Please be patient as the page loads
|
KCNN3_MOUSE
|
||||||
| SwissProt Accessions | P58391 | Gene names | Kcnn3, Sk3 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
KCNN1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.979626 (rank : 6) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q92952 | Gene names | KCNN1, SK | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCNN1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.979850 (rank : 5) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9EQR3, Q99JA9 | Gene names | Kcnn1, Sk1 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCNN2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.983771 (rank : 3) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9H2S1 | Gene names | KCNN2 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.982840 (rank : 4) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P58390 | Gene names | Kcnn2, Sk2 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN3_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.988439 (rank : 2) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 129 | |
| SwissProt Accessions | Q9UGI6, O43517 | Gene names | KCNN3, K3 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3) (SKCa3). | |||||
|
KCNN3_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 171 | |
| SwissProt Accessions | P58391 | Gene names | Kcnn3, Sk3 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3). | |||||
|
KCNN4_MOUSE
|
||||||
| θ value | 2.15548e-94 (rank : 7) | NC score | 0.946309 (rank : 7) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O89109, Q3U1J8, Q9CY39 | Gene names | Kcnn4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1). | |||||
|
KCNN4_HUMAN
|
||||||
| θ value | 2.01746e-92 (rank : 8) | NC score | 0.944375 (rank : 8) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O15554 | Gene names | KCNN4, IK1, IKCA1, KCA4, SK4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1) (IKCa1) (Putative Gardos channel). | |||||
|
KCNQ4_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 9) | NC score | 0.356350 (rank : 9) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P56696, O96025 | Gene names | KCNQ4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 4 (Voltage-gated potassium channel subunit Kv7.4) (Potassium channel subunit alpha KvLQT4) (KQT-like 4). | |||||
|
KCNQ5_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 10) | NC score | 0.351034 (rank : 10) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9NR82, Q9NRN0, Q9NYA6 | Gene names | KCNQ5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 5 (Voltage-gated potassium channel subunit Kv7.5) (Potassium channel subunit alpha KvLQT5) (KQT-like 5). | |||||
|
KCND2_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 11) | NC score | 0.296956 (rank : 18) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 12) | NC score | 0.296967 (rank : 17) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND1_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 13) | NC score | 0.296722 (rank : 19) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
KCND1_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 14) | NC score | 0.296703 (rank : 20) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KCNQ2_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 15) | NC score | 0.335066 (rank : 11) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9Z351, Q8R498, Q9QWN9, Q9Z342, Q9Z343, Q9Z344, Q9Z345, Q9Z346, Q9Z347, Q9Z348, Q9Z349, Q9Z350 | Gene names | Kcnq2, Kqt2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 2 (Voltage-gated potassium channel subunit Kv7.2) (Potassium channel subunit alpha KvLQT2) (KQT-like 2). | |||||
|
KCNQ2_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 16) | NC score | 0.332476 (rank : 12) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | O43526, O43796, O75580, O95845, Q5VYT8, Q96J59, Q99454 | Gene names | KCNQ2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 2 (Voltage-gated potassium channel subunit Kv7.2) (Neuroblastoma-specific potassium channel subunit alpha KvLQT2) (KQT-like 2). | |||||
|
KCND3_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 17) | NC score | 0.292704 (rank : 24) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND3_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 18) | NC score | 0.292929 (rank : 23) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCNS2_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 19) | NC score | 0.295846 (rank : 21) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O35174, Q543P3 | Gene names | Kcns2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNQ3_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 20) | NC score | 0.326684 (rank : 15) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O43525 | Gene names | KCNQ3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
KCNS2_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 21) | NC score | 0.294884 (rank : 22) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9ULS6 | Gene names | KCNS2, KIAA1144 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNQ3_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 22) | NC score | 0.323422 (rank : 16) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q8K3F6 | Gene names | Kcnq3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
ZN750_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 23) | NC score | 0.067649 (rank : 74) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BH05, Q66JP3, Q8C0L1 | Gene names | Znf750 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF750. | |||||
|
KCNA6_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 24) | NC score | 0.290862 (rank : 25) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P17658 | Gene names | KCNA6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (HBK2). | |||||
|
KCNA5_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 25) | NC score | 0.285621 (rank : 27) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q61762 | Gene names | Kcna5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (KV1-5). | |||||
|
KCNA6_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 26) | NC score | 0.288217 (rank : 26) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q61923 | Gene names | Kcna6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (MK1.6). | |||||
|
KCNB2_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 27) | NC score | 0.281574 (rank : 28) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
KCNA5_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 28) | NC score | 0.280793 (rank : 30) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P22460 | Gene names | KCNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (HK2) (HPCN1). | |||||
|
KCNA2_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 29) | NC score | 0.281286 (rank : 29) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P16389 | Gene names | KCNA2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (HBK5) (NGK1) (HUKIV). | |||||
|
KCNQ1_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 30) | NC score | 0.330709 (rank : 14) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P97414, O88702, Q7TNZ1, Q7TPL7, Q80VR7 | Gene names | Kcnq1, Kcna9, Kvlqt1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
EEA1_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 31) | NC score | 0.033019 (rank : 99) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
KCNA3_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 32) | NC score | 0.279367 (rank : 31) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P22001 | Gene names | KCNA3, HGK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (HPCN3) (HGK5) (HuKIII) (HLK3). | |||||
|
KCNA3_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 33) | NC score | 0.278769 (rank : 33) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P16390 | Gene names | Kcna3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (MK3). | |||||
|
KCNQ1_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 34) | NC score | 0.331766 (rank : 13) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P51787, O00347, O60607, O94787, Q7Z6G9, Q92960, Q9UMN8, Q9UMN9 | Gene names | KCNQ1, KCNA8, KCNA9, KVLQT1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
KCNA2_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 35) | NC score | 0.278932 (rank : 32) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P63141, P15386, Q02010, Q8C8W4 | Gene names | Kcna2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (MK2). | |||||
|
KCNB1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 36) | NC score | 0.273534 (rank : 39) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q14721, Q14193 | Gene names | KCNB1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (h-DRK1). | |||||
|
KCNB1_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 37) | NC score | 0.273577 (rank : 38) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q03717 | Gene names | Kcnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (mShab). | |||||
|
KCNA1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 38) | NC score | 0.274498 (rank : 35) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q09470 | Gene names | KCNA1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (HUKI) (HBK1). | |||||
|
KCNA1_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 39) | NC score | 0.274233 (rank : 37) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P16388 | Gene names | Kcna1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (MKI) (MBK1). | |||||
|
KCNS3_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 40) | NC score | 0.274409 (rank : 36) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9BQ31, O43651, Q96B56 | Gene names | KCNS3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily S member 3 (Voltage-gated potassium channel subunit Kv9.3) (Delayed-rectifier K(+) channel alpha subunit 3). | |||||
|
KCNA4_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 41) | NC score | 0.278531 (rank : 34) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
KCNG3_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 42) | NC score | 0.267266 (rank : 41) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8TAE7 | Gene names | KCNG3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
KCNG3_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 43) | NC score | 0.267081 (rank : 42) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P59053 | Gene names | Kcng3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
ZBT24_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 44) | NC score | 0.000214 (rank : 181) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 894 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80X44, Q3UIB7, Q7TPC4, Q8CJC9 | Gene names | Zbtb24, Bif1, Bsg1, Znf450 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 24 (Zinc finger protein 450) (Bone morphogenetic protein-induced factor 1) (Brain-specific protein 1). | |||||
|
AMOL1_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 45) | NC score | 0.042516 (rank : 90) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
NFAT5_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 46) | NC score | 0.037447 (rank : 93) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94916, O95693, Q9UN18 | Gene names | NFAT5, KIAA0827, TONEBP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells 5 (T-cell transcription factor NFAT5) (NF-AT5) (Tonicity-responsive enhancer-binding protein) (TonE- binding protein) (TonEBP). | |||||
