Please be patient as the page loads
|
PK2L2_HUMAN
|
||||||
| SwissProt Accessions | Q9NZM6, Q9UNJ0 | Gene names | PKD2L2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycystic kidney disease 2-like 2 protein (Polycystin-L2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PK2L2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q9NZM6, Q9UNJ0 | Gene names | PKD2L2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycystic kidney disease 2-like 2 protein (Polycystin-L2). | |||||
|
PK2L2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.989577 (rank : 2) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9JLG4 | Gene names | Pkd2l2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 2-like 2 protein (Polycystin-L2). | |||||
|
PK2L1_HUMAN
|
||||||
| θ value | 1.42583e-138 (rank : 3) | NC score | 0.961548 (rank : 4) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9P0L9, O75972, Q9UP35, Q9UPA2 | Gene names | PKD2L1, PKD2L, PKDL | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 2-like 1 protein (Polycystin-L) (Polycystin- 2 homolog). | |||||
|
PKD2_HUMAN
|
||||||
| θ value | 5.99046e-137 (rank : 4) | NC score | 0.962148 (rank : 3) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q13563, O60441, Q15764 | Gene names | PKD2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-2 (Polycystic kidney disease 2 protein homolog) (Autosomal dominant polycystic kidney disease type II protein) (Polycystwin) (R48321). | |||||
|
PKD2_MOUSE
|
||||||
| θ value | 2.78914e-126 (rank : 5) | NC score | 0.958962 (rank : 5) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | O35245, Q9QWP0, Q9Z193, Q9Z194 | Gene names | Pkd2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-2 (Polycystic kidney disease 2 protein homolog). | |||||
|
PKDRE_HUMAN
|
||||||
| θ value | 2.5182e-18 (rank : 6) | NC score | 0.587415 (rank : 6) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9NTG1, O95850 | Gene names | PKDREJ | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease and receptor for egg jelly-related protein precursor (PKD and REJ homolog). | |||||
|
PKDRE_MOUSE
|
||||||
| θ value | 5.60996e-18 (rank : 7) | NC score | 0.559714 (rank : 7) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Z0T6 | Gene names | Pkdrej | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease and receptor for egg jelly-related protein precursor (PKD and REJ homolog). | |||||
|
PKD1_HUMAN
|
||||||
| θ value | 1.58096e-12 (rank : 8) | NC score | 0.299181 (rank : 40) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P98161, Q15140, Q15141 | Gene names | PKD1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-1 precursor (Autosomal dominant polycystic kidney disease protein 1). | |||||
|
PK1L1_HUMAN
|
||||||
| θ value | 4.59992e-12 (rank : 9) | NC score | 0.475296 (rank : 8) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TDX9, Q6UWK1 | Gene names | PKD1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 1-like 1 protein (Polycystin-1L1). | |||||
|
CAC1F_MOUSE
|
||||||
| θ value | 1.25267e-09 (rank : 10) | NC score | 0.395944 (rank : 11) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JIS7 | Gene names | Cacna1f | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
MCLN2_MOUSE
|
||||||
| θ value | 1.25267e-09 (rank : 11) | NC score | 0.408413 (rank : 9) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8K595, Q3UCG4, Q8K2T6, Q9CQD3 | Gene names | Mcoln2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-2. | |||||
|
CAC1F_HUMAN
|
||||||
| θ value | 3.64472e-09 (rank : 12) | NC score | 0.392289 (rank : 14) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O60840, O43901 | Gene names | CACNA1F, CACNAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
MCLN3_HUMAN
|
||||||
| θ value | 3.64472e-09 (rank : 13) | NC score | 0.395519 (rank : 12) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TDD5, Q5T4H5, Q5T4H6, Q9NV09 | Gene names | MCOLN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-3. | |||||
|
MCLN3_MOUSE
|
||||||
| θ value | 4.76016e-09 (rank : 14) | NC score | 0.393347 (rank : 13) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8R4F0, Q8BS73, Q8BSG1, Q8CDB2 | Gene names | Mcoln3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-3. | |||||
|
CAC1C_MOUSE
|
||||||
| θ value | 1.38499e-08 (rank : 15) | NC score | 0.390830 (rank : 15) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q01815, Q04476, Q61242, Q99242 | Gene names | Cacna1c, Cach2, Cacn2, Cacnl1a1, Cchl1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle) (Mouse brain class C) (MBC) (MELC-CC). | |||||
|
CAC1D_HUMAN
|
||||||
| θ value | 1.38499e-08 (rank : 16) | NC score | 0.384016 (rank : 17) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q01668, Q13916, Q13931 | Gene names | CACNA1D, CACH3, CACN4, CACNL1A2, CCHL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1D (Voltage- gated calcium channel subunit alpha Cav1.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 2). | |||||
|
CAC1C_HUMAN
|
||||||
| θ value | 2.36244e-08 (rank : 17) | NC score | 0.389723 (rank : 16) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
MCLN2_HUMAN
|
||||||
| θ value | 3.08544e-08 (rank : 18) | NC score | 0.398726 (rank : 10) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8IZK6, Q8N9R3 | Gene names | MCOLN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-2. | |||||
|
CAC1E_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 19) | NC score | 0.382014 (rank : 18) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q61290 | Gene names | Cacna1e, Cach6, Cacnl1a6, Cchra1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent R-type calcium channel subunit alpha-1E (Voltage- gated calcium channel subunit alpha Cav2.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 6) (Brain calcium channel II) (BII). | |||||
|
SC11A_HUMAN
|
||||||
| θ value | 2.21117e-06 (rank : 20) | NC score | 0.339744 (rank : 28) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UI33, Q68K15, Q8NDX3, Q9UHE0, Q9UHM0 | Gene names | SCN11A, PN5, SCN12A, SNS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 11 subunit alpha (Sodium channel protein type XI subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.9) (Sensory neuron sodium channel 2) (Peripheral nerve sodium channel 5) (hNaN). | |||||
|
CAC1E_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 21) | NC score | 0.381345 (rank : 19) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q15878, Q14580, Q14581 | Gene names | CACNA1E, CACH6, CACNL1A6 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent R-type calcium channel subunit alpha-1E (Voltage- gated calcium channel subunit alpha Cav2.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 6) (Brain calcium channel II) (BII). | |||||
|
CAC1B_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 22) | NC score | 0.378169 (rank : 20) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q00975 | Gene names | CACNA1B, CACH5, CACNL1A5 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
TRPA1_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 23) | NC score | 0.122411 (rank : 61) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75762 | Gene names | TRPA1, ANKTM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily A member 1 (Ankyrin-like with transmembrane domains protein 1) (Transformation sensitive-protein p120). | |||||
|
CAC1S_HUMAN
|
||||||
| θ value | 1.87187e-05 (rank : 24) | NC score | 0.374344 (rank : 21) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q13698, Q12896, Q13934 | Gene names | CACNA1S, CACH1, CACN1, CACNL1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1S (Voltage- gated calcium channel subunit alpha Cav1.1) (Calcium channel, L type, alpha-1 polypeptide, isoform 3, skeletal muscle). | |||||
|
CAC1S_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 25) | NC score | 0.371891 (rank : 22) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q02789, Q99240 | Gene names | Cacna1s, Cach1, Cach1b, Cacn1, Cacnl1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1S (Voltage- gated calcium channel subunit alpha Cav1.1) (Calcium channel, L type, alpha-1 polypeptide, isoform 3, skeletal muscle). | |||||
|
MCLN1_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 26) | NC score | 0.357168 (rank : 24) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9GZU1, Q7Z4F7, Q9H292, Q9H4B3, Q9H4B5 | Gene names | MCOLN1, ML4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-1 (Mucolipidin) (MG-2). | |||||
|
MCLN1_MOUSE
|
||||||
| θ value | 3.19293e-05 (rank : 27) | NC score | 0.352552 (rank : 26) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99J21 | Gene names | Mcoln1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-1 (Mucolipidin). | |||||
|
CAC1B_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 28) | NC score | 0.368093 (rank : 23) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O55017, Q60609 | Gene names | Cacna1b, Cach5, Cacnl1a5, Cchn1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
