Please be patient as the page loads
|
SMRCD_MOUSE
|
||||||
| SwissProt Accessions | Q04692 | Gene names | Smarcad1, Etl1, Kiaa1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase SMARCAD1) (Enhancer trap locus homolog 1) (Etl-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SMRCD_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.995177 (rank : 2) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9H4L7, Q96SX1, Q9H017, Q9H860, Q9NPU9, Q9ULU7 | Gene names | SMARCAD1, KIAA1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase 1) (hHEL1). | |||||
|
SMRCD_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | Q04692 | Gene names | Smarcad1, Etl1, Kiaa1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase SMARCAD1) (Enhancer trap locus homolog 1) (Etl-1). | |||||
|
SMCA1_MOUSE
|
||||||
| θ value | 1.49618e-87 (rank : 3) | NC score | 0.950148 (rank : 6) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6PGB8, Q8BSS1, Q91Y58 | Gene names | Smarca1, Snf2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable global transcription activator SNF2L1 (EC 3.6.1.-) (Nucleosome remodeling factor subunit SNF2L) (ATP-dependent helicase SMARCA1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 1) (DNA-dependent ATPase SNF2L). | |||||
|
SMCA5_HUMAN
|
||||||
| θ value | 6.28603e-86 (rank : 4) | NC score | 0.953181 (rank : 4) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O60264 | Gene names | SMARCA5, SNF2H, WCRF135 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 5 (EC 3.6.1.-) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin A5) (Sucrose nonfermenting protein 2 homolog) (hSNF2H). | |||||
|
SMCA5_MOUSE
|
||||||
| θ value | 6.28603e-86 (rank : 5) | NC score | 0.953199 (rank : 3) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q91ZW3, Q8C791, Q8CA22, Q8VDG1, Q925M9 | Gene names | Smarca5, Snf2h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 5 (EC 3.6.1.-) (Sucrose nonfermenting protein 2 homolog) (mSnf2h). | |||||
|
SMCA1_HUMAN
|
||||||
| θ value | 9.07703e-85 (rank : 6) | NC score | 0.952115 (rank : 5) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P28370, Q5JV41, Q5JV42 | Gene names | SMARCA1, SNF2L, SNF2L1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L1 (EC 3.6.1.-) (Nucleosome remodeling factor subunit SNF2L) (ATP-dependent helicase SMARCA1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 1). | |||||
|
CHD1_MOUSE
|
||||||
| θ value | 2.82823e-78 (rank : 7) | NC score | 0.927774 (rank : 14) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CHD1_HUMAN
|
||||||
| θ value | 3.69378e-78 (rank : 8) | NC score | 0.923753 (rank : 16) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
SMCA2_HUMAN
|
||||||
| θ value | 3.12697e-77 (rank : 9) | NC score | 0.901220 (rank : 22) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
SMCA4_HUMAN
|
||||||
| θ value | 5.33382e-77 (rank : 10) | NC score | 0.890758 (rank : 23) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
CHD2_HUMAN
|
||||||
| θ value | 2.64714e-76 (rank : 11) | NC score | 0.929053 (rank : 11) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O14647 | Gene names | CHD2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 2 (EC 3.6.1.-) (ATP- dependent helicase CHD2) (CHD-2). | |||||
|
CHD6_HUMAN
|
||||||
| θ value | 2.09745e-73 (rank : 12) | NC score | 0.922064 (rank : 17) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8TD26, Q5JYQ0, Q5TGZ9, Q5TH00, Q5TH01, Q8IZR2, Q8WTY0, Q9H4H6, Q9H6D4, Q9NTT7, Q9P2L1 | Gene names | CHD6, CHD5, KIAA1335, RIGB | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 6 (EC 3.6.1.-) (ATP- dependent helicase CHD6) (CHD-6) (Radiation-induced gene B protein). | |||||
|
CHD8_HUMAN
|
||||||
| θ value | 1.35953e-72 (rank : 13) | NC score | 0.928051 (rank : 13) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9HCK8, Q68DQ0, Q8N3Z9, Q8NCY4, Q8TBR9, Q96F26 | Gene names | CHD8, HELSNF1, KIAA1564 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 8 (EC 3.6.1.-) (ATP- dependent helicase CHD8) (CHD-8) (Helicase with SNF2 domain 1). | |||||
|
CHD9_HUMAN
|
||||||
| θ value | 8.81223e-72 (rank : 14) | NC score | 0.916691 (rank : 20) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
CHD9_MOUSE
|
||||||
| θ value | 1.15091e-71 (rank : 15) | NC score | 0.911771 (rank : 21) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
CHD5_HUMAN
|
||||||
| θ value | 7.71989e-68 (rank : 16) | NC score | 0.856509 (rank : 27) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
CHD4_HUMAN
|
||||||
| θ value | 3.83135e-67 (rank : 17) | NC score | 0.859771 (rank : 26) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
CHD4_MOUSE
|
||||||
| θ value | 3.83135e-67 (rank : 18) | NC score | 0.855748 (rank : 28) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
CHD3_HUMAN
|
||||||
| θ value | 1.6097e-65 (rank : 19) | NC score | 0.860005 (rank : 25) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
ERCC6_HUMAN
|
||||||
| θ value | 1.50663e-63 (rank : 20) | NC score | 0.938587 (rank : 8) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q03468 | Gene names | ERCC6, CSB | |||
|
Domain Architecture |

|
|||||
| Description | DNA excision repair protein ERCC-6 (EC 3.6.1.-) (ATP-dependent helicase ERCC6) (Cockayne syndrome protein CSB). | |||||
|
RA54B_HUMAN
|
||||||
| θ value | 8.55514e-59 (rank : 21) | NC score | 0.935527 (rank : 9) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y620 | Gene names | RAD54B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54B (EC 3.6.1.-) (RAD54 homolog B). | |||||
|
BTAF1_HUMAN
|
||||||
| θ value | 2.7521e-57 (rank : 22) | NC score | 0.942626 (rank : 7) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O14981, O43578 | Gene names | BTAF1, TAF172 | |||
|
Domain Architecture |

