Please be patient as the page loads
|
NFIP2_HUMAN
|
||||||
| SwissProt Accessions | Q9NV92, Q7Z2H3, Q7Z428, Q8TAR3, Q9ULQ5 | Gene names | NDFIP2, KIAA1165, N4WBP5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD4 family-interacting protein 2 (NEDD4 WW domain-binding protein 5A) (Putative MAPK-activating protein PM04/PM05/PM06/PM07) (Putative NF-kappa-B-activating protein 413). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
NFIP2_HUMAN
|
||||||
| θ value | 2.35023e-141 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NV92, Q7Z2H3, Q7Z428, Q8TAR3, Q9ULQ5 | Gene names | NDFIP2, KIAA1165, N4WBP5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD4 family-interacting protein 2 (NEDD4 WW domain-binding protein 5A) (Putative MAPK-activating protein PM04/PM05/PM06/PM07) (Putative NF-kappa-B-activating protein 413). | |||||
|
NFIP1_HUMAN
|
||||||
| θ value | 8.87376e-48 (rank : 2) | NC score | 0.917261 (rank : 2) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BT67, Q658T8, Q8N2E3, Q8N2F9 | Gene names | NDFIP1, N4WBP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD4 family-interacting protein 1 (NEDD4 WW domain-binding protein 5) (Putative MAPK-activating protein PM13) (Putative NF-kappa-B- activating protein 164) (Putative NFKB and MAPK-activating protein) (Breast cancer-associated protein SGA-1M). | |||||
|
NFIP1_MOUSE
|
||||||
| θ value | 1.67352e-46 (rank : 3) | NC score | 0.915665 (rank : 3) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R0W6, Q9EQH8 | Gene names | Ndfip1, N4wbp5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD4 family-interacting protein 1 (NEDD4 WW domain-binding protein 5). | |||||
|
SELPL_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 4) | NC score | 0.104228 (rank : 4) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q62170 | Gene names | Selplg, Selp1, Selpl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | P-selectin glycoprotein ligand 1 precursor (PSGL-1) (Selectin P ligand). | |||||
|
MUC2_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 5) | NC score | 0.048936 (rank : 11) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | 0.21417 (rank : 6) | NC score | 0.060521 (rank : 5) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
TCRG1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 7) | NC score | 0.054789 (rank : 8) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 8) | NC score | 0.040629 (rank : 15) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
RERE_MOUSE
|
||||||
| θ value | 0.62314 (rank : 9) | NC score | 0.044051 (rank : 13) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
CN155_HUMAN
|
||||||
| θ value | 0.813845 (rank : 10) | NC score | 0.049365 (rank : 10) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
CUTL2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 11) | NC score | 0.029008 (rank : 33) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 771 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70298 | Gene names | Cutl2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein cut-like 2 (Homeobox protein Cux-2) (Cut-like 2). | |||||
|
LMO7_HUMAN
|
||||||
| θ value | 1.06291 (rank : 12) | NC score | 0.042776 (rank : 14) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
NFRKB_MOUSE
|
||||||
| θ value | 1.38821 (rank : 13) | NC score | 0.054818 (rank : 7) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
RXRB_HUMAN
|
||||||
| θ value | 1.38821 (rank : 14) | NC score | 0.011463 (rank : 68) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P28702, P28703 | Gene names | RXRB, NR2B2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoic acid receptor RXR-beta (Retinoid X receptor beta). | |||||
|
CCNK_MOUSE
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.036922 (rank : 19) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
NC6IP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.039538 (rank : 16) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96RS0, Q5GH23, Q8TDG9, Q96QU3, Q9H5V3 | Gene names | NCOA6IP, PIMT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA methyltransferase NCOA6IP (EC 2.1.1.-) (Nuclear receptor coactivator 6-interacting protein) (PRIP-interacting protein with methyltransferase motif) (PIPMT) (PIMT) (CLL-associated antigen KW-2) (Hepatocellular carcinoma-associated antigen 137) (HCA137). | |||||
|
NHERF_MOUSE
|
||||||
| θ value | 1.81305 (rank : 17) | NC score | 0.023864 (rank : 45) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70441 | Gene names | Slc9a3r1, Nherf | |||
|
Domain Architecture |
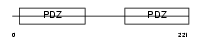
|
|||||
| Description | Ezrin-radixin-moesin-binding phosphoprotein 50 (EBP50) (Na(+)/H(+) exchange regulatory cofactor NHE-RF) (NHERF-1) (Regulatory cofactor of Na(+)/H(+) exchanger) (Sodium-hydrogen exchanger regulatory factor 1) (Solute carrier family 9 isoform 3 regulatory factor 1). | |||||
|
