Please be patient as the page loads
|
F120C_HUMAN
|
||||||
| SwissProt Accessions | Q9NX05 | Gene names | FAM120C, CXorf17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120C (Tumor antigen BJ-HCC-21). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
F120A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.966237 (rank : 3) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NZB2, O60649, Q14688, Q4VXF4, Q4VXF5, Q4VXG2, Q96I21, Q9NZB1 | Gene names | FAM120A, C9orf10, KIAA0183 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120A. | |||||
|
F120C_HUMAN
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9NX05 | Gene names | FAM120C, CXorf17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120C (Tumor antigen BJ-HCC-21). | |||||
|
F120C_MOUSE
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.982336 (rank : 2) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8C3F2 | Gene names | Fam120c, ORF34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120C. | |||||
|
TRAF4_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 4) | NC score | 0.066575 (rank : 8) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BUZ4, O75615, Q14848, Q2PJN8 | Gene names | TRAF4, CART1, MLN62, RNF83 | |||
|
Domain Architecture |
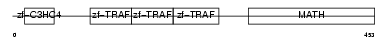
|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich domain associated with RING and Traf domains protein 1) (Malignant 62) (RING finger protein 83). | |||||
|
TRAF4_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 5) | NC score | 0.068908 (rank : 6) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61382 | Gene names | Traf4, Cart1 | |||
|
Domain Architecture |
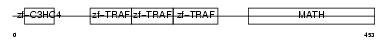
|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich motif associated to RING and Traf domains protein 1). | |||||
|
SF3A2_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 6) | NC score | 0.094189 (rank : 4) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |
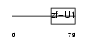
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
ZN282_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 7) | NC score | 0.006153 (rank : 82) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 796 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UDV7, O43691, Q6DKK0 | Gene names | ZNF282, HUB1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 282 (HTLV-I U5RE-binding protein 1) (HUB-1). | |||||
|
SPTN2_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 8) | NC score | 0.020862 (rank : 50) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
CIC_HUMAN
|
||||||
| θ value | 0.163984 (rank : 9) | NC score | 0.056502 (rank : 14) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
GORS2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 10) | NC score | 0.071361 (rank : 5) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99JX3, Q3U9D2, Q8CCD0, Q91W68 | Gene names | Gorasp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 2 (GRS2) (Golgi reassembly-stacking protein of 55 kDa) (GRASP55). | |||||
|
PEX5R_HUMAN
|
||||||
| θ value | 0.163984 (rank : 11) | NC score | 0.038873 (rank : 26) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IYB4, Q9NQD1, Q9P2U3, Q9P2U4 | Gene names | PEX5L, PEX5R, PXR2 | |||
|
Domain Architecture |

|
|||||
| Description | PEX5-related protein (Peroxin-5-related protein) (Pex5Rp) (PEX5-like protein) (PEX2-related protein). | |||||
|
AB1IP_MOUSE
|
||||||
| θ value | 0.365318 (rank : 12) | NC score | 0.062662 (rank : 11) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R5A3, O35329, Q8BRU0, Q99KV8 | Gene names | Apbb1ip, Prel1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1- interacting protein (APBB1-interacting protein 1) (Proline-rich EVH1 ligand 1) (PREL-1) (Proline-rich protein 48). | |||||
|
FBLI1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 13) | NC score | 0.037545 (rank : 29) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q71FD7, Q99J35 | Gene names | Fblim1, Cal | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-binding LIM protein 1 (CSX-associated LIM). | |||||
|
FGD1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 14) | NC score | 0.031054 (rank : 34) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P52734 | Gene names | Fgd1 | |||
|
Domain Architecture |

