Please be patient as the page loads
|
GORS2_HUMAN
|
||||||
| SwissProt Accessions | Q9H8Y8, Q96I74, Q96K84, Q9H946, Q9UFW4 | Gene names | GORASP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 2 (GRS2) (Golgi reassembly-stacking protein of 55 kDa) (GRASP55) (p59) (Golgi phosphoprotein 6) (GOLPH6). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
GORS2_MOUSE
|
||||||
| θ value | 1.15296e-180 (rank : 1) | NC score | 0.966946 (rank : 2) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99JX3, Q3U9D2, Q8CCD0, Q91W68 | Gene names | Gorasp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 2 (GRS2) (Golgi reassembly-stacking protein of 55 kDa) (GRASP55). | |||||
|
GORS2_HUMAN
|
||||||
| θ value | 6.14192e-166 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9H8Y8, Q96I74, Q96K84, Q9H946, Q9UFW4 | Gene names | GORASP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 2 (GRS2) (Golgi reassembly-stacking protein of 55 kDa) (GRASP55) (p59) (Golgi phosphoprotein 6) (GOLPH6). | |||||
|
GR65_HUMAN
|
||||||
| θ value | 1.54831e-84 (rank : 3) | NC score | 0.904480 (rank : 4) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BQQ3, Q8N272, Q96H42 | Gene names | GORASP1, GRASP65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 1 (Golgi reassembly-stacking protein of 65 kDa) (GRASP65) (Golgi peripheral membrane protein p65) (Golgi phosphoprotein 5) (GOLPH5). | |||||
|
GR65_MOUSE
|
||||||
| θ value | 1.71185e-83 (rank : 4) | NC score | 0.920037 (rank : 3) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q91X51, Q9D3L9 | Gene names | Gorasp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 1 (Golgi reassembly-stacking protein of 65 kDa) (GRASP65) (Golgi peripheral membrane protein p65). | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 5) | NC score | 0.100444 (rank : 11) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
KLF17_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 6) | NC score | 0.023585 (rank : 101) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 882 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5JT82, Q86VQ7, Q8N805 | Gene names | KLF17, ZNF393 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 17 (Zinc finger protein 393). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 7) | NC score | 0.114612 (rank : 8) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 8) | NC score | 0.074653 (rank : 23) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 9) | NC score | 0.100680 (rank : 10) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
CIC_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 10) | NC score | 0.090183 (rank : 15) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
CEL_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 11) | NC score | 0.048283 (rank : 78) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
PRAX_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 12) | NC score | 0.093031 (rank : 13) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BXM0, Q9BXL9, Q9HCF2 | Gene names | PRX, KIAA1620 | |||
|
Domain Architecture |
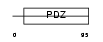
|
|||||
| Description | Periaxin. | |||||
|
NCOA6_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 13) | NC score | 0.080484 (rank : 20) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
ZEP1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 14) | NC score | 0.039637 (rank : 85) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
K1543_MOUSE
|
||||||
| θ value | 0.163984 (rank : 15) | NC score | 0.078113 (rank : 22) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80VC9, Q5DTW9, Q8BUZ0 | Gene names | Kiaa1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
NFRKB_MOUSE
|
||||||
| θ value | 0.163984 (rank : 16) | NC score | 0.092231 (rank : 14) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.049432 (rank : 75) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
PSMD9_HUMAN
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.173901 (rank : 6) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O00233, Q9BQ42 | Gene names | PSMD9 | |||
|
Domain Architecture |

|
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 9 (26S proteasome regulatory subunit p27). | |||||
|
SON_MOUSE
|
||||||
| θ value | 0.279714 (rank : 19) | NC score | 0.122171 (rank : 7) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
CSPG3_MOUSE
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.023193 (rank : 102) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P55066, Q6P1E3, Q8C4F8 | Gene names | Cspg3, Ncan | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
DCP1A_HUMAN
|
||||||
| θ value | 0.365318 (rank : 21) | NC score | 0.050952 (rank : 70) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NPI6 | Gene names | DCP1A, SMIF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (Smad4-interacting transcriptional co-activator). | |||||
|
PSMD9_MOUSE
|
||||||
| θ value | 0.365318 (rank : 22) | NC score | 0.184822 (rank : 5) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9CR00, Q3TG66 | Gene names | Psmd9 | |||
|
Domain Architecture |

