Please be patient as the page loads
|
SEN34_MOUSE
|
||||||
| SwissProt Accessions | Q8BMZ5, Q58EU2, Q99LV6, Q9CYW1 | Gene names | Tsen34, Leng5, Sen34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-splicing endonuclease subunit Sen34 (EC 3.1.27.9) (tRNA-intron endonuclease Sen34) (Leukocyte receptor cluster member 5 homolog). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SEN34_MOUSE
|
||||||
| θ value | 2.40964e-170 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 158 | |
| SwissProt Accessions | Q8BMZ5, Q58EU2, Q99LV6, Q9CYW1 | Gene names | Tsen34, Leng5, Sen34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-splicing endonuclease subunit Sen34 (EC 3.1.27.9) (tRNA-intron endonuclease Sen34) (Leukocyte receptor cluster member 5 homolog). | |||||
|
SEN34_HUMAN
|
||||||
| θ value | 4.13891e-146 (rank : 2) | NC score | 0.923826 (rank : 2) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BSV6, Q9BVT1, Q9H6H5 | Gene names | TSEN34, LENG5, SEN34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-splicing endonuclease subunit Sen34 (EC 3.1.27.9) (tRNA-intron endonuclease Sen34) (HsSen34) (Leukocyte receptor cluster member 5). | |||||
|
RING2_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 3) | NC score | 0.133890 (rank : 4) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99496, Q5TEN1, Q5TEN2 | Gene names | RNF2, BAP1, DING, HIPI3, RING1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein RING2 (EC 6.3.2.-) (RING finger protein 2) (RING finger protein 1B) (RING1b) (RING finger protein BAP-1) (DinG protein) (Huntingtin-interacting protein 2-interacting protein 3) (HIP2-interacting protein 3). | |||||
|
RING2_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 4) | NC score | 0.133902 (rank : 3) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CQJ4, O35699, O35729, Q4FJV5, Q8C1X8 | Gene names | Rnf2, DinG, Ring1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein RING2 (EC 6.3.2.-) (RING finger protein 2) (RING finger protein 1B) (RING1b). | |||||
|
NFM_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 5) | NC score | 0.040821 (rank : 64) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
TR150_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 6) | NC score | 0.100099 (rank : 9) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y2W1, Q5VTK6 | Gene names | THRAP3, TRAP150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
BMP6_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 7) | NC score | 0.028898 (rank : 92) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P20722 | Gene names | Bmp6, Bmp-6, Vgr1 | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein 6 precursor (BMP-6) (VG-1-related protein) (VGR-1). | |||||
|
LYAR_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 8) | NC score | 0.091128 (rank : 11) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q08288, Q9D9X2 | Gene names | Lyar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth-regulating nucleolar protein. | |||||
|
SRCA_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 9) | NC score | 0.070703 (rank : 18) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
MARCS_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 10) | NC score | 0.112301 (rank : 7) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
CLSPN_HUMAN
|
||||||
| θ value | 0.125558 (rank : 11) | NC score | 0.062741 (rank : 28) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9HAW4, Q5VYG0, Q6P6H5, Q8IWI1 | Gene names | CLSPN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin (hClaspin) (Hu-Claspin). | |||||
|
NFM_HUMAN
|
||||||
| θ value | 0.125558 (rank : 12) | NC score | 0.038596 (rank : 69) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
SCG1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 13) | NC score | 0.074737 (rank : 13) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P16014 | Gene names | Chgb, Scg-1, Scg1 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-1 precursor (Secretogranin I) (SgI) (Chromogranin B) (CgB). | |||||
|
SEN2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 14) | NC score | 0.126909 (rank : 5) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NCE0, Q8WTW7, Q9BPU7 | Gene names | TSEN2, SEN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-splicing endonuclease subunit Sen2 (EC 3.1.27.9) (tRNA-intron endonuclease Sen2) (HsSen2 protein). | |||||
|
TM16H_MOUSE
|
||||||
| θ value | 0.125558 (rank : 15) | NC score | 0.058202 (rank : 38) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PB70, Q69ZE4 | Gene names | Tmem16h, Kiaa1623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 16H. | |||||
|
TRI44_HUMAN
|
||||||
| θ value | 0.125558 (rank : 16) | NC score | 0.064446 (rank : 26) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96DX7, Q96QY2, Q9UGK0 | Gene names | TRIM44, DIPB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB). | |||||
|
SNPC4_HUMAN
|
||||||
| θ value | 0.163984 (rank : 17) | NC score | 0.061605 (rank : 33) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5SXM2, Q9Y6P7 | Gene names | SNAPC4, SNAP190 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit) (Proximal sequence element-binding transcription factor subunit alpha) (PSE-binding factor subunit alpha) (PTF subunit alpha). | |||||