BCL7B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.032947 (rank : 24) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BQE9, O43769, Q13845, Q6ZW75 | Gene names | BCL7B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell CLL/lymphoma 7 protein family member B. | |||||
|
CSPG3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 19) | NC score | 0.013975 (rank : 65) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P55066, Q6P1E3, Q8C4F8 | Gene names | Cspg3, Ncan | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
NID2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 20) | NC score | 0.014841 (rank : 63) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q14112, O43710 | Gene names | NID2 | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-2 precursor (NID-2) (Osteonidogen). | |||||
|
SFPQ_HUMAN
|
||||||
| θ value | 2.36792 (rank : 21) | NC score | 0.026643 (rank : 37) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
SHAN2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.035603 (rank : 22) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 446 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80Z38, Q3UTK4, Q5DU07 | Gene names | Shank2, Kiaa1022 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2) (Cortactin- binding protein 1) (CortBP1). | |||||
|
TCRG1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 23) | NC score | 0.048070 (rank : 12) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
UBP31_HUMAN
|
||||||
| θ value | 2.36792 (rank : 24) | NC score | 0.020260 (rank : 50) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
AP180_MOUSE
|
||||||
| θ value | 3.0926 (rank : 25) | NC score | 0.039273 (rank : 17) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61548, Q61547 | Gene names | Snap91 | |||
|
Domain Architecture |
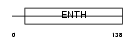
|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein) (Phosphoprotein F1- 20). | |||||
|
E2AK4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 26) | NC score | 0.006473 (rank : 71) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1012 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9P2K8, Q69YL7, Q6DC97, Q96GN6, Q9H5K1, Q9NSQ3, Q9NSZ5, Q9UJ56 | Gene names | EIF2AK4, GCN2, KIAA1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein). | |||||
|
MUC1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.055684 (rank : 6) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
MYCL1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.024543 (rank : 42) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
PTN23_HUMAN
|
||||||
| θ value | 3.0926 (rank : 29) | NC score | 0.013794 (rank : 66) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
FCRLA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 30) | NC score | 0.021161 (rank : 49) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7L513, Q5VXA1, Q5VXA2, Q5VXA3, Q5VXB0, Q8NEW4, Q8WXH3, Q96PC6, Q96PJ0, Q96PJ1, Q96PJ2, Q96PJ4, Q9BR57 | Gene names | FCRLA, FCRL, FCRL1, FCRLM1, FCRX, FREB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fc receptor-like and mucin-like 1 precursor (Fc receptor-like A) (Fc receptor homolog expressed in B-cells) (Fc receptor-related protein X) (FcRX) (Fc receptor-like protein). | |||||
|
PHF2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 31) | NC score | 0.027095 (rank : 35) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PO121_HUMAN
|
||||||
| θ value | 4.03905 (rank : 32) | NC score | 0.034281 (rank : 23) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
PXK_MOUSE
|
||||||
| θ value | 4.03905 (rank : 33) | NC score | 0.022685 (rank : 47) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 320 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BX57, Q3TD60, Q3TEL7, Q3TZZ1, Q3U197, Q3U773, Q3UCS5, Q4FBH7, Q91WB6 | Gene names | Pxk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PX domain-containing protein kinase-like protein (Modulator of Na,K- ATPase) (MONaKA). | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.024749 (rank : 40) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.035968 (rank : 21) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
ZIMP7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.025417 (rank : 38) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NF64, O94790, Q659A8, Q6JKL5, Q8WTX8, Q96Q01, Q9BQH7 | Gene names | ZIMP7, KIAA1886 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
ZN653_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.002221 (rank : 72) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6YND2, Q8BHT8, Q8CI02 | Gene names | Znf653, Zfp653, Zip67 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 653 (67 kDa zinc finger protein) (Zinc finger protein Zip67). | |||||
|