|
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 9 (26S proteasome regulatory subunit p27). | |||||
|
K1543_HUMAN
|
||||||
| θ value | 0.47712 (rank : 23) | NC score | 0.084522 (rank : 18) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
LEUK_HUMAN
|
||||||
| θ value | 0.47712 (rank : 24) | NC score | 0.069188 (rank : 27) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P16150 | Gene names | SPN, CD43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukosialin precursor (Leukocyte sialoglycoprotein) (Sialophorin) (Galactoglycoprotein) (GALGP) (CD43 antigen). | |||||
|
PRAX_MOUSE
|
||||||
| θ value | 0.47712 (rank : 25) | NC score | 0.103436 (rank : 9) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O55103, O55104 | Gene names | Prx | |||
|
Domain Architecture |
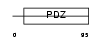
|
|||||
| Description | Periaxin. | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 26) | NC score | 0.057902 (rank : 43) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
PZRN4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.031253 (rank : 91) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZMN7, Q6N052, Q8IUU1, Q9NTP7 | Gene names | PDZRN4, LNX4, SEMCAP3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing RING finger protein 4 (Ligand of Numb-protein X 4) (SEMACAP3-like protein). | |||||
|
TRI66_HUMAN
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.055935 (rank : 50) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
U689_HUMAN
|
||||||
| θ value | 0.62314 (rank : 29) | NC score | 0.052795 (rank : 59) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6UX39 | Gene names | ames=UNQ689/PRO1329 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein UNQ689/PRO1329 precursor. | |||||
|
F120C_HUMAN
|
||||||
| θ value | 0.813845 (rank : 30) | NC score | 0.066729 (rank : 30) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NX05 | Gene names | FAM120C, CXorf17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120C (Tumor antigen BJ-HCC-21). | |||||
|
F120C_MOUSE
|
||||||
| θ value | 0.813845 (rank : 31) | NC score | 0.060631 (rank : 40) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C3F2 | Gene names | Fam120c, ORF34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120C. | |||||
|
MINT_MOUSE
|
||||||
| θ value | 1.38821 (rank : 32) | NC score | 0.049741 (rank : 74) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
SIGL7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 33) | NC score | 0.018506 (rank : 108) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y286, Q9NZQ1, Q9UJ86, Q9UJ87, Q9Y502 | Gene names | SIGLEC7, AIRM1 | |||
|
Domain Architecture |

|
|||||
| Description | Sialic acid-binding Ig-like lectin 7 precursor (Siglec-7) (QA79 membrane protein) (Adhesion inhibitory receptor molecule 1) (AIRM-1) (p75) (D-siglec) (CDw328 antigen). | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.017533 (rank : 109) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
LAP4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 35) | NC score | 0.020588 (rank : 104) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80U72, Q6P5H7, Q7TPH8, Q80VB1, Q8CI48, Q8VII1, Q922S3 | Gene names | Scrib, Kiaa0147, Lap4, Scrib1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog). | |||||
|
PER2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 36) | NC score | 0.031932 (rank : 90) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
SSBP4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 37) | NC score | 0.032303 (rank : 89) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BWG4 | Gene names | SSBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Single-stranded DNA-binding protein 4. | |||||
|
ZEP1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 38) | NC score | 0.035098 (rank : 88) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1063 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q03172 | Gene names | Hivep1, Cryabp1, Znf40 | |||
|
Domain Architecture |
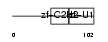
|
|||||
| Description | Zinc finger protein 40 (Transcription factor alphaA-CRYBP1) (Alpha A- crystallin-binding protein I) (Alpha A-CRYBP1). | |||||
|
ATF2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.043165 (rank : 83) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P15336, Q13000 | Gene names | ATF2, CREB2, CREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (HB16). | |||||
|
ATF2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.042032 (rank : 84) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P16951, Q64089, Q64090, Q64091 | Gene names | Atf2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (MXBP protein). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.057936 (rank : 42) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CP007_MOUSE
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.028246 (rank : 96) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C190, Q6PFD1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 homolog (5-day ovary-specific transcript 1 protein). | |||||
|
FOXL1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 43) | NC score | 0.008051 (rank : 124) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64731 | Gene names | Foxl1, Fkh6, Fkhl11 | |||
|
Domain Architecture |
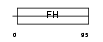
|
|||||
| Description | Forkhead box protein L1 (Forkhead-related protein FKHL11) (Transcription factor FKH-6). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.026960 (rank : 100) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SSPO_MOUSE
|
||||||
| θ value | 2.36792 (rank : 45) | NC score | 0.012704 (rank : 121) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
CMTA2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.044699 (rank : 82) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94983, Q7Z6M8, Q8N3V0, Q8NDG4, Q96G17 | Gene names | CAMTA2, KIAA0909 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-binding transcription activator 2. | |||||
|
CUTL1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.020389 (rank : 105) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.048117 (rank : 79) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
TAU_HUMAN
|
||||||
| θ value | 3.0926 (rank : 49) | NC score | 0.045139 (rank : 80) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
TM1L1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 50) | NC score | 0.028897 (rank : 94) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q923U0, Q99KE0, Q9D6Y5 | Gene names | Tom1l1, Srcasm | |||
|
Domain Architecture |
|
|||||
| Description | TOM1-like 1 protein (Target of myb-like 1 protein) (Src-activating and signaling molecule protein). | |||||
|
CIC_MOUSE
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.066359 (rank : 32) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
DAB2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.027767 (rank : 98) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P98082, Q13598, Q9BTY0, Q9UK04 | Gene names | DAB2, DOC2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (Differentially expressed protein 2) (DOC-2). | |||||
|
FINC_HUMAN
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.016857 (rank : 110) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P02751, O95609, O95610, Q14312, Q14325, Q14326, Q6LDP6, Q86T27, Q8IVI8, Q96KP7, Q96KP8, Q96KP9, Q9H1B8, Q9HAP3, Q9UMK2 | Gene names | FN1, FN | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN) (Cold-insoluble globulin) (CIG). | |||||
|
LAP4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.019517 (rank : 106) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 608 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14160, Q6P496, Q7Z5D1, Q8WWV8, Q96C69, Q96GG1 | Gene names | SCRIB, CRIB1, KIAA0147, LAP4, SCRB1, VARTUL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog) (hScrib). | |||||
|
ATX1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.035210 (rank : 87) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54254 | Gene names | Atxn1, Sca1 | |||
|
Domain Architecture |