|
HGS_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 47) | NC score | 0.050349 (rank : 88) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
KCNC4_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 48) | NC score | 0.258026 (rank : 46) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q03721, Q3MIM4, Q5TBI6 | Gene names | KCNC4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4) (KSHIIIC). | |||||
|
KCNC4_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 49) | NC score | 0.258786 (rank : 45) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8R1C0 | Gene names | Kcnc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4). | |||||
|
KCNA4_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 50) | NC score | 0.273401 (rank : 40) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q61423 | Gene names | Kcna4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4). | |||||
|
UACA_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 51) | NC score | 0.023696 (rank : 122) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1622 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9BZF9, Q8N3B8, Q9HCL1, Q9NWC6 | Gene names | UACA, KIAA1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein. | |||||
|
EMSY_HUMAN
|
||||||
| θ value | 0.125558 (rank : 52) | NC score | 0.053885 (rank : 84) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |
|
|||||
| Description | Protein EMSY. | |||||
|
KCNS1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 53) | NC score | 0.260365 (rank : 44) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96KK3, O43652, Q6DJU6 | Gene names | KCNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
HGS_HUMAN
|
||||||
| θ value | 0.163984 (rank : 54) | NC score | 0.048583 (rank : 89) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
KCMA1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 55) | NC score | 0.214460 (rank : 57) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
KCMA1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 56) | NC score | 0.214846 (rank : 56) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCNC1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 57) | NC score | 0.251003 (rank : 51) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P48547 | Gene names | KCNC1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
KCNC1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 58) | NC score | 0.251061 (rank : 50) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P15388 | Gene names | Kcnc1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
KCNK1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 59) | NC score | 0.131099 (rank : 58) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O00180, Q13307 | Gene names | KCNK1, HOHO1, KCNO1, TWIK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1) (Potassium channel KCNO1). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 0.163984 (rank : 60) | NC score | 0.029291 (rank : 106) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
AKAP9_HUMAN
|
||||||
| θ value | 0.21417 (rank : 61) | NC score | 0.032378 (rank : 101) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
BAG3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 62) | NC score | 0.027467 (rank : 109) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O95817, Q9NT20, Q9P120 | Gene names | BAG3, BIS | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis) (Docking protein CAIR-1). | |||||
|
KCNC3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 63) | NC score | 0.251195 (rank : 49) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q63959, Q62088 | Gene names | Kcnc3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
KCNG1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 64) | NC score | 0.257895 (rank : 47) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9UIX4, O43528, Q9BRC1 | Gene names | KCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 1 (Voltage-gated potassium channel subunit Kv6.1) (kH2). | |||||
|
MYO5A_MOUSE
|
||||||
| θ value | 0.21417 (rank : 65) | NC score | 0.022250 (rank : 127) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1117 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q99104 | Gene names | Myo5a, Dilute | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle). | |||||
|
EMSY_MOUSE
|
||||||
| θ value | 0.279714 (rank : 66) | NC score | 0.062440 (rank : 76) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BMB0, Q5FWK5, Q80XU1, Q8VDW9 | Gene names | Emsy | |||
|
Domain Architecture |
|
|||||
| Description | Protein EMSY. | |||||
|
GOGA1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 67) | NC score | 0.032736 (rank : 100) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 994 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CW79, Q80YB0 | Gene names | Golga1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
PHLB1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 68) | NC score | 0.034241 (rank : 97) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q86UU1, O75133, Q4KMF8, Q8TEQ2 | Gene names | PHLDB1, DLNB07, KIAA0638, LL5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
KCNC3_HUMAN
|
||||||
| θ value | 0.365318 (rank : 69) | NC score | 0.248675 (rank : 54) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q14003 | Gene names | KCNC3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
KCNG2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 70) | NC score | 0.263524 (rank : 43) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9UJ96 | Gene names | KCNG2, KCNF2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily G member 2 (Voltage-gated potassium channel subunit Kv6.2) (Cardiac potassium channel subunit). | |||||
|
KCNG4_HUMAN
|
||||||
| θ value | 0.365318 (rank : 71) | NC score | 0.249934 (rank : 53) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8TDN1, Q96H24 | Gene names | KCNG4, KCNG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 4 (Voltage-gated potassium channel subunit Kv6.4). | |||||
|
KCNK6_HUMAN
|
||||||
| θ value | 0.365318 (rank : 72) | NC score | 0.130119 (rank : 59) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Y257, Q9HB47 | Gene names | KCNK6, TOSS, TWIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 6 (Inward rectifying potassium channel protein TWIK-2) (TWIK-originated similarity sequence). | |||||
|
KCNK1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 73) | NC score | 0.116383 (rank : 60) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08581 | Gene names | Kcnk1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1). | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 0.47712 (rank : 74) | NC score | 0.002344 (rank : 176) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
PRGC2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 75) | NC score | 0.040514 (rank : 92) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
SYNE2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 76) | NC score | 0.027221 (rank : 110) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
ETV5_MOUSE
|
||||||
| θ value | 0.62314 (rank : 77) | NC score | 0.016480 (rank : 143) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
AMOL1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 78) | NC score | 0.040977 (rank : 91) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8IY63, Q63HK7, Q8NDN0, Q8WXD1, Q96CM5 | Gene names | AMOTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
KCNV2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 79) | NC score | 0.248249 (rank : 55) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8TDN2 | Gene names | KCNV2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily V member 2 (Voltage-gated potassium channel subunit Kv8.2). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 0.813845 (rank : 80) | NC score | 0.026168 (rank : 112) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
TARSH_HUMAN
|
||||||
| θ value | 0.813845 (rank : 81) | NC score | 0.025052 (rank : 116) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
ETV5_HUMAN
|
||||||
| θ value | 1.06291 (rank : 82) | NC score | 0.016020 (rank : 144) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
HCN1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 83) | NC score | 0.017732 (rank : 141) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O60741 | Gene names | HCN1, BCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1). | |||||
|
HCN1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 84) | NC score | 0.020338 (rank : 134) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O88704, O54899, Q9D613 | Gene names | Hcn1, Bcng1, Hac2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1) (Hyperpolarization-activated cation channel 2) (HAC-2). | |||||
|
HIPK1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 85) | NC score | 0.003599 (rank : 173) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 838 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O88904, Q80TV5, Q9QUQ8, Q9QZR3, Q9WVN7 | Gene names | Hipk1, Kiaa0630, Myak, Nbak2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeodomain-interacting protein kinase 1 (EC 2.7.11.1) (Protein kinase Myak) (Nuclear body-associated kinase 2). | |||||
|
KCNF1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 86) | NC score | 0.253192 (rank : 48) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9H3M0, O43527 | Gene names | KCNF1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily F member 1 (Voltage-gated potassium channel subunit Kv5.1) (kH1). | |||||
|
KCNS1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 87) | NC score | 0.250247 (rank : 52) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O35173 | Gene names | Kcns1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | 1.06291 (rank : 88) | NC score | 0.030865 (rank : 102) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
NUP88_HUMAN
|
||||||
| θ value | 1.06291 (rank : 89) | NC score | 0.036273 (rank : 94) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q99567, Q9BWE5 | Gene names | NUP88 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear pore complex protein Nup88 (Nucleoporin Nup88) (88 kDa nuclear pore complex protein). | |||||
|
PHF3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 90) | NC score | 0.022691 (rank : 125) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
SAPS1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 91) | NC score | 0.014675 (rank : 148) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
SN1L2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 92) | NC score | 0.001853 (rank : 177) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H0K1, O94878, Q6AZE2, Q76N03, Q8NCV7, Q96CZ8 | Gene names | SNF1LK2, KIAA0781, QIK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Qin- induced kinase). | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 93) | NC score | 0.030211 (rank : 104) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
MYO5A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 94) | NC score | 0.019345 (rank : 138) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y4I1, O60653, Q07902, Q16249, Q9UE30, Q9UE31 | Gene names | MYO5A, MYH12 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle) (Myosin heavy chain 12) (Myoxin). | |||||
|
DTNA_HUMAN
|
||||||
| θ value | 1.81305 (rank : 95) | NC score | 0.014135 (rank : 150) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Y4J8, O15332, O15333, O75697, Q13197, Q13198, Q13199, Q13498, Q13499, Q13500 | Gene names | DTNA, DRP3 | |||
|
