Please be patient as the page loads
|
UBP10_MOUSE
|
||||||
| SwissProt Accessions | P52479, Q91VY7 | Gene names | Usp10, Ode-1, Uchrp | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
UBP10_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.975313 (rank : 2) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q14694, Q9BWG7, Q9NSL7 | Gene names | USP10, KIAA0190 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
UBP10_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 150 | |
| SwissProt Accessions | P52479, Q91VY7 | Gene names | Usp10, Ode-1, Uchrp | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
UBP8_HUMAN
|
||||||
| θ value | 9.24701e-21 (rank : 3) | NC score | 0.709194 (rank : 20) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
UBP8_MOUSE
|
||||||
| θ value | 3.51386e-20 (rank : 4) | NC score | 0.741246 (rank : 6) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q80U87, Q80YP2, Q8R0D3, Q9EQU1, Q9WVP5 | Gene names | Usp8, Kiaa0055, Ubpy | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (mUBPy). | |||||
|
UB17L_HUMAN
|
||||||
| θ value | 8.95645e-16 (rank : 5) | NC score | 0.750550 (rank : 5) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q7RTZ2 | Gene names | USP17L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 17-like protein (EC 3.1.2.15) (Ubiquitin thioesterase 17-like) (Ubiquitin-specific-processing protease 17-like) (Deubiquitinating enzyme 17-like). | |||||
|
UBP42_HUMAN
|
||||||
| θ value | 1.16975e-15 (rank : 6) | NC score | 0.716165 (rank : 18) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q9H9J4, Q3C166, Q6P9B4 | Gene names | USP42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 42 (EC 3.1.2.15) (Ubiquitin thioesterase 42) (Ubiquitin-specific-processing protease 42) (Deubiquitinating enzyme 42). | |||||
|
UBP50_MOUSE
|
||||||
| θ value | 3.40345e-15 (rank : 7) | NC score | 0.753422 (rank : 3) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q6P8X6, Q9D558, Q9D9E7 | Gene names | Usp50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ubiquitin carboxyl-terminal hydrolase 50 (EC 3.1.2.15) (Ubiquitin thioesterase 50) (Ubiquitin-specific-processing protease 50) (Deubiquitinating enzyme 50). | |||||
|
UBP46_HUMAN
|
||||||
| θ value | 4.44505e-15 (rank : 8) | NC score | 0.739653 (rank : 7) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P62068, Q80V95, Q9H7U4, Q9H9T8 | Gene names | USP46 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 46 (EC 3.1.2.15) (Ubiquitin thioesterase 46) (Ubiquitin-specific-processing protease 46) (Deubiquitinating enzyme 46). | |||||
|
UBP46_MOUSE
|
||||||
| θ value | 4.44505e-15 (rank : 9) | NC score | 0.739653 (rank : 8) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P62069, Q80V95, Q9H7U4, Q9H9T8 | Gene names | Usp46 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 46 (EC 3.1.2.15) (Ubiquitin thioesterase 46) (Ubiquitin-specific-processing protease 46) (Deubiquitinating enzyme 46). | |||||
|
UBPW_MOUSE
|
||||||
| θ value | 4.91457e-14 (rank : 10) | NC score | 0.752280 (rank : 4) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q61068 | Gene names | Dub1, Dub-1 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase DUB-1 (EC 3.1.2.15) (Ubiquitin thioesterase DUB-1) (Ubiquitin-specific-processing protease DUB-1) (Deubiquitinating enzyme 1). | |||||
|
UBP36_HUMAN
|
||||||
| θ value | 2.43908e-13 (rank : 11) | NC score | 0.722768 (rank : 15) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9P275, Q8NDM8, Q9NVC8 | Gene names | USP36, KIAA1453 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 36 (EC 3.1.2.15) (Ubiquitin thioesterase 36) (Ubiquitin-specific-processing protease 36) (Deubiquitinating enzyme 36). | |||||
|
UBP3_MOUSE
|
||||||
| θ value | 3.18553e-13 (rank : 12) | NC score | 0.736033 (rank : 10) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q91W36 | Gene names | Usp3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 3 (EC 3.1.2.15) (Ubiquitin thioesterase 3) (Ubiquitin-specific-processing protease 3) (Deubiquitinating enzyme 3). | |||||
|
UBP15_HUMAN
|
||||||
| θ value | 5.43371e-13 (rank : 13) | NC score | 0.668979 (rank : 29) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9Y4E8, Q9HCA6, Q9UNP0, Q9Y5B5 | Gene names | USP15, KIAA0529 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 15 (EC 3.1.2.15) (Ubiquitin thioesterase 15) (Ubiquitin-specific-processing protease 15) (Deubiquitinating enzyme 15) (Unph-2) (Unph4). | |||||
|
UBP15_MOUSE
|
||||||
| θ value | 5.43371e-13 (rank : 14) | NC score | 0.669330 (rank : 27) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8R5H1, Q3UL25 | Gene names | Usp15 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 15 (EC 3.1.2.15) (Ubiquitin thioesterase 15) (Ubiquitin-specific-processing protease 15) (Deubiquitinating enzyme 15). | |||||
|
UBP12_HUMAN
|
||||||
| θ value | 1.2105e-12 (rank : 15) | NC score | 0.736049 (rank : 9) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O75317 | Gene names | USP12, UBH1, USP12L1 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 12 (EC 3.1.2.15) (Ubiquitin thioesterase 12) (Ubiquitin-specific-processing protease 12) (Deubiquitinating enzyme 12) (Ubiquitin-hydrolyzing enzyme 1). | |||||
|
UBP21_HUMAN
|
||||||
| θ value | 1.58096e-12 (rank : 16) | NC score | 0.729330 (rank : 12) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9UK80, Q9BTV1, Q9NYN4 | Gene names | USP21, USP23 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 21 (EC 3.1.2.15) (Ubiquitin thioesterase 21) (Ubiquitin-specific-processing protease 21) (Deubiquitinating enzyme 21) (NEDD8-specific protease). | |||||
|
UBP21_MOUSE
|
||||||
| θ value | 7.84624e-12 (rank : 17) | NC score | 0.726307 (rank : 14) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9QZL6, Q9D0R1 | Gene names | Usp21, Usp23 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 21 (EC 3.1.2.15) (Ubiquitin thioesterase 21) (Ubiquitin-specific-processing protease 21) (Deubiquitinating enzyme 21). | |||||
|
UBP11_HUMAN
|
||||||
| θ value | 1.02475e-11 (rank : 18) | NC score | 0.671024 (rank : 26) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P51784, Q8IUG6, Q9BWE1 | Gene names | USP11, UHX1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 11 (EC 3.1.2.15) (Ubiquitin thioesterase 11) (Ubiquitin-specific-processing protease 11) (Deubiquitinating enzyme 11). | |||||
|
UBP2_HUMAN
|
||||||
| θ value | 5.08577e-11 (rank : 19) | NC score | 0.721643 (rank : 16) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | O75604, Q96MB9, Q9BQ21 | Gene names | USP2, UBP41 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 2 (EC 3.1.2.15) (Ubiquitin thioesterase 2) (Ubiquitin-specific-processing protease 2) (Deubiquitinating enzyme 2) (41 kDa ubiquitin-specific protease). | |||||
|
UBP3_HUMAN
|
||||||
| θ value | 1.47974e-10 (rank : 20) | NC score | 0.729326 (rank : 13) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9Y6I4 | Gene names | USP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 3 (EC 3.1.2.15) (Ubiquitin thioesterase 3) (Ubiquitin-specific-processing protease 3) (Deubiquitinating enzyme 3). | |||||
|
UBP44_HUMAN
|
||||||
| θ value | 1.47974e-10 (rank : 21) | NC score | 0.690967 (rank : 23) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9H0E7 | Gene names | USP44 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 44 (EC 3.1.2.15) (Ubiquitin thioesterase 44) (Ubiquitin-specific-processing protease 44) (Deubiquitinating enzyme 44). | |||||
|
UBP50_HUMAN
|
||||||
| θ value | 1.9326e-10 (rank : 22) | NC score | 0.730346 (rank : 11) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q70EL3 | Gene names | USP50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive ubiquitin carboxyl-terminal hydrolase 50 (Inactive ubiquitin- specific peptidase 50). | |||||
|
UBP22_MOUSE
|
||||||
| θ value | 9.59137e-10 (rank : 23) | NC score | 0.711265 (rank : 19) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q5DU02, Q5SU81, Q66JV8 | Gene names | Usp22, Kiaa1063 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 22 (EC 3.1.2.15) (Ubiquitin thioesterase 22) (Ubiquitin-specific-processing protease 22) (Deubiquitinating enzyme 22). | |||||
|
UBP2_MOUSE
|
||||||
| θ value | 9.59137e-10 (rank : 24) | NC score | 0.716386 (rank : 17) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O88623, Q6X4T9, Q6X4U1, Q8VD74, Q9JJ87 | Gene names | Usp2, Ubp41 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 2 (EC 3.1.2.15) (Ubiquitin thioesterase 2) (Ubiquitin-specific-processing protease 2) (Deubiquitinating enzyme 2) (41 kDa ubiquitin-specific protease). | |||||
|
UBP48_HUMAN
|
||||||
| θ value | 1.63604e-09 (rank : 25) | NC score | 0.647255 (rank : 32) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q86UV5, Q2M3I4, Q5SZI4, Q5T3T5, Q6NX53, Q8N3F6, Q96F64, Q96IQ3, Q9H5N3, Q9H5T7, Q9NUJ6, Q9NXR0 | Gene names | USP48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 48 (EC 3.1.2.15) (Ubiquitin thioesterase 48) (Ubiquitin-specific-processing protease 48) (Deubiquitinating enzyme 48). | |||||
|
UBP48_MOUSE
|
||||||
| θ value | 1.63604e-09 (rank : 26) | NC score | 0.645152 (rank : 33) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q3V0C5, Q3TM39, Q571J9, Q8VDJ1 | Gene names | Usp48, Kiaa4202 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 48 (EC 3.1.2.15) (Ubiquitin thioesterase 48) (Ubiquitin-specific-processing protease 48) (Deubiquitinating enzyme 48). | |||||
|
UBP22_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 27) | NC score | 0.708998 (rank : 21) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9UPT9 | Gene names | USP22, KIAA1063, USP3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 22 (EC 3.1.2.15) (Ubiquitin thioesterase 22) (Ubiquitin-specific-processing protease 22) (Deubiquitinating enzyme 22). | |||||
|
UBP49_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 28) | NC score | 0.674604 (rank : 25) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q6P9L4 | Gene names | Usp49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 49 (EC 3.1.2.15) (Ubiquitin thioesterase 49) (Ubiquitin-specific-processing protease 49) (Deubiquitinating enzyme 49). | |||||
|
UBP49_HUMAN
|
||||||
| θ value | 3.64472e-09 (rank : 29) | NC score | 0.675131 (rank : 24) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q70CQ1, Q96CK4 | Gene names | USP49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 49 (EC 3.1.2.15) (Ubiquitin thioesterase 49) (Ubiquitin-specific-processing protease 49) (Deubiquitinating enzyme 49). | |||||
|
UBP14_HUMAN
|
||||||
| θ value | 1.38499e-08 (rank : 30) | NC score | 0.596021 (rank : 41) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P54578 | Gene names | USP14, TGT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 14 (EC 3.1.2.15) (Ubiquitin thioesterase 14) (Ubiquitin-specific-processing protease 14) (Deubiquitinating enzyme 14). | |||||
|
UBP14_MOUSE
|
||||||
| θ value | 1.38499e-08 (rank : 31) | NC score | 0.592282 (rank : 42) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9JMA1, Q543U5, Q923F2, Q9D0L0 | Gene names | Usp14 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 14 (EC 3.1.2.15) (Ubiquitin thioesterase 14) (Ubiquitin-specific-processing protease 14) (Deubiquitinating enzyme 14). | |||||
|
USP9X_MOUSE
|
||||||
| θ value | 1.80886e-08 (rank : 32) | NC score | 0.538301 (rank : 51) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
UBP4_HUMAN
|
||||||