|
TARA_HUMAN
|
||||||
| θ value | 0.163984 (rank : 18) | NC score | 0.072606 (rank : 15) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TBP_HUMAN
|
||||||
| θ value | 0.163984 (rank : 19) | NC score | 0.064495 (rank : 25) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20226, Q16845, Q9UC02 | Gene names | TBP, TF2D, TFIID | |||
|
Domain Architecture |
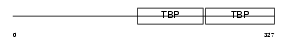
|
|||||
| Description | TATA-box-binding protein (TATA-box factor) (TATA-binding factor) (TATA sequence-binding protein) (Transcription initiation factor TFIID TBP subunit). | |||||
|
ARI1A_HUMAN
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.053736 (rank : 50) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 0.21417 (rank : 21) | NC score | 0.069938 (rank : 20) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
MARCS_HUMAN
|
||||||
| θ value | 0.21417 (rank : 22) | NC score | 0.101044 (rank : 8) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
CAMKV_HUMAN
|
||||||
| θ value | 0.365318 (rank : 23) | NC score | 0.016965 (rank : 135) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8NCB2, Q6FIB8, Q8NBS8, Q8NC85, Q8NDU4, Q8WTT8, Q9BQC9, Q9H0Q5 | Gene names | CAMKV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CaM kinase-like vesicle-associated protein. | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 24) | NC score | 0.062202 (rank : 32) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
SEN2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 25) | NC score | 0.116362 (rank : 6) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P7W5, Q3UYG6 | Gene names | Tsen2, Sen2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-splicing endonuclease subunit Sen2 (EC 3.1.27.9) (tRNA-intron endonuclease Sen2). | |||||
|
TRI44_MOUSE
|
||||||
| θ value | 0.365318 (rank : 26) | NC score | 0.062612 (rank : 29) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QXA7, Q80SV9, Q9JHR8 | Gene names | Trim44, Dipb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB) (Protein Mc7). | |||||
|
CA106_HUMAN
|
||||||
| θ value | 0.47712 (rank : 27) | NC score | 0.071484 (rank : 16) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q3KP66, Q9NV65, Q9NVI0 | Gene names | C1orf106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf106. | |||||
|
JHD2B_HUMAN
|
||||||
| θ value | 0.47712 (rank : 28) | NC score | 0.033153 (rank : 81) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7LBC6, Q9BVH6, Q9BW93, Q9BZ52, Q9NYF4, Q9UPS0 | Gene names | JMJD1B, C5orf7, JHDM2B, KIAA1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B) (Nuclear protein 5qNCA). | |||||
|
LNP_MOUSE
|
||||||
| θ value | 0.47712 (rank : 29) | NC score | 0.068684 (rank : 21) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TQ95, Q69ZC5, Q6PAQ1, Q6PEN8 | Gene names | Lnp, Kiaa1715, Uln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein lunapark (Protein ulnaless). | |||||
|
MYLK2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 30) | NC score | 0.012140 (rank : 147) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H1R3, Q96I84 | Gene names | MYLK2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin light chain kinase 2, skeletal/cardiac muscle (EC 2.7.11.18) (MLCK2). | |||||
|
STK23_MOUSE
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.025740 (rank : 100) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 572 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Z0G2 | Gene names | Stk23, Mssk1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase 23 (EC 2.7.11.1) (Muscle-specific serine kinase 1) (MSSK-1). | |||||
|
IPP2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 32) | NC score | 0.056663 (rank : 44) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DCL8, Q3UD25, Q8C1A6 | Gene names | Ppp1r2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase inhibitor 2 (IPP-2). | |||||
|
TTC28_HUMAN
|
||||||
| θ value | 0.62314 (rank : 33) | NC score | 0.039194 (rank : 67) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96AY4, O95928, O95929, Q9NTE4, Q9UG31, Q9UGG5, Q9UPV8, Q9Y3S5 | Gene names | TTC28, KIAA1043 | |||
|
Domain Architecture |
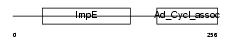
|
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.061233 (rank : 36) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CA106_MOUSE
|
||||||
| θ value | 0.813845 (rank : 35) | NC score | 0.068142 (rank : 22) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7TN12, Q7TN27, Q8BGK3 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf106 homolog. | |||||
|
LHX3_MOUSE
|
||||||
| θ value | 0.813845 (rank : 36) | NC score | 0.012488 (rank : 145) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50481, Q61800, Q61801 | Gene names | Lhx3, Lim-3, Lim3, Plim | |||
|
Domain Architecture |
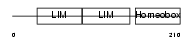
|
|||||
| Description | LIM/homeobox protein Lhx3 (Homeobox protein LIM-3) (Homeobox protein P-LIM) (LIM homeobox protein 3). | |||||
|
NFL_HUMAN
|
||||||
| θ value | 0.813845 (rank : 37) | NC score | 0.029518 (rank : 90) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 666 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P07196, Q16154, Q8IU72 | Gene names | NEFL, NF68, NFL | |||