|
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 1 (Faciogenital dysplasia 1 protein homolog) (Zinc finger FYVE domain-containing protein 3) (Rho/Rac guanine nucleotide exchange factor FGD1) (Rho/Rac GEF). | |||||
|
PDE1C_MOUSE
|
||||||
| θ value | 0.365318 (rank : 15) | NC score | 0.026029 (rank : 40) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64338, Q62045, Q8BSV6 | Gene names | Pde1c | |||
|
Domain Architecture |
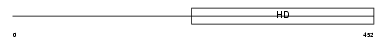
|
|||||
| Description | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C (EC 3.1.4.17) (Cam-PDE 1C). | |||||
|
ZEP2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 16) | NC score | 0.018297 (rank : 58) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3UHF7, O55140, Q3UVD4, Q3UVH5 | Gene names | Hivep2, Mibp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 homolog (Myc intron-binding protein 1) (MIBP-1). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 17) | NC score | 0.065396 (rank : 10) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
CIC_MOUSE
|
||||||
| θ value | 0.47712 (rank : 18) | NC score | 0.047942 (rank : 20) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
RXRB_MOUSE
|
||||||
| θ value | 0.47712 (rank : 19) | NC score | 0.018462 (rank : 56) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P28704, P33243 | Gene names | Rxrb, Nr2b2 | |||
|
Domain Architecture |
|
|||||
| Description | Retinoic acid receptor RXR-beta (Retinoid X receptor beta) (MHC class I regulatory element-binding protein H-2RIIBP). | |||||
|
US6NL_HUMAN
|
||||||
| θ value | 0.47712 (rank : 20) | NC score | 0.026810 (rank : 37) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92738, Q15400, Q7L0K9 | Gene names | USP6NL, KIAA0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein (Related to the N-terminus of tre) (RN- tre). | |||||
|
DPOD1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 21) | NC score | 0.043641 (rank : 22) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28340, Q8NER3, Q96H98 | Gene names | POLD1, POLD | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase delta catalytic subunit (EC 2.7.7.7) (DNA polymerase subunit delta p125). | |||||
|
GORS2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.066729 (rank : 7) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H8Y8, Q96I74, Q96K84, Q9H946, Q9UFW4 | Gene names | GORASP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 2 (GRS2) (Golgi reassembly-stacking protein of 55 kDa) (GRASP55) (p59) (Golgi phosphoprotein 6) (GOLPH6). | |||||
|
PER1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.032853 (rank : 32) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
RERE_MOUSE
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.049467 (rank : 18) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
FBRL_MOUSE
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.061547 (rank : 13) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35550, Q99L58 | Gene names | Fbl | |||
|
Domain Architecture |
|
|||||
| Description | rRNA 2'-O-methyltransferase fibrillarin (EC 2.1.1.-) (Nucleolar protein 1). | |||||
|
PRP2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.026751 (rank : 38) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
FBRL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.050825 (rank : 15) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P22087, O75259, Q9UPI6 | Gene names | FBL, FIB1, FLRN | |||
|
Domain Architecture |

|
|||||
| Description | rRNA 2'-O-methyltransferase fibrillarin (EC 2.1.1.-) (34 kDa nucleolar scleroderma antigen). | |||||
|
LNX2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.017594 (rank : 61) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N448, Q5W0P0, Q6ZMH2, Q96SH4 | Gene names | LNX2, PDZRN1 | |||
|
Domain Architecture |

|
|||||
| Description | Ligand of Numb-protein X 2 (Numb-binding protein 2) (PDZ domain- containing RING finger protein 1). | |||||
|
K22E_HUMAN
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.004824 (rank : 85) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 515 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35908 | Gene names | KRT2A, KRT2E | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type II cytoskeletal 2 epidermal (Cytokeratin-2e) (K2e) (CK 2e). | |||||
|
PININ_MOUSE
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.016998 (rank : 63) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
RSMN_HUMAN
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.049745 (rank : 16) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P63162, P14648, P17135 | Gene names | SNRPN, HCERN3, SMN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small nuclear ribonucleoprotein-associated protein N (snRNP-N) (Sm protein N) (Sm-N) (SmN) (Sm-D) (Tissue-specific-splicing protein). | |||||
|
RSMN_MOUSE
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.049745 (rank : 17) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P63163, P14648, P17135 | Gene names | Snrpn, Smn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small nuclear ribonucleoprotein-associated protein N (snRNP-N) (Sm protein N) (Sm-N) (SmN) (Sm-D) (Tissue-specific-splicing protein). | |||||
|
TCRG1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.023038 (rank : 45) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
TCRG1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.022910 (rank : 46) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
DIAP2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.030569 (rank : 36) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60879, O60878, Q8WX06, Q8WX48, Q9UJL2 | Gene names | DIAPH2, DIA | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2). | |||||
|
LATS1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.003159 (rank : 89) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O95835, Q6PKD0 | Gene names | LATS1, WARTS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase) (h-warts). | |||||
|
NOTC3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.003824 (rank : 88) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1062 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UM47, Q9UEB3, Q9UPL3, Q9Y6L8 | Gene names | NOTCH3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
PCTK3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.004023 (rank : 87) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 834 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q04899 | Gene names | Pctk3 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PCTAIRE-3 (EC 2.7.11.22) (PCTAIRE- motif protein kinase 3). | |||||
|
PER1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.025922 (rank : 41) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
RERE_HUMAN
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.042970 (rank : 23) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
WASF2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.048072 (rank : 19) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BH43 | Gene names | Wasf2, Wave2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2). | |||||
|
ZEP2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.015672 (rank : 65) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P31629, Q02646, Q5THT5, Q9NS05 | Gene names | HIVEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 (HIV- EP2) (MHC-binding protein 2) (MBP-2). | |||||
|
HORN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.019670 (rank : 54) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86YZ3 | Gene names | HRNR, S100A18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hornerin. | |||||
|
S23IP_MOUSE
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.025241 (rank : 42) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6NZC7, Q8BXY6 | Gene names | Sec23ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SEC23-interacting protein. | |||||
|
TR150_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.026420 (rank : 39) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2W1, Q5VTK6 | Gene names | THRAP3, TRAP150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
TR150_MOUSE
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.023251 (rank : 43) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
VMDL3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.014273 (rank : 68) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N1M1, Q8N356, Q8NFT9, Q9BR80 | Gene names | VMD2L3 | |||
|
Domain Architecture |

