Please be patient as the page loads
|
RGP2_HUMAN
|
||||||
| SwissProt Accessions | P47736 | Gene names | RAP1GAP, RAP1GA1 | |||
|
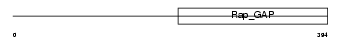
Domain Architecture |
|
|||||
| Description | Rap1 GTPase-activating protein 1 (Rap1GAP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
RGP2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P47736 | Gene names | RAP1GAP, RAP1GA1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-activating protein 1 (Rap1GAP). | |||||
|
SI1L3_HUMAN
|
||||||
| θ value | 1.45929e-58 (rank : 2) | NC score | 0.874658 (rank : 8) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60292, Q2TV87 | Gene names | SIPA1L3, KIAA0545, SPAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 3 (SPA-1-like protein 3). | |||||
|
SI1L1_MOUSE
|
||||||
| θ value | 5.54529e-58 (rank : 3) | NC score | 0.882084 (rank : 6) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8C0T5, Q6PDI8, Q80U02, Q8C026 | Gene names | Sipa1l1, Kiaa0440 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 1. | |||||
|
SI1L1_HUMAN
|
||||||
| θ value | 9.45883e-58 (rank : 4) | NC score | 0.883970 (rank : 5) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43166, O95321, Q9UDU4, Q9UNU4 | Gene names | SIPA1L1, E6TP1, KIAA0440 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 1 (High-risk human papilloma viruses E6 oncoproteins targeted protein 1) (E6- targeted protein 1). | |||||
|
SI1L2_HUMAN
|
||||||
| θ value | 1.61343e-57 (rank : 5) | NC score | 0.884649 (rank : 4) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9P2F8, Q5VXR7, Q5VXR8, Q641Q4, Q8NA38, Q96DZ3, Q9H9F6 | Gene names | SIPA1L2, KIAA1389 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 2. | |||||
|
SI1L2_MOUSE
|
||||||
| θ value | 2.32979e-56 (rank : 6) | NC score | 0.885562 (rank : 3) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80TE4, Q6PDY1 | Gene names | Sipa1l2, Kiaa0545 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 2. | |||||
|
SIPA1_MOUSE
|
||||||
| θ value | 6.77864e-56 (rank : 7) | NC score | 0.894291 (rank : 2) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P46062, P70204 | Gene names | Sipa1, Spa-1, Spa1 | |||
|
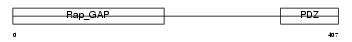
Domain Architecture |
|
|||||
| Description | Signal-induced proliferation-associated protein 1 (Sipa-1) (GTPase- activating protein Spa-1). | |||||
|
SIPA1_HUMAN
|
||||||
| θ value | 2.84797e-54 (rank : 8) | NC score | 0.880920 (rank : 7) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96FS4, O14518, O60484, O60618 | Gene names | SIPA1, SPA1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal-induced proliferation-associated protein 1 (Sipa-1) (GTPase- activating protein Spa-1) (p130 SPA-1). | |||||
|
GRIPE_HUMAN
|
||||||
| θ value | 4.91457e-14 (rank : 9) | NC score | 0.542469 (rank : 9) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6GYQ0, O94960, Q6GYP9, Q6ZT23, Q86YF3, Q86YF5, Q8ND69 | Gene names | GARNL1, KIAA0884, TULIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTPase-activating Rap/Ran-GAP domain-like 1 (GAP-related-interacting partner to E12) (GRIPE) (Tuberin-like protein 1). | |||||
|
GRIPE_MOUSE
|
||||||
| θ value | 1.42992e-13 (rank : 10) | NC score | 0.533043 (rank : 10) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6GYP7, Q6GYP6, Q6ZQ28, Q8BNJ5, Q8C9G8, Q8CIW4, Q9JMC4 | Gene names | Garnl1, Kiaa0884, Tulip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTPase-activating RapGAP domain-like 1 (GAP-related-interacting partner to E12) (GRIPE) (Tuberin-like protein 1). | |||||
|
TSC2_HUMAN
|
||||||
| θ value | 7.09661e-13 (rank : 11) | NC score | 0.530450 (rank : 11) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49815, O75275, Q8TAZ1 | Gene names | TSC2 | |||
|
Domain Architecture |

|
|||||
| Description | Tuberin (Tuberous sclerosis 2 protein). | |||||
|
TSC2_MOUSE
|
||||||
| θ value | 1.2105e-12 (rank : 12) | NC score | 0.512755 (rank : 12) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61037, P97723, P97724, P97725, P97727, Q61007, Q61008, Q9WUF6 | Gene names | Tsc2 | |||
|
Domain Architecture |

|
|||||
| Description | Tuberin (Tuberous sclerosis 2 homolog protein). | |||||
|