Domain Architecture |
|
|||||
| Description | Dystrobrevin alpha (Dystrobrevin-alpha) (Dystrophin-related protein 3). | |||||
|
DYH11_HUMAN
|
||||||
| θ value | 1.81305 (rank : 96) | NC score | 0.016753 (rank : 142) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96DT5, Q9UJ82 | Gene names | DNAH11 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 11 (Axonemal beta dynein heavy chain 11). | |||||
|
KCNK4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 97) | NC score | 0.076817 (rank : 71) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
KCNK9_HUMAN
|
||||||
| θ value | 1.81305 (rank : 98) | NC score | 0.104660 (rank : 64) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9NPC2, Q2M290, Q540F2 | Gene names | KCNK9, TASK3 | |||
|
Domain Architecture |
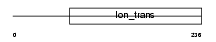
|
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3) (Two pore potassium channel KT3.2). | |||||
|
KCNK9_MOUSE
|
||||||
| θ value | 1.81305 (rank : 99) | NC score | 0.104262 (rank : 65) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q3LS21 | Gene names | Kcnk9, Task3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3). | |||||
|
LAMA2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 100) | NC score | 0.013190 (rank : 153) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
TCF7_HUMAN
|
||||||
| θ value | 1.81305 (rank : 101) | NC score | 0.015388 (rank : 147) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P36402, Q9UKI4 | Gene names | TCF7, TCF1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
ZN447_HUMAN
|
||||||
| θ value | 1.81305 (rank : 102) | NC score | 0.001213 (rank : 179) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TBC5, Q9BRK7, Q9H9A0 | Gene names | ZNF447 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 447. | |||||
|
EP300_HUMAN
|
||||||
| θ value | 2.36792 (rank : 103) | NC score | 0.022256 (rank : 126) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
HOME1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 104) | NC score | 0.033679 (rank : 98) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 630 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9Z2Y3, Q8K3E1, Q8K4M8, Q9Z0E9, Q9Z216 | Gene names | Homer1, Vesl1 | |||
|
Domain Architecture |
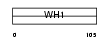
|
|||||
| Description | Homer protein homolog 1 (VASP/Ena-related gene up-regulated during seizure and LTP) (Vesl-1). | |||||
|
DESP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 105) | NC score | 0.025735 (rank : 113) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
KCNT1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 106) | NC score | 0.092407 (rank : 68) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q5JUK3, Q9P2C5 | Gene names | KCNT1, KIAA1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1 (KCa4.1). | |||||
|
LAMA4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 107) | NC score | 0.015836 (rank : 145) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q16363, Q14735, Q15335, Q4LE44, Q5SZG8, Q9UE18, Q9UJN9 | Gene names | LAMA4 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-4 chain precursor. | |||||
|
LMNB1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 108) | NC score | 0.020416 (rank : 133) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P20700, Q3SYN7, Q96EI6 | Gene names | LMNB1, LMN2, LMNB | |||
|
Domain Architecture |

|
|||||
| Description | Lamin-B1. | |||||
|
NAF1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 109) | NC score | 0.029275 (rank : 107) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q15025, O76008, Q96EL9, Q99833, Q9H1J3 | Gene names | TNIP1, KIAA0113, NAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (HIV-1 Nef-interacting protein) (Virion-associated nuclear shuttling protein) (VAN) (hVAN) (Nip40-1). | |||||
|
CP250_MOUSE
|
||||||
| θ value | 4.03905 (rank : 110) | NC score | 0.023964 (rank : 120) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
GOGA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 111) | NC score | 0.024529 (rank : 117) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
GOGA3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 112) | NC score | 0.023928 (rank : 121) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1437 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P55937, Q80VF5, Q8CCK4, Q9QYT2, Q9QYT3 | Gene names | Golga3, Mea2 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Male-enhanced antigen 2) (MEA-2). | |||||
|
HOME1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 113) | NC score | 0.029839 (rank : 105) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 564 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q86YM7, O96003, Q86YM5 | Gene names | HOMER1, SYN47 | |||
|
Domain Architecture |
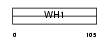
|
|||||
| Description | Homer protein homolog 1. | |||||
|
KCNH4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 114) | NC score | 0.034258 (rank : 96) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UQ05 | Gene names | KCNH4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 4 (Voltage-gated potassium channel subunit Kv12.3) (Ether-a-go-go-like potassium channel 1) (ELK channel 1) (ELK1) (Brain-specific eag-like channel 2) (BEC2). | |||||
|
LMNB1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 115) | NC score | 0.020093 (rank : 135) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 719 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P14733, Q61791 | Gene names | Lmnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Lamin-B1. | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 116) | NC score | 0.013033 (rank : 156) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
RPGR1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 117) | NC score | 0.034510 (rank : 95) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q96KN7, Q7Z2W6, Q8IXV5, Q96QA8, Q9HB94, Q9HB95, Q9HBK6, Q9NR40 | Gene names | RPGRIP1 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
SNTB1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 118) | NC score | 0.005778 (rank : 170) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99L88, O35925 | Gene names | Sntb1, Snt2b1 | |||
|
Domain Architecture |
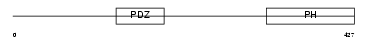
|
|||||
| Description | Beta-1-syntrophin (59 kDa dystrophin-associated protein A1 basic component 1) (DAPA1B) (Syntrophin 2). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 119) | NC score | 0.015591 (rank : 146) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
ZN366_HUMAN
|
||||||
| θ value | 4.03905 (rank : 120) | NC score | -0.001929 (rank : 183) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 732 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N895, Q7RTV4 | Gene names | ZNF366 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 366. | |||||
|
ZN462_HUMAN
|
||||||
| θ value | 4.03905 (rank : 121) | NC score | 0.000325 (rank : 180) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 740 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96JM2, Q8N408 | Gene names | ZNF462, KIAA1803 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 462. | |||||
|
ANGL6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 122) | NC score | 0.006875 (rank : 167) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NI99, Q9BZZ0 | Gene names | ANGPTL6, AGF, ARP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiopoietin-related protein 6 precursor (Angiopoietin-like 6) (Angiopoietin-related growth factor) (Angiopoietin-related protein 5). | |||||
|
C8AP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 123) | NC score | 0.019538 (rank : 137) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
CCD46_HUMAN
|
||||||
| θ value | 5.27518 (rank : 124) | NC score | 0.024160 (rank : 119) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
CF060_HUMAN
|
||||||
| θ value | 5.27518 (rank : 125) | NC score | 0.024280 (rank : 118) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1270 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8NB25, Q96GY8, Q9H851 | Gene names | C6orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf60. | |||||
|
FRAT2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 126) | NC score | 0.011207 (rank : 159) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75474, Q5JTI0, Q8WUL9, Q9BYG2 | Gene names | FRAT2 | |||
|
Domain Architecture |
|
|||||
| Description | GSK-3-binding protein FRAT2 (Frat-2). | |||||
|
GCC1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 127) | NC score | 0.030383 (rank : 103) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96CN9, Q9H6N7 | Gene names | GCC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP and coiled-coil domain-containing protein 1 (Golgi coiled coil protein 1). | |||||
|
GCC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 128) | NC score | 0.025192 (rank : 115) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
INVS_MOUSE
|
||||||
| θ value | 5.27518 (rank : 129) | NC score | 0.004394 (rank : 172) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
KCNK5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 130) | NC score | 0.109320 (rank : 63) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
KCNKD_HUMAN
|
||||||
| θ value | 5.27518 (rank : 131) | NC score | 0.063607 (rank : 75) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9HB14, Q96E79 | Gene names | KCNK13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
LIPS_HUMAN
|
||||||
| θ value | 5.27518 (rank : 132) | NC score | 0.019091 (rank : 139) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q05469, Q3LRT2, Q6NSL7 | Gene names | LIPE | |||
|
Domain Architecture |
|
|||||
| Description | Hormone-sensitive lipase (EC 3.1.1.79) (HSL). | |||||
|
NAF1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 133) | NC score | 0.026880 (rank : 111) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 852 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9WUU8, Q922A9, Q922F7, Q9EPP8, Q9R0X3 | Gene names | Tnip1, Abin, Naf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (A20- binding inhibitor of NF-kappa-B activation) (ABIN) (Virion-associated nuclear shuttling protein) (VAN) (mVAN). | |||||
|
RBP24_HUMAN
|
||||||
| θ value | 5.27518 (rank : 134) | NC score | 0.025229 (rank : 114) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
REST_MOUSE
|
||||||
| θ value | 5.27518 (rank : 135) | NC score | 0.020624 (rank : 132) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
SYCP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 136) | NC score | 0.021230 (rank : 130) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1456 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q62209, O09205, P70192, Q62329 | Gene names | Sycp1, Scp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
TNIK_HUMAN
|
||||||
| θ value | 5.27518 (rank : 137) | NC score | 0.002919 (rank : 175) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
TRI66_MOUSE
|
||||||
| θ value | 5.27518 (rank : 138) | NC score | 0.012987 (rank : 157) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
ATX2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 139) | NC score | 0.013867 (rank : 151) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O70305, P97421 | Gene names | Atxn2, Atx2, Sca2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein homolog). | |||||
|
CENPH_HUMAN
|
||||||
| θ value | 6.88961 (rank : 140) | NC score | 0.028642 (rank : 108) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9H3R5 | Gene names | CENPH, ICEN35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein H (CENP-H) (Interphase centromere complex protein 35). | |||||