|
|||||
| Description | Bestrophin-4 (Vitelliform macular dystrophy 2-like protein 3). | |||||
|
WASF2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.045488 (rank : 21) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y6W5, O60794, Q9UDY7 | Gene names | WASF2, WAVE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2) (Verprolin homology domain-containing protein 2). | |||||
|
ZBT46_MOUSE
|
||||||
| θ value | 3.0926 (rank : 49) | NC score | 0.005928 (rank : 84) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BID6, Q3U2R2, Q9D0G6, Q9D492 | Gene names | Zbtb46, Btbd4 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 46 (BTB/POZ domain- containing protein 4). | |||||
|
ANX11_HUMAN
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.010438 (rank : 76) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P50995 | Gene names | ANXA11, ANX11 | |||
|
Domain Architecture |
|
|||||
| Description | Annexin A11 (Annexin XI) (Calcyclin-associated annexin 50) (CAP-50) (56 kDa autoantigen). | |||||
|
CCNK_MOUSE
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.020427 (rank : 51) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
HORN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.017724 (rank : 60) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
M3K12_MOUSE
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.002926 (rank : 90) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60700, P70286, Q3TLL7, Q8C4N7, Q8CBX3, Q8CDL6 | Gene names | Map3k12, Zpk | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 12 (EC 2.7.11.25) (Mixed lineage kinase) (Leucine-zipper protein kinase) (ZPK) (Dual leucine zipper bearing kinase) (DLK) (MAPK-upstream kinase) (MUK). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.018869 (rank : 55) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
NCKX1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.023175 (rank : 44) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60721, O43485, O75184 | Gene names | SLC24A1, KIAA0702, NCKX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 1 (Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod Na-Ca+K exchanger). | |||||
|
PRCC_HUMAN
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.061807 (rank : 12) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92733, O00665, O00724 | Gene names | PRCC, TPRC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein PRCC (Papillary renal cell carcinoma translocation-associated gene protein). | |||||
|
SYNJ1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.012532 (rank : 72) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O43426, O43425, O94984 | Gene names | SYNJ1, KIAA0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
AB1IP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.038721 (rank : 27) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z5R6, Q8IWS8, Q8IZZ7 | Gene names | APBB1IP, PREL1, RARP1, RIAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1- interacting protein (APBB1-interacting protein 1) (Rap1-GTP- interacting adapter molecule) (RIAM) (Proline-rich EVH1 ligand 1) (PREL-1) (Proline-rich protein 73) (Retinoic acid-responsive proline- rich protein 1) (RARP-1). | |||||
|
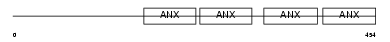
ANXA7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.012149 (rank : 74) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P20073 | Gene names | ANXA7, ANX7, SNX | |||
|
Domain Architecture |
|
|||||
| Description | Annexin A7 (Annexin VII) (Synexin). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.034720 (rank : 30) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.037571 (rank : 28) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CAC1G_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.014831 (rank : 66) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
CCDC9_MOUSE
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.016645 (rank : 64) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VC31 | Gene names | Ccdc9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 9. | |||||
|
CU063_HUMAN
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.017839 (rank : 59) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58658, Q8IXZ0 | Gene names | C21orf63 | |||
|
Domain Architecture |

|
|||||
| Description | Protein C21orf63 precursor (SUE21). | |||||
|
DLG5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.012161 (rank : 73) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TDM6, Q5T1H7, Q5T1H8, Q6DKG3, Q86WC0, Q8TDM7, Q9UE73, Q9Y4E3 | Gene names | DLG5, KIAA0583, PDLG | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 5 (Placenta and prostate DLG) (Discs large protein P-dlg). | |||||
|
FMN2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.017454 (rank : 62) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
RAI2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.041510 (rank : 24) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y5P3, Q8N6X7 | Gene names | RAI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2. | |||||
|
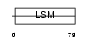
RSMB_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.032141 (rank : 33) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P14678, Q15490, Q9UIS5 | Gene names | SNRPB, COD, SNRPB1 | |||
|
Domain Architecture |
|
|||||
| Description | Small nuclear ribonucleoprotein-associated proteins B and B' (snRNP-B) (Sm protein B/B') (Sm-B/Sm-B') (SmB/SmB'). | |||||
|
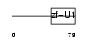
SF3A2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.066348 (rank : 9) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.018379 (rank : 57) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
ABP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.022784 (rank : 47) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8JZQ5, Q8R229, Q8VC36 | Gene names | Abp1 | |||
|
Domain Architecture |

