Please be patient as the page loads
|
X3CL1_HUMAN
|
||||||
| SwissProt Accessions | P78423, O00672 | Gene names | CX3CL1, A-152E5.2, FKN, NTT, SCYD1 | |||
|
Domain Architecture |
|
|||||
| Description | Fractalkine precursor (CX3CL1) (Neurotactin) (CX3C membrane-anchored chemokine) (Small inducible cytokine D1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
X3CL1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 152 | |
| SwissProt Accessions | P78423, O00672 | Gene names | CX3CL1, A-152E5.2, FKN, NTT, SCYD1 | |||
|
Domain Architecture |

|
|||||
| Description | Fractalkine precursor (CX3CL1) (Neurotactin) (CX3C membrane-anchored chemokine) (Small inducible cytokine D1). | |||||
|
X3CL1_MOUSE
|
||||||
| θ value | 8.41738e-115 (rank : 2) | NC score | 0.924591 (rank : 2) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O35188, O35933 | Gene names | Cx3cl1, Cx3c, Fkn, Scyd1 | |||
|
Domain Architecture |
|
|||||
| Description | Fractalkine precursor (CX3CL1) (Neurotactin) (CX3C membrane-anchored chemokine) (Small inducible cytokine D1). | |||||
|
CCL8_HUMAN
|
||||||
| θ value | 4.45536e-07 (rank : 3) | NC score | 0.581455 (rank : 4) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P80075, P78388 | Gene names | CCL8, MCP2, SCYA10, SCYA8 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A8 precursor (CCL8) (Monocyte chemotactic protein 2) (MCP-2) (Monocyte chemoattractant protein 2) (HC14) [Contains: MCP-2(6-76)]. | |||||
|
CCL11_MOUSE
|
||||||
| θ value | 3.77169e-06 (rank : 4) | NC score | 0.587529 (rank : 3) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P48298 | Gene names | Ccl11, Scya11 | |||
|
Domain Architecture |
|
|||||
| Description | Eotaxin precursor (Small inducible cytokine A11) (CCL11) (Eosinophil chemotactic protein). | |||||
|
CCL2_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 5) | NC score | 0.572884 (rank : 6) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P13500, Q9UDF3 | Gene names | CCL2, MCP1, SCYA2 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A2 precursor (CCL2) (Monocyte chemotactic protein 1) (MCP-1) (Monocyte chemoattractant protein 1) (Monocyte chemotactic and activating factor) (MCAF) (Monocyte secretory protein JE) (HC11). | |||||
|
CCL11_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 6) | NC score | 0.576582 (rank : 5) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P51671, P50877, Q92490, Q92491 | Gene names | CCL11, SCYA11 | |||
|
Domain Architecture |
|
|||||
| Description | Eotaxin precursor (Small inducible cytokine A11) (CCL11) (Eosinophil chemotactic protein). | |||||
|
CCL7_HUMAN
|
||||||
| θ value | 1.87187e-05 (rank : 7) | NC score | 0.563159 (rank : 7) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P80098 | Gene names | CCL7, MCP3, SCYA7 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A7 precursor (CCL7) (Monocyte chemotactic protein 3) (MCP-3) (Monocyte chemoattractant protein 3) (NC28). | |||||
|
CCL13_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 8) | NC score | 0.553909 (rank : 8) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99616, O95689 | Gene names | CCL13, MCP4, NCC1, SCYA13 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A13 precursor (CCL13) (Monocyte chemotactic protein 4) (MCP-4) (Monocyte chemoattractant protein 4) (CK-beta-10) (NCC-1) [Contains: Small inducible cytokine A13, long isoform; Small inducible cytokine A13, medium isoform; Small inducible cytokine A13, short isoform]. | |||||
|
CCL12_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 9) | NC score | 0.552873 (rank : 9) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q62401, Q9QYD6 | Gene names | Ccl12, Mcp5, Scya12 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A12 precursor (CCL12) (Monocyte chemotactic protein 5) (MCP-5) (MCP-1-related chemokine). | |||||
|
CCL8_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 10) | NC score | 0.548533 (rank : 10) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9Z121 | Gene names | Ccl8, Mcp2, Scya8 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A8 precursor (CCL8) (Monocyte chemotactic protein 2) (MCP-2) (Monocyte chemoattractant protein 2). | |||||
|
CCL2_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 11) | NC score | 0.546826 (rank : 11) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P10148, Q9QYD7 | Gene names | Ccl2, Je, Mcp1, Scya2 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A2 precursor (CCL2) (Monocyte chemotactic protein 1) (MCP-1) (Monocyte chemoattractant protein 1) (Platelet- derived growth factor-inducible protein JE). | |||||
|
CCL4_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 12) | NC score | 0.515469 (rank : 13) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P13236, P22617, Q13704, Q3SXL8, Q6FGI8 | Gene names | CCL4, LAG1, MIP1B, SCYA4 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A4 precursor (CCL4) (Macrophage inflammatory protein 1-beta) (MIP-1-beta) (MIP-1-beta(1-69)) (T-cell activation protein 2) (ACT-2) (PAT 744) (H400) (SIS-gamma) (Lymphocyte activation gene 1 protein) (LAG-1) (HC21) (G-26 T-lymphocyte-secreted protein) [Contains: MIP-1-beta(3-69)]. | |||||
|
CCL3_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 13) | NC score | 0.490797 (rank : 19) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P10855, P14096 | Gene names | Ccl3, Mip1a, Scya3 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A3 precursor (CCL3) (Macrophage inflammatory protein 1-alpha) (MIP-1-alpha) (TY-5) (SIS-alpha) (Heparin-binding chemotaxis protein) (L2G25B). | |||||
|
XCL1_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 14) | NC score | 0.482239 (rank : 25) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P47992, Q52MA8 | Gene names | XCL1, LTN, SCYC1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphotactin precursor (XCL1) (Cytokine SCM-1) (ATAC) (Lymphotaxin) (SCM-1-alpha) (Small inducible cytokine C1) (XC chemokine ligand 1). | |||||
|
XCL2_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 15) | NC score | 0.481180 (rank : 26) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UBD3 | Gene names | XCL2, SCYC2 | |||
|
Domain Architecture |
|
|||||
| Description | Cytokine SCM-1 beta precursor (XCL2) (XC chemokine ligand 2). | |||||
|
CCL24_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 16) | NC score | 0.542762 (rank : 12) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O00175 | Gene names | CCL24, MPIF2, SCYA24 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A24 precursor (CCL24) (Myeloid progenitor inhibitory factor 2) (MPIF-2) (CK-beta-6) (Eosinophil chemotactic protein 2) (Eotaxin-2). | |||||
|
GRIN1_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 17) | NC score | 0.119309 (rank : 47) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q3UNH4, Q6PGF9, Q9QZY2 | Gene names | Gprin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
CCL18_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 18) | NC score | 0.495383 (rank : 14) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P55774, Q53X71 | Gene names | CCL18, AMAC1, DCCK1, MIP4, PARC, SCYA18 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A18 precursor (CCL18) (Macrophage inflammatory protein 4) (MIP-4) (Pulmonary and activation-regulated chemokine) (CC chemokine PARC) (Alternative macrophage activation- associated CC chemokine 1) (AMAC-1) (Dendritic cell chemokine 1) (DC- CK1) [Contains: CCL18(1-68); CCL18(3-69); CCL18(4-69)]. | |||||
|
CCL3L_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 19) | NC score | 0.494925 (rank : 16) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P16619, Q53YA5, Q96I68 | Gene names | CCL3L1, G0S19-2, SCYA3L1 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A3-like 1 precursor (Tonsillar lymphocyte LD78 beta protein) (LD78-beta(1-70)) (G0/G1 switch regulatory protein 19-2) (G0S19-2 protein) (PAT 464.2) [Contains: LD78-beta(3-70); LD78- beta(5-70)]. | |||||
|
CCL3_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 20) | NC score | 0.490319 (rank : 20) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P10147 | Gene names | CCL3, G0S19-1, MIP1A, SCYA3 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A3 precursor (CCL3) (Macrophage inflammatory protein 1-alpha) (MIP-1-alpha) (Tonsillar lymphocyte LD78 alpha protein) (G0/G1 switch regulatory protein 19-1) (G0S19-1 protein) (SIS-beta) (PAT 464.1) [Contains: MIP-1-alpha(4-69) (LD78-alpha(4- 69))]. | |||||
|
CCL4_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 21) | NC score | 0.487162 (rank : 22) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P14097 | Gene names | Ccl4, Mip1b, Scya4 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A4 precursor (CCL4) (Macrophage inflammatory protein 1-beta) (MIP-1-beta) (H400 protein) (SIS-gamma) (ACT2). | |||||
|
CCL21_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 22) | NC score | 0.459273 (rank : 30) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00585 | Gene names | CCL21, SCYA21 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A21 precursor (CCL21) (Beta chemokine exodus- 2) (6Ckine) (Secondary lymphoid-tissue chemokine) (SLC). | |||||
|
CCL20_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 23) | NC score | 0.483782 (rank : 24) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O89093, Q9Z1X3 | Gene names | Ccl20, Larc, Scya20 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A20 precursor (CCL20) (Macrophage inflammatory protein 3 alpha) (MIP-3-alpha) (Liver and activation- regulated chemokine) (CC chemokine LARC) (Beta chemokine exodus-1) (CC chemokine ST38). | |||||
|
CCL23_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 24) | NC score | 0.449267 (rank : 31) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P55773, O00174, O75950, Q52LD4 | Gene names | CCL23, MIP3, MPIF1, SCYA23 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A23 precursor (CCL23) (Macrophage inflammatory protein 3) (MIP-3) (Myeloid progenitor inhibitory factor 1) (MPIF-1) (CK-beta-8) (CKB-8) [Contains: CCL23(19-99); CCL23(22-99); CCL23(27-99); CCL23(30-99)]. | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 25) | NC score | 0.080561 (rank : 53) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
CC21B_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 26) | NC score | 0.398933 (rank : 37) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P84444, O09002, O09006, Q91V84 | Gene names | Ccl21b, Scya21, Scya21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small inducible cytokine A21b precursor (CCL21) (Beta chemokine exodus-2) (6Ckine) (Thymus-derived chemotactic agent 4) (TCA4). | |||||
|
ARI1B_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 27) | NC score | 0.076226 (rank : 55) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
CCL5_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 28) | NC score | 0.493616 (rank : 18) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P30882 | Gene names | Ccl5, Scya5 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A5 precursor (CCL5) (T-cell-specific RANTES protein) (SIS-delta) (MuRantes). | |||||
|
CCL14_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 29) | NC score | 0.475484 (rank : 27) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q16627, Q13954 | Gene names | CCL14, NCC2, SCYA14 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A14 precursor (CCL14) (Chemokine CC-1/CC-3) (HCC-1/HCC-3) (HCC-1(1-74)) (NCC-2) [Contains: HCC-1(3-74); HCC-1(4- 74); HCC-1(9-74)]. | |||||
|
CCL17_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 30) | NC score | 0.488050 (rank : 21) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q92583, Q2M287 | Gene names | CCL17, A-152E5.3, SCYA17, TARC | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A17 precursor (CCL17) (Thymus and activation- regulated chemokine) (CC chemokine TARC). | |||||
|
CCL24_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 31) | NC score | 0.494654 (rank : 17) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9JKC0 | Gene names | Ccl24, Scya24 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A24 precursor (CCL24) (Eosinophil chemotactic protein 2) (Eotaxin-2). | |||||
|
CC21A_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 32) | NC score | 0.395268 (rank : 38) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P84443, O09002, O09006, Q91V84 | Gene names | Ccl21a, Scya21, Scya21a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small inducible cytokine A21a precursor (CCL21) (Beta chemokine exodus-2) (6Ckine) (Thymus-derived chemotactic agent 4) (TCA4). | |||||
|
CCL5_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 33) | NC score | 0.469039 (rank : 28) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P13501, O43646, Q4ZGJ1, Q9NYA2 | Gene names | CCL5, SCYA5 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A5 precursor (CCL5) (T-cell-specific RANTES protein) (SIS-delta) (T cell-specific protein P228) (TCP228) [Contains: RANTES(3-68); RANTES(4-68)]. | |||||
|
CCL15_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 34) | NC score | 0.431912 (rank : 33) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q16663, Q9UM74 | Gene names | CCL15, MIP5, NCC3, SCYA15 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A15 precursor (CCL15) (Macrophage inflammatory protein 5) (MIP-5) (Chemokine CC-2) (HCC-2) (NCC-3) (MIP- 1 delta) (Leukotactin-1) (LKN-1) (Mrp-2b) [Contains: CCL15(22-92); CCL15(25-92); CCL15(29-92)]. | |||||
|
CCL22_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 35) | NC score | 0.448655 (rank : 32) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O88430 | Gene names | Ccl22, Abcd1, Scya22 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A22 precursor (CCL22) (CC chemokine ABCD-1) (Activated B and dendritic cell-derived). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 36) | NC score | 0.095903 (rank : 52) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
CCL19_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 37) | NC score | 0.411875 (rank : 36) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q99731, O00697, O00736 | Gene names | CCL19, ELC, MIP3B, SCYA19 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A19 precursor (CCL19) (Macrophage inflammatory protein 3 beta) (MIP-3-beta) (EBI1-ligand chemokine) (ELC) (Beta chemokine exodus-3) (CK beta-11). | |||||
|
ANKS6_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 38) | NC score | 0.034990 (rank : 113) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 411 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q68DC2, Q5VSL0, Q5VSL2, Q5VSL3, Q5VSL4, Q68DB8, Q6P2R2, Q8N9L6, Q96D62 | Gene names | ANKS6, ANKRD14, SAMD6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 6 (Sterile alpha motif domain-containing protein 6) (Ankyrin repeat domain-containing protein 14). | |||||
|
CCL22_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 39) | NC score | 0.460762 (rank : 29) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00626 | Gene names | CCL22, A-152E5.1, MDC, SCYA22 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A22 precursor (CCL22) (Macrophage-derived chemokine) (MDC(1-69)) (Stimulated T-cell chemotactic protein 1) (CC chemokine STCP-1) [Contains: MDC(3-69); MDC(5-69); MDC(7-69)]. | |||||
|
CCL26_HUMAN
|
||||||
| θ value | 0.163984 (rank : 40) | NC score | 0.494997 (rank : 15) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Y258, Q52LV8 | Gene names | CCL26, SCYA26 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A26 precursor (CCL26) (Eotaxin-3) (Macrophage inflammatory protein 4-alpha) (MIP-4-alpha) (Thymic stroma chemokine- 1) (TSC-1) (CC chemokine IMAC). | |||||
|
K1802_HUMAN
|
||||||
| θ value | 0.163984 (rank : 41) | NC score | 0.053511 (rank : 75) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 0.163984 (rank : 42) | NC score | 0.052380 (rank : 77) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 43) | NC score | 0.020144 (rank : 141) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
CO7A1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 44) | NC score | 0.046210 (rank : 90) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
DCP1A_HUMAN
|
||||||
| θ value | 0.21417 (rank : 45) | NC score | 0.062413 (rank : 64) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NPI6 | Gene names | DCP1A, SMIF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (Smad4-interacting transcriptional co-activator). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 46) | NC score | 0.052886 (rank : 76) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
SEM4D_MOUSE
|
||||||
| θ value | 0.21417 (rank : 47) | NC score | 0.019207 (rank : 143) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O09126 | Gene names | Sema4d, Semacl2, Semaj | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4D precursor (Semaphorin J) (Sema J) (Semaphorin C-like 2) (M-Sema G). | |||||
|
CCL7_MOUSE
|
||||||
| θ value | 0.279714 (rank : 48) | NC score | 0.486377 (rank : 23) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q03366 | Gene names | Ccl7, Fic, Mcp3, Scya7 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A7 precursor (CCL7) (Monocyte chemotactic protein 3) (MCP-3) (Monocyte chemoattractant protein 3) (Intercrine/chemokine MARC) (FIC protein). | |||||
|
COCA1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 49) | NC score | 0.040939 (rank : 100) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q60847, P70322 | Gene names | Col12a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 50) | NC score | 0.070059 (rank : 57) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
MEGF9_HUMAN
|
||||||
| θ value | 0.279714 (rank : 51) | NC score | 0.029144 (rank : 124) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9H1U4, O75098 | Gene names | MEGF9, EGFL5, KIAA0818 | |||
|
Domain Architecture |
|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 0.365318 (rank : 52) | NC score | 0.056540 (rank : 71) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
RIN3_MOUSE
|
||||||
| θ value | 0.365318 (rank : 53) | NC score | 0.050112 (rank : 85) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P59729 | Gene names | Rin3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
TRIO_HUMAN
|
||||||
| θ value | 0.365318 (rank : 54) | NC score | 0.008141 (rank : 157) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1580 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O75962, Q13458 | Gene names | TRIO | |||
|
Domain Architecture |