|
|||||
| Description | TATA-binding protein-associated factor 172 (EC 3.6.1.-) (ATP-dependent helicase BTAF1) (TBP-associated factor 172) (TAF-172) (TAF(II)170) (B- TFIID transcription factor-associated 170 kDa subunit). | |||||
|
INOC1_MOUSE
|
||||||
| θ value | 1.36585e-56 (rank : 23) | NC score | 0.919634 (rank : 18) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6ZPV2, Q6P7V0, Q6PCP1, Q8C9T7 | Gene names | Inoc1, Kiaa1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-). | |||||
|
INOC1_HUMAN
|
||||||
| θ value | 1.78386e-56 (rank : 24) | NC score | 0.919261 (rank : 19) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9ULG1, Q9NTG6 | Gene names | INOC1, KIAA1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-) (hINO80). | |||||
|
RA54B_MOUSE
|
||||||
| θ value | 9.16162e-53 (rank : 25) | NC score | 0.932349 (rank : 10) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6PFE3 | Gene names | Rad54b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54B (EC 3.6.1.-) (RAD54 homolog B). | |||||
|
RAD54_HUMAN
|
||||||
| θ value | 1.97686e-47 (rank : 26) | NC score | 0.928853 (rank : 12) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q92698, Q6IUY3 | Gene names | RAD54L, RAD54A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54-like (EC 3.6.1.-) (RAD54 homolog) (hRAD54) (hHR54). | |||||
|
RAD54_MOUSE
|
||||||
| θ value | 7.03111e-45 (rank : 27) | NC score | 0.927665 (rank : 15) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P70270, Q8BSR5, Q8C2C4, Q8K3D4 | Gene names | Rad54l, Rad54 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54-like (EC 3.6.1.-) (RAD54 homolog) (mRAD54) (mHR54). | |||||
|
EP400_HUMAN
|
||||||
| θ value | 2.50074e-42 (rank : 28) | NC score | 0.830562 (rank : 30) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
EP400_MOUSE
|
||||||
| θ value | 1.16164e-39 (rank : 29) | NC score | 0.821176 (rank : 32) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
SMRA3_MOUSE
|
||||||
| θ value | 3.4976e-36 (rank : 30) | NC score | 0.768519 (rank : 34) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6PCN7, O35596, O35597, Q80VT6 | Gene names | Smarca3, Snf2l3, Zbu1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 3 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 3) (TNF-response element-binding protein) (P113). | |||||
|
SMRA3_HUMAN
|
||||||
| θ value | 1.12514e-34 (rank : 31) | NC score | 0.765114 (rank : 35) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q14527, Q14536, Q16051, Q7KYJ6, Q86YA5, Q92652, Q96KM9 | Gene names | SMARCA3, HIP116A, HLTF, SNF2L3, ZBU1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 3 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 3) (DNA-binding protein/plasminogen activator inhibitor 1 regulator) (Helicase-like transcription factor) (HIP116). | |||||
|
TTF2_HUMAN
|
||||||
| θ value | 3.3877e-31 (rank : 32) | NC score | 0.872378 (rank : 24) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9UNY4, O75921, Q5T2K7, Q5VVU8, Q8N6I8 | Gene names | TTF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2) (Factor 2) (F2) (HuF2) (Lodestar homolog). | |||||
|
TTF2_MOUSE
|
||||||
| θ value | 1.57365e-28 (rank : 33) | NC score | 0.824645 (rank : 31) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
SMAL1_MOUSE
|
||||||
| θ value | 9.54697e-26 (rank : 34) | NC score | 0.833836 (rank : 29) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BJL0, Q8BVK7, Q8BXW4, Q8K309, Q9EQK8, Q9QYC4 | Gene names | Smarcal1, Harp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A-like protein 1 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 1) (HepA-related protein) (mharp). | |||||
|
ATRX_MOUSE
|
||||||
| θ value | 2.97466e-19 (rank : 35) | NC score | 0.691618 (rank : 36) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
ATRX_HUMAN
|
||||||
| θ value | 3.88503e-19 (rank : 36) | NC score | 0.672366 (rank : 37) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
SMAL1_HUMAN
|
||||||
| θ value | 1.29331e-14 (rank : 37) | NC score | 0.818009 (rank : 33) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NZC9, Q96AY1, Q9NXQ5, Q9UFH3, Q9UI93 | Gene names | SMARCAL1, HARP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A-like protein 1 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 1) (HepA-related protein) (hHARP). | |||||
|
KMHN1_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 38) | NC score | 0.030320 (rank : 98) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q3V125, Q3TTQ8 | Gene names | Gm172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1 homolog. | |||||
|
DDX25_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 39) | NC score | 0.042688 (rank : 93) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QY15, Q8R1B6 | Gene names | Ddx25, Grth | |||
|
Domain Architecture |
|
|||||
| Description | ATP-dependent RNA helicase DDX25 (EC 3.6.1.-) (DEAD box protein 25) (Gonadotropin-regulated testicular RNA helicase). | |||||
|
ERCC3_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 40) | NC score | 0.056605 (rank : 78) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P19447 | Gene names | ERCC3, XPB, XPBC | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase XPB subunit (EC 3.6.1.-) (Basic transcription factor 2 89 kDa subunit) (BTF2-p89) (TFIIH 89 kDa subunit) (DNA-repair protein complementing XP-B cells) (Xeroderma pigmentosum group B-complementing protein) (DNA excision repair protein ERCC-3). | |||||
|
ERCC3_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 41) | NC score | 0.053062 (rank : 87) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P49135 | Gene names | Ercc3, Xpb, Xpbc | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase XPB subunit (EC 3.6.1.-) (Basic transcription factor 2 89 kDa subunit) (BTF2-p89) (TFIIH 89 kDa subunit) (DNA-repair protein complementing XP-B cells) (Xeroderma pigmentosum group B-complementing protein) (DNA excision repair protein ERCC-3). | |||||
|
ASPM_MOUSE
|
||||||
| θ value | 0.125558 (rank : 42) | NC score | 0.018479 (rank : 106) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CJ27, O88482, Q8BJI8, Q8BKT4 | Gene names | Aspm, Calmbp1, Sha1 | |||
|
Domain Architecture |