SCN9A_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 29) | NC score | 0.309094 (rank : 33) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q15858, Q6B4R9, Q6B4S0, Q6B4S1, Q70HX1, Q70HX2, Q8WTU1, Q8WWN4 | Gene names | SCN9A, NENA, PN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 9 subunit alpha (Sodium channel protein type IX subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.7) (Neuroendocrine sodium channel) (hNE-Na) (Peripheral sodium channel 1). | |||||
|
TRPA1_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 30) | NC score | 0.115220 (rank : 63) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BLA8 | Gene names | Trpa1, Anktm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily A member 1 (Ankyrin-like with transmembrane domains protein 1). | |||||
|
TRPV3_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 31) | NC score | 0.171489 (rank : 50) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8NET8, Q8NDW7, Q8NET9, Q8NFH2 | Gene names | TRPV3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3) (Vanilloid receptor-like 3) (VRL-3). | |||||
|
SCN5A_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 32) | NC score | 0.330244 (rank : 29) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q14524 | Gene names | SCN5A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 5 subunit alpha (Sodium channel protein type V subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.5) (Sodium channel protein, cardiac muscle alpha-subunit) (HH1). | |||||
|
TRPC3_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 33) | NC score | 0.175893 (rank : 49) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13507, O00593, Q15660 | Gene names | TRPC3, TRP3 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 3 (TrpC3) (Htrp-3) (Htrp3). | |||||
|
TRPC4_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 34) | NC score | 0.176229 (rank : 47) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UBN4, Q15721, Q9UIB0, Q9UIB1, Q9UIB2 | Gene names | TRPC4 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 4 (TrpC4) (trp-related protein 4) (hTrp-4) (hTrp4). | |||||
|
TRPC4_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 35) | NC score | 0.175985 (rank : 48) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QUQ5, Q62350, Q9QUQ9, Q9QZC0 | Gene names | Trpc4, Trrp4 | |||
|
Domain Architecture |
|
|||||
| Description | Short transient receptor potential channel 4 (TrpC4) (Receptor- activated cation channel TRP4) (Capacitative calcium entry channel Trp4). | |||||
|
CAC1A_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 36) | NC score | 0.350809 (rank : 27) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
TRPC3_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 37) | NC score | 0.177197 (rank : 46) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9QZC1, Q61058 | Gene names | Trpc3, Trp3, Trrp3 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 3 (TrpC3) (Transient receptor protein 3) (Mtrp3) (Receptor-activated cation channel TRP3) (trp-related protein 3). | |||||
|
CAC1A_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 38) | NC score | 0.353525 (rank : 25) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
CAC1I_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 39) | NC score | 0.328321 (rank : 30) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9P0X4, O95504, Q5JZ88, Q7Z6S9, Q8NFX6, Q9NZC8, Q9UH15, Q9UH30, Q9ULU9, Q9UNE6 | Gene names | CACNA1I, KIAA1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1I (Voltage- gated calcium channel subunit alpha Cav3.3) (Ca(v)3.3). | |||||
|
TRPV3_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 40) | NC score | 0.156029 (rank : 53) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8K424, Q5SV08 | Gene names | Trpv3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3). | |||||
|
SC10A_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 41) | NC score | 0.309203 (rank : 32) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q6QIY3, Q62243, Q6EWG7, Q6KCH7, Q703F9 | Gene names | Scn10a, Pn3, Sns | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 10 subunit alpha (Sodium channel protein type X subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.8) (Peripheral nerve sodium channel 3) (Sensory neuron sodium channel). | |||||
|
SC11A_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 42) | NC score | 0.316595 (rank : 31) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9R053, Q9JMD4 | Gene names | Scn11a, Nan, Nat, Sns2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 11 subunit alpha (Sodium channel protein type XI subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.9) (Sensory neuron sodium channel 2) (NaN). | |||||
|
SCN1A_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 43) | NC score | 0.304278 (rank : 36) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P35498, Q16172, Q96LA3, Q9C008 | Gene names | SCN1A, NAC1, SCN1 | |||
|
Domain Architecture |
|
|||||
| Description | Sodium channel protein type 1 subunit alpha (Sodium channel protein type I subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.1) (Sodium channel protein, brain I subunit alpha). | |||||
|
SCN4A_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 44) | NC score | 0.304462 (rank : 35) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P35499, Q15478, Q16447, Q7Z6B1 | Gene names | SCN4A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 4 subunit alpha (Sodium channel protein type IV subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.4) (Sodium channel protein, skeletal muscle subunit alpha) (SkM1). | |||||
|
TRPV5_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 45) | NC score | 0.146433 (rank : 54) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P69744 | Gene names | Trpv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 5 (TrpV5) (osm-9-like TRP channel 3) (OTRPC3) (Epithelial calcium channel 1) (ECaC1) (Calcium transport protein 2) (CaT2). | |||||
|
SCN2A_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 46) | NC score | 0.304022 (rank : 37) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99250, Q14472, Q9BZC9, Q9BZD0 | Gene names | SCN2A, NAC2, SCN2A2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 2 subunit alpha (Sodium channel protein type II subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.2) (Sodium channel protein, brain II subunit alpha) (HBSC II). | |||||
|
TRPM7_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 47) | NC score | 0.103347 (rank : 65) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q923J1, Q8C7S7, Q8CE54, Q921Y5, Q925B2, Q9CUT2, Q9JLQ1 | Gene names | Trpm7, Chak, Ltrpc7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 7 (EC 2.7.11.1) (Long transient receptor potential channel 7) (LTrpC7) (Channel-kinase 1) (Transient receptor potential-phospholipase C- interacting kinase) (TRP-PLIK). | |||||
|
SCN3A_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 48) | NC score | 0.303270 (rank : 38) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9NY46, Q16142, Q9BZB3, Q9C006, Q9NYK2, Q9UPD1, Q9Y6P4 | Gene names | SCN3A, NAC3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 3 subunit alpha (Sodium channel protein type III subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.3) (Sodium channel protein, brain III subunit alpha) (Voltage- gated sodium channel subtype III). | |||||
|
SC10A_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 49) | NC score | 0.308681 (rank : 34) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Y5Y9 | Gene names | SCN10A, PN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 10 subunit alpha (Sodium channel protein type X subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.8) (Peripheral nerve sodium channel 3) (hPN3). | |||||
|
TRPM7_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 50) | NC score | 0.099600 (rank : 70) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96QT4, Q6ZMF5, Q86VJ4, Q8NBW2, Q9BXB2, Q9NXQ2 | Gene names | TRPM7, CHAK1, LTRPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 7 (EC 2.7.11.1) (Long transient receptor potential channel 7) (LTrpC7) (Channel-kinase 1). | |||||
|
SCN8A_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 51) | NC score | 0.294444 (rank : 42) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9WTU3, Q60828, Q60858, Q62449 | Gene names | Scn8a, Nbna1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 8 subunit alpha (Sodium channel protein type VIII subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.6). | |||||
|
CAC1G_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 52) | NC score | 0.299840 (rank : 39) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
