Please be patient as the page loads
|
SKIV2_HUMAN
|
||||||
| SwissProt Accessions | Q15477, O15005, Q12902, Q15476 | Gene names | SKIV2L, DDX13, SKI2W, SKIV2, W | |||
|
Domain Architecture |

|
|||||
| Description | Helicase SKI2W (EC 3.6.1.-) (Helicase-like protein) (HLP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SKIV2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q15477, O15005, Q12902, Q15476 | Gene names | SKIV2L, DDX13, SKI2W, SKIV2, W | |||
|
Domain Architecture |

|
|||||
| Description | Helicase SKI2W (EC 3.6.1.-) (Helicase-like protein) (HLP). | |||||
|
SK2L2_MOUSE
|
||||||
| θ value | 3.26429e-159 (rank : 2) | NC score | 0.936298 (rank : 3) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9CZU3 | Gene names | Skiv2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Superkiller viralicidic activity 2-like 2 (EC 3.6.1.-) (ATP-dependent helicase SKIV2L2). | |||||
|
SK2L2_HUMAN
|
||||||
| θ value | 1.79118e-157 (rank : 3) | NC score | 0.937283 (rank : 2) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P42285, Q8TAG2 | Gene names | SKIV2L2, KIAA0052 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Superkiller viralicidic activity 2-like 2 (EC 3.6.1.-) (ATP-dependent helicase SKIV2L2). | |||||
|
U520_HUMAN
|
||||||
| θ value | 1.05554e-24 (rank : 4) | NC score | 0.560851 (rank : 5) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75643, O94884, Q6PX59, Q96IF2, Q9H7S0 | Gene names | ASCC3L1, HELIC2, KIAA0788 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U5 small nuclear ribonucleoprotein 200 kDa helicase (EC 3.6.1.-) (U5 snRNP-specific 200 kDa protein) (U5-200KD) (Activating signal cointegrator 1 complex subunit 3-like 1) (BRR2 homolog). | |||||
|
HELC1_HUMAN
|
||||||
| θ value | 2.87452e-22 (rank : 5) | NC score | 0.569619 (rank : 4) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N3C0, O43738, Q9H1I9, Q9NTR0 | Gene names | ASCC3, HELIC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 3 (EC 3.6.1.-) (ASC-1 complex subunit p200) (Trip4 complex subunit p200) (Helicase, ATP binding 1). | |||||
|
DDX54_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 6) | NC score | 0.087577 (rank : 12) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TDD1, Q86YT8, Q9BRZ1 | Gene names | DDX54 | |||
|
Domain Architecture |
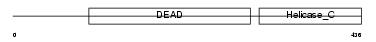
|
|||||
| Description | ATP-dependent RNA helicase DDX54 (EC 3.6.1.-) (DEAD box protein 54) (ATP-dependent RNA helicase DP97). | |||||
|
P3H3_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 7) | NC score | 0.052444 (rank : 25) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CG70, O88836 | Gene names | Leprel2, P3h3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 3 precursor (EC 1.14.11.7) (Leprecan-like protein 2) (Protein B). | |||||
|
DDX54_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 8) | NC score | 0.084729 (rank : 13) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8K4L0 | Gene names | Ddx54 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX54 (EC 3.6.1.-) (DEAD box protein 54). | |||||
|
RECQ1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 9) | NC score | 0.143147 (rank : 6) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P46063 | Gene names | RECQL, RECQL1 | |||
|
Domain Architecture |
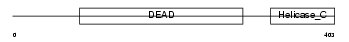
|
|||||
| Description | ATP-dependent DNA helicase Q1 (EC 3.6.1.-) (DNA-dependent ATPase Q1). | |||||
|
RECQ1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 10) | NC score | 0.125131 (rank : 7) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Z129, Q9Z128 | Gene names | Recql, Recql1 | |||
|
Domain Architecture |
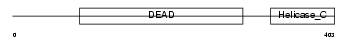
|
|||||
| Description | ATP-dependent DNA helicase Q1 (EC 3.6.1.-) (DNA-dependent ATPase Q1). | |||||
|
WRN_HUMAN
|
||||||
| θ value | 0.163984 (rank : 11) | NC score | 0.121814 (rank : 8) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14191 | Gene names | WRN, RECQ3, RECQL2 | |||
|
Domain Architecture |