ANKS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.008429 (rank : 70) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92625, Q5JYI9, Q5SYR2, Q86WQ7 | Gene names | ANKS1A, ANKS1, KIAA0229 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 1A (Odin). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.036948 (rank : 18) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
CCNK_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.036677 (rank : 20) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75909, Q96B63, Q9NNY9 | Gene names | CCNK, CPR4 | |||
|
Domain Architecture |
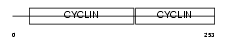
|
|||||
| Description | Cyclin-K. | |||||
|
DGKK_HUMAN
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.018911 (rank : 53) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.031568 (rank : 27) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
MUC13_HUMAN
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.032852 (rank : 25) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3R2, Q6UWD9, Q9NXT5 | Gene names | MUC13, DRCC1, RECC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-13 precursor (Down-regulated in colon cancer 1). | |||||
|
MUCDL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.024762 (rank : 39) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VHF2, Q8CEJ3, Q9D8I9 | Gene names | Mucdhl | |||
|
Domain Architecture |
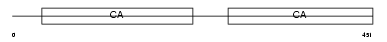
|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
PELP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.027633 (rank : 34) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
PHF2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.024654 (rank : 41) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
SETD2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.031019 (rank : 29) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
SHAN1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.018254 (rank : 54) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
X3CL1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.031023 (rank : 28) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P78423, O00672 | Gene names | CX3CL1, A-152E5.2, FKN, NTT, SCYD1 | |||
|
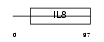
Domain Architecture |
|
|||||
| Description | Fractalkine precursor (CX3CL1) (Neurotactin) (CX3C membrane-anchored chemokine) (Small inducible cytokine D1). | |||||
|
ZFPL1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.030574 (rank : 31) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95159, O14616, Q9UID0 | Gene names | ZFPL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein-like 1 (Zinc finger protein MCG4). | |||||
|
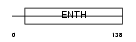
AP180_HUMAN
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.031924 (rank : 26) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60641, Q5JX13, Q68DL9, Q6P9D3, Q9NTY7 | Gene names | SNAP91, KIAA0656 | |||
|
Domain Architecture |
|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein). | |||||
|
CASC3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 52) | NC score | 0.030751 (rank : 30) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K3W3, Q8K219, Q99NF0 | Gene names | Casc3, Mln51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASC3 (Cancer susceptibility candidate gene 3 protein homolog) (Metastatic lymph node protein 51 homolog) (Protein MLN 51 homolog) (Protein barentsz) (Btz). | |||||
|
ENAH_MOUSE
|
||||||
| θ value | 6.88961 (rank : 53) | NC score | 0.015583 (rank : 62) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
EP300_HUMAN
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.024242 (rank : 43) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
GAH2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.022766 (rank : 46) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P63118 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-H_3q26 provirus ancestral Gag polyprotein (Gag polyprotein). | |||||
|
PHAR2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.017463 (rank : 55) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75167, Q68DM2 | Gene names | PHACTR2, C6orf56, KIAA0680 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatase and actin regulator 2. | |||||
|
PHC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.016445 (rank : 59) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QWH1, O88463, Q8K5D9 | Gene names | Phc2, Edr2, Ph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 2 (mPH2) (Early development regulatory protein 2) (p36). | |||||
|
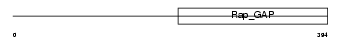
RGP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.017290 (rank : 57) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P47736 | Gene names | RAP1GAP, RAP1GA1 | |||
|
Domain Architecture |
|
|||||
| Description | Rap1 GTPase-activating protein 1 (Rap1GAP). | |||||
|
SFPQ_MOUSE
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.022215 (rank : 48) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