SCN8A_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 53) | NC score | 0.295015 (rank : 41) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UQD0, O95788, Q9NYX2, Q9UPB2 | Gene names | SCN8A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 8 subunit alpha (Sodium channel protein type VIII subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.6). | |||||
|
TRPV4_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 54) | NC score | 0.139589 (rank : 59) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9EPK8, Q91XR5, Q9EQZ4, Q9ERZ7, Q9ES76 | Gene names | Trpv4, Trp12, Vrl2, Vroac | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 4 (TrpV4) (osm-9-like TRP channel 4) (OTRPC4) (Vanilloid receptor-like channel 2) (Vanilloid receptor-like protein 2) (Vanilloid receptor- related osmotically-activated channel) (VR-OAC) (Transient receptor potential protein 12) (TRP12). | |||||
|
TRPV4_HUMAN
|
||||||
| θ value | 0.125558 (rank : 55) | NC score | 0.138619 (rank : 60) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HBA0, Q86YZ6, Q8NDY7, Q8NG64, Q96Q92, Q96RS7, Q9HBC0 | Gene names | TRPV4, VRL2, VROAC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 4 (TrpV4) (osm-9-like TRP channel 4) (OTRPC4) (Vanilloid receptor-like channel 2) (Vanilloid receptor-like protein 2) (VRL-2) (Vanilloid receptor-related osmotically-activated channel) (VR-OAC) (Transient receptor potential protein 12) (TRP12). | |||||
|
TRPC5_HUMAN
|
||||||
| θ value | 0.163984 (rank : 56) | NC score | 0.141764 (rank : 55) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UL62, O75233, Q5JXY8, Q9Y514 | Gene names | TRPC5, TRP5 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 5 (TrpC5) (Htrp-5) (Htrp5). | |||||
|
TRPC5_MOUSE
|
||||||
| θ value | 0.163984 (rank : 57) | NC score | 0.141594 (rank : 56) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QX29, Q61059, Q9QWT1, Q9R0D4 | Gene names | Trpc5, Trp5, Trrp5 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 5 (TrpC5) (Transient receptor protein 5) (Mtrp5) (trp-related protein 5) (Capacitative calcium entry channel 2) (CCE2). | |||||
|
CAC1H_HUMAN
|
||||||
| θ value | 0.21417 (rank : 58) | NC score | 0.289441 (rank : 43) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O95180, O95802, Q8WWI6, Q96QI6, Q96RZ9, Q9NYY4, Q9NYY5 | Gene names | CACNA1H | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2) (Low-voltage-activated calcium channel alpha1 3.2 subunit). | |||||
|
TRPC6_HUMAN
|
||||||
| θ value | 0.279714 (rank : 59) | NC score | 0.156782 (rank : 52) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y210, Q9HCW3, Q9NQA8, Q9NQA9 | Gene names | TRPC6, TRP6 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 6 (TrpC6). | |||||
|
TRPC6_MOUSE
|
||||||
| θ value | 0.279714 (rank : 60) | NC score | 0.157271 (rank : 51) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61143, Q9Z2J1 | Gene names | Trpc6, Trp6, Trrp6 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 6 (TrpC6) (Calcium entry channel). | |||||
|
MRP9_HUMAN
|
||||||
| θ value | 0.365318 (rank : 61) | NC score | 0.014956 (rank : 94) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96J65, Q49AL2, Q8TAF0, Q8TEY2 | Gene names | ABCC12, MRP9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multidrug resistance-associated protein 9 (ATP-binding cassette transporter sub-family C member 12). | |||||
|
KU70_HUMAN
|
||||||
| θ value | 0.47712 (rank : 62) | NC score | 0.053801 (rank : 83) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P12956, Q9UCQ2, Q9UCQ3 | Gene names | XRCC6, G22P1 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase 2 subunit 1 (EC 3.6.1.-) (ATP-dependent DNA helicase II 70 kDa subunit) (Lupus Ku autoantigen protein p70) (Ku70) (70 kDa subunit of Ku antigen) (Thyroid-lupus autoantigen) (TLAA) (CTC box-binding factor 75 kDa subunit) (CTCBF) (CTC75) (DNA-repair protein XRCC6). | |||||
|
KU70_MOUSE
|
||||||
| θ value | 0.813845 (rank : 63) | NC score | 0.044837 (rank : 85) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P23475, O88212, Q62027, Q62382, Q62453 | Gene names | Xrcc6, G22p1, Ku70 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase 2 subunit 1 (EC 3.6.1.-) (ATP-dependent DNA helicase II 70 kDa subunit) (Ku autoantigen protein p70 homolog) (Ku70) (CTC box-binding factor 75 kDa subunit) (CTCBF) (CTC75) (DNA- repair protein XRCC6). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 64) | NC score | 0.015152 (rank : 93) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
TRPV5_HUMAN
|
||||||
| θ value | 1.06291 (rank : 65) | NC score | 0.118258 (rank : 62) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NQA5, Q8NDW5, Q8NDX7, Q8NDX8, Q96PM6 | Gene names | TRPV5, ECAC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 5 (TrpV5) (osm-9-like TRP channel 3) (OTRPC3) (Epithelial calcium channel 1) (ECaC) (ECaC1) (Calcium transport protein 2) (CaT2). | |||||
|
ENPL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 66) | NC score | 0.035363 (rank : 86) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P14625, Q96A97 | Gene names | HSP90B1, TRA1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (gp96 homolog) (Tumor rejection antigen 1). | |||||
|
ENPL_MOUSE
|
||||||
| θ value | 1.38821 (rank : 67) | NC score | 0.034047 (rank : 87) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P08113, P11427 | Gene names | Hsp90b1, Tra-1, Tra1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (ERP99) (Polymorphic tumor rejection antigen 1) (Tumor rejection antigen gp96). | |||||
|
TRPV6_MOUSE
|
||||||
| θ value | 1.38821 (rank : 68) | NC score | 0.101312 (rank : 66) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q91WD2 | Gene names | Trpv6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 6 (TrpV6) (Epithelial calcium channel 2) (ECaC2) (Calcium transport protein 1) (CaT1). | |||||
|
TSH1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 69) | NC score | 0.017653 (rank : 90) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 558 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZSZ6, O60534, Q4LE29, Q53EU4 | Gene names | TSHZ1, SDCCAG33, TSH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 1 (Serologically defined colon cancer antigen 3) (Antigen NY-CO-33). | |||||
|
TSH1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 70) | NC score | 0.017065 (rank : 91) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5DTH5, Q6PE60, Q8VD19, Q9JLD5 | Gene names | Tshz1, Kiaa4206, Sdccag33, Tsh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 1 (Serologically defined colon cancer antigen 3 homolog). | |||||
|
SCN7A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 71) | NC score | 0.286844 (rank : 44) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q01118 | Gene names | SCN7A, SCN6A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 7 subunit alpha (Sodium channel protein type VII subunit alpha) (Putative voltage-gated sodium channel alpha subunit Nax) (Sodium channel protein, cardiac and skeletal muscle alpha-subunit). | |||||
|
BAZ1B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.009705 (rank : 97) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
CAC1H_MOUSE
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.284728 (rank : 45) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O88427, Q80TJ2, Q9JKU5 | Gene names | Cacna1h, Kiaa1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2). | |||||
|
TRPV6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 74) | NC score | 0.108153 (rank : 64) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9H1D0, Q8TDL3, Q8WXR8, Q96LC5, Q9H1D1, Q9H296 | Gene names | TRPV6, ECAC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 6 (TrpV6) (Epithelial calcium channel 2) (ECaC2) (Calcium transport protein 1) (CaT1) (CaT-like) (CaT-L). | |||||
|
ANR26_MOUSE
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.009262 (rank : 98) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
ITPR1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.019153 (rank : 88) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14643, Q14660, Q99897 | Gene names | ITPR1, INSP3R1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 1 (Type 1 inositol 1,4,5- trisphosphate receptor) (Type 1 InsP3 receptor) (IP3 receptor isoform 1) (InsP3R1) (IP3R). | |||||
|
ITPR1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.019150 (rank : 89) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P11881, P20943, Q99LG5 | Gene names | Itpr1, Insp3r, Pcd6, Pcp1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 1 (Type 1 inositol 1,4,5- trisphosphate receptor) (Type 1 InsP3 receptor) (IP3 receptor isoform 1) (InsP3R1) (Inositol 1,4,5-trisphosphate-binding protein P400) (Purkinje cell protein 1) (Protein PCD-6). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.009748 (rank : 96) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
