Please be patient as the page loads
|
AXN1_HUMAN
|
||||||
| SwissProt Accessions | O15169, Q4TT26, Q4TT27, Q86YA7, Q8WVW6, Q96S28 | Gene names | AXIN1, AXIN | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (hAxin). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
AXN1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 99 | |
| SwissProt Accessions | O15169, Q4TT26, Q4TT27, Q86YA7, Q8WVW6, Q96S28 | Gene names | AXIN1, AXIN | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (hAxin). | |||||
|
AXN1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.992127 (rank : 2) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O35625 | Gene names | Axin1, Axin, Fu | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (Fused protein). | |||||
|
AXN2_HUMAN
|
||||||
| θ value | 8.33966e-147 (rank : 3) | NC score | 0.953621 (rank : 3) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Y2T1, Q9H3M6, Q9UH84 | Gene names | AXIN2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
AXN2_MOUSE
|
||||||
| θ value | 2.77624e-142 (rank : 4) | NC score | 0.948670 (rank : 4) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 9.90251e-15 (rank : 5) | NC score | 0.504799 (rank : 35) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
RGS10_HUMAN
|
||||||
| θ value | 1.68911e-14 (rank : 6) | NC score | 0.712747 (rank : 5) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O43665 | Gene names | RGS10 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 4.91457e-14 (rank : 7) | NC score | 0.444140 (rank : 40) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
RGS10_MOUSE
|
||||||
| θ value | 1.42992e-13 (rank : 8) | NC score | 0.705571 (rank : 6) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9CQE5, Q9D3L2 | Gene names | Rgs10 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS18_MOUSE
|
||||||
| θ value | 1.42992e-13 (rank : 9) | NC score | 0.697968 (rank : 7) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99PG4 | Gene names | Rgs18 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS5_HUMAN
|
||||||
| θ value | 1.86753e-13 (rank : 10) | NC score | 0.692757 (rank : 9) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O15539 | Gene names | RGS5 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS18_HUMAN
|
||||||
| θ value | 3.18553e-13 (rank : 11) | NC score | 0.697833 (rank : 8) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NS28 | Gene names | RGS18, RGS13 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS1_HUMAN
|
||||||
| θ value | 4.16044e-13 (rank : 12) | NC score | 0.690311 (rank : 10) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q08116, Q07918, Q9H1W2 | Gene names | RGS1, 1R20, BL34 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1) (Early response protein 1R20) (B-cell activation protein BL34). | |||||
|
RGS5_MOUSE
|
||||||
| θ value | 1.2105e-12 (rank : 13) | NC score | 0.685938 (rank : 13) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O08850, Q543B1, Q9D0Z2 | Gene names | Rgs5 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS8_HUMAN
|
||||||
| θ value | 1.58096e-12 (rank : 14) | NC score | 0.687206 (rank : 12) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P57771, Q3SYD2 | Gene names | RGS8 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS8_MOUSE
|
||||||
| θ value | 1.58096e-12 (rank : 15) | NC score | 0.688579 (rank : 11) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8BXT1 | Gene names | Rgs8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS17_MOUSE
|
||||||
| θ value | 3.52202e-12 (rank : 16) | NC score | 0.663019 (rank : 23) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB0, Q543T9 | Gene names | Rgs17, Rgsz2 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17) (Regulator of Gz-selective protein signaling 2). | |||||
|
RGS4_MOUSE
|
||||||
| θ value | 1.33837e-11 (rank : 17) | NC score | 0.683634 (rank : 16) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08899, Q99L30 | Gene names | Rgs4 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4). | |||||
|
RGS2_MOUSE
|
||||||
| θ value | 1.74796e-11 (rank : 18) | NC score | 0.682622 (rank : 17) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O08849, Q544S7, Q91WX1, Q9JL24 | Gene names | Rgs2 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2). | |||||
|
RGS4_HUMAN
|
||||||
| θ value | 1.74796e-11 (rank : 19) | NC score | 0.682085 (rank : 18) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49798 | Gene names | RGS4 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4) (RGP4). | |||||
|
RGS1_MOUSE
|
||||||
| θ value | 2.28291e-11 (rank : 20) | NC score | 0.684743 (rank : 15) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9JL25, Q3TBD1, Q3TC18, Q3TD56 | Gene names | Rgs1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1). | |||||
|
RGS2_HUMAN
|
||||||
| θ value | 2.28291e-11 (rank : 21) | NC score | 0.685857 (rank : 14) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P41220 | Gene names | RGS2, G0S8 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2) (G0/G1 switch regulatory protein 8). | |||||
|
RGS16_MOUSE
|
||||||
| θ value | 3.89403e-11 (rank : 22) | NC score | 0.676076 (rank : 19) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P97428, O09091, P97420, Q80V16 | Gene names | Rgs16, Rgsr | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS20_MOUSE
|
||||||
| θ value | 2.52405e-10 (rank : 23) | NC score | 0.658041 (rank : 24) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB1, Q3TY63, Q9CUV8, Q9QZB2 | Gene names | Rgs20, Rgsz1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of G-protein signaling Z1). | |||||
|
RGS13_MOUSE
|
||||||
| θ value | 3.29651e-10 (rank : 24) | NC score | 0.675775 (rank : 20) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8K443 | Gene names | Rgs13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS16_HUMAN
|
||||||