ASXL1_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 13) | NC score | 0.069635 (rank : 13) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IXJ9, Q5JWS9, Q8IYY7, Q9H466, Q9NQF8, Q9UFJ0, Q9UFP8, Q9Y2I4 | Gene names | ASXL1, KIAA0978 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative Polycomb group protein ASXL1 (Additional sex combs-like protein 1). | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 14) | NC score | 0.047529 (rank : 21) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
MUC5B_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 15) | NC score | 0.025623 (rank : 30) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
NC6IP_HUMAN
|
||||||
| θ value | 0.125558 (rank : 16) | NC score | 0.048827 (rank : 20) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96RS0, Q5GH23, Q8TDG9, Q96QU3, Q9H5V3 | Gene names | NCOA6IP, PIMT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA methyltransferase NCOA6IP (EC 2.1.1.-) (Nuclear receptor coactivator 6-interacting protein) (PRIP-interacting protein with methyltransferase motif) (PIPMT) (PIMT) (CLL-associated antigen KW-2) (Hepatocellular carcinoma-associated antigen 137) (HCA137). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 17) | NC score | 0.057034 (rank : 17) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
BLM_HUMAN
|
||||||
| θ value | 0.365318 (rank : 18) | NC score | 0.024281 (rank : 34) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P54132 | Gene names | BLM, RECQ2, RECQL3 | |||
|
Domain Architecture |

|
|||||
| Description | Bloom syndrome protein (EC 3.6.1.-) (RecQ protein-like 3) (DNA helicase, RecQ-like type 2). | |||||
|
CAC1G_HUMAN
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.018434 (rank : 39) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
PGCA_MOUSE
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.017376 (rank : 41) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
MARK2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.004000 (rank : 60) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q05512, Q6PDR4, Q8BR95 | Gene names | Mark2, Emk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL Motif Kinase) (EMK1). | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.005547 (rank : 56) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
SALL1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.004499 (rank : 57) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NSC2, Q99881, Q9NSC3, Q9P1R0 | Gene names | SALL1, SAL1 | |||
|
Domain Architecture |

|
|||||
| Description | Sal-like protein 1 (Zinc finger protein SALL1) (Spalt-like transcription factor 1) (HSal1). | |||||
|
TCF20_MOUSE
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.030670 (rank : 27) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
GPSM2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.025108 (rank : 32) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P81274, Q8N0Z5 | Gene names | GPSM2, LGN | |||
|
Domain Architecture |

|
|||||
| Description | G-protein signaling modulator 2 (Mosaic protein LGN). | |||||
|
KI67_HUMAN
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.036574 (rank : 24) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
CJ012_HUMAN
|
||||||
| θ value | 1.81305 (rank : 27) | NC score | 0.034902 (rank : 25) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
IF16_HUMAN
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.023343 (rank : 36) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q16666, Q9UH78 | Gene names | IFI16, IFNGIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-interferon-inducible protein Ifi-16 (Interferon-inducible myeloid differentiation transcriptional activator) (IFI 16). | |||||
|
MAP4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.030796 (rank : 26) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
PHF22_HUMAN
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.028558 (rank : 28) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96CB8, Q9HD71 | Gene names | INTS12, PHF22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.037513 (rank : 23) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
PHLPP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.011741 (rank : 48) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
4ET_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.020420 (rank : 38) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.025410 (rank : 31) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
MILK2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.013562 (rank : 46) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
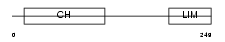