|
|||||
| Description | Werner syndrome ATP-dependent helicase (EC 3.6.1.-). | |||||
|
EPN3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 12) | NC score | 0.032846 (rank : 33) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91W69, Q9CV55 | Gene names | Epn3 | |||
|
Domain Architecture |
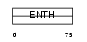
|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
RECQ4_HUMAN
|
||||||
| θ value | 0.279714 (rank : 13) | NC score | 0.118705 (rank : 9) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O94761, Q3Y424, Q96DW2, Q96F55 | Gene names | RECQL4, RECQ4 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4) (RecQ4) (RTS). | |||||
|
RECQ4_MOUSE
|
||||||
| θ value | 0.365318 (rank : 14) | NC score | 0.114891 (rank : 10) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q75NR7, Q76MT1, Q99PV9 | Gene names | Recql4, Recq4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4). | |||||
|
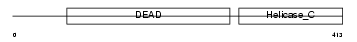
DDX10_HUMAN
|
||||||
| θ value | 0.47712 (rank : 15) | NC score | 0.061552 (rank : 17) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q13206, Q5BJD8 | Gene names | DDX10 | |||
|
Domain Architecture |
|
|||||
| Description | Probable ATP-dependent RNA helicase DDX10 (EC 3.6.1.-) (DEAD box protein 10). | |||||
|
DDX50_HUMAN
|
||||||
| θ value | 0.47712 (rank : 16) | NC score | 0.056538 (rank : 20) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BQ39, Q8WV76, Q9BWI8 | Gene names | DDX50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX50 (EC 3.6.1.-) (DEAD box protein 50) (Nucleolar protein Gu2) (Gu-beta). | |||||
|
DDX50_MOUSE
|
||||||
| θ value | 0.47712 (rank : 17) | NC score | 0.056481 (rank : 21) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q99MJ9 | Gene names | Ddx50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX50 (EC 3.6.1.-) (DEAD box protein 50) (Nucleolar protein Gu2) (Gu-beta). | |||||
|
DHX36_HUMAN
|
||||||
| θ value | 0.62314 (rank : 18) | NC score | 0.033472 (rank : 32) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H2U1, Q70JU3, Q8IYE5, Q9P240 | Gene names | DHX36, DDX36, KIAA1488, MLEL1, RHAU | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DHX36 (EC 3.6.1.-) (DEAH box protein 36) (MLE-like protein 1) (RNA helicase associated with AU-rich element ARE). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.62314 (rank : 19) | NC score | 0.015782 (rank : 40) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
DDX18_HUMAN
|
||||||
| θ value | 1.38821 (rank : 20) | NC score | 0.049685 (rank : 30) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NVP1, Q6IAU4, Q92732, Q9BQB7 | Gene names | DDX18 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX18 (EC 3.6.1.-) (DEAD box protein 18) (Myc-regulated DEAD box protein) (MrDb). | |||||
|
DDX28_HUMAN
|
||||||
| θ value | 1.38821 (rank : 21) | NC score | 0.046568 (rank : 31) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NUL7 | Gene names | DDX28, MDDX28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX28 (EC 3.6.1.-) (Mitochondrial DEAD box protein 28). | |||||
|
DDX18_MOUSE
|
||||||
| θ value | 1.81305 (rank : 22) | NC score | 0.052821 (rank : 24) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8K363, Q3MIB0, Q8BVZ2, Q9D2E0 | Gene names | Ddx18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX18 (EC 3.6.1.-) (DEAD box protein 18). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.015749 (rank : 41) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NSD1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.020008 (rank : 36) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
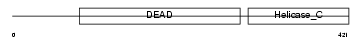
DDX6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.058121 (rank : 19) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P26196 | Gene names | DDX6, HLR2, RCK | |||
|
Domain Architecture |
|
|||||
| Description | Probable ATP-dependent RNA helicase DDX6 (EC 3.6.1.-) (DEAD box protein 6) (ATP-dependent RNA helicase p54) (Oncogene RCK). | |||||
|
DDX6_MOUSE
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.058132 (rank : 18) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P54823 | Gene names | Ddx6, Hlr2, Rck | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX6 (EC 3.6.1.-) (DEAD box protein 6) (ATP-dependent RNA helicase p54) (Oncogene RCK homolog). | |||||
|
CHK2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.000869 (rank : 53) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z265 | Gene names | Chek2, Chk2, Rad53 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Chk2 (EC 2.7.11.1). | |||||
|
WRN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.091737 (rank : 11) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O09053, O09050, Q9JKD4, Q9Z241, Q9Z242 | Gene names | Wrn | |||
|
Domain Architecture |

