Please be patient as the page loads
|
HD_MOUSE
|
||||||
| SwissProt Accessions | P42859 | Gene names | Hd, Hdh | |||
|
Domain Architecture |
|
|||||
| Description | Huntingtin (Huntington disease protein homolog) (HD protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HD_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.977056 (rank : 2) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
HD_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 116 | |
| SwissProt Accessions | P42859 | Gene names | Hd, Hdh | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein homolog) (HD protein). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 3) | NC score | 0.099636 (rank : 4) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 4) | NC score | 0.113414 (rank : 3) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
MKL1_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 5) | NC score | 0.065362 (rank : 8) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
TSC2_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 6) | NC score | 0.054327 (rank : 15) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61037, P97723, P97724, P97725, P97727, Q61007, Q61008, Q9WUF6 | Gene names | Tsc2 | |||
|
Domain Architecture |

|
|||||
| Description | Tuberin (Tuberous sclerosis 2 homolog protein). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 7) | NC score | 0.050168 (rank : 19) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
2AAA_MOUSE
|
||||||
| θ value | 0.163984 (rank : 8) | NC score | 0.075300 (rank : 5) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q76MZ3 | Gene names | Ppp2r1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform (PP2A, subunit A, PR65-alpha isoform) (PP2A, subunit A, R1-alpha isoform). | |||||
|
AKAP6_HUMAN
|
||||||
| θ value | 0.279714 (rank : 9) | NC score | 0.046316 (rank : 27) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13023, O15028 | Gene names | AKAP6, AKAP100, KIAA0311 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 6 (Protein kinase A-anchoring protein 6) (PRKA6) (A-kinase anchor protein 100 kDa) (AKAP 100) (mAKAP). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.279714 (rank : 10) | NC score | 0.043549 (rank : 33) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
2AAA_HUMAN
|
||||||
| θ value | 0.365318 (rank : 11) | NC score | 0.068315 (rank : 7) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P30153, Q13773 | Gene names | PPP2R1A | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform (PP2A, subunit A, PR65-alpha isoform) (PP2A, subunit A, R1-alpha isoform) (Medium tumor antigen-associated 61 kDa protein). | |||||
|
AN32B_MOUSE
|
||||||
| θ value | 0.365318 (rank : 12) | NC score | 0.038636 (rank : 41) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9EST5 | Gene names | Anp32b, Pal31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member B (Proliferation-related acidic leucine-rich protein PAL31). | |||||
|
JPH2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 13) | NC score | 0.039917 (rank : 37) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
SRCA_HUMAN
|
||||||
| θ value | 0.365318 (rank : 14) | NC score | 0.034207 (rank : 49) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
GATA6_MOUSE
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.049329 (rank : 21) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61169, P97729, Q9QZK3 | Gene names | Gata6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor GATA-6 (GATA-binding factor 6). | |||||
|
DUET_HUMAN
|
||||||
| θ value | 0.813845 (rank : 16) | NC score | 0.007714 (rank : 108) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1220 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2A5, Q8TBQ5, Q9NSZ4 | Gene names | DUET, TRAD | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Duet (EC 2.7.11.1) (Serine/threonine kinase with Dbl- and pleckstrin homology domain). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 17) | NC score | 0.060416 (rank : 12) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
LRC47_MOUSE
|
||||||
| θ value | 0.813845 (rank : 18) | NC score | 0.035166 (rank : 46) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q505F5, Q3U380, Q6ZPW2, Q9CUZ0 | Gene names | Lrrc47, Kiaa1185 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 47. | |||||
|
SF3B1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 19) | NC score | 0.062822 (rank : 10) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75533 | Gene names | SF3B1, SAP155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
SF3B1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 20) | NC score | 0.062753 (rank : 11) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99NB9, Q9CSK5 | Gene names | Sf3b1, Sap155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
TCF7_MOUSE
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.034246 (rank : 48) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q00417 | Gene names | Tcf7, Tcf-1, Tcf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
GAK5_HUMAN
|
||||||
| θ value | 1.06291 (rank : 22) | NC score | 0.050179 (rank : 18) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
PPRB_MOUSE
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.063524 (rank : 9) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
PSL1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.054200 (rank : 16) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
TOPRS_HUMAN
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.037388 (rank : 42) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NS56, O43273, Q6P987, Q9NS55, Q9UNR9 | Gene names | TOPORS, LUN, TP53BPL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
ATX7_MOUSE
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.048272 (rank : 22) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8R4I1, Q8BL17 | Gene names | Atxn7, Sca7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-7 (Spinocerebellar ataxia type 7 protein homolog). | |||||
|
CD2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.075044 (rank : 6) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
GATA6_HUMAN
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.039921 (rank : 36) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92908, P78327 | Gene names | GATA6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor GATA-6 (GATA-binding factor 6). | |||||
|
ILDR1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 29) | NC score | 0.032383 (rank : 52) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86SU0, Q6ZP61, Q7Z578 | Gene names | ILDR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immunoglobulin-like domain-containing receptor 1 precursor. | |||||
|
AMBN_HUMAN
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.043800 (rank : 32) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NP70, Q9H2X1, Q9H4L1 | Gene names | AMBN | |||