|
|||||
| Description | Triple functional domain protein (PTPRF-interacting protein). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 55) | NC score | 0.050423 (rank : 84) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 0.47712 (rank : 56) | NC score | 0.054514 (rank : 73) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
XCL1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 57) | NC score | 0.419461 (rank : 35) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P47993 | Gene names | Xcl1, Lptn, Ltn, Scyc1 | |||
|
Domain Architecture |
|
|||||
| Description | Lymphotactin precursor (XCL1) (Cytokine SCM-1) (Lymphotaxin) (Small inducible cytokine C1). | |||||
|
ANKS6_MOUSE
|
||||||
| θ value | 0.62314 (rank : 58) | NC score | 0.031020 (rank : 120) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6GQX6 | Gene names | Anks6, Samd6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 6 (Sterile alpha motif domain-containing protein 6). | |||||
|
SYN3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 59) | NC score | 0.037643 (rank : 105) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |
|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
TM131_MOUSE
|
||||||
| θ value | 0.813845 (rank : 60) | NC score | 0.055756 (rank : 72) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O70472 | Gene names | Tmem131, D1Bwg0491e, Rw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 131 (Protein RW1). | |||||
|
WIRE_HUMAN
|
||||||
| θ value | 0.813845 (rank : 61) | NC score | 0.061255 (rank : 66) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
BASP_MOUSE
|
||||||
| θ value | 1.06291 (rank : 62) | NC score | 0.079502 (rank : 54) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
COCA1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 63) | NC score | 0.038277 (rank : 104) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q99715, Q99716 | Gene names | COL12A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
M3K9_HUMAN
|
||||||
| θ value | 1.06291 (rank : 64) | NC score | 0.005647 (rank : 162) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P80192, Q6EH31, Q9H2N5 | Gene names | MAP3K9, MLK1, PRKE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 9 (EC 2.7.11.25) (Mixed lineage kinase 1). | |||||
|
NECP1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 65) | NC score | 0.037057 (rank : 106) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CR95, Q3TH13, Q78JW3, Q8C4P1 | Gene names | Necap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adaptin ear-binding coat-associated protein 1 (NECAP-1). | |||||
|
UBP10_MOUSE
|
||||||
| θ value | 1.06291 (rank : 66) | NC score | 0.029056 (rank : 125) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P52479, Q91VY7 | Gene names | Usp10, Ode-1, Uchrp | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
ASH2L_HUMAN
|
||||||
| θ value | 1.38821 (rank : 67) | NC score | 0.046297 (rank : 89) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBL3, O60659, O60660, Q96B62 | Gene names | ASH2L | |||
|
Domain Architecture |