|
|||||
| Description | Werner syndrome ATP-dependent helicase homolog (EC 3.6.1.-). | |||||
|
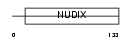
AP4A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 29) | NC score | 0.028350 (rank : 34) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50583 | Gene names | NUDT2, APAH1 | |||
|
Domain Architecture |
|
|||||
| Description | Bis(5'-nucleosyl)-tetraphosphatase [Asymmetrical] (EC 3.6.1.17) (Diadenosine 5',5'''-P1,P4-tetraphosphate asymmetrical hydrolase) (Diadenosine tetraphosphatase) (Ap4A hydrolase) (Ap4Aase) (Nucleoside diphosphate-linked moiety X motif 2) (Nudix motif 2). | |||||
|
TTF2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 30) | NC score | 0.010941 (rank : 44) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
ATX2L_HUMAN
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.014189 (rank : 43) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WWM7, O95135, Q6NVJ8, Q6PJW6, Q8IU61, Q8IU95, Q8WWM3, Q8WWM4, Q8WWM5, Q8WWM6, Q99703 | Gene names | ATXN2L, A2D, A2LG, A2LP, A2RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein (Ataxin-2 domain protein) (Ataxin-2-related protein). | |||||
|
CAD18_HUMAN
|
||||||
| θ value | 5.27518 (rank : 32) | NC score | 0.004012 (rank : 51) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13634 | Gene names | CDH18, CDH14 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-18 precursor (Cadherin-14). | |||||
|
DDX10_MOUSE
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.050558 (rank : 29) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q80Y44 | Gene names | Ddx10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX10 (EC 3.6.1.-) (DEAD box protein 10). | |||||
|
ZC11A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 34) | NC score | 0.017700 (rank : 38) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
ATG9A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 35) | NC score | 0.016910 (rank : 39) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z3C6, Q6P0N7, Q7Z317, Q7Z320, Q8NDK6, Q8WU65, Q9BVL5, Q9H6L1, Q9HAG7 | Gene names | ATG9A, APG9L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Autophagy-related protein 9A (APG9-like 1). | |||||
|
EVX1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.004167 (rank : 50) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23683 | Gene names | Evx1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox even-skipped homolog protein 1 (EVX-1). | |||||
|
HES6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.015293 (rank : 42) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHE6, Q3U2D7 | Gene names | Hes6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6). | |||||
|
MN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.018349 (rank : 37) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q10571 | Gene names | MN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable tumor suppressor protein MN1. | |||||
|
PTPRT_HUMAN
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.006104 (rank : 47) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14522, O43655, O75664, Q5W0X9, Q5W0Y1, Q9BR24, Q9BR28, Q9H0Y8, Q9NTL1, Q9NU72, Q9UBD2, Q9UJL7 | Gene names | PTPRT, KIAA0283 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase T precursor (EC 3.1.3.48) (R-PTP-T) (RPTP-rho). | |||||
|
PTPRT_MOUSE
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.006099 (rank : 48) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99M80, Q99M81, Q99M82, Q9JIZ1, Q9JIZ2, Q9JKC2, Q9JLP0 | Gene names | Ptprt | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase T precursor (EC 3.1.3.48) (R-PTP-T) (RPTP-rho) (mRPTPrho) (RPTPmam4). | |||||
|
BAZ2A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 41) | NC score | 0.007803 (rank : 46) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
CADH3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.003401 (rank : 52) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P22223 | Gene names | CDH3, CDHP | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-3 precursor (Placental-cadherin) (P-cadherin). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.009686 (rank : 45) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
TRPV5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.005109 (rank : 49) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P69744 | Gene names | Trpv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 5 (TrpV5) (osm-9-like TRP channel 3) (OTRPC3) (Epithelial calcium channel 1) (ECaC1) (Calcium transport protein 2) (CaT2). | |||||
|
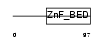
ZBED3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.020285 (rank : 35) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96IU2 | Gene names | ZBED3 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger BED domain-containing protein 3. | |||||
|
BLM_HUMAN
|
||||||
| θ value | θ > 10 (rank : 46) | NC score | 0.067791 (rank : 15) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P54132 | Gene names | BLM, RECQ2, RECQL3 | |||
|
Domain Architecture |