|
Domain Architecture |
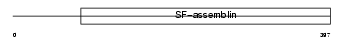
|
|||||
| Description | Neurofilament triplet L protein (68 kDa neurofilament protein) (Neurofilament light polypeptide) (NF-L). | |||||
|
TR150_MOUSE
|
||||||
| θ value | 0.813845 (rank : 38) | NC score | 0.093600 (rank : 10) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
CAC1C_HUMAN
|
||||||
| θ value | 1.06291 (rank : 39) | NC score | 0.021782 (rank : 122) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
CD2AP_MOUSE
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.023366 (rank : 115) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9JLQ0, O88903, Q8K4Z1, Q8VCI9 | Gene names | Cd2ap, Mets1 | |||
|
Domain Architecture |
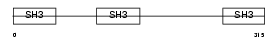
|
|||||
| Description | CD2-associated protein (Mesenchyme-to-epithelium transition protein with SH3 domains 1) (METS-1). | |||||
|
HPS4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.045697 (rank : 59) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NQG7, Q5H8V6, Q96LX6, Q9BY93, Q9UH37, Q9UH38 | Gene names | HPS4, KIAA1667 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 4 protein (Light-ear protein homolog). | |||||
|
PDXL2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.052152 (rank : 52) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8CAE9, Q8CFW3 | Gene names | Podxl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
RERE_HUMAN
|
||||||
| θ value | 1.06291 (rank : 43) | NC score | 0.035607 (rank : 74) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
RP1L1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 44) | NC score | 0.042459 (rank : 62) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CGM2, Q8K0I4 | Gene names | Rp1l1, Rp1hl1 | |||
|
Domain Architecture |
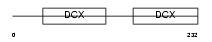
|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein (Retinitis pigmentosa 1-like protein 1). | |||||
|
SRCA_MOUSE
|
||||||
| θ value | 1.06291 (rank : 45) | NC score | 0.061345 (rank : 34) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7TQ48, Q8BS43 | Gene names | Srl, Sar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
TIF1G_HUMAN
|
||||||
| θ value | 1.06291 (rank : 46) | NC score | 0.022896 (rank : 119) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
UBF1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 47) | NC score | 0.038139 (rank : 70) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P25976 | Gene names | Ubtf, Tcfubf, Ubf-1, Ubf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1). | |||||
|
CMTA2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 48) | NC score | 0.032869 (rank : 82) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80Y50, Q3TFK0, Q5SX68, Q80TP1, Q8R0D9, Q8R2N5 | Gene names | Camta2, Kiaa0909 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-binding transcription activator 2. | |||||
|
M4K4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 49) | NC score | 0.009430 (rank : 152) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1560 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P97820 | Gene names | Map4k4, Nik | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase kinase 4 (EC 2.7.11.1) (MAPK/ERK kinase kinase kinase 4) (MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase HGK) (Nck-interacting kinase). | |||||
|
MYT1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 50) | NC score | 0.025438 (rank : 102) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
NPHP1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 51) | NC score | 0.027974 (rank : 94) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QY53, Q9D7G8, Q9QZP0, Q9WUZ2 | Gene names | Nphp1, Nph1 | |||
|
Domain Architecture |

|
|||||
| Description | Nephrocystin-1. | |||||
|
PO121_MOUSE
|
||||||
| θ value | 1.38821 (rank : 52) | NC score | 0.039920 (rank : 66) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
RALY_MOUSE
|
||||||
| θ value | 1.38821 (rank : 53) | NC score | 0.031602 (rank : 86) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q64012, Q99K76, Q9CXH8, Q9QZX6 | Gene names | Raly, Merc | |||
|
Domain Architecture |
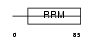
|
|||||
| Description | RNA-binding protein Raly (hnRNP associated with lethal yellow protein) (Maternally expressed hnRNP C-related protein). | |||||
|
SHAN2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 54) | NC score | 0.030976 (rank : 87) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 446 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80Z38, Q3UTK4, Q5DU07 | Gene names | Shank2, Kiaa1022 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2) (Cortactin- binding protein 1) (CortBP1). | |||||
|
SPEG_HUMAN
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.014194 (rank : 141) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
ZN414_HUMAN
|
||||||
| θ value | 1.38821 (rank : 56) | NC score | 0.026716 (rank : 97) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96IQ9 | Gene names | ZNF414 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 414. | |||||
|