|
|||||
| Description | Amiloride-sensitive amine oxidase [copper-containing] precursor (EC 1.4.3.6) (Diamine oxidase) (DAO) (Amiloride-binding protein) (ABP) (Histaminase). | |||||
|
DHX57_MOUSE
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.007212 (rank : 81) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P5D3, Q3TS93, Q6NZK4, Q6P1B4, Q8BI63, Q8BIA2, Q8R360 | Gene names | Dhx57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
FUS_MOUSE
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.010174 (rank : 77) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56959 | Gene names | Fus | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein FUS (Pigpen protein). | |||||
|
LYRIC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.014110 (rank : 69) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80WJ7, Q3U9F8, Q3UAQ8, Q8BN67, Q8CBT9, Q8CDL0, Q8CGI7, Q9D052 | Gene names | Mtdh, Lyric | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/LYRIC) (Metastasis adhesion protein) (Metadherin). | |||||
|
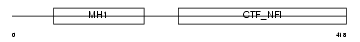
NFIX_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.012855 (rank : 71) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14938, O60413, Q12859, Q13050, Q13052, Q9UPH1, Q9Y6R8 | Gene names | NFIX | |||
|
Domain Architecture |
|
|||||
| Description | Nuclear factor 1 X-type (Nuclear factor 1/X) (NF1-X) (NFI-X) (NF-I/X) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||
|
NFIX_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.012885 (rank : 70) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70257, O08519, P70258, Q64192, Q99L78, Q9R1G2 | Gene names | Nfix | |||
|
Domain Architecture |
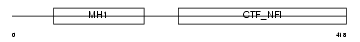
|
|||||
| Description | Nuclear factor 1 X-type (Nuclear factor 1/X) (NF1-X) (NFI-X) (NF-I/X) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||
|
PAK7_MOUSE
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.000666 (rank : 91) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C015, Q3TQJ7, Q6RWS7 | Gene names | Pak7, Pak5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase PAK 7 (EC 2.7.11.1) (p21-activated kinase 7) (PAK-7) (p21-activated kinase 5). | |||||
|
RBM22_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.020196 (rank : 52) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NW64, O95607 | Gene names | RBM22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor RBM22 (RNA-binding motif protein 22). | |||||
|
RBM22_MOUSE
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.020196 (rank : 53) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BHS3, Q3TJB0, Q9CXA0 | Gene names | Rbm22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor RBM22 (RNA-binding motif protein 22). | |||||
|
RXRB_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.011027 (rank : 75) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P28702, P28703 | Gene names | RXRB, NR2B2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoic acid receptor RXR-beta (Retinoid X receptor beta). | |||||
|
WASF3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.033401 (rank : 31) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8VHI6 | Gene names | Wasf3, Wave3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3). | |||||
|
AP2C_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.008184 (rank : 80) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92754, O00685, O00730, Q86V30, Q8IVB6, Q9P1X2 | Gene names | TFAP2C | |||
|
Domain Architecture |
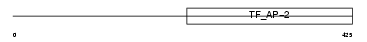
|
|||||
| Description | Transcription factor AP-2 gamma (AP2-gamma) (Activating enhancer- binding protein 2 gamma) (Transcription factor ERF-1). | |||||
|
DIAP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.014741 (rank : 67) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 839 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O08808 | Gene names | Diaph1, Diap1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1) (mDIA1) (p140mDIA). | |||||
|
HNRPR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.005968 (rank : 83) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43390 | Gene names | HNRPR | |||
|
Domain Architecture |
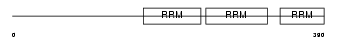
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein R (hnRNP R). | |||||
|
NEURL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.009913 (rank : 78) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q923S6, Q91ZT6, Q9CWF1, Q9QWF1 | Gene names | Neurl, Neurl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuralized-like protein 1 (m-neuralized 1) (m-neu1). | |||||
|
PHF3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.021768 (rank : 49) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
RBM12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.009282 (rank : 79) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NTZ6, O94865, Q8N3B1, Q9H196 | Gene names | RBM12, KIAA0765 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
RBM16_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.022129 (rank : 48) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UPN6, Q5TBU6, Q9BQN8, Q9BX43 | Gene names | RBM16, KIAA1116 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein 16 (RNA-binding motif protein 16). | |||||
|
RTP3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.030582 (rank : 35) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5QGU6, Q8VDC2 | Gene names | Rtp3, Tmem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-transporting protein 3 (Transmembrane protein 7). | |||||
|
WASF4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.039038 (rank : 25) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8IV90 | Gene names | WASF4, SCAR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 4 (WASP-family protein member 4). | |||||
|
ZBT46_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.004572 (rank : 86) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86UZ6, Q5JWJ3, Q9BQK3, Q9H3Z8, Q9H3Z9 | Gene names | ZBTB46, BTBD4, ZNF340 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 46 (BTB/POZ domain- containing protein 4) (Zinc finger protein 340). | |||||
|
F120C_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9NX05 | Gene names | FAM120C, CXorf17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120C (Tumor antigen BJ-HCC-21). | |||||
|
F120C_MOUSE
|
||||||
| NC score | 0.982336 (rank : 2) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8C3F2 | Gene names | Fam120c, ORF34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120C. | |||||
|
F120A_HUMAN
|
||||||
| NC score | 0.966237 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NZB2, O60649, Q14688, Q4VXF4, Q4VXF5, Q4VXG2, Q96I21, Q9NZB1 | Gene names | FAM120A, C9orf10, KIAA0183 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120A. | |||||
|
SF3A2_MOUSE
|
||||||
| NC score | 0.094189 (rank : 4) | θ value | 0.0330416 (rank : 6) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
GORS2_MOUSE
|
||||||
| NC score | 0.071361 (rank : 5) | θ value | 0.163984 (rank : 10) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99JX3, Q3U9D2, Q8CCD0, Q91W68 | Gene names | Gorasp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 2 (GRS2) (Golgi reassembly-stacking protein of 55 kDa) (GRASP55). | |||||
|
TRAF4_MOUSE
|
||||||
| NC score | 0.068908 (rank : 6) | θ value | 0.00390308 (rank : 5) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61382 | Gene names | Traf4, Cart1 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich motif associated to RING and Traf domains protein 1). | |||||
|
GORS2_HUMAN
|
||||||
| NC score | 0.066729 (rank : 7) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H8Y8, Q96I74, Q96K84, Q9H946, Q9UFW4 | Gene names | GORASP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 2 (GRS2) (Golgi reassembly-stacking protein of 55 kDa) (GRASP55) (p59) (Golgi phosphoprotein 6) (GOLPH6). | |||||
|
TRAF4_HUMAN
|
||||||
| NC score | 0.066575 (rank : 8) | θ value | 0.00390308 (rank : 4) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BUZ4, O75615, Q14848, Q2PJN8 | Gene names | TRAF4, CART1, MLN62, RNF83 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich domain associated with RING and Traf domains protein 1) (Malignant 62) (RING finger protein 83). | |||||
|
SF3A2_HUMAN
|
||||||
| NC score | 0.066348 (rank : 9) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.065396 (rank : 10) | θ value | 0.47712 (rank : 17) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
AB1IP_MOUSE
|
||||||
| NC score | 0.062662 (rank : 11) | θ value | 0.365318 (rank : 12) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R5A3, O35329, Q8BRU0, Q99KV8 | Gene names | Apbb1ip, Prel1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1- interacting protein (APBB1-interacting protein 1) (Proline-rich EVH1 ligand 1) (PREL-1) (Proline-rich protein 48). | |||||
|
PRCC_HUMAN
|
||||||
| NC score | 0.061807 (rank : 12) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92733, O00665, O00724 | Gene names | PRCC, TPRC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein PRCC (Papillary renal cell carcinoma translocation-associated gene protein). | |||||
|
FBRL_MOUSE
|
||||||
| NC score | 0.061547 (rank : 13) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35550, Q99L58 | Gene names | Fbl | |||
|
Domain Architecture |