| θ value | 5.26297e-08 (rank : 33) | NC score | 0.645034 (rank : 34) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q13107, O43452, O43453 | Gene names | USP4, UNP, UNPH | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 4 (EC 3.1.2.15) (Ubiquitin thioesterase 4) (Ubiquitin-specific-processing protease 4) (Deubiquitinating enzyme 4) (Ubiquitous nuclear protein homolog). | |||||
|
UBP4_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 34) | NC score | 0.643611 (rank : 35) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P35123, O54704 | Gene names | Usp4, Unp | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 4 (EC 3.1.2.15) (Ubiquitin thioesterase 4) (Ubiquitin-specific-processing protease 4) (Deubiquitinating enzyme 4) (Ubiquitous nuclear protein). | |||||
|
UBP33_HUMAN
|
||||||
| θ value | 6.87365e-08 (rank : 35) | NC score | 0.669310 (rank : 28) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8TEY7, Q8TEY6, Q96AV6, Q9H9F0, Q9UPQ5 | Gene names | USP33, KIAA1097, VDU1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 33 (EC 3.1.2.15) (Ubiquitin thioesterase 33) (Ubiquitin-specific-processing protease 33) (Deubiquitinating enzyme 33) (VHL-interacting deubiquitinating enzyme 1). | |||||
|
USP9X_HUMAN
|
||||||
| θ value | 8.97725e-08 (rank : 36) | NC score | 0.535439 (rank : 53) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
UBP51_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 37) | NC score | 0.699301 (rank : 22) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q70EK9, Q8IWJ8 | Gene names | USP51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 51 (EC 3.1.2.15) (Ubiquitin thioesterase 51) (Ubiquitin-specific-processing protease 51) (Deubiquitinating enzyme 51). | |||||
|
USP9Y_HUMAN
|
||||||
| θ value | 1.69304e-06 (rank : 38) | NC score | 0.530778 (rank : 54) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O00507, O14601 | Gene names | USP9Y, DFFRY, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-Y (EC 3.1.2.15) (Ubiquitin thioesterase FAF-Y) (Ubiquitin-specific-processing protease FAF-Y) (Deubiquitinating enzyme FAF-Y) (Fat facets protein-related, Y- linked) (Ubiquitin-specific protease 9, Y chromosome). | |||||
|
UBP20_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 39) | NC score | 0.661555 (rank : 31) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9Y2K6, Q96LG5, Q9UQN8, Q9UQP0 | Gene names | USP20, KIAA1003, LSFR3A | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 20 (EC 3.1.2.15) (Ubiquitin thioesterase 20) (Ubiquitin-specific-processing protease 20) (Deubiquitinating enzyme 20). | |||||
|
UBP37_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 40) | NC score | 0.550982 (rank : 47) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q86T82, Q9HCH8 | Gene names | USP37, KIAA1594 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 37 (EC 3.1.2.15) (Ubiquitin thioesterase 37) (Ubiquitin-specific-processing protease 37) (Deubiquitinating enzyme 37). | |||||
|
UBP43_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 41) | NC score | 0.618843 (rank : 36) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q8BUM9, Q8VDP5 | Gene names | Usp43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 43 (EC 3.1.2.15) (Ubiquitin thioesterase 43) (Ubiquitin-specific-processing protease 43) (Deubiquitinating enzyme 43). | |||||
|
UBP16_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 42) | NC score | 0.663391 (rank : 30) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9Y5T5, Q8NEL3 | Gene names | USP16 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 16 (EC 3.1.2.15) (Ubiquitin thioesterase 16) (Ubiquitin-specific-processing protease 16) (Deubiquitinating enzyme 16) (Ubiquitin-processing protease UBP-M). | |||||
|
UBP43_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 43) | NC score | 0.615127 (rank : 38) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q70EL4, Q8N2C5, Q96DQ6 | Gene names | USP43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 43 (EC 3.1.2.15) (Ubiquitin thioesterase 43) (Ubiquitin-specific-processing protease 43) (Deubiquitinating enzyme 43). | |||||
|
UBP7_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 44) | NC score | 0.587265 (rank : 43) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q93009 | Gene names | USP7, HAUSP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 7 (EC 3.1.2.15) (Ubiquitin thioesterase 7) (Ubiquitin-specific-processing protease 7) (Deubiquitinating enzyme 7) (Herpesvirus-associated ubiquitin-specific protease). | |||||
|
UBP18_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 45) | NC score | 0.615357 (rank : 37) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9UMW8, Q9NY71 | Gene names | USP18, ISG43 | |||
|
Domain Architecture |
|
|||||
| Description | Ubl carboxyl-terminal hydrolase 18 (EC 3.1.2.-) (Ubl thioesterase 18) (ISG15-specific-processing protease) (43 kDa ISG15-specific protease) (hUBP43). | |||||
|
UBP1_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 46) | NC score | 0.611951 (rank : 39) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | O94782, Q9UFR0, Q9UNJ3 | Gene names | USP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 1 (EC 3.1.2.15) (Ubiquitin thioesterase 1) (Ubiquitin-specific-processing protease 1) (Deubiquitinating enzyme 1) (hUBP). | |||||
|
SNUT2_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 47) | NC score | 0.565106 (rank : 46) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q3TIX9, Q3TI55, Q8BLI5, Q8BZZ5, Q9CQR1 | Gene names | Usp39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 2 (Inactive ubiquitin-specific peptidase 39). | |||||
|
SNUT2_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 48) | NC score | 0.548050 (rank : 48) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q53GS9, Q6NX47, Q96RK9, Q9BV89, Q9H381, Q9P050, Q9Y310 | Gene names | USP39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 2 (U4/U6.U5 tri-snRNP-associated 65 kDa protein) (65K) (Inactive ubiquitin-specific peptidase 39) (SAD1 homolog). | |||||
|
UBP31_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 49) | NC score | 0.607191 (rank : 40) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
UBP24_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 50) | NC score | 0.567658 (rank : 44) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9UPU5, Q6ZSY2, Q8N2Y4, Q9NXD1 | Gene names | USP24, KIAA1057 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 24 (EC 3.1.2.15) (Ubiquitin thioesterase 24) (Ubiquitin-specific-processing protease 24) (Deubiquitinating enzyme 24). | |||||
|
UBP26_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 51) | NC score | 0.436234 (rank : 66) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q99MX1 | Gene names | Usp26 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 26 (EC 3.1.2.15) (Ubiquitin thioesterase 26) (Ubiquitin-specific-processing protease 26) (Deubiquitinating enzyme 26). | |||||
|
UBP26_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 52) | NC score | 0.484729 (rank : 61) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9BXU7 | Gene names | USP26 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 26 (EC 3.1.2.15) (Ubiquitin thioesterase 26) (Ubiquitin-specific-processing protease 26) (Deubiquitinating enzyme 26). | |||||
|
UBP47_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 53) | NC score | 0.542144 (rank : 50) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q8BY87, Q5EBP2, Q6KAR9, Q80V06, Q8BHU1, Q8BI15, Q8BI16, Q8BUW4, Q91X25 | Gene names | Usp47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
UBP29_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 54) | NC score | 0.493668 (rank : 59) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9HBJ7 | Gene names | USP29 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 29 (EC 3.1.2.15) (Ubiquitin thioesterase 29) (Ubiquitin-specific-processing protease 29) (Deubiquitinating enzyme 29). | |||||
|
UBP47_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 55) | NC score | 0.538127 (rank : 52) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q96K76, Q658U0, Q86Y73, Q8TEP6, Q9BWI0, Q9NWN1 | Gene names | USP47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
UBP29_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 56) | NC score | 0.470823 (rank : 63) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9ES63 | Gene names | Usp29 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 29 (EC 3.1.2.15) (Ubiquitin thioesterase 29) (Ubiquitin-specific-processing protease 29) (Deubiquitinating enzyme 29). | |||||
|
BASP_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 57) | NC score | 0.068600 (rank : 77) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
UBP35_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 58) | NC score | 0.524328 (rank : 55) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9P2H5 | Gene names | USP35, KIAA1372, USP34 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 35 (EC 3.1.2.15) (Ubiquitin thioesterase 35) (Ubiquitin-specific-processing protease 35) (Deubiquitinating enzyme 35). | |||||
|
RBM19_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 59) | NC score | 0.027602 (rank : 102) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R3C6, Q8BHR0, Q9CW63 | Gene names | Rbm19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 19 (RNA-binding motif protein 19). | |||||
|
UBP30_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 60) | NC score | 0.566845 (rank : 45) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q70CQ3, Q8WTU7, Q96JX4, Q9BSS3 | Gene names | USP30 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 30 (EC 3.1.2.15) (Ubiquitin thioesterase 30) (Ubiquitin-specific-processing protease 30) (Deubiquitinating enzyme 30). | |||||
|
UBP5_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 61) | NC score | 0.423973 (rank : 67) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P45974, Q96J22 | Gene names | USP5, ISOT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP38_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 62) | NC score | 0.492363 (rank : 60) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8NB14, Q8NDF5, Q96DK6, Q96PZ6, Q9BY55 | Gene names | USP38, KIAA1891 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 38 (EC 3.1.2.15) (Ubiquitin thioesterase 38) (Ubiquitin-specific-processing protease 38) (Deubiquitinating enzyme 38) (HP43.8KD). | |||||
|
UBP13_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 63) | NC score | 0.406520 (rank : 69) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q92995 | Gene names | USP13, ISOT3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 13 (EC 3.1.2.15) (Ubiquitin thioesterase 13) (Ubiquitin-specific-processing protease 13) (Deubiquitinating enzyme 13) (Isopeptidase T-3) (ISOT-3). | |||||
|
UBP32_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 64) | NC score | 0.523349 (rank : 56) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8NFA0, Q9BX85, Q9Y591 | Gene names | USP32, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 32 (EC 3.1.2.15) (Ubiquitin thioesterase 32) (Ubiquitin-specific-processing protease 32) (Deubiquitinating enzyme 32) (NY-REN-60 antigen). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 65) | NC score | 0.035488 (rank : 94) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
UBP25_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 66) | NC score | 0.297859 (rank : 70) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9UHP3, Q9H9W1 | Gene names | USP25, USP21 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (USP on chromosome 21). | |||||
|