|
|||||
| Description | Abnormal spindle-like microcephaly-associated protein homolog (Calmodulin-binding protein 1) (Spindle and hydroxyurea checkpoint abnormal protein) (Calmodulin-binding protein Sha1). | |||||
|
CE290_HUMAN
|
||||||
| θ value | 0.279714 (rank : 43) | NC score | 0.013384 (rank : 116) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
DD19A_HUMAN
|
||||||
| θ value | 0.279714 (rank : 44) | NC score | 0.040083 (rank : 95) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NUU7, Q53FM0 | Gene names | DDX19A, DDX19L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX19A (EC 3.6.1.-) (DEAD box protein 19A) (DDX19-like protein). | |||||
|
DD19A_MOUSE
|
||||||
| θ value | 0.279714 (rank : 45) | NC score | 0.040177 (rank : 94) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61655, Q921R0 | Gene names | Ddx19a, Ddx19, Eif4a-rs1 | |||
|
Domain Architecture |
|
|||||
| Description | ATP-dependent RNA helicase DDX19A (EC 3.6.1.-) (DEAD box protein 19A) (DEAD box RNA helicase DEAD5) (mDEAD5) (Eukaryotic translation initiation factor 4A-related sequence 1). | |||||
|
DD19B_HUMAN
|
||||||
| θ value | 0.279714 (rank : 46) | NC score | 0.040054 (rank : 96) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UMR2, Q6FIB7, Q6IAE0 | Gene names | DDX19B, DBP5, DDX19 | |||
|
Domain Architecture |
|
|||||
| Description | ATP-dependent RNA helicase DDX19B (EC 3.6.1.-) (DEAD box protein 19B) (DEAD box RNA helicase DEAD5). | |||||
|
DDX25_HUMAN
|
||||||
| θ value | 0.279714 (rank : 47) | NC score | 0.027489 (rank : 99) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UHL0, Q86W81, Q8IYP1 | Gene names | DDX25, GRTH | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX25 (EC 3.6.1.-) (DEAD box protein 25) (Gonadotropin-regulated testicular RNA helicase). | |||||
|
GCC2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 48) | NC score | 0.016477 (rank : 107) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 49) | NC score | 0.070283 (rank : 55) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
RBP24_HUMAN
|
||||||
| θ value | 0.47712 (rank : 50) | NC score | 0.016406 (rank : 108) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
AKAP9_HUMAN
|
||||||
| θ value | 0.62314 (rank : 51) | NC score | 0.011891 (rank : 122) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
CF170_HUMAN
|
||||||
| θ value | 0.62314 (rank : 52) | NC score | 0.024759 (rank : 102) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96NH3, Q6ZMY4, Q6ZUR7, Q8NB47 | Gene names | C6orf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf170. | |||||
|
DDX47_HUMAN
|
||||||
| θ value | 0.813845 (rank : 53) | NC score | 0.022545 (rank : 103) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H0S4, Q96GM0, Q96NV8 | Gene names | DDX47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX47 (EC 3.6.1.-) (DEAD box protein 47). | |||||
|
PRP40_MOUSE
|
||||||
| θ value | 0.813845 (rank : 54) | NC score | 0.032157 (rank : 97) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
RTN4_MOUSE
|
||||||
| θ value | 0.813845 (rank : 55) | NC score | 0.013581 (rank : 114) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99P72, Q5DTK9, Q7TNB7, Q80W95, Q8BGK7, Q8BGM9, Q8K290, Q8K3G8, Q9CTE3 | Gene names | Rtn4, Kiaa0886, Nogo | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein). | |||||
|
CCD21_HUMAN
|
||||||
| θ value | 1.06291 (rank : 56) | NC score | 0.015163 (rank : 111) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
DDX47_MOUSE
|
||||||
| θ value | 1.06291 (rank : 57) | NC score | 0.021201 (rank : 105) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9CWX9 | Gene names | Ddx47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX47 (EC 3.6.1.-) (DEAD box protein 47). | |||||
|
EMIL2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 58) | NC score | 0.011953 (rank : 121) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
PER2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 59) | NC score | 0.015657 (rank : 110) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
RYR2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 60) | NC score | 0.009881 (rank : 128) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92736, Q15411, Q546N8 | Gene names | RYR2 | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 2 (Cardiac muscle-type ryanodine receptor) (RyR2) (RYR-2) (Cardiac muscle ryanodine receptor-calcium release channel) (hRYR-2). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | 1.06291 (rank : 61) | NC score | 0.024782 (rank : 101) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
MELK_HUMAN
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | -0.004665 (rank : 145) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14680, Q7L3C3 | Gene names | MELK, KIAA0175 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Maternal embryonic leucine zipper kinase (EC 2.7.11.1) (hMELK) (Protein kinase PK38) (hPK38). | |||||
|
TPM2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.010396 (rank : 126) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 677 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P07951, P06468, Q13894, Q53FM4, Q5TCU4, Q5TCU7, Q9UH67 | Gene names | TPM2, TMSB | |||
|
Domain Architecture |
|
|||||
| Description | Tropomyosin beta chain (Tropomyosin 2) (Beta-tropomyosin). | |||||
|
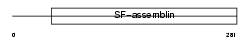
TPM2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.010568 (rank : 125) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P58774, P02560, P46901 | Gene names | Tpm2, Tpm-2 | |||
|
Domain Architecture |
|
|||||
| Description | Tropomyosin beta chain (Tropomyosin 2) (Beta-tropomyosin). | |||||
|
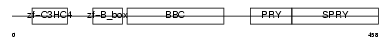
TRI39_MOUSE
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.007110 (rank : 136) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 691 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9ESN2, Q8BPR5, Q8K0F7 | Gene names | Trim39, Rnf23, Tfp | |||
|
Domain Architecture |
|
|||||
| Description | Tripartite motif-containing protein 39 (RING finger protein 23) (Testis-abundant finger protein). | |||||
|
EMIL2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 66) | NC score | 0.011342 (rank : 123) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BXX0, Q8NBH3, Q96JQ4 | Gene names | EMILIN2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Protein FOAP-10). | |||||
|
UBP34_HUMAN
|
||||||
| θ value | 3.0926 (rank : 67) | NC score | 0.007200 (rank : 135) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q70CQ2, O60316, O94834, Q3B777, Q6P6C9, Q7L8P6, Q8N3T9, Q8TBW2, Q9UGA1 | Gene names | USP34, KIAA0570, KIAA0729 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
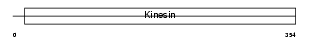
KIF11_HUMAN
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.003220 (rank : 143) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 583 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P52732, Q15716, Q5VWX0 | Gene names | KIF11, EG5, KNSL1 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF11 (Kinesin-related motor protein Eg5) (Kinesin-like spindle protein HKSP) (Thyroid receptor-interacting protein 5) (TRIP5) (Kinesin-like protein 1). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.064775 (rank : 66) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
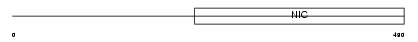
NUP93_MOUSE
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.014167 (rank : 113) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BJ71, Q3TED9, Q8BIK7, Q8K1R4, Q8R592 | Gene names | Nup93, Cip4, Kiaa0095 | |||
|
Domain Architecture |
|
|||||
| Description | Nuclear pore complex protein Nup93 (Nucleoporin Nup93) (93 kDa nucleoporin) (CBP-interacting protein 4). | |||||
|
PPR3A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.010734 (rank : 124) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99MR9, Q8BUJ4, Q8BUL0 | Gene names | Ppp1r3a, Pp1g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase 1 regulatory subunit 3A (Protein phosphatase 1 glycogen-associated regulatory subunit) (Protein phosphatase type-1 glycogen targeting subunit) (RGL). | |||||
|
ANR26_HUMAN
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.014740 (rank : 112) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1614 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UPS8, Q2TAZ3, Q6ZR14, Q9H1Q1, Q9NSK9, Q9NTD5, Q9NW69 | Gene names | ANKRD26, KIAA1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
FANCM_HUMAN
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.049260 (rank : 92) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8IYD8, Q3YFH9, Q8N9X6, Q9HCH6 | Gene names | FANCM, KIAA1596 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group M protein (EC 3.6.1.-) (ATP-dependent RNA helicase FANCM) (Protein FACM) (Fanconi anemia-associated polypeptide of 250 kDa) (FAAP250) (Protein Hef ortholog). | |||||
|
GRAM3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.012721 (rank : 119) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PEM6 | Gene names | Gramd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRAM domain-containing protein 3. | |||||
|
IFT81_MOUSE
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.013206 (rank : 117) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O35594, Q8R4W2, Q9CSA1, Q9EQM1 | Gene names | Ift81, Cdv-1, Cdv1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 81 (Carnitine deficiency-associated protein expressed in ventricle 1) (CDV-1 protein). | |||||
|
MYCL1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.007691 (rank : 133) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
NUP93_HUMAN
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.013410 (rank : 115) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N1F7, Q14705 | Gene names | NUP93, KIAA0095 | |||
|
Domain Architecture |
|
|||||
| Description | Nuclear pore complex protein Nup93 (Nucleoporin Nup93) (93 kDa nucleoporin). | |||||
|
RHG18_HUMAN
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.006536 (rank : 138) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N392, Q58EZ3, Q6P679, Q6PJD7, Q96S64 | Gene names | ARHGAP18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18 (MacGAP). | |||||
|
BCLF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.015692 (rank : 109) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NYF8, Q14673, Q86WU6, Q86WY0 | Gene names | BCLAF1, BTF, KIAA0164 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bcl-2-associated transcription factor 1 (Btf). | |||||
|
BSCL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.009202 (rank : 129) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96G97, Q96SV1, Q9BSQ0 | Gene names | BSCL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Seipin (Bernardinelli-Seip congenital lipodystrophy type 2 protein). | |||||
|
INADL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.005183 (rank : 142) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63ZW7, O70471, Q5PRG3, Q6P6J1, Q80YR8, Q8BPB9 | Gene names | Inadl, Cipp, Patj | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | InaD-like protein (Inadl protein) (Pals1-associated tight junction protein) (Protein associated to tight junctions) (Channel-interacting PDZ domain-containing protein). | |||||
|
RECQ5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.012481 (rank : 120) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
REPS1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.005955 (rank : 141) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O54916, Q8C9J9, Q99LR8 | Gene names | Reps1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 1 (RalBP1-interacting protein 1). | |||||
|
TE2IP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.013180 (rank : 118) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NYB0, Q8WYZ3, Q9NWR2 | Gene names | TERF2IP, RAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomeric repeat-binding factor 2-interacting protein 1 (TRF2- interacting telomeric protein Rap1) (hRap1). | |||||
|
WRN_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.021344 (rank : 104) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O09053, O09050, Q9JKD4, Q9Z241, Q9Z242 | Gene names | Wrn | |||
|
Domain Architecture |