| θ value | 5.62301e-10 (rank : 25) | NC score | 0.670429 (rank : 22) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O15492, Q99701 | Gene names | RGS16, RGSR | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS14_HUMAN
|
||||||
| θ value | 7.34386e-10 (rank : 26) | NC score | 0.621802 (rank : 30) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O43566, O43565, Q8TD62 | Gene names | RGS14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14). | |||||
|
RGS17_HUMAN
|
||||||
| θ value | 7.34386e-10 (rank : 27) | NC score | 0.652308 (rank : 27) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UGC6, Q5TF49, Q8TD61, Q9UJS8 | Gene names | RGS17 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17). | |||||
|
RGS20_HUMAN
|
||||||
| θ value | 9.59137e-10 (rank : 28) | NC score | 0.646598 (rank : 28) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O76081, Q96BG9 | Gene names | RGS20, RGSZ1, ZGAP1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of Gz-selective protein signaling 1) (Gz-selective GTPase-activating protein) (G(z)GAP). | |||||
|
RGS19_HUMAN
|
||||||
| θ value | 1.63604e-09 (rank : 29) | NC score | 0.654391 (rank : 26) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P49795, Q53XN0, Q8TD60 | Gene names | RGS19, GAIP, GNAI3IP | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19) (G-alpha-interacting protein) (GAIP protein). | |||||
|
RGS14_MOUSE
|
||||||
| θ value | 2.13673e-09 (rank : 30) | NC score | 0.629383 (rank : 29) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P97492, Q9DCD1 | Gene names | Rgs14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14) (RAP1/RAP2-interacting protein). | |||||
|
RGS19_MOUSE
|
||||||
| θ value | 4.76016e-09 (rank : 31) | NC score | 0.654540 (rank : 25) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CX84, Q99L50 | Gene names | Rgs19 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19). | |||||
|
RGS13_HUMAN
|
||||||
| θ value | 6.21693e-09 (rank : 32) | NC score | 0.673757 (rank : 21) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O14921, Q6PGR2, Q8TD63, Q9BX45 | Gene names | RGS13 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS12_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 33) | NC score | 0.526789 (rank : 34) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O14924, O14922, O14923, O43510, O75338 | Gene names | RGS12 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 12 (RGS12). | |||||
|
DVL2_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 34) | NC score | 0.230057 (rank : 44) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O14641 | Gene names | DVL2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL2_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 35) | NC score | 0.230702 (rank : 43) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q60838 | Gene names | Dvl2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL3_HUMAN
|
||||||
| θ value | 1.87187e-05 (rank : 36) | NC score | 0.237050 (rank : 42) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92997, O14642, Q13531, Q8N5E9, Q92607 | Gene names | DVL3, KIAA0208 | |||
|
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL3_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 37) | NC score | 0.237240 (rank : 41) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q61062 | Gene names | Dvl3 | |||
|
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
RGS11_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 38) | NC score | 0.553728 (rank : 31) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O94810, O75883 | Gene names | RGS11 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 11 (RGS11). | |||||
|
RGS9_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 39) | NC score | 0.530727 (rank : 32) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O54828, Q9Z0S0 | Gene names | Rgs9 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS9_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 40) | NC score | 0.529709 (rank : 33) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75916, O75573, Q696R2, Q8TD64, Q8TD65, Q9HC32, Q9HC33 | Gene names | RGS9 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
DVL1L_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 41) | NC score | 0.213538 (rank : 47) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54792 | Gene names | DVL1L1, DVL, DVL1 | |||
|
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1-like (Dishevelled- 1-like) (DSH homolog 1-like). | |||||
|
DVL1_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 42) | NC score | 0.211332 (rank : 49) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O14640 | Gene names | DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
DVL1_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 43) | NC score | 0.211388 (rank : 48) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
Domain Architecture |
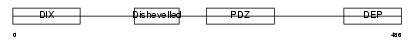
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
SNX14_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 44) | NC score | 0.152386 (rank : 53) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y5W7, Q9BSD1 | Gene names | SNX14 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-14. | |||||
|
SNX14_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 45) | NC score | 0.152639 (rank : 52) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BHY8 | Gene names | Snx14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-14. | |||||
|
RGS7_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 46) | NC score | 0.499228 (rank : 36) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P49802, Q8TD66, Q8TD67, Q9UNU7, Q9Y6B9 | Gene names | RGS7 | |||
|
Domain Architecture |
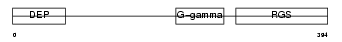
|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