SMRC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.015605 (rank : 61) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
TF3C1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.019314 (rank : 52) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q12789, Q12838, Q9Y4W9 | Gene names | GTF3C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 1 (Transcription factor IIIC-subunit alpha) (TF3C-alpha) (TFIIIC 220 kDa subunit) (TFIIIC220) (TFIIIC box B-binding subunit). | |||||
|
WASF4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.016198 (rank : 60) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IV90 | Gene names | WASF4, SCAR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 4 (WASP-family protein member 4). | |||||
|
YETS2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.026702 (rank : 36) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q3TUF7, Q6PGF8, Q80TI2, Q8CG86 | Gene names | Yeats2, Kiaa1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.029024 (rank : 32) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BL1S3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.024115 (rank : 44) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6QNY0 | Gene names | BLOC1S3, BLOS3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Biogenesis of lysosome-related organelles complex-1 subunit 3 (BLOC subunit 3). | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.013735 (rank : 67) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
EAF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.017300 (rank : 56) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D4C5 | Gene names | Eaf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELL-associated factor 1. | |||||
|
ELF4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.019677 (rank : 51) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
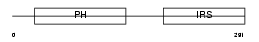
IRS2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.014222 (rank : 64) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y4H2, Q96RR2, Q9BZG0, Q9Y6I5 | Gene names | IRS2 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin receptor substrate 2 (IRS-2). | |||||
|
S3TC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.016579 (rank : 58) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TE82 | Gene names | SH3TC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 domain and tetratricopeptide repeats-containing protein 1. | |||||
|
SIX2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.009165 (rank : 69) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NPC8, Q9BXH7 | Gene names | SIX2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein SIX2 (Sine oculis homeobox homolog 2). | |||||
|
SLK_MOUSE
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.001782 (rank : 73) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1798 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O54988, Q80U65, Q8CAU2, Q8CDW2, Q9WU41 | Gene names | Slk, Kiaa0204, Stk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | STE20-like serine/threonine-protein kinase (EC 2.7.11.1) (STE20-like kinase) (STE20-related serine/threonine-protein kinase) (STE20-related kinase) (mSLK) (Serine/threonine-protein kinase 2) (STE20-related kinase SMAK) (Etk4). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.051483 (rank : 9) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
NFIP2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 2.35023e-141 (rank : 1) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NV92, Q7Z2H3, Q7Z428, Q8TAR3, Q9ULQ5 | Gene names | NDFIP2, KIAA1165, N4WBP5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD4 family-interacting protein 2 (NEDD4 WW domain-binding protein 5A) (Putative MAPK-activating protein PM04/PM05/PM06/PM07) (Putative NF-kappa-B-activating protein 413). | |||||
|
NFIP1_HUMAN
|
||||||
| NC score | 0.917261 (rank : 2) | θ value | 8.87376e-48 (rank : 2) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BT67, Q658T8, Q8N2E3, Q8N2F9 | Gene names | NDFIP1, N4WBP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD4 family-interacting protein 1 (NEDD4 WW domain-binding protein 5) (Putative MAPK-activating protein PM13) (Putative NF-kappa-B- activating protein 164) (Putative NFKB and MAPK-activating protein) (Breast cancer-associated protein SGA-1M). | |||||
|
NFIP1_MOUSE
|
||||||
| NC score | 0.915665 (rank : 3) | θ value | 1.67352e-46 (rank : 3) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R0W6, Q9EQH8 | Gene names | Ndfip1, N4wbp5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD4 family-interacting protein 1 (NEDD4 WW domain-binding protein 5). | |||||
|
SELPL_MOUSE
|
||||||