|
|||||
| Description | Werner syndrome ATP-dependent helicase homolog (EC 3.6.1.-). | |||||
|
BIM_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.006255 (rank : 140) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54918, O54919, O54920 | Gene names | Bcl2l11, Bim | |||
|
Domain Architecture |

|
|||||
| Description | Bcl-2-like protein 11 (Bcl2-interacting mediator of cell death). | |||||
|
CM007_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.025225 (rank : 100) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5W0B1, Q8TBY2, Q9H0T2, Q9H8M0 | Gene names | C13orf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C13orf7. | |||||
|
DLG5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.007772 (rank : 131) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8TDM6, Q5T1H7, Q5T1H8, Q6DKG3, Q86WC0, Q8TDM7, Q9UE73, Q9Y4E3 | Gene names | DLG5, KIAA0583, PDLG | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 5 (Placenta and prostate DLG) (Discs large protein P-dlg). | |||||
|
DYDC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.007827 (rank : 130) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WWB3, Q5QP03, Q5QP04, Q6WNP4, Q6ZU87 | Gene names | DYDC1, DPY30D1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DPY30 domain-containing protein 1 (RSD-9). | |||||
|
IF4G3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.006914 (rank : 137) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O43432, Q15597, Q8NEN1 | Gene names | EIF4G3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
LAMC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.000558 (rank : 144) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P11047 | Gene names | LAMC1, LAMB2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.061687 (rank : 70) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
OMP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.007751 (rank : 132) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P47874 | Gene names | OMP | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory marker protein (Olfactory neuronal-specific protein). | |||||
|
PKHA6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.010278 (rank : 127) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
SOX13_MOUSE
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.007684 (rank : 134) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q04891, Q922L3 | Gene names | Sox13, Sox-13 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-13 protein. | |||||
|
UTRO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.006391 (rank : 139) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
AIRE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.092124 (rank : 39) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
AIRE_MOUSE
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.093736 (rank : 38) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
BAZ1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.069182 (rank : 57) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
BAZ1B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.064884 (rank : 65) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
BAZ1B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.072526 (rank : 53) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
BAZ2B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.054614 (rank : 81) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
BPTF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.087659 (rank : 40) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
CBX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.065627 (rank : 63) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P83916, P23197 | Gene names | CBX1, CBX | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25) (HP1Hsbeta) (p25beta). | |||||
|
CBX1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.065627 (rank : 64) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P83917, P23197 | Gene names | Cbx1, Cbx | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25). | |||||
|
CBX3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.056738 (rank : 77) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13185, Q96CD7, Q99409, Q9BVS3, Q9P0Z6 | Gene names | CBX3 | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (HECH). | |||||
|
CBX3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.059841 (rank : 72) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P23198, Q921C4 | Gene names | Cbx3 | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (M32). | |||||
|
CBX5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.078695 (rank : 48) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P45973 | Gene names | CBX5, HP1A | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha) (Antigen p25). | |||||
|
CBX5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.069988 (rank : 56) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61686, Q9CS35, Q9CXD1 | Gene names | Cbx5, Hp1a | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha). | |||||
|
CBX7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.057253 (rank : 76) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O95931, Q86T17 | Gene names | CBX7 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 7. | |||||
|
CBX7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.060561 (rank : 71) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VDS3 | Gene names | Cbx7, D15Ertd417e | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 7. | |||||
|
DDX41_MOUSE
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.051225 (rank : 88) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q91VN6 | Gene names | Ddx41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX41 (EC 3.6.1.-) (DEAD box protein 41). | |||||
|
DPF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.051137 (rank : 89) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.072836 (rank : 52) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.063444 (rank : 69) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
LY10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.063671 (rank : 68) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.050655 (rank : 91) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.058137 (rank : 75) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
NSD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.055674 (rank : 80) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
NSD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.058547 (rank : 74) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
PB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.053691 (rank : 84) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86U86, Q9H2T3, Q9H2T4, Q9H2T5, Q9H301, Q9H314 | Gene names | PB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein polybromo-1 (hPB1) (Polybromo-1D) (BRG1-associated factor 180) (BAF180). | |||||
|
PF21A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.079177 (rank : 47) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
PF21A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.080547 (rank : 46) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
PF21B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.081283 (rank : 45) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PF21B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.082152 (rank : 44) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PHF12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.086937 (rank : 42) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.087018 (rank : 41) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PKCB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.083362 (rank : 43) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
PL10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.053521 (rank : 85) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P16381 | Gene names | D1Pas1, Pl10 | |||
|
Domain Architecture |

|
|||||
| Description | Putative ATP-dependent RNA helicase Pl10 (EC 3.6.1.-). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.053883 (rank : 83) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.054505 (rank : 82) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
RSF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.073549 (rank : 51) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
SP110_HUMAN
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.071001 (rank : 54) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9HB58, Q14976, Q14977, Q8WUZ6, Q9HCT8 | Gene names | SP110 | |||
|
Domain Architecture |

|
|||||
| Description | Sp110 nuclear body protein (Speckled 110 kDa) (Transcriptional coactivator Sp110) (Interferon-induced protein 41/75). | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.050773 (rank : 90) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.066928 (rank : 60) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
TIF1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.063718 (rank : 67) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
TIF1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.066623 (rank : 61) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
TIF1G_HUMAN
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.058771 (rank : 73) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
TIF1G_MOUSE
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.065944 (rank : 62) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99PP7, Q6SI71, Q6ZPX5 | Gene names | Trim33, Kiaa1113 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33). | |||||
|
TRI66_HUMAN
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.067478 (rank : 58) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
TRI66_MOUSE
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.067177 (rank : 59) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
TXND2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.056107 (rank : 79) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
UHRF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.053518 (rank : 86) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96PU4, Q5VYR1, Q5VYR3, Q659C8, Q8TAG7 | Gene names | UHRF2, NIRF, RNF107 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95/ICBP90-like RING finger protein) (Np95-like RING finger protein) (Nuclear zinc finger protein Np97) (RING finger protein 107). | |||||
|
ZMY11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.077561 (rank : 49) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15326 | Gene names | ZMYND11, BS69 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11 (Adenovirus 5 E1A- binding protein) (BS69 protein). | |||||
|
ZMY11_MOUSE
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.075045 (rank : 50) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8R5C8, Q8C155 | Gene names | Zmynd11 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11. | |||||
|
SMRCD_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | Q04692 | Gene names | Smarcad1, Etl1, Kiaa1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase SMARCAD1) (Enhancer trap locus homolog 1) (Etl-1). | |||||
|
SMRCD_HUMAN
|
||||||
| NC score | 0.995177 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9H4L7, Q96SX1, Q9H017, Q9H860, Q9NPU9, Q9ULU7 | Gene names | SMARCAD1, KIAA1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase 1) (hHEL1). | |||||
|
SMCA5_MOUSE
|
||||||
| NC score | 0.953199 (rank : 3) | θ value | 6.28603e-86 (rank : 5) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q91ZW3, Q8C791, Q8CA22, Q8VDG1, Q925M9 | Gene names | Smarca5, Snf2h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 5 (EC 3.6.1.-) (Sucrose nonfermenting protein 2 homolog) (mSnf2h). | |||||
|
SMCA5_HUMAN
|
||||||
| NC score | 0.953181 (rank : 4) | θ value | 6.28603e-86 (rank : 4) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O60264 | Gene names | SMARCA5, SNF2H, WCRF135 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 5 (EC 3.6.1.-) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin A5) (Sucrose nonfermenting protein 2 homolog) (hSNF2H). | |||||
|
SMCA1_HUMAN
|
||||||
| NC score | 0.952115 (rank : 5) | θ value | 9.07703e-85 (rank : 6) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P28370, Q5JV41, Q5JV42 | Gene names | SMARCA1, SNF2L, SNF2L1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L1 (EC 3.6.1.-) (Nucleosome remodeling factor subunit SNF2L) (ATP-dependent helicase SMARCA1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 1). | |||||
|
SMCA1_MOUSE
|
||||||
| NC score | 0.950148 (rank : 6) | θ value | 1.49618e-87 (rank : 3) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6PGB8, Q8BSS1, Q91Y58 | Gene names | Smarca1, Snf2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable global transcription activator SNF2L1 (EC 3.6.1.-) (Nucleosome remodeling factor subunit SNF2L) (ATP-dependent helicase SMARCA1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 1) (DNA-dependent ATPase SNF2L). | |||||
|
BTAF1_HUMAN
|
||||||
| NC score | 0.942626 (rank : 7) | θ value | 2.7521e-57 (rank : 22) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O14981, O43578 | Gene names | BTAF1, TAF172 | |||
|
Domain Architecture |