UBP5_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 67) | NC score | 0.415108 (rank : 68) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P56399 | Gene names | Usp5, Isot | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP6_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 68) | NC score | 0.496556 (rank : 58) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P35125, Q15634, Q86WP6, Q8IWT4 | Gene names | USP6, TRE2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 6 (EC 3.1.2.15) (Ubiquitin thioesterase 6) (Ubiquitin-specific-processing protease 6) (Deubiquitinating enzyme 6) (Proto-oncogene TRE-2). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 0.125558 (rank : 69) | NC score | 0.047925 (rank : 84) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 0.125558 (rank : 70) | NC score | 0.022152 (rank : 113) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
HSP74_MOUSE
|
||||||
| θ value | 0.163984 (rank : 71) | NC score | 0.021122 (rank : 117) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61316 | Gene names | Hspa4, Hsp110 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4 (Heat shock 70-related protein APG-2). | |||||
|
AP180_HUMAN
|
||||||
| θ value | 0.21417 (rank : 72) | NC score | 0.055674 (rank : 81) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60641, Q5JX13, Q68DL9, Q6P9D3, Q9NTY7 | Gene names | SNAP91, KIAA0656 | |||
|
Domain Architecture |
|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein). | |||||
|
RD23B_HUMAN
|
||||||
| θ value | 0.21417 (rank : 73) | NC score | 0.045129 (rank : 86) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P54727, Q8WUB0 | Gene names | RAD23B | |||
|
Domain Architecture |
|
|||||
| Description | UV excision repair protein RAD23 homolog B (hHR23B) (XP-C repair- complementing complex 58 kDa protein) (p58). | |||||
|
HCN1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 74) | NC score | 0.021725 (rank : 114) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60741 | Gene names | HCN1, BCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1). | |||||
|
OKL38_HUMAN
|
||||||
| θ value | 0.365318 (rank : 75) | NC score | 0.040523 (rank : 88) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJX0, Q86UQ1, Q96S88, Q9BZ70 | Gene names | OKL38 | |||
|
Domain Architecture |
|
|||||
| Description | Pregnancy-induced growth inhibitor OKL38 (huOKL38) (Ovary, kidney and liver protein 38). | |||||
|
CJ030_MOUSE
|
||||||
| θ value | 0.47712 (rank : 76) | NC score | 0.036353 (rank : 93) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BSV3, Q4VAD9, Q6PHN3, Q8BPJ4, Q8C973 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C10orf30 homolog. | |||||
|
HCN1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 77) | NC score | 0.020935 (rank : 118) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O88704, O54899, Q9D613 | Gene names | Hcn1, Bcng1, Hac2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1) (Hyperpolarization-activated cation channel 2) (HAC-2). | |||||
|
LDB3_MOUSE
|
||||||
| θ value | 0.47712 (rank : 78) | NC score | 0.015162 (rank : 127) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9JKS4, Q6A038, Q811P2, Q811P3, Q811P4, Q811P5, Q9D130, Q9JKS3, Q9R0Z1, Q9WVH1, Q9WVH2 | Gene names | Ldb3, Kiaa0613 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher) (Protein oracle). | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 79) | NC score | 0.036514 (rank : 92) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
NFRKB_MOUSE
|
||||||
| θ value | 0.47712 (rank : 80) | NC score | 0.042238 (rank : 87) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
TBX6_HUMAN
|
||||||
| θ value | 0.47712 (rank : 81) | NC score | 0.012917 (rank : 131) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95947, Q8TAS4 | Gene names | TBX6 | |||
|
Domain Architecture |
|
|||||
| Description | T-box transcription factor TBX6 (T-box protein 6). | |||||
|
TIAM1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 82) | NC score | 0.018540 (rank : 120) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60610 | Gene names | Tiam1, Tiam-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
UBP38_MOUSE
|
||||||
| θ value | 0.62314 (rank : 83) | NC score | 0.477954 (rank : 62) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8BW70, Q8BWL1, Q8BX03 | Gene names | Usp38 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 38 (EC 3.1.2.15) (Ubiquitin thioesterase 38) (Ubiquitin-specific-processing protease 38) (Deubiquitinating enzyme 38). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | 0.813845 (rank : 84) | NC score | 0.037748 (rank : 90) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
PO2F1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 85) | NC score | 0.013480 (rank : 130) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P25425, Q61994, Q63891, Q6Y681, Q7TSD0, Q8BT04, Q8K570, Q99JH0, Q9WTZ4, Q9WTZ5, Q9WTZ6 | Gene names | Pou2f1, Oct-1, Otf-1, Otf1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 2, transcription factor 1 (Octamer-binding transcription factor 1) (Oct-1) (OTF-1) (NF-A1). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 86) | NC score | 0.046320 (rank : 85) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
AP180_MOUSE
|
||||||
| θ value | 1.06291 (rank : 87) | NC score | 0.048232 (rank : 83) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61548, Q61547 | Gene names | Snap91 | |||
|
Domain Architecture |
|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein) (Phosphoprotein F1- 20). | |||||
|
FMN2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 88) | NC score | 0.021701 (rank : 115) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
PKHA6_HUMAN
|
||||||
| θ value | 1.06291 (rank : 89) | NC score | 0.031409 (rank : 97) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
X3CL1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 90) | NC score | 0.029056 (rank : 99) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P78423, O00672 | Gene names | CX3CL1, A-152E5.2, FKN, NTT, SCYD1 | |||
|
Domain Architecture |
|
|||||
| Description | Fractalkine precursor (CX3CL1) (Neurotactin) (CX3C membrane-anchored chemokine) (Small inducible cytokine D1). | |||||
|
GPR98_HUMAN
|
||||||
| θ value | 1.38821 (rank : 91) | NC score | 0.016156 (rank : 124) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WXG9, O75171, Q9H0X5, Q9UL61 | Gene names | GPR98, KIAA0686, MASS1, VLGR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor 98 precursor (Monogenic audiogenic seizure susceptibility protein 1 homolog) (Very large G-protein coupled receptor 1) (Usher syndrome type-2C protein). | |||||
|
LAP4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 92) | NC score | 0.007786 (rank : 143) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80U72, Q6P5H7, Q7TPH8, Q80VB1, Q8CI48, Q8VII1, Q922S3 | Gene names | Scrib, Kiaa0147, Lap4, Scrib1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 1.38821 (rank : 93) | NC score | 0.028678 (rank : 100) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
UBP25_MOUSE
|
||||||
| θ value | 1.38821 (rank : 94) | NC score | 0.288665 (rank : 73) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P57080, Q80ZT9 | Gene names | Usp25 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (mUSP25). | |||||
|
DDX24_HUMAN
|
||||||
| θ value | 1.81305 (rank : 95) | NC score | 0.009318 (rank : 139) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9GZR7 | Gene names | DDX24 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX24 (EC 3.6.1.-) (DEAD box protein 24). | |||||
|
K0460_MOUSE
|
||||||
| θ value | 1.81305 (rank : 96) | NC score | 0.027624 (rank : 101) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6NXI6, Q3U3L8, Q6ZQA7 | Gene names | Kiaa0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
MAP4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 97) | NC score | 0.034179 (rank : 95) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
RUFY2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 98) | NC score | 0.007755 (rank : 144) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
SLIK6_MOUSE
|
||||||
| θ value | 1.81305 (rank : 99) | NC score | 0.004746 (rank : 152) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C110, Q8BLL0 | Gene names | Slitrk6, Sltk6 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT and NTRK-like protein 6 precursor. | |||||
|
HSP74_HUMAN
|
||||||
| θ value | 2.36792 (rank : 100) | NC score | 0.016758 (rank : 122) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P34932, O95756 | Gene names | HSPA4 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4 (Heat shock 70-related protein APG-2) (HSP70RY). | |||||
|
INT1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 101) | NC score | 0.021522 (rank : 116) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6P4S8, Q80UQ7, Q91Z01, Q9CTF7 | Gene names | Ints1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 1 (Int1). | |||||
|
PK3C3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 102) | NC score | 0.014134 (rank : 128) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PF93, Q8R3S8 | Gene names | Pik3c3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3). | |||||
|
THAP8_HUMAN
|
||||||
| θ value | 2.36792 (rank : 103) | NC score | 0.026216 (rank : 106) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NA92, Q96M21 | Gene names | THAP8 | |||
|
Domain Architecture |
|
|||||
| Description | THAP domain-containing protein 8. | |||||
|
UBP28_HUMAN
|
||||||
| θ value | 2.36792 (rank : 104) | NC score | 0.296849 (rank : 71) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q96RU2, Q9P213 | Gene names | USP28, KIAA1515 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
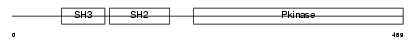
ABL1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 105) | NC score | -0.002546 (rank : 158) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P00520, P97896, Q61252, Q61253, Q61254, Q61255, Q61256, Q61257, Q61258, Q61259, Q61260, Q61261, Q6PCM5, Q8C1X4 | Gene names | Abl1, Abl | |||
|
Domain Architecture |
|
|||||
| Description | Proto-oncogene tyrosine-protein kinase ABL1 (EC 2.7.10.2) (p150) (c- ABL) (Abelson murine leukemia viral oncogene homolog 1). | |||||
|
BASP_MOUSE
|
||||||
| θ value | 3.0926 (rank : 106) | NC score | 0.050824 (rank : 82) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 107) | NC score | 0.026588 (rank : 104) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 108) | NC score | 0.026473 (rank : 105) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CD2L2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 109) | NC score | -0.001840 (rank : 157) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1439 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UQ88, O95227, O95228, O96012, Q12821, Q12853, Q12854, Q5QPR0, Q5QPR1, Q5QPR2, Q9UBC4, Q9UBI3, Q9UEI1, Q9UEI2, Q9UP53, Q9UP54, Q9UP55, Q9UP56, Q9UQ86, Q9UQ87, Q9UQ89 | Gene names | CDC2L2, PITSLREB | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L2 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 2) (CDK11). | |||||
|
CIZ1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 110) | NC score | 0.026030 (rank : 108) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9ULV3, Q9NYM8, Q9UHK4, Q9Y3F9, Q9Y3G0 | Gene names | CIZ1, LSFR1, NP94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cip1-interacting zinc finger protein (Nuclear protein NP94). | |||||
|
GNAS1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 111) | NC score | 0.009601 (rank : 137) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6R0H7, Q6R0H4, Q6R0H5, Q6R2J5, Q9JJX0, Q9Z1N8 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