|
|||||
| Description | Ataxin-1 (Spinocerebellar ataxia type 1 protein homolog). | |||||
|
CBPD_MOUSE
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.008972 (rank : 123) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O89001 | Gene names | Cpd | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase D precursor (EC 3.4.17.22) (Metallocarboxypeptidase D) (gp180). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.056948 (rank : 47) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CYLD_HUMAN
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.016501 (rank : 112) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NQC7, O94934, Q7L3N6, Q96EH0, Q9NZX9 | Gene names | CYLD, CYLD1, KIAA0849 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase CYLD (EC 3.1.2.15) (Ubiquitin thioesterase CYLD) (Ubiquitin-specific-processing protease CYLD) (Deubiquitinating enzyme CYLD). | |||||
|
GLI3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.006141 (rank : 125) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61602 | Gene names | Gli3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI3. | |||||
|
HCFC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.048962 (rank : 76) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61191, Q684R1, Q7TSB0, Q8C2D0, Q9QWH2 | Gene names | Hcfc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N- terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C-terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
MBN3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.014424 (rank : 117) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NUK0, Q5JXN8, Q5JXN9, Q5JXP4, Q6UDQ1, Q8TAD9, Q8TAF4, Q9H0Z7, Q9UF37 | Gene names | MBNL3, CHCR, MBLX39, MBXL | |||
|
Domain Architecture |

|
|||||
| Description | Muscleblind-like X-linked protein (Muscleblind-like protein 3) (Cys3His CCG1-required protein) (Protein HCHCR). | |||||
|
MPDZ_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.016256 (rank : 113) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75970, O43798, Q5CZ80, Q5JTX3, Q5JTX6, Q5JTX7, Q5JUC3, Q5JUC4, Q5VZ62, Q8N790 | Gene names | MPDZ, MUPP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple PDZ domain protein (Multi PDZ domain protein 1) (Multi-PDZ- domain protein 1). | |||||
|
SIG12_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.014410 (rank : 118) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96PQ1, Q8IYH7 | Gene names | SIGLEC12, SIGLECL1, SLG | |||
|
Domain Architecture |

|
|||||
| Description | Sialic acid-binding Ig-like lectin 12 precursor (Siglec-12) (Sialic acid-binding Ig-like lectin-like 1) (Siglec-L1). | |||||
|
YTHD3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.016198 (rank : 114) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BYK6, Q3UVI5, Q6NXJ6, Q6NXJ8, Q8BKB6 | Gene names | Ythdf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YTH domain family protein 3. | |||||
|
ZN385_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.027314 (rank : 99) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
ZN687_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.012965 (rank : 120) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
AMBN_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.030109 (rank : 93) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NP70, Q9H2X1, Q9H4L1 | Gene names | AMBN | |||
|
Domain Architecture |
|
|||||
| Description | Ameloblastin precursor. | |||||
|
AMBN_MOUSE
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.028236 (rank : 97) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O55189 | Gene names | Ambn | |||
|
Domain Architecture |
|
|||||
| Description | Ameloblastin precursor. | |||||
|
CJ012_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.028261 (rank : 95) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
CK054_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.016786 (rank : 111) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 3 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H0W9, Q6FI88, Q6XYB0, Q96EI3, Q96IX1, Q9Y6B4 | Gene names | C11orf54 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ester hydrolase C11orf54 (EC 3.1.-.-). | |||||
|
CO039_MOUSE
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.037742 (rank : 86) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
HCFC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.048476 (rank : 77) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P51610, Q6P4G5 | Gene names | HCFC1, HCF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) (VP16 accessory protein) (VCAF) (CFF) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N-terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C- terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
ARID2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.030210 (rank : 92) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q68CP9, Q5EB51, Q645I3, Q6ZRY5, Q7Z3I5, Q86T28, Q96SJ6, Q9HCL5 | Gene names | ARID2, KIAA1557 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 2 (ARID domain- containing protein 2) (BRG1-associated factor 200) (BAF200). | |||||
|
BLNK_HUMAN
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.014745 (rank : 116) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WV28, O75498, O75499 | Gene names | BLNK, BASH, SLP65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell linker protein (Cytoplasmic adapter protein) (B-cell adapter containing SH2 domain protein) (B-cell adapter containing Src homology 2 domain protein) (Src homology 2 domain-containing leukocyte protein of 65 kDa) (SLP-65). | |||||
|
COCA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.012489 (rank : 122) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99715, Q99716 | Gene names | COL12A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
DOT1L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.044723 (rank : 81) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
FINC_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.015928 (rank : 115) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P11276, Q61567, Q61568, Q61569, Q64233, Q80UI4 | Gene names | Fn1 | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.062695 (rank : 33) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SMPX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.018827 (rank : 107) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UHP9 | Gene names | SMPX, SRMX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small muscular protein (Stretch-responsive skeletal muscle protein). | |||||
|
TENS4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.021145 (rank : 103) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IZW8, Q71RB7, Q8WV64, Q96JV4 | Gene names | TNS4, CTEN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-4 precursor (C-terminal tensin-like protein). | |||||
|
TNFC_MOUSE
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.013238 (rank : 119) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P41155 | Gene names | Ltb, Tnfc, Tnfsf3 | |||
|
Domain Architecture |
|
|||||
| Description | Lymphotoxin-beta (LT-beta) (Tumor necrosis factor C) (TNF-C) (Tumor necrosis factor ligand superfamily member 3). | |||||
|
ZN438_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.005627 (rank : 126) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z4V0, Q5T426, Q658Q4, Q6ZN65 | Gene names | ZNF438 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 438. | |||||
|
ATN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.052242 (rank : 65) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
CF015_HUMAN
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.067016 (rank : 29) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6UXA7, Q5SQ81, Q86Z05, Q9UIG3 | Gene names | C6orf15, STG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf15 precursor (Protein STG). | |||||
|
CN032_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.056848 (rank : 48) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
CN032_MOUSE
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.055069 (rank : 54) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BH93, Q3TF12, Q3TGL0, Q8CC90 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32 homolog. | |||||
|
CN155_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.073660 (rank : 25) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
CO002_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.055847 (rank : 51) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
CPSF6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.051371 (rank : 69) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
CS016_HUMAN
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.052558 (rank : 62) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8ND99 | Gene names | C19orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf16. | |||||
|
ENAH_MOUSE
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.059276 (rank : 41) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
EP300_HUMAN
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.054950 (rank : 55) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
FMN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.050049 (rank : 72) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
FMN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.052344 (rank : 64) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JL04 | Gene names | Fmn2 | |||
|
Domain Architecture |