|
|||||
| Description | Set1/Ash2 histone methyltransferase complex subunit ASH2 (ASH2-like protein). | |||||
|
CO3A1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 68) | NC score | 0.043249 (rank : 96) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
CO5A1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 69) | NC score | 0.049597 (rank : 86) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P20908, Q15094, Q5SUX4 | Gene names | COL5A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
CO5A1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 70) | NC score | 0.049596 (rank : 87) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O88207 | Gene names | Col5a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
CSPG5_MOUSE
|
||||||
| θ value | 1.38821 (rank : 71) | NC score | 0.071027 (rank : 56) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q71M36, Q71M37, Q7TNT8, Q8BPJ5, Q9QY32 | Gene names | Cspg5, Caleb, Ngc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
MAP2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 72) | NC score | 0.036393 (rank : 110) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P20357 | Gene names | Map2, Mtap2 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 2 (MAP 2). | |||||
|
PODXL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 73) | NC score | 0.057622 (rank : 69) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O00592 | Gene names | PODXL, PCLP, PCLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Podocalyxin-like protein 1 precursor. | |||||
|
PYRG2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 74) | NC score | 0.027160 (rank : 131) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70303, Q3TPQ2, Q3TT59 | Gene names | Ctps2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTP synthase 2 (EC 6.3.4.2) (UTP--ammonia ligase 2) (CTP synthetase 2) (CTPsH). | |||||
|
SCYBD_MOUSE
|
||||||
| θ value | 1.38821 (rank : 75) | NC score | 0.216084 (rank : 45) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O55038 | Gene names | Cxcl13, Blc, Scyb13 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine B13 precursor (CXCL13) (B lymphocyte chemoattractant) (CXC chemokine BLC). | |||||
|
SELPL_MOUSE
|
||||||
| θ value | 1.38821 (rank : 76) | NC score | 0.062915 (rank : 63) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q62170 | Gene names | Selplg, Selp1, Selpl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | P-selectin glycoprotein ligand 1 precursor (PSGL-1) (Selectin P ligand). | |||||
|
AATK_HUMAN
|
||||||
| θ value | 1.81305 (rank : 77) | NC score | 0.010032 (rank : 155) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1243 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6ZMQ8, O75136, Q6ZN31, Q86X28 | Gene names | AATK, KIAA0641 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis-associated tyrosine-protein kinase (EC 2.7.10.2) (AATYK) (Brain apoptosis-associated tyrosine kinase) (CDK5-binding protein) (p35-binding protein) (p35BP). | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 78) | NC score | 0.063764 (rank : 60) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CCL6_MOUSE
|
||||||
| θ value | 1.81305 (rank : 79) | NC score | 0.316739 (rank : 43) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P27784, Q3U3W3, Q5QNW1, Q99M24, Q9D830 | Gene names | Ccl6, C10, Scya6 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A6 precursor (CCL6) (C10 protein) [Contains: CCL6(22-95); CCL6(23-95)]. | |||||
|
CJ078_MOUSE
|
||||||
| θ value | 1.81305 (rank : 80) | NC score | 0.051311 (rank : 81) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 559 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BP27, Q3UIQ6, Q8R3W0, Q9CRT7, Q9D0D7, Q9D116, Q9D4W4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf78 homolog. | |||||
|
EP300_HUMAN
|
||||||
| θ value | 1.81305 (rank : 81) | NC score | 0.047974 (rank : 88) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 82) | NC score | 0.063674 (rank : 61) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
MUC13_MOUSE
|
||||||
| θ value | 1.81305 (rank : 83) | NC score | 0.051549 (rank : 79) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
CCL16_HUMAN
|
||||||
| θ value | 2.36792 (rank : 84) | NC score | 0.423257 (rank : 34) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O15467 | Gene names | CCL16, ILINCK, NCC4, SCYA16 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A16 precursor (CCL16) (IL-10-inducible chemokine) (Chemokine LEC) (Liver-expressed chemokine) (Monotactin-1) (MTN-1) (Chemokine CC-4) (HCC-4) (NCC-4) (Lymphocyte and monocyte chemoattractant) (LMC) (LCC-1). | |||||
|
CD2AP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 85) | NC score | 0.018706 (rank : 144) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y5K6, Q9UG97 | Gene names | CD2AP | |||
|
Domain Architecture |
|
|||||
| Description | CD2-associated protein (Cas ligand with multiple SH3 domains) (Adapter protein CMS). | |||||
|
COFA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 86) | NC score | 0.041543 (rank : 98) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P39059, Q5T6J4, Q9Y4W4 | Gene names | COL15A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV) (Restin) (Related to endostatin)]. | |||||
|
EMID2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 87) | NC score | 0.045005 (rank : 92) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q91VF6, Q8K4P3 | Gene names | Emid2, Col26a, Col26a1, Emu2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXVI) chain precursor (EMI domain-containing protein 2) (Protein Emu2) (Emilin and multimerin domain-containing protein 2). | |||||
|
GGA3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 88) | NC score | 0.024166 (rank : 134) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BMI3, Q80U71 | Gene names | Gga3, Kiaa0154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-binding protein GGA3 (Golgi-localized, gamma ear-containing, ARF-binding protein 3). | |||||
|
MARCO_HUMAN
|
||||||
| θ value | 2.36792 (rank : 89) | NC score | 0.032156 (rank : 116) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UEW3, Q9Y5S3 | Gene names | MARCO, SCARA2 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage receptor MARCO (Macrophage receptor with collagenous structure) (Scavenger receptor class A member 2). | |||||
|
TAF3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 90) | NC score | 0.036094 (rank : 111) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
ZSCA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 91) | NC score | 0.002372 (rank : 164) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 724 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NBB4, Q6WLH8, Q86WS8 | Gene names | ZSCAN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and SCAN domain-containing protein 1. | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 92) | NC score | 0.063013 (rank : 62) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BAT3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 93) | NC score | 0.060516 (rank : 67) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Z1R2, Q8SNA3 | Gene names | Bat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3). | |||||
|
BCAR1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 94) | NC score | 0.041445 (rank : 99) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
CIC_MOUSE
|
||||||
| θ value | 3.0926 (rank : 95) | NC score | 0.034516 (rank : 115) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
FOG1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 96) | NC score | 0.014876 (rank : 150) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
KI67_HUMAN
|
||||||
| θ value | 3.0926 (rank : 97) | NC score | 0.034762 (rank : 114) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
RERE_HUMAN
|
||||||
| θ value | 3.0926 (rank : 98) | NC score | 0.043888 (rank : 94) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
S3TC1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 99) | NC score | 0.025245 (rank : 133) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8TE82 | Gene names | SH3TC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 domain and tetratricopeptide repeats-containing protein 1. | |||||
|
TAF1C_MOUSE
|
||||||
| θ value | 3.0926 (rank : 100) | NC score | 0.028270 (rank : 127) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
ANR11_HUMAN
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.021243 (rank : 138) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
BAT3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.059754 (rank : 68) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P46379, O95874, Q5SQ35, Q5SQ36, Q5SQ37, Q5SRP8, Q96SA6, Q9BCN4 | Gene names | BAT3, G3 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3) (Protein G3). | |||||
|
BCAR1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 103) | NC score | 0.035929 (rank : 112) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61140, Q60869 | Gene names | Bcar1, Cas, Crkas | |||
|
Domain Architecture |
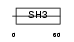
|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
BPTF_HUMAN
|
||||||
| θ value | 4.03905 (rank : 104) | NC score | 0.027950 (rank : 128) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
CCL20_HUMAN
|
||||||
| θ value | 4.03905 (rank : 105) | NC score | 0.377607 (rank : 40) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P78556, Q53S51, Q99664 | Gene names | CCL20, LARC, MIP3A, SCYA20 | |||
|
Domain Architecture |
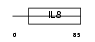
|
|||||
| Description | Small inducible cytokine A20 precursor (CCL20) (Macrophage inflammatory protein 3 alpha) (MIP-3-alpha) (Liver and activation- regulated chemokine) (CC chemokine LARC) (Beta chemokine exodus-1) [Contains: CCL20(1-67); CCL20(1-64); CCL20(2-70)]. | |||||
|
F100A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 106) | NC score | 0.029925 (rank : 123) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TB05, Q71MF6 | Gene names | FAM100A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM100A. | |||||
|
GLI2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 107) | NC score | 0.002416 (rank : 163) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 971 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P10070, O60252, O60253, O60254, O60255, Q15590, Q15591 | Gene names | GLI2, THP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI2 (Tax helper protein). | |||||
|
IRS2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 108) | NC score | 0.030432 (rank : 122) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y4H2, Q96RR2, Q9BZG0, Q9Y6I5 | Gene names | IRS2 | |||
|
Domain Architecture |
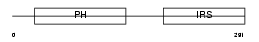
|
|||||
| Description | Insulin receptor substrate 2 (IRS-2). | |||||
|
KLF12_MOUSE
|
||||||
| θ value | 4.03905 (rank : 109) | NC score | 0.000379 (rank : 166) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35738 | Gene names | Klf12, Ap2rep | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 12 (Transcriptional repressor AP-2rep). | |||||
|
L1CAM_HUMAN
|
||||||
| θ value | 4.03905 (rank : 110) | NC score | 0.009156 (rank : 156) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P32004, Q8TA87 | Gene names | L1CAM, CAML1, MIC5 | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule L1 precursor (N-CAM L1) (CD171 antigen). | |||||
|
MAP4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 111) | NC score | 0.046150 (rank : 91) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
MYO15_HUMAN
|
||||||
| θ value | 4.03905 (rank : 112) | NC score | 0.015033 (rank : 149) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
NOS1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 113) | NC score | 0.014624 (rank : 151) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P29475 | Gene names | NOS1 | |||
|
Domain Architecture |

|
|||||
| Description | Nitric-oxide synthase, brain (EC 1.14.13.39) (NOS type I) (Neuronal NOS) (N-NOS) (nNOS) (Constitutive NOS) (NC-NOS) (bNOS). | |||||
|
OS9_HUMAN
|
||||||
| θ value | 4.03905 (rank : 114) | NC score | 0.031148 (rank : 118) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13438, O00579 | Gene names | OS9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein OS-9 precursor (Amplified in osteosarcoma 9). | |||||
|
TBCD3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 115) | NC score | 0.018598 (rank : 145) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IZP1, Q9H0B9, Q9UDD4 | Gene names | TBC1D3, PRC17 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 3 (Rab GTPase-activating protein PRC17) (Prostate cancer gene 17 protein) (TRE17 alpha protein). | |||||
|
TENS1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 116) | NC score | 0.027624 (rank : 129) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9HBL0 | Gene names | TNS1, TNS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-1. | |||||
|
ASH2L_MOUSE
|
||||||
| θ value | 5.27518 (rank : 117) | NC score | 0.040893 (rank : 101) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91X20, Q3UIF9, Q9Z2X4 | Gene names | Ash2l | |||
|
Domain Architecture |

|
|||||
| Description | Set1/Ash2 histone methyltransferase complex subunit ASH2 (ASH2-like protein). | |||||
|
CMTA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 118) | NC score | 0.019768 (rank : 142) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y6Y1, Q5VUE1 | Gene names | CAMTA1, KIAA0833 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-binding transcription activator 1. | |||||
|
CO5A3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 119) | NC score | 0.044351 (rank : 93) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P25940, Q9NZQ6 | Gene names | COL5A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(V) chain precursor. | |||||
|
EMID2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 120) | NC score | 0.040714 (rank : 102) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96A83 | Gene names | EMID2, COL26A1, EMU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXVI) chain precursor (EMI domain-containing protein 2) (Protein Emu2) (Emilin and multimerin domain-containing protein 2). | |||||
|
ENAH_MOUSE
|
||||||
| θ value | 5.27518 (rank : 121) | NC score | 0.040252 (rank : 103) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
FHOD1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 122) | NC score | 0.023543 (rank : 135) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
JIP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 123) | NC score | 0.021808 (rank : 137) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13387, Q96G62, Q99771, Q9NZ59, Q9UKQ4 | Gene names | MAPK8IP2, IB2, JIP2 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 2 (JNK-interacting protein 2) (JIP-2) (JNK MAP kinase scaffold protein 2) (Islet-brain-2) (IB-2) (Mitogen-activated protein kinase 8-interacting protein 2). | |||||
|
NFIP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 124) | NC score | 0.031023 (rank : 119) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NV92, Q7Z2H3, Q7Z428, Q8TAR3, Q9ULQ5 | Gene names | NDFIP2, KIAA1165, N4WBP5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD4 family-interacting protein 2 (NEDD4 WW domain-binding protein 5A) (Putative MAPK-activating protein PM04/PM05/PM06/PM07) (Putative NF-kappa-B-activating protein 413). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 5.27518 (rank : 125) | NC score | 0.062346 (rank : 65) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
RHBD1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 126) | NC score | 0.016065 (rank : 148) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75783, Q9NQ85 | Gene names | RHBDL1, RHBDL | |||
|
Domain Architecture |