LATS1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 112) | NC score | -0.000148 (rank : 156) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1027 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BYR2, Q9Z0W4 | Gene names | Lats1, Warts | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase). | |||||
|
RERE_HUMAN
|
||||||
| θ value | 3.0926 (rank : 113) | NC score | 0.024727 (rank : 109) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
RERE_MOUSE
|
||||||
| θ value | 3.0926 (rank : 114) | NC score | 0.026176 (rank : 107) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
UBP28_MOUSE
|
||||||
| θ value | 3.0926 (rank : 115) | NC score | 0.292529 (rank : 72) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q5I043, Q6NZP3, Q6ZPP1, Q8BWI1 | Gene names | Usp28, Kiaa1515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
ZN335_HUMAN
|
||||||
| θ value | 3.0926 (rank : 116) | NC score | -0.002669 (rank : 159) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H4Z2, Q9H684 | Gene names | ZNF335 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 335. | |||||
|
ZN592_MOUSE
|
||||||
| θ value | 3.0926 (rank : 117) | NC score | 0.002677 (rank : 154) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BHZ4, Q80XM1 | Gene names | Znf592, Kiaa0211, Zfp592 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592 (Zfp-592). | |||||
|
ANC5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 118) | NC score | 0.016079 (rank : 125) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJX4, Q8N4H7, Q9BQD4 | Gene names | ANAPC5, APC5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Anaphase-promoting complex subunit 5 (APC5) (Cyclosome subunit 5). | |||||
|
LAP4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 119) | NC score | 0.007232 (rank : 146) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 608 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14160, Q6P496, Q7Z5D1, Q8WWV8, Q96C69, Q96GG1 | Gene names | SCRIB, CRIB1, KIAA0147, LAP4, SCRB1, VARTUL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog) (hScrib). | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 120) | NC score | 0.022531 (rank : 112) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
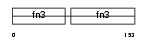
NCA11_MOUSE
|
||||||
| θ value | 4.03905 (rank : 121) | NC score | 0.008723 (rank : 141) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P13595, Q61949 | Gene names | Ncam1, Ncam | |||
|
Domain Architecture |
|
|||||
| Description | Neural cell adhesion molecule 1, 180 kDa isoform precursor (N-CAM 180) (NCAM-180). | |||||
|
NFRKB_HUMAN
|
||||||
| θ value | 4.03905 (rank : 122) | NC score | 0.036774 (rank : 91) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6P4R8, Q12869, Q15312, Q9H048 | Gene names | NFRKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
PKHA6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 123) | NC score | 0.026924 (rank : 103) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
SAPS2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 124) | NC score | 0.011259 (rank : 134) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75170, Q9UGB9 | Gene names | SAPS2, KIAA0685 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 2. | |||||
|
SREC_HUMAN
|
||||||
| θ value | 4.03905 (rank : 125) | NC score | 0.003138 (rank : 153) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 126) | NC score | 0.022684 (rank : 111) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
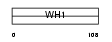
EVL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 127) | NC score | 0.023208 (rank : 110) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P70429, Q9ERU8 | Gene names | Evl | |||
|
Domain Architecture |
|
|||||
| Description | Ena/VASP-like protein (Ena/vasodilator-stimulated phosphoprotein- like). | |||||
|
FGD5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 128) | NC score | 0.009173 (rank : 140) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80UZ0, Q8BHM5 | Gene names | Fgd5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5. | |||||
|
RUFY2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 129) | NC score | 0.007208 (rank : 147) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8R4C2, Q69ZH1 | Gene names | Rufy2, Kiaa1537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Leucine zipper FYVE-finger protein) (LZ-FYVE). | |||||
|
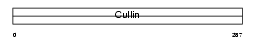
CUL4B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 130) | NC score | 0.006017 (rank : 149) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13620, Q6PIE4, Q6UP07, Q7Z673, Q9BY37, Q9UEB7, Q9UED7 | Gene names | CUL4B, KIAA0695 | |||
|
Domain Architecture |
|
|||||
| Description | Cullin-4B (CUL-4B). | |||||
|
DDX51_HUMAN
|
||||||
| θ value | 6.88961 (rank : 131) | NC score | 0.007891 (rank : 142) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
IL4RA_MOUSE
|
||||||
| θ value | 6.88961 (rank : 132) | NC score | 0.012786 (rank : 132) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P16382, O54690, Q60583, Q8CBW5 | Gene names | Il4r, Il4ra | |||
|
Domain Architecture |
|
|||||
| Description | Interleukin-4 receptor alpha chain precursor (IL-4R-alpha) (CD124 antigen) [Contains: Soluble interleukin-4 receptor alpha chain (IL-4- binding protein) (IL4-BP)]. | |||||
|
KCC2B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 133) | NC score | -0.004346 (rank : 160) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 866 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13554, O95437, O95438, O95599, Q9UGH7, Q9UGH8, Q9UGH9, Q9UNX0, Q9UNX7, Q9UP00, Q9Y5N4, Q9Y6F4 | Gene names | CAMK2B, CAM2, CAMKB | |||
|
Domain Architecture |

|
|||||
| Description | Calcium/calmodulin-dependent protein kinase type II beta chain (EC 2.7.11.17) (CaM-kinase II beta chain) (CaM kinase II subunit beta) (CaMK-II subunit beta). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 6.88961 (rank : 134) | NC score | 0.030200 (rank : 98) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
SYNJ1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 135) | NC score | 0.013986 (rank : 129) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
ZN592_HUMAN
|
||||||
| θ value | 6.88961 (rank : 136) | NC score | 0.001549 (rank : 155) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
AMOT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.009524 (rank : 138) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
ASXL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.018339 (rank : 121) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P59598 | Gene names | Asxl1, Kiaa0978 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative Polycomb group protein ASXL1 (Additional sex combs-like protein 1). | |||||
|
BICD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.005206 (rank : 151) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TD16, O75181, Q5TBQ2, Q5TBQ3, Q96LH2, Q9BT84, Q9H561 | Gene names | BICD2, KIAA0699 | |||
|
Domain Architecture |
|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
BICD2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.005661 (rank : 150) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q921C5, Q80TU1, Q8BTE3, Q9DCL3 | Gene names | Bicd2, Kiaa0699 | |||
|
Domain Architecture |
|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
CENPB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.007742 (rank : 145) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P07199 | Gene names | CENPB | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
ELK4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.006305 (rank : 148) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P28324, P28323 | Gene names | ELK4, SAP1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.020021 (rank : 119) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
NU214_HUMAN
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.039742 (rank : 89) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
OXA1L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.015437 (rank : 126) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15070 | Gene names | OXA1L | |||
|
Domain Architecture |
|
|||||
| Description | Inner membrane protein OXA1L, mitochondrial precursor (Oxidase assembly 1-like protein) (OXA1-like protein) (OXA1Hs) (Hsa). | |||||
|
PK3C3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | 0.010843 (rank : 135) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NEB9, Q15134 | Gene names | PIK3C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3) (Phosphatidylinositol 3-kinase p100 subunit). | |||||
|
RANB3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 147) | NC score | 0.016573 (rank : 123) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9H6Z4, O60405, O75759, O75760, Q9BT47, Q9UG74 | Gene names | RANBP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 3 (RanBP3). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 148) | NC score | 0.033277 (rank : 96) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TBCD4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 149) | NC score | 0.011908 (rank : 133) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BYJ6, Q5DU23, Q66JU2, Q6P2M2, Q8BMH6, Q8BXM2 | Gene names | Tbc1d4, Kiaa0603 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 4 (Akt substrate of 160 kDa) (AS160). | |||||
|
TENS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 150) | NC score | 0.010044 (rank : 136) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9HBL0 | Gene names | TNS1, TNS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-1. | |||||
|
BRAP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.074548 (rank : 76) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7Z569, O43238, O75341 | Gene names | BRAP, RNF52 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP) (RING finger protein 52) (NY-REN-63 antigen). | |||||
|
BRAP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.086237 (rank : 74) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q99MP8, Q8CC00, Q8CHX1, Q99MP7, Q9CXX8 | Gene names | Brap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP). | |||||
|
GGNB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.062788 (rank : 78) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5YKI7, Q5YKI8 | Gene names | GGNBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
GGNB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.078273 (rank : 75) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6K1E7, Q8C5T9 | Gene names | Ggnbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
HDAC6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.058105 (rank : 80) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UBN7 | Gene names | HDAC6 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 6 (HD6). | |||||
|
HDAC6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.060666 (rank : 79) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z2V5 | Gene names | Hdac6 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 6 (HD6) (Histone deacetylase mHDA2). | |||||
|
UBP18_MOUSE
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.543926 (rank : 49) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9WTV6 | Gene names | Usp18, Ubp43 | |||
|
Domain Architecture |
|
|||||
| Description | Ubl carboxyl-terminal hydrolase 18 (EC 3.1.2.-) (Ubl thioesterase 18) (ISG15-specific-processing protease) (43 kDa ISG15-specific protease). | |||||
|