ARBK1_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 47) | NC score | 0.023388 (rank : 69) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P25098, Q13837 | Gene names | ADRBK1, BARK, BARK1, GRK2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 1 (EC 2.7.11.15) (Beta-ARK-1) (G- protein coupled receptor kinase 2). | |||||
|
ARBK1_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 48) | NC score | 0.020146 (rank : 71) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99MK8, Q99LL8 | Gene names | Adrbk1, Grk2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 1 (EC 2.7.11.15) (Beta-ARK-1) (G- protein-coupled receptor kinase 2). | |||||
|
CT151_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 49) | NC score | 0.067573 (rank : 55) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NC74, Q8N4Z9, Q9BR75, Q9H0Y9 | Gene names | C20orf151 | |||
|
Domain Architecture |
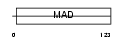
|
|||||
| Description | Uncharacterized protein C20orf151. | |||||
|
BAZ1B_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 50) | NC score | 0.041399 (rank : 59) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
RGS7_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 51) | NC score | 0.491638 (rank : 37) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O54829 | Gene names | Rgs7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
MARCS_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 52) | NC score | 0.054881 (rank : 56) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
SNX13_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 53) | NC score | 0.226296 (rank : 45) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Y5W8, O94821, Q8WVZ2, Q8WXH8 | Gene names | SNX13, KIAA0713 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-13 (RGS domain- and PHOX domain-containing protein) (RGS-PX1). | |||||
|
SNX13_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 54) | NC score | 0.226263 (rank : 46) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6PHS6 | Gene names | Snx13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-13. | |||||
|
CD2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 55) | NC score | 0.045414 (rank : 57) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
ES8L1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 56) | NC score | 0.030856 (rank : 64) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TE68, Q71RE2, Q8NC10, Q96BB7, Q9BSQ2, Q9GZQ2, Q9NXH0 | Gene names | EPS8L1, DRC3, EPS8R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 1 (Epidermal growth factor receptor pathway substrate 8-related protein 1) (EPS8-like protein 1). | |||||
|
RGS6_MOUSE
|
||||||
| θ value | 0.279714 (rank : 57) | NC score | 0.478737 (rank : 38) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Z2H2, Q8BGI8 | Gene names | Rgs6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 6 (RGS6). | |||||
|
EP300_HUMAN
|
||||||
| θ value | 0.62314 (rank : 58) | NC score | 0.036376 (rank : 60) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
ES8L1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 59) | NC score | 0.027935 (rank : 67) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R5F8, Q8R0D6, Q9D2M6 | Gene names | Eps8l1, Eps8r1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 1 (Epidermal growth factor receptor pathway substrate 8-related protein 1) (EPS8-like protein 1). | |||||
|
CDSN_MOUSE
|
||||||
| θ value | 0.813845 (rank : 60) | NC score | 0.031287 (rank : 63) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TPC1 | Gene names | Cdsn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor. | |||||
|
PDZD2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 61) | NC score | 0.045217 (rank : 58) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 62) | NC score | 0.022364 (rank : 70) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
ZN333_HUMAN
|
||||||
| θ value | 1.06291 (rank : 63) | NC score | -0.002316 (rank : 100) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96JL9, Q8TDL0 | Gene names | ZNF333, KIAA1806 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 333. | |||||
|
ARBK2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 64) | NC score | 0.019144 (rank : 72) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P35626, Q9UGW9 | Gene names | ADRBK2, BARK2, GRK3 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 2 (EC 2.7.11.15) (Beta-ARK-2) (G- protein-coupled receptor kinase 3). | |||||
|
RGS6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.448389 (rank : 39) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P49758, O75576, O75577, Q8TE13, Q8TE14, Q8TE15, Q8TE16, Q8TE17, Q8TE18, Q8TE19, Q8TE20, Q8TE21, Q8TE22, Q9Y245, Q9Y647 | Gene names | RGS6 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 6 (RGS6) (S914). | |||||
|
CAC1C_HUMAN
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.007143 (rank : 85) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
SRCA_HUMAN
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.025487 (rank : 68) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
GATA3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 68) | NC score | 0.009830 (rank : 81) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23772 | Gene names | Gata3, Gata-3 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-acting T-cell-specific transcription factor GATA-3 (GATA-binding factor 3). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.017771 (rank : 75) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PK3C3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.018624 (rank : 74) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEB9, Q15134 | Gene names | PIK3C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3) (Phosphatidylinositol 3-kinase p100 subunit). | |||||
|
PK3C3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.018634 (rank : 73) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PF93, Q8R3S8 | Gene names | Pik3c3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.031655 (rank : 62) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.028563 (rank : 65) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