OR6C1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 79) | NC score | -0.002596 (rank : 107) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 740 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96RD1 | Gene names | OR6C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Olfactory receptor 6C1 (OST267). | |||||
|
TAAR8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | -0.000392 (rank : 105) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q969N4, Q5VUQ0 | Gene names | TAAR8, GPR102, TA5, TAR5, TRAR5 | |||
|
Domain Architecture |
|
|||||
| Description | Trace amine-associated receptor 8 (Trace amine receptor 5) (TaR-5) (G- protein coupled receptor 102). | |||||
|
CCD93_MOUSE
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.007916 (rank : 100) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7TQK5, Q3TX53 | Gene names | Ccdc93 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 93. | |||||
|
DYHC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.005312 (rank : 103) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 729 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JHU4 | Gene names | Dync1h1, Dnch1, Dnchc1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynein heavy chain, cytosolic (DYHC) (Cytoplasmic dynein heavy chain 1) (DHC1) (Dynein heavy chain 1, cytoplasmic 1). | |||||
|
OR9I1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | -0.002760 (rank : 108) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NGQ6, Q96RA8 | Gene names | OR9I1 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 9I1. | |||||
|
PRS6B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.006356 (rank : 102) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P43686, Q96FV5, Q9UBM3, Q9UEX3 | Gene names | PSMC4, TBP7 | |||
|
Domain Architecture |
|
|||||
| Description | 26S protease regulatory subunit 6B (Proteasome 26S subunit ATPase 4) (MIP224) (MB67-interacting protein) (TAT-binding protein 7) (TBP-7). | |||||
|
PRS6B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.006405 (rank : 101) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P54775 | Gene names | Psmc4, Tbp7 | |||
|
Domain Architecture |
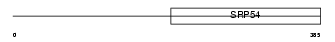
|
|||||
| Description | 26S protease regulatory subunit 6B (Proteasome 26S subunit ATPase 4) (MIP224) (MB67-interacting protein) (TAT-binding protein 7) (TBP-7) (CIP21). | |||||
|
TAA8B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | -0.001204 (rank : 106) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5QD06 | Gene names | Taar8b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trace amine-associated receptor 8b. | |||||
|
FYB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.015489 (rank : 92) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O35601, Q9Z2H3 | Gene names | Fyb | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
ANR26_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.008378 (rank : 99) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1614 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UPS8, Q2TAZ3, Q6ZR14, Q9H1Q1, Q9NSK9, Q9NTD5, Q9NW69 | Gene names | ANKRD26, KIAA1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
FYB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.010714 (rank : 95) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15117, O00359 | Gene names | FYB, SLAP130 | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
RCN1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.004802 (rank : 104) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15293 | Gene names | RCN1, RCN | |||
|
Domain Architecture |
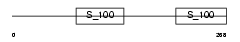
|
|||||
| Description | Reticulocalbin-1 precursor. | |||||
|
KCNF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.056082 (rank : 81) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9H3M0, O43527 | Gene names | KCNF1 | |||
|
Domain Architecture |
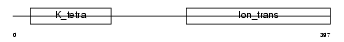
|
|||||
| Description | Potassium voltage-gated channel subfamily F member 1 (Voltage-gated potassium channel subunit Kv5.1) (kH1). | |||||
|
KCNV2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.051344 (rank : 84) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8TDN2 | Gene names | KCNV2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily V member 2 (Voltage-gated potassium channel subunit Kv8.2). | |||||
|
TRPC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.100888 (rank : 68) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P48995 | Gene names | TRPC1, TRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 1 (TrpC1) (TRP-1 protein). | |||||
|
TRPC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.100942 (rank : 67) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61056, O35722 | Gene names | Trpc1, Trp1, Trrp1 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 1 (TrpC1) (Transient receptor protein 1) (Mtrp1) (Trp-related protein 1). | |||||
|
TRPC2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.099821 (rank : 69) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R244, Q9ES59, Q9ES60, Q9R243 | Gene names | Trpc2, Trp2, Trrp2 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 2 (TrpC2) (mTrp2). | |||||
|
TRPC7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.140417 (rank : 58) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9HCX4 | Gene names | TRPC7, TRP7 | |||
|
Domain Architecture |
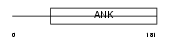
|
|||||
| Description | Short transient receptor potential channel 7 (TrpC7) (TRP7 protein). | |||||
|
TRPC7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.140507 (rank : 57) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9WVC5 | Gene names | Trpc7, Trp7, Trrp8 | |||
|
Domain Architecture |
|
|||||
| Description | Short transient receptor potential channel 7 (TrpC7) (mTRP7). | |||||
|
TRPM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.056724 (rank : 80) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O94759, Q5KTC2, Q6J3P5, Q96KN6, Q96Q93 | Gene names | TRPM2, EREG1, KNP3, LTRPC2, TRPC7 | |||
|
Domain Architecture |

|
|||||
| Description | Transient receptor potential cation channel subfamily M member 2 (EC 3.6.1.13) (Long transient receptor potential channel 2) (LTrpC2) (LTrpC-2) (Transient receptor potential channel 7) (TrpC7) (Estrogen- responsive element-associated gene 1 protein). | |||||
|
TRPM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.057117 (rank : 79) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q91YD4 | Gene names | Trpm2, Ltrpc2, Trpc7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 2 (EC 3.6.1.13) (Long transient receptor potential channel 2) (LTrpC2) (LTrpC-2) (Transient receptor potential channel 7) (TrpC7). | |||||
|
TRPM3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.058195 (rank : 77) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9HCF6, Q5VW02, Q5VW03, Q5VW04, Q5W5T7, Q86SH0, Q86SH6, Q86UL0, Q86WK1, Q86WK2, Q86WK3, Q86WK4, Q86YZ9, Q86Z00, Q86Z01 | Gene names | TRPM3, KIAA1616, LTRPC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 3 (Long transient receptor potential channel 3) (LTrpC3) (Melastatin-2) (MLSN2). | |||||
|
TRPM6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.055959 (rank : 82) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BX84, Q6VPR8, Q6VPR9, Q6VPS0, Q6VPS1, Q6VPS2 | Gene names | TRPM6, CHAK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 6 (EC 2.7.11.1) (Channel kinase 2) (Melastatin-related TRP cation channel 6). | |||||
|
TRPM6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.057296 (rank : 78) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CIR4 | Gene names | Trpm6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 6 (EC 2.7.11.1) (Channel kinase 2) (Melastatin-related TRP cation channel 6). | |||||
|
TRPM8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.078610 (rank : 74) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7Z2W7, Q6QNH9, Q8TAC3, Q8TDX8, Q9BVK1 | Gene names | TRPM8, LTRPC6, TRPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 8 (Transient receptor potential-p8) (Trp-p8) (Long transient receptor potential channel 6) (LTrpC6). | |||||
|
TRPM8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.079663 (rank : 73) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8R4D5 | Gene names | Trpm8, Trpp8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 8. | |||||
|
TRPV1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.087570 (rank : 71) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8NER1, Q9H0G9, Q9H303, Q9H304, Q9NQ74, Q9NY22 | Gene names | TRPV1, VR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 1 (TrpV1) (osm-9-like TRP channel 1) (OTRPC1) (Vanilloid receptor 1) (Capsaicin receptor). | |||||
|