|
|||||
| Description | Formin-2. | |||||
|
FMNL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.055945 (rank : 49) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
Domain Architecture |
|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
GGN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.052477 (rank : 63) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
GSCR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.089550 (rank : 16) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
MBD6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.067340 (rank : 28) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.050351 (rank : 71) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
MUC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.052202 (rank : 66) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
MUC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.062044 (rank : 36) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.089294 (rank : 17) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.061358 (rank : 38) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
NFRKB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.061949 (rank : 37) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6P4R8, Q12869, Q15312, Q9H048 | Gene names | NFRKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
NU214_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.052132 (rank : 67) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
PO121_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.057867 (rank : 44) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.055480 (rank : 52) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.069584 (rank : 26) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PRG4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.055338 (rank : 53) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PRP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.057222 (rank : 46) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
PRP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.066686 (rank : 31) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
PRR12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.073804 (rank : 24) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.057628 (rank : 45) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
SF01_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.060658 (rank : 39) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q15637, Q14818, Q14819, Q15913, Q8IY00, Q92744, Q92745, Q969H7, Q9BW01, Q9UEI0 | Gene names | SF1, ZFM1, ZNF162 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (Zinc finger gene in MEN1 locus) (Mammalian branch point-binding protein mBBP) (BBP). | |||||
|
SF3A2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.053710 (rank : 57) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
SF3A2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.052956 (rank : 58) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
SFR15_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.052644 (rank : 61) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O95104, Q6P1M5, Q8N3I8, Q9UFM1, Q9ULP8 | Gene names | SFRS15, KIAA1172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 15 (CTD-binding SR-like protein RA4). | |||||
|
T22D2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.053812 (rank : 56) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
TAF4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.062306 (rank : 35) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.096127 (rank : 12) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
WASIP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.083547 (rank : 19) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
WBP11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.051567 (rank : 68) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y2W2, Q96AY8 | Gene names | WBP11, NPWBP, SNP70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11) (SH3 domain-binding protein SNP70) (Npw38-binding protein) (NpwBP). | |||||
|
WIRE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.052698 (rank : 60) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
WIRE_MOUSE
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.050028 (rank : 73) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.062690 (rank : 34) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
ZN207_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.078702 (rank : 21) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
GORS2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 6.14192e-166 (rank : 2) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9H8Y8, Q96I74, Q96K84, Q9H946, Q9UFW4 | Gene names | GORASP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 2 (GRS2) (Golgi reassembly-stacking protein of 55 kDa) (GRASP55) (p59) (Golgi phosphoprotein 6) (GOLPH6). | |||||
|
GORS2_MOUSE
|
||||||
| NC score | 0.966946 (rank : 2) | θ value | 1.15296e-180 (rank : 1) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99JX3, Q3U9D2, Q8CCD0, Q91W68 | Gene names | Gorasp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 2 (GRS2) (Golgi reassembly-stacking protein of 55 kDa) (GRASP55). | |||||
|
GR65_MOUSE
|
||||||
| NC score | 0.920037 (rank : 3) | θ value | 1.71185e-83 (rank : 4) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q91X51, Q9D3L9 | Gene names | Gorasp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 1 (Golgi reassembly-stacking protein of 65 kDa) (GRASP65) (Golgi peripheral membrane protein p65). | |||||
|
GR65_HUMAN
|
||||||
| NC score | 0.904480 (rank : 4) | θ value | 1.54831e-84 (rank : 3) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BQQ3, Q8N272, Q96H42 | Gene names | GORASP1, GRASP65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 1 (Golgi reassembly-stacking protein of 65 kDa) (GRASP65) (Golgi peripheral membrane protein p65) (Golgi phosphoprotein 5) (GOLPH5). | |||||
|
PSMD9_MOUSE
|
||||||
| NC score | 0.184822 (rank : 5) | θ value | 0.365318 (rank : 22) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9CR00, Q3TG66 | Gene names | Psmd9 | |||
|
Domain Architecture |