PPRB_HUMAN
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.015127 (rank : 76) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
RGP2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.011589 (rank : 78) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P47736 | Gene names | RAP1GAP, RAP1GA1 | |||
|
Domain Architecture |
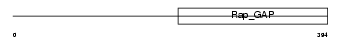
|
|||||
| Description | Rap1 GTPase-activating protein 1 (Rap1GAP). | |||||
|
SPAG1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.013258 (rank : 77) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80ZX8, Q8CCK7, Q9QZJ3, Q9QZJ4 | Gene names | Spag1, Tpis | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 1 (Infertility-related sperm protein Spag-1) (TPR-containing protein involved in spermatogenesis) (TPIS). | |||||
|
FOXJ3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.006290 (rank : 89) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UPW0, Q9NSS7 | Gene names | FOXJ3, KIAA1041 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein J3. | |||||
|
K1949_HUMAN
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.035163 (rank : 61) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6NYC8, Q68CK8, Q6ZUJ6, Q8NDQ4, Q8TF52, Q9BRL9 | Gene names | KIAA1949 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949. | |||||
|
MSX2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.000802 (rank : 95) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q03358, Q63856 | Gene names | Msx2, Hox-8.1, Msx-2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein MSX-2 (Hox-8.1). | |||||
|
SNX25_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.115270 (rank : 54) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9H3E2 | Gene names | SNX25 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-25. | |||||
|
SVIL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.010507 (rank : 79) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8K4L3 | Gene names | Svil | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
ZHX2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.006922 (rank : 86) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6X8 | Gene names | ZHX2, AFR1, KIAA0854, RAF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 2 (Zinc finger and homeodomain protein 2) (Alpha-fetoprotein regulator 1) (AFP regulator 1) (Regulator of AFP). | |||||
|
ZN358_HUMAN
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | -0.001066 (rank : 97) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 840 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NW07, Q9BTM7 | Gene names | ZNF358 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 358. | |||||
|
DDX49_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.006809 (rank : 87) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6V7, Q53FJ1 | Gene names | DDX49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX49 (EC 3.6.1.-) (DEAD box protein 49). | |||||
|
FOXJ3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.005839 (rank : 90) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BUR3, Q3UTD8 | Gene names | Foxj3, Kiaa1041 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein J3. | |||||
|
K1949_MOUSE
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.028397 (rank : 66) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BQ30, Q3UDS6, Q8CEY9, Q8R3D8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949 homolog. | |||||
|
LRP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | -0.000299 (rank : 96) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q07954, Q2PP12, Q8IVG8 | Gene names | LRP1, A2MR | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 1 precursor (LRP) (Alpha-2-macroglobulin receptor) (A2MR) (Apolipoprotein E receptor) (APOER) (CD91 antigen). | |||||
|
MAST4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.004278 (rank : 92) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
MRP6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.002358 (rank : 94) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9R1S7, Q80YB6 | Gene names | Abcc6, Mrp6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multidrug resistance-associated protein 6 (ATP-binding cassette sub- family C member 6). | |||||
|
MYH6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | -0.001417 (rank : 98) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
RASA2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.008179 (rank : 82) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P58069 | Gene names | Rasa2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
RRBP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.004914 (rank : 91) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.002731 (rank : 93) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
CEBPA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.006770 (rank : 88) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P53566, Q91XB6 | Gene names | Cebpa | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein alpha (C/EBP alpha). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.010459 (rank : 80) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MYH6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | -0.001548 (rank : 99) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
PTN23_MOUSE
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.008117 (rank : 83) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
SGPP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.007903 (rank : 84) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JI99 | Gene names | Sgpp1, Spp1, Spph1 | |||
|
Domain Architecture |
|
|||||
| Description | Sphingosine-1-phosphate phosphatase 1 (EC 3.1.3.-) (Sphingosine-1- phosphatase 1) (SPPase1) (SPP) (mSPP1). | |||||