Domain Architecture |
|
|||||
| Description | MICAL-like protein 2. | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.047016 (rank : 22) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TARA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.024089 (rank : 35) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
ATBF1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.010043 (rank : 53) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
ATBF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.009989 (rank : 54) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
AXN1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.011589 (rank : 49) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15169, Q4TT26, Q4TT27, Q86YA7, Q8WVW6, Q96S28 | Gene names | AXIN1, AXIN | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (hAxin). | |||||
|
DPOLZ_HUMAN
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.014581 (rank : 44) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
MELK_MOUSE
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.001457 (rank : 64) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61846, Q61804, Q6ZQH6 | Gene names | Melk, Kiaa0175, Pk38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Maternal embryonic leucine zipper kinase (EC 2.7.11.1) (Protein kinase PK38) (mPK38). | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.027248 (rank : 29) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
PHLPP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.011231 (rank : 50) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CHE4, Q6PCN0, Q8QZU8, Q8R5E5, Q99KL6 | Gene names | Phlpp, Kiaa0606, Plekhe1, Scop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein). | |||||
|
PI51C_HUMAN
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.010950 (rank : 52) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60331, Q7LE07 | Gene names | PIP5K1C, KIAA0589 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 5-kinase type-1 gamma (EC 2.7.1.68) (Phosphatidylinositol-4-phosphate 5-kinase type I gamma) (PtdIns(4)P- 5-kinase gamma) (PtdInsPKIgamma) (PIP5KIgamma). | |||||
|
TCOF_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.024999 (rank : 33) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
HXA2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.003718 (rank : 61) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43364 | Gene names | HOXA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein Hox-A2. | |||||
|
MYST4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.014784 (rank : 43) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
T240L_MOUSE
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.013810 (rank : 45) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6JPI3, Q3TRF4, Q3UQI8, Q80TM3, Q80WQ0 | Gene names | Thrap2, Kiaa1025, Trap240l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 2 (Thyroid hormone receptor-associated protein complex 240 kDa component-like). | |||||
|
ATX2L_MOUSE
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.011032 (rank : 51) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7TQH0, Q80XN9, Q8K059 | Gene names | Atxn2l, A2lp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein. | |||||
|
CAD23_MOUSE
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.002029 (rank : 63) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99PF4, Q99NH1, Q9D4N9 | Gene names | Cdh23 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-23 precursor (Otocadherin). | |||||
|
K0310_HUMAN
|
||||||
| θ value | 6.88961 (rank : 52) | NC score | 0.022180 (rank : 37) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
NFIP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 53) | NC score | 0.017290 (rank : 42) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NV92, Q7Z2H3, Q7Z428, Q8TAR3, Q9ULQ5 | Gene names | NDFIP2, KIAA1165, N4WBP5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD4 family-interacting protein 2 (NEDD4 WW domain-binding protein 5A) (Putative MAPK-activating protein PM04/PM05/PM06/PM07) (Putative NF-kappa-B-activating protein 413). | |||||
|
NRK_HUMAN
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.001111 (rank : 65) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7Z2Y5, Q32ND6, Q5H9K2, Q6ZMP2 | Gene names | NRK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nik-related protein kinase (EC 2.7.11.1). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.012470 (rank : 47) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