|
CI093_HUMAN
|
||||||
| θ value | 6.88961 (rank : 141) | NC score | 0.021829 (rank : 128) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6TFL3, Q5SU58, Q6P1W1, Q6ZR13, Q8N1Z4, Q8N8L3, Q8TBR2 | Gene names | C9orf93 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf93. | |||||
|
CUTL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 142) | NC score | 0.013112 (rank : 154) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O14529 | Gene names | CUTL2, KIAA0293 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 2 (Homeobox protein Cux-2) (Cut-like 2). | |||||
|
HIPK1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 143) | NC score | 0.001423 (rank : 178) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86Z02, O75125, Q8IYD7, Q8NDN5, Q8NEB6, Q8TBZ1 | Gene names | HIPK1, KIAA0630 | |||
|
Domain Architecture |

|
|||||
| Description | Homeodomain-interacting protein kinase 1 (EC 2.7.11.1). | |||||
|
IRF3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 144) | NC score | 0.006756 (rank : 168) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14653 | Gene names | IRF3 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 3 (IRF-3). | |||||
|
KCNKA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 145) | NC score | 0.109602 (rank : 62) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P57789, Q8TDK7, Q8TDK8, Q9HB59 | Gene names | KCNK10, TREK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 10 (Outward rectifying potassium channel protein TREK-2) (TREK-2 K(+) channel subunit). | |||||
|
KCNKD_MOUSE
|
||||||
| θ value | 6.88961 (rank : 146) | NC score | 0.056743 (rank : 77) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8R1P5 | Gene names | Kcnk13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNKG_HUMAN
|
||||||
| θ value | 6.88961 (rank : 147) | NC score | 0.082813 (rank : 70) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q96T55, Q5TCF3, Q9H591 | Gene names | KCNK16, TALK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 16 (TWIK-related alkaline pH- activated K(+) channel 1) (2P domain potassium channel Talk-1). | |||||
|
LRSM1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 148) | NC score | 0.013090 (rank : 155) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q80ZI6 | Gene names | Lrsam1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase). | |||||
|
MA1B1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 149) | NC score | 0.006938 (rank : 166) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UKM7, Q5VSG3, Q9BRS9, Q9Y5K7 | Gene names | MAN1B1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase (EC 3.2.1.113) (ER alpha-1,2-mannosidase) (Mannosidase alpha class 1B member 1) (Man9GlcNAc2-specific-processing alpha-mannosidase). | |||||
|
NF2L2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 150) | NC score | 0.006570 (rank : 169) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60795 | Gene names | Nfe2l2, Nrf-2, Nrf2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor erythroid 2-related factor 2 (NF-E2-related factor 2) (NFE2-related factor 2) (Nuclear factor, erythroid derived 2, like 2). | |||||
|
RP3A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 151) | NC score | 0.004609 (rank : 171) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P47708 | Gene names | Rph3a | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
ZBTB4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 152) | NC score | -0.000449 (rank : 182) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 926 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P1Z0, Q7Z697, Q86XJ4, Q8N4V8 | Gene names | ZBTB4, KIAA1538 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 4 (KAISO-like zinc finger protein 1) (KAISO-L1). | |||||
|
ASPP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 153) | NC score | 0.010908 (rank : 160) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q13625, Q12892, Q86X75, Q96KQ3 | Gene names | TP53BP2, ASPP2, BBP | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 2 (Tumor suppressor p53-binding protein 2) (p53-binding protein 2) (p53BP2) (53BP2) (Bcl2-binding protein) (Bbp) (NY-REN-51 antigen). | |||||
|
BCAR1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 154) | NC score | 0.012147 (rank : 158) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 155) | NC score | 0.023257 (rank : 123) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 156) | NC score | 0.018069 (rank : 140) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 157) | NC score | 0.009215 (rank : 163) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
GA2L2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 158) | NC score | 0.009402 (rank : 162) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5SSG4 | Gene names | Gas2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 159) | NC score | 0.019621 (rank : 136) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
IFT81_HUMAN
|
||||||
| θ value | 8.99809 (rank : 160) | NC score | 0.020979 (rank : 131) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8WYA0, Q8NB51, Q9BSV2, Q9UNY8 | Gene names | IFT81, CDV1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 81 (Carnitine deficiency-associated protein expressed in ventricle 1) (CDV-1 protein). | |||||
|
K1914_HUMAN
|
||||||
| θ value | 8.99809 (rank : 161) | NC score | 0.014282 (rank : 149) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N4X5, Q8TB54, Q96PX4, Q96SY5 | Gene names | KIAA1914 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1914. | |||||
|
KCNK3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 162) | NC score | 0.092991 (rank : 66) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O14649 | Gene names | KCNK3, TASK, TASK1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Two pore potassium channel KT3.1). | |||||
|
KCNK3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 163) | NC score | 0.092918 (rank : 67) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O35111, O35163 | Gene names | Kcnk3, Ctbak, Task, Task1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Cardiac two-pore background K(+) channel) (cTBAK-1) (Two pore potassium channel KT3.1). | |||||
|
KCNKF_HUMAN
|
||||||
| θ value | 8.99809 (rank : 164) | NC score | 0.110366 (rank : 61) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9H427, Q9HBC8 | Gene names | KCNK15, TASK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 15 (Acid-sensitive potassium channel protein TASK-5) (TWIK-related acid-sensitive K(+) channel 5) (Two pore potassium channel KT3.3). | |||||
|
LCP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 165) | NC score | 0.008222 (rank : 165) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13094 | Gene names | LCP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte cytosolic protein 2 (SH2 domain-containing leukocyte protein of 76 kDa) (SLP-76 tyrosine phosphoprotein) (SLP76). | |||||
|
MAVS_HUMAN
|
||||||
| θ value | 8.99809 (rank : 166) | NC score | 0.008618 (rank : 164) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7Z434, Q3I0Y2, Q5T7I6, Q86VY7, Q9H1H3, Q9H4Y1, Q9H8D3, Q9ULE9 | Gene names | MAVS, IPS1, KIAA1271, VISA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif) (Putative NF-kappa-B- activating protein 031N). | |||||
|
MTSS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 167) | NC score | 0.013575 (rank : 152) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O43312, Q8TCA2, Q96RX2 | Gene names | MTSS1, KIAA0429, MIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein) (Metastasis suppressor YGL-1). | |||||
|
PRGC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 168) | NC score | 0.023159 (rank : 124) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86YN6, Q86YN3, Q86YN4, Q86YN5, Q8N1N9, Q8TDE4, Q8TDE5 | Gene names | PPARGC1B, PERC, PGC1, PGC1B, PPARGC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta) (PGC-1-related estrogen receptor alpha coactivator) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
R3HD2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 169) | NC score | 0.021559 (rank : 129) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
RFFL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 170) | NC score | 0.010471 (rank : 161) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8WZ73, Q8NHW0, Q8TBY7, Q96BE6 | Gene names | RFFL, RNF189, RNF34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (FYVE-RING finger protein Sakura) (Fring) (Caspases-8 and -10- associated RING finger protein 2) (CARP-2) (Caspase regulator CARP2) (RING finger protein 189) (RING finger protein 34-like). | |||||
|
SHC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 171) | NC score | 0.003323 (rank : 174) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P29353, O15290, Q8N4K5, Q96CL1 | Gene names | SHC1, SHC, SHCA | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 1 (SH2 domain protein C1) (Src homology 2 domain-containing-transforming protein C1). | |||||
|
KCD15_HUMAN
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.054783 (rank : 83) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96SI1, Q9BVI6 | Gene names | KCTD15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD15. | |||||
|
KCD15_MOUSE
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.052431 (rank : 85) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8K0E1 | Gene names | Kctd15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD15. | |||||
|
KCNK2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.069076 (rank : 73) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O95069, Q9UNE3 | Gene names | KCNK2, TREK, TREK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (TREK-1 K(+) channel subunit) (Two-pore potassium channel TPKC1). | |||||
|
KCNK2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.073286 (rank : 72) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P97438 | Gene names | Kcnk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (Two-pore potassium channel TPKC1) (TREK-1 K(+) channel subunit). | |||||
|
KCNK4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.055083 (rank : 81) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NYG8, Q96T94 | Gene names | KCNK4, TRAAK | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK) (Two pore K(+) channel KT4.1). | |||||
|
KCNKH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.084719 (rank : 69) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q96T54, Q5TCF4, Q8TAW4, Q9BXD1, Q9H592 | Gene names | KCNK17, TALK2, TASK4 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 17 (TWIK-related alkaline pH- activated K(+) channel 2) (2P domain potassium channel Talk-2) (TWIK- related acid-sensitive K(+) channel 4) (TASK-4). | |||||
|
KCTD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.055976 (rank : 78) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q719H9 | Gene names | KCTD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD1. | |||||
|
KCTD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.055976 (rank : 79) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5M956, Q5EER9 | Gene names | Kctd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD1. | |||||
|
KCTD4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 180) | NC score | 0.050457 (rank : 87) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WVF5 | Gene names | KCTD4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD4. | |||||