IF4G3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.025794 (rank : 99) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80XI3, Q80Y69, Q8BJ68, Q8BQF3, Q8C6Q3, Q8CIH0 | Gene names | Eif4g3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.055162 (rank : 48) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MRP_MOUSE
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.074458 (rank : 14) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P28667, Q3TEZ4, Q91W07 | Gene names | Marcksl1, Mlp, Mrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MARCKS-related protein (MARCKS-like protein 1) (Macrophage myristoylated alanine-rich C kinase substrate) (Mac-MARCKS) (MacMARCKS) (Brain protein F52). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 60) | NC score | 0.048777 (rank : 53) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
PTMS_HUMAN
|
||||||
| θ value | 1.81305 (rank : 61) | NC score | 0.070654 (rank : 19) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
SAM14_HUMAN
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.057952 (rank : 39) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IZD0, Q8N2X0 | Gene names | SAMD14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha motif domain-containing protein 14. | |||||
|
SET1A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.055566 (rank : 47) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
SOX12_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.015207 (rank : 138) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15370, Q9NUD4 | Gene names | SOX12, SOX22 | |||
|
Domain Architecture |
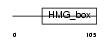
|
|||||
| Description | SOX-12 protein (SOX-22 protein). | |||||
|
TRHY_HUMAN
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.023292 (rank : 116) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
ACRBP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.034723 (rank : 76) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3V140, Q62253, Q62254, Q8C621, Q91VQ1 | Gene names | Acrbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acrosin-binding protein precursor (Proacrosin-binding protein sp32). | |||||
|
ANDR_HUMAN
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.010864 (rank : 149) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P10275 | Gene names | AR, DHTR, NR3C4 | |||
|
Domain Architecture |

|
|||||
| Description | Androgen receptor (Dihydrotestosterone receptor). | |||||
|
BCOR_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.021223 (rank : 126) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CGN4, Q6PDK5, Q6ZPM3, Q8BKF5, Q8CGN1, Q8CGN2, Q8CGN3 | Gene names | Bcor, Kiaa1575 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BCoR protein (BCL-6 corepressor). | |||||
|
BSPRY_MOUSE
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.024809 (rank : 105) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80YW5, Q3TU74, Q8BZF0, Q99KV7, Q9ER70 | Gene names | Bspry | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B box and SPRY domain-containing protein. | |||||
|
DDX51_HUMAN
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.022847 (rank : 120) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
HNRLL_MOUSE
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.021515 (rank : 124) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q921F4, Q8BIP6, Q99J40 | Gene names | Hnrpll | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L-like. | |||||
|
MARH4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.033584 (rank : 78) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80TE3 | Gene names | March4, Kiaa1399 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated RING finger protein 4 (Membrane-associated RING-CH protein IV) (MARCH-IV). | |||||
|
MCRS1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.034884 (rank : 75) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99L90, O35255, Q32P11 | Gene names | Mcrs1, Msp58 | |||
|
Domain Architecture |

|
|||||
| Description | Microspherule protein 1 (58 kDa microspherule protein). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 2.36792 (rank : 74) | NC score | 0.055936 (rank : 46) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NASP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 75) | NC score | 0.079961 (rank : 12) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
PDC6I_HUMAN
|
||||||
| θ value | 2.36792 (rank : 76) | NC score | 0.029062 (rank : 91) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WUM4, Q9BX86, Q9NUN0, Q9P2H2, Q9UKL5 | Gene names | PDCD6IP, AIP1, ALIX, KIAA1375 | |||
|
Domain Architecture |
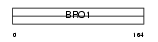
|
|||||
| Description | Programmed cell death 6-interacting protein (PDCD6-interacting protein) (ALG-2-interacting protein 1) (Hp95). | |||||
|
PLK2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 77) | NC score | 0.007528 (rank : 158) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P53351 | Gene names | Plk2, Snk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase PLK2 (EC 2.7.11.21) (Polo-like kinase 1) (PLK-2) (Serine/threonine-protein kinase SNK) (Serum-inducible kinase). | |||||
|
SAM14_MOUSE
|
||||||
