIP receptor). | |||||
|
KCNS1_HUMAN
|
||||||
| NC score | 0.020655 (rank : 59) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96KK3, O43652, Q6DJU6 | Gene names | KCNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
TRPV3_HUMAN
|
||||||
| NC score | 0.010332 (rank : 60) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NET8, Q8NDW7, Q8NET9, Q8NFH2 | Gene names | TRPV3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3) (Vanilloid receptor-like 3) (VRL-3). | |||||
|
TRPV3_MOUSE
|
||||||
| NC score | 0.009380 (rank : 61) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K424, Q5SV08 | Gene names | Trpv3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3). | |||||