|
|||||
| Description | rRNA 2'-O-methyltransferase fibrillarin (EC 2.1.1.-) (Nucleolar protein 1). | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.056502 (rank : 14) | θ value | 0.163984 (rank : 9) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
FBRL_HUMAN
|
||||||
| NC score | 0.050825 (rank : 15) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P22087, O75259, Q9UPI6 | Gene names | FBL, FIB1, FLRN | |||
|
Domain Architecture |

|
|||||
| Description | rRNA 2'-O-methyltransferase fibrillarin (EC 2.1.1.-) (34 kDa nucleolar scleroderma antigen). | |||||
|
RSMN_HUMAN
|
||||||
| NC score | 0.049745 (rank : 16) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P63162, P14648, P17135 | Gene names | SNRPN, HCERN3, SMN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small nuclear ribonucleoprotein-associated protein N (snRNP-N) (Sm protein N) (Sm-N) (SmN) (Sm-D) (Tissue-specific-splicing protein). | |||||
|
RSMN_MOUSE
|
||||||
| NC score | 0.049745 (rank : 17) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P63163, P14648, P17135 | Gene names | Snrpn, Smn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small nuclear ribonucleoprotein-associated protein N (snRNP-N) (Sm protein N) (Sm-N) (SmN) (Sm-D) (Tissue-specific-splicing protein). | |||||
|
RERE_MOUSE
|
||||||
| NC score | 0.049467 (rank : 18) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
WASF2_MOUSE
|
||||||
| NC score | 0.048072 (rank : 19) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BH43 | Gene names | Wasf2, Wave2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2). | |||||
|
CIC_MOUSE
|
||||||
| NC score | 0.047942 (rank : 20) | θ value | 0.47712 (rank : 18) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
WASF2_HUMAN
|
||||||
| NC score | 0.045488 (rank : 21) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y6W5, O60794, Q9UDY7 | Gene names | WASF2, WAVE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2) (Verprolin homology domain-containing protein 2). | |||||
|
DPOD1_HUMAN
|
||||||
| NC score | 0.043641 (rank : 22) | θ value | 0.62314 (rank : 21) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28340, Q8NER3, Q96H98 | Gene names | POLD1, POLD | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase delta catalytic subunit (EC 2.7.7.7) (DNA polymerase subunit delta p125). | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.042970 (rank : 23) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
RAI2_HUMAN
|
||||||
| NC score | 0.041510 (rank : 24) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y5P3, Q8N6X7 | Gene names | RAI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2. | |||||
|
WASF4_HUMAN
|
||||||
| NC score | 0.039038 (rank : 25) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8IV90 | Gene names | WASF4, SCAR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 4 (WASP-family protein member 4). | |||||
|
PEX5R_HUMAN
|
||||||
| NC score | 0.038873 (rank : 26) | θ value | 0.163984 (rank : 11) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IYB4, Q9NQD1, Q9P2U3, Q9P2U4 | Gene names | PEX5L, PEX5R, PXR2 | |||
|
Domain Architecture |