DDX51_HUMAN
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.005947 (rank : 55) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
GPSM2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.018079 (rank : 40) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VDU0, Q8BLX3 | Gene names | Gpsm2, Lgn, Pins | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein signaling modulator 2 (Pins homolog). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.004235 (rank : 59) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
SRPK1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.004263 (rank : 58) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96SB4, Q12890, Q5R364, Q5R365, Q8IY12 | Gene names | SRPK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK1 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 1) (SR-protein-specific kinase 1) (SFRS protein kinase 1). | |||||
|
ZEP2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.002923 (rank : 62) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3UHF7, O55140, Q3UVD4, Q3UVH5 | Gene names | Hivep2, Mibp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 homolog (Myc intron-binding protein 1) (MIBP-1). | |||||
|
APBA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.058182 (rank : 16) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99767, O60571 | Gene names | APBA2, MINT2 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 2 (Neuron- specific X11L protein) (Neuronal Munc18-1-interacting protein 2) (Mint-2) (Adapter protein X11beta). | |||||
|
APBA2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.058526 (rank : 15) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P98084, Q6PAJ2 | Gene names | Apba2, X11l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 2 (Neuron- specific X11L protein) (Neuronal Munc18-1-interacting protein 2) (Mint-2) (Adapter protein X11beta). | |||||
|
APBA3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.056976 (rank : 18) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O88888 | Gene names | Apba3, Mint3 | |||
|
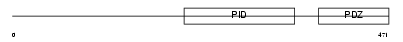
Domain Architecture |
|
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 3 (Neuron- specific X11L2 protein) (Neuronal Munc18-1-interacting protein 3) (Mint-3) (Adapter protein X11gamma). | |||||
|
SHAN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.063012 (rank : 14) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UPX8, Q3Y8G9, Q52LK2, Q9UKP1 | Gene names | SHANK2, KIAA1022 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2). | |||||
|
SHAN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.053860 (rank : 19) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 446 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80Z38, Q3UTK4, Q5DU07 | Gene names | Shank2, Kiaa1022 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2) (Cortactin- binding protein 1) (CortBP1). | |||||
|
RGP2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P47736 | Gene names | RAP1GAP, RAP1GA1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-activating protein 1 (Rap1GAP). | |||||
|
SIPA1_MOUSE
|
||||||
| NC score | 0.894291 (rank : 2) | θ value | 6.77864e-56 (rank : 7) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P46062, P70204 | Gene names | Sipa1, Spa-1, Spa1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal-induced proliferation-associated protein 1 (Sipa-1) (GTPase- activating protein Spa-1). | |||||
|
SI1L2_MOUSE
|
||||||
| NC score | 0.885562 (rank : 3) | θ value | 2.32979e-56 (rank : 6) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80TE4, Q6PDY1 | Gene names | Sipa1l2, Kiaa0545 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 2. | |||||
|
SI1L2_HUMAN
|
||||||
| NC score | 0.884649 (rank : 4) | θ value | 1.61343e-57 (rank : 5) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9P2F8, Q5VXR7, Q5VXR8, Q641Q4, Q8NA38, Q96DZ3, Q9H9F6 | Gene names | SIPA1L2, KIAA1389 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 2. | |||||
|
SI1L1_HUMAN
|
||||||
| NC score | 0.883970 (rank : 5) | θ value | 9.45883e-58 (rank : 4) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43166, O95321, Q9UDU4, Q9UNU4 | Gene names | SIPA1L1, E6TP1, KIAA0440 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 1 (High-risk human papilloma viruses E6 oncoproteins targeted protein 1) (E6- targeted protein 1). | |||||
