Please be patient as the page loads
|
AMBN_HUMAN
|
||||||
| SwissProt Accessions | Q9NP70, Q9H2X1, Q9H4L1 | Gene names | AMBN | |||
|
Domain Architecture |
|
|||||
| Description | Ameloblastin precursor. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
AMBN_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 107 | |
| SwissProt Accessions | Q9NP70, Q9H2X1, Q9H4L1 | Gene names | AMBN | |||
|
Domain Architecture |

|
|||||
| Description | Ameloblastin precursor. | |||||
|
AMBN_MOUSE
|
||||||
| θ value | 1.34075e-120 (rank : 2) | NC score | 0.930525 (rank : 2) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O55189 | Gene names | Ambn | |||
|
Domain Architecture |
|
|||||
| Description | Ameloblastin precursor. | |||||
|
ASPP1_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 3) | NC score | 0.054497 (rank : 36) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q62415 | Gene names | Ppp1r13b, Aspp1 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 1 (Protein phosphatase 1 regulatory subunit 13B). | |||||
|
DIAP2_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 4) | NC score | 0.104314 (rank : 5) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O70566, Q8C2G8 | Gene names | Diaph2, Diap2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2) (mDia3). | |||||
|
FMN2_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 5) | NC score | 0.122818 (rank : 3) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9JL04 | Gene names | Fmn2 | |||
|
Domain Architecture |

|
|||||
| Description | Formin-2. | |||||
|
AIRE_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 6) | NC score | 0.057290 (rank : 30) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
ZN687_HUMAN
|
||||||
| θ value | 0.125558 (rank : 7) | NC score | 0.032045 (rank : 68) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
FMN2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 8) | NC score | 0.112416 (rank : 4) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
IF4G1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 9) | NC score | 0.043698 (rank : 49) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
SHAN2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 10) | NC score | 0.043254 (rank : 50) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 446 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80Z38, Q3UTK4, Q5DU07 | Gene names | Shank2, Kiaa1022 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2) (Cortactin- binding protein 1) (CortBP1). | |||||
|
ZFHX2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 11) | NC score | 0.032408 (rank : 67) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9C0A1 | Gene names | ZFHX2, KIAA1762 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger homeobox protein 2. | |||||
|
SYP2L_MOUSE
|
||||||
| θ value | 0.62314 (rank : 12) | NC score | 0.039539 (rank : 54) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BWB1 | Gene names | Synpo2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin 2-like protein. | |||||
|
AMELX_HUMAN
|
||||||
| θ value | 0.813845 (rank : 13) | NC score | 0.068980 (rank : 17) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99217, Q96NW6, Q9UCA7 | Gene names | AMELX, AMG, AMGX | |||
|
Domain Architecture |

|
|||||
| Description | Amelogenin, X isoform precursor. | |||||
|
DRD4_HUMAN
|
||||||
| θ value | 0.813845 (rank : 14) | NC score | 0.009505 (rank : 108) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P21917 | Gene names | DRD4 | |||
|
Domain Architecture |
|
|||||
| Description | D(4) dopamine receptor (Dopamine D4 receptor) (D(2C) dopamine receptor). | |||||
|
FMNL_HUMAN
|
||||||
| θ value | 0.813845 (rank : 15) | NC score | 0.097068 (rank : 8) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
Domain Architecture |
|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
MBD6_HUMAN
|
||||||
| θ value | 0.813845 (rank : 16) | NC score | 0.070420 (rank : 15) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
TACC3_MOUSE
|
||||||
| θ value | 0.813845 (rank : 17) | NC score | 0.047238 (rank : 44) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JJ11, Q9WVK9 | Gene names | Tacc3, Aint | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ARNT-interacting protein). | |||||
|
DIAP1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.085218 (rank : 11) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O60610, Q9UC76 | Gene names | DIAPH1, DIAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1). | |||||
|
NFAC1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 19) | NC score | 0.034376 (rank : 64) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O95644, Q12865, Q15793 | Gene names | NFATC1, NFAT2, NFATC | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 1 (NFAT transcription complex cytosolic component) (NF-ATc1) (NF-ATc). | |||||
|
PRAX_HUMAN
|
||||||
| θ value | 1.06291 (rank : 20) | NC score | 0.062528 (rank : 24) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BXM0, Q9BXL9, Q9HCF2 | Gene names | PRX, KIAA1620 | |||
|
Domain Architecture |
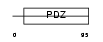
|
|||||
| Description | Periaxin. | |||||
|
R3HD2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 21) | NC score | 0.053028 (rank : 39) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
SYN1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 22) | NC score | 0.067713 (rank : 18) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
SYN1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.066879 (rank : 19) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O88935, Q62279, Q8QZT8 | Gene names | Syn1, Syn-1 | |||
|
Domain Architecture |
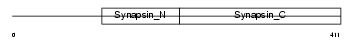
|
|||||
| Description | Synapsin-1 (Synapsin I). | |||||
|
3BP2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 24) | NC score | 0.055571 (rank : 34) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P78314, O00500, O15373, P78315 | Gene names | SH3BP2, 3BP2 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 2 (3BP-2). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.053774 (rank : 38) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CO4A3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.026680 (rank : 77) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q01955, Q9BQT2 | Gene names | COL4A3 | |||
|
Domain Architecture |
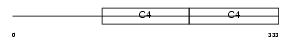
|
|||||
| Description | Collagen alpha-3(IV) chain precursor (Goodpasture antigen). | |||||
|
DIAP2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.092459 (rank : 9) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60879, O60878, Q8WX06, Q8WX48, Q9UJL2 | Gene names | DIAPH2, DIA | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2). | |||||
|
FMNL_MOUSE
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.091990 (rank : 10) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9JL26, Q6KAN4, Q9Z2V7 | Gene names | Fmnl1, Frl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-like protein 1 (Formin-related protein). | |||||
|
MTF1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 29) | NC score | 0.007865 (rank : 112) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 814 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14872, Q96CB1 | Gene names | MTF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
ZN206_HUMAN
|
||||||
| θ value | 1.38821 (rank : 30) | NC score | 0.004970 (rank : 120) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 853 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96SZ4 | Gene names | ZNF206, ZSCAN10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 206 (Zinc finger and SCAN domain-containing protein 10). | |||||
|
CSPG3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.014039 (rank : 102) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P55066, Q6P1E3, Q8C4F8 | Gene names | Cspg3, Ncan | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
ENAH_MOUSE
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.030083 (rank : 70) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
HD_MOUSE
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.043800 (rank : 48) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P42859 | Gene names | Hd, Hdh | |||
|
Domain Architecture |
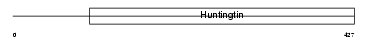
|
|||||
| Description | Huntingtin (Huntington disease protein homolog) (HD protein). | |||||
|
SYN2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.060683 (rank : 26) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64332, Q6NZR0, Q9QWV7 | Gene names | Syn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
AMELX_MOUSE
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.059373 (rank : 27) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P63277, P45559, P70592, Q61293 | Gene names | Amelx, Amel | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amelogenin, X isoform precursor (Leucine-rich amelogenin peptide) (LRAP). | |||||
|
BASP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.035159 (rank : 63) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
K0562_HUMAN
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.036218 (rank : 60) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60308, Q5JSQ3, Q5SR24, Q5SR25, Q6PKF5, Q86W32, Q86X14 | Gene names | KIAA0562 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0562. | |||||
|
PRAX_MOUSE
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.058870 (rank : 28) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O55103, O55104 | Gene names | Prx | |||
|
Domain Architecture |
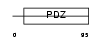
|
|||||
| Description | Periaxin. | |||||
|
PRB4S_HUMAN
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.034215 (rank : 65) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
PRDM2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.017329 (rank : 95) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.032978 (rank : 66) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
RBM12_MOUSE
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.039099 (rank : 56) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8R4X3, Q8K373, Q8R302, Q9CS80 | Gene names | Rbm12 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
RX_HUMAN
|
||||||
| θ value | 2.36792 (rank : 43) | NC score | 0.012343 (rank : 104) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2V3 | Gene names | RAX, RX | |||
|
Domain Architecture |