|
Domain Architecture |
|
|||||
| Description | Ameloblastin precursor. | |||||
|
BTBDC_HUMAN
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.034661 (rank : 47) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IY92, Q96JP1 | Gene names | BTBD12, KIAA1784 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 12. | |||||
|
ITA4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.011966 (rank : 99) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P13612 | Gene names | ITGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-4 precursor (Integrin alpha-IV) (VLA-4) (CD49d antigen). | |||||
|
NFAT5_MOUSE
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.020315 (rank : 81) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WV30 | Gene names | Nfat5 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells 5 (T cell transcription factor NFAT5) (NF-AT5) (Rel domain-containing transcription factor NFAT5). | |||||
|
PPRB_HUMAN
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.044893 (rank : 31) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
CC021_MOUSE
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.045552 (rank : 30) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3U4G3, Q3TLX5, Q8K2I0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C3orf21 homolog. | |||||
|
GCN1L_HUMAN
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.053938 (rank : 17) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92616, O95001, O95651, Q6P2S3, Q86X65, Q8N5I5, Q8WU80, Q99736, Q9UE60 | Gene names | GCN1L1, KIAA0219 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GCN1-like protein 1 (HsGCN1). | |||||
|
IRF3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.023956 (rank : 72) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14653 | Gene names | IRF3 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 3 (IRF-3). | |||||
|
KCNH8_HUMAN
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.011008 (rank : 102) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96L42 | Gene names | KCNH8 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 8 (Voltage-gated potassium channel subunit Kv12.1) (Ether-a-go-go-like potassium channel 3) (ELK channel 3) (ELK3) (ELK1) (hElk1). | |||||
|
M4K1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.002527 (rank : 117) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 858 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70218, Q8R3R0, Q99K62 | Gene names | Map4k1, Hpk1 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase kinase 1 (EC 2.7.11.1) (MAPK/ERK kinase kinase kinase 1) (MEK kinase kinase 1) (MEKKK 1) (Hematopoietic progenitor kinase) (HPK). | |||||
|
MOCOS_HUMAN
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.025682 (rank : 66) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96EN8, Q53GP5, Q8N3A4, Q9NWM7 | Gene names | MOCOS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Molybdenum cofactor sulfurase (EC 4.4.-.-) (MoCo sulfurase) (MOS) (HMCS). | |||||
|
PF21B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.021416 (rank : 77) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
TOPRS_MOUSE
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.038940 (rank : 39) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80Z37, Q3U5B5, Q3UPZ1, Q3USN3, Q3UZ55, Q8BXP2, Q8CFF5, Q8CGC8, Q920L3 | Gene names | Topors | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 2.36792 (rank : 43) | NC score | 0.006570 (rank : 111) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
ZCH10_MOUSE
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.049413 (rank : 20) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CX48 | Gene names | Zcchc10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 10. | |||||
|
4ET_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.036893 (rank : 44) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
CRSP7_HUMAN
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.027870 (rank : 59) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
DZIP3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.017131 (rank : 90) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7TPV2, Q80TU4, Q8BYK7 | Gene names | Dzip3, Kiaa0675 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) (DAZ-interacting protein 3 homolog). | |||||
|
GAK11_HUMAN
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.047207 (rank : 25) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK12_HUMAN
|
||||||
| θ value | 3.0926 (rank : 49) | NC score | 0.047493 (rank : 23) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 50) | NC score | 0.040558 (rank : 35) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P62683 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 51) | NC score | 0.047038 (rank : 26) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.047271 (rank : 24) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.046276 (rank : 28) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
PEX14_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.029314 (rank : 56) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75381, Q8WX51 | Gene names | PEX14 | |||
|
Domain Architecture |
|
|||||
| Description | Peroxisomal membrane protein PEX14 (Peroxin-14) (Peroxisomal membrane anchor protein PEX14) (PTS1 receptor docking protein). | |||||
|
PHF14_MOUSE
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.018835 (rank : 85) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.024108 (rank : 71) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
TARSH_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.038661 (rank : 40) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
TF65_HUMAN
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.017586 (rank : 89) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q04206, Q6SLK1 | Gene names | RELA, NFKB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor p65 (Nuclear factor NF-kappa-B p65 subunit). | |||||
|
TSKS_MOUSE
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.022790 (rank : 75) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54887 | Gene names | Stk22s1, Tsks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-specific serine kinase substrate (Testis-specific kinase substrate) (STK22 substrate 1). | |||||
|
CD3E_MOUSE
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.028809 (rank : 57) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22646 | Gene names | Cd3e | |||
|
Domain Architecture |
|
|||||
| Description | T-cell surface glycoprotein CD3 epsilon chain precursor (T-cell surface antigen T3/Leu-4 epsilon chain). | |||||
|
CO039_HUMAN