|
SI1L1_MOUSE
|
||||||
| NC score | 0.882084 (rank : 6) | θ value | 5.54529e-58 (rank : 3) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8C0T5, Q6PDI8, Q80U02, Q8C026 | Gene names | Sipa1l1, Kiaa0440 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 1. | |||||
|
SIPA1_HUMAN
|
||||||
| NC score | 0.880920 (rank : 7) | θ value | 2.84797e-54 (rank : 8) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96FS4, O14518, O60484, O60618 | Gene names | SIPA1, SPA1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal-induced proliferation-associated protein 1 (Sipa-1) (GTPase- activating protein Spa-1) (p130 SPA-1). | |||||
|
SI1L3_HUMAN
|
||||||
| NC score | 0.874658 (rank : 8) | θ value | 1.45929e-58 (rank : 2) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60292, Q2TV87 | Gene names | SIPA1L3, KIAA0545, SPAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 3 (SPA-1-like protein 3). | |||||
|
GRIPE_HUMAN
|
||||||
| NC score | 0.542469 (rank : 9) | θ value | 4.91457e-14 (rank : 9) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6GYQ0, O94960, Q6GYP9, Q6ZT23, Q86YF3, Q86YF5, Q8ND69 | Gene names | GARNL1, KIAA0884, TULIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTPase-activating Rap/Ran-GAP domain-like 1 (GAP-related-interacting partner to E12) (GRIPE) (Tuberin-like protein 1). | |||||
|
GRIPE_MOUSE
|
||||||
| NC score | 0.533043 (rank : 10) | θ value | 1.42992e-13 (rank : 10) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6GYP7, Q6GYP6, Q6ZQ28, Q8BNJ5, Q8C9G8, Q8CIW4, Q9JMC4 | Gene names | Garnl1, Kiaa0884, Tulip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTPase-activating RapGAP domain-like 1 (GAP-related-interacting partner to E12) (GRIPE) (Tuberin-like protein 1). | |||||
|
TSC2_HUMAN
|
||||||
| NC score | 0.530450 (rank : 11) | θ value | 7.09661e-13 (rank : 11) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49815, O75275, Q8TAZ1 | Gene names | TSC2 | |||
|
Domain Architecture |

|
|||||
| Description | Tuberin (Tuberous sclerosis 2 protein). | |||||
|
TSC2_MOUSE
|
||||||
| NC score | 0.512755 (rank : 12) | θ value | 1.2105e-12 (rank : 12) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61037, P97723, P97724, P97725, P97727, Q61007, Q61008, Q9WUF6 | Gene names | Tsc2 | |||
|
Domain Architecture |

|
|||||
| Description | Tuberin (Tuberous sclerosis 2 homolog protein). | |||||
|
ASXL1_HUMAN
|
||||||
| NC score | 0.069635 (rank : 13) | θ value | 0.00509761 (rank : 13) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IXJ9, Q5JWS9, Q8IYY7, Q9H466, Q9NQF8, Q9UFJ0, Q9UFP8, Q9Y2I4 | Gene names | ASXL1, KIAA0978 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative Polycomb group protein ASXL1 (Additional sex combs-like protein 1). | |||||
|
SHAN2_HUMAN
|
||||||
| NC score | 0.063012 (rank : 14) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UPX8, Q3Y8G9, Q52LK2, Q9UKP1 | Gene names | SHANK2, KIAA1022 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2). | |||||
|
APBA2_MOUSE
|
||||||
| NC score | 0.058526 (rank : 15) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P98084, Q6PAJ2 | Gene names | Apba2, X11l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 2 (Neuron- specific X11L protein) (Neuronal Munc18-1-interacting protein 2) (Mint-2) (Adapter protein X11beta). | |||||
|
APBA2_HUMAN
|
||||||
| NC score | 0.058182 (rank : 16) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99767, O60571 | Gene names | APBA2, MINT2 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 2 (Neuron- specific X11L protein) (Neuronal Munc18-1-interacting protein 2) (Mint-2) (Adapter protein X11beta). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.057034 (rank : 17) | θ value | 0.125558 (rank : 17) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
APBA3_MOUSE
|
||||||
| NC score | 0.056976 (rank : 18) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O88888 | Gene names | Apba3, Mint3 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 3 (Neuron- specific X11L2 protein) (Neuronal Munc18-1-interacting protein 3) (Mint-3) (Adapter protein X11gamma). | |||||
|
SHAN2_MOUSE
|
||||||
| NC score | 0.053860 (rank : 19) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 446 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80Z38, Q3UTK4, Q5DU07 | Gene names | Shank2, Kiaa1022 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2) (Cortactin- binding protein 1) (CortBP1). | |||||