|
|||||
| Description | Retinal homeobox protein Rx (Retina and anterior neural fold homeobox protein). | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.028905 (rank : 73) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
DCP1A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.035269 (rank : 62) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NPI6 | Gene names | DCP1A, SMIF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (Smad4-interacting transcriptional co-activator). | |||||
|
PLXB1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.018715 (rank : 93) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43157, Q6NY20, Q9UIV7, Q9UJ92, Q9UJ93 | Gene names | PLXNB1, KIAA0407, SEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor (Semaphorin receptor SEP). | |||||
|
WBP11_HUMAN
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.045432 (rank : 46) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y2W2, Q96AY8 | Gene names | WBP11, NPWBP, SNP70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11) (SH3 domain-binding protein SNP70) (Npw38-binding protein) (NpwBP). | |||||
|
WBP11_MOUSE
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.048598 (rank : 42) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q923D5, O88539, Q3UMD8, Q8VDI0 | Gene names | Wbp11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11). | |||||
|
DAB1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.036080 (rank : 61) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P97318, P97316, P97317 | Gene names | Dab1 | |||
|
Domain Architecture |
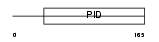
|
|||||
| Description | Disabled homolog 1. | |||||
|
DDEF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.015013 (rank : 99) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QWY8, O08612, Q80TG8, Q80UV6, Q99LV8, Q9Z2B6 | Gene names | Ddef1, Asap1, Kiaa1249, Shag1 | |||
|
Domain Architecture |

|
|||||
| Description | 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase- activating protein (PIP2-dependent ARF1 GAP) (ADP-ribosylation factor- directed GTPase-activating protein 1) (ARF GTPase-activating protein 1) (Development and differentiation-enhancing factor 1) (Differentiation-enhancing factor 1) (DEF-1). | |||||
|
FMN1A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.098104 (rank : 7) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q05860 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoforms I/II/III (Limb deformity protein). | |||||
|
FMN1B_MOUSE
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.099317 (rank : 6) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q05859 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoform IV (Limb deformity protein). | |||||
|
KLF2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.005263 (rank : 118) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q60843 | Gene names | Klf2, Lklf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 2 (Lung krueppel-like factor). | |||||
|
PRR12_HUMAN
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.045514 (rank : 45) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
SFR14_HUMAN
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.024286 (rank : 81) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IX01, O15071, O60369, Q5JPH7, Q8WUF7 | Gene names | SFRS14, KIAA0365 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
SPON1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.022048 (rank : 89) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HCB6, O94862, Q8NCD7, Q8WUR5 | Gene names | SPON1, KIAA0762, VSGP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spondin-1 precursor (F-spondin) (Vascular smooth muscle cell growth- promoting factor). | |||||
|
SPON1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.021980 (rank : 90) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VCC9, Q8K2Q8 | Gene names | Spon1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spondin-1 precursor (F-spondin). | |||||
|
SYN2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.057797 (rank : 29) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92777 | Gene names | SYN2 | |||
|
Domain Architecture |
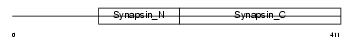
|
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
TNFB_MOUSE
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.027254 (rank : 76) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P09225 | Gene names | Lta, Tnfb, Tnfsf1 | |||
|
Domain Architecture |
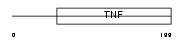
|
|||||
| Description | Lymphotoxin-alpha precursor (LT-alpha) (TNF-beta) (Tumor necrosis factor ligand superfamily member 1). | |||||
|
AAK1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.008711 (rank : 110) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q3UHJ0, Q3TY53, Q6ZPZ6, Q80XP6 | Gene names | Aak1, Kiaa1048 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP2-associated protein kinase 1 (EC 2.7.11.1) (Adaptor-associated kinase 1). | |||||
|
BSCL2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.026465 (rank : 78) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z2E9, Q810B0, Q9JJC2, Q9JMF1 | Gene names | Bscl2, Gng3lg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Seipin (Bernardinelli-Seip congenital lipodystrophy type 2 protein homolog). | |||||
|
CO4A2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.023655 (rank : 84) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 454 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P08572 | Gene names | COL4A2 | |||
|
Domain Architecture |
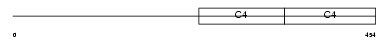
|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
CO5A1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.023078 (rank : 86) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P20908, Q15094, Q5SUX4 | Gene names | COL5A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
DIAP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.078577 (rank : 13) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 839 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O08808 | Gene names | Diaph1, Diap1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1) (mDIA1) (p140mDIA). | |||||
|
DREB_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.023108 (rank : 85) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q16643 | Gene names | DBN1 | |||
|
Domain Architecture |