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.026348 (rank : 62) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6ZRI6, Q71JB1, Q7L3S0, Q8N3F2, Q96FB6, Q9NTU5 | Gene names | C15orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39. | |||||
|
COFA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.014275 (rank : 96) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P39059, Q5T6J4, Q9Y4W4 | Gene names | COL15A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV) (Restin) (Related to endostatin)]. | |||||
|
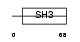
CT032_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.025794 (rank : 65) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQ75, Q96K09, Q9BYL5 | Gene names | HEFL, C20orf32 | |||
|
Domain Architecture |
|
|||||
| Description | HEF-like protein. | |||||
|
DOHH_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.024895 (rank : 69) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BU89, O75265 | Gene names | DOHH, HLRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deoxyhypusine hydroxylase (EC 1.14.99.29) (Deoxyhypusine monooxygenase) (hDOHH) (HEAT-like repeat-containing protein 1). | |||||
|
FBX18_HUMAN
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.027189 (rank : 61) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NFZ0, Q7Z4Q6, Q7Z4R0, Q8N1P5, Q8N586, Q96E82, Q96K67, Q96SW7, Q9UFB2 | Gene names | FBXO18, FBH1, FBX18 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 18 (EC 3.6.1.-) (F-box DNA helicase 1). | |||||
|
MYO15_HUMAN
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.008533 (rank : 105) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
OAS3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.016435 (rank : 93) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y6K5, Q9H3P5 | Gene names | OAS3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 2'-5'-oligoadenylate synthetase 3 (EC 2.7.7.-) ((2-5')oligo(A) synthetase 3) (2-5A synthetase 3) (p100 OAS) (p100OAS). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.021136 (rank : 79) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
RPGF5_MOUSE
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.012307 (rank : 98) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C0Q9, Q8BJJ9, Q8C0R5 | Gene names | Rapgef5, Gfr, Kiaa0277, Mrgef | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 5 (Guanine nucleotide exchange factor for Rap1) (M-Ras-regulated Rap GEF) (MR-GEF). | |||||
|
SL9A8_MOUSE
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.025912 (rank : 64) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R4D1, Q3UPR4, Q5WA59, Q8BIH8, Q8BJ27 | Gene names | Slc9a8, Nhe8 | |||
|
Domain Architecture |
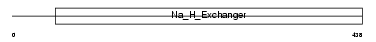
|
|||||
| Description | Sodium/hydrogen exchanger 8 (Na(+)/H(+) exchanger 8) (NHE-8) (Solute carrier family 9 member 8). | |||||
|
SPR1A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.045868 (rank : 29) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62266 | Gene names | Sprr1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cornifin A (Small proline-rich protein 1A) (SPR1A) (SPR1 A). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.036430 (rank : 45) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.021754 (rank : 76) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
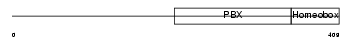
TFE2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.014416 (rank : 95) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P15923, P15883, Q14208, Q14635, Q14636, Q9UPI9 | Gene names | TCF3, E2A, ITF1 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Immunoglobulin transcription factor 1) (Transcription factor ITF-1) (Kappa-E2-binding factor). | |||||
|
ADA12_MOUSE
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.005244 (rank : 115) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61824 | Gene names | Adam12, Mltna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
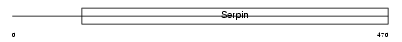
ANGT_MOUSE
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.011307 (rank : 101) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P11859 | Gene names | Agt, Serpina8 | |||
|
Domain Architecture |
|
|||||
| Description | Angiotensinogen precursor [Contains: Angiotensin-1 (Angiotensin I) (Ang I); Angiotensin-2 (Angiotensin II) (Ang II); Angiotensin-3 (Angiotensin III) (Ang III) (Des-Asp[1]-angiotensin II)]. | |||||
|
AUTS2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.039052 (rank : 38) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WXX7, Q9Y4F2 | Gene names | AUTS2, KIAA0442 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Autism susceptibility gene 2 protein. | |||||
|
CABIN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.029565 (rank : 55) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
CAND2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.021005 (rank : 80) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZQ73, Q6PHT7 | Gene names | Cand2, Kiaa0667, Tip120b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cullin-associated NEDD8-dissociated protein 2 (Cullin-associated and neddylation-dissociated protein 2) (p120 CAND2) (TBP-interacting protein TIP120B) (TBP-interacting protein of 120 kDa B). | |||||
|
FOXN4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.018585 (rank : 87) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96NZ1, Q6ZMR4, Q96NZ0 | Gene names | FOXN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein N4. | |||||
|
HGS_HUMAN
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.030253 (rank : 53) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.028263 (rank : 58) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MTF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | 0.000500 (rank : 118) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 814 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14872, Q96CB1 | Gene names | MTF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
NUFP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.030110 (rank : 54) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UHK0, Q8WVM5, Q96SG1 | Gene names | NUFIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 1 (Nuclear FMRP-interacting protein 1). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.023680 (rank : 74) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PIGG_HUMAN
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | 0.019189 (rank : 83) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5H8A4, Q2TAK5, Q6UX31, Q7L5Y4, Q8N866, Q8NCC9, Q96SY9, Q9BVT7, Q9NXG5 | Gene names | PIGG, GPI7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI ethanolamine phosphate transferase 2 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class G protein) (PIG-G) (GPI7 homolog) (hGPI7). | |||||
|
SL9A8_HUMAN
|
||||||