|
|||||
| Description | Rhomboid-related protein 1 (EC 3.4.21.105) (RRP) (Rhomboid-like protein 1). | |||||
|
SC6A5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 127) | NC score | 0.007165 (rank : 158) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q761V0, Q8CFM5, Q91ZQ2 | Gene names | Slc6a5, Glyt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium- and chloride-dependent glycine transporter 2 (GlyT2) (GlyT-2) (Solute carrier family 6 member 5). | |||||
|
SDC3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 128) | NC score | 0.036584 (rank : 109) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q64519, Q3UDD9, Q6ZQA4, Q7TQD4 | Gene names | Sdc3, Kiaa0468 | |||
|
Domain Architecture |
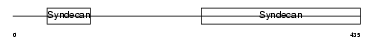
|
|||||
| Description | Syndecan-3 precursor (SYND3). | |||||
|
BCOR_MOUSE
|
||||||
| θ value | 6.88961 (rank : 129) | NC score | 0.016677 (rank : 147) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8CGN4, Q6PDK5, Q6ZPM3, Q8BKF5, Q8CGN1, Q8CGN2, Q8CGN3 | Gene names | Bcor, Kiaa1575 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BCoR protein (BCL-6 corepressor). | |||||
|
CGRE1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 130) | NC score | 0.017541 (rank : 146) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R1U2, Q9D1K1 | Gene names | Cgref1, Cgr11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
FOXP3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 131) | NC score | 0.007128 (rank : 159) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BZS1, O60827, Q4ZH51 | Gene names | FOXP3, IPEX | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P3 (Scurfin). | |||||
|
K0310_HUMAN
|
||||||
| θ value | 6.88961 (rank : 132) | NC score | 0.036790 (rank : 107) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
MINT_MOUSE
|
||||||
| θ value | 6.88961 (rank : 133) | NC score | 0.031495 (rank : 117) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PHF2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 134) | NC score | 0.020347 (rank : 140) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
SNTB2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 135) | NC score | 0.010506 (rank : 154) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13425, Q9BY09 | Gene names | SNTB2, SNT2B2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-2-syntrophin (59 kDa dystrophin-associated protein A1 basic component 2) (Syntrophin 3) (SNT3) (Syntrophin-like) (SNTL). | |||||
|
C1QR1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.006078 (rank : 160) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NPY3, O00274 | Gene names | CD93, C1QR1, MXRA4 | |||
|
Domain Architecture |
|
|||||
| Description | Complement component C1q receptor precursor (Complement component 1 q subcomponent receptor 1) (C1qR) (C1qRp) (C1qR(p)) (C1q/MBL/SPA receptor) (CD93 antigen) (CDw93). | |||||
|
CJ012_HUMAN
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.026743 (rank : 132) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
CO3A1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.042925 (rank : 97) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 466 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P08121, Q61429, Q9CRN7 | Gene names | Col3a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
COBA2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.043477 (rank : 95) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
COGA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.036646 (rank : 108) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q07092 | Gene names | COL16A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVI) chain precursor. | |||||
|
CTRO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | -0.001831 (rank : 168) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
DPOLL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.013938 (rank : 152) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QXE2, Q9CTJ1 | Gene names | Poll, Polk | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase lambda (EC 2.7.7.7) (EC 4.2.99.-) (Pol Lambda) (DNA polymerase kappa). | |||||
|
EFS_HUMAN
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.028920 (rank : 126) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O43281, O43282 | Gene names | EFS | |||
|
Domain Architecture |
|
|||||
| Description | Embryonal Fyn-associated substrate (HEFS). | |||||
|
EMIL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.023276 (rank : 136) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99K41 | Gene names | Emilin1 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-1 precursor (Elastin microfibril interface-located protein 1) (Elastin microfibril interfacer 1). | |||||
|
IL8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.138337 (rank : 46) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P10145, Q6FGF6, Q6LAE6, Q96RG6, Q9C077 | Gene names | IL8 | |||
|
Domain Architecture |
|
|||||
| Description | Interleukin-8 precursor (IL-8) (CXCL8) (Monocyte-derived neutrophil chemotactic factor) (MDNCF) (T-cell chemotactic factor) (Neutrophil- activating protein 1) (NAP-1) (Protein 3-10C) (Granulocyte chemotactic protein 1) (GCP-1) (Monocyte-derived neutrophil-activating peptide) (MONAP) (Emoctakin) [Contains: MDNCF-a (IL8/NAP1 form I) (GCP/IL-8 protein IV); Interleukin-8 (IL-8(1-77)) (MDNCF-b) (IL8/NAP1 form II) (GCP/IL-8 protein II) ((Ala-IL-8)77); IL-8(5-77); IL-8(6-77) (Lymphocyte-derived neutrophil-activating factor) (LYNAP) (Neutrophil- activating factor) (NAF) (MDNCF-c) (IL8/NAP1 form III) (GCP/IL-8 protein I) ((Ser-IL-8)72); IL-8(7-77) (IL8/NAP1 form IV) (GCP/IL-8 protein V); IL-8(8-77) (IL8/NAP1 form V) (GCP/IL-8 protein VI); IL- 8(9-77) (IL8/NAP1 form VI) (GCP/IL-8 protein III)]. | |||||
|
M3K1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | 0.001366 (rank : 165) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P53349, Q60831, Q9R0U3, Q9R256 | Gene names | Map3k1, Mekk, Mekk1 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 1 (EC 2.7.11.25) (MAPK/ERK kinase kinase 1) (MEK kinase 1) (MEKK 1). | |||||
|
MEGF9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 147) | NC score | 0.030588 (rank : 121) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
NAF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 148) | NC score | 0.005907 (rank : 161) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15025, O76008, Q96EL9, Q99833, Q9H1J3 | Gene names | TNIP1, KIAA0113, NAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (HIV-1 Nef-interacting protein) (Virion-associated nuclear shuttling protein) (VAN) (hVAN) (Nip40-1). | |||||
|
SVIL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 149) | NC score | 0.011161 (rank : 153) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95425, O60611, O60612, Q5VZK5, Q5VZK6, Q9H1R7 | Gene names | SVIL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
TCRG1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 150) | NC score | 0.020920 (rank : 139) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
ZN598_HUMAN
|
||||||
| θ value | 8.99809 (rank : 151) | NC score | 0.027488 (rank : 130) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86UK7, Q8IW49, Q8N3D9, Q96FG3, Q9H7J3 | Gene names | ZNF598 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 598. | |||||
|
ZN688_HUMAN
|
||||||
| θ value | 8.99809 (rank : 152) | NC score | -0.000854 (rank : 167) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 873 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WV14, O75701, Q8IW91, Q96MN0 | Gene names | ZNF688 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 688. | |||||
|
ARI1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.054329 (rank : 74) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
CCL19_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.383858 (rank : 39) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O70460 | Gene names | Ccl19, Elc, Scya19 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A19 precursor (CCL19) (Epstein-Barr virus- induced molecule 1 ligand chemokine) (EBI1-ligand chemokine) (ELC). | |||||
|
CCL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.349421 (rank : 41) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P22362 | Gene names | CCL1, SCYA1 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A1 precursor (CCL1) (T lymphocyte-secreted protein I-309). | |||||
|
CCL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.331488 (rank : 42) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P10146, P14098, Q9QUR9, Q9QUS5 | Gene names | Ccl1, Scya1, Tca3 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A1 precursor (CCL1) (T-cell activation protein TCA3) (TCA-3) (SIS-epsilon) (P500). | |||||
|
CCL25_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.114071 (rank : 48) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O15444, Q96KJ7 | Gene names | CCL25, SCYA25, TECK | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A25 precursor (CCL25) (Chemokine TECK) (Thymus expressed chemokine). | |||||
|
CCL9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.310905 (rank : 44) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P51670, Q5QNW2 | Gene names | Ccl9, Mrp2, Scya10, Scya9 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A9 precursor (CCL9) (Macrophage inflammatory protein 1-gamma) (MIP-1-gamma) (Macrophage inflammatory protein- related protein 2) (MRP-2) (CCF18) [Contains: CCL9(29-101); CCL9(30- 101); CCL9(31-101)]. | |||||
|
GRIN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.066839 (rank : 58) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.065380 (rank : 59) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.051986 (rank : 78) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.057502 (rank : 70) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.051418 (rank : 80) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.050614 (rank : 82) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
SCYBD_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.101443 (rank : 51) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43927 | Gene names | CXCL13, BCA1, BLC, SCYB13 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine B13 precursor (CXCL13) (B lymphocyte chemoattractant) (CXC chemokine BLC) (B cell-attracting chemokine 1) (BCA-1) (ANGIE). | |||||
|
SDF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.108740 (rank : 50) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P48061 | Gene names | CXCL12, SDF1 | |||
|
Domain Architecture |
|
|||||
| Description | Stromal cell-derived factor 1 precursor (SDF-1) (CXCL12) (Pre-B cell growth-stimulating factor) (PBSF) (hIRH) [Contains: SDF-1-beta(3-72); SDF-1-alpha(3-67)]. | |||||
|
SDF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.108814 (rank : 49) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P40224, Q543V6 | Gene names | Cxcl12, Sdf1 | |||
|
Domain Architecture |
|
|||||
| Description | Stromal cell-derived factor 1 precursor (SDF-1) (CXCL12) (Pre-B cell growth-stimulating factor) (PBSF) (12-O-tetradecanoylphorbol 13- acetate repressed protein 1) (TPAR1) (Thymic lymphoma cell-stimulating factor) (TLSF). | |||||
|
SON_HUMAN
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.050436 (rank : 83) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
X3CL1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 152 | |
| SwissProt Accessions | P78423, O00672 | Gene names | CX3CL1, A-152E5.2, FKN, NTT, SCYD1 | |||
|
Domain Architecture |

|
|||||
| Description | Fractalkine precursor (CX3CL1) (Neurotactin) (CX3C membrane-anchored chemokine) (Small inducible cytokine D1). | |||||
|
X3CL1_MOUSE
|
||||||
| NC score | 0.924591 (rank : 2) | θ value | 8.41738e-115 (rank : 2) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O35188, O35933 | Gene names | Cx3cl1, Cx3c, Fkn, Scyd1 | |||
|
Domain Architecture |

|
|||||
| Description | Fractalkine precursor (CX3CL1) (Neurotactin) (CX3C membrane-anchored chemokine) (Small inducible cytokine D1). | |||||
|
CCL11_MOUSE
|
||||||
| NC score | 0.587529 (rank : 3) | θ value | 3.77169e-06 (rank : 4) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P48298 | Gene names | Ccl11, Scya11 | |||
|
Domain Architecture |

|
|||||
| Description | Eotaxin precursor (Small inducible cytokine A11) (CCL11) (Eosinophil chemotactic protein). | |||||
|
CCL8_HUMAN
|
||||||
| NC score | 0.581455 (rank : 4) | θ value | 4.45536e-07 (rank : 3) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P80075, P78388 | Gene names | CCL8, MCP2, SCYA10, SCYA8 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A8 precursor (CCL8) (Monocyte chemotactic protein 2) (MCP-2) (Monocyte chemoattractant protein 2) (HC14) [Contains: MCP-2(6-76)]. | |||||
|
CCL11_HUMAN
|
||||||
| NC score | 0.576582 (rank : 5) | θ value | 8.40245e-06 (rank : 6) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P51671, P50877, Q92490, Q92491 | Gene names | CCL11, SCYA11 | |||
|
Domain Architecture |