|
|||||
| Description | Drebrin (Developmentally-regulated brain protein). | |||||
|
HGS_MOUSE
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.023945 (rank : 83) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
MTF1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.006243 (rank : 117) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q07243 | Gene names | Mtf1, Mtf-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
MTMR7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.012036 (rank : 106) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y216, Q68DX4 | Gene names | MTMR7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
MTSS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.047948 (rank : 43) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O43312, Q8TCA2, Q96RX2 | Gene names | MTSS1, KIAA0429, MIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein) (Metastasis suppressor YGL-1). | |||||
|
SNX18_MOUSE
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.016898 (rank : 96) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91ZR2 | Gene names | Snag1, Snx18 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-18 (Sorting nexin-associated Golgi protein 1). | |||||
|
SSBP4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.069824 (rank : 16) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9BWG4 | Gene names | SSBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Single-stranded DNA-binding protein 4. | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.041915 (rank : 51) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.043874 (rank : 47) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
B3A2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.008950 (rank : 109) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P04920, Q45EY5, Q969L3 | Gene names | SLC4A2, AE2, EPB3L1, HKB3, MPB3L | |||
|
Domain Architecture |

|
|||||
| Description | Anion exchange protein 2 (Non-erythroid band 3-like protein) (AE2 anion exchanger) (Solute carrier family 4 member 2) (BND3L). | |||||
|
CPSF6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.036964 (rank : 58) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q16630, Q53ES1, Q9BSJ7, Q9BW18 | Gene names | CPSF6, CFIM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6 (Cleavage and polyadenylation specificity factor 68 kDa subunit) (CPSF 68 kDa subunit) (Pre-mRNA cleavage factor Im 68 kDa subunit) (Protein HPBRII- 4/7). | |||||
|
CPSF6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.037058 (rank : 57) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
DAB1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.036933 (rank : 59) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75553, Q9NYA8 | Gene names | DAB1 | |||
|
Domain Architecture |
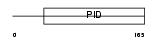
|
|||||
| Description | Disabled homolog 1. | |||||
|
GORS2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.030109 (rank : 69) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H8Y8, Q96I74, Q96K84, Q9H946, Q9UFW4 | Gene names | GORASP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 2 (GRS2) (Golgi reassembly-stacking protein of 55 kDa) (GRASP55) (p59) (Golgi phosphoprotein 6) (GOLPH6). | |||||
|
HD_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.039209 (rank : 55) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
IL3RB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.028237 (rank : 75) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 355 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P26955 | Gene names | Csf2rb, Aic2b, Csf2rb1, Il3rb1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytokine receptor common beta chain precursor (GM-CSF/IL-3/IL-5 receptor common beta-chain) (CD131 antigen). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.040827 (rank : 52) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MUC7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.029041 (rank : 72) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.024461 (rank : 80) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
R51A1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.024183 (rank : 82) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96B01, O43403, Q7Z779 | Gene names | RAD51AP1, PIR51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAD51-associated protein 1 (RAD51-interacting protein). | |||||
|
TIF1B_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.014152 (rank : 101) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62318, P70391, Q8C283, Q99PN4 | Gene names | Trim28, Krip1, Tif1b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (KRAB-A-interacting protein) (KRIP-1). | |||||
|
TSK_MOUSE
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.006674 (rank : 115) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CBR6, Q4W655 | Gene names | Lrrc54, Tsk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tsukushi precursor (Leucine-rich repeat-containing protein 54). | |||||
|
USP9X_MOUSE
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.011072 (rank : 107) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
WASP_MOUSE
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.022959 (rank : 87) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70315 | Gene names | Was, Wasp | |||
|
Domain Architecture |
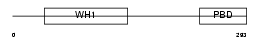
|
|||||
| Description | Wiskott-Aldrich syndrome protein homolog (WASp). | |||||
|
AAK1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.006521 (rank : 116) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1077 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q2M2I8, Q4ZFZ3, Q53RX6, Q9UPV4 | Gene names | AAK1, KIAA1048 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP2-associated protein kinase 1 (EC 2.7.11.1) (Adaptor-associated kinase 1). | |||||
|
ABI3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.013927 (rank : 103) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2A4, Q9H0P6 | Gene names | ABI3, NESH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABI gene family member 3 (New molecule including SH3) (Nesh). | |||||
|
CO3A1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.020628 (rank : 91) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
CO4A5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.024683 (rank : 79) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P29400, Q16006, Q16126, Q6LD84 | Gene names | COL4A5 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-5(IV) chain precursor. | |||||
|
CO9A1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.022820 (rank : 88) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q05722, Q61269, Q61270, Q61433, Q61940 | Gene names | Col9a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
CO9A2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.018935 (rank : 92) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q07643 | Gene names | Col9a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(IX) chain precursor. | |||||
|
DEN1A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.016338 (rank : 98) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TEH3, Q5VWF0, Q8IVD6, Q9H796 | Gene names | DENND1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
FBX46_MOUSE
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.014750 (rank : 100) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BG80 | Gene names | Fbxo46, Fbx46, Fbxo34l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
KI13B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.005208 (rank : 119) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NQT8, O75134, Q9BYJ6 | Gene names | KIF13B, GAKIN, KIAA0639 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13B (Kinesin-like protein GAKIN). | |||||
|
LHX1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.006899 (rank : 114) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P63006, P36199 | Gene names | Lhx1, Lim-1, Lim1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM/homeobox protein Lhx1 (LIM homeobox protein 1) (Homeobox protein Lim-1). | |||||
|
PCNT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.007621 (rank : 113) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.029203 (rank : 71) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
RBM16_HUMAN
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.028519 (rank : 74) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UPN6, Q5TBU6, Q9BQN8, Q9BX43 | Gene names | RBM16, KIAA1116 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein 16 (RNA-binding motif protein 16). | |||||
|
RIN2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 102) | NC score | 0.012195 (rank : 105) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WYP3, Q00425, Q5TFT8, Q9BQL3, Q9H071 | Gene names | RIN2, RASSF4 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2) (Ras inhibitor JC265) (Ras association domain family 4). | |||||
|
SMRC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 103) | NC score | 0.017840 (rank : 94) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
SOX17_HUMAN
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.008159 (rank : 111) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H6I2 | Gene names | SOX17 | |||
|
Domain Architecture |
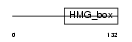
|
|||||
| Description | Transcription factor SOX-17. | |||||
|
STRN4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.004945 (rank : 121) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NRL3 | Gene names | STRN4, ZIN | |||
|
Domain Architecture |