| θ value | 5.27518 (rank : 87) | NC score | 0.023724 (rank : 73) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2E8, Q68CZ8, Q9BX15, Q9Y507 | Gene names | SLC9A8, KIAA0939, NHE8 | |||
|
Domain Architecture |
|
|||||
| Description | Sodium/hydrogen exchanger 8 (Na(+)/H(+) exchanger 8) (NHE-8) (Solute carrier family 9 member 8). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | 0.007174 (rank : 110) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
SPR1B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.034065 (rank : 50) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62267 | Gene names | Sprr1b | |||
|
Domain Architecture |

|
|||||
| Description | Cornifin B (Small proline-rich protein 1B) (SPR1B) (SPR1 B). | |||||
|
TPO_MOUSE
|
||||||
| θ value | 5.27518 (rank : 90) | NC score | 0.032998 (rank : 51) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P40226 | Gene names | Thpo | |||
|
Domain Architecture |

|
|||||
| Description | Thrombopoietin precursor (Megakaryocyte colony-stimulating factor) (Myeloproliferative leukemia virus oncogene ligand) (C-mpl ligand) (ML) (Megakaryocyte growth and development factor) (MGDF). | |||||
|
AKAP2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.019278 (rank : 82) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
BAG3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.017681 (rank : 88) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
CEL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.008432 (rank : 106) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
CJ026_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.037253 (rank : 43) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NX94, Q2HIY7, Q5F2G6 | Gene names | C10orf26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C10orf26. | |||||
|
DTX1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.024346 (rank : 70) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86Y01, O60630, Q9BS04 | Gene names | DTX1 | |||
|
Domain Architecture |
|
|||||
| Description | Protein deltex-1 (Deltex-1) (Deltex1) (hDTX1). | |||||
|
DTX1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.025599 (rank : 67) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61010, Q3TER5, Q8C2H2, Q9ER09 | Gene names | Dtx1 | |||
|
Domain Architecture |
|
|||||
| Description | Protein deltex-1 (Deltex-1) (Deltex1) (mDTX1) (FXI-T1). | |||||
|
ETV5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.012641 (rank : 97) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
RFXK_HUMAN
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.004941 (rank : 116) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O14593, O95839 | Gene names | RFXANK, ANKRA1, RFXB | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein RFXANK (Regulatory factor X subunit B) (RFX-B) (Ankyrin repeat family A protein 1). | |||||
|
SMG1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.018928 (rank : 84) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96Q15, O43305, Q13284, Q8NFX2, Q96QV0, Q96RW3 | Gene names | SMG1, ATX, KIAA0421, LIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1) (hSMG-1) (Lambda/iota protein kinase C-interacting protein) (Lambda-interacting protein) (61E3.4). | |||||
|
SMG1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | 0.018824 (rank : 86) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BKX6, Q5DU42, Q6ZQC0, Q8BLU4, Q8BWJ5, Q8BXD3 | Gene names | Smg1, Kiaa0421 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1). | |||||
|
SON_MOUSE
|
||||||
| θ value | 6.88961 (rank : 101) | NC score | 0.043144 (rank : 34) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
SPAG5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.008358 (rank : 107) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96R06, O95213, Q9BWE8, Q9NT17, Q9UFE6 | Gene names | SPAG5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 5 (Astrin) (Mitotic spindle-associated protein p126) (MAP126) (Deepest). | |||||
|
TCF7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.027738 (rank : 60) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P36402, Q9UKI4 | Gene names | TCF7, TCF1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
AKP8L_MOUSE
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.015634 (rank : 94) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R0L7 | Gene names | Akap8l, Nakap95 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 8-like (AKAP8-like protein) (Neighbor of A- kinase-anchoring protein 95) (Neighbor of AKAP95). | |||||
|
APS_MOUSE
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.008858 (rank : 104) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JID9, O88936 | Gene names | Aps | |||
|
Domain Architecture |
|
|||||
| Description | SH2 and PH domain-containing adapter protein APS (Adapter protein with pleckstrin homology and Src homology 2 domains). | |||||
|
CHD9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.005319 (rank : 114) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
DNMBP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.007693 (rank : 109) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 413 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6XZF7, Q8IVY3, Q9Y2L3 | Gene names | DNMBP, KIAA1010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dynamin-binding protein (Scaffold protein Tuba). | |||||
|
IMA7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.009483 (rank : 103) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.026318 (rank : 63) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MNT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.021267 (rank : 78) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
NADAP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.016676 (rank : 92) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BWU0, Q4KMT1, Q4KMX0, Q7Z5Q9, Q9NVN2 | Gene names | SLC4A1AP, HLC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kanadaptin (Kidney anion exchanger adapter protein) (Solute carrier family 4 anion exchanger member 1 adapter protein) (Lung cancer oncogene 3 protein). | |||||
|
PDLI7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.006102 (rank : 113) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 547 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NR12, Q14250, Q5XG82, Q6NVZ5, Q96C91, Q9BXB8, Q9BXB9 | Gene names | PDLIM7, ENIGMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ and LIM domain protein 7 (LIM mineralization protein) (LMP) (Protein enigma). | |||||
|
PDXL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.016862 (rank : 91) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NZ53, Q6UVY4, Q8WUV6 | Gene names | PODXL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