|
|||||
| Description | Bloom syndrome protein (EC 3.6.1.-) (RecQ protein-like 3) (DNA helicase, RecQ-like type 2). | |||||
|
BLM_MOUSE
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.063479 (rank : 16) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O88700, O88198 | Gene names | Blm | |||
|
Domain Architecture |

|
|||||
| Description | Bloom syndrome protein homolog (EC 3.6.1.-) (mBLM). | |||||
|
DDX31_HUMAN
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.051526 (rank : 26) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H8H2, Q5K6N2, Q5K6N3, Q5K6N4, Q5VZJ4, Q5VZJ9, Q96E91, Q96NY2, Q96SX5, Q9H5K6 | Gene names | DDX31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX31 (EC 3.6.1.-) (DEAD box protein 31) (Helicain). | |||||
|
DDX46_HUMAN
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.050793 (rank : 28) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7L014, O94894, Q96EI0, Q9Y658 | Gene names | DDX46, KIAA0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46) (PRP5 homolog). | |||||
|
DDX46_MOUSE
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.050919 (rank : 27) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q569Z5, Q6ZQ42, Q8R0R6 | Gene names | Ddx46, Kiaa0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46). | |||||
|
DDX52_HUMAN
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.055636 (rank : 22) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Y2R4, Q86YG1, Q8N213, Q9NVE0, Q9Y482 | Gene names | DDX52, ROK1 | |||
|
Domain Architecture |
|
|||||
| Description | Probable ATP-dependent RNA helicase DDX52 (EC 3.6.1.-) (DEAD box protein 52) (ATP-dependent RNA helicase ROK1-like). | |||||
|
DDX52_MOUSE
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.053729 (rank : 23) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8K301, Q8BV29 | Gene names | Ddx52, Rok1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX52 (EC 3.6.1.-) (DEAD box protein 52) (ATP-dependent RNA helicase ROK1-like). | |||||
|
RECQ5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.077722 (rank : 14) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
SKIV2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q15477, O15005, Q12902, Q15476 | Gene names | SKIV2L, DDX13, SKI2W, SKIV2, W | |||
|
Domain Architecture |

|
|||||
| Description | Helicase SKI2W (EC 3.6.1.-) (Helicase-like protein) (HLP). | |||||
|
SK2L2_HUMAN
|
||||||
| NC score | 0.937283 (rank : 2) | θ value | 1.79118e-157 (rank : 3) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P42285, Q8TAG2 | Gene names | SKIV2L2, KIAA0052 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Superkiller viralicidic activity 2-like 2 (EC 3.6.1.-) (ATP-dependent helicase SKIV2L2). | |||||
|
SK2L2_MOUSE
|
||||||
| NC score | 0.936298 (rank : 3) | θ value | 3.26429e-159 (rank : 2) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9CZU3 | Gene names | Skiv2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Superkiller viralicidic activity 2-like 2 (EC 3.6.1.-) (ATP-dependent helicase SKIV2L2). | |||||
|
HELC1_HUMAN
|
||||||
| NC score | 0.569619 (rank : 4) | θ value | 2.87452e-22 (rank : 5) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N3C0, O43738, Q9H1I9, Q9NTR0 | Gene names | ASCC3, HELIC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 3 (EC 3.6.1.-) (ASC-1 complex subunit p200) (Trip4 complex subunit p200) (Helicase, ATP binding 1). | |||||
|
U520_HUMAN
|
||||||
| NC score | 0.560851 (rank : 5) | θ value | 1.05554e-24 (rank : 4) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75643, O94884, Q6PX59, Q96IF2, Q9H7S0 | Gene names | ASCC3L1, HELIC2, KIAA0788 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U5 small nuclear ribonucleoprotein 200 kDa helicase (EC 3.6.1.-) (U5 snRNP-specific 200 kDa protein) (U5-200KD) (Activating signal cointegrator 1 complex subunit 3-like 1) (BRR2 homolog). | |||||
|
RECQ1_HUMAN
|
||||||
| NC score | 0.143147 (rank : 6) | θ value | 0.0431538 (rank : 9) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P46063 | Gene names | RECQL, RECQL1 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q1 (EC 3.6.1.-) (DNA-dependent ATPase Q1). | |||||
|
RECQ1_MOUSE
|
||||||
| NC score | 0.125131 (rank : 7) | θ value | 0.163984 (rank : 10) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Z129, Q9Z128 | Gene names | Recql, Recql1 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q1 (EC 3.6.1.-) (DNA-dependent ATPase Q1). | |||||
|
WRN_HUMAN
|
||||||
| NC score | 0.121814 (rank : 8) | θ value | 0.163984 (rank : 11) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14191 | Gene names | WRN, RECQ3, RECQL2 | |||
|
Domain Architecture |