|
|||||
| Description | Striatin-4 (Zinedin). | |||||
|
TJAP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.016812 (rank : 97) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5JTD0, Q5JTD1, Q5JWW1, Q68DB2, Q6P2P3, Q9H7V7 | Gene names | TJAP1, PILT, TJP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tight junction-associated protein 1 (Tight junction protein 4) (Protein incorporated later into tight junctions). | |||||
|
ZIM10_HUMAN
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.039590 (rank : 53) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9ULJ6, Q7Z7E6 | Gene names | RAI17, KIAA1224, ZIMP10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 17 (PIAS-like protein Zimp10). | |||||
|
CEL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.050013 (rank : 41) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
DAAM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.066055 (rank : 20) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 483 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y4D1, Q86U34, Q8N1Z8, Q8TB39 | Gene names | DAAM1, KIAA0666 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 1. | |||||
|
DAAM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.064924 (rank : 21) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BPM0, Q80Y68, Q9CQQ2 | Gene names | Daam1 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 1. | |||||
|
DAAM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.061829 (rank : 25) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86T65, Q5T4U0, Q9NQI5, Q9Y4G0 | Gene names | DAAM2, KIAA0381 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 2. | |||||
|
DAAM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.062635 (rank : 23) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 371 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80U19, Q810J5 | Gene names | Daam2, Kiaa0381 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 2. | |||||
|
DIAP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.078761 (rank : 12) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NSV4, Q5T2S7, Q5XKF6, Q6MZF0, Q6NUP0, Q86VS4, Q8NAV4 | Gene names | DIAPH3, DIAP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein diaphanous homolog 3 (Diaphanous-related formin-3) (DRF3). | |||||
|
DIAP3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.075284 (rank : 14) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Z207 | Gene names | Diaph3, Diap3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 3 (Diaphanous-related formin-3) (DRF3) (mDIA2) (p134mDIA2). | |||||
|
FHOD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.054310 (rank : 37) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
FHOD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.057204 (rank : 31) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6P9Q4, Q8BMK2 | Gene names | Fhod1, Fhos1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.057114 (rank : 33) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.057198 (rank : 32) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.054843 (rank : 35) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.052604 (rank : 40) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
SON_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.063783 (rank : 22) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
AMBN_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 107 | |
| SwissProt Accessions | Q9NP70, Q9H2X1, Q9H4L1 | Gene names | AMBN | |||
|
Domain Architecture |

|
|||||
| Description | Ameloblastin precursor. | |||||
|
AMBN_MOUSE
|
||||||
| NC score | 0.930525 (rank : 2) | θ value | 1.34075e-120 (rank : 2) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O55189 | Gene names | Ambn | |||
|
Domain Architecture |

|
|||||
| Description | Ameloblastin precursor. | |||||
|
FMN2_MOUSE
|
||||||
| NC score | 0.122818 (rank : 3) | θ value | 0.0431538 (rank : 5) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9JL04 | Gene names | Fmn2 | |||
|
Domain Architecture |

|
|||||
| Description | Formin-2. | |||||
|
FMN2_HUMAN
|
||||||
| NC score | 0.112416 (rank : 4) | θ value | 0.163984 (rank : 8) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
DIAP2_MOUSE
|
||||||
| NC score | 0.104314 (rank : 5) | θ value | 0.0431538 (rank : 4) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O70566, Q8C2G8 | Gene names | Diaph2, Diap2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2) (mDia3). | |||||
|
FMN1B_MOUSE
|
||||||
| NC score | 0.099317 (rank : 6) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q05859 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoform IV (Limb deformity protein). | |||||
|
FMN1A_MOUSE
|
||||||
| NC score | 0.098104 (rank : 7) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q05860 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoforms I/II/III (Limb deformity protein). | |||||
|
FMNL_HUMAN
|
||||||
| NC score | 0.097068 (rank : 8) | θ value | 0.813845 (rank : 15) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
Domain Architecture |