|
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 9 (26S proteasome regulatory subunit p27). | |||||
|
PSMD9_HUMAN
|
||||||
| NC score | 0.173901 (rank : 6) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O00233, Q9BQ42 | Gene names | PSMD9 | |||
|
Domain Architecture |

|
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 9 (26S proteasome regulatory subunit p27). | |||||
|
SON_MOUSE
|
||||||
| NC score | 0.122171 (rank : 7) | θ value | 0.279714 (rank : 19) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.114612 (rank : 8) | θ value | 0.00020696 (rank : 7) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
PRAX_MOUSE
|
||||||
| NC score | 0.103436 (rank : 9) | θ value | 0.47712 (rank : 25) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O55103, O55104 | Gene names | Prx | |||
|
Domain Architecture |

|
|||||
| Description | Periaxin. | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.100680 (rank : 10) | θ value | 0.0193708 (rank : 9) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.100444 (rank : 11) | θ value | 2.88788e-06 (rank : 5) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.096127 (rank : 12) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
PRAX_HUMAN
|
||||||
| NC score | 0.093031 (rank : 13) | θ value | 0.0431538 (rank : 12) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BXM0, Q9BXL9, Q9HCF2 | Gene names | PRX, KIAA1620 | |||
|
Domain Architecture |

|
|||||
| Description | Periaxin. | |||||
|
NFRKB_MOUSE
|
||||||
| NC score | 0.092231 (rank : 14) | θ value | 0.163984 (rank : 16) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.090183 (rank : 15) | θ value | 0.0252991 (rank : 10) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
GSCR1_HUMAN
|
||||||
| NC score | 0.089550 (rank : 16) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.089294 (rank : 17) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
K1543_HUMAN
|
||||||
| NC score | 0.084522 (rank : 18) | θ value | 0.47712 (rank : 23) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
WASIP_MOUSE
|
||||||
| NC score | 0.083547 (rank : 19) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
NCOA6_MOUSE
|
||||||
| NC score | 0.080484 (rank : 20) | θ value | 0.0563607 (rank : 13) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
ZN207_HUMAN
|
||||||
| NC score | 0.078702 (rank : 21) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
K1543_MOUSE
|
||||||
| NC score | 0.078113 (rank : 22) | θ value | 0.163984 (rank : 15) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80VC9, Q5DTW9, Q8BUZ0 | Gene names | Kiaa1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.074653 (rank : 23) | θ value | 0.00175202 (rank : 8) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.073804 (rank : 24) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
CN155_HUMAN
|
||||||
| NC score | 0.073660 (rank : 25) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.069584 (rank : 26) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
LEUK_HUMAN
|
||||||
| NC score | 0.069188 (rank : 27) | θ value | 0.47712 (rank : 24) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P16150 | Gene names | SPN, CD43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukosialin precursor (Leukocyte sialoglycoprotein) (Sialophorin) (Galactoglycoprotein) (GALGP) (CD43 antigen). | |||||
|
MBD6_HUMAN
|
||||||
| NC score | 0.067340 (rank : 28) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
CF015_HUMAN
|
||||||
| NC score | 0.067016 (rank : 29) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6UXA7, Q5SQ81, Q86Z05, Q9UIG3 | Gene names | C6orf15, STG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf15 precursor (Protein STG). | |||||
|
F120C_HUMAN
|
||||||
| NC score | 0.066729 (rank : 30) | θ value | 0.813845 (rank : 30) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NX05 | Gene names | FAM120C, CXorf17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120C (Tumor antigen BJ-HCC-21). | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.066686 (rank : 31) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
CIC_MOUSE
|
||||||
| NC score | 0.066359 (rank : 32) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.062695 (rank : 33) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.062690 (rank : 34) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.062306 (rank : 35) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
MUC2_HUMAN
|
||||||
| NC score | 0.062044 (rank : 36) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
NFRKB_HUMAN
|
||||||
| NC score | 0.061949 (rank : 37) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6P4R8, Q12869, Q15312, Q9H048 | Gene names | NFRKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.061358 (rank : 38) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
SF01_HUMAN
|
||||||
| NC score | 0.060658 (rank : 39) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q15637, Q14818, Q14819, Q15913, Q8IY00, Q92744, Q92745, Q969H7, Q9BW01, Q9UEI0 | Gene names | SF1, ZFM1, ZNF162 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (Zinc finger gene in MEN1 locus) (Mammalian branch point-binding protein mBBP) (BBP). | |||||
|
F120C_MOUSE
|
||||||
| NC score | 0.060631 (rank : 40) | θ value | 0.813845 (rank : 31) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C3F2 | Gene names | Fam120c, ORF34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120C. | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.059276 (rank : 41) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.057936 (rank : 42) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.057902 (rank : 43) | θ value | 0.62314 (rank : 26) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
PO121_HUMAN
|
||||||
| NC score | 0.057867 (rank : 44) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.057628 (rank : 45) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.057222 (rank : 46) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.056948 (rank : 47) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CN032_HUMAN
|
||||||
| NC score | 0.056848 (rank : 48) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
FMNL_HUMAN
|
||||||
| NC score | 0.055945 (rank : 49) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
Domain Architecture |