| θ value | 2.36792 (rank : 78) | NC score | 0.056379 (rank : 45) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K070, Q5SWB7, Q8BHE2, Q8C8N5 | Gene names | Samd14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha motif domain-containing protein 14. | |||||
|
SMAL1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 79) | NC score | 0.019590 (rank : 128) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BJL0, Q8BVK7, Q8BXW4, Q8K309, Q9EQK8, Q9QYC4 | Gene names | Smarcal1, Harp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A-like protein 1 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 1) (HepA-related protein) (mharp). | |||||
|
TAF10_HUMAN
|
||||||
| θ value | 2.36792 (rank : 80) | NC score | 0.044785 (rank : 61) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12962, O00703, Q13175 | Gene names | TAF10, TAF2H, TAFII30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 10 (Transcription initiation factor TFIID 30 kDa subunit) (TAF(II)30) (TAFII-30) (TAFII30) (STAF28). | |||||
|
TEX2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 81) | NC score | 0.048243 (rank : 55) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IWB9, Q6AHZ5, Q8N3L0, Q9C0C5 | Gene names | TEX2, KIAA1738 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 2 protein. | |||||
|
TEX2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 82) | NC score | 0.048089 (rank : 56) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZPJ0 | Gene names | Tex2, Kiaa1738 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 2 protein. | |||||
|
ZCH10_HUMAN
|
||||||
| θ value | 2.36792 (rank : 83) | NC score | 0.054636 (rank : 49) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8TBK6, Q9NXR4 | Gene names | ZCCHC10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 10. | |||||
|
FAM9A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 84) | NC score | 0.039950 (rank : 65) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IZU1 | Gene names | FAM9A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM9A. | |||||
|
KI67_HUMAN
|
||||||
| θ value | 3.0926 (rank : 85) | NC score | 0.058525 (rank : 37) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
MK07_HUMAN
|
||||||
| θ value | 3.0926 (rank : 86) | NC score | 0.007913 (rank : 157) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1087 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13164, Q16634 | Gene names | MAPK7, ERK4, ERK5, PRKM7 | |||
|
Domain Architecture |
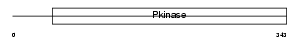
|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (ERK4) (BMK1 kinase). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 87) | NC score | 0.028344 (rank : 93) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
NEUM_HUMAN
|
||||||
| θ value | 3.0926 (rank : 88) | NC score | 0.048243 (rank : 54) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P17677 | Gene names | GAP43 | |||
|
Domain Architecture |
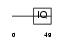
|
|||||
| Description | Neuromodulin (Axonal membrane protein GAP-43) (Growth-associated protein 43) (PP46) (Neural phosphoprotein B-50). | |||||
|
RBM28_MOUSE
|
||||||
| θ value | 3.0926 (rank : 89) | NC score | 0.023716 (rank : 113) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8CGC6, Q8BG25, Q8VEJ8, Q9CS22, Q9CSE6 | Gene names | Rbm28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 28 (RNA-binding motif protein 28). | |||||
|
TMC2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 90) | NC score | 0.021749 (rank : 123) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8TDI7, Q5JXT0, Q5JXT1, Q6UWW4, Q6ZS41, Q8N9F3, Q9BYN2, Q9BYN3, Q9BYN4, Q9BYN5 | Gene names | TMC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane cochlear-expressed protein 2. | |||||
|
UBP42_HUMAN
|
||||||
| θ value | 3.0926 (rank : 91) | NC score | 0.022531 (rank : 121) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H9J4, Q3C166, Q6P9B4 | Gene names | USP42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 42 (EC 3.1.2.15) (Ubiquitin thioesterase 42) (Ubiquitin-specific-processing protease 42) (Deubiquitinating enzyme 42). | |||||
|
ZN696_HUMAN
|
||||||
| θ value | 3.0926 (rank : 92) | NC score | -0.001814 (rank : 166) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 743 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H7X3 | Gene names | ZNF696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 696. | |||||
|
ASTL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.024572 (rank : 108) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6HA08 | Gene names | ASTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Astacin-like metalloendopeptidase precursor (EC 3.4.-.-) (Oocyte astacin) (Ovastacin). | |||||
|
BAZ2A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 94) | NC score | 0.024911 (rank : 104) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
CAF1B_MOUSE
|
||||||
| θ value | 4.03905 (rank : 95) | NC score | 0.012288 (rank : 146) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D0N7 | Gene names | Chaf1b | |||
|
Domain Architecture |
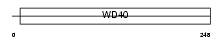
|
|||||
| Description | Chromatin assembly factor 1 subunit B (CAF-1 subunit B) (Chromatin assembly factor I p60 subunit) (CAF-I 60 kDa subunit) (CAF-Ip60). | |||||
|