| NC score | 0.104228 (rank : 4) | θ value | 0.0330416 (rank : 4) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q62170 | Gene names | Selplg, Selp1, Selpl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | P-selectin glycoprotein ligand 1 precursor (PSGL-1) (Selectin P ligand). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.060521 (rank : 5) | θ value | 0.21417 (rank : 6) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
MUC1_MOUSE
|
||||||
| NC score | 0.055684 (rank : 6) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
NFRKB_MOUSE
|
||||||
| NC score | 0.054818 (rank : 7) | θ value | 1.38821 (rank : 13) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
TCRG1_HUMAN
|
||||||
| NC score | 0.054789 (rank : 8) | θ value | 0.21417 (rank : 7) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.051483 (rank : 9) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
CN155_HUMAN
|
||||||
| NC score | 0.049365 (rank : 10) | θ value | 0.813845 (rank : 10) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
MUC2_HUMAN
|
||||||
| NC score | 0.048936 (rank : 11) | θ value | 0.0431538 (rank : 5) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
TCRG1_MOUSE
|
||||||
| NC score | 0.048070 (rank : 12) | θ value | 2.36792 (rank : 23) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
RERE_MOUSE
|
||||||
| NC score | 0.044051 (rank : 13) | θ value | 0.62314 (rank : 9) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
LMO7_HUMAN
|
||||||
| NC score | 0.042776 (rank : 14) | θ value | 1.06291 (rank : 12) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.040629 (rank : 15) | θ value | 0.365318 (rank : 8) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
NC6IP_HUMAN
|
||||||
| NC score | 0.039538 (rank : 16) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96RS0, Q5GH23, Q8TDG9, Q96QU3, Q9H5V3 | Gene names | NCOA6IP, PIMT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA methyltransferase NCOA6IP (EC 2.1.1.-) (Nuclear receptor coactivator 6-interacting protein) (PRIP-interacting protein with methyltransferase motif) (PIPMT) (PIMT) (CLL-associated antigen KW-2) (Hepatocellular carcinoma-associated antigen 137) (HCA137). | |||||
|
AP180_MOUSE
|
||||||
| NC score | 0.039273 (rank : 17) | θ value | 3.0926 (rank : 25) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61548, Q61547 | Gene names | Snap91 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein) (Phosphoprotein F1- 20). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.036948 (rank : 18) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
CCNK_MOUSE
|
||||||
| NC score | 0.036922 (rank : 19) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
CCNK_HUMAN
|
||||||
| NC score | 0.036677 (rank : 20) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75909, Q96B63, Q9NNY9 | Gene names | CCNK, CPR4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-K. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.035968 (rank : 21) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
SHAN2_MOUSE
|
||||||
| NC score | 0.035603 (rank : 22) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 446 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80Z38, Q3UTK4, Q5DU07 | Gene names | Shank2, Kiaa1022 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2) (Cortactin- binding protein 1) (CortBP1). | |||||
|
PO121_HUMAN
|
||||||
| NC score | 0.034281 (rank : 23) | θ value | 4.03905 (rank : 32) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
BCL7B_HUMAN
|
||||||
| NC score | 0.032947 (rank : 24) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BQE9, O43769, Q13845, Q6ZW75 | Gene names | BCL7B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell CLL/lymphoma 7 protein family member B. | |||||
|
MUC13_HUMAN
|
||||||
| NC score | 0.032852 (rank : 25) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3R2, Q6UWD9, Q9NXT5 | Gene names | MUC13, DRCC1, RECC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-13 precursor (Down-regulated in colon cancer 1). | |||||
|
AP180_HUMAN
|
||||||
| NC score | 0.031924 (rank : 26) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60641, Q5JX13, Q68DL9, Q6P9D3, Q9NTY7 | Gene names | SNAP91, KIAA0656 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.031568 (rank : 27) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
X3CL1_HUMAN
|
||||||
| NC score | 0.031023 (rank : 28) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P78423, O00672 | Gene names | CX3CL1, A-152E5.2, FKN, NTT, SCYD1 | |||
|
Domain Architecture |

|
|||||
| Description | Fractalkine precursor (CX3CL1) (Neurotactin) (CX3C membrane-anchored chemokine) (Small inducible cytokine D1). | |||||
|
SETD2_HUMAN
|
||||||
| NC score | 0.031019 (rank : 29) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
CASC3_MOUSE
|
||||||
| NC score | 0.030751 (rank : 30) | θ value | 6.88961 (rank : 52) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K3W3, Q8K219, Q99NF0 | Gene names | Casc3, Mln51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASC3 (Cancer susceptibility candidate gene 3 protein homolog) (Metastatic lymph node protein 51 homolog) (Protein MLN 51 homolog) (Protein barentsz) (Btz). | |||||