|
KCTD4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 181) | NC score | 0.050682 (rank : 86) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9D7X1, Q8CCQ3, Q9CYK4 | Gene names | Kctd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD4. | |||||
|
KCTD6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 182) | NC score | 0.054948 (rank : 82) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8NC69, Q8NBS6, Q8TCA6 | Gene names | KCTD6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD6. | |||||
|
KCTD6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 183) | NC score | 0.055383 (rank : 80) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BNL5, Q3TXX1 | Gene names | Kctd6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD6. | |||||
|
KCNN3_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 171 | |
| SwissProt Accessions | P58391 | Gene names | Kcnn3, Sk3 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3). | |||||
|
KCNN3_HUMAN
|
||||||
| NC score | 0.988439 (rank : 2) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 129 | |
| SwissProt Accessions | Q9UGI6, O43517 | Gene names | KCNN3, K3 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3) (SKCa3). | |||||
|
KCNN2_HUMAN
|
||||||
| NC score | 0.983771 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9H2S1 | Gene names | KCNN2 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN2_MOUSE
|
||||||
| NC score | 0.982840 (rank : 4) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P58390 | Gene names | Kcnn2, Sk2 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN1_MOUSE
|
||||||
| NC score | 0.979850 (rank : 5) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9EQR3, Q99JA9 | Gene names | Kcnn1, Sk1 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCNN1_HUMAN
|
||||||
| NC score | 0.979626 (rank : 6) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q92952 | Gene names | KCNN1, SK | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCNN4_MOUSE
|
||||||
| NC score | 0.946309 (rank : 7) | θ value | 2.15548e-94 (rank : 7) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O89109, Q3U1J8, Q9CY39 | Gene names | Kcnn4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1). | |||||
|
KCNN4_HUMAN
|
||||||
| NC score | 0.944375 (rank : 8) | θ value | 2.01746e-92 (rank : 8) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O15554 | Gene names | KCNN4, IK1, IKCA1, KCA4, SK4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1) (IKCa1) (Putative Gardos channel). | |||||
|
KCNQ4_HUMAN
|
||||||
| NC score | 0.356350 (rank : 9) | θ value | 4.1701e-05 (rank : 9) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P56696, O96025 | Gene names | KCNQ4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 4 (Voltage-gated potassium channel subunit Kv7.4) (Potassium channel subunit alpha KvLQT4) (KQT-like 4). | |||||
|
KCNQ5_HUMAN
|
||||||
| NC score | 0.351034 (rank : 10) | θ value | 4.1701e-05 (rank : 10) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9NR82, Q9NRN0, Q9NYA6 | Gene names | KCNQ5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 5 (Voltage-gated potassium channel subunit Kv7.5) (Potassium channel subunit alpha KvLQT5) (KQT-like 5). | |||||
|
KCNQ2_MOUSE
|
||||||
| NC score | 0.335066 (rank : 11) | θ value | 0.000461057 (rank : 15) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9Z351, Q8R498, Q9QWN9, Q9Z342, Q9Z343, Q9Z344, Q9Z345, Q9Z346, Q9Z347, Q9Z348, Q9Z349, Q9Z350 | Gene names | Kcnq2, Kqt2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 2 (Voltage-gated potassium channel subunit Kv7.2) (Potassium channel subunit alpha KvLQT2) (KQT-like 2). | |||||
|
KCNQ2_HUMAN
|
||||||
| NC score | 0.332476 (rank : 12) | θ value | 0.000602161 (rank : 16) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | O43526, O43796, O75580, O95845, Q5VYT8, Q96J59, Q99454 | Gene names | KCNQ2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 2 (Voltage-gated potassium channel subunit Kv7.2) (Neuroblastoma-specific potassium channel subunit alpha KvLQT2) (KQT-like 2). | |||||
|
KCNQ1_HUMAN
|
||||||
| NC score | 0.331766 (rank : 13) | θ value | 0.0193708 (rank : 34) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P51787, O00347, O60607, O94787, Q7Z6G9, Q92960, Q9UMN8, Q9UMN9 | Gene names | KCNQ1, KCNA8, KCNA9, KVLQT1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
KCNQ1_MOUSE
|
||||||
| NC score | 0.330709 (rank : 14) | θ value | 0.0148317 (rank : 30) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P97414, O88702, Q7TNZ1, Q7TPL7, Q80VR7 | Gene names | Kcnq1, Kcna9, Kvlqt1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
KCNQ3_HUMAN
|
||||||
| NC score | 0.326684 (rank : 15) | θ value | 0.00102713 (rank : 20) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O43525 | Gene names | KCNQ3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
KCNQ3_MOUSE
|
||||||
| NC score | 0.323422 (rank : 16) | θ value | 0.00228821 (rank : 22) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q8K3F6 | Gene names | Kcnq3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
KCND2_MOUSE
|
||||||
| NC score | 0.296967 (rank : 17) | θ value | 0.00035302 (rank : 12) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_HUMAN
|
||||||
| NC score | 0.296956 (rank : 18) | θ value | 0.00035302 (rank : 11) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND1_HUMAN
|
||||||
| NC score | 0.296722 (rank : 19) | θ value | 0.000461057 (rank : 13) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
KCND1_MOUSE
|
||||||
| NC score | 0.296703 (rank : 20) | θ value | 0.000461057 (rank : 14) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KCNS2_MOUSE
|
||||||
| NC score | 0.295846 (rank : 21) | θ value | 0.000786445 (rank : 19) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O35174, Q543P3 | Gene names | Kcns2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNS2_HUMAN
|
||||||
| NC score | 0.294884 (rank : 22) | θ value | 0.00102713 (rank : 21) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9ULS6 | Gene names | KCNS2, KIAA1144 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCND3_MOUSE
|
||||||
| NC score | 0.292929 (rank : 23) | θ value | 0.000786445 (rank : 18) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND3_HUMAN
|
||||||
| NC score | 0.292704 (rank : 24) | θ value | 0.000786445 (rank : 17) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCNA6_HUMAN
|
||||||
| NC score | 0.290862 (rank : 25) | θ value | 0.00298849 (rank : 24) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P17658 | Gene names | KCNA6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (HBK2). | |||||
|
KCNA6_MOUSE
|
||||||
| NC score | 0.288217 (rank : 26) | θ value | 0.00509761 (rank : 26) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q61923 | Gene names | Kcna6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (MK1.6). | |||||
|
KCNA5_MOUSE
|
||||||
| NC score | 0.285621 (rank : 27) | θ value | 0.00509761 (rank : 25) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q61762 | Gene names | Kcna5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (KV1-5). | |||||
|
KCNB2_HUMAN
|
||||||
| NC score | 0.281574 (rank : 28) | θ value | 0.00869519 (rank : 27) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
KCNA2_HUMAN
|
||||||
| NC score | 0.281286 (rank : 29) | θ value | 0.0148317 (rank : 29) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P16389 | Gene names | KCNA2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (HBK5) (NGK1) (HUKIV). | |||||
|
KCNA5_HUMAN
|
||||||
| NC score | 0.280793 (rank : 30) | θ value | 0.0113563 (rank : 28) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P22460 | Gene names | KCNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (HK2) (HPCN1). | |||||
|
KCNA3_HUMAN
|
||||||
| NC score | 0.279367 (rank : 31) | θ value | 0.0193708 (rank : 32) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P22001 | Gene names | KCNA3, HGK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (HPCN3) (HGK5) (HuKIII) (HLK3). | |||||
|
KCNA2_MOUSE
|
||||||
| NC score | 0.278932 (rank : 32) | θ value | 0.0252991 (rank : 35) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P63141, P15386, Q02010, Q8C8W4 | Gene names | Kcna2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (MK2). | |||||
|
KCNA3_MOUSE
|
||||||
| NC score | 0.278769 (rank : 33) | θ value | 0.0193708 (rank : 33) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P16390 | Gene names | Kcna3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (MK3). | |||||
|
KCNA4_HUMAN
|
||||||
| NC score | 0.278531 (rank : 34) | θ value | 0.0431538 (rank : 41) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
KCNA1_HUMAN
|
||||||
| NC score | 0.274498 (rank : 35) | θ value | 0.0330416 (rank : 38) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q09470 | Gene names | KCNA1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (HUKI) (HBK1). | |||||
|
KCNS3_HUMAN
|
||||||
| NC score | 0.274409 (rank : 36) | θ value | 0.0330416 (rank : 40) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9BQ31, O43651, Q96B56 | Gene names | KCNS3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 3 (Voltage-gated potassium channel subunit Kv9.3) (Delayed-rectifier K(+) channel alpha subunit 3). | |||||
|
KCNA1_MOUSE
|
||||||
| NC score | 0.274233 (rank : 37) | θ value | 0.0330416 (rank : 39) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P16388 | Gene names | Kcna1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (MKI) (MBK1). | |||||
|
KCNB1_MOUSE
|
||||||
| NC score | 0.273577 (rank : 38) | θ value | 0.0252991 (rank : 37) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q03717 | Gene names | Kcnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (mShab). | |||||
|
KCNB1_HUMAN
|
||||||
| NC score | 0.273534 (rank : 39) | θ value | 0.0252991 (rank : 36) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q14721, Q14193 | Gene names | KCNB1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (h-DRK1). | |||||
|
KCNA4_MOUSE
|
||||||
| NC score | 0.273401 (rank : 40) | θ value | 0.0961366 (rank : 50) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q61423 | Gene names | Kcna4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4). | |||||
|
KCNG3_HUMAN
|
||||||
| NC score | 0.267266 (rank : 41) | θ value | 0.0431538 (rank : 42) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8TAE7 | Gene names | KCNG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
KCNG3_MOUSE
|
||||||
| NC score | 0.267081 (rank : 42) | θ value | 0.0431538 (rank : 43) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P59053 | Gene names | Kcng3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
KCNG2_HUMAN
|
||||||
| NC score | 0.263524 (rank : 43) | θ value | 0.365318 (rank : 70) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9UJ96 | Gene names | KCNG2, KCNF2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 2 (Voltage-gated potassium channel subunit Kv6.2) (Cardiac potassium channel subunit). | |||||
|
KCNS1_HUMAN
|
||||||
| NC score | 0.260365 (rank : 44) | θ value | 0.125558 (rank : 53) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96KK3, O43652, Q6DJU6 | Gene names | KCNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
KCNC4_MOUSE
|
||||||
| NC score | 0.258786 (rank : 45) | θ value | 0.0736092 (rank : 49) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8R1C0 | Gene names | Kcnc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4). | |||||
|
KCNC4_HUMAN
|
||||||