|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
DIAP2_HUMAN
|
||||||
| NC score | 0.092459 (rank : 9) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60879, O60878, Q8WX06, Q8WX48, Q9UJL2 | Gene names | DIAPH2, DIA | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2). | |||||
|
FMNL_MOUSE
|
||||||
| NC score | 0.091990 (rank : 10) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9JL26, Q6KAN4, Q9Z2V7 | Gene names | Fmnl1, Frl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-like protein 1 (Formin-related protein). | |||||
|
DIAP1_HUMAN
|
||||||
| NC score | 0.085218 (rank : 11) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O60610, Q9UC76 | Gene names | DIAPH1, DIAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1). | |||||
|
DIAP3_HUMAN
|
||||||
| NC score | 0.078761 (rank : 12) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NSV4, Q5T2S7, Q5XKF6, Q6MZF0, Q6NUP0, Q86VS4, Q8NAV4 | Gene names | DIAPH3, DIAP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein diaphanous homolog 3 (Diaphanous-related formin-3) (DRF3). | |||||
|
DIAP1_MOUSE
|
||||||
| NC score | 0.078577 (rank : 13) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 839 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O08808 | Gene names | Diaph1, Diap1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1) (mDIA1) (p140mDIA). | |||||
|
DIAP3_MOUSE
|
||||||
| NC score | 0.075284 (rank : 14) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Z207 | Gene names | Diaph3, Diap3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 3 (Diaphanous-related formin-3) (DRF3) (mDIA2) (p134mDIA2). | |||||
|
MBD6_HUMAN
|
||||||
| NC score | 0.070420 (rank : 15) | θ value | 0.813845 (rank : 16) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
SSBP4_HUMAN
|
||||||
| NC score | 0.069824 (rank : 16) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9BWG4 | Gene names | SSBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Single-stranded DNA-binding protein 4. | |||||
|
AMELX_HUMAN
|
||||||
| NC score | 0.068980 (rank : 17) | θ value | 0.813845 (rank : 13) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99217, Q96NW6, Q9UCA7 | Gene names | AMELX, AMG, AMGX | |||
|
Domain Architecture |

|
|||||
| Description | Amelogenin, X isoform precursor. | |||||
|
SYN1_HUMAN
|
||||||
| NC score | 0.067713 (rank : 18) | θ value | 1.06291 (rank : 22) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
SYN1_MOUSE
|
||||||
| NC score | 0.066879 (rank : 19) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O88935, Q62279, Q8QZT8 | Gene names | Syn1, Syn-1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I). | |||||
|
DAAM1_HUMAN
|
||||||
| NC score | 0.066055 (rank : 20) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 483 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y4D1, Q86U34, Q8N1Z8, Q8TB39 | Gene names | DAAM1, KIAA0666 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 1. | |||||
|
DAAM1_MOUSE
|
||||||
| NC score | 0.064924 (rank : 21) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BPM0, Q80Y68, Q9CQQ2 | Gene names | Daam1 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 1. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.063783 (rank : 22) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
DAAM2_MOUSE
|
||||||
| NC score | 0.062635 (rank : 23) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 371 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80U19, Q810J5 | Gene names | Daam2, Kiaa0381 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 2. | |||||
|
PRAX_HUMAN
|
||||||
| NC score | 0.062528 (rank : 24) | θ value | 1.06291 (rank : 20) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BXM0, Q9BXL9, Q9HCF2 | Gene names | PRX, KIAA1620 | |||
|
Domain Architecture |

|
|||||
| Description | Periaxin. | |||||
|
DAAM2_HUMAN
|
||||||
| NC score | 0.061829 (rank : 25) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86T65, Q5T4U0, Q9NQI5, Q9Y4G0 | Gene names | DAAM2, KIAA0381 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 2. | |||||
|
SYN2_MOUSE
|
||||||
| NC score | 0.060683 (rank : 26) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64332, Q6NZR0, Q9QWV7 | Gene names | Syn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
AMELX_MOUSE
|
||||||
| NC score | 0.059373 (rank : 27) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P63277, P45559, P70592, Q61293 | Gene names | Amelx, Amel | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amelogenin, X isoform precursor (Leucine-rich amelogenin peptide) (LRAP). | |||||
|
PRAX_MOUSE
|
||||||
| NC score | 0.058870 (rank : 28) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O55103, O55104 | Gene names | Prx | |||
|
Domain Architecture |