|
|||||
| Description | Werner syndrome ATP-dependent helicase (EC 3.6.1.-). | |||||
|
RECQ4_HUMAN
|
||||||
| NC score | 0.118705 (rank : 9) | θ value | 0.279714 (rank : 13) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O94761, Q3Y424, Q96DW2, Q96F55 | Gene names | RECQL4, RECQ4 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4) (RecQ4) (RTS). | |||||
|
RECQ4_MOUSE
|
||||||
| NC score | 0.114891 (rank : 10) | θ value | 0.365318 (rank : 14) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q75NR7, Q76MT1, Q99PV9 | Gene names | Recql4, Recq4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4). | |||||
|
WRN_MOUSE
|
||||||
| NC score | 0.091737 (rank : 11) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O09053, O09050, Q9JKD4, Q9Z241, Q9Z242 | Gene names | Wrn | |||
|
Domain Architecture |

|
|||||
| Description | Werner syndrome ATP-dependent helicase homolog (EC 3.6.1.-). | |||||
|
DDX54_HUMAN
|
||||||
| NC score | 0.087577 (rank : 12) | θ value | 0.0113563 (rank : 6) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TDD1, Q86YT8, Q9BRZ1 | Gene names | DDX54 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX54 (EC 3.6.1.-) (DEAD box protein 54) (ATP-dependent RNA helicase DP97). | |||||
|
DDX54_MOUSE
|
||||||
| NC score | 0.084729 (rank : 13) | θ value | 0.0252991 (rank : 8) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8K4L0 | Gene names | Ddx54 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX54 (EC 3.6.1.-) (DEAD box protein 54). | |||||
|
RECQ5_HUMAN
|
||||||
| NC score | 0.077722 (rank : 14) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
BLM_HUMAN
|
||||||
| NC score | 0.067791 (rank : 15) | θ value | θ > 10 (rank : 46) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P54132 | Gene names | BLM, RECQ2, RECQL3 | |||
|
Domain Architecture |

|
|||||
| Description | Bloom syndrome protein (EC 3.6.1.-) (RecQ protein-like 3) (DNA helicase, RecQ-like type 2). | |||||
|
BLM_MOUSE
|
||||||
| NC score | 0.063479 (rank : 16) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O88700, O88198 | Gene names | Blm | |||
|
Domain Architecture |