UBP34_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.441461 (rank : 64) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q70CQ2, O60316, O94834, Q3B777, Q6P6C9, Q7L8P6, Q8N3T9, Q8TBW2, Q9UGA1 | Gene names | USP34, KIAA0570, KIAA0729 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
UBP34_MOUSE
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.441285 (rank : 65) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q6ZQ93, Q3UPN0, Q6P563, Q7TMJ6, Q8CCH0 | Gene names | Usp34, Kiaa0570 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
UBP40_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.511225 (rank : 57) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9NVE5, Q6NX38, Q70EL0 | Gene names | USP40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 40 (EC 3.1.2.15) (Ubiquitin thioesterase 40) (Ubiquitin-specific-processing protease 40) (Deubiquitinating enzyme 40). | |||||
|
UBP10_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 150 | |
| SwissProt Accessions | P52479, Q91VY7 | Gene names | Usp10, Ode-1, Uchrp | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
UBP10_HUMAN
|
||||||
| NC score | 0.975313 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q14694, Q9BWG7, Q9NSL7 | Gene names | USP10, KIAA0190 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
UBP50_MOUSE
|
||||||
| NC score | 0.753422 (rank : 3) | θ value | 3.40345e-15 (rank : 7) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q6P8X6, Q9D558, Q9D9E7 | Gene names | Usp50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ubiquitin carboxyl-terminal hydrolase 50 (EC 3.1.2.15) (Ubiquitin thioesterase 50) (Ubiquitin-specific-processing protease 50) (Deubiquitinating enzyme 50). | |||||
|
UBPW_MOUSE
|
||||||
| NC score | 0.752280 (rank : 4) | θ value | 4.91457e-14 (rank : 10) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q61068 | Gene names | Dub1, Dub-1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase DUB-1 (EC 3.1.2.15) (Ubiquitin thioesterase DUB-1) (Ubiquitin-specific-processing protease DUB-1) (Deubiquitinating enzyme 1). | |||||
|
UB17L_HUMAN
|
||||||
| NC score | 0.750550 (rank : 5) | θ value | 8.95645e-16 (rank : 5) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q7RTZ2 | Gene names | USP17L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 17-like protein (EC 3.1.2.15) (Ubiquitin thioesterase 17-like) (Ubiquitin-specific-processing protease 17-like) (Deubiquitinating enzyme 17-like). | |||||
|
UBP8_MOUSE
|
||||||
| NC score | 0.741246 (rank : 6) | θ value | 3.51386e-20 (rank : 4) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q80U87, Q80YP2, Q8R0D3, Q9EQU1, Q9WVP5 | Gene names | Usp8, Kiaa0055, Ubpy | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (mUBPy). | |||||
|
UBP46_HUMAN
|
||||||
| NC score | 0.739653 (rank : 7) | θ value | 4.44505e-15 (rank : 8) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P62068, Q80V95, Q9H7U4, Q9H9T8 | Gene names | USP46 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 46 (EC 3.1.2.15) (Ubiquitin thioesterase 46) (Ubiquitin-specific-processing protease 46) (Deubiquitinating enzyme 46). | |||||
|
UBP46_MOUSE
|
||||||
| NC score | 0.739653 (rank : 8) | θ value | 4.44505e-15 (rank : 9) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P62069, Q80V95, Q9H7U4, Q9H9T8 | Gene names | Usp46 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 46 (EC 3.1.2.15) (Ubiquitin thioesterase 46) (Ubiquitin-specific-processing protease 46) (Deubiquitinating enzyme 46). | |||||
|
UBP12_HUMAN
|
||||||
| NC score | 0.736049 (rank : 9) | θ value | 1.2105e-12 (rank : 15) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O75317 | Gene names | USP12, UBH1, USP12L1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 12 (EC 3.1.2.15) (Ubiquitin thioesterase 12) (Ubiquitin-specific-processing protease 12) (Deubiquitinating enzyme 12) (Ubiquitin-hydrolyzing enzyme 1). | |||||
|
UBP3_MOUSE
|
||||||
| NC score | 0.736033 (rank : 10) | θ value | 3.18553e-13 (rank : 12) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q91W36 | Gene names | Usp3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 3 (EC 3.1.2.15) (Ubiquitin thioesterase 3) (Ubiquitin-specific-processing protease 3) (Deubiquitinating enzyme 3). | |||||
|
UBP50_HUMAN
|
||||||
| NC score | 0.730346 (rank : 11) | θ value | 1.9326e-10 (rank : 22) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q70EL3 | Gene names | USP50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive ubiquitin carboxyl-terminal hydrolase 50 (Inactive ubiquitin- specific peptidase 50). | |||||
|
UBP21_HUMAN
|
||||||
| NC score | 0.729330 (rank : 12) | θ value | 1.58096e-12 (rank : 16) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9UK80, Q9BTV1, Q9NYN4 | Gene names | USP21, USP23 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 21 (EC 3.1.2.15) (Ubiquitin thioesterase 21) (Ubiquitin-specific-processing protease 21) (Deubiquitinating enzyme 21) (NEDD8-specific protease). | |||||
|
UBP3_HUMAN
|
||||||
| NC score | 0.729326 (rank : 13) | θ value | 1.47974e-10 (rank : 20) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9Y6I4 | Gene names | USP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 3 (EC 3.1.2.15) (Ubiquitin thioesterase 3) (Ubiquitin-specific-processing protease 3) (Deubiquitinating enzyme 3). | |||||
|
UBP21_MOUSE
|
||||||
| NC score | 0.726307 (rank : 14) | θ value | 7.84624e-12 (rank : 17) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9QZL6, Q9D0R1 | Gene names | Usp21, Usp23 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 21 (EC 3.1.2.15) (Ubiquitin thioesterase 21) (Ubiquitin-specific-processing protease 21) (Deubiquitinating enzyme 21). | |||||
|
UBP36_HUMAN
|
||||||
| NC score | 0.722768 (rank : 15) | θ value | 2.43908e-13 (rank : 11) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9P275, Q8NDM8, Q9NVC8 | Gene names | USP36, KIAA1453 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 36 (EC 3.1.2.15) (Ubiquitin thioesterase 36) (Ubiquitin-specific-processing protease 36) (Deubiquitinating enzyme 36). | |||||
|
UBP2_HUMAN
|
||||||
| NC score | 0.721643 (rank : 16) | θ value | 5.08577e-11 (rank : 19) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | O75604, Q96MB9, Q9BQ21 | Gene names | USP2, UBP41 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 2 (EC 3.1.2.15) (Ubiquitin thioesterase 2) (Ubiquitin-specific-processing protease 2) (Deubiquitinating enzyme 2) (41 kDa ubiquitin-specific protease). | |||||
|
UBP2_MOUSE
|
||||||
| NC score | 0.716386 (rank : 17) | θ value | 9.59137e-10 (rank : 24) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O88623, Q6X4T9, Q6X4U1, Q8VD74, Q9JJ87 | Gene names | Usp2, Ubp41 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 2 (EC 3.1.2.15) (Ubiquitin thioesterase 2) (Ubiquitin-specific-processing protease 2) (Deubiquitinating enzyme 2) (41 kDa ubiquitin-specific protease). | |||||
|
UBP42_HUMAN
|
||||||
| NC score | 0.716165 (rank : 18) | θ value | 1.16975e-15 (rank : 6) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q9H9J4, Q3C166, Q6P9B4 | Gene names | USP42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 42 (EC 3.1.2.15) (Ubiquitin thioesterase 42) (Ubiquitin-specific-processing protease 42) (Deubiquitinating enzyme 42). | |||||
|
UBP22_MOUSE
|
||||||
| NC score | 0.711265 (rank : 19) | θ value | 9.59137e-10 (rank : 23) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q5DU02, Q5SU81, Q66JV8 | Gene names | Usp22, Kiaa1063 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 22 (EC 3.1.2.15) (Ubiquitin thioesterase 22) (Ubiquitin-specific-processing protease 22) (Deubiquitinating enzyme 22). | |||||
|
UBP8_HUMAN
|
||||||
| NC score | 0.709194 (rank : 20) | θ value | 9.24701e-21 (rank : 3) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
UBP22_HUMAN
|
||||||
| NC score | 0.708998 (rank : 21) | θ value | 2.79066e-09 (rank : 27) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9UPT9 | Gene names | USP22, KIAA1063, USP3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 22 (EC 3.1.2.15) (Ubiquitin thioesterase 22) (Ubiquitin-specific-processing protease 22) (Deubiquitinating enzyme 22). | |||||
|
UBP51_HUMAN
|
||||||
| NC score | 0.699301 (rank : 22) | θ value | 3.41135e-07 (rank : 37) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q70EK9, Q8IWJ8 | Gene names | USP51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 51 (EC 3.1.2.15) (Ubiquitin thioesterase 51) (Ubiquitin-specific-processing protease 51) (Deubiquitinating enzyme 51). | |||||
|
UBP44_HUMAN
|
||||||
| NC score | 0.690967 (rank : 23) | θ value | 1.47974e-10 (rank : 21) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9H0E7 | Gene names | USP44 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 44 (EC 3.1.2.15) (Ubiquitin thioesterase 44) (Ubiquitin-specific-processing protease 44) (Deubiquitinating enzyme 44). | |||||
|
UBP49_HUMAN
|
||||||
| NC score | 0.675131 (rank : 24) | θ value | 3.64472e-09 (rank : 29) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q70CQ1, Q96CK4 | Gene names | USP49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 49 (EC 3.1.2.15) (Ubiquitin thioesterase 49) (Ubiquitin-specific-processing protease 49) (Deubiquitinating enzyme 49). | |||||
|
UBP49_MOUSE
|
||||||
| NC score | 0.674604 (rank : 25) | θ value | 2.79066e-09 (rank : 28) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q6P9L4 | Gene names | Usp49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 49 (EC 3.1.2.15) (Ubiquitin thioesterase 49) (Ubiquitin-specific-processing protease 49) (Deubiquitinating enzyme 49). | |||||
|
UBP11_HUMAN
|
||||||
| NC score | 0.671024 (rank : 26) | θ value | 1.02475e-11 (rank : 18) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P51784, Q8IUG6, Q9BWE1 | Gene names | USP11, UHX1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 11 (EC 3.1.2.15) (Ubiquitin thioesterase 11) (Ubiquitin-specific-processing protease 11) (Deubiquitinating enzyme 11). | |||||
|
UBP15_MOUSE
|
||||||
| NC score | 0.669330 (rank : 27) | θ value | 5.43371e-13 (rank : 14) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8R5H1, Q3UL25 | Gene names | Usp15 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 15 (EC 3.1.2.15) (Ubiquitin thioesterase 15) (Ubiquitin-specific-processing protease 15) (Deubiquitinating enzyme 15). | |||||
|
UBP33_HUMAN
|
||||||
| NC score | 0.669310 (rank : 28) | θ value | 6.87365e-08 (rank : 35) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8TEY7, Q8TEY6, Q96AV6, Q9H9F0, Q9UPQ5 | Gene names | USP33, KIAA1097, VDU1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 33 (EC 3.1.2.15) (Ubiquitin thioesterase 33) (Ubiquitin-specific-processing protease 33) (Deubiquitinating enzyme 33) (VHL-interacting deubiquitinating enzyme 1). | |||||
|
UBP15_HUMAN
|
||||||
| NC score | 0.668979 (rank : 29) | θ value | 5.43371e-13 (rank : 13) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9Y4E8, Q9HCA6, Q9UNP0, Q9Y5B5 | Gene names | USP15, KIAA0529 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 15 (EC 3.1.2.15) (Ubiquitin thioesterase 15) (Ubiquitin-specific-processing protease 15) (Deubiquitinating enzyme 15) (Unph-2) (Unph4). | |||||
|
UBP16_HUMAN
|
||||||
| NC score | 0.663391 (rank : 30) | θ value | 1.43324e-05 (rank : 42) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9Y5T5, Q8NEL3 | Gene names | USP16 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 16 (EC 3.1.2.15) (Ubiquitin thioesterase 16) (Ubiquitin-specific-processing protease 16) (Deubiquitinating enzyme 16) (Ubiquitin-processing protease UBP-M). | |||||
|
UBP20_HUMAN
|
||||||