|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
TRI66_HUMAN
|
||||||
| NC score | 0.055935 (rank : 50) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
CO002_HUMAN
|
||||||
| NC score | 0.055847 (rank : 51) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.055480 (rank : 52) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.055338 (rank : 53) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
CN032_MOUSE
|
||||||
| NC score | 0.055069 (rank : 54) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BH93, Q3TF12, Q3TGL0, Q8CC90 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32 homolog. | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.054950 (rank : 55) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
T22D2_HUMAN
|
||||||
| NC score | 0.053812 (rank : 56) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
SF3A2_HUMAN
|
||||||
| NC score | 0.053710 (rank : 57) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
SF3A2_MOUSE
|
||||||
| NC score | 0.052956 (rank : 58) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
U689_HUMAN
|
||||||
| NC score | 0.052795 (rank : 59) | θ value | 0.62314 (rank : 29) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6UX39 | Gene names | ames=UNQ689/PRO1329 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein UNQ689/PRO1329 precursor. | |||||
|
WIRE_HUMAN
|
||||||
| NC score | 0.052698 (rank : 60) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
SFR15_HUMAN
|
||||||
| NC score | 0.052644 (rank : 61) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O95104, Q6P1M5, Q8N3I8, Q9UFM1, Q9ULP8 | Gene names | SFRS15, KIAA1172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 15 (CTD-binding SR-like protein RA4). | |||||
|
CS016_HUMAN
|
||||||
| NC score | 0.052558 (rank : 62) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8ND99 | Gene names | C19orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf16. | |||||
|
GGN_MOUSE
|
||||||
| NC score | 0.052477 (rank : 63) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
FMN2_MOUSE
|
||||||
| NC score | 0.052344 (rank : 64) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JL04 | Gene names | Fmn2 | |||
|
Domain Architecture |

|
|||||
| Description | Formin-2. | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.052242 (rank : 65) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
MUC1_MOUSE
|
||||||
| NC score | 0.052202 (rank : 66) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
NU214_HUMAN
|
||||||
| NC score | 0.052132 (rank : 67) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
WBP11_HUMAN
|
||||||
| NC score | 0.051567 (rank : 68) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y2W2, Q96AY8 | Gene names | WBP11, NPWBP, SNP70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11) (SH3 domain-binding protein SNP70) (Npw38-binding protein) (NpwBP). | |||||
|
CPSF6_MOUSE
|
||||||
| NC score | 0.051371 (rank : 69) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
DCP1A_HUMAN
|
||||||
| NC score | 0.050952 (rank : 70) | θ value | 0.365318 (rank : 21) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NPI6 | Gene names | DCP1A, SMIF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (Smad4-interacting transcriptional co-activator). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.050351 (rank : 71) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
FMN2_HUMAN
|
||||||
| NC score | 0.050049 (rank : 72) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
WIRE_MOUSE
|
||||||
| NC score | 0.050028 (rank : 73) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.049741 (rank : 74) | θ value | 1.38821 (rank : 32) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.049432 (rank : 75) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
HCFC1_MOUSE
|
||||||
| NC score | 0.048962 (rank : 76) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61191, Q684R1, Q7TSB0, Q8C2D0, Q9QWH2 | Gene names | Hcfc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N- terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C-terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
HCFC1_HUMAN
|
||||||
| NC score | 0.048476 (rank : 77) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P51610, Q6P4G5 | Gene names | HCFC1, HCF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) (VP16 accessory protein) (VCAF) (CFF) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N-terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C- terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.048283 (rank : 78) | θ value | 0.0330416 (rank : 11) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.048117 (rank : 79) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
TAU_HUMAN
|
||||||
| NC score | 0.045139 (rank : 80) | θ value | 3.0926 (rank : 49) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
DOT1L_HUMAN
|
||||||
| NC score | 0.044723 (rank : 81) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
CMTA2_HUMAN
|
||||||
| NC score | 0.044699 (rank : 82) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94983, Q7Z6M8, Q8N3V0, Q8NDG4, Q96G17 | Gene names | CAMTA2, KIAA0909 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-binding transcription activator 2. | |||||
|
ATF2_HUMAN
|
||||||
| NC score | 0.043165 (rank : 83) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P15336, Q13000 | Gene names | ATF2, CREB2, CREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (HB16). | |||||
|
ATF2_MOUSE
|
||||||
| NC score | 0.042032 (rank : 84) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P16951, Q64089, Q64090, Q64091 | Gene names | Atf2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (MXBP protein). | |||||
|
ZEP1_HUMAN
|
||||||
| NC score | 0.039637 (rank : 85) | θ value | 0.0736092 (rank : 14) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
CO039_MOUSE
|
||||||
| NC score | 0.037742 (rank : 86) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
ATX1_MOUSE
|
||||||
| NC score | 0.035210 (rank : 87) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54254 | Gene names | Atxn1, Sca1 | |||
|
Domain Architecture |