|
|||||
| Description | TATA-binding protein-associated factor 172 (EC 3.6.1.-) (ATP-dependent helicase BTAF1) (TBP-associated factor 172) (TAF-172) (TAF(II)170) (B- TFIID transcription factor-associated 170 kDa subunit). | |||||
|
ERCC6_HUMAN
|
||||||
| NC score | 0.938587 (rank : 8) | θ value | 1.50663e-63 (rank : 20) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q03468 | Gene names | ERCC6, CSB | |||
|
Domain Architecture |

|
|||||
| Description | DNA excision repair protein ERCC-6 (EC 3.6.1.-) (ATP-dependent helicase ERCC6) (Cockayne syndrome protein CSB). | |||||
|
RA54B_HUMAN
|
||||||
| NC score | 0.935527 (rank : 9) | θ value | 8.55514e-59 (rank : 21) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y620 | Gene names | RAD54B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54B (EC 3.6.1.-) (RAD54 homolog B). | |||||
|
RA54B_MOUSE
|
||||||
| NC score | 0.932349 (rank : 10) | θ value | 9.16162e-53 (rank : 25) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6PFE3 | Gene names | Rad54b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54B (EC 3.6.1.-) (RAD54 homolog B). | |||||
|
CHD2_HUMAN
|
||||||
| NC score | 0.929053 (rank : 11) | θ value | 2.64714e-76 (rank : 11) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O14647 | Gene names | CHD2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 2 (EC 3.6.1.-) (ATP- dependent helicase CHD2) (CHD-2). | |||||
|
RAD54_HUMAN
|
||||||
| NC score | 0.928853 (rank : 12) | θ value | 1.97686e-47 (rank : 26) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q92698, Q6IUY3 | Gene names | RAD54L, RAD54A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54-like (EC 3.6.1.-) (RAD54 homolog) (hRAD54) (hHR54). | |||||
|
CHD8_HUMAN
|
||||||
| NC score | 0.928051 (rank : 13) | θ value | 1.35953e-72 (rank : 13) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9HCK8, Q68DQ0, Q8N3Z9, Q8NCY4, Q8TBR9, Q96F26 | Gene names | CHD8, HELSNF1, KIAA1564 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 8 (EC 3.6.1.-) (ATP- dependent helicase CHD8) (CHD-8) (Helicase with SNF2 domain 1). | |||||
|
CHD1_MOUSE
|
||||||
| NC score | 0.927774 (rank : 14) | θ value | 2.82823e-78 (rank : 7) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
RAD54_MOUSE
|
||||||
| NC score | 0.927665 (rank : 15) | θ value | 7.03111e-45 (rank : 27) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P70270, Q8BSR5, Q8C2C4, Q8K3D4 | Gene names | Rad54l, Rad54 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54-like (EC 3.6.1.-) (RAD54 homolog) (mRAD54) (mHR54). | |||||
|
CHD1_HUMAN
|
||||||
| NC score | 0.923753 (rank : 16) | θ value | 3.69378e-78 (rank : 8) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CHD6_HUMAN
|
||||||
| NC score | 0.922064 (rank : 17) | θ value | 2.09745e-73 (rank : 12) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8TD26, Q5JYQ0, Q5TGZ9, Q5TH00, Q5TH01, Q8IZR2, Q8WTY0, Q9H4H6, Q9H6D4, Q9NTT7, Q9P2L1 | Gene names | CHD6, CHD5, KIAA1335, RIGB | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 6 (EC 3.6.1.-) (ATP- dependent helicase CHD6) (CHD-6) (Radiation-induced gene B protein). | |||||
|
INOC1_MOUSE
|
||||||
| NC score | 0.919634 (rank : 18) | θ value | 1.36585e-56 (rank : 23) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6ZPV2, Q6P7V0, Q6PCP1, Q8C9T7 | Gene names | Inoc1, Kiaa1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-). | |||||
|
INOC1_HUMAN
|
||||||
| NC score | 0.919261 (rank : 19) | θ value | 1.78386e-56 (rank : 24) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9ULG1, Q9NTG6 | Gene names | INOC1, KIAA1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-) (hINO80). | |||||
|
CHD9_HUMAN
|
||||||
| NC score | 0.916691 (rank : 20) | θ value | 8.81223e-72 (rank : 14) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
CHD9_MOUSE
|
||||||
| NC score | 0.911771 (rank : 21) | θ value | 1.15091e-71 (rank : 15) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
SMCA2_HUMAN
|
||||||
| NC score | 0.901220 (rank : 22) | θ value | 3.12697e-77 (rank : 9) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
SMCA4_HUMAN
|
||||||
| NC score | 0.890758 (rank : 23) | θ value | 5.33382e-77 (rank : 10) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
TTF2_HUMAN
|
||||||
| NC score | 0.872378 (rank : 24) | θ value | 3.3877e-31 (rank : 32) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9UNY4, O75921, Q5T2K7, Q5VVU8, Q8N6I8 | Gene names | TTF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2) (Factor 2) (F2) (HuF2) (Lodestar homolog). | |||||
|
CHD3_HUMAN
|
||||||
| NC score | 0.860005 (rank : 25) | θ value | 1.6097e-65 (rank : 19) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
CHD4_HUMAN
|
||||||
| NC score | 0.859771 (rank : 26) | θ value | 3.83135e-67 (rank : 17) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
CHD5_HUMAN
|
||||||
| NC score | 0.856509 (rank : 27) | θ value | 7.71989e-68 (rank : 16) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
CHD4_MOUSE
|
||||||
| NC score | 0.855748 (rank : 28) | θ value | 3.83135e-67 (rank : 18) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
SMAL1_MOUSE
|
||||||
| NC score | 0.833836 (rank : 29) | θ value | 9.54697e-26 (rank : 34) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BJL0, Q8BVK7, Q8BXW4, Q8K309, Q9EQK8, Q9QYC4 | Gene names | Smarcal1, Harp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A-like protein 1 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 1) (HepA-related protein) (mharp). | |||||
|
EP400_HUMAN
|
||||||
| NC score | 0.830562 (rank : 30) | θ value | 2.50074e-42 (rank : 28) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
TTF2_MOUSE
|
||||||
| NC score | 0.824645 (rank : 31) | θ value | 1.57365e-28 (rank : 33) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
EP400_MOUSE
|
||||||
| NC score | 0.821176 (rank : 32) | θ value | 1.16164e-39 (rank : 29) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
SMAL1_HUMAN
|
||||||
| NC score | 0.818009 (rank : 33) | θ value | 1.29331e-14 (rank : 37) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NZC9, Q96AY1, Q9NXQ5, Q9UFH3, Q9UI93 | Gene names | SMARCAL1, HARP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A-like protein 1 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 1) (HepA-related protein) (hHARP). | |||||
|
SMRA3_MOUSE
|
||||||
| NC score | 0.768519 (rank : 34) | θ value | 3.4976e-36 (rank : 30) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6PCN7, O35596, O35597, Q80VT6 | Gene names | Smarca3, Snf2l3, Zbu1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 3 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 3) (TNF-response element-binding protein) (P113). | |||||
|
SMRA3_HUMAN
|
||||||
| NC score | 0.765114 (rank : 35) | θ value | 1.12514e-34 (rank : 31) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q14527, Q14536, Q16051, Q7KYJ6, Q86YA5, Q92652, Q96KM9 | Gene names | SMARCA3, HIP116A, HLTF, SNF2L3, ZBU1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 3 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 3) (DNA-binding protein/plasminogen activator inhibitor 1 regulator) (Helicase-like transcription factor) (HIP116). | |||||
|
ATRX_MOUSE
|
||||||
| NC score | 0.691618 (rank : 36) | θ value | 2.97466e-19 (rank : 35) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
ATRX_HUMAN
|
||||||
| NC score | 0.672366 (rank : 37) | θ value | 3.88503e-19 (rank : 36) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
AIRE_MOUSE
|
||||||
| NC score | 0.093736 (rank : 38) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
AIRE_HUMAN
|
||||||
| NC score | 0.092124 (rank : 39) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
BPTF_HUMAN
|
||||||
| NC score | 0.087659 (rank : 40) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
PHF12_MOUSE
|
||||||
| NC score | 0.087018 (rank : 41) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF12_HUMAN
|
||||||
| NC score | 0.086937 (rank : 42) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PKCB1_HUMAN
|
||||||
| NC score | 0.083362 (rank : 43) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
PF21B_MOUSE
|
||||||
| NC score | 0.082152 (rank : 44) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PF21B_HUMAN
|
||||||
| NC score | 0.081283 (rank : 45) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PF21A_MOUSE
|
||||||
| NC score | 0.080547 (rank : 46) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
PF21A_HUMAN
|
||||||
| NC score | 0.079177 (rank : 47) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
CBX5_HUMAN
|
||||||
| NC score | 0.078695 (rank : 48) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P45973 | Gene names | CBX5, HP1A | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha) (Antigen p25). | |||||
|
ZMY11_HUMAN
|
||||||
| NC score | 0.077561 (rank : 49) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15326 | Gene names | ZMYND11, BS69 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11 (Adenovirus 5 E1A- binding protein) (BS69 protein). | |||||
|
ZMY11_MOUSE
|
||||||
| NC score | 0.075045 (rank : 50) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8R5C8, Q8C155 | Gene names | Zmynd11 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11. | |||||
|
RSF1_HUMAN
|
||||||
| NC score | 0.073549 (rank : 51) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.072836 (rank : 52) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
BAZ1B_MOUSE
|
||||||
| NC score | 0.072526 (rank : 53) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
SP110_HUMAN
|
||||||
| NC score | 0.071001 (rank : 54) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9HB58, Q14976, Q14977, Q8WUZ6, Q9HCT8 | Gene names | SP110 | |||
|
Domain Architecture |