|
|||||
| Description | Bloom syndrome protein homolog (EC 3.6.1.-) (mBLM). | |||||
|
DDX10_HUMAN
|
||||||
| NC score | 0.061552 (rank : 17) | θ value | 0.47712 (rank : 15) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q13206, Q5BJD8 | Gene names | DDX10 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX10 (EC 3.6.1.-) (DEAD box protein 10). | |||||
|
DDX6_MOUSE
|
||||||
| NC score | 0.058132 (rank : 18) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P54823 | Gene names | Ddx6, Hlr2, Rck | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX6 (EC 3.6.1.-) (DEAD box protein 6) (ATP-dependent RNA helicase p54) (Oncogene RCK homolog). | |||||
|
DDX6_HUMAN
|
||||||
| NC score | 0.058121 (rank : 19) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P26196 | Gene names | DDX6, HLR2, RCK | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX6 (EC 3.6.1.-) (DEAD box protein 6) (ATP-dependent RNA helicase p54) (Oncogene RCK). | |||||
|
DDX50_HUMAN
|
||||||
| NC score | 0.056538 (rank : 20) | θ value | 0.47712 (rank : 16) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BQ39, Q8WV76, Q9BWI8 | Gene names | DDX50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX50 (EC 3.6.1.-) (DEAD box protein 50) (Nucleolar protein Gu2) (Gu-beta). | |||||
|
DDX50_MOUSE
|
||||||
| NC score | 0.056481 (rank : 21) | θ value | 0.47712 (rank : 17) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q99MJ9 | Gene names | Ddx50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX50 (EC 3.6.1.-) (DEAD box protein 50) (Nucleolar protein Gu2) (Gu-beta). | |||||
|
DDX52_HUMAN
|
||||||
| NC score | 0.055636 (rank : 22) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Y2R4, Q86YG1, Q8N213, Q9NVE0, Q9Y482 | Gene names | DDX52, ROK1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX52 (EC 3.6.1.-) (DEAD box protein 52) (ATP-dependent RNA helicase ROK1-like). | |||||
|
DDX52_MOUSE
|
||||||
| NC score | 0.053729 (rank : 23) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8K301, Q8BV29 | Gene names | Ddx52, Rok1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX52 (EC 3.6.1.-) (DEAD box protein 52) (ATP-dependent RNA helicase ROK1-like). | |||||
|
DDX18_MOUSE
|
||||||
| NC score | 0.052821 (rank : 24) | θ value | 1.81305 (rank : 22) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8K363, Q3MIB0, Q8BVZ2, Q9D2E0 | Gene names | Ddx18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX18 (EC 3.6.1.-) (DEAD box protein 18). | |||||
|
P3H3_MOUSE
|
||||||
| NC score | 0.052444 (rank : 25) | θ value | 0.0113563 (rank : 7) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CG70, O88836 | Gene names | Leprel2, P3h3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 3 precursor (EC 1.14.11.7) (Leprecan-like protein 2) (Protein B). | |||||
|
DDX31_HUMAN
|
||||||
| NC score | 0.051526 (rank : 26) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H8H2, Q5K6N2, Q5K6N3, Q5K6N4, Q5VZJ4, Q5VZJ9, Q96E91, Q96NY2, Q96SX5, Q9H5K6 | Gene names | DDX31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX31 (EC 3.6.1.-) (DEAD box protein 31) (Helicain). | |||||
|
DDX46_MOUSE
|
||||||
| NC score | 0.050919 (rank : 27) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q569Z5, Q6ZQ42, Q8R0R6 | Gene names | Ddx46, Kiaa0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46). | |||||
|
DDX46_HUMAN
|
||||||
| NC score | 0.050793 (rank : 28) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7L014, O94894, Q96EI0, Q9Y658 | Gene names | DDX46, KIAA0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46) (PRP5 homolog). | |||||
|
DDX10_MOUSE
|
||||||
| NC score | 0.050558 (rank : 29) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q80Y44 | Gene names | Ddx10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX10 (EC 3.6.1.-) (DEAD box protein 10). | |||||
|
DDX18_HUMAN
|
||||||
| NC score | 0.049685 (rank : 30) | θ value | 1.38821 (rank : 20) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NVP1, Q6IAU4, Q92732, Q9BQB7 | Gene names | DDX18 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX18 (EC 3.6.1.-) (DEAD box protein 18) (Myc-regulated DEAD box protein) (MrDb). | |||||
|
DDX28_HUMAN
|
||||||
| NC score | 0.046568 (rank : 31) | θ value | 1.38821 (rank : 21) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NUL7 | Gene names | DDX28, MDDX28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX28 (EC 3.6.1.-) (Mitochondrial DEAD box protein 28). | |||||
|
DHX36_HUMAN
|
||||||
| NC score | 0.033472 (rank : 32) | θ value | 0.62314 (rank : 18) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H2U1, Q70JU3, Q8IYE5, Q9P240 | Gene names | DHX36, DDX36, KIAA1488, MLEL1, RHAU | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DHX36 (EC 3.6.1.-) (DEAH box protein 36) (MLE-like protein 1) (RNA helicase associated with AU-rich element ARE). | |||||
|
EPN3_MOUSE
|
||||||
| NC score | 0.032846 (rank : 33) | θ value | 0.21417 (rank : 12) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91W69, Q9CV55 | Gene names | Epn3 | |||
|
Domain Architecture |

|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
AP4A_HUMAN
|
||||||
| NC score | 0.028350 (rank : 34) | θ value | 4.03905 (rank : 29) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50583 | Gene names | NUDT2, APAH1 | |||
|
Domain Architecture |