|
|||||
| Description | Ataxin-1 (Spinocerebellar ataxia type 1 protein homolog). | |||||
|
ZEP1_MOUSE
|
||||||
| NC score | 0.035098 (rank : 88) | θ value | 1.81305 (rank : 38) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1063 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q03172 | Gene names | Hivep1, Cryabp1, Znf40 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 40 (Transcription factor alphaA-CRYBP1) (Alpha A- crystallin-binding protein I) (Alpha A-CRYBP1). | |||||
|
SSBP4_HUMAN
|
||||||
| NC score | 0.032303 (rank : 89) | θ value | 1.81305 (rank : 37) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BWG4 | Gene names | SSBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Single-stranded DNA-binding protein 4. | |||||
|
PER2_MOUSE
|
||||||
| NC score | 0.031932 (rank : 90) | θ value | 1.81305 (rank : 36) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
PZRN4_HUMAN
|
||||||
| NC score | 0.031253 (rank : 91) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZMN7, Q6N052, Q8IUU1, Q9NTP7 | Gene names | PDZRN4, LNX4, SEMCAP3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing RING finger protein 4 (Ligand of Numb-protein X 4) (SEMACAP3-like protein). | |||||
|
ARID2_HUMAN
|
||||||
| NC score | 0.030210 (rank : 92) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q68CP9, Q5EB51, Q645I3, Q6ZRY5, Q7Z3I5, Q86T28, Q96SJ6, Q9HCL5 | Gene names | ARID2, KIAA1557 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 2 (ARID domain- containing protein 2) (BRG1-associated factor 200) (BAF200). | |||||
|
AMBN_HUMAN
|
||||||
| NC score | 0.030109 (rank : 93) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NP70, Q9H2X1, Q9H4L1 | Gene names | AMBN | |||
|
Domain Architecture |

|
|||||
| Description | Ameloblastin precursor. | |||||
|
TM1L1_MOUSE
|
||||||
| NC score | 0.028897 (rank : 94) | θ value | 3.0926 (rank : 50) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q923U0, Q99KE0, Q9D6Y5 | Gene names | Tom1l1, Srcasm | |||
|
Domain Architecture |

|
|||||
| Description | TOM1-like 1 protein (Target of myb-like 1 protein) (Src-activating and signaling molecule protein). | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.028261 (rank : 95) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
CP007_MOUSE
|
||||||
| NC score | 0.028246 (rank : 96) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C190, Q6PFD1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 homolog (5-day ovary-specific transcript 1 protein). | |||||
|
AMBN_MOUSE
|
||||||
| NC score | 0.028236 (rank : 97) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O55189 | Gene names | Ambn | |||
|
Domain Architecture |

|
|||||
| Description | Ameloblastin precursor. | |||||
|
DAB2_HUMAN
|
||||||
| NC score | 0.027767 (rank : 98) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P98082, Q13598, Q9BTY0, Q9UK04 | Gene names | DAB2, DOC2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (Differentially expressed protein 2) (DOC-2). | |||||
|
ZN385_HUMAN
|
||||||
| NC score | 0.027314 (rank : 99) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.026960 (rank : 100) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
KLF17_HUMAN
|
||||||
| NC score | 0.023585 (rank : 101) | θ value | 0.000121331 (rank : 6) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 882 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5JT82, Q86VQ7, Q8N805 | Gene names | KLF17, ZNF393 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 17 (Zinc finger protein 393). | |||||
|
CSPG3_MOUSE
|
||||||
| NC score | 0.023193 (rank : 102) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P55066, Q6P1E3, Q8C4F8 | Gene names | Cspg3, Ncan | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
TENS4_HUMAN
|
||||||
| NC score | 0.021145 (rank : 103) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IZW8, Q71RB7, Q8WV64, Q96JV4 | Gene names | TNS4, CTEN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-4 precursor (C-terminal tensin-like protein). | |||||
|
LAP4_MOUSE
|
||||||
| NC score | 0.020588 (rank : 104) | θ value | 1.81305 (rank : 35) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80U72, Q6P5H7, Q7TPH8, Q80VB1, Q8CI48, Q8VII1, Q922S3 | Gene names | Scrib, Kiaa0147, Lap4, Scrib1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog). | |||||
|
CUTL1_HUMAN
|
||||||
| NC score | 0.020389 (rank : 105) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
LAP4_HUMAN
|
||||||
| NC score | 0.019517 (rank : 106) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 608 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14160, Q6P496, Q7Z5D1, Q8WWV8, Q96C69, Q96GG1 | Gene names | SCRIB, CRIB1, KIAA0147, LAP4, SCRB1, VARTUL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog) (hScrib). | |||||
|
SMPX_HUMAN
|
||||||
| NC score | 0.018827 (rank : 107) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UHP9 | Gene names | SMPX, SRMX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small muscular protein (Stretch-responsive skeletal muscle protein). | |||||
|
SIGL7_HUMAN
|
||||||
| NC score | 0.018506 (rank : 108) | θ value | 1.38821 (rank : 33) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y286, Q9NZQ1, Q9UJ86, Q9UJ87, Q9Y502 | Gene names | SIGLEC7, AIRM1 | |||
|
Domain Architecture |