TRPV1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.084097 (rank : 72) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q704Y3, Q5SSE1, Q5SSE2, Q5SSE4, Q5WPV5, Q68SW0 | Gene names | Trpv1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 1 (TrpV1) (osm-9-like TRP channel 1) (OTRPC1) (Vanilloid receptor 1). | |||||
|
TRPV2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.074807 (rank : 75) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y5S1, Q9Y670 | Gene names | TRPV2, VRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 2 (TrpV2) (osm-9-like TRP channel 2) (OTRPC2) (Vanilloid receptor-like protein 1) (VRL-1). | |||||
|
TRPV2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.073706 (rank : 76) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9WTR1, Q99K71 | Gene names | Trpv2, Grc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 2 (TrpV2) (osm-9-like TRP channel 2) (OTRPC2) (Growth factor-regulated calcium channel) (GRC). | |||||
|
PK2L2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q9NZM6, Q9UNJ0 | Gene names | PKD2L2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycystic kidney disease 2-like 2 protein (Polycystin-L2). | |||||
|
PK2L2_MOUSE
|
||||||
| NC score | 0.989577 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9JLG4 | Gene names | Pkd2l2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 2-like 2 protein (Polycystin-L2). | |||||
|
PKD2_HUMAN
|
||||||
| NC score | 0.962148 (rank : 3) | θ value | 5.99046e-137 (rank : 4) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q13563, O60441, Q15764 | Gene names | PKD2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-2 (Polycystic kidney disease 2 protein homolog) (Autosomal dominant polycystic kidney disease type II protein) (Polycystwin) (R48321). | |||||
|
PK2L1_HUMAN
|
||||||
| NC score | 0.961548 (rank : 4) | θ value | 1.42583e-138 (rank : 3) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9P0L9, O75972, Q9UP35, Q9UPA2 | Gene names | PKD2L1, PKD2L, PKDL | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 2-like 1 protein (Polycystin-L) (Polycystin- 2 homolog). | |||||
|
PKD2_MOUSE
|
||||||
| NC score | 0.958962 (rank : 5) | θ value | 2.78914e-126 (rank : 5) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | O35245, Q9QWP0, Q9Z193, Q9Z194 | Gene names | Pkd2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-2 (Polycystic kidney disease 2 protein homolog). | |||||
|
PKDRE_HUMAN
|
||||||
| NC score | 0.587415 (rank : 6) | θ value | 2.5182e-18 (rank : 6) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9NTG1, O95850 | Gene names | PKDREJ | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease and receptor for egg jelly-related protein precursor (PKD and REJ homolog). | |||||
|
PKDRE_MOUSE
|
||||||
| NC score | 0.559714 (rank : 7) | θ value | 5.60996e-18 (rank : 7) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Z0T6 | Gene names | Pkdrej | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease and receptor for egg jelly-related protein precursor (PKD and REJ homolog). | |||||
|
PK1L1_HUMAN
|
||||||
| NC score | 0.475296 (rank : 8) | θ value | 4.59992e-12 (rank : 9) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TDX9, Q6UWK1 | Gene names | PKD1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 1-like 1 protein (Polycystin-1L1). | |||||
|
MCLN2_MOUSE
|
||||||
| NC score | 0.408413 (rank : 9) | θ value | 1.25267e-09 (rank : 11) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8K595, Q3UCG4, Q8K2T6, Q9CQD3 | Gene names | Mcoln2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-2. | |||||
|
MCLN2_HUMAN
|
||||||
| NC score | 0.398726 (rank : 10) | θ value | 3.08544e-08 (rank : 18) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8IZK6, Q8N9R3 | Gene names | MCOLN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-2. | |||||
|
CAC1F_MOUSE
|
||||||
| NC score | 0.395944 (rank : 11) | θ value | 1.25267e-09 (rank : 10) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JIS7 | Gene names | Cacna1f | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
MCLN3_HUMAN
|
||||||
| NC score | 0.395519 (rank : 12) | θ value | 3.64472e-09 (rank : 13) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TDD5, Q5T4H5, Q5T4H6, Q9NV09 | Gene names | MCOLN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-3. | |||||
|
MCLN3_MOUSE
|
||||||
| NC score | 0.393347 (rank : 13) | θ value | 4.76016e-09 (rank : 14) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8R4F0, Q8BS73, Q8BSG1, Q8CDB2 | Gene names | Mcoln3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-3. | |||||
|
CAC1F_HUMAN
|
||||||
| NC score | 0.392289 (rank : 14) | θ value | 3.64472e-09 (rank : 12) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O60840, O43901 | Gene names | CACNA1F, CACNAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
CAC1C_MOUSE
|
||||||
| NC score | 0.390830 (rank : 15) | θ value | 1.38499e-08 (rank : 15) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q01815, Q04476, Q61242, Q99242 | Gene names | Cacna1c, Cach2, Cacn2, Cacnl1a1, Cchl1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle) (Mouse brain class C) (MBC) (MELC-CC). | |||||
|
CAC1C_HUMAN
|
||||||
| NC score | 0.389723 (rank : 16) | θ value | 2.36244e-08 (rank : 17) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
CAC1D_HUMAN
|
||||||
| NC score | 0.384016 (rank : 17) | θ value | 1.38499e-08 (rank : 16) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q01668, Q13916, Q13931 | Gene names | CACNA1D, CACH3, CACN4, CACNL1A2, CCHL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1D (Voltage- gated calcium channel subunit alpha Cav1.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 2). | |||||
|
CAC1E_MOUSE
|
||||||
| NC score | 0.382014 (rank : 18) | θ value | 2.21117e-06 (rank : 19) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q61290 | Gene names | Cacna1e, Cach6, Cacnl1a6, Cchra1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent R-type calcium channel subunit alpha-1E (Voltage- gated calcium channel subunit alpha Cav2.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 6) (Brain calcium channel II) (BII). | |||||
|
CAC1E_HUMAN
|
||||||
| NC score | 0.381345 (rank : 19) | θ value | 2.88788e-06 (rank : 21) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q15878, Q14580, Q14581 | Gene names | CACNA1E, CACH6, CACNL1A6 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent R-type calcium channel subunit alpha-1E (Voltage- gated calcium channel subunit alpha Cav2.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 6) (Brain calcium channel II) (BII). | |||||
|
CAC1B_HUMAN
|
||||||
| NC score | 0.378169 (rank : 20) | θ value | 6.43352e-06 (rank : 22) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q00975 | Gene names | CACNA1B, CACH5, CACNL1A5 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
CAC1S_HUMAN
|
||||||
| NC score | 0.374344 (rank : 21) | θ value | 1.87187e-05 (rank : 24) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q13698, Q12896, Q13934 | Gene names | CACNA1S, CACH1, CACN1, CACNL1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1S (Voltage- gated calcium channel subunit alpha Cav1.1) (Calcium channel, L type, alpha-1 polypeptide, isoform 3, skeletal muscle). | |||||
|
CAC1S_MOUSE
|
||||||
| NC score | 0.371891 (rank : 22) | θ value | 1.87187e-05 (rank : 25) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q02789, Q99240 | Gene names | Cacna1s, Cach1, Cach1b, Cacn1, Cacnl1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1S (Voltage- gated calcium channel subunit alpha Cav1.1) (Calcium channel, L type, alpha-1 polypeptide, isoform 3, skeletal muscle). | |||||
|
CAC1B_MOUSE
|
||||||
| NC score | 0.368093 (rank : 23) | θ value | 4.1701e-05 (rank : 28) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O55017, Q60609 | Gene names | Cacna1b, Cach5, Cacnl1a5, Cchn1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
MCLN1_HUMAN
|
||||||
| NC score | 0.357168 (rank : 24) | θ value | 2.44474e-05 (rank : 26) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9GZU1, Q7Z4F7, Q9H292, Q9H4B3, Q9H4B5 | Gene names | MCOLN1, ML4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-1 (Mucolipidin) (MG-2). | |||||
|
CAC1A_MOUSE
|
||||||
| NC score | 0.353525 (rank : 25) | θ value | 0.00228821 (rank : 38) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
MCLN1_MOUSE
|
||||||
| NC score | 0.352552 (rank : 26) | θ value | 3.19293e-05 (rank : 27) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99J21 | Gene names | Mcoln1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucolipin-1 (Mucolipidin). | |||||
|
CAC1A_HUMAN
|
||||||
| NC score | 0.350809 (rank : 27) | θ value | 0.00175202 (rank : 36) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
SC11A_HUMAN
|
||||||