|
|||||
| Description | Eotaxin precursor (Small inducible cytokine A11) (CCL11) (Eosinophil chemotactic protein). | |||||
|
CCL2_HUMAN
|
||||||
| NC score | 0.572884 (rank : 6) | θ value | 6.43352e-06 (rank : 5) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P13500, Q9UDF3 | Gene names | CCL2, MCP1, SCYA2 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A2 precursor (CCL2) (Monocyte chemotactic protein 1) (MCP-1) (Monocyte chemoattractant protein 1) (Monocyte chemotactic and activating factor) (MCAF) (Monocyte secretory protein JE) (HC11). | |||||
|
CCL7_HUMAN
|
||||||
| NC score | 0.563159 (rank : 7) | θ value | 1.87187e-05 (rank : 7) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P80098 | Gene names | CCL7, MCP3, SCYA7 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A7 precursor (CCL7) (Monocyte chemotactic protein 3) (MCP-3) (Monocyte chemoattractant protein 3) (NC28). | |||||
|
CCL13_HUMAN
|
||||||
| NC score | 0.553909 (rank : 8) | θ value | 3.19293e-05 (rank : 8) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99616, O95689 | Gene names | CCL13, MCP4, NCC1, SCYA13 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A13 precursor (CCL13) (Monocyte chemotactic protein 4) (MCP-4) (Monocyte chemoattractant protein 4) (CK-beta-10) (NCC-1) [Contains: Small inducible cytokine A13, long isoform; Small inducible cytokine A13, medium isoform; Small inducible cytokine A13, short isoform]. | |||||
|
CCL12_MOUSE
|
||||||
| NC score | 0.552873 (rank : 9) | θ value | 7.1131e-05 (rank : 9) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q62401, Q9QYD6 | Gene names | Ccl12, Mcp5, Scya12 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A12 precursor (CCL12) (Monocyte chemotactic protein 5) (MCP-5) (MCP-1-related chemokine). | |||||
|
CCL8_MOUSE
|
||||||
| NC score | 0.548533 (rank : 10) | θ value | 7.1131e-05 (rank : 10) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9Z121 | Gene names | Ccl8, Mcp2, Scya8 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A8 precursor (CCL8) (Monocyte chemotactic protein 2) (MCP-2) (Monocyte chemoattractant protein 2). | |||||
|
CCL2_MOUSE
|
||||||
| NC score | 0.546826 (rank : 11) | θ value | 9.29e-05 (rank : 11) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P10148, Q9QYD7 | Gene names | Ccl2, Je, Mcp1, Scya2 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A2 precursor (CCL2) (Monocyte chemotactic protein 1) (MCP-1) (Monocyte chemoattractant protein 1) (Platelet- derived growth factor-inducible protein JE). | |||||
|
CCL24_HUMAN
|
||||||
| NC score | 0.542762 (rank : 12) | θ value | 0.000461057 (rank : 16) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O00175 | Gene names | CCL24, MPIF2, SCYA24 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A24 precursor (CCL24) (Myeloid progenitor inhibitory factor 2) (MPIF-2) (CK-beta-6) (Eosinophil chemotactic protein 2) (Eotaxin-2). | |||||
|
CCL4_HUMAN
|
||||||
| NC score | 0.515469 (rank : 13) | θ value | 9.29e-05 (rank : 12) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P13236, P22617, Q13704, Q3SXL8, Q6FGI8 | Gene names | CCL4, LAG1, MIP1B, SCYA4 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A4 precursor (CCL4) (Macrophage inflammatory protein 1-beta) (MIP-1-beta) (MIP-1-beta(1-69)) (T-cell activation protein 2) (ACT-2) (PAT 744) (H400) (SIS-gamma) (Lymphocyte activation gene 1 protein) (LAG-1) (HC21) (G-26 T-lymphocyte-secreted protein) [Contains: MIP-1-beta(3-69)]. | |||||
|
CCL18_HUMAN
|
||||||
| NC score | 0.495383 (rank : 14) | θ value | 0.00134147 (rank : 18) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P55774, Q53X71 | Gene names | CCL18, AMAC1, DCCK1, MIP4, PARC, SCYA18 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A18 precursor (CCL18) (Macrophage inflammatory protein 4) (MIP-4) (Pulmonary and activation-regulated chemokine) (CC chemokine PARC) (Alternative macrophage activation- associated CC chemokine 1) (AMAC-1) (Dendritic cell chemokine 1) (DC- CK1) [Contains: CCL18(1-68); CCL18(3-69); CCL18(4-69)]. | |||||
|
CCL26_HUMAN
|
||||||
| NC score | 0.494997 (rank : 15) | θ value | 0.163984 (rank : 40) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Y258, Q52LV8 | Gene names | CCL26, SCYA26 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A26 precursor (CCL26) (Eotaxin-3) (Macrophage inflammatory protein 4-alpha) (MIP-4-alpha) (Thymic stroma chemokine- 1) (TSC-1) (CC chemokine IMAC). | |||||
|
CCL3L_HUMAN
|
||||||
| NC score | 0.494925 (rank : 16) | θ value | 0.00134147 (rank : 19) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P16619, Q53YA5, Q96I68 | Gene names | CCL3L1, G0S19-2, SCYA3L1 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A3-like 1 precursor (Tonsillar lymphocyte LD78 beta protein) (LD78-beta(1-70)) (G0/G1 switch regulatory protein 19-2) (G0S19-2 protein) (PAT 464.2) [Contains: LD78-beta(3-70); LD78- beta(5-70)]. | |||||
|
CCL24_MOUSE
|
||||||
| NC score | 0.494654 (rank : 17) | θ value | 0.0330416 (rank : 31) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9JKC0 | Gene names | Ccl24, Scya24 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A24 precursor (CCL24) (Eosinophil chemotactic protein 2) (Eotaxin-2). | |||||
|
CCL5_MOUSE
|
||||||
| NC score | 0.493616 (rank : 18) | θ value | 0.0252991 (rank : 28) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P30882 | Gene names | Ccl5, Scya5 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A5 precursor (CCL5) (T-cell-specific RANTES protein) (SIS-delta) (MuRantes). | |||||
|
CCL3_MOUSE
|
||||||
| NC score | 0.490797 (rank : 19) | θ value | 0.00035302 (rank : 13) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P10855, P14096 | Gene names | Ccl3, Mip1a, Scya3 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A3 precursor (CCL3) (Macrophage inflammatory protein 1-alpha) (MIP-1-alpha) (TY-5) (SIS-alpha) (Heparin-binding chemotaxis protein) (L2G25B). | |||||
|
CCL3_HUMAN
|
||||||
| NC score | 0.490319 (rank : 20) | θ value | 0.00134147 (rank : 20) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P10147 | Gene names | CCL3, G0S19-1, MIP1A, SCYA3 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A3 precursor (CCL3) (Macrophage inflammatory protein 1-alpha) (MIP-1-alpha) (Tonsillar lymphocyte LD78 alpha protein) (G0/G1 switch regulatory protein 19-1) (G0S19-1 protein) (SIS-beta) (PAT 464.1) [Contains: MIP-1-alpha(4-69) (LD78-alpha(4- 69))]. | |||||
|
CCL17_HUMAN
|
||||||
| NC score | 0.488050 (rank : 21) | θ value | 0.0330416 (rank : 30) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q92583, Q2M287 | Gene names | CCL17, A-152E5.3, SCYA17, TARC | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A17 precursor (CCL17) (Thymus and activation- regulated chemokine) (CC chemokine TARC). | |||||
|
CCL4_MOUSE
|
||||||
| NC score | 0.487162 (rank : 22) | θ value | 0.00134147 (rank : 21) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P14097 | Gene names | Ccl4, Mip1b, Scya4 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A4 precursor (CCL4) (Macrophage inflammatory protein 1-beta) (MIP-1-beta) (H400 protein) (SIS-gamma) (ACT2). | |||||
|
CCL7_MOUSE
|
||||||
| NC score | 0.486377 (rank : 23) | θ value | 0.279714 (rank : 48) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q03366 | Gene names | Ccl7, Fic, Mcp3, Scya7 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A7 precursor (CCL7) (Monocyte chemotactic protein 3) (MCP-3) (Monocyte chemoattractant protein 3) (Intercrine/chemokine MARC) (FIC protein). | |||||
|
CCL20_MOUSE
|
||||||
| NC score | 0.483782 (rank : 24) | θ value | 0.00298849 (rank : 23) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O89093, Q9Z1X3 | Gene names | Ccl20, Larc, Scya20 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A20 precursor (CCL20) (Macrophage inflammatory protein 3 alpha) (MIP-3-alpha) (Liver and activation- regulated chemokine) (CC chemokine LARC) (Beta chemokine exodus-1) (CC chemokine ST38). | |||||
|
XCL1_HUMAN
|
||||||
| NC score | 0.482239 (rank : 25) | θ value | 0.00035302 (rank : 14) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P47992, Q52MA8 | Gene names | XCL1, LTN, SCYC1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphotactin precursor (XCL1) (Cytokine SCM-1) (ATAC) (Lymphotaxin) (SCM-1-alpha) (Small inducible cytokine C1) (XC chemokine ligand 1). | |||||
|
XCL2_HUMAN
|
||||||
| NC score | 0.481180 (rank : 26) | θ value | 0.00035302 (rank : 15) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UBD3 | Gene names | XCL2, SCYC2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytokine SCM-1 beta precursor (XCL2) (XC chemokine ligand 2). | |||||
|
CCL14_HUMAN
|
||||||
| NC score | 0.475484 (rank : 27) | θ value | 0.0330416 (rank : 29) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q16627, Q13954 | Gene names | CCL14, NCC2, SCYA14 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A14 precursor (CCL14) (Chemokine CC-1/CC-3) (HCC-1/HCC-3) (HCC-1(1-74)) (NCC-2) [Contains: HCC-1(3-74); HCC-1(4- 74); HCC-1(9-74)]. | |||||
|
CCL5_HUMAN
|
||||||
| NC score | 0.469039 (rank : 28) | θ value | 0.0431538 (rank : 33) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P13501, O43646, Q4ZGJ1, Q9NYA2 | Gene names | CCL5, SCYA5 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A5 precursor (CCL5) (T-cell-specific RANTES protein) (SIS-delta) (T cell-specific protein P228) (TCP228) [Contains: RANTES(3-68); RANTES(4-68)]. | |||||
|
CCL22_HUMAN
|
||||||
| NC score | 0.460762 (rank : 29) | θ value | 0.0961366 (rank : 39) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00626 | Gene names | CCL22, A-152E5.1, MDC, SCYA22 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A22 precursor (CCL22) (Macrophage-derived chemokine) (MDC(1-69)) (Stimulated T-cell chemotactic protein 1) (CC chemokine STCP-1) [Contains: MDC(3-69); MDC(5-69); MDC(7-69)]. | |||||
|
CCL21_HUMAN
|
||||||
| NC score | 0.459273 (rank : 30) | θ value | 0.00175202 (rank : 22) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00585 | Gene names | CCL21, SCYA21 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A21 precursor (CCL21) (Beta chemokine exodus- 2) (6Ckine) (Secondary lymphoid-tissue chemokine) (SLC). | |||||
|
CCL23_HUMAN
|
||||||
| NC score | 0.449267 (rank : 31) | θ value | 0.0113563 (rank : 24) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P55773, O00174, O75950, Q52LD4 | Gene names | CCL23, MIP3, MPIF1, SCYA23 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A23 precursor (CCL23) (Macrophage inflammatory protein 3) (MIP-3) (Myeloid progenitor inhibitory factor 1) (MPIF-1) (CK-beta-8) (CKB-8) [Contains: CCL23(19-99); CCL23(22-99); CCL23(27-99); CCL23(30-99)]. | |||||
|
CCL22_MOUSE
|
||||||
| NC score | 0.448655 (rank : 32) | θ value | 0.0563607 (rank : 35) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O88430 | Gene names | Ccl22, Abcd1, Scya22 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A22 precursor (CCL22) (CC chemokine ABCD-1) (Activated B and dendritic cell-derived). | |||||
|
CCL15_HUMAN
|
||||||
| NC score | 0.431912 (rank : 33) | θ value | 0.0563607 (rank : 34) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q16663, Q9UM74 | Gene names | CCL15, MIP5, NCC3, SCYA15 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A15 precursor (CCL15) (Macrophage inflammatory protein 5) (MIP-5) (Chemokine CC-2) (HCC-2) (NCC-3) (MIP- 1 delta) (Leukotactin-1) (LKN-1) (Mrp-2b) [Contains: CCL15(22-92); CCL15(25-92); CCL15(29-92)]. | |||||
|
CCL16_HUMAN
|
||||||
| NC score | 0.423257 (rank : 34) | θ value | 2.36792 (rank : 84) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O15467 | Gene names | CCL16, ILINCK, NCC4, SCYA16 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A16 precursor (CCL16) (IL-10-inducible chemokine) (Chemokine LEC) (Liver-expressed chemokine) (Monotactin-1) (MTN-1) (Chemokine CC-4) (HCC-4) (NCC-4) (Lymphocyte and monocyte chemoattractant) (LMC) (LCC-1). | |||||
|
XCL1_MOUSE
|
||||||
| NC score | 0.419461 (rank : 35) | θ value | 0.47712 (rank : 57) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P47993 | Gene names | Xcl1, Lptn, Ltn, Scyc1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphotactin precursor (XCL1) (Cytokine SCM-1) (Lymphotaxin) (Small inducible cytokine C1). | |||||
|
CCL19_HUMAN
|
||||||
| NC score | 0.411875 (rank : 36) | θ value | 0.0736092 (rank : 37) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q99731, O00697, O00736 | Gene names | CCL19, ELC, MIP3B, SCYA19 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A19 precursor (CCL19) (Macrophage inflammatory protein 3 beta) (MIP-3-beta) (EBI1-ligand chemokine) (ELC) (Beta chemokine exodus-3) (CK beta-11). | |||||
|
CC21B_MOUSE
|
||||||
| NC score | 0.398933 (rank : 37) | θ value | 0.0193708 (rank : 26) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P84444, O09002, O09006, Q91V84 | Gene names | Ccl21b, Scya21, Scya21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small inducible cytokine A21b precursor (CCL21) (Beta chemokine exodus-2) (6Ckine) (Thymus-derived chemotactic agent 4) (TCA4). | |||||
|
CC21A_MOUSE
|
||||||
| NC score | 0.395268 (rank : 38) | θ value | 0.0431538 (rank : 32) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P84443, O09002, O09006, Q91V84 | Gene names | Ccl21a, Scya21, Scya21a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small inducible cytokine A21a precursor (CCL21) (Beta chemokine exodus-2) (6Ckine) (Thymus-derived chemotactic agent 4) (TCA4). | |||||
|
CCL19_MOUSE
|
||||||
| NC score | 0.383858 (rank : 39) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O70460 | Gene names | Ccl19, Elc, Scya19 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A19 precursor (CCL19) (Epstein-Barr virus- induced molecule 1 ligand chemokine) (EBI1-ligand chemokine) (ELC). | |||||
|
CCL20_HUMAN
|
||||||
| NC score | 0.377607 (rank : 40) | θ value | 4.03905 (rank : 105) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P78556, Q53S51, Q99664 | Gene names | CCL20, LARC, MIP3A, SCYA20 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A20 precursor (CCL20) (Macrophage inflammatory protein 3 alpha) (MIP-3-alpha) (Liver and activation- regulated chemokine) (CC chemokine LARC) (Beta chemokine exodus-1) [Contains: CCL20(1-67); CCL20(1-64); CCL20(2-70)]. | |||||
|
CCL1_HUMAN
|
||||||
| NC score | 0.349421 (rank : 41) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P22362 | Gene names | CCL1, SCYA1 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A1 precursor (CCL1) (T lymphocyte-secreted protein I-309). | |||||
|
CCL1_MOUSE
|
||||||
| NC score | 0.331488 (rank : 42) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P10146, P14098, Q9QUR9, Q9QUS5 | Gene names | Ccl1, Scya1, Tca3 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A1 precursor (CCL1) (T-cell activation protein TCA3) (TCA-3) (SIS-epsilon) (P500). | |||||
|
CCL6_MOUSE
|
||||||
| NC score | 0.316739 (rank : 43) | θ value | 1.81305 (rank : 79) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P27784, Q3U3W3, Q5QNW1, Q99M24, Q9D830 | Gene names | Ccl6, C10, Scya6 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A6 precursor (CCL6) (C10 protein) [Contains: CCL6(22-95); CCL6(23-95)]. | |||||
|
CCL9_MOUSE
|
||||||
| NC score | 0.310905 (rank : 44) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P51670, Q5QNW2 | Gene names | Ccl9, Mrp2, Scya10, Scya9 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A9 precursor (CCL9) (Macrophage inflammatory protein 1-gamma) (MIP-1-gamma) (Macrophage inflammatory protein- related protein 2) (MRP-2) (CCF18) [Contains: CCL9(29-101); CCL9(30- 101); CCL9(31-101)]. | |||||
|
SCYBD_MOUSE
|
||||||
| NC score | 0.216084 (rank : 45) | θ value | 1.38821 (rank : 75) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O55038 | Gene names | Cxcl13, Blc, Scyb13 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine B13 precursor (CXCL13) (B lymphocyte chemoattractant) (CXC chemokine BLC). | |||||
|
IL8_HUMAN
|
||||||
| NC score | 0.138337 (rank : 46) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P10145, Q6FGF6, Q6LAE6, Q96RG6, Q9C077 | Gene names | IL8 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-8 precursor (IL-8) (CXCL8) (Monocyte-derived neutrophil chemotactic factor) (MDNCF) (T-cell chemotactic factor) (Neutrophil- activating protein 1) (NAP-1) (Protein 3-10C) (Granulocyte chemotactic protein 1) (GCP-1) (Monocyte-derived neutrophil-activating peptide) (MONAP) (Emoctakin) [Contains: MDNCF-a (IL8/NAP1 form I) (GCP/IL-8 protein IV); Interleukin-8 (IL-8(1-77)) (MDNCF-b) (IL8/NAP1 form II) (GCP/IL-8 protein II) ((Ala-IL-8)77); IL-8(5-77); IL-8(6-77) (Lymphocyte-derived neutrophil-activating factor) (LYNAP) (Neutrophil- activating factor) (NAF) (MDNCF-c) (IL8/NAP1 form III) (GCP/IL-8 protein I) ((Ser-IL-8)72); IL-8(7-77) (IL8/NAP1 form IV) (GCP/IL-8 protein V); IL-8(8-77) (IL8/NAP1 form V) (GCP/IL-8 protein VI); IL- 8(9-77) (IL8/NAP1 form VI) (GCP/IL-8 protein III)]. | |||||
|
GRIN1_MOUSE
|
||||||
| NC score | 0.119309 (rank : 47) | θ value | 0.000602161 (rank : 17) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q3UNH4, Q6PGF9, Q9QZY2 | Gene names | Gprin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
CCL25_HUMAN
|
||||||
| NC score | 0.114071 (rank : 48) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O15444, Q96KJ7 | Gene names | CCL25, SCYA25, TECK | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A25 precursor (CCL25) (Chemokine TECK) (Thymus expressed chemokine). | |||||
|
SDF1_MOUSE
|
||||||
| NC score | 0.108814 (rank : 49) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P40224, Q543V6 | Gene names | Cxcl12, Sdf1 | |||
|
Domain Architecture |