CENG1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 96) | NC score | 0.010944 (rank : 148) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3UHD9, Q5DU45 | Gene names | Centg1, Agap2, Kiaa0167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-gamma 1 (ARF-GAP with GTP-binding protein-like, ankyrin repeat and pleckstrin homology domains 2) (AGAP-2) (Phosphatidylinositol-3-kinase enhancer) (PIKE). | |||||
|
DACH1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 97) | NC score | 0.023882 (rank : 111) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QYB2, O88716, Q8BPQ0, Q8C8D7, Q9QY47, Q9R218, Q9Z0Y5 | Gene names | Dach1, Dach | |||
|
Domain Architecture |

|
|||||
| Description | Dachshund homolog 1 (Dach1). | |||||
|
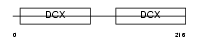
DCDC2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 98) | NC score | 0.035743 (rank : 73) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UHG0, Q5VTR8, Q5VTR9, Q86W35, Q9UFD1, Q9ULR6 | Gene names | DCDC2, KIAA1154 | |||
|
Domain Architecture |
|
|||||
| Description | Doublecortin domain-containing protein 2 (RU2S protein). | |||||
|
ELL3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 99) | NC score | 0.027528 (rank : 96) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80VR2 | Gene names | Ell3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL3. | |||||
|
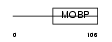
MYRIP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 100) | NC score | 0.032254 (rank : 84) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8NFW9, Q8IUF5, Q9Y3V4 | Gene names | MYRIP, SLAC2C | |||
|
Domain Architecture |
|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.036346 (rank : 72) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
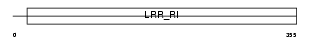
RGP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.023811 (rank : 112) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46061, Q60801 | Gene names | Rangap1, Fug1 | |||
|
Domain Architecture |
|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
RIN3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 103) | NC score | 0.024017 (rank : 109) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TB24, Q8NF30, Q8TEE8, Q8WYP4, Q9H6A5, Q9HAG1 | Gene names | RIN3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
SNRK_HUMAN
|
||||||
| θ value | 4.03905 (rank : 104) | NC score | 0.006781 (rank : 164) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 855 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NRH2, Q14706, Q68D15, Q6IQ46, Q9NXI7 | Gene names | SNRK, KIAA0096, SNFRK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SNF-related serine/threonine-protein kinase (EC 2.7.11.1) (SNF1- related kinase). | |||||
|
TNC6B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 105) | NC score | 0.034010 (rank : 77) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
ZN447_HUMAN
|
||||||
| θ value | 4.03905 (rank : 106) | NC score | 0.008574 (rank : 155) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TBC5, Q9BRK7, Q9H9A0 | Gene names | ZNF447 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 447. | |||||
|
BASP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.046580 (rank : 57) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BL1S3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.031931 (rank : 85) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6QNY0 | Gene names | BLOC1S3, BLOS3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Biogenesis of lysosome-related organelles complex-1 subunit 3 (BLOC subunit 3). | |||||
|
CENPB_MOUSE
|
||||||
| θ value | 5.27518 (rank : 109) | NC score | 0.025454 (rank : 101) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P27790 | Gene names | Cenpb, Cenp-b | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
CS007_HUMAN
|
||||||
| θ value | 5.27518 (rank : 110) | NC score | 0.024594 (rank : 107) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
CTDP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 111) | NC score | 0.030660 (rank : 89) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
DAXX_MOUSE
|
||||||
| θ value | 5.27518 (rank : 112) | NC score | 0.023994 (rank : 110) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O35613, Q9QWT8, Q9QWV3 | Gene names | Daxx | |||
|
Domain Architecture |

|
|||||
| Description | Death domain-associated protein 6 (Daxx). | |||||
|
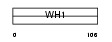
ENAH_HUMAN
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | 0.018372 (rank : 132) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8N8S7, Q502W5, Q5T5M7, Q5VTQ9, Q5VTR0, Q9NVF3, Q9UFB8 | Gene names | ENAH, MENA | |||
|
Domain Architecture |
|
|||||
| Description | Protein enabled homolog. | |||||
|
HNF3A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 114) | NC score | 0.007010 (rank : 160) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P55317, Q9H2A0 | Gene names | FOXA1, HNF3A, TCF3A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-alpha (HNF-3A) (Forkhead box protein A1). | |||||
|
IQEC3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 115) | NC score | 0.015374 (rank : 137) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