| NC score | 0.339744 (rank : 28) | θ value | 2.21117e-06 (rank : 20) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UI33, Q68K15, Q8NDX3, Q9UHE0, Q9UHM0 | Gene names | SCN11A, PN5, SCN12A, SNS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 11 subunit alpha (Sodium channel protein type XI subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.9) (Sensory neuron sodium channel 2) (Peripheral nerve sodium channel 5) (hNaN). | |||||
|
SCN5A_HUMAN
|
||||||
| NC score | 0.330244 (rank : 29) | θ value | 0.000461057 (rank : 32) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q14524 | Gene names | SCN5A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 5 subunit alpha (Sodium channel protein type V subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.5) (Sodium channel protein, cardiac muscle alpha-subunit) (HH1). | |||||
|
CAC1I_HUMAN
|
||||||
| NC score | 0.328321 (rank : 30) | θ value | 0.00228821 (rank : 39) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9P0X4, O95504, Q5JZ88, Q7Z6S9, Q8NFX6, Q9NZC8, Q9UH15, Q9UH30, Q9ULU9, Q9UNE6 | Gene names | CACNA1I, KIAA1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1I (Voltage- gated calcium channel subunit alpha Cav3.3) (Ca(v)3.3). | |||||
|
SC11A_MOUSE
|
||||||
| NC score | 0.316595 (rank : 31) | θ value | 0.00509761 (rank : 42) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9R053, Q9JMD4 | Gene names | Scn11a, Nan, Nat, Sns2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 11 subunit alpha (Sodium channel protein type XI subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.9) (Sensory neuron sodium channel 2) (NaN). | |||||
|
SC10A_MOUSE
|
||||||
| NC score | 0.309203 (rank : 32) | θ value | 0.00509761 (rank : 41) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q6QIY3, Q62243, Q6EWG7, Q6KCH7, Q703F9 | Gene names | Scn10a, Pn3, Sns | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 10 subunit alpha (Sodium channel protein type X subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.8) (Peripheral nerve sodium channel 3) (Sensory neuron sodium channel). | |||||
|
SCN9A_HUMAN
|
||||||
| NC score | 0.309094 (rank : 33) | θ value | 0.000158464 (rank : 29) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q15858, Q6B4R9, Q6B4S0, Q6B4S1, Q70HX1, Q70HX2, Q8WTU1, Q8WWN4 | Gene names | SCN9A, NENA, PN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 9 subunit alpha (Sodium channel protein type IX subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.7) (Neuroendocrine sodium channel) (hNE-Na) (Peripheral sodium channel 1). | |||||
|
SC10A_HUMAN
|
||||||
| NC score | 0.308681 (rank : 34) | θ value | 0.0431538 (rank : 49) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Y5Y9 | Gene names | SCN10A, PN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 10 subunit alpha (Sodium channel protein type X subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.8) (Peripheral nerve sodium channel 3) (hPN3). | |||||
|
SCN4A_HUMAN
|
||||||
| NC score | 0.304462 (rank : 35) | θ value | 0.0113563 (rank : 44) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P35499, Q15478, Q16447, Q7Z6B1 | Gene names | SCN4A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 4 subunit alpha (Sodium channel protein type IV subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.4) (Sodium channel protein, skeletal muscle subunit alpha) (SkM1). | |||||
|
SCN1A_HUMAN
|
||||||
| NC score | 0.304278 (rank : 36) | θ value | 0.0113563 (rank : 43) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P35498, Q16172, Q96LA3, Q9C008 | Gene names | SCN1A, NAC1, SCN1 | |||
|
Domain Architecture |
|
|||||
| Description | Sodium channel protein type 1 subunit alpha (Sodium channel protein type I subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.1) (Sodium channel protein, brain I subunit alpha). | |||||
|
SCN2A_HUMAN
|
||||||
| NC score | 0.304022 (rank : 37) | θ value | 0.0193708 (rank : 46) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99250, Q14472, Q9BZC9, Q9BZD0 | Gene names | SCN2A, NAC2, SCN2A2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 2 subunit alpha (Sodium channel protein type II subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.2) (Sodium channel protein, brain II subunit alpha) (HBSC II). | |||||
|
SCN3A_HUMAN
|
||||||
| NC score | 0.303270 (rank : 38) | θ value | 0.0252991 (rank : 48) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9NY46, Q16142, Q9BZB3, Q9C006, Q9NYK2, Q9UPD1, Q9Y6P4 | Gene names | SCN3A, NAC3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 3 subunit alpha (Sodium channel protein type III subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.3) (Sodium channel protein, brain III subunit alpha) (Voltage- gated sodium channel subtype III). | |||||
|
CAC1G_HUMAN
|
||||||
| NC score | 0.299840 (rank : 39) | θ value | 0.0736092 (rank : 52) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
PKD1_HUMAN
|
||||||
| NC score | 0.299181 (rank : 40) | θ value | 1.58096e-12 (rank : 8) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P98161, Q15140, Q15141 | Gene names | PKD1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-1 precursor (Autosomal dominant polycystic kidney disease protein 1). | |||||
|
SCN8A_HUMAN
|
||||||
| NC score | 0.295015 (rank : 41) | θ value | 0.0961366 (rank : 53) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UQD0, O95788, Q9NYX2, Q9UPB2 | Gene names | SCN8A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 8 subunit alpha (Sodium channel protein type VIII subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.6). | |||||
|
SCN8A_MOUSE
|
||||||
| NC score | 0.294444 (rank : 42) | θ value | 0.0563607 (rank : 51) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9WTU3, Q60828, Q60858, Q62449 | Gene names | Scn8a, Nbna1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 8 subunit alpha (Sodium channel protein type VIII subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.6). | |||||
|
CAC1H_HUMAN
|
||||||
| NC score | 0.289441 (rank : 43) | θ value | 0.21417 (rank : 58) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O95180, O95802, Q8WWI6, Q96QI6, Q96RZ9, Q9NYY4, Q9NYY5 | Gene names | CACNA1H | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2) (Low-voltage-activated calcium channel alpha1 3.2 subunit). | |||||
|
SCN7A_HUMAN
|
||||||
| NC score | 0.286844 (rank : 44) | θ value | 1.81305 (rank : 71) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q01118 | Gene names | SCN7A, SCN6A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 7 subunit alpha (Sodium channel protein type VII subunit alpha) (Putative voltage-gated sodium channel alpha subunit Nax) (Sodium channel protein, cardiac and skeletal muscle alpha-subunit). | |||||
|
CAC1H_MOUSE
|
||||||
| NC score | 0.284728 (rank : 45) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O88427, Q80TJ2, Q9JKU5 | Gene names | Cacna1h, Kiaa1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2). | |||||
|
TRPC3_MOUSE
|
||||||
| NC score | 0.177197 (rank : 46) | θ value | 0.00175202 (rank : 37) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9QZC1, Q61058 | Gene names | Trpc3, Trp3, Trrp3 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 3 (TrpC3) (Transient receptor protein 3) (Mtrp3) (Receptor-activated cation channel TRP3) (trp-related protein 3). | |||||
|
TRPC4_HUMAN
|
||||||
| NC score | 0.176229 (rank : 47) | θ value | 0.00102713 (rank : 34) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UBN4, Q15721, Q9UIB0, Q9UIB1, Q9UIB2 | Gene names | TRPC4 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 4 (TrpC4) (trp-related protein 4) (hTrp-4) (hTrp4). | |||||
|
TRPC4_MOUSE
|
||||||
| NC score | 0.175985 (rank : 48) | θ value | 0.00102713 (rank : 35) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QUQ5, Q62350, Q9QUQ9, Q9QZC0 | Gene names | Trpc4, Trrp4 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 4 (TrpC4) (Receptor- activated cation channel TRP4) (Capacitative calcium entry channel Trp4). | |||||
|
TRPC3_HUMAN
|
||||||
| NC score | 0.175893 (rank : 49) | θ value | 0.000461057 (rank : 33) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13507, O00593, Q15660 | Gene names | TRPC3, TRP3 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 3 (TrpC3) (Htrp-3) (Htrp3). | |||||
|
TRPV3_HUMAN
|
||||||
| NC score | 0.171489 (rank : 50) | θ value | 0.00035302 (rank : 31) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8NET8, Q8NDW7, Q8NET9, Q8NFH2 | Gene names | TRPV3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3) (Vanilloid receptor-like 3) (VRL-3). | |||||
|
TRPC6_MOUSE
|
||||||
| NC score | 0.157271 (rank : 51) | θ value | 0.279714 (rank : 60) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61143, Q9Z2J1 | Gene names | Trpc6, Trp6, Trrp6 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 6 (TrpC6) (Calcium entry channel). | |||||
|
TRPC6_HUMAN
|
||||||