| NC score | 0.661555 (rank : 31) | θ value | 6.43352e-06 (rank : 39) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9Y2K6, Q96LG5, Q9UQN8, Q9UQP0 | Gene names | USP20, KIAA1003, LSFR3A | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 20 (EC 3.1.2.15) (Ubiquitin thioesterase 20) (Ubiquitin-specific-processing protease 20) (Deubiquitinating enzyme 20). | |||||
|
UBP48_HUMAN
|
||||||
| NC score | 0.647255 (rank : 32) | θ value | 1.63604e-09 (rank : 25) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q86UV5, Q2M3I4, Q5SZI4, Q5T3T5, Q6NX53, Q8N3F6, Q96F64, Q96IQ3, Q9H5N3, Q9H5T7, Q9NUJ6, Q9NXR0 | Gene names | USP48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 48 (EC 3.1.2.15) (Ubiquitin thioesterase 48) (Ubiquitin-specific-processing protease 48) (Deubiquitinating enzyme 48). | |||||
|
UBP48_MOUSE
|
||||||
| NC score | 0.645152 (rank : 33) | θ value | 1.63604e-09 (rank : 26) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q3V0C5, Q3TM39, Q571J9, Q8VDJ1 | Gene names | Usp48, Kiaa4202 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 48 (EC 3.1.2.15) (Ubiquitin thioesterase 48) (Ubiquitin-specific-processing protease 48) (Deubiquitinating enzyme 48). | |||||
|
UBP4_HUMAN
|
||||||
| NC score | 0.645034 (rank : 34) | θ value | 5.26297e-08 (rank : 33) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q13107, O43452, O43453 | Gene names | USP4, UNP, UNPH | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 4 (EC 3.1.2.15) (Ubiquitin thioesterase 4) (Ubiquitin-specific-processing protease 4) (Deubiquitinating enzyme 4) (Ubiquitous nuclear protein homolog). | |||||
|
UBP4_MOUSE
|
||||||
| NC score | 0.643611 (rank : 35) | θ value | 5.26297e-08 (rank : 34) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P35123, O54704 | Gene names | Usp4, Unp | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 4 (EC 3.1.2.15) (Ubiquitin thioesterase 4) (Ubiquitin-specific-processing protease 4) (Deubiquitinating enzyme 4) (Ubiquitous nuclear protein). | |||||
|
UBP43_MOUSE
|
||||||
| NC score | 0.618843 (rank : 36) | θ value | 1.09739e-05 (rank : 41) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q8BUM9, Q8VDP5 | Gene names | Usp43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 43 (EC 3.1.2.15) (Ubiquitin thioesterase 43) (Ubiquitin-specific-processing protease 43) (Deubiquitinating enzyme 43). | |||||
|
UBP18_HUMAN
|
||||||
| NC score | 0.615357 (rank : 37) | θ value | 3.19293e-05 (rank : 45) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9UMW8, Q9NY71 | Gene names | USP18, ISG43 | |||
|
Domain Architecture |

|
|||||
| Description | Ubl carboxyl-terminal hydrolase 18 (EC 3.1.2.-) (Ubl thioesterase 18) (ISG15-specific-processing protease) (43 kDa ISG15-specific protease) (hUBP43). | |||||
|
UBP43_HUMAN
|
||||||
| NC score | 0.615127 (rank : 38) | θ value | 1.43324e-05 (rank : 43) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q70EL4, Q8N2C5, Q96DQ6 | Gene names | USP43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 43 (EC 3.1.2.15) (Ubiquitin thioesterase 43) (Ubiquitin-specific-processing protease 43) (Deubiquitinating enzyme 43). | |||||
|
UBP1_HUMAN
|
||||||
| NC score | 0.611951 (rank : 39) | θ value | 3.19293e-05 (rank : 46) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | O94782, Q9UFR0, Q9UNJ3 | Gene names | USP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 1 (EC 3.1.2.15) (Ubiquitin thioesterase 1) (Ubiquitin-specific-processing protease 1) (Deubiquitinating enzyme 1) (hUBP). | |||||
|
UBP31_HUMAN
|
||||||
| NC score | 0.607191 (rank : 40) | θ value | 0.00020696 (rank : 49) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
UBP14_HUMAN
|
||||||
| NC score | 0.596021 (rank : 41) | θ value | 1.38499e-08 (rank : 30) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P54578 | Gene names | USP14, TGT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 14 (EC 3.1.2.15) (Ubiquitin thioesterase 14) (Ubiquitin-specific-processing protease 14) (Deubiquitinating enzyme 14). | |||||
|
UBP14_MOUSE
|
||||||
| NC score | 0.592282 (rank : 42) | θ value | 1.38499e-08 (rank : 31) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9JMA1, Q543U5, Q923F2, Q9D0L0 | Gene names | Usp14 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 14 (EC 3.1.2.15) (Ubiquitin thioesterase 14) (Ubiquitin-specific-processing protease 14) (Deubiquitinating enzyme 14). | |||||
|
UBP7_HUMAN
|
||||||
| NC score | 0.587265 (rank : 43) | θ value | 1.43324e-05 (rank : 44) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q93009 | Gene names | USP7, HAUSP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 7 (EC 3.1.2.15) (Ubiquitin thioesterase 7) (Ubiquitin-specific-processing protease 7) (Deubiquitinating enzyme 7) (Herpesvirus-associated ubiquitin-specific protease). | |||||
|
UBP24_HUMAN
|
||||||
| NC score | 0.567658 (rank : 44) | θ value | 0.000602161 (rank : 50) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9UPU5, Q6ZSY2, Q8N2Y4, Q9NXD1 | Gene names | USP24, KIAA1057 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 24 (EC 3.1.2.15) (Ubiquitin thioesterase 24) (Ubiquitin-specific-processing protease 24) (Deubiquitinating enzyme 24). | |||||
|
UBP30_HUMAN
|
||||||
| NC score | 0.566845 (rank : 45) | θ value | 0.0193708 (rank : 60) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q70CQ3, Q8WTU7, Q96JX4, Q9BSS3 | Gene names | USP30 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 30 (EC 3.1.2.15) (Ubiquitin thioesterase 30) (Ubiquitin-specific-processing protease 30) (Deubiquitinating enzyme 30). | |||||
|
SNUT2_MOUSE
|
||||||
| NC score | 0.565106 (rank : 46) | θ value | 4.1701e-05 (rank : 47) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q3TIX9, Q3TI55, Q8BLI5, Q8BZZ5, Q9CQR1 | Gene names | Usp39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 2 (Inactive ubiquitin-specific peptidase 39). | |||||
|
UBP37_HUMAN
|
||||||
| NC score | 0.550982 (rank : 47) | θ value | 1.09739e-05 (rank : 40) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q86T82, Q9HCH8 | Gene names | USP37, KIAA1594 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 37 (EC 3.1.2.15) (Ubiquitin thioesterase 37) (Ubiquitin-specific-processing protease 37) (Deubiquitinating enzyme 37). | |||||
|
SNUT2_HUMAN
|
||||||
| NC score | 0.548050 (rank : 48) | θ value | 0.000158464 (rank : 48) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q53GS9, Q6NX47, Q96RK9, Q9BV89, Q9H381, Q9P050, Q9Y310 | Gene names | USP39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 2 (U4/U6.U5 tri-snRNP-associated 65 kDa protein) (65K) (Inactive ubiquitin-specific peptidase 39) (SAD1 homolog). | |||||
|
UBP18_MOUSE
|
||||||
| NC score | 0.543926 (rank : 49) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9WTV6 | Gene names | Usp18, Ubp43 | |||
|
Domain Architecture |

|
|||||
| Description | Ubl carboxyl-terminal hydrolase 18 (EC 3.1.2.-) (Ubl thioesterase 18) (ISG15-specific-processing protease) (43 kDa ISG15-specific protease). | |||||
|
UBP47_MOUSE
|
||||||
| NC score | 0.542144 (rank : 50) | θ value | 0.00102713 (rank : 53) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q8BY87, Q5EBP2, Q6KAR9, Q80V06, Q8BHU1, Q8BI15, Q8BI16, Q8BUW4, Q91X25 | Gene names | Usp47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
USP9X_MOUSE
|
||||||
| NC score | 0.538301 (rank : 51) | θ value | 1.80886e-08 (rank : 32) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
UBP47_HUMAN
|
||||||
| NC score | 0.538127 (rank : 52) | θ value | 0.00390308 (rank : 55) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q96K76, Q658U0, Q86Y73, Q8TEP6, Q9BWI0, Q9NWN1 | Gene names | USP47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
USP9X_HUMAN
|
||||||
| NC score | 0.535439 (rank : 53) | θ value | 8.97725e-08 (rank : 36) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
USP9Y_HUMAN
|
||||||
| NC score | 0.530778 (rank : 54) | θ value | 1.69304e-06 (rank : 38) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O00507, O14601 | Gene names | USP9Y, DFFRY, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-Y (EC 3.1.2.15) (Ubiquitin thioesterase FAF-Y) (Ubiquitin-specific-processing protease FAF-Y) (Deubiquitinating enzyme FAF-Y) (Fat facets protein-related, Y- linked) (Ubiquitin-specific protease 9, Y chromosome). | |||||
|
UBP35_HUMAN
|
||||||
| NC score | 0.524328 (rank : 55) | θ value | 0.0148317 (rank : 58) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9P2H5 | Gene names | USP35, KIAA1372, USP34 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 35 (EC 3.1.2.15) (Ubiquitin thioesterase 35) (Ubiquitin-specific-processing protease 35) (Deubiquitinating enzyme 35). | |||||
|
UBP32_HUMAN
|
||||||
| NC score | 0.523349 (rank : 56) | θ value | 0.0563607 (rank : 64) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8NFA0, Q9BX85, Q9Y591 | Gene names | USP32, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 32 (EC 3.1.2.15) (Ubiquitin thioesterase 32) (Ubiquitin-specific-processing protease 32) (Deubiquitinating enzyme 32) (NY-REN-60 antigen). | |||||
|
UBP40_HUMAN
|
||||||
| NC score | 0.511225 (rank : 57) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9NVE5, Q6NX38, Q70EL0 | Gene names | USP40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 40 (EC 3.1.2.15) (Ubiquitin thioesterase 40) (Ubiquitin-specific-processing protease 40) (Deubiquitinating enzyme 40). | |||||
|
UBP6_HUMAN
|
||||||
| NC score | 0.496556 (rank : 58) | θ value | 0.0736092 (rank : 68) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P35125, Q15634, Q86WP6, Q8IWT4 | Gene names | USP6, TRE2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 6 (EC 3.1.2.15) (Ubiquitin thioesterase 6) (Ubiquitin-specific-processing protease 6) (Deubiquitinating enzyme 6) (Proto-oncogene TRE-2). | |||||
|
UBP29_HUMAN
|
||||||
| NC score | 0.493668 (rank : 59) | θ value | 0.00390308 (rank : 54) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9HBJ7 | Gene names | USP29 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 29 (EC 3.1.2.15) (Ubiquitin thioesterase 29) (Ubiquitin-specific-processing protease 29) (Deubiquitinating enzyme 29). | |||||
|
UBP38_HUMAN
|
||||||
| NC score | 0.492363 (rank : 60) | θ value | 0.0330416 (rank : 62) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8NB14, Q8NDF5, Q96DK6, Q96PZ6, Q9BY55 | Gene names | USP38, KIAA1891 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 38 (EC 3.1.2.15) (Ubiquitin thioesterase 38) (Ubiquitin-specific-processing protease 38) (Deubiquitinating enzyme 38) (HP43.8KD). | |||||
|
UBP26_HUMAN
|
||||||
| NC score | 0.484729 (rank : 61) | θ value | 0.00102713 (rank : 52) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9BXU7 | Gene names | USP26 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 26 (EC 3.1.2.15) (Ubiquitin thioesterase 26) (Ubiquitin-specific-processing protease 26) (Deubiquitinating enzyme 26). | |||||
|
UBP38_MOUSE
|
||||||
| NC score | 0.477954 (rank : 62) | θ value | 0.62314 (rank : 83) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8BW70, Q8BWL1, Q8BX03 | Gene names | Usp38 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 38 (EC 3.1.2.15) (Ubiquitin thioesterase 38) (Ubiquitin-specific-processing protease 38) (Deubiquitinating enzyme 38). | |||||
|
UBP29_MOUSE
|
||||||
| NC score | 0.470823 (rank : 63) | θ value | 0.0113563 (rank : 56) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9ES63 | Gene names | Usp29 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 29 (EC 3.1.2.15) (Ubiquitin thioesterase 29) (Ubiquitin-specific-processing protease 29) (Deubiquitinating enzyme 29). | |||||
|
UBP34_HUMAN
|
||||||