|
|||||
| Description | Sp110 nuclear body protein (Speckled 110 kDa) (Transcriptional coactivator Sp110) (Interferon-induced protein 41/75). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.070283 (rank : 55) | θ value | 0.47712 (rank : 49) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
CBX5_MOUSE
|
||||||
| NC score | 0.069988 (rank : 56) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61686, Q9CS35, Q9CXD1 | Gene names | Cbx5, Hp1a | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha). | |||||
|
BAZ1A_HUMAN
|
||||||
| NC score | 0.069182 (rank : 57) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
TRI66_HUMAN
|
||||||
| NC score | 0.067478 (rank : 58) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
TRI66_MOUSE
|
||||||
| NC score | 0.067177 (rank : 59) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.066928 (rank : 60) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
TIF1A_MOUSE
|
||||||
| NC score | 0.066623 (rank : 61) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
TIF1G_MOUSE
|
||||||
| NC score | 0.065944 (rank : 62) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99PP7, Q6SI71, Q6ZPX5 | Gene names | Trim33, Kiaa1113 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33). | |||||
|
CBX1_HUMAN
|
||||||
| NC score | 0.065627 (rank : 63) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P83916, P23197 | Gene names | CBX1, CBX | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25) (HP1Hsbeta) (p25beta). | |||||
|
CBX1_MOUSE
|
||||||
| NC score | 0.065627 (rank : 64) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P83917, P23197 | Gene names | Cbx1, Cbx | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25). | |||||
|
BAZ1B_HUMAN
|
||||||
| NC score | 0.064884 (rank : 65) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.064775 (rank : 66) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
TIF1A_HUMAN
|
||||||
| NC score | 0.063718 (rank : 67) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
LY10_HUMAN
|
||||||
| NC score | 0.063671 (rank : 68) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.063444 (rank : 69) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.061687 (rank : 70) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
CBX7_MOUSE
|
||||||
| NC score | 0.060561 (rank : 71) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VDS3 | Gene names | Cbx7, D15Ertd417e | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 7. | |||||
|
CBX3_MOUSE
|
||||||
| NC score | 0.059841 (rank : 72) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P23198, Q921C4 | Gene names | Cbx3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (M32). | |||||
|
TIF1G_HUMAN
|
||||||
| NC score | 0.058771 (rank : 73) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
NSD1_MOUSE
|
||||||
| NC score | 0.058547 (rank : 74) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.058137 (rank : 75) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
CBX7_HUMAN
|
||||||
| NC score | 0.057253 (rank : 76) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O95931, Q86T17 | Gene names | CBX7 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 7. | |||||
|
CBX3_HUMAN
|
||||||
| NC score | 0.056738 (rank : 77) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13185, Q96CD7, Q99409, Q9BVS3, Q9P0Z6 | Gene names | CBX3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (HECH). | |||||
|
ERCC3_HUMAN
|
||||||
| NC score | 0.056605 (rank : 78) | θ value | 0.0431538 (rank : 40) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P19447 | Gene names | ERCC3, XPB, XPBC | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase XPB subunit (EC 3.6.1.-) (Basic transcription factor 2 89 kDa subunit) (BTF2-p89) (TFIIH 89 kDa subunit) (DNA-repair protein complementing XP-B cells) (Xeroderma pigmentosum group B-complementing protein) (DNA excision repair protein ERCC-3). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.056107 (rank : 79) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.055674 (rank : 80) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
BAZ2B_HUMAN
|
||||||
| NC score | 0.054614 (rank : 81) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.054505 (rank : 82) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.053883 (rank : 83) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
PB1_HUMAN
|
||||||
| NC score | 0.053691 (rank : 84) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86U86, Q9H2T3, Q9H2T4, Q9H2T5, Q9H301, Q9H314 | Gene names | PB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein polybromo-1 (hPB1) (Polybromo-1D) (BRG1-associated factor 180) (BAF180). | |||||
|
PL10_MOUSE
|
||||||
| NC score | 0.053521 (rank : 85) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P16381 | Gene names | D1Pas1, Pl10 | |||
|
Domain Architecture |