SORC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.006406 (rank : 112) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JLC4, Q8VI45, Q922I1, Q9QY21 | Gene names | Sorcs1, Sorcs | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS1 precursor (mSorCS). | |||||
|
ZBED1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.025108 (rank : 68) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O96006, Q96BY4 | Gene names | ZBED1, ALTE, DREF, KIAA0785, TRAMP | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger BED domain-containing protein 1 (dREF homolog) (Putative Ac-like transposable element). | |||||
|
ZC11A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.011548 (rank : 100) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
2AAB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.056896 (rank : 14) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P30154, O75620 | Gene names | PPP2R1B | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A beta isoform (PP2A, subunit A, PR65-beta isoform) (PP2A, subunit A, R1-beta isoform). | |||||
|
2AAB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.058764 (rank : 13) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TNP2 | Gene names | Ppp2r1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A beta isoform (PP2A, subunit A, PR65-beta isoform) (PP2A, subunit A, R1-beta isoform). | |||||
|
HD_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 116 | |
| SwissProt Accessions | P42859 | Gene names | Hd, Hdh | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein homolog) (HD protein). | |||||
|
HD_HUMAN
|
||||||
| NC score | 0.977056 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.113414 (rank : 3) | θ value | 0.00390308 (rank : 4) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.099636 (rank : 4) | θ value | 0.00175202 (rank : 3) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
2AAA_MOUSE
|
||||||
| NC score | 0.075300 (rank : 5) | θ value | 0.163984 (rank : 8) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q76MZ3 | Gene names | Ppp2r1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform (PP2A, subunit A, PR65-alpha isoform) (PP2A, subunit A, R1-alpha isoform). | |||||
|
CD2_HUMAN
|
||||||
| NC score | 0.075044 (rank : 6) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
2AAA_HUMAN
|
||||||
| NC score | 0.068315 (rank : 7) | θ value | 0.365318 (rank : 11) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P30153, Q13773 | Gene names | PPP2R1A | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform (PP2A, subunit A, PR65-alpha isoform) (PP2A, subunit A, R1-alpha isoform) (Medium tumor antigen-associated 61 kDa protein). | |||||
|
MKL1_MOUSE
|
||||||
| NC score | 0.065362 (rank : 8) | θ value | 0.0113563 (rank : 5) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
PPRB_MOUSE
|
||||||
| NC score | 0.063524 (rank : 9) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
SF3B1_HUMAN
|
||||||
| NC score | 0.062822 (rank : 10) | θ value | 0.813845 (rank : 19) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75533 | Gene names | SF3B1, SAP155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
SF3B1_MOUSE
|
||||||
| NC score | 0.062753 (rank : 11) | θ value | 0.813845 (rank : 20) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99NB9, Q9CSK5 | Gene names | Sf3b1, Sap155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.060416 (rank : 12) | θ value | 0.813845 (rank : 17) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
2AAB_MOUSE
|
||||||
| NC score | 0.058764 (rank : 13) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TNP2 | Gene names | Ppp2r1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A beta isoform (PP2A, subunit A, PR65-beta isoform) (PP2A, subunit A, R1-beta isoform). | |||||
|
2AAB_HUMAN
|
||||||
| NC score | 0.056896 (rank : 14) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P30154, O75620 | Gene names | PPP2R1B | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A beta isoform (PP2A, subunit A, PR65-beta isoform) (PP2A, subunit A, R1-beta isoform). | |||||
|
TSC2_MOUSE
|
||||||
| NC score | 0.054327 (rank : 15) | θ value | 0.0330416 (rank : 6) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61037, P97723, P97724, P97725, P97727, Q61007, Q61008, Q9WUF6 | Gene names | Tsc2 | |||
|
Domain Architecture |

|
|||||
| Description | Tuberin (Tuberous sclerosis 2 homolog protein). | |||||
|
PSL1_HUMAN
|
||||||
| NC score | 0.054200 (rank : 16) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
GCN1L_HUMAN
|
||||||
| NC score | 0.053938 (rank : 17) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92616, O95001, O95651, Q6P2S3, Q86X65, Q8N5I5, Q8WU80, Q99736, Q9UE60 | Gene names | GCN1L1, KIAA0219 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GCN1-like protein 1 (HsGCN1). | |||||
|
GAK5_HUMAN
|
||||||
| NC score | 0.050179 (rank : 18) | θ value | 1.06291 (rank : 22) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.050168 (rank : 19) | θ value | 0.0961366 (rank : 7) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
ZCH10_MOUSE
|
||||||
| NC score | 0.049413 (rank : 20) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CX48 | Gene names | Zcchc10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 10. | |||||
|
GATA6_MOUSE
|
||||||
| NC score | 0.049329 (rank : 21) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61169, P97729, Q9QZK3 | Gene names | Gata6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor GATA-6 (GATA-binding factor 6). | |||||
|
ATX7_MOUSE
|
||||||
| NC score | 0.048272 (rank : 22) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8R4I1, Q8BL17 | Gene names | Atxn7, Sca7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-7 (Spinocerebellar ataxia type 7 protein homolog). | |||||
|
GAK12_HUMAN
|
||||||
| NC score | 0.047493 (rank : 23) | θ value | 3.0926 (rank : 49) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK4_HUMAN
|
||||||
| NC score | 0.047271 (rank : 24) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK11_HUMAN
|
||||||
| NC score | 0.047207 (rank : 25) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK2_HUMAN
|
||||||
| NC score | 0.047038 (rank : 26) | θ value | 3.0926 (rank : 51) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
AKAP6_HUMAN
|
||||||
| NC score | 0.046316 (rank : 27) | θ value | 0.279714 (rank : 9) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13023, O15028 | Gene names | AKAP6, AKAP100, KIAA0311 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 6 (Protein kinase A-anchoring protein 6) (PRKA6) (A-kinase anchor protein 100 kDa) (AKAP 100) (mAKAP). | |||||
|
GAK6_HUMAN
|
||||||
| NC score | 0.046276 (rank : 28) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
SPR1A_MOUSE
|
||||||
| NC score | 0.045868 (rank : 29) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62266 | Gene names | Sprr1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cornifin A (Small proline-rich protein 1A) (SPR1A) (SPR1 A). | |||||