|
ZN426_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | -0.002982 (rank : 101) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BUY5 | Gene names | ZNF426 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 426. | |||||
|
AKA10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.197124 (rank : 50) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O43572, Q96AJ7 | Gene names | AKAP10 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
AKA10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.196854 (rank : 51) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O88845, Q5SUB5, Q5SUB7, Q7TPE7, Q8BQL6 | Gene names | Akap10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
AXN1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 99 | |
| SwissProt Accessions | O15169, Q4TT26, Q4TT27, Q86YA7, Q8WVW6, Q96S28 | Gene names | AXIN1, AXIN | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (hAxin). | |||||
|
AXN1_MOUSE
|
||||||
| NC score | 0.992127 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O35625 | Gene names | Axin1, Axin, Fu | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (Fused protein). | |||||
|
AXN2_HUMAN
|
||||||
| NC score | 0.953621 (rank : 3) | θ value | 8.33966e-147 (rank : 3) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Y2T1, Q9H3M6, Q9UH84 | Gene names | AXIN2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
AXN2_MOUSE
|
||||||
| NC score | 0.948670 (rank : 4) | θ value | 2.77624e-142 (rank : 4) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
RGS10_HUMAN
|
||||||
| NC score | 0.712747 (rank : 5) | θ value | 1.68911e-14 (rank : 6) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O43665 | Gene names | RGS10 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS10_MOUSE
|
||||||
| NC score | 0.705571 (rank : 6) | θ value | 1.42992e-13 (rank : 8) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9CQE5, Q9D3L2 | Gene names | Rgs10 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS18_MOUSE
|
||||||
| NC score | 0.697968 (rank : 7) | θ value | 1.42992e-13 (rank : 9) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99PG4 | Gene names | Rgs18 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS18_HUMAN
|
||||||
| NC score | 0.697833 (rank : 8) | θ value | 3.18553e-13 (rank : 11) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NS28 | Gene names | RGS18, RGS13 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS5_HUMAN
|
||||||
| NC score | 0.692757 (rank : 9) | θ value | 1.86753e-13 (rank : 10) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O15539 | Gene names | RGS5 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS1_HUMAN
|
||||||
| NC score | 0.690311 (rank : 10) | θ value | 4.16044e-13 (rank : 12) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q08116, Q07918, Q9H1W2 | Gene names | RGS1, 1R20, BL34 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1) (Early response protein 1R20) (B-cell activation protein BL34). | |||||
|
RGS8_MOUSE
|
||||||
| NC score | 0.688579 (rank : 11) | θ value | 1.58096e-12 (rank : 15) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8BXT1 | Gene names | Rgs8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS8_HUMAN
|
||||||
| NC score | 0.687206 (rank : 12) | θ value | 1.58096e-12 (rank : 14) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P57771, Q3SYD2 | Gene names | RGS8 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS5_MOUSE
|
||||||
| NC score | 0.685938 (rank : 13) | θ value | 1.2105e-12 (rank : 13) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O08850, Q543B1, Q9D0Z2 | Gene names | Rgs5 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS2_HUMAN
|
||||||
| NC score | 0.685857 (rank : 14) | θ value | 2.28291e-11 (rank : 21) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P41220 | Gene names | RGS2, G0S8 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2) (G0/G1 switch regulatory protein 8). | |||||
|
RGS1_MOUSE
|
||||||
| NC score | 0.684743 (rank : 15) | θ value | 2.28291e-11 (rank : 20) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9JL25, Q3TBD1, Q3TC18, Q3TD56 | Gene names | Rgs1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1). | |||||
|
RGS4_MOUSE
|
||||||
| NC score | 0.683634 (rank : 16) | θ value | 1.33837e-11 (rank : 17) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08899, Q99L30 | Gene names | Rgs4 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4). | |||||
|
RGS2_MOUSE
|
||||||
| NC score | 0.682622 (rank : 17) | θ value | 1.74796e-11 (rank : 18) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O08849, Q544S7, Q91WX1, Q9JL24 | Gene names | Rgs2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2). | |||||
|
RGS4_HUMAN
|
||||||
| NC score | 0.682085 (rank : 18) | θ value | 1.74796e-11 (rank : 19) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49798 | Gene names | RGS4 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4) (RGP4). | |||||
|
RGS16_MOUSE
|
||||||
| NC score | 0.676076 (rank : 19) | θ value | 3.89403e-11 (rank : 22) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P97428, O09091, P97420, Q80V16 | Gene names | Rgs16, Rgsr | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS13_MOUSE
|
||||||
| NC score | 0.675775 (rank : 20) | θ value | 3.29651e-10 (rank : 24) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8K443 | Gene names | Rgs13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS13_HUMAN
|
||||||
| NC score | 0.673757 (rank : 21) | θ value | 6.21693e-09 (rank : 32) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O14921, Q6PGR2, Q8TD63, Q9BX45 | Gene names | RGS13 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS16_HUMAN
|
||||||
| NC score | 0.670429 (rank : 22) | θ value | 5.62301e-10 (rank : 25) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O15492, Q99701 | Gene names | RGS16, RGSR | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS17_MOUSE