|
|||||
| Description | Putative ATP-dependent RNA helicase Pl10 (EC 3.6.1.-). | |||||
|
UHRF2_HUMAN
|
||||||
| NC score | 0.053518 (rank : 86) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96PU4, Q5VYR1, Q5VYR3, Q659C8, Q8TAG7 | Gene names | UHRF2, NIRF, RNF107 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95/ICBP90-like RING finger protein) (Np95-like RING finger protein) (Nuclear zinc finger protein Np97) (RING finger protein 107). | |||||
|
ERCC3_MOUSE
|
||||||
| NC score | 0.053062 (rank : 87) | θ value | 0.0431538 (rank : 41) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P49135 | Gene names | Ercc3, Xpb, Xpbc | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase XPB subunit (EC 3.6.1.-) (Basic transcription factor 2 89 kDa subunit) (BTF2-p89) (TFIIH 89 kDa subunit) (DNA-repair protein complementing XP-B cells) (Xeroderma pigmentosum group B-complementing protein) (DNA excision repair protein ERCC-3). | |||||
|
DDX41_MOUSE
|
||||||
| NC score | 0.051225 (rank : 88) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q91VN6 | Gene names | Ddx41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX41 (EC 3.6.1.-) (DEAD box protein 41). | |||||
|
DPF1_HUMAN
|
||||||
| NC score | 0.051137 (rank : 89) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.050773 (rank : 90) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.050655 (rank : 91) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
FANCM_HUMAN
|
||||||
| NC score | 0.049260 (rank : 92) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8IYD8, Q3YFH9, Q8N9X6, Q9HCH6 | Gene names | FANCM, KIAA1596 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group M protein (EC 3.6.1.-) (ATP-dependent RNA helicase FANCM) (Protein FACM) (Fanconi anemia-associated polypeptide of 250 kDa) (FAAP250) (Protein Hef ortholog). | |||||
|
DDX25_MOUSE
|
||||||
| NC score | 0.042688 (rank : 93) | θ value | 0.0330416 (rank : 39) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QY15, Q8R1B6 | Gene names | Ddx25, Grth | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX25 (EC 3.6.1.-) (DEAD box protein 25) (Gonadotropin-regulated testicular RNA helicase). | |||||
|
DD19A_MOUSE
|
||||||
| NC score | 0.040177 (rank : 94) | θ value | 0.279714 (rank : 45) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61655, Q921R0 | Gene names | Ddx19a, Ddx19, Eif4a-rs1 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX19A (EC 3.6.1.-) (DEAD box protein 19A) (DEAD box RNA helicase DEAD5) (mDEAD5) (Eukaryotic translation initiation factor 4A-related sequence 1). | |||||
|
DD19A_HUMAN
|
||||||
| NC score | 0.040083 (rank : 95) | θ value | 0.279714 (rank : 44) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NUU7, Q53FM0 | Gene names | DDX19A, DDX19L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX19A (EC 3.6.1.-) (DEAD box protein 19A) (DDX19-like protein). | |||||
|
DD19B_HUMAN
|
||||||
| NC score | 0.040054 (rank : 96) | θ value | 0.279714 (rank : 46) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UMR2, Q6FIB7, Q6IAE0 | Gene names | DDX19B, DBP5, DDX19 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX19B (EC 3.6.1.-) (DEAD box protein 19B) (DEAD box RNA helicase DEAD5). | |||||
|
PRP40_MOUSE
|
||||||
| NC score | 0.032157 (rank : 97) | θ value | 0.813845 (rank : 54) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
KMHN1_MOUSE
|
||||||
| NC score | 0.030320 (rank : 98) | θ value | 0.000461057 (rank : 38) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q3V125, Q3TTQ8 | Gene names | Gm172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1 homolog. | |||||
|
DDX25_HUMAN
|
||||||
| NC score | 0.027489 (rank : 99) | θ value | 0.279714 (rank : 47) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UHL0, Q86W81, Q8IYP1 | Gene names | DDX25, GRTH | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX25 (EC 3.6.1.-) (DEAD box protein 25) (Gonadotropin-regulated testicular RNA helicase). | |||||
|
CM007_HUMAN
|
||||||
| NC score | 0.025225 (rank : 100) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5W0B1, Q8TBY2, Q9H0T2, Q9H8M0 | Gene names | C13orf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C13orf7. | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.024782 (rank : 101) | θ value | 1.06291 (rank : 61) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
CF170_HUMAN
|
||||||
| NC score | 0.024759 (rank : 102) | θ value | 0.62314 (rank : 52) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96NH3, Q6ZMY4, Q6ZUR7, Q8NB47 | Gene names | C6orf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf170. | |||||
|
DDX47_HUMAN
|
||||||
| NC score | 0.022545 (rank : 103) | θ value | 0.813845 (rank : 53) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H0S4, Q96GM0, Q96NV8 | Gene names | DDX47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX47 (EC 3.6.1.-) (DEAD box protein 47). | |||||
|
WRN_MOUSE
|
||||||
| NC score | 0.021344 (rank : 104) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O09053, O09050, Q9JKD4, Q9Z241, Q9Z242 | Gene names | Wrn | |||
|
Domain Architecture |

|
|||||
| Description | Werner syndrome ATP-dependent helicase homolog (EC 3.6.1.-). | |||||
|
DDX47_MOUSE
|
||||||
| NC score | 0.021201 (rank : 105) | θ value | 1.06291 (rank : 57) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9CWX9 | Gene names | Ddx47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX47 (EC 3.6.1.-) (DEAD box protein 47). | |||||
|
ASPM_MOUSE
|
||||||
| NC score | 0.018479 (rank : 106) | θ value | 0.125558 (rank : 42) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CJ27, O88482, Q8BJI8, Q8BKT4 | Gene names | Aspm, Calmbp1, Sha1 | |||
|
Domain Architecture |