|
|||||
| Description | Stromal cell-derived factor 1 precursor (SDF-1) (CXCL12) (Pre-B cell growth-stimulating factor) (PBSF) (12-O-tetradecanoylphorbol 13- acetate repressed protein 1) (TPAR1) (Thymic lymphoma cell-stimulating factor) (TLSF). | |||||
|
SDF1_HUMAN
|
||||||
| NC score | 0.108740 (rank : 50) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P48061 | Gene names | CXCL12, SDF1 | |||
|
Domain Architecture |

|
|||||
| Description | Stromal cell-derived factor 1 precursor (SDF-1) (CXCL12) (Pre-B cell growth-stimulating factor) (PBSF) (hIRH) [Contains: SDF-1-beta(3-72); SDF-1-alpha(3-67)]. | |||||
|
SCYBD_HUMAN
|
||||||
| NC score | 0.101443 (rank : 51) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43927 | Gene names | CXCL13, BCA1, BLC, SCYB13 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine B13 precursor (CXCL13) (B lymphocyte chemoattractant) (CXC chemokine BLC) (B cell-attracting chemokine 1) (BCA-1) (ANGIE). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.095903 (rank : 52) | θ value | 0.0563607 (rank : 36) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.080561 (rank : 53) | θ value | 0.0113563 (rank : 25) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.079502 (rank : 54) | θ value | 1.06291 (rank : 62) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
ARI1B_HUMAN
|
||||||
| NC score | 0.076226 (rank : 55) | θ value | 0.0252991 (rank : 27) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
CSPG5_MOUSE
|
||||||
| NC score | 0.071027 (rank : 56) | θ value | 1.38821 (rank : 71) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q71M36, Q71M37, Q7TNT8, Q8BPJ5, Q9QY32 | Gene names | Cspg5, Caleb, Ngc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.070059 (rank : 57) | θ value | 0.279714 (rank : 50) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
GRIN1_HUMAN
|
||||||
| NC score | 0.066839 (rank : 58) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.065380 (rank : 59) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.063764 (rank : 60) | θ value | 1.81305 (rank : 78) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.063674 (rank : 61) | θ value | 1.81305 (rank : 82) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.063013 (rank : 62) | θ value | 3.0926 (rank : 92) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
SELPL_MOUSE
|
||||||
| NC score | 0.062915 (rank : 63) | θ value | 1.38821 (rank : 76) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q62170 | Gene names | Selplg, Selp1, Selpl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | P-selectin glycoprotein ligand 1 precursor (PSGL-1) (Selectin P ligand). | |||||
|
DCP1A_HUMAN
|
||||||
| NC score | 0.062413 (rank : 64) | θ value | 0.21417 (rank : 45) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NPI6 | Gene names | DCP1A, SMIF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (Smad4-interacting transcriptional co-activator). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.062346 (rank : 65) | θ value | 5.27518 (rank : 125) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
WIRE_HUMAN
|
||||||
| NC score | 0.061255 (rank : 66) | θ value | 0.813845 (rank : 61) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
BAT3_MOUSE
|
||||||
| NC score | 0.060516 (rank : 67) | θ value | 3.0926 (rank : 93) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Z1R2, Q8SNA3 | Gene names | Bat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3). | |||||
|
BAT3_HUMAN
|
||||||
| NC score | 0.059754 (rank : 68) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P46379, O95874, Q5SQ35, Q5SQ36, Q5SQ37, Q5SRP8, Q96SA6, Q9BCN4 | Gene names | BAT3, G3 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3) (Protein G3). | |||||
|
PODXL_HUMAN
|
||||||
| NC score | 0.057622 (rank : 69) | θ value | 1.38821 (rank : 73) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O00592 | Gene names | PODXL, PCLP, PCLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Podocalyxin-like protein 1 precursor. | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.057502 (rank : 70) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.056540 (rank : 71) | θ value | 0.365318 (rank : 52) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
TM131_MOUSE
|
||||||
| NC score | 0.055756 (rank : 72) | θ value | 0.813845 (rank : 60) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O70472 | Gene names | Tmem131, D1Bwg0491e, Rw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 131 (Protein RW1). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.054514 (rank : 73) | θ value | 0.47712 (rank : 56) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
ARI1A_HUMAN
|
||||||
| NC score | 0.054329 (rank : 74) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
K1802_HUMAN
|
||||||
| NC score | 0.053511 (rank : 75) | θ value | 0.163984 (rank : 41) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.052886 (rank : 76) | θ value | 0.21417 (rank : 46) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.052380 (rank : 77) | θ value | 0.163984 (rank : 42) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.051986 (rank : 78) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
MUC13_MOUSE
|
||||||
| NC score | 0.051549 (rank : 79) | θ value | 1.81305 (rank : 83) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.051418 (rank : 80) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
CJ078_MOUSE
|
||||||
| NC score | 0.051311 (rank : 81) | θ value | 1.81305 (rank : 80) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 559 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BP27, Q3UIQ6, Q8R3W0, Q9CRT7, Q9D0D7, Q9D116, Q9D4W4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf78 homolog. | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.050614 (rank : 82) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.050436 (rank : 83) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.050423 (rank : 84) | θ value | 0.47712 (rank : 55) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
RIN3_MOUSE
|
||||||
| NC score | 0.050112 (rank : 85) | θ value | 0.365318 (rank : 53) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P59729 | Gene names | Rin3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
CO5A1_HUMAN
|
||||||
| NC score | 0.049597 (rank : 86) | θ value | 1.38821 (rank : 69) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P20908, Q15094, Q5SUX4 | Gene names | COL5A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
CO5A1_MOUSE
|
||||||
| NC score | 0.049596 (rank : 87) | θ value | 1.38821 (rank : 70) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O88207 | Gene names | Col5a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.047974 (rank : 88) | θ value | 1.81305 (rank : 81) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
ASH2L_HUMAN
|
||||||
| NC score | 0.046297 (rank : 89) | θ value | 1.38821 (rank : 67) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBL3, O60659, O60660, Q96B62 | Gene names | ASH2L | |||
|
Domain Architecture |