| NC score | 0.441461 (rank : 64) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q70CQ2, O60316, O94834, Q3B777, Q6P6C9, Q7L8P6, Q8N3T9, Q8TBW2, Q9UGA1 | Gene names | USP34, KIAA0570, KIAA0729 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
UBP34_MOUSE
|
||||||
| NC score | 0.441285 (rank : 65) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q6ZQ93, Q3UPN0, Q6P563, Q7TMJ6, Q8CCH0 | Gene names | Usp34, Kiaa0570 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
UBP26_MOUSE
|
||||||
| NC score | 0.436234 (rank : 66) | θ value | 0.000786445 (rank : 51) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q99MX1 | Gene names | Usp26 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 26 (EC 3.1.2.15) (Ubiquitin thioesterase 26) (Ubiquitin-specific-processing protease 26) (Deubiquitinating enzyme 26). | |||||
|
UBP5_HUMAN
|
||||||
| NC score | 0.423973 (rank : 67) | θ value | 0.0252991 (rank : 61) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P45974, Q96J22 | Gene names | USP5, ISOT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP5_MOUSE
|
||||||
| NC score | 0.415108 (rank : 68) | θ value | 0.0736092 (rank : 67) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P56399 | Gene names | Usp5, Isot | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP13_HUMAN
|
||||||
| NC score | 0.406520 (rank : 69) | θ value | 0.0563607 (rank : 63) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q92995 | Gene names | USP13, ISOT3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 13 (EC 3.1.2.15) (Ubiquitin thioesterase 13) (Ubiquitin-specific-processing protease 13) (Deubiquitinating enzyme 13) (Isopeptidase T-3) (ISOT-3). | |||||
|
UBP25_HUMAN
|
||||||
| NC score | 0.297859 (rank : 70) | θ value | 0.0736092 (rank : 66) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9UHP3, Q9H9W1 | Gene names | USP25, USP21 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (USP on chromosome 21). | |||||
|
UBP28_HUMAN
|
||||||
| NC score | 0.296849 (rank : 71) | θ value | 2.36792 (rank : 104) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q96RU2, Q9P213 | Gene names | USP28, KIAA1515 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
UBP28_MOUSE
|
||||||
| NC score | 0.292529 (rank : 72) | θ value | 3.0926 (rank : 115) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q5I043, Q6NZP3, Q6ZPP1, Q8BWI1 | Gene names | Usp28, Kiaa1515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
UBP25_MOUSE
|
||||||
| NC score | 0.288665 (rank : 73) | θ value | 1.38821 (rank : 94) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P57080, Q80ZT9 | Gene names | Usp25 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (mUSP25). | |||||
|
BRAP_MOUSE
|
||||||
| NC score | 0.086237 (rank : 74) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q99MP8, Q8CC00, Q8CHX1, Q99MP7, Q9CXX8 | Gene names | Brap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP). | |||||
|
GGNB1_MOUSE
|
||||||
| NC score | 0.078273 (rank : 75) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6K1E7, Q8C5T9 | Gene names | Ggnbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
BRAP_HUMAN
|
||||||
| NC score | 0.074548 (rank : 76) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7Z569, O43238, O75341 | Gene names | BRAP, RNF52 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP) (RING finger protein 52) (NY-REN-63 antigen). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.068600 (rank : 77) | θ value | 0.0148317 (rank : 57) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
GGNB1_HUMAN
|
||||||
| NC score | 0.062788 (rank : 78) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5YKI7, Q5YKI8 | Gene names | GGNBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
HDAC6_MOUSE
|
||||||
| NC score | 0.060666 (rank : 79) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z2V5 | Gene names | Hdac6 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 6 (HD6) (Histone deacetylase mHDA2). | |||||
|
HDAC6_HUMAN
|
||||||
| NC score | 0.058105 (rank : 80) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UBN7 | Gene names | HDAC6 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 6 (HD6). | |||||
|
AP180_HUMAN
|
||||||
| NC score | 0.055674 (rank : 81) | θ value | 0.21417 (rank : 72) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60641, Q5JX13, Q68DL9, Q6P9D3, Q9NTY7 | Gene names | SNAP91, KIAA0656 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.050824 (rank : 82) | θ value | 3.0926 (rank : 106) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
AP180_MOUSE
|
||||||
| NC score | 0.048232 (rank : 83) | θ value | 1.06291 (rank : 87) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61548, Q61547 | Gene names | Snap91 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein) (Phosphoprotein F1- 20). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.047925 (rank : 84) | θ value | 0.125558 (rank : 69) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.046320 (rank : 85) | θ value | 0.813845 (rank : 86) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
RD23B_HUMAN
|
||||||
| NC score | 0.045129 (rank : 86) | θ value | 0.21417 (rank : 73) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P54727, Q8WUB0 | Gene names | RAD23B | |||
|
Domain Architecture |

|
|||||
| Description | UV excision repair protein RAD23 homolog B (hHR23B) (XP-C repair- complementing complex 58 kDa protein) (p58). | |||||
|
NFRKB_MOUSE
|
||||||
| NC score | 0.042238 (rank : 87) | θ value | 0.47712 (rank : 80) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
OKL38_HUMAN
|
||||||
| NC score | 0.040523 (rank : 88) | θ value | 0.365318 (rank : 75) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJX0, Q86UQ1, Q96S88, Q9BZ70 | Gene names | OKL38 | |||
|
Domain Architecture |

|
|||||
| Description | Pregnancy-induced growth inhibitor OKL38 (huOKL38) (Ovary, kidney and liver protein 38). | |||||
|
NU214_HUMAN
|
||||||
| NC score | 0.039742 (rank : 89) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.037748 (rank : 90) | θ value | 0.813845 (rank : 84) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
NFRKB_HUMAN
|
||||||
| NC score | 0.036774 (rank : 91) | θ value | 4.03905 (rank : 122) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6P4R8, Q12869, Q15312, Q9H048 | Gene names | NFRKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.036514 (rank : 92) | θ value | 0.47712 (rank : 79) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
CJ030_MOUSE
|
||||||
| NC score | 0.036353 (rank : 93) | θ value | 0.47712 (rank : 76) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BSV3, Q4VAD9, Q6PHN3, Q8BPJ4, Q8C973 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C10orf30 homolog. | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.035488 (rank : 94) | θ value | 0.0736092 (rank : 65) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
MAP4_MOUSE
|
||||||
| NC score | 0.034179 (rank : 95) | θ value | 1.81305 (rank : 97) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.033277 (rank : 96) | θ value | 8.99809 (rank : 148) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
PKHA6_HUMAN
|
||||||
| NC score | 0.031409 (rank : 97) | θ value | 1.06291 (rank : 89) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.030200 (rank : 98) | θ value | 6.88961 (rank : 134) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
X3CL1_HUMAN
|
||||||
| NC score | 0.029056 (rank : 99) | θ value | 1.06291 (rank : 90) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P78423, O00672 | Gene names | CX3CL1, A-152E5.2, FKN, NTT, SCYD1 | |||
|
Domain Architecture |

|
|||||
| Description | Fractalkine precursor (CX3CL1) (Neurotactin) (CX3C membrane-anchored chemokine) (Small inducible cytokine D1). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.028678 (rank : 100) | θ value | 1.38821 (rank : 93) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
K0460_MOUSE
|
||||||
| NC score | 0.027624 (rank : 101) | θ value | 1.81305 (rank : 96) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6NXI6, Q3U3L8, Q6ZQA7 | Gene names | Kiaa0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
RBM19_MOUSE
|
||||||
| NC score | 0.027602 (rank : 102) | θ value | 0.0193708 (rank : 59) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R3C6, Q8BHR0, Q9CW63 | Gene names | Rbm19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 19 (RNA-binding motif protein 19). | |||||
|
PKHA6_MOUSE
|
||||||
| NC score | 0.026924 (rank : 103) | θ value | 4.03905 (rank : 123) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.026588 (rank : 104) | θ value | 3.0926 (rank : 107) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.026473 (rank : 105) | θ value | 3.0926 (rank : 108) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
THAP8_HUMAN
|
||||||
| NC score | 0.026216 (rank : 106) | θ value | 2.36792 (rank : 103) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NA92, Q96M21 | Gene names | THAP8 | |||
|
Domain Architecture |

|
|||||
| Description | THAP domain-containing protein 8. | |||||
|
RERE_MOUSE
|
||||||
| NC score | 0.026176 (rank : 107) | θ value | 3.0926 (rank : 114) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
CIZ1_HUMAN
|
||||||
| NC score | 0.026030 (rank : 108) | θ value | 3.0926 (rank : 110) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9ULV3, Q9NYM8, Q9UHK4, Q9Y3F9, Q9Y3G0 | Gene names | CIZ1, LSFR1, NP94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cip1-interacting zinc finger protein (Nuclear protein NP94). | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.024727 (rank : 109) | θ value | 3.0926 (rank : 113) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
EVL_MOUSE
|
||||||
| NC score | 0.023208 (rank : 110) | θ value | 5.27518 (rank : 127) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P70429, Q9ERU8 | Gene names | Evl | |||
|
Domain Architecture |

|
|||||
| Description | Ena/VASP-like protein (Ena/vasodilator-stimulated phosphoprotein- like). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.022684 (rank : 111) | θ value | 4.03905 (rank : 126) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.022531 (rank : 112) | θ value | 4.03905 (rank : 120) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.022152 (rank : 113) | θ value | 0.125558 (rank : 70) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
HCN1_HUMAN
|
||||||
| NC score | 0.021725 (rank : 114) | θ value | 0.279714 (rank : 74) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60741 | Gene names | HCN1, BCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1). | |||||
|
FMN2_HUMAN
|
||||||
| NC score | 0.021701 (rank : 115) | θ value | 1.06291 (rank : 88) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
INT1_MOUSE
|
||||||
| NC score | 0.021522 (rank : 116) | θ value | 2.36792 (rank : 101) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6P4S8, Q80UQ7, Q91Z01, Q9CTF7 | Gene names | Ints1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 1 (Int1). | |||||
|
HSP74_MOUSE
|
||||||
| NC score | 0.021122 (rank : 117) | θ value | 0.163984 (rank : 71) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61316 | Gene names | Hspa4, Hsp110 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4 (Heat shock 70-related protein APG-2). | |||||
|
HCN1_MOUSE
|
||||||
| NC score | 0.020935 (rank : 118) | θ value | 0.47712 (rank : 77) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O88704, O54899, Q9D613 | Gene names | Hcn1, Bcng1, Hac2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 1 (Brain cyclic nucleotide-gated channel 1) (BCNG-1) (Hyperpolarization-activated cation channel 2) (HAC-2). | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.020021 (rank : 119) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