|
NC6IP_HUMAN
|
||||||
| NC score | 0.048827 (rank : 20) | θ value | 0.125558 (rank : 16) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96RS0, Q5GH23, Q8TDG9, Q96QU3, Q9H5V3 | Gene names | NCOA6IP, PIMT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA methyltransferase NCOA6IP (EC 2.1.1.-) (Nuclear receptor coactivator 6-interacting protein) (PRIP-interacting protein with methyltransferase motif) (PIPMT) (PIMT) (CLL-associated antigen KW-2) (Hepatocellular carcinoma-associated antigen 137) (HCA137). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.047529 (rank : 21) | θ value | 0.0330416 (rank : 14) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.047016 (rank : 22) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.037513 (rank : 23) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.036574 (rank : 24) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.034902 (rank : 25) | θ value | 1.81305 (rank : 27) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
MAP4_MOUSE
|
||||||
| NC score | 0.030796 (rank : 26) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
TCF20_MOUSE
|
||||||
| NC score | 0.030670 (rank : 27) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
PHF22_HUMAN
|
||||||
| NC score | 0.028558 (rank : 28) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96CB8, Q9HD71 | Gene names | INTS12, PHF22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.027248 (rank : 29) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
MUC5B_HUMAN
|
||||||
| NC score | 0.025623 (rank : 30) | θ value | 0.0961366 (rank : 15) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.025410 (rank : 31) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
GPSM2_HUMAN
|
||||||
| NC score | 0.025108 (rank : 32) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P81274, Q8N0Z5 | Gene names | GPSM2, LGN | |||
|
Domain Architecture |

|
|||||
| Description | G-protein signaling modulator 2 (Mosaic protein LGN). | |||||
|
TCOF_MOUSE
|
||||||
| NC score | 0.024999 (rank : 33) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
BLM_HUMAN
|
||||||
| NC score | 0.024281 (rank : 34) | θ value | 0.365318 (rank : 18) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P54132 | Gene names | BLM, RECQ2, RECQL3 | |||
|
Domain Architecture |

|
|||||
| Description | Bloom syndrome protein (EC 3.6.1.-) (RecQ protein-like 3) (DNA helicase, RecQ-like type 2). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.024089 (rank : 35) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
IF16_HUMAN
|
||||||
| NC score | 0.023343 (rank : 36) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q16666, Q9UH78 | Gene names | IFI16, IFNGIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-interferon-inducible protein Ifi-16 (Interferon-inducible myeloid differentiation transcriptional activator) (IFI 16). | |||||
|
K0310_HUMAN
|
||||||
| NC score | 0.022180 (rank : 37) | θ value | 6.88961 (rank : 52) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
4ET_HUMAN
|
||||||
| NC score | 0.020420 (rank : 38) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
CAC1G_HUMAN
|
||||||
| NC score | 0.018434 (rank : 39) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
GPSM2_MOUSE
|
||||||
| NC score | 0.018079 (rank : 40) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VDU0, Q8BLX3 | Gene names | Gpsm2, Lgn, Pins | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein signaling modulator 2 (Pins homolog). | |||||
|
PGCA_MOUSE
|
||||||
| NC score | 0.017376 (rank : 41) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
NFIP2_HUMAN
|
||||||
| NC score | 0.017290 (rank : 42) | θ value | 6.88961 (rank : 53) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NV92, Q7Z2H3, Q7Z428, Q8TAR3, Q9ULQ5 | Gene names | NDFIP2, KIAA1165, N4WBP5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD4 family-interacting protein 2 (NEDD4 WW domain-binding protein 5A) (Putative MAPK-activating protein PM04/PM05/PM06/PM07) (Putative NF-kappa-B-activating protein 413). | |||||
|
MYST4_HUMAN
|
||||||
| NC score | 0.014784 (rank : 43) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
DPOLZ_HUMAN
|
||||||
| NC score | 0.014581 (rank : 44) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
T240L_MOUSE
|
||||||