|
||||||
| NC score | 0.663019 (rank : 23) | θ value | 3.52202e-12 (rank : 16) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB0, Q543T9 | Gene names | Rgs17, Rgsz2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17) (Regulator of Gz-selective protein signaling 2). | |||||
|
RGS20_MOUSE
|
||||||
| NC score | 0.658041 (rank : 24) | θ value | 2.52405e-10 (rank : 23) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB1, Q3TY63, Q9CUV8, Q9QZB2 | Gene names | Rgs20, Rgsz1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of G-protein signaling Z1). | |||||
|
RGS19_MOUSE
|
||||||
| NC score | 0.654540 (rank : 25) | θ value | 4.76016e-09 (rank : 31) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CX84, Q99L50 | Gene names | Rgs19 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19). | |||||
|
RGS19_HUMAN
|
||||||
| NC score | 0.654391 (rank : 26) | θ value | 1.63604e-09 (rank : 29) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P49795, Q53XN0, Q8TD60 | Gene names | RGS19, GAIP, GNAI3IP | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19) (G-alpha-interacting protein) (GAIP protein). | |||||
|
RGS17_HUMAN
|
||||||
| NC score | 0.652308 (rank : 27) | θ value | 7.34386e-10 (rank : 27) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UGC6, Q5TF49, Q8TD61, Q9UJS8 | Gene names | RGS17 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17). | |||||
|
RGS20_HUMAN
|
||||||
| NC score | 0.646598 (rank : 28) | θ value | 9.59137e-10 (rank : 28) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O76081, Q96BG9 | Gene names | RGS20, RGSZ1, ZGAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of Gz-selective protein signaling 1) (Gz-selective GTPase-activating protein) (G(z)GAP). | |||||
|
RGS14_MOUSE
|
||||||
| NC score | 0.629383 (rank : 29) | θ value | 2.13673e-09 (rank : 30) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P97492, Q9DCD1 | Gene names | Rgs14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14) (RAP1/RAP2-interacting protein). | |||||
|
RGS14_HUMAN
|
||||||
| NC score | 0.621802 (rank : 30) | θ value | 7.34386e-10 (rank : 26) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O43566, O43565, Q8TD62 | Gene names | RGS14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14). | |||||
|
RGS11_HUMAN
|
||||||
| NC score | 0.553728 (rank : 31) | θ value | 5.44631e-05 (rank : 38) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O94810, O75883 | Gene names | RGS11 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 11 (RGS11). | |||||
|
RGS9_MOUSE
|
||||||
| NC score | 0.530727 (rank : 32) | θ value | 7.1131e-05 (rank : 39) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O54828, Q9Z0S0 | Gene names | Rgs9 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS9_HUMAN
|
||||||
| NC score | 0.529709 (rank : 33) | θ value | 0.000158464 (rank : 40) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75916, O75573, Q696R2, Q8TD64, Q8TD65, Q9HC32, Q9HC33 | Gene names | RGS9 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS12_HUMAN
|
||||||
| NC score | 0.526789 (rank : 34) | θ value | 6.43352e-06 (rank : 33) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O14924, O14922, O14923, O43510, O75338 | Gene names | RGS12 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 12 (RGS12). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.504799 (rank : 35) | θ value | 9.90251e-15 (rank : 5) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
RGS7_HUMAN
|
||||||
| NC score | 0.499228 (rank : 36) | θ value | 0.00665767 (rank : 46) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P49802, Q8TD66, Q8TD67, Q9UNU7, Q9Y6B9 | Gene names | RGS7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS7_MOUSE
|
||||||
| NC score | 0.491638 (rank : 37) | θ value | 0.0431538 (rank : 51) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O54829 | Gene names | Rgs7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS6_MOUSE
|
||||||
| NC score | 0.478737 (rank : 38) | θ value | 0.279714 (rank : 57) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Z2H2, Q8BGI8 | Gene names | Rgs6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 6 (RGS6). | |||||
|
RGS6_HUMAN
|
||||||
| NC score | 0.448389 (rank : 39) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P49758, O75576, O75577, Q8TE13, Q8TE14, Q8TE15, Q8TE16, Q8TE17, Q8TE18, Q8TE19, Q8TE20, Q8TE21, Q8TE22, Q9Y245, Q9Y647 | Gene names | RGS6 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 6 (RGS6) (S914). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.444140 (rank : 40) | θ value | 4.91457e-14 (rank : 7) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
DVL3_MOUSE
|
||||||
| NC score | 0.237240 (rank : 41) | θ value | 1.87187e-05 (rank : 37) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q61062 | Gene names | Dvl3 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL3_HUMAN
|
||||||
| NC score | 0.237050 (rank : 42) | θ value | 1.87187e-05 (rank : 36) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92997, O14642, Q13531, Q8N5E9, Q92607 | Gene names | DVL3, KIAA0208 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL2_MOUSE
|
||||||
| NC score | 0.230702 (rank : 43) | θ value | 1.43324e-05 (rank : 35) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q60838 | Gene names | Dvl2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL2_HUMAN
|
||||||
| NC score | 0.230057 (rank : 44) | θ value | 1.43324e-05 (rank : 34) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O14641 | Gene names | DVL2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
SNX13_HUMAN
|
||||||
| NC score | 0.226296 (rank : 45) | θ value | 0.0961366 (rank : 53) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Y5W8, O94821, Q8WVZ2, Q8WXH8 | Gene names | SNX13, KIAA0713 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-13 (RGS domain- and PHOX domain-containing protein) (RGS-PX1). | |||||