| NC score | 0.156782 (rank : 52) | θ value | 0.279714 (rank : 59) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y210, Q9HCW3, Q9NQA8, Q9NQA9 | Gene names | TRPC6, TRP6 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 6 (TrpC6). | |||||
|
TRPV3_MOUSE
|
||||||
| NC score | 0.156029 (rank : 53) | θ value | 0.00228821 (rank : 40) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8K424, Q5SV08 | Gene names | Trpv3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3). | |||||
|
TRPV5_MOUSE
|
||||||
| NC score | 0.146433 (rank : 54) | θ value | 0.0113563 (rank : 45) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P69744 | Gene names | Trpv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 5 (TrpV5) (osm-9-like TRP channel 3) (OTRPC3) (Epithelial calcium channel 1) (ECaC1) (Calcium transport protein 2) (CaT2). | |||||
|
TRPC5_HUMAN
|
||||||
| NC score | 0.141764 (rank : 55) | θ value | 0.163984 (rank : 56) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UL62, O75233, Q5JXY8, Q9Y514 | Gene names | TRPC5, TRP5 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 5 (TrpC5) (Htrp-5) (Htrp5). | |||||
|
TRPC5_MOUSE
|
||||||
| NC score | 0.141594 (rank : 56) | θ value | 0.163984 (rank : 57) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QX29, Q61059, Q9QWT1, Q9R0D4 | Gene names | Trpc5, Trp5, Trrp5 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 5 (TrpC5) (Transient receptor protein 5) (Mtrp5) (trp-related protein 5) (Capacitative calcium entry channel 2) (CCE2). | |||||
|
TRPC7_MOUSE
|
||||||
| NC score | 0.140507 (rank : 57) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9WVC5 | Gene names | Trpc7, Trp7, Trrp8 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 7 (TrpC7) (mTRP7). | |||||
|
TRPC7_HUMAN
|
||||||
| NC score | 0.140417 (rank : 58) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9HCX4 | Gene names | TRPC7, TRP7 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 7 (TrpC7) (TRP7 protein). | |||||
|
TRPV4_MOUSE
|
||||||
| NC score | 0.139589 (rank : 59) | θ value | 0.0961366 (rank : 54) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9EPK8, Q91XR5, Q9EQZ4, Q9ERZ7, Q9ES76 | Gene names | Trpv4, Trp12, Vrl2, Vroac | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 4 (TrpV4) (osm-9-like TRP channel 4) (OTRPC4) (Vanilloid receptor-like channel 2) (Vanilloid receptor-like protein 2) (Vanilloid receptor- related osmotically-activated channel) (VR-OAC) (Transient receptor potential protein 12) (TRP12). | |||||
|
TRPV4_HUMAN
|
||||||
| NC score | 0.138619 (rank : 60) | θ value | 0.125558 (rank : 55) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HBA0, Q86YZ6, Q8NDY7, Q8NG64, Q96Q92, Q96RS7, Q9HBC0 | Gene names | TRPV4, VRL2, VROAC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 4 (TrpV4) (osm-9-like TRP channel 4) (OTRPC4) (Vanilloid receptor-like channel 2) (Vanilloid receptor-like protein 2) (VRL-2) (Vanilloid receptor-related osmotically-activated channel) (VR-OAC) (Transient receptor potential protein 12) (TRP12). | |||||
|
TRPA1_HUMAN
|
||||||
| NC score | 0.122411 (rank : 61) | θ value | 1.43324e-05 (rank : 23) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75762 | Gene names | TRPA1, ANKTM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily A member 1 (Ankyrin-like with transmembrane domains protein 1) (Transformation sensitive-protein p120). | |||||
|
TRPV5_HUMAN
|
||||||
| NC score | 0.118258 (rank : 62) | θ value | 1.06291 (rank : 65) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NQA5, Q8NDW5, Q8NDX7, Q8NDX8, Q96PM6 | Gene names | TRPV5, ECAC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 5 (TrpV5) (osm-9-like TRP channel 3) (OTRPC3) (Epithelial calcium channel 1) (ECaC) (ECaC1) (Calcium transport protein 2) (CaT2). | |||||
|
TRPA1_MOUSE
|
||||||
| NC score | 0.115220 (rank : 63) | θ value | 0.00020696 (rank : 30) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BLA8 | Gene names | Trpa1, Anktm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily A member 1 (Ankyrin-like with transmembrane domains protein 1). | |||||
|
TRPV6_HUMAN
|
||||||
| NC score | 0.108153 (rank : 64) | θ value | 2.36792 (rank : 74) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9H1D0, Q8TDL3, Q8WXR8, Q96LC5, Q9H1D1, Q9H296 | Gene names | TRPV6, ECAC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 6 (TrpV6) (Epithelial calcium channel 2) (ECaC2) (Calcium transport protein 1) (CaT1) (CaT-like) (CaT-L). | |||||
|
TRPM7_MOUSE
|
||||||
| NC score | 0.103347 (rank : 65) | θ value | 0.0193708 (rank : 47) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q923J1, Q8C7S7, Q8CE54, Q921Y5, Q925B2, Q9CUT2, Q9JLQ1 | Gene names | Trpm7, Chak, Ltrpc7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 7 (EC 2.7.11.1) (Long transient receptor potential channel 7) (LTrpC7) (Channel-kinase 1) (Transient receptor potential-phospholipase C- interacting kinase) (TRP-PLIK). | |||||
|
TRPV6_MOUSE
|
||||||
| NC score | 0.101312 (rank : 66) | θ value | 1.38821 (rank : 68) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q91WD2 | Gene names | Trpv6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 6 (TrpV6) (Epithelial calcium channel 2) (ECaC2) (Calcium transport protein 1) (CaT1). | |||||
|
TRPC1_MOUSE
|
||||||
| NC score | 0.100942 (rank : 67) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61056, O35722 | Gene names | Trpc1, Trp1, Trrp1 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 1 (TrpC1) (Transient receptor protein 1) (Mtrp1) (Trp-related protein 1). | |||||
|
TRPC1_HUMAN
|
||||||
| NC score | 0.100888 (rank : 68) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P48995 | Gene names | TRPC1, TRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 1 (TrpC1) (TRP-1 protein). | |||||
|
TRPC2_MOUSE
|
||||||
| NC score | 0.099821 (rank : 69) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R244, Q9ES59, Q9ES60, Q9R243 | Gene names | Trpc2, Trp2, Trrp2 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 2 (TrpC2) (mTrp2). | |||||
|
TRPM7_HUMAN
|
||||||
| NC score | 0.099600 (rank : 70) | θ value | 0.0431538 (rank : 50) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96QT4, Q6ZMF5, Q86VJ4, Q8NBW2, Q9BXB2, Q9NXQ2 | Gene names | TRPM7, CHAK1, LTRPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 7 (EC 2.7.11.1) (Long transient receptor potential channel 7) (LTrpC7) (Channel-kinase 1). | |||||
|
TRPV1_HUMAN
|
||||||
| NC score | 0.087570 (rank : 71) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8NER1, Q9H0G9, Q9H303, Q9H304, Q9NQ74, Q9NY22 | Gene names | TRPV1, VR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 1 (TrpV1) (osm-9-like TRP channel 1) (OTRPC1) (Vanilloid receptor 1) (Capsaicin receptor). | |||||
|
TRPV1_MOUSE
|
||||||
| NC score | 0.084097 (rank : 72) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q704Y3, Q5SSE1, Q5SSE2, Q5SSE4, Q5WPV5, Q68SW0 | Gene names | Trpv1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 1 (TrpV1) (osm-9-like TRP channel 1) (OTRPC1) (Vanilloid receptor 1). | |||||
|
TRPM8_MOUSE
|
||||||
| NC score | 0.079663 (rank : 73) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8R4D5 | Gene names | Trpm8, Trpp8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 8. | |||||
|
TRPM8_HUMAN
|
||||||
| NC score | 0.078610 (rank : 74) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7Z2W7, Q6QNH9, Q8TAC3, Q8TDX8, Q9BVK1 | Gene names | TRPM8, LTRPC6, TRPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 8 (Transient receptor potential-p8) (Trp-p8) (Long transient receptor potential channel 6) (LTrpC6). | |||||
|
TRPV2_HUMAN
|
||||||
| NC score | 0.074807 (rank : 75) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y5S1, Q9Y670 | Gene names | TRPV2, VRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 2 (TrpV2) (osm-9-like TRP channel 2) (OTRPC2) (Vanilloid receptor-like protein 1) (VRL-1). | |||||
|
TRPV2_MOUSE
|
||||||
| NC score | 0.073706 (rank : 76) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9WTR1, Q99K71 | Gene names | Trpv2, Grc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 2 (TrpV2) (osm-9-like TRP channel 2) (OTRPC2) (Growth factor-regulated calcium channel) (GRC). | |||||
|
TRPM3_HUMAN
|
||||||
| NC score | 0.058195 (rank : 77) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9HCF6, Q5VW02, Q5VW03, Q5VW04, Q5W5T7, Q86SH0, Q86SH6, Q86UL0, Q86WK1, Q86WK2, Q86WK3, Q86WK4, Q86YZ9, Q86Z00, Q86Z01 | Gene names | TRPM3, KIAA1616, LTRPC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 3 (Long transient receptor potential channel 3) (LTrpC3) (Melastatin-2) (MLSN2). | |||||
|
TRPM6_MOUSE
|
||||||
| NC score | 0.057296 (rank : 78) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CIR4 | Gene names | Trpm6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 6 (EC 2.7.11.1) (Channel kinase 2) (Melastatin-related TRP cation channel 6). | |||||
|
TRPM2_MOUSE
|
||||||