|
CC021_MOUSE
|
||||||
| NC score | 0.045552 (rank : 30) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3U4G3, Q3TLX5, Q8K2I0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C3orf21 homolog. | |||||
|
PPRB_HUMAN
|
||||||
| NC score | 0.044893 (rank : 31) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
AMBN_HUMAN
|
||||||
| NC score | 0.043800 (rank : 32) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NP70, Q9H2X1, Q9H4L1 | Gene names | AMBN | |||
|
Domain Architecture |

|
|||||
| Description | Ameloblastin precursor. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.043549 (rank : 33) | θ value | 0.279714 (rank : 10) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SON_MOUSE
|
||||||
| NC score | 0.043144 (rank : 34) | θ value | 6.88961 (rank : 101) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
GAK1_HUMAN
|
||||||
| NC score | 0.040558 (rank : 35) | θ value | 3.0926 (rank : 50) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P62683 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GATA6_HUMAN
|
||||||
| NC score | 0.039921 (rank : 36) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92908, P78327 | Gene names | GATA6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor GATA-6 (GATA-binding factor 6). | |||||
|
JPH2_HUMAN
|
||||||
| NC score | 0.039917 (rank : 37) | θ value | 0.365318 (rank : 13) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
AUTS2_HUMAN
|
||||||
| NC score | 0.039052 (rank : 38) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WXX7, Q9Y4F2 | Gene names | AUTS2, KIAA0442 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Autism susceptibility gene 2 protein. | |||||
|
TOPRS_MOUSE
|
||||||
| NC score | 0.038940 (rank : 39) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80Z37, Q3U5B5, Q3UPZ1, Q3USN3, Q3UZ55, Q8BXP2, Q8CFF5, Q8CGC8, Q920L3 | Gene names | Topors | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
TARSH_HUMAN
|
||||||
| NC score | 0.038661 (rank : 40) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
AN32B_MOUSE
|
||||||
| NC score | 0.038636 (rank : 41) | θ value | 0.365318 (rank : 12) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9EST5 | Gene names | Anp32b, Pal31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member B (Proliferation-related acidic leucine-rich protein PAL31). | |||||
|
TOPRS_HUMAN
|
||||||
| NC score | 0.037388 (rank : 42) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NS56, O43273, Q6P987, Q9NS55, Q9UNR9 | Gene names | TOPORS, LUN, TP53BPL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
CJ026_HUMAN
|
||||||
| NC score | 0.037253 (rank : 43) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NX94, Q2HIY7, Q5F2G6 | Gene names | C10orf26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C10orf26. | |||||
|
4ET_HUMAN
|
||||||
| NC score | 0.036893 (rank : 44) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.036430 (rank : 45) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
LRC47_MOUSE
|
||||||
| NC score | 0.035166 (rank : 46) | θ value | 0.813845 (rank : 18) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q505F5, Q3U380, Q6ZPW2, Q9CUZ0 | Gene names | Lrrc47, Kiaa1185 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 47. | |||||
|
BTBDC_HUMAN
|
||||||
| NC score | 0.034661 (rank : 47) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IY92, Q96JP1 | Gene names | BTBD12, KIAA1784 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 12. | |||||
|
TCF7_MOUSE
|
||||||
| NC score | 0.034246 (rank : 48) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q00417 | Gene names | Tcf7, Tcf-1, Tcf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
SRCA_HUMAN
|
||||||
| NC score | 0.034207 (rank : 49) | θ value | 0.365318 (rank : 14) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
SPR1B_MOUSE
|
||||||
| NC score | 0.034065 (rank : 50) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62267 | Gene names | Sprr1b | |||
|
Domain Architecture |

|
|||||
| Description | Cornifin B (Small proline-rich protein 1B) (SPR1B) (SPR1 B). | |||||
|
TPO_MOUSE
|
||||||
| NC score | 0.032998 (rank : 51) | θ value | 5.27518 (rank : 90) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P40226 | Gene names | Thpo | |||
|
Domain Architecture |

|
|||||
| Description | Thrombopoietin precursor (Megakaryocyte colony-stimulating factor) (Myeloproliferative leukemia virus oncogene ligand) (C-mpl ligand) (ML) (Megakaryocyte growth and development factor) (MGDF). | |||||
|
ILDR1_HUMAN
|
||||||
| NC score | 0.032383 (rank : 52) | θ value | 1.38821 (rank : 29) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86SU0, Q6ZP61, Q7Z578 | Gene names | ILDR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immunoglobulin-like domain-containing receptor 1 precursor. | |||||
|
HGS_HUMAN
|
||||||
| NC score | 0.030253 (rank : 53) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
NUFP1_HUMAN
|
||||||
| NC score | 0.030110 (rank : 54) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UHK0, Q8WVM5, Q96SG1 | Gene names | NUFIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 1 (Nuclear FMRP-interacting protein 1). | |||||
|
CABIN_HUMAN
|
||||||
| NC score | 0.029565 (rank : 55) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
PEX14_HUMAN
|
||||||
| NC score | 0.029314 (rank : 56) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75381, Q8WX51 | Gene names | PEX14 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal membrane protein PEX14 (Peroxin-14) (Peroxisomal membrane anchor protein PEX14) (PTS1 receptor docking protein). | |||||
|
CD3E_MOUSE
|
||||||
| NC score | 0.028809 (rank : 57) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22646 | Gene names | Cd3e | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface glycoprotein CD3 epsilon chain precursor (T-cell surface antigen T3/Leu-4 epsilon chain). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.028263 (rank : 58) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
CRSP7_HUMAN
|
||||||
| NC score | 0.027870 (rank : 59) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
TCF7_HUMAN
|
||||||
| NC score | 0.027738 (rank : 60) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P36402, Q9UKI4 | Gene names | TCF7, TCF1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
FBX18_HUMAN
|
||||||
| NC score | 0.027189 (rank : 61) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NFZ0, Q7Z4Q6, Q7Z4R0, Q8N1P5, Q8N586, Q96E82, Q96K67, Q96SW7, Q9UFB2 | Gene names | FBXO18, FBH1, FBX18 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 18 (EC 3.6.1.-) (F-box DNA helicase 1). | |||||
|
CO039_HUMAN
|
||||||