|
|||||
| Description | Periaxin. | |||||
|
SYN2_HUMAN
|
||||||
| NC score | 0.057797 (rank : 29) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92777 | Gene names | SYN2 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
AIRE_HUMAN
|
||||||
| NC score | 0.057290 (rank : 30) | θ value | 0.0563607 (rank : 6) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
FHOD1_MOUSE
|
||||||
| NC score | 0.057204 (rank : 31) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6P9Q4, Q8BMK2 | Gene names | Fhod1, Fhos1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.057198 (rank : 32) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.057114 (rank : 33) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
3BP2_HUMAN
|
||||||
| NC score | 0.055571 (rank : 34) | θ value | 1.38821 (rank : 24) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P78314, O00500, O15373, P78315 | Gene names | SH3BP2, 3BP2 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 2 (3BP-2). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.054843 (rank : 35) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
ASPP1_MOUSE
|
||||||
| NC score | 0.054497 (rank : 36) | θ value | 0.00665767 (rank : 3) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q62415 | Gene names | Ppp1r13b, Aspp1 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 1 (Protein phosphatase 1 regulatory subunit 13B). | |||||
|
FHOD1_HUMAN
|
||||||
| NC score | 0.054310 (rank : 37) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.053774 (rank : 38) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
R3HD2_MOUSE
|
||||||
| NC score | 0.053028 (rank : 39) | θ value | 1.06291 (rank : 21) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.052604 (rank : 40) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.050013 (rank : 41) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
WBP11_MOUSE
|
||||||
| NC score | 0.048598 (rank : 42) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q923D5, O88539, Q3UMD8, Q8VDI0 | Gene names | Wbp11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11). | |||||
|
MTSS1_HUMAN
|
||||||
| NC score | 0.047948 (rank : 43) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O43312, Q8TCA2, Q96RX2 | Gene names | MTSS1, KIAA0429, MIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein) (Metastasis suppressor YGL-1). | |||||
|
TACC3_MOUSE
|
||||||
| NC score | 0.047238 (rank : 44) | θ value | 0.813845 (rank : 17) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JJ11, Q9WVK9 | Gene names | Tacc3, Aint | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ARNT-interacting protein). | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.045514 (rank : 45) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
WBP11_HUMAN
|
||||||
| NC score | 0.045432 (rank : 46) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y2W2, Q96AY8 | Gene names | WBP11, NPWBP, SNP70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11) (SH3 domain-binding protein SNP70) (Npw38-binding protein) (NpwBP). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.043874 (rank : 47) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
HD_MOUSE
|
||||||
| NC score | 0.043800 (rank : 48) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P42859 | Gene names | Hd, Hdh | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein homolog) (HD protein). | |||||
|
IF4G1_HUMAN
|
||||||
| NC score | 0.043698 (rank : 49) | θ value | 0.279714 (rank : 9) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
SHAN2_MOUSE
|
||||||
| NC score | 0.043254 (rank : 50) | θ value | 0.279714 (rank : 10) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 446 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80Z38, Q3UTK4, Q5DU07 | Gene names | Shank2, Kiaa1022 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2) (Cortactin- binding protein 1) (CortBP1). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.041915 (rank : 51) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.040827 (rank : 52) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
ZIM10_HUMAN
|
||||||
| NC score | 0.039590 (rank : 53) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9ULJ6, Q7Z7E6 | Gene names | RAI17, KIAA1224, ZIMP10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 17 (PIAS-like protein Zimp10). | |||||
|
SYP2L_MOUSE
|
||||||
| NC score | 0.039539 (rank : 54) | θ value | 0.62314 (rank : 12) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BWB1 | Gene names | Synpo2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin 2-like protein. | |||||
|
HD_HUMAN
|
||||||
| NC score | 0.039209 (rank : 55) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
RBM12_MOUSE
|
||||||
| NC score | 0.039099 (rank : 56) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8R4X3, Q8K373, Q8R302, Q9CS80 | Gene names | Rbm12 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
CPSF6_MOUSE
|
||||||
| NC score | 0.037058 (rank : 57) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
CPSF6_HUMAN
|
||||||
| NC score | 0.036964 (rank : 58) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q16630, Q53ES1, Q9BSJ7, Q9BW18 | Gene names | CPSF6, CFIM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6 (Cleavage and polyadenylation specificity factor 68 kDa subunit) (CPSF 68 kDa subunit) (Pre-mRNA cleavage factor Im 68 kDa subunit) (Protein HPBRII- 4/7). | |||||
|
DAB1_HUMAN
|
||||||
| NC score | 0.036933 (rank : 59) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75553, Q9NYA8 | Gene names | DAB1 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 1. | |||||
|
K0562_HUMAN
|
||||||
| NC score | 0.036218 (rank : 60) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60308, Q5JSQ3, Q5SR24, Q5SR25, Q6PKF5, Q86W32, Q86X14 | Gene names | KIAA0562 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0562. | |||||
|
DAB1_MOUSE
|
||||||
| NC score | 0.036080 (rank : 61) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P97318, P97316, P97317 | Gene names | Dab1 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 1. | |||||
|
DCP1A_HUMAN
|
||||||
| NC score | 0.035269 (rank : 62) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NPI6 | Gene names | DCP1A, SMIF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (Smad4-interacting transcriptional co-activator). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.035159 (rank : 63) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
NFAC1_HUMAN
|
||||||
| NC score | 0.034376 (rank : 64) | θ value | 1.06291 (rank : 19) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O95644, Q12865, Q15793 | Gene names | NFATC1, NFAT2, NFATC | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 1 (NFAT transcription complex cytosolic component) (NF-ATc1) (NF-ATc). | |||||
|
PRB4S_HUMAN
|
||||||
| NC score | 0.034215 (rank : 65) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.032978 (rank : 66) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
ZFHX2_HUMAN
|
||||||
| NC score | 0.032408 (rank : 67) | θ value | 0.47712 (rank : 11) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9C0A1 | Gene names | ZFHX2, KIAA1762 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger homeobox protein 2. | |||||
|
ZN687_HUMAN
|
||||||
| NC score | 0.032045 (rank : 68) | θ value | 0.125558 (rank : 7) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
GORS2_HUMAN
|
||||||
| NC score | 0.030109 (rank : 69) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H8Y8, Q96I74, Q96K84, Q9H946, Q9UFW4 | Gene names | GORASP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 2 (GRS2) (Golgi reassembly-stacking protein of 55 kDa) (GRASP55) (p59) (Golgi phosphoprotein 6) (GOLPH6). | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.030083 (rank : 70) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.029203 (rank : 71) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.029041 (rank : 72) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.028905 (rank : 73) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
RBM16_HUMAN
|
||||||
| NC score | 0.028519 (rank : 74) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UPN6, Q5TBU6, Q9BQN8, Q9BX43 | Gene names | RBM16, KIAA1116 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein 16 (RNA-binding motif protein 16). | |||||
|
IL3RB_MOUSE
|
||||||
| NC score | 0.028237 (rank : 75) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 355 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P26955 | Gene names | Csf2rb, Aic2b, Csf2rb1, Il3rb1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytokine receptor common beta chain precursor (GM-CSF/IL-3/IL-5 receptor common beta-chain) (CD131 antigen). | |||||
|
TNFB_MOUSE
|
||||||
| NC score | 0.027254 (rank : 76) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P09225 | Gene names | Lta, Tnfb, Tnfsf1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphotoxin-alpha precursor (LT-alpha) (TNF-beta) (Tumor necrosis factor ligand superfamily member 1). | |||||
|
CO4A3_HUMAN
|
||||||
| NC score | 0.026680 (rank : 77) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q01955, Q9BQT2 | Gene names | COL4A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(IV) chain precursor (Goodpasture antigen). | |||||
|
BSCL2_MOUSE
|
||||||
| NC score | 0.026465 (rank : 78) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z2E9, Q810B0, Q9JJC2, Q9JMF1 | Gene names | Bscl2, Gng3lg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Seipin (Bernardinelli-Seip congenital lipodystrophy type 2 protein homolog). | |||||
|
CO4A5_HUMAN
|
||||||
| NC score | 0.024683 (rank : 79) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P29400, Q16006, Q16126, Q6LD84 | Gene names | COL4A5 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-5(IV) chain precursor. | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.024461 (rank : 80) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SFR14_HUMAN
|
||||||
| NC score | 0.024286 (rank : 81) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IX01, O15071, O60369, Q5JPH7, Q8WUF7 | Gene names | SFRS14, KIAA0365 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
R51A1_HUMAN
|
||||||
| NC score | 0.024183 (rank : 82) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96B01, O43403, Q7Z779 | Gene names | RAD51AP1, PIR51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAD51-associated protein 1 (RAD51-interacting protein). | |||||
|
HGS_MOUSE
|
||||||
| NC score | 0.023945 (rank : 83) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
CO4A2_HUMAN
|
||||||
| NC score | 0.023655 (rank : 84) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 454 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P08572 | Gene names | COL4A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
DREB_HUMAN
|
||||||
| NC score | 0.023108 (rank : 85) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q16643 | Gene names | DBN1 | |||
|
Domain Architecture |