|
|||||
| Description | PEX5-related protein (Peroxin-5-related protein) (Pex5Rp) (PEX5-like protein) (PEX2-related protein). | |||||
|
AB1IP_HUMAN
|
||||||
| NC score | 0.038721 (rank : 27) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z5R6, Q8IWS8, Q8IZZ7 | Gene names | APBB1IP, PREL1, RARP1, RIAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1- interacting protein (APBB1-interacting protein 1) (Rap1-GTP- interacting adapter molecule) (RIAM) (Proline-rich EVH1 ligand 1) (PREL-1) (Proline-rich protein 73) (Retinoic acid-responsive proline- rich protein 1) (RARP-1). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.037571 (rank : 28) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
FBLI1_MOUSE
|
||||||
| NC score | 0.037545 (rank : 29) | θ value | 0.365318 (rank : 13) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q71FD7, Q99J35 | Gene names | Fblim1, Cal | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-binding LIM protein 1 (CSX-associated LIM). | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.034720 (rank : 30) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
WASF3_MOUSE
|
||||||
| NC score | 0.033401 (rank : 31) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8VHI6 | Gene names | Wasf3, Wave3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3). | |||||
|
PER1_MOUSE
|
||||||
| NC score | 0.032853 (rank : 32) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
RSMB_HUMAN
|
||||||
| NC score | 0.032141 (rank : 33) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P14678, Q15490, Q9UIS5 | Gene names | SNRPB, COD, SNRPB1 | |||
|
Domain Architecture |

|
|||||
| Description | Small nuclear ribonucleoprotein-associated proteins B and B' (snRNP-B) (Sm protein B/B') (Sm-B/Sm-B') (SmB/SmB'). | |||||
|
FGD1_MOUSE
|
||||||
| NC score | 0.031054 (rank : 34) | θ value | 0.365318 (rank : 14) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P52734 | Gene names | Fgd1 | |||
|
Domain Architecture |

|
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 1 (Faciogenital dysplasia 1 protein homolog) (Zinc finger FYVE domain-containing protein 3) (Rho/Rac guanine nucleotide exchange factor FGD1) (Rho/Rac GEF). | |||||
|
RTP3_MOUSE
|
||||||
| NC score | 0.030582 (rank : 35) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5QGU6, Q8VDC2 | Gene names | Rtp3, Tmem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-transporting protein 3 (Transmembrane protein 7). | |||||
|
DIAP2_HUMAN
|
||||||
| NC score | 0.030569 (rank : 36) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60879, O60878, Q8WX06, Q8WX48, Q9UJL2 | Gene names | DIAPH2, DIA | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2). | |||||
|
US6NL_HUMAN
|
||||||
| NC score | 0.026810 (rank : 37) | θ value | 0.47712 (rank : 20) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92738, Q15400, Q7L0K9 | Gene names | USP6NL, KIAA0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein (Related to the N-terminus of tre) (RN- tre). | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.026751 (rank : 38) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
TR150_HUMAN
|
||||||
| NC score | 0.026420 (rank : 39) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2W1, Q5VTK6 | Gene names | THRAP3, TRAP150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
PDE1C_MOUSE
|
||||||
| NC score | 0.026029 (rank : 40) | θ value | 0.365318 (rank : 15) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64338, Q62045, Q8BSV6 | Gene names | Pde1c | |||
|
Domain Architecture |

|
|||||
| Description | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C (EC 3.1.4.17) (Cam-PDE 1C). | |||||
|
PER1_HUMAN
|
||||||
| NC score | 0.025922 (rank : 41) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
S23IP_MOUSE
|
||||||
| NC score | 0.025241 (rank : 42) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6NZC7, Q8BXY6 | Gene names | Sec23ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SEC23-interacting protein. | |||||
|
TR150_MOUSE
|
||||||
| NC score | 0.023251 (rank : 43) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
NCKX1_HUMAN
|
||||||
| NC score | 0.023175 (rank : 44) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60721, O43485, O75184 | Gene names | SLC24A1, KIAA0702, NCKX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 1 (Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod Na-Ca+K exchanger). | |||||
|
TCRG1_HUMAN
|
||||||
| NC score | 0.023038 (rank : 45) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
TCRG1_MOUSE
|
||||||
| NC score | 0.022910 (rank : 46) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
ABP1_MOUSE
|
||||||
| NC score | 0.022784 (rank : 47) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8JZQ5, Q8R229, Q8VC36 | Gene names | Abp1 | |||
|
Domain Architecture |