LNP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 116) | NC score | 0.052721 (rank : 51) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9C0E8, Q2M2V8, Q2YD99, Q658W8, Q8N5V9, Q96MS5 | Gene names | LNP, KIAA1715 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein lunapark. | |||||
|
MAGB6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 117) | NC score | 0.019009 (rank : 129) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8N7X4, Q6GS19, Q9H219 | Gene names | MAGEB6 | |||
|
Domain Architecture |
|
|||||
| Description | Melanoma-associated antigen B6 (MAGE-B6 antigen). | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 118) | NC score | 0.033162 (rank : 80) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
SHOC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 119) | NC score | 0.007393 (rank : 159) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UQ13, O76063 | Gene names | SHOC2, KIAA0862 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat protein SHOC-2 (Ras-binding protein Sur-8). | |||||
|
TBCD2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 120) | NC score | 0.027549 (rank : 95) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BYX2 | Gene names | TBC1D2, PARIS1 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 2 (Prostate antigen recognized and identified by SEREX) (PARIS-1). | |||||
|
TCF20_MOUSE
|
||||||
| θ value | 5.27518 (rank : 121) | NC score | 0.038677 (rank : 68) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
B3A3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 122) | NC score | 0.008789 (rank : 154) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P48751 | Gene names | SLC4A3, AE3 | |||
|
Domain Architecture |

|
|||||
| Description | Anion exchange protein 3 (Neuronal band 3-like protein) (Solute carrier family 4 member 3) (Cardiac/brain band 3-like protein) (CAE3/BAE3). | |||||
|
CAC1C_MOUSE
|
||||||
| θ value | 6.88961 (rank : 123) | NC score | 0.016198 (rank : 136) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q01815, Q04476, Q61242, Q99242 | Gene names | Cacna1c, Cach2, Cacn2, Cacnl1a1, Cchl1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle) (Mouse brain class C) (MBC) (MELC-CC). | |||||
|
FARP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 124) | NC score | 0.006880 (rank : 162) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O94887, Q53QM5, Q8WU27, Q9UFE7 | Gene names | FARP2, KIAA0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
HNRPU_HUMAN
|
||||||
| θ value | 6.88961 (rank : 125) | NC score | 0.019750 (rank : 127) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q00839, O75507, Q96HY9 | Gene names | HNRPU, SAFA, U21.1 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U (hnRNP U) (Scaffold attachment factor A) (SAF-A) (p120) (pp120). | |||||
|
HTSF1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 126) | NC score | 0.033231 (rank : 79) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
IF2P_HUMAN
|
||||||
| θ value | 6.88961 (rank : 127) | NC score | 0.025209 (rank : 103) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
MRP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 128) | NC score | 0.062478 (rank : 30) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P49006, Q5TEE6, Q6NXS5 | Gene names | MARCKSL1, MLP, MRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MARCKS-related protein (MARCKS-like protein 1) (Macrophage myristoylated alanine-rich C kinase substrate) (Mac-MARCKS) (MacMARCKS). | |||||
|
SHOC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 129) | NC score | 0.006892 (rank : 161) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88520, Q3UJH6, Q8BVL0 | Gene names | Shoc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat protein SHOC-2 (Ras-binding protein Sur-8). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 130) | NC score | 0.045415 (rank : 60) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TAF5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 131) | NC score | 0.003860 (rank : 165) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15542, Q86UZ7, Q9Y4K5 | Gene names | TAF5, TAF2D | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 5 (Transcription initiation factor TFIID 100 kDa subunit) (TAF(II)100) (TAFII-100) (TAFII100). | |||||
|
UN119_MOUSE
|
||||||
| θ value | 6.88961 (rank : 132) | NC score | 0.023022 (rank : 117) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z2R6 | Gene names | Unc119, Unc119h | |||
|
Domain Architecture |
|
|||||
| Description | Unc-119 protein homolog (Retinal protein 4) (MRG4). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.064687 (rank : 24) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
ARS2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | 0.026134 (rank : 98) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99MR6, Q99MR4, Q99MR5, Q99MR7 | Gene names | Ars2, Asr2 | |||
|
Domain Architecture |

|
|||||
| Description | Arsenite-resistance protein 2. | |||||
|
BAZ1A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 135) | NC score | 0.018147 (rank : 133) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