TIAM1_MOUSE
|
||||||
| NC score | 0.018540 (rank : 120) | θ value | 0.47712 (rank : 82) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60610 | Gene names | Tiam1, Tiam-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
ASXL1_MOUSE
|
||||||
| NC score | 0.018339 (rank : 121) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P59598 | Gene names | Asxl1, Kiaa0978 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative Polycomb group protein ASXL1 (Additional sex combs-like protein 1). | |||||
|
HSP74_HUMAN
|
||||||
| NC score | 0.016758 (rank : 122) | θ value | 2.36792 (rank : 100) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P34932, O95756 | Gene names | HSPA4 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4 (Heat shock 70-related protein APG-2) (HSP70RY). | |||||
|
RANB3_HUMAN
|
||||||
| NC score | 0.016573 (rank : 123) | θ value | 8.99809 (rank : 147) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9H6Z4, O60405, O75759, O75760, Q9BT47, Q9UG74 | Gene names | RANBP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 3 (RanBP3). | |||||
|
GPR98_HUMAN
|
||||||
| NC score | 0.016156 (rank : 124) | θ value | 1.38821 (rank : 91) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WXG9, O75171, Q9H0X5, Q9UL61 | Gene names | GPR98, KIAA0686, MASS1, VLGR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor 98 precursor (Monogenic audiogenic seizure susceptibility protein 1 homolog) (Very large G-protein coupled receptor 1) (Usher syndrome type-2C protein). | |||||
|
ANC5_HUMAN
|
||||||
| NC score | 0.016079 (rank : 125) | θ value | 4.03905 (rank : 118) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJX4, Q8N4H7, Q9BQD4 | Gene names | ANAPC5, APC5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Anaphase-promoting complex subunit 5 (APC5) (Cyclosome subunit 5). | |||||
|
OXA1L_HUMAN
|
||||||
| NC score | 0.015437 (rank : 126) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15070 | Gene names | OXA1L | |||
|
Domain Architecture |

|
|||||
| Description | Inner membrane protein OXA1L, mitochondrial precursor (Oxidase assembly 1-like protein) (OXA1-like protein) (OXA1Hs) (Hsa). | |||||
|
LDB3_MOUSE
|
||||||
| NC score | 0.015162 (rank : 127) | θ value | 0.47712 (rank : 78) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9JKS4, Q6A038, Q811P2, Q811P3, Q811P4, Q811P5, Q9D130, Q9JKS3, Q9R0Z1, Q9WVH1, Q9WVH2 | Gene names | Ldb3, Kiaa0613 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher) (Protein oracle). | |||||
|
PK3C3_MOUSE
|
||||||
| NC score | 0.014134 (rank : 128) | θ value | 2.36792 (rank : 102) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PF93, Q8R3S8 | Gene names | Pik3c3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3). | |||||
|
SYNJ1_MOUSE
|
||||||
| NC score | 0.013986 (rank : 129) | θ value | 6.88961 (rank : 135) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
PO2F1_MOUSE
|
||||||
| NC score | 0.013480 (rank : 130) | θ value | 0.813845 (rank : 85) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P25425, Q61994, Q63891, Q6Y681, Q7TSD0, Q8BT04, Q8K570, Q99JH0, Q9WTZ4, Q9WTZ5, Q9WTZ6 | Gene names | Pou2f1, Oct-1, Otf-1, Otf1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 2, transcription factor 1 (Octamer-binding transcription factor 1) (Oct-1) (OTF-1) (NF-A1). | |||||
|
TBX6_HUMAN
|
||||||
| NC score | 0.012917 (rank : 131) | θ value | 0.47712 (rank : 81) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95947, Q8TAS4 | Gene names | TBX6 | |||
|
Domain Architecture |

|
|||||
| Description | T-box transcription factor TBX6 (T-box protein 6). | |||||
|
IL4RA_MOUSE
|
||||||
| NC score | 0.012786 (rank : 132) | θ value | 6.88961 (rank : 132) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P16382, O54690, Q60583, Q8CBW5 | Gene names | Il4r, Il4ra | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-4 receptor alpha chain precursor (IL-4R-alpha) (CD124 antigen) [Contains: Soluble interleukin-4 receptor alpha chain (IL-4- binding protein) (IL4-BP)]. | |||||
|
TBCD4_MOUSE
|
||||||
| NC score | 0.011908 (rank : 133) | θ value | 8.99809 (rank : 149) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BYJ6, Q5DU23, Q66JU2, Q6P2M2, Q8BMH6, Q8BXM2 | Gene names | Tbc1d4, Kiaa0603 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 4 (Akt substrate of 160 kDa) (AS160). | |||||
|
SAPS2_HUMAN
|
||||||
| NC score | 0.011259 (rank : 134) | θ value | 4.03905 (rank : 124) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75170, Q9UGB9 | Gene names | SAPS2, KIAA0685 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 2. | |||||
|
PK3C3_HUMAN
|
||||||
| NC score | 0.010843 (rank : 135) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NEB9, Q15134 | Gene names | PIK3C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3) (Phosphatidylinositol 3-kinase p100 subunit). | |||||
|
TENS1_HUMAN
|
||||||
| NC score | 0.010044 (rank : 136) | θ value | 8.99809 (rank : 150) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9HBL0 | Gene names | TNS1, TNS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-1. | |||||
|
GNAS1_MOUSE
|
||||||
| NC score | 0.009601 (rank : 137) | θ value | 3.0926 (rank : 111) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6R0H7, Q6R0H4, Q6R0H5, Q6R2J5, Q9JJX0, Q9Z1N8 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
AMOT_HUMAN
|
||||||
| NC score | 0.009524 (rank : 138) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
DDX24_HUMAN
|
||||||
| NC score | 0.009318 (rank : 139) | θ value | 1.81305 (rank : 95) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9GZR7 | Gene names | DDX24 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX24 (EC 3.6.1.-) (DEAD box protein 24). | |||||
|
FGD5_MOUSE
|
||||||
| NC score | 0.009173 (rank : 140) | θ value | 5.27518 (rank : 128) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80UZ0, Q8BHM5 | Gene names | Fgd5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5. | |||||
|
NCA11_MOUSE
|
||||||
| NC score | 0.008723 (rank : 141) | θ value | 4.03905 (rank : 121) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P13595, Q61949 | Gene names | Ncam1, Ncam | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule 1, 180 kDa isoform precursor (N-CAM 180) (NCAM-180). | |||||
|
DDX51_HUMAN
|
||||||
| NC score | 0.007891 (rank : 142) | θ value | 6.88961 (rank : 131) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
LAP4_MOUSE
|
||||||
| NC score | 0.007786 (rank : 143) | θ value | 1.38821 (rank : 92) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80U72, Q6P5H7, Q7TPH8, Q80VB1, Q8CI48, Q8VII1, Q922S3 | Gene names | Scrib, Kiaa0147, Lap4, Scrib1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog). | |||||
|
RUFY2_HUMAN
|
||||||
| NC score | 0.007755 (rank : 144) | θ value | 1.81305 (rank : 98) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
CENPB_HUMAN
|
||||||
| NC score | 0.007742 (rank : 145) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P07199 | Gene names | CENPB | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
LAP4_HUMAN
|
||||||
| NC score | 0.007232 (rank : 146) | θ value | 4.03905 (rank : 119) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 608 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14160, Q6P496, Q7Z5D1, Q8WWV8, Q96C69, Q96GG1 | Gene names | SCRIB, CRIB1, KIAA0147, LAP4, SCRB1, VARTUL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog) (hScrib). | |||||
|
RUFY2_MOUSE
|
||||||
| NC score | 0.007208 (rank : 147) | θ value | 5.27518 (rank : 129) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8R4C2, Q69ZH1 | Gene names | Rufy2, Kiaa1537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Leucine zipper FYVE-finger protein) (LZ-FYVE). | |||||
|
ELK4_HUMAN
|
||||||
| NC score | 0.006305 (rank : 148) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P28324, P28323 | Gene names | ELK4, SAP1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
CUL4B_HUMAN
|
||||||
| NC score | 0.006017 (rank : 149) | θ value | 6.88961 (rank : 130) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13620, Q6PIE4, Q6UP07, Q7Z673, Q9BY37, Q9UEB7, Q9UED7 | Gene names | CUL4B, KIAA0695 | |||
|
Domain Architecture |

|
|||||
| Description | Cullin-4B (CUL-4B). | |||||
|
BICD2_MOUSE
|
||||||
| NC score | 0.005661 (rank : 150) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q921C5, Q80TU1, Q8BTE3, Q9DCL3 | Gene names | Bicd2, Kiaa0699 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
BICD2_HUMAN
|
||||||
| NC score | 0.005206 (rank : 151) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TD16, O75181, Q5TBQ2, Q5TBQ3, Q96LH2, Q9BT84, Q9H561 | Gene names | BICD2, KIAA0699 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
SLIK6_MOUSE
|
||||||
| NC score | 0.004746 (rank : 152) | θ value | 1.81305 (rank : 99) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C110, Q8BLL0 | Gene names | Slitrk6, Sltk6 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT and NTRK-like protein 6 precursor. | |||||
|
SREC_HUMAN
|
||||||
| NC score | 0.003138 (rank : 153) | θ value | 4.03905 (rank : 125) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
ZN592_MOUSE
|
||||||
| NC score | 0.002677 (rank : 154) | θ value | 3.0926 (rank : 117) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BHZ4, Q80XM1 | Gene names | Znf592, Kiaa0211, Zfp592 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592 (Zfp-592). | |||||
|
ZN592_HUMAN
|
||||||
| NC score | 0.001549 (rank : 155) | θ value | 6.88961 (rank : 136) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
LATS1_MOUSE
|
||||||
| NC score | -0.000148 (rank : 156) | θ value | 3.0926 (rank : 112) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1027 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BYR2, Q9Z0W4 | Gene names | Lats1, Warts | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase). | |||||
|
CD2L2_HUMAN
|
||||||
| NC score | -0.001840 (rank : 157) | θ value | 3.0926 (rank : 109) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1439 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UQ88, O95227, O95228, O96012, Q12821, Q12853, Q12854, Q5QPR0, Q5QPR1, Q5QPR2, Q9UBC4, Q9UBI3, Q9UEI1, Q9UEI2, Q9UP53, Q9UP54, Q9UP55, Q9UP56, Q9UQ86, Q9UQ87, Q9UQ89 | Gene names | CDC2L2, PITSLREB | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L2 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 2) (CDK11). | |||||
|
ABL1_MOUSE
|
||||||
| NC score | -0.002546 (rank : 158) | θ value | 3.0926 (rank : 105) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P00520, P97896, Q61252, Q61253, Q61254, Q61255, Q61256, Q61257, Q61258, Q61259, Q61260, Q61261, Q6PCM5, Q8C1X4 | Gene names | Abl1, Abl | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene tyrosine-protein kinase ABL1 (EC 2.7.10.2) (p150) (c- ABL) (Abelson murine leukemia viral oncogene homolog 1). | |||||
|
ZN335_HUMAN
|
||||||
| NC score | -0.002669 (rank : 159) | θ value | 3.0926 (rank : 116) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H4Z2, Q9H684 | Gene names | ZNF335 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 335. | |||||
|
KCC2B_HUMAN
|
||||||
| NC score | -0.004346 (rank : 160) | θ value | 6.88961 (rank : 133) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 866 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13554, O95437, O95438, O95599, Q9UGH7, Q9UGH8, Q9UGH9, Q9UNX0, Q9UNX7, Q9UP00, Q9Y5N4, Q9Y6F4 | Gene names | CAMK2B, CAM2, CAMKB | |||
|
Domain Architecture |

|
|||||
| Description | Calcium/calmodulin-dependent protein kinase type II beta chain (EC 2.7.11.17) (CaM-kinase II beta chain) (CaM kinase II subunit beta) (CaMK-II subunit beta). | |||||