|
|||||
| Description | Amiloride-sensitive amine oxidase [copper-containing] precursor (EC 1.4.3.6) (Diamine oxidase) (DAO) (Amiloride-binding protein) (ABP) (Histaminase). | |||||
|
RBM16_HUMAN
|
||||||
| NC score | 0.022129 (rank : 48) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UPN6, Q5TBU6, Q9BQN8, Q9BX43 | Gene names | RBM16, KIAA1116 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein 16 (RNA-binding motif protein 16). | |||||
|
PHF3_HUMAN
|
||||||
| NC score | 0.021768 (rank : 49) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
SPTN2_HUMAN
|
||||||
| NC score | 0.020862 (rank : 50) | θ value | 0.0736092 (rank : 8) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
CCNK_MOUSE
|
||||||
| NC score | 0.020427 (rank : 51) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
RBM22_HUMAN
|
||||||
| NC score | 0.020196 (rank : 52) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NW64, O95607 | Gene names | RBM22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor RBM22 (RNA-binding motif protein 22). | |||||
|
RBM22_MOUSE
|
||||||
| NC score | 0.020196 (rank : 53) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BHS3, Q3TJB0, Q9CXA0 | Gene names | Rbm22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor RBM22 (RNA-binding motif protein 22). | |||||
|
HORN_HUMAN
|
||||||
| NC score | 0.019670 (rank : 54) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86YZ3 | Gene names | HRNR, S100A18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hornerin. | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.018869 (rank : 55) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
RXRB_MOUSE
|
||||||
| NC score | 0.018462 (rank : 56) | θ value | 0.47712 (rank : 19) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P28704, P33243 | Gene names | Rxrb, Nr2b2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoic acid receptor RXR-beta (Retinoid X receptor beta) (MHC class I regulatory element-binding protein H-2RIIBP). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.018379 (rank : 57) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
ZEP2_MOUSE
|
||||||
| NC score | 0.018297 (rank : 58) | θ value | 0.365318 (rank : 16) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3UHF7, O55140, Q3UVD4, Q3UVH5 | Gene names | Hivep2, Mibp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 homolog (Myc intron-binding protein 1) (MIBP-1). | |||||
|
CU063_HUMAN
|
||||||
| NC score | 0.017839 (rank : 59) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58658, Q8IXZ0 | Gene names | C21orf63 | |||
|
Domain Architecture |

|
|||||
| Description | Protein C21orf63 precursor (SUE21). | |||||
|
HORN_MOUSE
|
||||||
| NC score | 0.017724 (rank : 60) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
LNX2_HUMAN
|
||||||
| NC score | 0.017594 (rank : 61) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N448, Q5W0P0, Q6ZMH2, Q96SH4 | Gene names | LNX2, PDZRN1 | |||
|
Domain Architecture |

|
|||||
| Description | Ligand of Numb-protein X 2 (Numb-binding protein 2) (PDZ domain- containing RING finger protein 1). | |||||
|
FMN2_HUMAN
|
||||||
| NC score | 0.017454 (rank : 62) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
PININ_MOUSE
|
||||||
| NC score | 0.016998 (rank : 63) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
CCDC9_MOUSE
|
||||||
| NC score | 0.016645 (rank : 64) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VC31 | Gene names | Ccdc9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 9. | |||||
|
ZEP2_HUMAN
|
||||||
| NC score | 0.015672 (rank : 65) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P31629, Q02646, Q5THT5, Q9NS05 | Gene names | HIVEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 (HIV- EP2) (MHC-binding protein 2) (MBP-2). | |||||
|
CAC1G_HUMAN
|
||||||
| NC score | 0.014831 (rank : 66) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
DIAP1_MOUSE
|
||||||
| NC score | 0.014741 (rank : 67) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 839 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O08808 | Gene names | Diaph1, Diap1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1) (mDIA1) (p140mDIA). | |||||
|
VMDL3_HUMAN
|
||||||
| NC score | 0.014273 (rank : 68) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N1M1, Q8N356, Q8NFT9, Q9BR80 | Gene names | VMD2L3 | |||
|
Domain Architecture |