| NC score | 0.258026 (rank : 46) | θ value | 0.0736092 (rank : 48) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q03721, Q3MIM4, Q5TBI6 | Gene names | KCNC4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4) (KSHIIIC). | |||||
|
KCNG1_HUMAN
|
||||||
| NC score | 0.257895 (rank : 47) | θ value | 0.21417 (rank : 64) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9UIX4, O43528, Q9BRC1 | Gene names | KCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 1 (Voltage-gated potassium channel subunit Kv6.1) (kH2). | |||||
|
KCNF1_HUMAN
|
||||||
| NC score | 0.253192 (rank : 48) | θ value | 1.06291 (rank : 86) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9H3M0, O43527 | Gene names | KCNF1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily F member 1 (Voltage-gated potassium channel subunit Kv5.1) (kH1). | |||||
|
KCNC3_MOUSE
|
||||||
| NC score | 0.251195 (rank : 49) | θ value | 0.21417 (rank : 63) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q63959, Q62088 | Gene names | Kcnc3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
KCNC1_MOUSE
|
||||||
| NC score | 0.251061 (rank : 50) | θ value | 0.163984 (rank : 58) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P15388 | Gene names | Kcnc1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
KCNC1_HUMAN
|
||||||
| NC score | 0.251003 (rank : 51) | θ value | 0.163984 (rank : 57) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P48547 | Gene names | KCNC1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
KCNS1_MOUSE
|
||||||
| NC score | 0.250247 (rank : 52) | θ value | 1.06291 (rank : 87) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O35173 | Gene names | Kcns1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
KCNG4_HUMAN
|
||||||
| NC score | 0.249934 (rank : 53) | θ value | 0.365318 (rank : 71) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8TDN1, Q96H24 | Gene names | KCNG4, KCNG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 4 (Voltage-gated potassium channel subunit Kv6.4). | |||||
|
KCNC3_HUMAN
|
||||||
| NC score | 0.248675 (rank : 54) | θ value | 0.365318 (rank : 69) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q14003 | Gene names | KCNC3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
KCNV2_HUMAN
|
||||||
| NC score | 0.248249 (rank : 55) | θ value | 0.813845 (rank : 79) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8TDN2 | Gene names | KCNV2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily V member 2 (Voltage-gated potassium channel subunit Kv8.2). | |||||
|
KCMA1_MOUSE
|
||||||
| NC score | 0.214846 (rank : 56) | θ value | 0.163984 (rank : 56) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCMA1_HUMAN
|
||||||
| NC score | 0.214460 (rank : 57) | θ value | 0.163984 (rank : 55) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
KCNK1_HUMAN
|
||||||
| NC score | 0.131099 (rank : 58) | θ value | 0.163984 (rank : 59) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O00180, Q13307 | Gene names | KCNK1, HOHO1, KCNO1, TWIK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1) (Potassium channel KCNO1). | |||||
|
KCNK6_HUMAN
|
||||||
| NC score | 0.130119 (rank : 59) | θ value | 0.365318 (rank : 72) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Y257, Q9HB47 | Gene names | KCNK6, TOSS, TWIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 6 (Inward rectifying potassium channel protein TWIK-2) (TWIK-originated similarity sequence). | |||||
|
KCNK1_MOUSE
|
||||||
| NC score | 0.116383 (rank : 60) | θ value | 0.47712 (rank : 73) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08581 | Gene names | Kcnk1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1). | |||||
|
KCNKF_HUMAN
|
||||||
| NC score | 0.110366 (rank : 61) | θ value | 8.99809 (rank : 164) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9H427, Q9HBC8 | Gene names | KCNK15, TASK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 15 (Acid-sensitive potassium channel protein TASK-5) (TWIK-related acid-sensitive K(+) channel 5) (Two pore potassium channel KT3.3). | |||||
|
KCNKA_HUMAN
|
||||||
| NC score | 0.109602 (rank : 62) | θ value | 6.88961 (rank : 145) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P57789, Q8TDK7, Q8TDK8, Q9HB59 | Gene names | KCNK10, TREK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 10 (Outward rectifying potassium channel protein TREK-2) (TREK-2 K(+) channel subunit). | |||||
|
KCNK5_HUMAN
|
||||||
| NC score | 0.109320 (rank : 63) | θ value | 5.27518 (rank : 130) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
KCNK9_HUMAN
|
||||||
| NC score | 0.104660 (rank : 64) | θ value | 1.81305 (rank : 98) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9NPC2, Q2M290, Q540F2 | Gene names | KCNK9, TASK3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3) (Two pore potassium channel KT3.2). | |||||
|
KCNK9_MOUSE
|
||||||
| NC score | 0.104262 (rank : 65) | θ value | 1.81305 (rank : 99) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q3LS21 | Gene names | Kcnk9, Task3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3). | |||||
|
KCNK3_HUMAN
|
||||||
| NC score | 0.092991 (rank : 66) | θ value | 8.99809 (rank : 162) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O14649 | Gene names | KCNK3, TASK, TASK1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Two pore potassium channel KT3.1). | |||||
|
KCNK3_MOUSE
|
||||||
| NC score | 0.092918 (rank : 67) | θ value | 8.99809 (rank : 163) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O35111, O35163 | Gene names | Kcnk3, Ctbak, Task, Task1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Cardiac two-pore background K(+) channel) (cTBAK-1) (Two pore potassium channel KT3.1). | |||||
|
KCNT1_HUMAN
|
||||||
| NC score | 0.092407 (rank : 68) | θ value | 3.0926 (rank : 106) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q5JUK3, Q9P2C5 | Gene names | KCNT1, KIAA1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1 (KCa4.1). | |||||
|
KCNKH_HUMAN
|
||||||
| NC score | 0.084719 (rank : 69) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q96T54, Q5TCF4, Q8TAW4, Q9BXD1, Q9H592 | Gene names | KCNK17, TALK2, TASK4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 17 (TWIK-related alkaline pH- activated K(+) channel 2) (2P domain potassium channel Talk-2) (TWIK- related acid-sensitive K(+) channel 4) (TASK-4). | |||||
|
KCNKG_HUMAN
|
||||||
| NC score | 0.082813 (rank : 70) | θ value | 6.88961 (rank : 147) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q96T55, Q5TCF3, Q9H591 | Gene names | KCNK16, TALK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 16 (TWIK-related alkaline pH- activated K(+) channel 1) (2P domain potassium channel Talk-1). | |||||
|
KCNK4_MOUSE
|
||||||
| NC score | 0.076817 (rank : 71) | θ value | 1.81305 (rank : 97) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
KCNK2_MOUSE
|
||||||
| NC score | 0.073286 (rank : 72) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P97438 | Gene names | Kcnk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (Two-pore potassium channel TPKC1) (TREK-1 K(+) channel subunit). | |||||
|
KCNK2_HUMAN
|
||||||
| NC score | 0.069076 (rank : 73) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O95069, Q9UNE3 | Gene names | KCNK2, TREK, TREK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (TREK-1 K(+) channel subunit) (Two-pore potassium channel TPKC1). | |||||
|
ZN750_MOUSE
|
||||||
| NC score | 0.067649 (rank : 74) | θ value | 0.00228821 (rank : 23) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BH05, Q66JP3, Q8C0L1 | Gene names | Znf750 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF750. | |||||
|
KCNKD_HUMAN
|
||||||
| NC score | 0.063607 (rank : 75) | θ value | 5.27518 (rank : 131) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9HB14, Q96E79 | Gene names | KCNK13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
EMSY_MOUSE
|
||||||
| NC score | 0.062440 (rank : 76) | θ value | 0.279714 (rank : 66) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BMB0, Q5FWK5, Q80XU1, Q8VDW9 | Gene names | Emsy | |||
|
Domain Architecture |

|
|||||
| Description | Protein EMSY. | |||||
|
KCNKD_MOUSE
|
||||||
| NC score | 0.056743 (rank : 77) | θ value | 6.88961 (rank : 146) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8R1P5 | Gene names | Kcnk13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCTD1_HUMAN
|
||||||
| NC score | 0.055976 (rank : 78) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q719H9 | Gene names | KCTD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD1. | |||||
|
KCTD1_MOUSE
|
||||||
| NC score | 0.055976 (rank : 79) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5M956, Q5EER9 | Gene names | Kctd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD1. | |||||
|
KCTD6_MOUSE
|
||||||
| NC score | 0.055383 (rank : 80) | θ value | θ > 10 (rank : 183) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BNL5, Q3TXX1 | Gene names | Kctd6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD6. | |||||
|
KCNK4_HUMAN
|
||||||
| NC score | 0.055083 (rank : 81) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NYG8, Q96T94 | Gene names | KCNK4, TRAAK | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK) (Two pore K(+) channel KT4.1). | |||||
|
KCTD6_HUMAN
|
||||||
| NC score | 0.054948 (rank : 82) | θ value | θ > 10 (rank : 182) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8NC69, Q8NBS6, Q8TCA6 | Gene names | KCTD6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD6. | |||||
|
KCD15_HUMAN
|
||||||
| NC score | 0.054783 (rank : 83) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96SI1, Q9BVI6 | Gene names | KCTD15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD15. | |||||
|
EMSY_HUMAN
|
||||||
| NC score | 0.053885 (rank : 84) | θ value | 0.125558 (rank : 52) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |

|
|||||
| Description | Protein EMSY. | |||||
|
KCD15_MOUSE
|
||||||
| NC score | 0.052431 (rank : 85) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8K0E1 | Gene names | Kctd15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD15. | |||||
|
KCTD4_MOUSE
|
||||||
| NC score | 0.050682 (rank : 86) | θ value | θ > 10 (rank : 181) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9D7X1, Q8CCQ3, Q9CYK4 | Gene names | Kctd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD4. | |||||
|
KCTD4_HUMAN
|
||||||
| NC score | 0.050457 (rank : 87) | θ value | θ > 10 (rank : 180) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WVF5 | Gene names | KCTD4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD4. | |||||
|
HGS_MOUSE
|
||||||
| NC score | 0.050349 (rank : 88) | θ value | 0.0736092 (rank : 47) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
HGS_HUMAN
|
||||||
| NC score | 0.048583 (rank : 89) | θ value | 0.163984 (rank : 54) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
AMOL1_MOUSE
|
||||||
| NC score | 0.042516 (rank : 90) | θ value | 0.0563607 (rank : 45) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
AMOL1_HUMAN
|
||||||
| NC score | 0.040977 (rank : 91) | θ value | 0.813845 (rank : 78) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8IY63, Q63HK7, Q8NDN0, Q8WXD1, Q96CM5 | Gene names | AMOTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
PRGC2_MOUSE
|
||||||
| NC score | 0.040514 (rank : 92) | θ value | 0.47712 (rank : 75) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
NFAT5_HUMAN
|
||||||