| NC score | 0.026348 (rank : 62) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6ZRI6, Q71JB1, Q7L3S0, Q8N3F2, Q96FB6, Q9NTU5 | Gene names | C15orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39. | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.026318 (rank : 63) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
SL9A8_MOUSE
|
||||||
| NC score | 0.025912 (rank : 64) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R4D1, Q3UPR4, Q5WA59, Q8BIH8, Q8BJ27 | Gene names | Slc9a8, Nhe8 | |||
|
Domain Architecture |
|
|||||
| Description | Sodium/hydrogen exchanger 8 (Na(+)/H(+) exchanger 8) (NHE-8) (Solute carrier family 9 member 8). | |||||
|
CT032_HUMAN
|
||||||
| NC score | 0.025794 (rank : 65) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQ75, Q96K09, Q9BYL5 | Gene names | HEFL, C20orf32 | |||
|
Domain Architecture |

|
|||||
| Description | HEF-like protein. | |||||
|
MOCOS_HUMAN
|
||||||
| NC score | 0.025682 (rank : 66) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96EN8, Q53GP5, Q8N3A4, Q9NWM7 | Gene names | MOCOS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Molybdenum cofactor sulfurase (EC 4.4.-.-) (MoCo sulfurase) (MOS) (HMCS). | |||||
|
DTX1_MOUSE
|
||||||
| NC score | 0.025599 (rank : 67) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61010, Q3TER5, Q8C2H2, Q9ER09 | Gene names | Dtx1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein deltex-1 (Deltex-1) (Deltex1) (mDTX1) (FXI-T1). | |||||
|
ZBED1_HUMAN
|
||||||
| NC score | 0.025108 (rank : 68) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O96006, Q96BY4 | Gene names | ZBED1, ALTE, DREF, KIAA0785, TRAMP | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger BED domain-containing protein 1 (dREF homolog) (Putative Ac-like transposable element). | |||||
|
DOHH_HUMAN
|
||||||
| NC score | 0.024895 (rank : 69) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BU89, O75265 | Gene names | DOHH, HLRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deoxyhypusine hydroxylase (EC 1.14.99.29) (Deoxyhypusine monooxygenase) (hDOHH) (HEAT-like repeat-containing protein 1). | |||||
|
DTX1_HUMAN
|
||||||
| NC score | 0.024346 (rank : 70) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86Y01, O60630, Q9BS04 | Gene names | DTX1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein deltex-1 (Deltex-1) (Deltex1) (hDTX1). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.024108 (rank : 71) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
IRF3_HUMAN
|
||||||
| NC score | 0.023956 (rank : 72) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14653 | Gene names | IRF3 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 3 (IRF-3). | |||||
|
SL9A8_HUMAN
|
||||||
| NC score | 0.023724 (rank : 73) | θ value | 5.27518 (rank : 87) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2E8, Q68CZ8, Q9BX15, Q9Y507 | Gene names | SLC9A8, KIAA0939, NHE8 | |||
|
Domain Architecture |
|
|||||
| Description | Sodium/hydrogen exchanger 8 (Na(+)/H(+) exchanger 8) (NHE-8) (Solute carrier family 9 member 8). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.023680 (rank : 74) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
TSKS_MOUSE
|
||||||
| NC score | 0.022790 (rank : 75) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54887 | Gene names | Stk22s1, Tsks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-specific serine kinase substrate (Testis-specific kinase substrate) (STK22 substrate 1). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.021754 (rank : 76) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
PF21B_MOUSE
|
||||||
| NC score | 0.021416 (rank : 77) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
MNT_HUMAN
|
||||||
| NC score | 0.021267 (rank : 78) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.021136 (rank : 79) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
CAND2_MOUSE
|
||||||
| NC score | 0.021005 (rank : 80) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZQ73, Q6PHT7 | Gene names | Cand2, Kiaa0667, Tip120b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cullin-associated NEDD8-dissociated protein 2 (Cullin-associated and neddylation-dissociated protein 2) (p120 CAND2) (TBP-interacting protein TIP120B) (TBP-interacting protein of 120 kDa B). | |||||
|
NFAT5_MOUSE
|
||||||
| NC score | 0.020315 (rank : 81) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WV30 | Gene names | Nfat5 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells 5 (T cell transcription factor NFAT5) (NF-AT5) (Rel domain-containing transcription factor NFAT5). | |||||
|
AKAP2_MOUSE
|
||||||
| NC score | 0.019278 (rank : 82) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
PIGG_HUMAN
|
||||||
| NC score | 0.019189 (rank : 83) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5H8A4, Q2TAK5, Q6UX31, Q7L5Y4, Q8N866, Q8NCC9, Q96SY9, Q9BVT7, Q9NXG5 | Gene names | PIGG, GPI7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI ethanolamine phosphate transferase 2 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class G protein) (PIG-G) (GPI7 homolog) (hGPI7). | |||||
|
SMG1_HUMAN
|
||||||
| NC score | 0.018928 (rank : 84) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96Q15, O43305, Q13284, Q8NFX2, Q96QV0, Q96RW3 | Gene names | SMG1, ATX, KIAA0421, LIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1) (hSMG-1) (Lambda/iota protein kinase C-interacting protein) (Lambda-interacting protein) (61E3.4). | |||||
|
PHF14_MOUSE
|
||||||
| NC score | 0.018835 (rank : 85) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
SMG1_MOUSE
|
||||||
| NC score | 0.018824 (rank : 86) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BKX6, Q5DU42, Q6ZQC0, Q8BLU4, Q8BWJ5, Q8BXD3 | Gene names | Smg1, Kiaa0421 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1). | |||||
|
FOXN4_HUMAN
|
||||||
| NC score | 0.018585 (rank : 87) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96NZ1, Q6ZMR4, Q96NZ0 | Gene names | FOXN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein N4. | |||||
|
BAG3_MOUSE
|
||||||
| NC score | 0.017681 (rank : 88) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
TF65_HUMAN
|
||||||
| NC score | 0.017586 (rank : 89) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q04206, Q6SLK1 | Gene names | RELA, NFKB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor p65 (Nuclear factor NF-kappa-B p65 subunit). | |||||
|
DZIP3_MOUSE
|
||||||
| NC score | 0.017131 (rank : 90) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7TPV2, Q80TU4, Q8BYK7 | Gene names | Dzip3, Kiaa0675 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) (DAZ-interacting protein 3 homolog). | |||||
|
PDXL2_HUMAN
|
||||||
| NC score | 0.016862 (rank : 91) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NZ53, Q6UVY4, Q8WUV6 | Gene names | PODXL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
NADAP_HUMAN
|
||||||