|
|||||
| Description | Drebrin (Developmentally-regulated brain protein). | |||||
|
CO5A1_HUMAN
|
||||||
| NC score | 0.023078 (rank : 86) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P20908, Q15094, Q5SUX4 | Gene names | COL5A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
WASP_MOUSE
|
||||||
| NC score | 0.022959 (rank : 87) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70315 | Gene names | Was, Wasp | |||
|
Domain Architecture |

|
|||||
| Description | Wiskott-Aldrich syndrome protein homolog (WASp). | |||||
|
CO9A1_MOUSE
|
||||||
| NC score | 0.022820 (rank : 88) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q05722, Q61269, Q61270, Q61433, Q61940 | Gene names | Col9a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
SPON1_HUMAN
|
||||||
| NC score | 0.022048 (rank : 89) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HCB6, O94862, Q8NCD7, Q8WUR5 | Gene names | SPON1, KIAA0762, VSGP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spondin-1 precursor (F-spondin) (Vascular smooth muscle cell growth- promoting factor). | |||||
|
SPON1_MOUSE
|
||||||
| NC score | 0.021980 (rank : 90) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VCC9, Q8K2Q8 | Gene names | Spon1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spondin-1 precursor (F-spondin). | |||||
|
CO3A1_HUMAN
|
||||||
| NC score | 0.020628 (rank : 91) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
CO9A2_MOUSE
|
||||||
| NC score | 0.018935 (rank : 92) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q07643 | Gene names | Col9a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(IX) chain precursor. | |||||
|
PLXB1_HUMAN
|
||||||
| NC score | 0.018715 (rank : 93) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43157, Q6NY20, Q9UIV7, Q9UJ92, Q9UJ93 | Gene names | PLXNB1, KIAA0407, SEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor (Semaphorin receptor SEP). | |||||
|
SMRC2_HUMAN
|
||||||
| NC score | 0.017840 (rank : 94) | θ value | 8.99809 (rank : 103) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
PRDM2_HUMAN
|
||||||
| NC score | 0.017329 (rank : 95) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
SNX18_MOUSE
|
||||||
| NC score | 0.016898 (rank : 96) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91ZR2 | Gene names | Snag1, Snx18 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-18 (Sorting nexin-associated Golgi protein 1). | |||||
|
TJAP1_HUMAN
|
||||||
| NC score | 0.016812 (rank : 97) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5JTD0, Q5JTD1, Q5JWW1, Q68DB2, Q6P2P3, Q9H7V7 | Gene names | TJAP1, PILT, TJP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tight junction-associated protein 1 (Tight junction protein 4) (Protein incorporated later into tight junctions). | |||||
|
DEN1A_HUMAN
|
||||||
| NC score | 0.016338 (rank : 98) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TEH3, Q5VWF0, Q8IVD6, Q9H796 | Gene names | DENND1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
DDEF1_MOUSE
|
||||||
| NC score | 0.015013 (rank : 99) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QWY8, O08612, Q80TG8, Q80UV6, Q99LV8, Q9Z2B6 | Gene names | Ddef1, Asap1, Kiaa1249, Shag1 | |||
|
Domain Architecture |