|
|||||
| Description | Set1/Ash2 histone methyltransferase complex subunit ASH2 (ASH2-like protein). | |||||
|
CO7A1_HUMAN
|
||||||
| NC score | 0.046210 (rank : 90) | θ value | 0.21417 (rank : 44) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
MAP4_MOUSE
|
||||||
| NC score | 0.046150 (rank : 91) | θ value | 4.03905 (rank : 111) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
EMID2_MOUSE
|
||||||
| NC score | 0.045005 (rank : 92) | θ value | 2.36792 (rank : 87) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q91VF6, Q8K4P3 | Gene names | Emid2, Col26a, Col26a1, Emu2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXVI) chain precursor (EMI domain-containing protein 2) (Protein Emu2) (Emilin and multimerin domain-containing protein 2). | |||||
|
CO5A3_HUMAN
|
||||||
| NC score | 0.044351 (rank : 93) | θ value | 5.27518 (rank : 119) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P25940, Q9NZQ6 | Gene names | COL5A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(V) chain precursor. | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.043888 (rank : 94) | θ value | 3.0926 (rank : 98) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
COBA2_HUMAN
|
||||||
| NC score | 0.043477 (rank : 95) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
CO3A1_HUMAN
|
||||||
| NC score | 0.043249 (rank : 96) | θ value | 1.38821 (rank : 68) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
CO3A1_MOUSE
|
||||||
| NC score | 0.042925 (rank : 97) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 466 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P08121, Q61429, Q9CRN7 | Gene names | Col3a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
COFA1_HUMAN
|
||||||
| NC score | 0.041543 (rank : 98) | θ value | 2.36792 (rank : 86) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P39059, Q5T6J4, Q9Y4W4 | Gene names | COL15A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV) (Restin) (Related to endostatin)]. | |||||
|
BCAR1_HUMAN
|
||||||
| NC score | 0.041445 (rank : 99) | θ value | 3.0926 (rank : 94) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
COCA1_MOUSE
|
||||||
| NC score | 0.040939 (rank : 100) | θ value | 0.279714 (rank : 49) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q60847, P70322 | Gene names | Col12a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
ASH2L_MOUSE
|
||||||
| NC score | 0.040893 (rank : 101) | θ value | 5.27518 (rank : 117) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91X20, Q3UIF9, Q9Z2X4 | Gene names | Ash2l | |||
|
Domain Architecture |

|
|||||
| Description | Set1/Ash2 histone methyltransferase complex subunit ASH2 (ASH2-like protein). | |||||
|
EMID2_HUMAN
|
||||||
| NC score | 0.040714 (rank : 102) | θ value | 5.27518 (rank : 120) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96A83 | Gene names | EMID2, COL26A1, EMU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXVI) chain precursor (EMI domain-containing protein 2) (Protein Emu2) (Emilin and multimerin domain-containing protein 2). | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.040252 (rank : 103) | θ value | 5.27518 (rank : 121) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
COCA1_HUMAN
|
||||||
| NC score | 0.038277 (rank : 104) | θ value | 1.06291 (rank : 63) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q99715, Q99716 | Gene names | COL12A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
SYN3_HUMAN
|
||||||
| NC score | 0.037643 (rank : 105) | θ value | 0.62314 (rank : 59) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
NECP1_MOUSE
|
||||||
| NC score | 0.037057 (rank : 106) | θ value | 1.06291 (rank : 65) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CR95, Q3TH13, Q78JW3, Q8C4P1 | Gene names | Necap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adaptin ear-binding coat-associated protein 1 (NECAP-1). | |||||
|
K0310_HUMAN
|
||||||
| NC score | 0.036790 (rank : 107) | θ value | 6.88961 (rank : 132) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
COGA1_HUMAN
|
||||||
| NC score | 0.036646 (rank : 108) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q07092 | Gene names | COL16A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVI) chain precursor. | |||||
|
SDC3_MOUSE
|
||||||
| NC score | 0.036584 (rank : 109) | θ value | 5.27518 (rank : 128) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q64519, Q3UDD9, Q6ZQA4, Q7TQD4 | Gene names | Sdc3, Kiaa0468 | |||
|
Domain Architecture |

|
|||||
| Description | Syndecan-3 precursor (SYND3). | |||||
|
MAP2_MOUSE
|
||||||
| NC score | 0.036393 (rank : 110) | θ value | 1.38821 (rank : 72) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P20357 | Gene names | Map2, Mtap2 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 2 (MAP 2). | |||||
|
TAF3_HUMAN
|
||||||
| NC score | 0.036094 (rank : 111) | θ value | 2.36792 (rank : 90) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
BCAR1_MOUSE
|
||||||
| NC score | 0.035929 (rank : 112) | θ value | 4.03905 (rank : 103) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61140, Q60869 | Gene names | Bcar1, Cas, Crkas | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
ANKS6_HUMAN
|
||||||
| NC score | 0.034990 (rank : 113) | θ value | 0.0961366 (rank : 38) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 411 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q68DC2, Q5VSL0, Q5VSL2, Q5VSL3, Q5VSL4, Q68DB8, Q6P2R2, Q8N9L6, Q96D62 | Gene names | ANKS6, ANKRD14, SAMD6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 6 (Sterile alpha motif domain-containing protein 6) (Ankyrin repeat domain-containing protein 14). | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.034762 (rank : 114) | θ value | 3.0926 (rank : 97) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
CIC_MOUSE
|
||||||
| NC score | 0.034516 (rank : 115) | θ value | 3.0926 (rank : 95) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
MARCO_HUMAN
|
||||||
| NC score | 0.032156 (rank : 116) | θ value | 2.36792 (rank : 89) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UEW3, Q9Y5S3 | Gene names | MARCO, SCARA2 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage receptor MARCO (Macrophage receptor with collagenous structure) (Scavenger receptor class A member 2). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.031495 (rank : 117) | θ value | 6.88961 (rank : 133) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
OS9_HUMAN
|
||||||
| NC score | 0.031148 (rank : 118) | θ value | 4.03905 (rank : 114) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13438, O00579 | Gene names | OS9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein OS-9 precursor (Amplified in osteosarcoma 9). | |||||
|
NFIP2_HUMAN
|
||||||
| NC score | 0.031023 (rank : 119) | θ value | 5.27518 (rank : 124) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NV92, Q7Z2H3, Q7Z428, Q8TAR3, Q9ULQ5 | Gene names | NDFIP2, KIAA1165, N4WBP5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD4 family-interacting protein 2 (NEDD4 WW domain-binding protein 5A) (Putative MAPK-activating protein PM04/PM05/PM06/PM07) (Putative NF-kappa-B-activating protein 413). | |||||
|
ANKS6_MOUSE
|
||||||
| NC score | 0.031020 (rank : 120) | θ value | 0.62314 (rank : 58) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6GQX6 | Gene names | Anks6, Samd6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 6 (Sterile alpha motif domain-containing protein 6). | |||||
|
MEGF9_MOUSE
|
||||||
| NC score | 0.030588 (rank : 121) | θ value | 8.99809 (rank : 147) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
IRS2_HUMAN
|
||||||
| NC score | 0.030432 (rank : 122) | θ value | 4.03905 (rank : 108) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y4H2, Q96RR2, Q9BZG0, Q9Y6I5 | Gene names | IRS2 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin receptor substrate 2 (IRS-2). | |||||
|
F100A_HUMAN
|
||||||
| NC score | 0.029925 (rank : 123) | θ value | 4.03905 (rank : 106) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TB05, Q71MF6 | Gene names | FAM100A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM100A. | |||||
|
MEGF9_HUMAN
|
||||||
| NC score | 0.029144 (rank : 124) | θ value | 0.279714 (rank : 51) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9H1U4, O75098 | Gene names | MEGF9, EGFL5, KIAA0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
UBP10_MOUSE
|
||||||
| NC score | 0.029056 (rank : 125) | θ value | 1.06291 (rank : 66) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P52479, Q91VY7 | Gene names | Usp10, Ode-1, Uchrp | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
EFS_HUMAN
|
||||||
| NC score | 0.028920 (rank : 126) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O43281, O43282 | Gene names | EFS | |||
|
Domain Architecture |