| NC score | 0.016676 (rank : 92) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BWU0, Q4KMT1, Q4KMX0, Q7Z5Q9, Q9NVN2 | Gene names | SLC4A1AP, HLC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kanadaptin (Kidney anion exchanger adapter protein) (Solute carrier family 4 anion exchanger member 1 adapter protein) (Lung cancer oncogene 3 protein). | |||||
|
OAS3_HUMAN
|
||||||
| NC score | 0.016435 (rank : 93) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y6K5, Q9H3P5 | Gene names | OAS3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 2'-5'-oligoadenylate synthetase 3 (EC 2.7.7.-) ((2-5')oligo(A) synthetase 3) (2-5A synthetase 3) (p100 OAS) (p100OAS). | |||||
|
AKP8L_MOUSE
|
||||||
| NC score | 0.015634 (rank : 94) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R0L7 | Gene names | Akap8l, Nakap95 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 8-like (AKAP8-like protein) (Neighbor of A- kinase-anchoring protein 95) (Neighbor of AKAP95). | |||||
|
TFE2_HUMAN
|
||||||
| NC score | 0.014416 (rank : 95) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P15923, P15883, Q14208, Q14635, Q14636, Q9UPI9 | Gene names | TCF3, E2A, ITF1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Immunoglobulin transcription factor 1) (Transcription factor ITF-1) (Kappa-E2-binding factor). | |||||
|
COFA1_HUMAN
|
||||||
| NC score | 0.014275 (rank : 96) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P39059, Q5T6J4, Q9Y4W4 | Gene names | COL15A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV) (Restin) (Related to endostatin)]. | |||||
|
ETV5_HUMAN
|
||||||
| NC score | 0.012641 (rank : 97) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
RPGF5_MOUSE
|
||||||
| NC score | 0.012307 (rank : 98) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C0Q9, Q8BJJ9, Q8C0R5 | Gene names | Rapgef5, Gfr, Kiaa0277, Mrgef | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 5 (Guanine nucleotide exchange factor for Rap1) (M-Ras-regulated Rap GEF) (MR-GEF). | |||||
|
ITA4_HUMAN
|
||||||
| NC score | 0.011966 (rank : 99) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P13612 | Gene names | ITGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-4 precursor (Integrin alpha-IV) (VLA-4) (CD49d antigen). | |||||
|
ZC11A_MOUSE
|
||||||
| NC score | 0.011548 (rank : 100) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
ANGT_MOUSE
|
||||||
| NC score | 0.011307 (rank : 101) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P11859 | Gene names | Agt, Serpina8 | |||
|
Domain Architecture |

|
|||||
| Description | Angiotensinogen precursor [Contains: Angiotensin-1 (Angiotensin I) (Ang I); Angiotensin-2 (Angiotensin II) (Ang II); Angiotensin-3 (Angiotensin III) (Ang III) (Des-Asp[1]-angiotensin II)]. | |||||
|
KCNH8_HUMAN
|
||||||
| NC score | 0.011008 (rank : 102) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96L42 | Gene names | KCNH8 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 8 (Voltage-gated potassium channel subunit Kv12.1) (Ether-a-go-go-like potassium channel 3) (ELK channel 3) (ELK3) (ELK1) (hElk1). | |||||
|
IMA7_HUMAN
|
||||||
| NC score | 0.009483 (rank : 103) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
APS_MOUSE
|
||||||
| NC score | 0.008858 (rank : 104) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JID9, O88936 | Gene names | Aps | |||
|
Domain Architecture |

|
|||||
| Description | SH2 and PH domain-containing adapter protein APS (Adapter protein with pleckstrin homology and Src homology 2 domains). | |||||
|
MYO15_HUMAN
|
||||||
| NC score | 0.008533 (rank : 105) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.008432 (rank : 106) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
SPAG5_HUMAN
|
||||||
| NC score | 0.008358 (rank : 107) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96R06, O95213, Q9BWE8, Q9NT17, Q9UFE6 | Gene names | SPAG5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 5 (Astrin) (Mitotic spindle-associated protein p126) (MAP126) (Deepest). | |||||
|
DUET_HUMAN
|
||||||
| NC score | 0.007714 (rank : 108) | θ value | 0.813845 (rank : 16) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1220 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2A5, Q8TBQ5, Q9NSZ4 | Gene names | DUET, TRAD | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Duet (EC 2.7.11.1) (Serine/threonine kinase with Dbl- and pleckstrin homology domain). | |||||
|
DNMBP_HUMAN
|
||||||
| NC score | 0.007693 (rank : 109) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 413 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6XZF7, Q8IVY3, Q9Y2L3 | Gene names | DNMBP, KIAA1010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dynamin-binding protein (Scaffold protein Tuba). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.007174 (rank : 110) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.006570 (rank : 111) | θ value | 2.36792 (rank : 43) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
SORC1_MOUSE
|
||||||
| NC score | 0.006406 (rank : 112) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JLC4, Q8VI45, Q922I1, Q9QY21 | Gene names | Sorcs1, Sorcs | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS1 precursor (mSorCS). | |||||
|
PDLI7_HUMAN
|
||||||
| NC score | 0.006102 (rank : 113) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 547 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NR12, Q14250, Q5XG82, Q6NVZ5, Q96C91, Q9BXB8, Q9BXB9 | Gene names | PDLIM7, ENIGMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ and LIM domain protein 7 (LIM mineralization protein) (LMP) (Protein enigma). | |||||
|
CHD9_MOUSE
|
||||||
| NC score | 0.005319 (rank : 114) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
ADA12_MOUSE
|
||||||
| NC score | 0.005244 (rank : 115) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61824 | Gene names | Adam12, Mltna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
RFXK_HUMAN
|
||||||
| NC score | 0.004941 (rank : 116) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O14593, O95839 | Gene names | RFXANK, ANKRA1, RFXB | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein RFXANK (Regulatory factor X subunit B) (RFX-B) (Ankyrin repeat family A protein 1). | |||||
|
M4K1_MOUSE
|
||||||
| NC score | 0.002527 (rank : 117) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 858 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70218, Q8R3R0, Q99K62 | Gene names | Map4k1, Hpk1 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase kinase 1 (EC 2.7.11.1) (MAPK/ERK kinase kinase kinase 1) (MEK kinase kinase 1) (MEKKK 1) (Hematopoietic progenitor kinase) (HPK). | |||||
|
MTF1_HUMAN
|
||||||
| NC score | 0.000500 (rank : 118) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 814 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14872, Q96CB1 | Gene names | MTF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||