|
ZFPL1_HUMAN
|
||||||
| NC score | 0.030574 (rank : 31) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95159, O14616, Q9UID0 | Gene names | ZFPL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein-like 1 (Zinc finger protein MCG4). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.029024 (rank : 32) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CUTL2_MOUSE
|
||||||
| NC score | 0.029008 (rank : 33) | θ value | 0.813845 (rank : 11) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 771 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70298 | Gene names | Cutl2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein cut-like 2 (Homeobox protein Cux-2) (Cut-like 2). | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.027633 (rank : 34) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
PHF2_MOUSE
|
||||||
| NC score | 0.027095 (rank : 35) | θ value | 4.03905 (rank : 31) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
YETS2_MOUSE
|
||||||
| NC score | 0.026702 (rank : 36) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q3TUF7, Q6PGF8, Q80TI2, Q8CG86 | Gene names | Yeats2, Kiaa1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
SFPQ_HUMAN
|
||||||
| NC score | 0.026643 (rank : 37) | θ value | 2.36792 (rank : 21) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
ZIMP7_HUMAN
|
||||||
| NC score | 0.025417 (rank : 38) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NF64, O94790, Q659A8, Q6JKL5, Q8WTX8, Q96Q01, Q9BQH7 | Gene names | ZIMP7, KIAA1886 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
MUCDL_MOUSE
|
||||||
| NC score | 0.024762 (rank : 39) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VHF2, Q8CEJ3, Q9D8I9 | Gene names | Mucdhl | |||
|
Domain Architecture |

|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.024749 (rank : 40) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
PHF2_HUMAN
|
||||||
| NC score | 0.024654 (rank : 41) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
MYCL1_MOUSE
|
||||||
| NC score | 0.024543 (rank : 42) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.024242 (rank : 43) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
BL1S3_HUMAN
|
||||||
| NC score | 0.024115 (rank : 44) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6QNY0 | Gene names | BLOC1S3, BLOS3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Biogenesis of lysosome-related organelles complex-1 subunit 3 (BLOC subunit 3). | |||||
|
NHERF_MOUSE
|
||||||
| NC score | 0.023864 (rank : 45) | θ value | 1.81305 (rank : 17) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70441 | Gene names | Slc9a3r1, Nherf | |||
|
Domain Architecture |

|
|||||
| Description | Ezrin-radixin-moesin-binding phosphoprotein 50 (EBP50) (Na(+)/H(+) exchange regulatory cofactor NHE-RF) (NHERF-1) (Regulatory cofactor of Na(+)/H(+) exchanger) (Sodium-hydrogen exchanger regulatory factor 1) (Solute carrier family 9 isoform 3 regulatory factor 1). | |||||
|
GAH2_HUMAN
|
||||||
| NC score | 0.022766 (rank : 46) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P63118 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-H_3q26 provirus ancestral Gag polyprotein (Gag polyprotein). | |||||
|
PXK_MOUSE
|
||||||
| NC score | 0.022685 (rank : 47) | θ value | 4.03905 (rank : 33) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 320 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BX57, Q3TD60, Q3TEL7, Q3TZZ1, Q3U197, Q3U773, Q3UCS5, Q4FBH7, Q91WB6 | Gene names | Pxk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PX domain-containing protein kinase-like protein (Modulator of Na,K- ATPase) (MONaKA). | |||||
|
SFPQ_MOUSE
|
||||||
| NC score | 0.022215 (rank : 48) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
FCRLA_HUMAN
|
||||||
| NC score | 0.021161 (rank : 49) | θ value | 4.03905 (rank : 30) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7L513, Q5VXA1, Q5VXA2, Q5VXA3, Q5VXB0, Q8NEW4, Q8WXH3, Q96PC6, Q96PJ0, Q96PJ1, Q96PJ2, Q96PJ4, Q9BR57 | Gene names | FCRLA, FCRL, FCRL1, FCRLM1, FCRX, FREB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fc receptor-like and mucin-like 1 precursor (Fc receptor-like A) (Fc receptor homolog expressed in B-cells) (Fc receptor-related protein X) (FcRX) (Fc receptor-like protein). | |||||
|
UBP31_HUMAN
|
||||||
| NC score | 0.020260 (rank : 50) | θ value | 2.36792 (rank : 24) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
ELF4_MOUSE
|
||||||
| NC score | 0.019677 (rank : 51) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
TF3C1_HUMAN
|
||||||
| NC score | 0.019314 (rank : 52) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q12789, Q12838, Q9Y4W9 | Gene names | GTF3C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 1 (Transcription factor IIIC-subunit alpha) (TF3C-alpha) (TFIIIC 220 kDa subunit) (TFIIIC220) (TFIIIC box B-binding subunit). | |||||