|
|||||
| Description | Sialic acid-binding Ig-like lectin 7 precursor (Siglec-7) (QA79 membrane protein) (Adhesion inhibitory receptor molecule 1) (AIRM-1) (p75) (D-siglec) (CDw328 antigen). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.017533 (rank : 109) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
FINC_HUMAN
|
||||||
| NC score | 0.016857 (rank : 110) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P02751, O95609, O95610, Q14312, Q14325, Q14326, Q6LDP6, Q86T27, Q8IVI8, Q96KP7, Q96KP8, Q96KP9, Q9H1B8, Q9HAP3, Q9UMK2 | Gene names | FN1, FN | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN) (Cold-insoluble globulin) (CIG). | |||||
|
CK054_HUMAN
|
||||||
| NC score | 0.016786 (rank : 111) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 3 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H0W9, Q6FI88, Q6XYB0, Q96EI3, Q96IX1, Q9Y6B4 | Gene names | C11orf54 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ester hydrolase C11orf54 (EC 3.1.-.-). | |||||
|
CYLD_HUMAN
|
||||||
| NC score | 0.016501 (rank : 112) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NQC7, O94934, Q7L3N6, Q96EH0, Q9NZX9 | Gene names | CYLD, CYLD1, KIAA0849 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase CYLD (EC 3.1.2.15) (Ubiquitin thioesterase CYLD) (Ubiquitin-specific-processing protease CYLD) (Deubiquitinating enzyme CYLD). | |||||
|
MPDZ_HUMAN
|
||||||
| NC score | 0.016256 (rank : 113) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75970, O43798, Q5CZ80, Q5JTX3, Q5JTX6, Q5JTX7, Q5JUC3, Q5JUC4, Q5VZ62, Q8N790 | Gene names | MPDZ, MUPP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple PDZ domain protein (Multi PDZ domain protein 1) (Multi-PDZ- domain protein 1). | |||||
|
YTHD3_MOUSE
|
||||||
| NC score | 0.016198 (rank : 114) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BYK6, Q3UVI5, Q6NXJ6, Q6NXJ8, Q8BKB6 | Gene names | Ythdf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YTH domain family protein 3. | |||||
|
FINC_MOUSE
|
||||||
| NC score | 0.015928 (rank : 115) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P11276, Q61567, Q61568, Q61569, Q64233, Q80UI4 | Gene names | Fn1 | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN). | |||||
|
BLNK_HUMAN
|
||||||
| NC score | 0.014745 (rank : 116) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WV28, O75498, O75499 | Gene names | BLNK, BASH, SLP65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell linker protein (Cytoplasmic adapter protein) (B-cell adapter containing SH2 domain protein) (B-cell adapter containing Src homology 2 domain protein) (Src homology 2 domain-containing leukocyte protein of 65 kDa) (SLP-65). | |||||
|
MBN3_HUMAN
|
||||||
| NC score | 0.014424 (rank : 117) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NUK0, Q5JXN8, Q5JXN9, Q5JXP4, Q6UDQ1, Q8TAD9, Q8TAF4, Q9H0Z7, Q9UF37 | Gene names | MBNL3, CHCR, MBLX39, MBXL | |||
|
Domain Architecture |

|
|||||
| Description | Muscleblind-like X-linked protein (Muscleblind-like protein 3) (Cys3His CCG1-required protein) (Protein HCHCR). | |||||
|
SIG12_HUMAN
|
||||||
| NC score | 0.014410 (rank : 118) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96PQ1, Q8IYH7 | Gene names | SIGLEC12, SIGLECL1, SLG | |||
|
Domain Architecture |

|
|||||
| Description | Sialic acid-binding Ig-like lectin 12 precursor (Siglec-12) (Sialic acid-binding Ig-like lectin-like 1) (Siglec-L1). | |||||
|
TNFC_MOUSE
|
||||||
| NC score | 0.013238 (rank : 119) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P41155 | Gene names | Ltb, Tnfc, Tnfsf3 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphotoxin-beta (LT-beta) (Tumor necrosis factor C) (TNF-C) (Tumor necrosis factor ligand superfamily member 3). | |||||
|
ZN687_HUMAN
|
||||||
| NC score | 0.012965 (rank : 120) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
SSPO_MOUSE
|
||||||
| NC score | 0.012704 (rank : 121) | θ value | 2.36792 (rank : 45) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
COCA1_HUMAN
|
||||||
| NC score | 0.012489 (rank : 122) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99715, Q99716 | Gene names | COL12A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
CBPD_MOUSE
|
||||||
| NC score | 0.008972 (rank : 123) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O89001 | Gene names | Cpd | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase D precursor (EC 3.4.17.22) (Metallocarboxypeptidase D) (gp180). | |||||
|
FOXL1_MOUSE
|
||||||
| NC score | 0.008051 (rank : 124) | θ value | 2.36792 (rank : 43) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64731 | Gene names | Foxl1, Fkh6, Fkhl11 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein L1 (Forkhead-related protein FKHL11) (Transcription factor FKH-6). | |||||
|
GLI3_MOUSE
|
||||||
| NC score | 0.006141 (rank : 125) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61602 | Gene names | Gli3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI3. | |||||
|
ZN438_HUMAN
|
||||||
| NC score | 0.005627 (rank : 126) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z4V0, Q5T426, Q658Q4, Q6ZN65 | Gene names | ZNF438 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 438. | |||||