|
SNX13_MOUSE
|
||||||
| NC score | 0.226263 (rank : 46) | θ value | 0.0961366 (rank : 54) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6PHS6 | Gene names | Snx13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-13. | |||||
|
DVL1L_HUMAN
|
||||||
| NC score | 0.213538 (rank : 47) | θ value | 0.000602161 (rank : 41) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54792 | Gene names | DVL1L1, DVL, DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1-like (Dishevelled- 1-like) (DSH homolog 1-like). | |||||
|
DVL1_MOUSE
|
||||||
| NC score | 0.211388 (rank : 48) | θ value | 0.000602161 (rank : 43) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
DVL1_HUMAN
|
||||||
| NC score | 0.211332 (rank : 49) | θ value | 0.000602161 (rank : 42) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O14640 | Gene names | DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
AKA10_HUMAN
|
||||||
| NC score | 0.197124 (rank : 50) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O43572, Q96AJ7 | Gene names | AKAP10 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
AKA10_MOUSE
|
||||||
| NC score | 0.196854 (rank : 51) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O88845, Q5SUB5, Q5SUB7, Q7TPE7, Q8BQL6 | Gene names | Akap10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
SNX14_MOUSE
|
||||||
| NC score | 0.152639 (rank : 52) | θ value | 0.00175202 (rank : 45) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BHY8 | Gene names | Snx14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-14. | |||||
|
SNX14_HUMAN
|
||||||
| NC score | 0.152386 (rank : 53) | θ value | 0.00175202 (rank : 44) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y5W7, Q9BSD1 | Gene names | SNX14 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-14. | |||||
|
SNX25_HUMAN
|
||||||
| NC score | 0.115270 (rank : 54) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9H3E2 | Gene names | SNX25 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-25. | |||||
|
CT151_HUMAN
|
||||||
| NC score | 0.067573 (rank : 55) | θ value | 0.0193708 (rank : 49) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NC74, Q8N4Z9, Q9BR75, Q9H0Y9 | Gene names | C20orf151 | |||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf151. | |||||
|
MARCS_MOUSE
|
||||||
| NC score | 0.054881 (rank : 56) | θ value | 0.0961366 (rank : 52) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
CD2_HUMAN
|
||||||
| NC score | 0.045414 (rank : 57) | θ value | 0.125558 (rank : 55) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
PDZD2_HUMAN
|
||||||
| NC score | 0.045217 (rank : 58) | θ value | 0.813845 (rank : 61) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
BAZ1B_HUMAN
|
||||||
| NC score | 0.041399 (rank : 59) | θ value | 0.0330416 (rank : 50) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.036376 (rank : 60) | θ value | 0.62314 (rank : 58) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
K1949_HUMAN
|
||||||
| NC score | 0.035163 (rank : 61) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6NYC8, Q68CK8, Q6ZUJ6, Q8NDQ4, Q8TF52, Q9BRL9 | Gene names | KIAA1949 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949. | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.031655 (rank : 62) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CDSN_MOUSE
|
||||||
| NC score | 0.031287 (rank : 63) | θ value | 0.813845 (rank : 60) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TPC1 | Gene names | Cdsn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor. | |||||
|
ES8L1_HUMAN
|
||||||
| NC score | 0.030856 (rank : 64) | θ value | 0.21417 (rank : 56) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TE68, Q71RE2, Q8NC10, Q96BB7, Q9BSQ2, Q9GZQ2, Q9NXH0 | Gene names | EPS8L1, DRC3, EPS8R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 1 (Epidermal growth factor receptor pathway substrate 8-related protein 1) (EPS8-like protein 1). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.028563 (rank : 65) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
K1949_MOUSE
|
||||||
| NC score | 0.028397 (rank : 66) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BQ30, Q3UDS6, Q8CEY9, Q8R3D8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949 homolog. | |||||
|
ES8L1_MOUSE
|
||||||
| NC score | 0.027935 (rank : 67) | θ value | 0.62314 (rank : 59) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R5F8, Q8R0D6, Q9D2M6 | Gene names | Eps8l1, Eps8r1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 1 (Epidermal growth factor receptor pathway substrate 8-related protein 1) (EPS8-like protein 1). | |||||
|
SRCA_HUMAN
|
||||||
| NC score | 0.025487 (rank : 68) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
ARBK1_HUMAN
|
||||||
| NC score | 0.023388 (rank : 69) | θ value | 0.0113563 (rank : 47) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P25098, Q13837 | Gene names | ADRBK1, BARK, BARK1, GRK2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 1 (EC 2.7.11.15) (Beta-ARK-1) (G- protein coupled receptor kinase 2). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.022364 (rank : 70) | θ value | 1.06291 (rank : 62) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
ARBK1_MOUSE
|
||||||
| NC score | 0.020146 (rank : 71) | θ value | 0.0113563 (rank : 48) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99MK8, Q99LL8 | Gene names | Adrbk1, Grk2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 1 (EC 2.7.11.15) (Beta-ARK-1) (G- protein-coupled receptor kinase 2). | |||||
|
ARBK2_HUMAN
|
||||||
| NC score | 0.019144 (rank : 72) | θ value | 1.38821 (rank : 64) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P35626, Q9UGW9 | Gene names | ADRBK2, BARK2, GRK3 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 2 (EC 2.7.11.15) (Beta-ARK-2) (G- protein-coupled receptor kinase 3). | |||||
|