|
DGKK_HUMAN
|
||||||
| NC score | 0.018911 (rank : 53) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
SHAN1_HUMAN
|
||||||
| NC score | 0.018254 (rank : 54) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
PHAR2_HUMAN
|
||||||
| NC score | 0.017463 (rank : 55) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75167, Q68DM2 | Gene names | PHACTR2, C6orf56, KIAA0680 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatase and actin regulator 2. | |||||
|
EAF1_MOUSE
|
||||||
| NC score | 0.017300 (rank : 56) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D4C5 | Gene names | Eaf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELL-associated factor 1. | |||||
|
RGP2_HUMAN
|
||||||
| NC score | 0.017290 (rank : 57) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P47736 | Gene names | RAP1GAP, RAP1GA1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-activating protein 1 (Rap1GAP). | |||||
|
S3TC1_HUMAN
|
||||||
| NC score | 0.016579 (rank : 58) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TE82 | Gene names | SH3TC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 domain and tetratricopeptide repeats-containing protein 1. | |||||
|
PHC2_MOUSE
|
||||||
| NC score | 0.016445 (rank : 59) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QWH1, O88463, Q8K5D9 | Gene names | Phc2, Edr2, Ph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 2 (mPH2) (Early development regulatory protein 2) (p36). | |||||
|
WASF4_HUMAN
|
||||||
| NC score | 0.016198 (rank : 60) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IV90 | Gene names | WASF4, SCAR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 4 (WASP-family protein member 4). | |||||
|
SMRC2_MOUSE
|
||||||
| NC score | 0.015605 (rank : 61) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.015583 (rank : 62) | θ value | 6.88961 (rank : 53) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
NID2_HUMAN
|
||||||
| NC score | 0.014841 (rank : 63) | θ value | 2.36792 (rank : 20) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q14112, O43710 | Gene names | NID2 | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-2 precursor (NID-2) (Osteonidogen). | |||||
|
IRS2_HUMAN
|
||||||
| NC score | 0.014222 (rank : 64) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y4H2, Q96RR2, Q9BZG0, Q9Y6I5 | Gene names | IRS2 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin receptor substrate 2 (IRS-2). | |||||
|
CSPG3_MOUSE
|
||||||
| NC score | 0.013975 (rank : 65) | θ value | 2.36792 (rank : 19) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P55066, Q6P1E3, Q8C4F8 | Gene names | Cspg3, Ncan | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
PTN23_HUMAN
|
||||||
| NC score | 0.013794 (rank : 66) | θ value | 3.0926 (rank : 29) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.013735 (rank : 67) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
RXRB_HUMAN
|
||||||
| NC score | 0.011463 (rank : 68) | θ value | 1.38821 (rank : 14) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P28702, P28703 | Gene names | RXRB, NR2B2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoic acid receptor RXR-beta (Retinoid X receptor beta). | |||||
|
SIX2_HUMAN
|
||||||
| NC score | 0.009165 (rank : 69) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NPC8, Q9BXH7 | Gene names | SIX2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein SIX2 (Sine oculis homeobox homolog 2). | |||||
|
ANKS1_HUMAN
|
||||||
| NC score | 0.008429 (rank : 70) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92625, Q5JYI9, Q5SYR2, Q86WQ7 | Gene names | ANKS1A, ANKS1, KIAA0229 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 1A (Odin). | |||||
|
E2AK4_HUMAN
|
||||||
| NC score | 0.006473 (rank : 71) | θ value | 3.0926 (rank : 26) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1012 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9P2K8, Q69YL7, Q6DC97, Q96GN6, Q9H5K1, Q9NSQ3, Q9NSZ5, Q9UJ56 | Gene names | EIF2AK4, GCN2, KIAA1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein). | |||||
|
ZN653_MOUSE
|
||||||
| NC score | 0.002221 (rank : 72) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6YND2, Q8BHT8, Q8CI02 | Gene names | Znf653, Zfp653, Zip67 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 653 (67 kDa zinc finger protein) (Zinc finger protein Zip67). | |||||
|
SLK_MOUSE
|
||||||
| NC score | 0.001782 (rank : 73) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1798 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O54988, Q80U65, Q8CAU2, Q8CDW2, Q9WU41 | Gene names | Slk, Kiaa0204, Stk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | STE20-like serine/threonine-protein kinase (EC 2.7.11.1) (STE20-like kinase) (STE20-related serine/threonine-protein kinase) (STE20-related kinase) (mSLK) (Serine/threonine-protein kinase 2) (STE20-related kinase SMAK) (Etk4). | |||||