|
|||||
| Description | Abnormal spindle-like microcephaly-associated protein homolog (Calmodulin-binding protein 1) (Spindle and hydroxyurea checkpoint abnormal protein) (Calmodulin-binding protein Sha1). | |||||
|
GCC2_HUMAN
|
||||||
| NC score | 0.016477 (rank : 107) | θ value | 0.47712 (rank : 48) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
RBP24_HUMAN
|
||||||
| NC score | 0.016406 (rank : 108) | θ value | 0.47712 (rank : 50) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
BCLF1_HUMAN
|
||||||
| NC score | 0.015692 (rank : 109) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NYF8, Q14673, Q86WU6, Q86WY0 | Gene names | BCLAF1, BTF, KIAA0164 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bcl-2-associated transcription factor 1 (Btf). | |||||
|
PER2_HUMAN
|
||||||
| NC score | 0.015657 (rank : 110) | θ value | 1.06291 (rank : 59) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
CCD21_HUMAN
|
||||||
| NC score | 0.015163 (rank : 111) | θ value | 1.06291 (rank : 56) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
ANR26_HUMAN
|
||||||
| NC score | 0.014740 (rank : 112) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1614 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UPS8, Q2TAZ3, Q6ZR14, Q9H1Q1, Q9NSK9, Q9NTD5, Q9NW69 | Gene names | ANKRD26, KIAA1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
NUP93_MOUSE
|
||||||
| NC score | 0.014167 (rank : 113) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BJ71, Q3TED9, Q8BIK7, Q8K1R4, Q8R592 | Gene names | Nup93, Cip4, Kiaa0095 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup93 (Nucleoporin Nup93) (93 kDa nucleoporin) (CBP-interacting protein 4). | |||||
|
RTN4_MOUSE
|
||||||
| NC score | 0.013581 (rank : 114) | θ value | 0.813845 (rank : 55) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99P72, Q5DTK9, Q7TNB7, Q80W95, Q8BGK7, Q8BGM9, Q8K290, Q8K3G8, Q9CTE3 | Gene names | Rtn4, Kiaa0886, Nogo | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein). | |||||
|
NUP93_HUMAN
|
||||||
| NC score | 0.013410 (rank : 115) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N1F7, Q14705 | Gene names | NUP93, KIAA0095 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup93 (Nucleoporin Nup93) (93 kDa nucleoporin). | |||||
|
CE290_HUMAN
|
||||||
| NC score | 0.013384 (rank : 116) | θ value | 0.279714 (rank : 43) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
IFT81_MOUSE
|
||||||
| NC score | 0.013206 (rank : 117) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O35594, Q8R4W2, Q9CSA1, Q9EQM1 | Gene names | Ift81, Cdv-1, Cdv1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 81 (Carnitine deficiency-associated protein expressed in ventricle 1) (CDV-1 protein). | |||||
|
TE2IP_HUMAN
|
||||||
| NC score | 0.013180 (rank : 118) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NYB0, Q8WYZ3, Q9NWR2 | Gene names | TERF2IP, RAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomeric repeat-binding factor 2-interacting protein 1 (TRF2- interacting telomeric protein Rap1) (hRap1). | |||||
|
GRAM3_MOUSE
|
||||||
| NC score | 0.012721 (rank : 119) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PEM6 | Gene names | Gramd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRAM domain-containing protein 3. | |||||
|
RECQ5_HUMAN
|
||||||
| NC score | 0.012481 (rank : 120) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
EMIL2_MOUSE
|
||||||
| NC score | 0.011953 (rank : 121) | θ value | 1.06291 (rank : 58) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
AKAP9_HUMAN
|
||||||
| NC score | 0.011891 (rank : 122) | θ value | 0.62314 (rank : 51) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
EMIL2_HUMAN
|
||||||
| NC score | 0.011342 (rank : 123) | θ value | 3.0926 (rank : 66) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BXX0, Q8NBH3, Q96JQ4 | Gene names | EMILIN2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Protein FOAP-10). | |||||
|
PPR3A_MOUSE
|
||||||
| NC score | 0.010734 (rank : 124) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99MR9, Q8BUJ4, Q8BUL0 | Gene names | Ppp1r3a, Pp1g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase 1 regulatory subunit 3A (Protein phosphatase 1 glycogen-associated regulatory subunit) (Protein phosphatase type-1 glycogen targeting subunit) (RGL). | |||||
|
TPM2_MOUSE
|
||||||
| NC score | 0.010568 (rank : 125) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P58774, P02560, P46901 | Gene names | Tpm2, Tpm-2 | |||
|
Domain Architecture |

|
|||||
| Description | Tropomyosin beta chain (Tropomyosin 2) (Beta-tropomyosin). | |||||
|
TPM2_HUMAN
|
||||||
| NC score | 0.010396 (rank : 126) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 677 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P07951, P06468, Q13894, Q53FM4, Q5TCU4, Q5TCU7, Q9UH67 | Gene names | TPM2, TMSB | |||
|
Domain Architecture |

|
|||||
| Description | Tropomyosin beta chain (Tropomyosin 2) (Beta-tropomyosin). | |||||
|
PKHA6_MOUSE
|
||||||
| NC score | 0.010278 (rank : 127) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
RYR2_HUMAN
|
||||||
| NC score | 0.009881 (rank : 128) | θ value | 1.06291 (rank : 60) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92736, Q15411, Q546N8 | Gene names | RYR2 | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 2 (Cardiac muscle-type ryanodine receptor) (RyR2) (RYR-2) (Cardiac muscle ryanodine receptor-calcium release channel) (hRYR-2). | |||||
|
BSCL2_HUMAN
|
||||||
| NC score | 0.009202 (rank : 129) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96G97, Q96SV1, Q9BSQ0 | Gene names | BSCL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Seipin (Bernardinelli-Seip congenital lipodystrophy type 2 protein). | |||||
|
DYDC1_HUMAN
|
||||||
| NC score | 0.007827 (rank : 130) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WWB3, Q5QP03, Q5QP04, Q6WNP4, Q6ZU87 | Gene names | DYDC1, DPY30D1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DPY30 domain-containing protein 1 (RSD-9). | |||||
|
DLG5_HUMAN
|
||||||
| NC score | 0.007772 (rank : 131) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8TDM6, Q5T1H7, Q5T1H8, Q6DKG3, Q86WC0, Q8TDM7, Q9UE73, Q9Y4E3 | Gene names | DLG5, KIAA0583, PDLG | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 5 (Placenta and prostate DLG) (Discs large protein P-dlg). | |||||
|
OMP_HUMAN
|
||||||
| NC score | 0.007751 (rank : 132) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P47874 | Gene names | OMP | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory marker protein (Olfactory neuronal-specific protein). | |||||
|
MYCL1_HUMAN
|
||||||
| NC score | 0.007691 (rank : 133) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
SOX13_MOUSE
|
||||||
| NC score | 0.007684 (rank : 134) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q04891, Q922L3 | Gene names | Sox13, Sox-13 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-13 protein. | |||||
|
UBP34_HUMAN
|
||||||
| NC score | 0.007200 (rank : 135) | θ value | 3.0926 (rank : 67) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q70CQ2, O60316, O94834, Q3B777, Q6P6C9, Q7L8P6, Q8N3T9, Q8TBW2, Q9UGA1 | Gene names | USP34, KIAA0570, KIAA0729 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
TRI39_MOUSE
|
||||||
| NC score | 0.007110 (rank : 136) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 691 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9ESN2, Q8BPR5, Q8K0F7 | Gene names | Trim39, Rnf23, Tfp | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 39 (RING finger protein 23) (Testis-abundant finger protein). | |||||
|
IF4G3_HUMAN
|
||||||
| NC score | 0.006914 (rank : 137) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O43432, Q15597, Q8NEN1 | Gene names | EIF4G3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
RHG18_HUMAN
|
||||||
| NC score | 0.006536 (rank : 138) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N392, Q58EZ3, Q6P679, Q6PJD7, Q96S64 | Gene names | ARHGAP18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18 (MacGAP). | |||||
|
UTRO_HUMAN
|
||||||
| NC score | 0.006391 (rank : 139) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
BIM_MOUSE
|
||||||
| NC score | 0.006255 (rank : 140) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54918, O54919, O54920 | Gene names | Bcl2l11, Bim | |||
|
Domain Architecture |

|
|||||
| Description | Bcl-2-like protein 11 (Bcl2-interacting mediator of cell death). | |||||
|
REPS1_MOUSE
|
||||||
| NC score | 0.005955 (rank : 141) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O54916, Q8C9J9, Q99LR8 | Gene names | Reps1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 1 (RalBP1-interacting protein 1). | |||||
|
INADL_MOUSE
|
||||||
| NC score | 0.005183 (rank : 142) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63ZW7, O70471, Q5PRG3, Q6P6J1, Q80YR8, Q8BPB9 | Gene names | Inadl, Cipp, Patj | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | InaD-like protein (Inadl protein) (Pals1-associated tight junction protein) (Protein associated to tight junctions) (Channel-interacting PDZ domain-containing protein). | |||||
|
KIF11_HUMAN
|
||||||
| NC score | 0.003220 (rank : 143) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 583 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P52732, Q15716, Q5VWX0 | Gene names | KIF11, EG5, KNSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF11 (Kinesin-related motor protein Eg5) (Kinesin-like spindle protein HKSP) (Thyroid receptor-interacting protein 5) (TRIP5) (Kinesin-like protein 1). | |||||
|
LAMC1_HUMAN
|
||||||
| NC score | 0.000558 (rank : 144) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P11047 | Gene names | LAMC1, LAMB2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
MELK_HUMAN
|
||||||
| NC score | -0.004665 (rank : 145) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14680, Q7L3C3 | Gene names | MELK, KIAA0175 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Maternal embryonic leucine zipper kinase (EC 2.7.11.1) (hMELK) (Protein kinase PK38) (hPK38). | |||||