|
|||||
| Description | Embryonal Fyn-associated substrate (HEFS). | |||||
|
TAF1C_MOUSE
|
||||||
| NC score | 0.028270 (rank : 127) | θ value | 3.0926 (rank : 100) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
BPTF_HUMAN
|
||||||
| NC score | 0.027950 (rank : 128) | θ value | 4.03905 (rank : 104) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
TENS1_HUMAN
|
||||||
| NC score | 0.027624 (rank : 129) | θ value | 4.03905 (rank : 116) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9HBL0 | Gene names | TNS1, TNS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-1. | |||||
|
ZN598_HUMAN
|
||||||
| NC score | 0.027488 (rank : 130) | θ value | 8.99809 (rank : 151) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86UK7, Q8IW49, Q8N3D9, Q96FG3, Q9H7J3 | Gene names | ZNF598 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 598. | |||||
|
PYRG2_MOUSE
|
||||||
| NC score | 0.027160 (rank : 131) | θ value | 1.38821 (rank : 74) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70303, Q3TPQ2, Q3TT59 | Gene names | Ctps2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTP synthase 2 (EC 6.3.4.2) (UTP--ammonia ligase 2) (CTP synthetase 2) (CTPsH). | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.026743 (rank : 132) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
S3TC1_HUMAN
|
||||||
| NC score | 0.025245 (rank : 133) | θ value | 3.0926 (rank : 99) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8TE82 | Gene names | SH3TC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 domain and tetratricopeptide repeats-containing protein 1. | |||||
|
GGA3_MOUSE
|
||||||
| NC score | 0.024166 (rank : 134) | θ value | 2.36792 (rank : 88) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BMI3, Q80U71 | Gene names | Gga3, Kiaa0154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-binding protein GGA3 (Golgi-localized, gamma ear-containing, ARF-binding protein 3). | |||||
|
FHOD1_HUMAN
|
||||||
| NC score | 0.023543 (rank : 135) | θ value | 5.27518 (rank : 122) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
EMIL1_MOUSE
|
||||||
| NC score | 0.023276 (rank : 136) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99K41 | Gene names | Emilin1 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-1 precursor (Elastin microfibril interface-located protein 1) (Elastin microfibril interfacer 1). | |||||
|
JIP2_HUMAN
|
||||||
| NC score | 0.021808 (rank : 137) | θ value | 5.27518 (rank : 123) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13387, Q96G62, Q99771, Q9NZ59, Q9UKQ4 | Gene names | MAPK8IP2, IB2, JIP2 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 2 (JNK-interacting protein 2) (JIP-2) (JNK MAP kinase scaffold protein 2) (Islet-brain-2) (IB-2) (Mitogen-activated protein kinase 8-interacting protein 2). | |||||
|
ANR11_HUMAN
|
||||||
| NC score | 0.021243 (rank : 138) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
TCRG1_HUMAN
|
||||||
| NC score | 0.020920 (rank : 139) | θ value | 8.99809 (rank : 150) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
PHF2_MOUSE
|
||||||
| NC score | 0.020347 (rank : 140) | θ value | 6.88961 (rank : 134) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
WNK2_HUMAN
|
||||||
| NC score | 0.020144 (rank : 141) | θ value | 0.163984 (rank : 43) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
CMTA1_HUMAN
|
||||||
| NC score | 0.019768 (rank : 142) | θ value | 5.27518 (rank : 118) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y6Y1, Q5VUE1 | Gene names | CAMTA1, KIAA0833 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-binding transcription activator 1. | |||||
|
SEM4D_MOUSE
|
||||||
| NC score | 0.019207 (rank : 143) | θ value | 0.21417 (rank : 47) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O09126 | Gene names | Sema4d, Semacl2, Semaj | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4D precursor (Semaphorin J) (Sema J) (Semaphorin C-like 2) (M-Sema G). | |||||
|
CD2AP_HUMAN
|
||||||
| NC score | 0.018706 (rank : 144) | θ value | 2.36792 (rank : 85) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y5K6, Q9UG97 | Gene names | CD2AP | |||
|
Domain Architecture |

|
|||||
| Description | CD2-associated protein (Cas ligand with multiple SH3 domains) (Adapter protein CMS). | |||||
|
TBCD3_HUMAN
|
||||||
| NC score | 0.018598 (rank : 145) | θ value | 4.03905 (rank : 115) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IZP1, Q9H0B9, Q9UDD4 | Gene names | TBC1D3, PRC17 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 3 (Rab GTPase-activating protein PRC17) (Prostate cancer gene 17 protein) (TRE17 alpha protein). | |||||
|
CGRE1_MOUSE
|
||||||
| NC score | 0.017541 (rank : 146) | θ value | 6.88961 (rank : 130) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R1U2, Q9D1K1 | Gene names | Cgref1, Cgr11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
BCOR_MOUSE
|
||||||
| NC score | 0.016677 (rank : 147) | θ value | 6.88961 (rank : 129) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8CGN4, Q6PDK5, Q6ZPM3, Q8BKF5, Q8CGN1, Q8CGN2, Q8CGN3 | Gene names | Bcor, Kiaa1575 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BCoR protein (BCL-6 corepressor). | |||||
|
RHBD1_HUMAN
|
||||||
| NC score | 0.016065 (rank : 148) | θ value | 5.27518 (rank : 126) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75783, Q9NQ85 | Gene names | RHBDL1, RHBDL | |||
|
Domain Architecture |

|
|||||
| Description | Rhomboid-related protein 1 (EC 3.4.21.105) (RRP) (Rhomboid-like protein 1). | |||||
|
MYO15_HUMAN
|
||||||
| NC score | 0.015033 (rank : 149) | θ value | 4.03905 (rank : 112) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
FOG1_MOUSE
|
||||||
| NC score | 0.014876 (rank : 150) | θ value | 3.0926 (rank : 96) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
NOS1_HUMAN
|
||||||
| NC score | 0.014624 (rank : 151) | θ value | 4.03905 (rank : 113) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P29475 | Gene names | NOS1 | |||
|
Domain Architecture |

|
|||||
| Description | Nitric-oxide synthase, brain (EC 1.14.13.39) (NOS type I) (Neuronal NOS) (N-NOS) (nNOS) (Constitutive NOS) (NC-NOS) (bNOS). | |||||
|
DPOLL_MOUSE
|
||||||
| NC score | 0.013938 (rank : 152) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QXE2, Q9CTJ1 | Gene names | Poll, Polk | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase lambda (EC 2.7.7.7) (EC 4.2.99.-) (Pol Lambda) (DNA polymerase kappa). | |||||
|
SVIL_HUMAN
|
||||||
| NC score | 0.011161 (rank : 153) | θ value | 8.99809 (rank : 149) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95425, O60611, O60612, Q5VZK5, Q5VZK6, Q9H1R7 | Gene names | SVIL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
SNTB2_HUMAN
|
||||||
| NC score | 0.010506 (rank : 154) | θ value | 6.88961 (rank : 135) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13425, Q9BY09 | Gene names | SNTB2, SNT2B2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-2-syntrophin (59 kDa dystrophin-associated protein A1 basic component 2) (Syntrophin 3) (SNT3) (Syntrophin-like) (SNTL). | |||||
|
AATK_HUMAN
|
||||||
| NC score | 0.010032 (rank : 155) | θ value | 1.81305 (rank : 77) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1243 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6ZMQ8, O75136, Q6ZN31, Q86X28 | Gene names | AATK, KIAA0641 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis-associated tyrosine-protein kinase (EC 2.7.10.2) (AATYK) (Brain apoptosis-associated tyrosine kinase) (CDK5-binding protein) (p35-binding protein) (p35BP). | |||||
|
L1CAM_HUMAN
|
||||||
| NC score | 0.009156 (rank : 156) | θ value | 4.03905 (rank : 110) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P32004, Q8TA87 | Gene names | L1CAM, CAML1, MIC5 | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule L1 precursor (N-CAM L1) (CD171 antigen). | |||||
|
TRIO_HUMAN
|
||||||
| NC score | 0.008141 (rank : 157) | θ value | 0.365318 (rank : 54) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1580 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O75962, Q13458 | Gene names | TRIO | |||
|
Domain Architecture |

|
|||||
| Description | Triple functional domain protein (PTPRF-interacting protein). | |||||
|
SC6A5_MOUSE
|
||||||
| NC score | 0.007165 (rank : 158) | θ value | 5.27518 (rank : 127) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q761V0, Q8CFM5, Q91ZQ2 | Gene names | Slc6a5, Glyt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium- and chloride-dependent glycine transporter 2 (GlyT2) (GlyT-2) (Solute carrier family 6 member 5). | |||||
|
FOXP3_HUMAN
|
||||||
| NC score | 0.007128 (rank : 159) | θ value | 6.88961 (rank : 131) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BZS1, O60827, Q4ZH51 | Gene names | FOXP3, IPEX | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P3 (Scurfin). | |||||
|
C1QR1_HUMAN
|
||||||
| NC score | 0.006078 (rank : 160) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NPY3, O00274 | Gene names | CD93, C1QR1, MXRA4 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C1q receptor precursor (Complement component 1 q subcomponent receptor 1) (C1qR) (C1qRp) (C1qR(p)) (C1q/MBL/SPA receptor) (CD93 antigen) (CDw93). | |||||
|
NAF1_HUMAN
|
||||||
| NC score | 0.005907 (rank : 161) | θ value | 8.99809 (rank : 148) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15025, O76008, Q96EL9, Q99833, Q9H1J3 | Gene names | TNIP1, KIAA0113, NAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (HIV-1 Nef-interacting protein) (Virion-associated nuclear shuttling protein) (VAN) (hVAN) (Nip40-1). | |||||
|
M3K9_HUMAN
|
||||||
| NC score | 0.005647 (rank : 162) | θ value | 1.06291 (rank : 64) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P80192, Q6EH31, Q9H2N5 | Gene names | MAP3K9, MLK1, PRKE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 9 (EC 2.7.11.25) (Mixed lineage kinase 1). | |||||
|
GLI2_HUMAN
|
||||||
| NC score | 0.002416 (rank : 163) | θ value | 4.03905 (rank : 107) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 971 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P10070, O60252, O60253, O60254, O60255, Q15590, Q15591 | Gene names | GLI2, THP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI2 (Tax helper protein). | |||||
|
ZSCA1_HUMAN
|
||||||
| NC score | 0.002372 (rank : 164) | θ value | 2.36792 (rank : 91) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 724 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NBB4, Q6WLH8, Q86WS8 | Gene names | ZSCAN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and SCAN domain-containing protein 1. | |||||
|
M3K1_MOUSE
|
||||||
| NC score | 0.001366 (rank : 165) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P53349, Q60831, Q9R0U3, Q9R256 | Gene names | Map3k1, Mekk, Mekk1 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 1 (EC 2.7.11.25) (MAPK/ERK kinase kinase 1) (MEK kinase 1) (MEKK 1). | |||||
|
KLF12_MOUSE
|
||||||
| NC score | 0.000379 (rank : 166) | θ value | 4.03905 (rank : 109) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35738 | Gene names | Klf12, Ap2rep | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 12 (Transcriptional repressor AP-2rep). | |||||
|
ZN688_HUMAN
|
||||||
| NC score | -0.000854 (rank : 167) | θ value | 8.99809 (rank : 152) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 873 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WV14, O75701, Q8IW91, Q96MN0 | Gene names | ZNF688 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 688. | |||||
|
CTRO_HUMAN
|
||||||
| NC score | -0.001831 (rank : 168) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 152 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||