| NC score | 0.037447 (rank : 93) | θ value | 0.0563607 (rank : 46) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94916, O95693, Q9UN18 | Gene names | NFAT5, KIAA0827, TONEBP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells 5 (T-cell transcription factor NFAT5) (NF-AT5) (Tonicity-responsive enhancer-binding protein) (TonE- binding protein) (TonEBP). | |||||
|
NUP88_HUMAN
|
||||||
| NC score | 0.036273 (rank : 94) | θ value | 1.06291 (rank : 89) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q99567, Q9BWE5 | Gene names | NUP88 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear pore complex protein Nup88 (Nucleoporin Nup88) (88 kDa nuclear pore complex protein). | |||||
|
RPGR1_HUMAN
|
||||||
| NC score | 0.034510 (rank : 95) | θ value | 4.03905 (rank : 117) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q96KN7, Q7Z2W6, Q8IXV5, Q96QA8, Q9HB94, Q9HB95, Q9HBK6, Q9NR40 | Gene names | RPGRIP1 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
KCNH4_HUMAN
|
||||||
| NC score | 0.034258 (rank : 96) | θ value | 4.03905 (rank : 114) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UQ05 | Gene names | KCNH4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 4 (Voltage-gated potassium channel subunit Kv12.3) (Ether-a-go-go-like potassium channel 1) (ELK channel 1) (ELK1) (Brain-specific eag-like channel 2) (BEC2). | |||||
|
PHLB1_HUMAN
|
||||||
| NC score | 0.034241 (rank : 97) | θ value | 0.279714 (rank : 68) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q86UU1, O75133, Q4KMF8, Q8TEQ2 | Gene names | PHLDB1, DLNB07, KIAA0638, LL5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
HOME1_MOUSE
|
||||||
| NC score | 0.033679 (rank : 98) | θ value | 2.36792 (rank : 104) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 630 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9Z2Y3, Q8K3E1, Q8K4M8, Q9Z0E9, Q9Z216 | Gene names | Homer1, Vesl1 | |||
|
Domain Architecture |

|
|||||
| Description | Homer protein homolog 1 (VASP/Ena-related gene up-regulated during seizure and LTP) (Vesl-1). | |||||
|
EEA1_HUMAN
|
||||||
| NC score | 0.033019 (rank : 99) | θ value | 0.0193708 (rank : 31) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
GOGA1_MOUSE
|
||||||
| NC score | 0.032736 (rank : 100) | θ value | 0.279714 (rank : 67) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 994 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CW79, Q80YB0 | Gene names | Golga1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
AKAP9_HUMAN
|
||||||
| NC score | 0.032378 (rank : 101) | θ value | 0.21417 (rank : 61) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.030865 (rank : 102) | θ value | 1.06291 (rank : 88) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
GCC1_HUMAN
|
||||||
| NC score | 0.030383 (rank : 103) | θ value | 5.27518 (rank : 127) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96CN9, Q9H6N7 | Gene names | GCC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP and coiled-coil domain-containing protein 1 (Golgi coiled coil protein 1). | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.030211 (rank : 104) | θ value | 1.38821 (rank : 93) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
HOME1_HUMAN
|
||||||
| NC score | 0.029839 (rank : 105) | θ value | 4.03905 (rank : 113) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 564 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q86YM7, O96003, Q86YM5 | Gene names | HOMER1, SYN47 | |||
|
Domain Architecture |

|
|||||
| Description | Homer protein homolog 1. | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.029291 (rank : 106) | θ value | 0.163984 (rank : 60) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
NAF1_HUMAN
|
||||||
| NC score | 0.029275 (rank : 107) | θ value | 3.0926 (rank : 109) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q15025, O76008, Q96EL9, Q99833, Q9H1J3 | Gene names | TNIP1, KIAA0113, NAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (HIV-1 Nef-interacting protein) (Virion-associated nuclear shuttling protein) (VAN) (hVAN) (Nip40-1). | |||||
|
CENPH_HUMAN
|
||||||
| NC score | 0.028642 (rank : 108) | θ value | 6.88961 (rank : 140) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9H3R5 | Gene names | CENPH, ICEN35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein H (CENP-H) (Interphase centromere complex protein 35). | |||||
|
BAG3_HUMAN
|
||||||
| NC score | 0.027467 (rank : 109) | θ value | 0.21417 (rank : 62) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O95817, Q9NT20, Q9P120 | Gene names | BAG3, BIS | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis) (Docking protein CAIR-1). | |||||
|
SYNE2_HUMAN
|
||||||
| NC score | 0.027221 (rank : 110) | θ value | 0.47712 (rank : 76) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
NAF1_MOUSE
|
||||||
| NC score | 0.026880 (rank : 111) | θ value | 5.27518 (rank : 133) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 852 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9WUU8, Q922A9, Q922F7, Q9EPP8, Q9R0X3 | Gene names | Tnip1, Abin, Naf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (A20- binding inhibitor of NF-kappa-B activation) (ABIN) (Virion-associated nuclear shuttling protein) (VAN) (mVAN). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.026168 (rank : 112) | θ value | 0.813845 (rank : 80) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
DESP_HUMAN
|
||||||
| NC score | 0.025735 (rank : 113) | θ value | 3.0926 (rank : 105) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
RBP24_HUMAN
|
||||||
| NC score | 0.025229 (rank : 114) | θ value | 5.27518 (rank : 134) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
GCC2_HUMAN
|
||||||
| NC score | 0.025192 (rank : 115) | θ value | 5.27518 (rank : 128) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
TARSH_HUMAN
|
||||||
| NC score | 0.025052 (rank : 116) | θ value | 0.813845 (rank : 81) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
GOGA1_HUMAN
|
||||||
| NC score | 0.024529 (rank : 117) | θ value | 4.03905 (rank : 111) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
CF060_HUMAN
|
||||||
| NC score | 0.024280 (rank : 118) | θ value | 5.27518 (rank : 125) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1270 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8NB25, Q96GY8, Q9H851 | Gene names | C6orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf60. | |||||
|
CCD46_HUMAN
|
||||||
| NC score | 0.024160 (rank : 119) | θ value | 5.27518 (rank : 124) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
CP250_MOUSE
|
||||||
| NC score | 0.023964 (rank : 120) | θ value | 4.03905 (rank : 110) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
GOGA3_MOUSE
|
||||||
| NC score | 0.023928 (rank : 121) | θ value | 4.03905 (rank : 112) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1437 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P55937, Q80VF5, Q8CCK4, Q9QYT2, Q9QYT3 | Gene names | Golga3, Mea2 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Male-enhanced antigen 2) (MEA-2). | |||||
|
UACA_HUMAN
|
||||||
| NC score | 0.023696 (rank : 122) | θ value | 0.0961366 (rank : 51) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1622 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9BZF9, Q8N3B8, Q9HCL1, Q9NWC6 | Gene names | UACA, KIAA1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein. | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.023257 (rank : 123) | θ value | 8.99809 (rank : 155) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
PRGC2_HUMAN
|
||||||
| NC score | 0.023159 (rank : 124) | θ value | 8.99809 (rank : 168) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86YN6, Q86YN3, Q86YN4, Q86YN5, Q8N1N9, Q8TDE4, Q8TDE5 | Gene names | PPARGC1B, PERC, PGC1, PGC1B, PPARGC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta) (PGC-1-related estrogen receptor alpha coactivator) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
PHF3_HUMAN
|
||||||
| NC score | 0.022691 (rank : 125) | θ value | 1.06291 (rank : 90) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.022256 (rank : 126) | θ value | 2.36792 (rank : 103) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
MYO5A_MOUSE
|
||||||
| NC score | 0.022250 (rank : 127) | θ value | 0.21417 (rank : 65) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1117 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q99104 | Gene names | Myo5a, Dilute | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle). | |||||
|
CI093_HUMAN
|
||||||
| NC score | 0.021829 (rank : 128) | θ value | 6.88961 (rank : 141) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6TFL3, Q5SU58, Q6P1W1, Q6ZR13, Q8N1Z4, Q8N8L3, Q8TBR2 | Gene names | C9orf93 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf93. | |||||
|
R3HD2_MOUSE
|
||||||
| NC score | 0.021559 (rank : 129) | θ value | 8.99809 (rank : 169) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
SYCP1_MOUSE
|
||||||
| NC score | 0.021230 (rank : 130) | θ value | 5.27518 (rank : 136) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1456 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q62209, O09205, P70192, Q62329 | Gene names | Sycp1, Scp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
IFT81_HUMAN
|
||||||
| NC score | 0.020979 (rank : 131) | θ value | 8.99809 (rank : 160) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8WYA0, Q8NB51, Q9BSV2, Q9UNY8 | Gene names | IFT81, CDV1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 81 (Carnitine deficiency-associated protein expressed in ventricle 1) (CDV-1 protein). | |||||
|
REST_MOUSE
|
||||||
| NC score | 0.020624 (rank : 132) | θ value | 5.27518 (rank : 135) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
LMNB1_HUMAN
|
||||||
| NC score | 0.020416 (rank : 133) | θ value | 3.0926 (rank : 108) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P20700, Q3SYN7, Q96EI6 | Gene names | LMNB1, LMN2, LMNB | |||
|
Domain Architecture |

|
|||||
| Description | Lamin-B1. | |||||
|
HCN1_MOUSE
|
||||||
| NC score | 0.020338 (rank : 134) | θ value | 1.06291 (rank : 84) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O88704, O54899, Q9D613 | Gene names | Hcn1, Bcng1, Hac2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1) (Hyperpolarization-activated cation channel 2) (HAC-2). | |||||
|
LMNB1_MOUSE
|
||||||
| NC score | 0.020093 (rank : 135) | θ value | 4.03905 (rank : 115) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 719 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P14733, Q61791 | Gene names | Lmnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Lamin-B1. | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.019621 (rank : 136) | θ value | 8.99809 (rank : 159) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
C8AP2_MOUSE
|
||||||
| NC score | 0.019538 (rank : 137) | θ value | 5.27518 (rank : 123) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
MYO5A_HUMAN
|
||||||
| NC score | 0.019345 (rank : 138) | θ value | 1.38821 (rank : 94) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y4I1, O60653, Q07902, Q16249, Q9UE30, Q9UE31 | Gene names | MYO5A, MYH12 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle) (Myosin heavy chain 12) (Myoxin). | |||||
|
LIPS_HUMAN
|
||||||
| NC score | 0.019091 (rank : 139) | θ value | 5.27518 (rank : 132) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q05469, Q3LRT2, Q6NSL7 | Gene names | LIPE | |||
|
Domain Architecture |

|
|||||
| Description | Hormone-sensitive lipase (EC 3.1.1.79) (HSL). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.018069 (rank : 140) | θ value | 8.99809 (rank : 156) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
HCN1_HUMAN
|
||||||