|
|||||
| Description | Bis(5'-nucleosyl)-tetraphosphatase [Asymmetrical] (EC 3.6.1.17) (Diadenosine 5',5'''-P1,P4-tetraphosphate asymmetrical hydrolase) (Diadenosine tetraphosphatase) (Ap4A hydrolase) (Ap4Aase) (Nucleoside diphosphate-linked moiety X motif 2) (Nudix motif 2). | |||||
|
ZBED3_HUMAN
|
||||||
| NC score | 0.020285 (rank : 35) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96IU2 | Gene names | ZBED3 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger BED domain-containing protein 3. | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.020008 (rank : 36) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
MN1_HUMAN
|
||||||
| NC score | 0.018349 (rank : 37) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q10571 | Gene names | MN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable tumor suppressor protein MN1. | |||||
|
ZC11A_MOUSE
|
||||||
| NC score | 0.017700 (rank : 38) | θ value | 5.27518 (rank : 34) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
ATG9A_HUMAN
|
||||||
| NC score | 0.016910 (rank : 39) | θ value | 6.88961 (rank : 35) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z3C6, Q6P0N7, Q7Z317, Q7Z320, Q8NDK6, Q8WU65, Q9BVL5, Q9H6L1, Q9HAG7 | Gene names | ATG9A, APG9L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Autophagy-related protein 9A (APG9-like 1). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.015782 (rank : 40) | θ value | 0.62314 (rank : 19) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.015749 (rank : 41) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
HES6_MOUSE
|
||||||
| NC score | 0.015293 (rank : 42) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHE6, Q3U2D7 | Gene names | Hes6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6). | |||||
|
ATX2L_HUMAN
|
||||||
| NC score | 0.014189 (rank : 43) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WWM7, O95135, Q6NVJ8, Q6PJW6, Q8IU61, Q8IU95, Q8WWM3, Q8WWM4, Q8WWM5, Q8WWM6, Q99703 | Gene names | ATXN2L, A2D, A2LG, A2LP, A2RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein (Ataxin-2 domain protein) (Ataxin-2-related protein). | |||||
|
TTF2_MOUSE
|
||||||
| NC score | 0.010941 (rank : 44) | θ value | 4.03905 (rank : 30) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.009686 (rank : 45) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
BAZ2A_HUMAN
|
||||||
| NC score | 0.007803 (rank : 46) | θ value | 8.99809 (rank : 41) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
PTPRT_HUMAN
|
||||||
| NC score | 0.006104 (rank : 47) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14522, O43655, O75664, Q5W0X9, Q5W0Y1, Q9BR24, Q9BR28, Q9H0Y8, Q9NTL1, Q9NU72, Q9UBD2, Q9UJL7 | Gene names | PTPRT, KIAA0283 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase T precursor (EC 3.1.3.48) (R-PTP-T) (RPTP-rho). | |||||
|
PTPRT_MOUSE
|
||||||
| NC score | 0.006099 (rank : 48) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99M80, Q99M81, Q99M82, Q9JIZ1, Q9JIZ2, Q9JKC2, Q9JLP0 | Gene names | Ptprt | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase T precursor (EC 3.1.3.48) (R-PTP-T) (RPTP-rho) (mRPTPrho) (RPTPmam4). | |||||
|
TRPV5_MOUSE
|
||||||
| NC score | 0.005109 (rank : 49) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P69744 | Gene names | Trpv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 5 (TrpV5) (osm-9-like TRP channel 3) (OTRPC3) (Epithelial calcium channel 1) (ECaC1) (Calcium transport protein 2) (CaT2). | |||||
|
EVX1_MOUSE
|
||||||
| NC score | 0.004167 (rank : 50) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23683 | Gene names | Evx1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox even-skipped homolog protein 1 (EVX-1). | |||||
|
CAD18_HUMAN
|
||||||
| NC score | 0.004012 (rank : 51) | θ value | 5.27518 (rank : 32) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13634 | Gene names | CDH18, CDH14 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-18 precursor (Cadherin-14). | |||||
|
CADH3_HUMAN
|
||||||
| NC score | 0.003401 (rank : 52) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P22223 | Gene names | CDH3, CDHP | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-3 precursor (Placental-cadherin) (P-cadherin). | |||||
|
CHK2_MOUSE
|
||||||
| NC score | 0.000869 (rank : 53) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z265 | Gene names | Chek2, Chk2, Rad53 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Chk2 (EC 2.7.11.1). | |||||