|
|||||
| Description | 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase- activating protein (PIP2-dependent ARF1 GAP) (ADP-ribosylation factor- directed GTPase-activating protein 1) (ARF GTPase-activating protein 1) (Development and differentiation-enhancing factor 1) (Differentiation-enhancing factor 1) (DEF-1). | |||||
|
FBX46_MOUSE
|
||||||
| NC score | 0.014750 (rank : 100) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BG80 | Gene names | Fbxo46, Fbx46, Fbxo34l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
TIF1B_MOUSE
|
||||||
| NC score | 0.014152 (rank : 101) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62318, P70391, Q8C283, Q99PN4 | Gene names | Trim28, Krip1, Tif1b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (KRAB-A-interacting protein) (KRIP-1). | |||||
|
CSPG3_MOUSE
|
||||||
| NC score | 0.014039 (rank : 102) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P55066, Q6P1E3, Q8C4F8 | Gene names | Cspg3, Ncan | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
ABI3_HUMAN
|
||||||
| NC score | 0.013927 (rank : 103) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2A4, Q9H0P6 | Gene names | ABI3, NESH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABI gene family member 3 (New molecule including SH3) (Nesh). | |||||
|
RX_HUMAN
|
||||||
| NC score | 0.012343 (rank : 104) | θ value | 2.36792 (rank : 43) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2V3 | Gene names | RAX, RX | |||
|
Domain Architecture |

|
|||||
| Description | Retinal homeobox protein Rx (Retina and anterior neural fold homeobox protein). | |||||
|
RIN2_HUMAN
|
||||||
| NC score | 0.012195 (rank : 105) | θ value | 8.99809 (rank : 102) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WYP3, Q00425, Q5TFT8, Q9BQL3, Q9H071 | Gene names | RIN2, RASSF4 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2) (Ras inhibitor JC265) (Ras association domain family 4). | |||||
|
MTMR7_HUMAN
|
||||||
| NC score | 0.012036 (rank : 106) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y216, Q68DX4 | Gene names | MTMR7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
USP9X_MOUSE
|
||||||
| NC score | 0.011072 (rank : 107) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
DRD4_HUMAN
|
||||||
| NC score | 0.009505 (rank : 108) | θ value | 0.813845 (rank : 14) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P21917 | Gene names | DRD4 | |||
|
Domain Architecture |

|
|||||
| Description | D(4) dopamine receptor (Dopamine D4 receptor) (D(2C) dopamine receptor). | |||||
|
B3A2_HUMAN
|
||||||
| NC score | 0.008950 (rank : 109) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P04920, Q45EY5, Q969L3 | Gene names | SLC4A2, AE2, EPB3L1, HKB3, MPB3L | |||
|
Domain Architecture |

|
|||||
| Description | Anion exchange protein 2 (Non-erythroid band 3-like protein) (AE2 anion exchanger) (Solute carrier family 4 member 2) (BND3L). | |||||
|
AAK1_MOUSE
|
||||||
| NC score | 0.008711 (rank : 110) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q3UHJ0, Q3TY53, Q6ZPZ6, Q80XP6 | Gene names | Aak1, Kiaa1048 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP2-associated protein kinase 1 (EC 2.7.11.1) (Adaptor-associated kinase 1). | |||||
|
SOX17_HUMAN
|
||||||
| NC score | 0.008159 (rank : 111) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H6I2 | Gene names | SOX17 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-17. | |||||
|
MTF1_HUMAN
|
||||||
| NC score | 0.007865 (rank : 112) | θ value | 1.38821 (rank : 29) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 814 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14872, Q96CB1 | Gene names | MTF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
PCNT_HUMAN
|
||||||
| NC score | 0.007621 (rank : 113) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
LHX1_MOUSE
|
||||||
| NC score | 0.006899 (rank : 114) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P63006, P36199 | Gene names | Lhx1, Lim-1, Lim1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM/homeobox protein Lhx1 (LIM homeobox protein 1) (Homeobox protein Lim-1). | |||||
|
TSK_MOUSE
|
||||||
| NC score | 0.006674 (rank : 115) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CBR6, Q4W655 | Gene names | Lrrc54, Tsk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tsukushi precursor (Leucine-rich repeat-containing protein 54). | |||||
|
AAK1_HUMAN
|
||||||
| NC score | 0.006521 (rank : 116) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 1077 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q2M2I8, Q4ZFZ3, Q53RX6, Q9UPV4 | Gene names | AAK1, KIAA1048 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP2-associated protein kinase 1 (EC 2.7.11.1) (Adaptor-associated kinase 1). | |||||
|
MTF1_MOUSE
|
||||||
| NC score | 0.006243 (rank : 117) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q07243 | Gene names | Mtf1, Mtf-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
KLF2_MOUSE
|
||||||
| NC score | 0.005263 (rank : 118) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q60843 | Gene names | Klf2, Lklf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 2 (Lung krueppel-like factor). | |||||
|
KI13B_HUMAN
|
||||||
| NC score | 0.005208 (rank : 119) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NQT8, O75134, Q9BYJ6 | Gene names | KIF13B, GAKIN, KIAA0639 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13B (Kinesin-like protein GAKIN). | |||||
|
ZN206_HUMAN
|
||||||
| NC score | 0.004970 (rank : 120) | θ value | 1.38821 (rank : 30) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 853 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96SZ4 | Gene names | ZNF206, ZSCAN10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 206 (Zinc finger and SCAN domain-containing protein 10). | |||||
|
STRN4_HUMAN
|
||||||
| NC score | 0.004945 (rank : 121) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 107 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NRL3 | Gene names | STRN4, ZIN | |||
|
Domain Architecture |

|
|||||
| Description | Striatin-4 (Zinedin). | |||||