|
|||||
| Description | Bestrophin-4 (Vitelliform macular dystrophy 2-like protein 3). | |||||
|
LYRIC_MOUSE
|
||||||
| NC score | 0.014110 (rank : 69) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80WJ7, Q3U9F8, Q3UAQ8, Q8BN67, Q8CBT9, Q8CDL0, Q8CGI7, Q9D052 | Gene names | Mtdh, Lyric | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/LYRIC) (Metastasis adhesion protein) (Metadherin). | |||||
|
NFIX_MOUSE
|
||||||
| NC score | 0.012885 (rank : 70) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70257, O08519, P70258, Q64192, Q99L78, Q9R1G2 | Gene names | Nfix | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor 1 X-type (Nuclear factor 1/X) (NF1-X) (NFI-X) (NF-I/X) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||
|
NFIX_HUMAN
|
||||||
| NC score | 0.012855 (rank : 71) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14938, O60413, Q12859, Q13050, Q13052, Q9UPH1, Q9Y6R8 | Gene names | NFIX | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor 1 X-type (Nuclear factor 1/X) (NF1-X) (NFI-X) (NF-I/X) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||
|
SYNJ1_HUMAN
|
||||||
| NC score | 0.012532 (rank : 72) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O43426, O43425, O94984 | Gene names | SYNJ1, KIAA0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
DLG5_HUMAN
|
||||||
| NC score | 0.012161 (rank : 73) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TDM6, Q5T1H7, Q5T1H8, Q6DKG3, Q86WC0, Q8TDM7, Q9UE73, Q9Y4E3 | Gene names | DLG5, KIAA0583, PDLG | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 5 (Placenta and prostate DLG) (Discs large protein P-dlg). | |||||
|
ANXA7_HUMAN
|
||||||
| NC score | 0.012149 (rank : 74) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P20073 | Gene names | ANXA7, ANX7, SNX | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A7 (Annexin VII) (Synexin). | |||||
|
RXRB_HUMAN
|
||||||
| NC score | 0.011027 (rank : 75) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P28702, P28703 | Gene names | RXRB, NR2B2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoic acid receptor RXR-beta (Retinoid X receptor beta). | |||||
|
ANX11_HUMAN
|
||||||
| NC score | 0.010438 (rank : 76) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P50995 | Gene names | ANXA11, ANX11 | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A11 (Annexin XI) (Calcyclin-associated annexin 50) (CAP-50) (56 kDa autoantigen). | |||||
|
FUS_MOUSE
|
||||||
| NC score | 0.010174 (rank : 77) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56959 | Gene names | Fus | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein FUS (Pigpen protein). | |||||
|
NEURL_MOUSE
|
||||||
| NC score | 0.009913 (rank : 78) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q923S6, Q91ZT6, Q9CWF1, Q9QWF1 | Gene names | Neurl, Neurl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuralized-like protein 1 (m-neuralized 1) (m-neu1). | |||||
|
RBM12_HUMAN
|
||||||
| NC score | 0.009282 (rank : 79) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NTZ6, O94865, Q8N3B1, Q9H196 | Gene names | RBM12, KIAA0765 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
AP2C_HUMAN
|
||||||
| NC score | 0.008184 (rank : 80) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92754, O00685, O00730, Q86V30, Q8IVB6, Q9P1X2 | Gene names | TFAP2C | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-2 gamma (AP2-gamma) (Activating enhancer- binding protein 2 gamma) (Transcription factor ERF-1). | |||||
|
DHX57_MOUSE
|
||||||
| NC score | 0.007212 (rank : 81) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P5D3, Q3TS93, Q6NZK4, Q6P1B4, Q8BI63, Q8BIA2, Q8R360 | Gene names | Dhx57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
ZN282_HUMAN
|
||||||
| NC score | 0.006153 (rank : 82) | θ value | 0.0563607 (rank : 7) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 796 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UDV7, O43691, Q6DKK0 | Gene names | ZNF282, HUB1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 282 (HTLV-I U5RE-binding protein 1) (HUB-1). | |||||
|
HNRPR_HUMAN
|
||||||
| NC score | 0.005968 (rank : 83) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43390 | Gene names | HNRPR | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein R (hnRNP R). | |||||
|
ZBT46_MOUSE
|
||||||
| NC score | 0.005928 (rank : 84) | θ value | 3.0926 (rank : 49) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BID6, Q3U2R2, Q9D0G6, Q9D492 | Gene names | Zbtb46, Btbd4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 46 (BTB/POZ domain- containing protein 4). | |||||
|
K22E_HUMAN
|
||||||
| NC score | 0.004824 (rank : 85) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 515 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35908 | Gene names | KRT2A, KRT2E | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type II cytoskeletal 2 epidermal (Cytokeratin-2e) (K2e) (CK 2e). | |||||
|
ZBT46_HUMAN
|
||||||
| NC score | 0.004572 (rank : 86) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86UZ6, Q5JWJ3, Q9BQK3, Q9H3Z8, Q9H3Z9 | Gene names | ZBTB46, BTBD4, ZNF340 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 46 (BTB/POZ domain- containing protein 4) (Zinc finger protein 340). | |||||
|
PCTK3_MOUSE
|
||||||
| NC score | 0.004023 (rank : 87) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 834 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q04899 | Gene names | Pctk3 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PCTAIRE-3 (EC 2.7.11.22) (PCTAIRE- motif protein kinase 3). | |||||
|
NOTC3_HUMAN
|
||||||
| NC score | 0.003824 (rank : 88) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1062 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UM47, Q9UEB3, Q9UPL3, Q9Y6L8 | Gene names | NOTCH3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
LATS1_HUMAN
|
||||||
| NC score | 0.003159 (rank : 89) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O95835, Q6PKD0 | Gene names | LATS1, WARTS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase) (h-warts). | |||||
|
M3K12_MOUSE
|
||||||
| NC score | 0.002926 (rank : 90) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60700, P70286, Q3TLL7, Q8C4N7, Q8CBX3, Q8CDL6 | Gene names | Map3k12, Zpk | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 12 (EC 2.7.11.25) (Mixed lineage kinase) (Leucine-zipper protein kinase) (ZPK) (Dual leucine zipper bearing kinase) (DLK) (MAPK-upstream kinase) (MUK). | |||||
|
PAK7_MOUSE
|
||||||
| NC score | 0.000666 (rank : 91) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C015, Q3TQJ7, Q6RWS7 | Gene names | Pak7, Pak5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase PAK 7 (EC 2.7.11.1) (p21-activated kinase 7) (PAK-7) (p21-activated kinase 5). | |||||