| NC score | 0.017732 (rank : 141) | θ value | 1.06291 (rank : 83) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O60741 | Gene names | HCN1, BCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1). | |||||
|
DYH11_HUMAN
|
||||||
| NC score | 0.016753 (rank : 142) | θ value | 1.81305 (rank : 96) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96DT5, Q9UJ82 | Gene names | DNAH11 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 11 (Axonemal beta dynein heavy chain 11). | |||||
|
ETV5_MOUSE
|
||||||
| NC score | 0.016480 (rank : 143) | θ value | 0.62314 (rank : 77) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
ETV5_HUMAN
|
||||||
| NC score | 0.016020 (rank : 144) | θ value | 1.06291 (rank : 82) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
LAMA4_HUMAN
|
||||||
| NC score | 0.015836 (rank : 145) | θ value | 3.0926 (rank : 107) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q16363, Q14735, Q15335, Q4LE44, Q5SZG8, Q9UE18, Q9UJN9 | Gene names | LAMA4 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-4 chain precursor. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.015591 (rank : 146) | θ value | 4.03905 (rank : 119) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TCF7_HUMAN
|
||||||
| NC score | 0.015388 (rank : 147) | θ value | 1.81305 (rank : 101) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P36402, Q9UKI4 | Gene names | TCF7, TCF1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
SAPS1_HUMAN
|
||||||
| NC score | 0.014675 (rank : 148) | θ value | 1.06291 (rank : 91) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
K1914_HUMAN
|
||||||
| NC score | 0.014282 (rank : 149) | θ value | 8.99809 (rank : 161) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N4X5, Q8TB54, Q96PX4, Q96SY5 | Gene names | KIAA1914 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1914. | |||||
|
DTNA_HUMAN
|
||||||
| NC score | 0.014135 (rank : 150) | θ value | 1.81305 (rank : 95) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Y4J8, O15332, O15333, O75697, Q13197, Q13198, Q13199, Q13498, Q13499, Q13500 | Gene names | DTNA, DRP3 | |||
|
Domain Architecture |

|
|||||
| Description | Dystrobrevin alpha (Dystrobrevin-alpha) (Dystrophin-related protein 3). | |||||
|
ATX2_MOUSE
|
||||||
| NC score | 0.013867 (rank : 151) | θ value | 6.88961 (rank : 139) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O70305, P97421 | Gene names | Atxn2, Atx2, Sca2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein homolog). | |||||
|
MTSS1_HUMAN
|
||||||
| NC score | 0.013575 (rank : 152) | θ value | 8.99809 (rank : 167) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O43312, Q8TCA2, Q96RX2 | Gene names | MTSS1, KIAA0429, MIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein) (Metastasis suppressor YGL-1). | |||||
|
LAMA2_HUMAN
|
||||||
| NC score | 0.013190 (rank : 153) | θ value | 1.81305 (rank : 100) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
CUTL2_HUMAN
|
||||||
| NC score | 0.013112 (rank : 154) | θ value | 6.88961 (rank : 142) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O14529 | Gene names | CUTL2, KIAA0293 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 2 (Homeobox protein Cux-2) (Cut-like 2). | |||||
|
LRSM1_MOUSE
|
||||||
| NC score | 0.013090 (rank : 155) | θ value | 6.88961 (rank : 148) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q80ZI6 | Gene names | Lrsam1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.013033 (rank : 156) | θ value | 4.03905 (rank : 116) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
TRI66_MOUSE
|
||||||
| NC score | 0.012987 (rank : 157) | θ value | 5.27518 (rank : 138) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
BCAR1_HUMAN
|
||||||
| NC score | 0.012147 (rank : 158) | θ value | 8.99809 (rank : 154) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
FRAT2_HUMAN
|
||||||
| NC score | 0.011207 (rank : 159) | θ value | 5.27518 (rank : 126) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75474, Q5JTI0, Q8WUL9, Q9BYG2 | Gene names | FRAT2 | |||
|
Domain Architecture |

|
|||||
| Description | GSK-3-binding protein FRAT2 (Frat-2). | |||||
|
ASPP2_HUMAN
|
||||||
| NC score | 0.010908 (rank : 160) | θ value | 8.99809 (rank : 153) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q13625, Q12892, Q86X75, Q96KQ3 | Gene names | TP53BP2, ASPP2, BBP | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 2 (Tumor suppressor p53-binding protein 2) (p53-binding protein 2) (p53BP2) (53BP2) (Bcl2-binding protein) (Bbp) (NY-REN-51 antigen). | |||||
|
RFFL_HUMAN
|
||||||
| NC score | 0.010471 (rank : 161) | θ value | 8.99809 (rank : 170) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8WZ73, Q8NHW0, Q8TBY7, Q96BE6 | Gene names | RFFL, RNF189, RNF34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (FYVE-RING finger protein Sakura) (Fring) (Caspases-8 and -10- associated RING finger protein 2) (CARP-2) (Caspase regulator CARP2) (RING finger protein 189) (RING finger protein 34-like). | |||||
|
GA2L2_MOUSE
|
||||||
| NC score | 0.009402 (rank : 162) | θ value | 8.99809 (rank : 158) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5SSG4 | Gene names | Gas2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2). | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.009215 (rank : 163) | θ value | 8.99809 (rank : 157) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
MAVS_HUMAN
|
||||||
| NC score | 0.008618 (rank : 164) | θ value | 8.99809 (rank : 166) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7Z434, Q3I0Y2, Q5T7I6, Q86VY7, Q9H1H3, Q9H4Y1, Q9H8D3, Q9ULE9 | Gene names | MAVS, IPS1, KIAA1271, VISA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif) (Putative NF-kappa-B- activating protein 031N). | |||||
|
LCP2_HUMAN
|
||||||
| NC score | 0.008222 (rank : 165) | θ value | 8.99809 (rank : 165) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13094 | Gene names | LCP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte cytosolic protein 2 (SH2 domain-containing leukocyte protein of 76 kDa) (SLP-76 tyrosine phosphoprotein) (SLP76). | |||||
|
MA1B1_HUMAN
|
||||||
| NC score | 0.006938 (rank : 166) | θ value | 6.88961 (rank : 149) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UKM7, Q5VSG3, Q9BRS9, Q9Y5K7 | Gene names | MAN1B1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase (EC 3.2.1.113) (ER alpha-1,2-mannosidase) (Mannosidase alpha class 1B member 1) (Man9GlcNAc2-specific-processing alpha-mannosidase). | |||||
|
ANGL6_HUMAN
|
||||||
| NC score | 0.006875 (rank : 167) | θ value | 5.27518 (rank : 122) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NI99, Q9BZZ0 | Gene names | ANGPTL6, AGF, ARP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiopoietin-related protein 6 precursor (Angiopoietin-like 6) (Angiopoietin-related growth factor) (Angiopoietin-related protein 5). | |||||
|
IRF3_HUMAN
|
||||||
| NC score | 0.006756 (rank : 168) | θ value | 6.88961 (rank : 144) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14653 | Gene names | IRF3 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 3 (IRF-3). | |||||
|
NF2L2_MOUSE
|
||||||
| NC score | 0.006570 (rank : 169) | θ value | 6.88961 (rank : 150) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60795 | Gene names | Nfe2l2, Nrf-2, Nrf2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor erythroid 2-related factor 2 (NF-E2-related factor 2) (NFE2-related factor 2) (Nuclear factor, erythroid derived 2, like 2). | |||||
|
SNTB1_MOUSE
|
||||||
| NC score | 0.005778 (rank : 170) | θ value | 4.03905 (rank : 118) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99L88, O35925 | Gene names | Sntb1, Snt2b1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1-syntrophin (59 kDa dystrophin-associated protein A1 basic component 1) (DAPA1B) (Syntrophin 2). | |||||
|
RP3A_MOUSE
|
||||||
| NC score | 0.004609 (rank : 171) | θ value | 6.88961 (rank : 151) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P47708 | Gene names | Rph3a | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
INVS_MOUSE
|
||||||
| NC score | 0.004394 (rank : 172) | θ value | 5.27518 (rank : 129) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
HIPK1_MOUSE
|
||||||
| NC score | 0.003599 (rank : 173) | θ value | 1.06291 (rank : 85) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 838 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O88904, Q80TV5, Q9QUQ8, Q9QZR3, Q9WVN7 | Gene names | Hipk1, Kiaa0630, Myak, Nbak2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeodomain-interacting protein kinase 1 (EC 2.7.11.1) (Protein kinase Myak) (Nuclear body-associated kinase 2). | |||||
|
SHC1_HUMAN
|
||||||
| NC score | 0.003323 (rank : 174) | θ value | 8.99809 (rank : 171) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P29353, O15290, Q8N4K5, Q96CL1 | Gene names | SHC1, SHC, SHCA | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 1 (SH2 domain protein C1) (Src homology 2 domain-containing-transforming protein C1). | |||||
|
TNIK_HUMAN
|
||||||
| NC score | 0.002919 (rank : 175) | θ value | 5.27518 (rank : 137) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | 0.002344 (rank : 176) | θ value | 0.47712 (rank : 74) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
SN1L2_HUMAN
|
||||||
| NC score | 0.001853 (rank : 177) | θ value | 1.06291 (rank : 92) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H0K1, O94878, Q6AZE2, Q76N03, Q8NCV7, Q96CZ8 | Gene names | SNF1LK2, KIAA0781, QIK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Qin- induced kinase). | |||||
|
HIPK1_HUMAN
|
||||||
| NC score | 0.001423 (rank : 178) | θ value | 6.88961 (rank : 143) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86Z02, O75125, Q8IYD7, Q8NDN5, Q8NEB6, Q8TBZ1 | Gene names | HIPK1, KIAA0630 | |||
|
Domain Architecture |

|
|||||
| Description | Homeodomain-interacting protein kinase 1 (EC 2.7.11.1). | |||||
|
ZN447_HUMAN
|
||||||
| NC score | 0.001213 (rank : 179) | θ value | 1.81305 (rank : 102) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TBC5, Q9BRK7, Q9H9A0 | Gene names | ZNF447 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 447. | |||||
|
ZN462_HUMAN
|
||||||
| NC score | 0.000325 (rank : 180) | θ value | 4.03905 (rank : 121) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 740 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96JM2, Q8N408 | Gene names | ZNF462, KIAA1803 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 462. | |||||
|
ZBT24_MOUSE
|
||||||
| NC score | 0.000214 (rank : 181) | θ value | 0.0431538 (rank : 44) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 894 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80X44, Q3UIB7, Q7TPC4, Q8CJC9 | Gene names | Zbtb24, Bif1, Bsg1, Znf450 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 24 (Zinc finger protein 450) (Bone morphogenetic protein-induced factor 1) (Brain-specific protein 1). | |||||
|
ZBTB4_HUMAN
|
||||||
| NC score | -0.000449 (rank : 182) | θ value | 6.88961 (rank : 152) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 926 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P1Z0, Q7Z697, Q86XJ4, Q8N4V8 | Gene names | ZBTB4, KIAA1538 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 4 (KAISO-like zinc finger protein 1) (KAISO-L1). | |||||
|
ZN366_HUMAN
|
||||||
| NC score | -0.001929 (rank : 183) | θ value | 4.03905 (rank : 120) | |||
| Query Neighborhood Hits | 171 | Target Neighborhood Hits | 732 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N895, Q7RTV4 | Gene names | ZNF366 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 366. | |||||