| NC score | 0.057117 (rank : 79) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q91YD4 | Gene names | Trpm2, Ltrpc2, Trpc7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 2 (EC 3.6.1.13) (Long transient receptor potential channel 2) (LTrpC2) (LTrpC-2) (Transient receptor potential channel 7) (TrpC7). | |||||
|
TRPM2_HUMAN
|
||||||
| NC score | 0.056724 (rank : 80) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O94759, Q5KTC2, Q6J3P5, Q96KN6, Q96Q93 | Gene names | TRPM2, EREG1, KNP3, LTRPC2, TRPC7 | |||
|
Domain Architecture |

|
|||||
| Description | Transient receptor potential cation channel subfamily M member 2 (EC 3.6.1.13) (Long transient receptor potential channel 2) (LTrpC2) (LTrpC-2) (Transient receptor potential channel 7) (TrpC7) (Estrogen- responsive element-associated gene 1 protein). | |||||
|
KCNF1_HUMAN
|
||||||
| NC score | 0.056082 (rank : 81) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9H3M0, O43527 | Gene names | KCNF1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily F member 1 (Voltage-gated potassium channel subunit Kv5.1) (kH1). | |||||
|
TRPM6_HUMAN
|
||||||
| NC score | 0.055959 (rank : 82) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BX84, Q6VPR8, Q6VPR9, Q6VPS0, Q6VPS1, Q6VPS2 | Gene names | TRPM6, CHAK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 6 (EC 2.7.11.1) (Channel kinase 2) (Melastatin-related TRP cation channel 6). | |||||
|
KU70_HUMAN
|
||||||
| NC score | 0.053801 (rank : 83) | θ value | 0.47712 (rank : 62) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P12956, Q9UCQ2, Q9UCQ3 | Gene names | XRCC6, G22P1 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase 2 subunit 1 (EC 3.6.1.-) (ATP-dependent DNA helicase II 70 kDa subunit) (Lupus Ku autoantigen protein p70) (Ku70) (70 kDa subunit of Ku antigen) (Thyroid-lupus autoantigen) (TLAA) (CTC box-binding factor 75 kDa subunit) (CTCBF) (CTC75) (DNA-repair protein XRCC6). | |||||
|
KCNV2_HUMAN
|
||||||
| NC score | 0.051344 (rank : 84) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8TDN2 | Gene names | KCNV2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily V member 2 (Voltage-gated potassium channel subunit Kv8.2). | |||||
|
KU70_MOUSE
|
||||||
| NC score | 0.044837 (rank : 85) | θ value | 0.813845 (rank : 63) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P23475, O88212, Q62027, Q62382, Q62453 | Gene names | Xrcc6, G22p1, Ku70 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase 2 subunit 1 (EC 3.6.1.-) (ATP-dependent DNA helicase II 70 kDa subunit) (Ku autoantigen protein p70 homolog) (Ku70) (CTC box-binding factor 75 kDa subunit) (CTCBF) (CTC75) (DNA- repair protein XRCC6). | |||||
|
ENPL_HUMAN
|
||||||
| NC score | 0.035363 (rank : 86) | θ value | 1.38821 (rank : 66) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P14625, Q96A97 | Gene names | HSP90B1, TRA1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (gp96 homolog) (Tumor rejection antigen 1). | |||||
|
ENPL_MOUSE
|
||||||
| NC score | 0.034047 (rank : 87) | θ value | 1.38821 (rank : 67) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P08113, P11427 | Gene names | Hsp90b1, Tra-1, Tra1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (ERP99) (Polymorphic tumor rejection antigen 1) (Tumor rejection antigen gp96). | |||||
|
ITPR1_HUMAN
|
||||||
| NC score | 0.019153 (rank : 88) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14643, Q14660, Q99897 | Gene names | ITPR1, INSP3R1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 1 (Type 1 inositol 1,4,5- trisphosphate receptor) (Type 1 InsP3 receptor) (IP3 receptor isoform 1) (InsP3R1) (IP3R). | |||||
|
ITPR1_MOUSE
|
||||||
| NC score | 0.019150 (rank : 89) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P11881, P20943, Q99LG5 | Gene names | Itpr1, Insp3r, Pcd6, Pcp1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 1 (Type 1 inositol 1,4,5- trisphosphate receptor) (Type 1 InsP3 receptor) (IP3 receptor isoform 1) (InsP3R1) (Inositol 1,4,5-trisphosphate-binding protein P400) (Purkinje cell protein 1) (Protein PCD-6). | |||||
|
TSH1_HUMAN
|
||||||
| NC score | 0.017653 (rank : 90) | θ value | 1.38821 (rank : 69) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 558 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZSZ6, O60534, Q4LE29, Q53EU4 | Gene names | TSHZ1, SDCCAG33, TSH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 1 (Serologically defined colon cancer antigen 3) (Antigen NY-CO-33). | |||||
|
TSH1_MOUSE
|
||||||
| NC score | 0.017065 (rank : 91) | θ value | 1.38821 (rank : 70) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5DTH5, Q6PE60, Q8VD19, Q9JLD5 | Gene names | Tshz1, Kiaa4206, Sdccag33, Tsh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 1 (Serologically defined colon cancer antigen 3 homolog). | |||||
|
FYB_MOUSE
|
||||||
| NC score | 0.015489 (rank : 92) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O35601, Q9Z2H3 | Gene names | Fyb | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.015152 (rank : 93) | θ value | 1.06291 (rank : 64) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
MRP9_HUMAN
|
||||||
| NC score | 0.014956 (rank : 94) | θ value | 0.365318 (rank : 61) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96J65, Q49AL2, Q8TAF0, Q8TEY2 | Gene names | ABCC12, MRP9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multidrug resistance-associated protein 9 (ATP-binding cassette transporter sub-family C member 12). | |||||
|
FYB_HUMAN
|
||||||
| NC score | 0.010714 (rank : 95) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15117, O00359 | Gene names | FYB, SLAP130 | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.009748 (rank : 96) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
BAZ1B_MOUSE
|
||||||
| NC score | 0.009705 (rank : 97) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
ANR26_MOUSE
|
||||||
| NC score | 0.009262 (rank : 98) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
ANR26_HUMAN
|
||||||
| NC score | 0.008378 (rank : 99) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1614 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UPS8, Q2TAZ3, Q6ZR14, Q9H1Q1, Q9NSK9, Q9NTD5, Q9NW69 | Gene names | ANKRD26, KIAA1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
CCD93_MOUSE
|
||||||
| NC score | 0.007916 (rank : 100) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7TQK5, Q3TX53 | Gene names | Ccdc93 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 93. | |||||
|
PRS6B_MOUSE
|
||||||
| NC score | 0.006405 (rank : 101) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P54775 | Gene names | Psmc4, Tbp7 | |||
|
Domain Architecture |

|
|||||
| Description | 26S protease regulatory subunit 6B (Proteasome 26S subunit ATPase 4) (MIP224) (MB67-interacting protein) (TAT-binding protein 7) (TBP-7) (CIP21). | |||||
|
PRS6B_HUMAN
|
||||||
| NC score | 0.006356 (rank : 102) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P43686, Q96FV5, Q9UBM3, Q9UEX3 | Gene names | PSMC4, TBP7 | |||
|
Domain Architecture |

|
|||||
| Description | 26S protease regulatory subunit 6B (Proteasome 26S subunit ATPase 4) (MIP224) (MB67-interacting protein) (TAT-binding protein 7) (TBP-7). | |||||
|
DYHC_MOUSE
|
||||||
| NC score | 0.005312 (rank : 103) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 729 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JHU4 | Gene names | Dync1h1, Dnch1, Dnchc1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynein heavy chain, cytosolic (DYHC) (Cytoplasmic dynein heavy chain 1) (DHC1) (Dynein heavy chain 1, cytoplasmic 1). | |||||
|
RCN1_HUMAN
|
||||||
| NC score | 0.004802 (rank : 104) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15293 | Gene names | RCN1, RCN | |||
|
Domain Architecture |

|
|||||
| Description | Reticulocalbin-1 precursor. | |||||
|
TAAR8_HUMAN
|
||||||
| NC score | -0.000392 (rank : 105) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q969N4, Q5VUQ0 | Gene names | TAAR8, GPR102, TA5, TAR5, TRAR5 | |||
|
Domain Architecture |

|
|||||
| Description | Trace amine-associated receptor 8 (Trace amine receptor 5) (TaR-5) (G- protein coupled receptor 102). | |||||
|
TAA8B_MOUSE
|
||||||
| NC score | -0.001204 (rank : 106) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5QD06 | Gene names | Taar8b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trace amine-associated receptor 8b. | |||||
|
OR6C1_HUMAN
|
||||||
| NC score | -0.002596 (rank : 107) | θ value | 4.03905 (rank : 79) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 740 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96RD1 | Gene names | OR6C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Olfactory receptor 6C1 (OST267). | |||||
|
OR9I1_HUMAN
|
||||||
| NC score | -0.002760 (rank : 108) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NGQ6, Q96RA8 | Gene names | OR9I1 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 9I1. | |||||