| NC score | 0.013810 (rank : 45) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6JPI3, Q3TRF4, Q3UQI8, Q80TM3, Q80WQ0 | Gene names | Thrap2, Kiaa1025, Trap240l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 2 (Thyroid hormone receptor-associated protein complex 240 kDa component-like). | |||||
|
MILK2_HUMAN
|
||||||
| NC score | 0.013562 (rank : 46) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | MICAL-like protein 2. | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.012470 (rank : 47) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
PHLPP_HUMAN
|
||||||
| NC score | 0.011741 (rank : 48) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
AXN1_HUMAN
|
||||||
| NC score | 0.011589 (rank : 49) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15169, Q4TT26, Q4TT27, Q86YA7, Q8WVW6, Q96S28 | Gene names | AXIN1, AXIN | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (hAxin). | |||||
|
PHLPP_MOUSE
|
||||||
| NC score | 0.011231 (rank : 50) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CHE4, Q6PCN0, Q8QZU8, Q8R5E5, Q99KL6 | Gene names | Phlpp, Kiaa0606, Plekhe1, Scop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein). | |||||
|
ATX2L_MOUSE
|
||||||
| NC score | 0.011032 (rank : 51) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7TQH0, Q80XN9, Q8K059 | Gene names | Atxn2l, A2lp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein. | |||||
|
PI51C_HUMAN
|
||||||
| NC score | 0.010950 (rank : 52) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60331, Q7LE07 | Gene names | PIP5K1C, KIAA0589 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 5-kinase type-1 gamma (EC 2.7.1.68) (Phosphatidylinositol-4-phosphate 5-kinase type I gamma) (PtdIns(4)P- 5-kinase gamma) (PtdInsPKIgamma) (PIP5KIgamma). | |||||
|
ATBF1_HUMAN
|
||||||
| NC score | 0.010043 (rank : 53) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
ATBF1_MOUSE
|
||||||
| NC score | 0.009989 (rank : 54) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
DDX51_HUMAN
|
||||||
| NC score | 0.005947 (rank : 55) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | 0.005547 (rank : 56) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
SALL1_HUMAN
|
||||||
| NC score | 0.004499 (rank : 57) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NSC2, Q99881, Q9NSC3, Q9P1R0 | Gene names | SALL1, SAL1 | |||
|
Domain Architecture |

|
|||||
| Description | Sal-like protein 1 (Zinc finger protein SALL1) (Spalt-like transcription factor 1) (HSal1). | |||||
|
SRPK1_HUMAN
|
||||||
| NC score | 0.004263 (rank : 58) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96SB4, Q12890, Q5R364, Q5R365, Q8IY12 | Gene names | SRPK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK1 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 1) (SR-protein-specific kinase 1) (SFRS protein kinase 1). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.004235 (rank : 59) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
MARK2_MOUSE
|
||||||
| NC score | 0.004000 (rank : 60) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q05512, Q6PDR4, Q8BR95 | Gene names | Mark2, Emk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL Motif Kinase) (EMK1). | |||||
|
HXA2_HUMAN
|
||||||
| NC score | 0.003718 (rank : 61) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43364 | Gene names | HOXA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein Hox-A2. | |||||
|
ZEP2_MOUSE
|
||||||
| NC score | 0.002923 (rank : 62) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3UHF7, O55140, Q3UVD4, Q3UVH5 | Gene names | Hivep2, Mibp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 homolog (Myc intron-binding protein 1) (MIBP-1). | |||||
|
CAD23_MOUSE
|
||||||
| NC score | 0.002029 (rank : 63) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99PF4, Q99NH1, Q9D4N9 | Gene names | Cdh23 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-23 precursor (Otocadherin). | |||||
|
MELK_MOUSE
|
||||||
| NC score | 0.001457 (rank : 64) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61846, Q61804, Q6ZQH6 | Gene names | Melk, Kiaa0175, Pk38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Maternal embryonic leucine zipper kinase (EC 2.7.11.1) (Protein kinase PK38) (mPK38). | |||||
|
NRK_HUMAN
|
||||||
| NC score | 0.001111 (rank : 65) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7Z2Y5, Q32ND6, Q5H9K2, Q6ZMP2 | Gene names | NRK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nik-related protein kinase (EC 2.7.11.1). | |||||