PK3C3_MOUSE
|
||||||
| NC score | 0.018634 (rank : 73) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PF93, Q8R3S8 | Gene names | Pik3c3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3). | |||||
|
PK3C3_HUMAN
|
||||||
| NC score | 0.018624 (rank : 74) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEB9, Q15134 | Gene names | PIK3C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3) (Phosphatidylinositol 3-kinase p100 subunit). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.017771 (rank : 75) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PPRB_HUMAN
|
||||||
| NC score | 0.015127 (rank : 76) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
SPAG1_MOUSE
|
||||||
| NC score | 0.013258 (rank : 77) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80ZX8, Q8CCK7, Q9QZJ3, Q9QZJ4 | Gene names | Spag1, Tpis | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 1 (Infertility-related sperm protein Spag-1) (TPR-containing protein involved in spermatogenesis) (TPIS). | |||||
|
RGP2_HUMAN
|
||||||
| NC score | 0.011589 (rank : 78) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P47736 | Gene names | RAP1GAP, RAP1GA1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-activating protein 1 (Rap1GAP). | |||||
|
SVIL_MOUSE
|
||||||
| NC score | 0.010507 (rank : 79) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8K4L3 | Gene names | Svil | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.010459 (rank : 80) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
GATA3_MOUSE
|
||||||
| NC score | 0.009830 (rank : 81) | θ value | 3.0926 (rank : 68) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23772 | Gene names | Gata3, Gata-3 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-acting T-cell-specific transcription factor GATA-3 (GATA-binding factor 3). | |||||
|
RASA2_MOUSE
|
||||||
| NC score | 0.008179 (rank : 82) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P58069 | Gene names | Rasa2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
PTN23_MOUSE
|
||||||
| NC score | 0.008117 (rank : 83) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
SGPP1_MOUSE
|
||||||
| NC score | 0.007903 (rank : 84) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JI99 | Gene names | Sgpp1, Spp1, Spph1 | |||
|
Domain Architecture |

|
|||||
| Description | Sphingosine-1-phosphate phosphatase 1 (EC 3.1.3.-) (Sphingosine-1- phosphatase 1) (SPPase1) (SPP) (mSPP1). | |||||
|
CAC1C_HUMAN
|
||||||
| NC score | 0.007143 (rank : 85) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
ZHX2_HUMAN
|
||||||
| NC score | 0.006922 (rank : 86) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6X8 | Gene names | ZHX2, AFR1, KIAA0854, RAF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 2 (Zinc finger and homeodomain protein 2) (Alpha-fetoprotein regulator 1) (AFP regulator 1) (Regulator of AFP). | |||||
|
DDX49_HUMAN
|
||||||
| NC score | 0.006809 (rank : 87) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6V7, Q53FJ1 | Gene names | DDX49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX49 (EC 3.6.1.-) (DEAD box protein 49). | |||||
|
CEBPA_MOUSE
|
||||||
| NC score | 0.006770 (rank : 88) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P53566, Q91XB6 | Gene names | Cebpa | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein alpha (C/EBP alpha). | |||||
|
FOXJ3_HUMAN
|
||||||
| NC score | 0.006290 (rank : 89) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UPW0, Q9NSS7 | Gene names | FOXJ3, KIAA1041 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J3. | |||||
|
FOXJ3_MOUSE
|
||||||
| NC score | 0.005839 (rank : 90) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BUR3, Q3UTD8 | Gene names | Foxj3, Kiaa1041 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J3. | |||||
|
RRBP1_HUMAN
|
||||||
| NC score | 0.004914 (rank : 91) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
MAST4_HUMAN
|
||||||
| NC score | 0.004278 (rank : 92) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.002731 (rank : 93) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
MRP6_MOUSE
|
||||||
| NC score | 0.002358 (rank : 94) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9R1S7, Q80YB6 | Gene names | Abcc6, Mrp6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multidrug resistance-associated protein 6 (ATP-binding cassette sub- family C member 6). | |||||
|
MSX2_MOUSE
|
||||||
| NC score | 0.000802 (rank : 95) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q03358, Q63856 | Gene names | Msx2, Hox-8.1, Msx-2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein MSX-2 (Hox-8.1). | |||||
|
LRP1_HUMAN
|
||||||
| NC score | -0.000299 (rank : 96) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q07954, Q2PP12, Q8IVG8 | Gene names | LRP1, A2MR | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 1 precursor (LRP) (Alpha-2-macroglobulin receptor) (A2MR) (Apolipoprotein E receptor) (APOER) (CD91 antigen). | |||||
|
ZN358_HUMAN
|
||||||
| NC score | -0.001066 (rank : 97) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 840 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NW07, Q9BTM7 | Gene names | ZNF358 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 358. | |||||
|
MYH6_MOUSE
|
||||||
| NC score | -0.001417 (rank : 98) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH6_HUMAN
|
||||||
| NC score | -0.001548 (rank : 99) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
ZN333_HUMAN
|
||||||
| NC score | -0.002316 (rank : 100) | θ value | 1.06291 (rank : 63) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96JL9, Q8TDL0 | Gene names | ZNF333, KIAA1806 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 333. | |||||
|
ZN426_HUMAN
|
||||||
| NC score | -0.002982 (rank : 101) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BUY5 | Gene names | ZNF426 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 426. | |||||