CD2L2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.006865 (rank : 163) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1439 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UQ88, O95227, O95228, O96012, Q12821, Q12853, Q12854, Q5QPR0, Q5QPR1, Q5QPR2, Q9UBC4, Q9UBI3, Q9UEI1, Q9UEI2, Q9UP53, Q9UP54, Q9UP55, Q9UP56, Q9UQ86, Q9UQ87, Q9UQ89 | Gene names | CDC2L2, PITSLREB | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L2 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 2) (CDK11). | |||||
|
CDSN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.024787 (rank : 106) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TPC1 | Gene names | Cdsn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor. | |||||
|
CEP35_HUMAN
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.021390 (rank : 125) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
CTR9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.017991 (rank : 134) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62018, Q3UFF5, Q3UY40, Q66JX4, Q7TPS6, Q8BND9, Q8BRD1, Q8C9W7, Q8C9Y3, Q8CHI1 | Gene names | Ctr9, Kiaa0155, Sh2bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein CTR9 homolog (SH2 domain-binding protein 1) (Tetratricopeptide repeat-containing, SH2-binding phosphoprotein of 150 kDa) (TPR-containing, SH2-binding phosphoprotein of 150 kDa) (p150TSP). | |||||
|
DHX37_HUMAN
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.008924 (rank : 153) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IY37, Q9BUI7, Q9P211 | Gene names | DHX37, DDX37, KIAA1517 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DHX37 (EC 3.6.1.-) (DEAH box protein 37). | |||||
|
DMRT2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.018477 (rank : 131) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y5R5 | Gene names | DMRT2, DSXL2 | |||
|
Domain Architecture |

|
|||||
| Description | Doublesex- and mab-3-related transcription factor 2 (Doublesex-like 2 protein) (DSXL-2). | |||||
|
HEMGN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.022973 (rank : 118) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ERZ0, Q3TAE6, Q80YT2, Q8CF29 | Gene names | Hemgn, Ndr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemogen (Hemopoietic gene protein) (Negative differentiation regulator protein) (mNDR). | |||||
|
HMGB1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.012850 (rank : 143) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P63158, P07155, P27109, P27428 | Gene names | Hmgb1, Hmg-1, Hmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein B1 (High mobility group protein 1) (HMG- 1). | |||||
|
JIP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.014752 (rank : 140) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UQF2, O43407 | Gene names | MAPK8IP1, IB1, JIP1 | |||
|
Domain Architecture |
|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 1 (JNK-interacting protein 1) (JIP-1) (JNK MAP kinase scaffold protein 1) (Islet-brain 1) (IB-1) (Mitogen-activated protein kinase 8-interacting protein 1). | |||||
|
JPH2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.012549 (rank : 144) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
NCOA5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | 0.018939 (rank : 130) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91W39 | Gene names | Ncoa5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 5 (NCoA-5) (Coactivator independent of AF-2) (CIA). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 147) | NC score | 0.065071 (rank : 23) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
OFD1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 148) | NC score | 0.015167 (rank : 139) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75665, O75666 | Gene names | OFD1, CXorf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oral-facial-digital syndrome 1 protein (Protein 71-7A). | |||||
|
PROX1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 149) | NC score | 0.013465 (rank : 142) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92786, Q5SW76, Q8TB91 | Gene names | PROX1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox prospero-like protein PROX1 (PROX 1). | |||||
|
PTMS_MOUSE
|
||||||
| θ value | 8.99809 (rank : 150) | NC score | 0.045774 (rank : 58) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
RBM19_MOUSE
|
||||||
| θ value | 8.99809 (rank : 151) | NC score | 0.009987 (rank : 151) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R3C6, Q8BHR0, Q9CW63 | Gene names | Rbm19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 19 (RNA-binding motif protein 19). | |||||
|
RHG06_HUMAN
|
||||||
| θ value | 8.99809 (rank : 152) | NC score | 0.010413 (rank : 150) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43182, O43437, Q9P1B3, Q9UK81, Q9UK82 | Gene names | ARHGAP6, RHOGAP6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
SFR17_HUMAN
|
||||||
| θ value | 8.99809 (rank : 153) | NC score | 0.030896 (rank : 88) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 154) | NC score | 0.041828 (rank : 63) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
STAR8_MOUSE
|
||||||
| θ value | 8.99809 (rank : 155) | NC score | 0.008427 (rank : 156) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K031, Q3UZC7, Q6A0A8, Q6P5E0, Q8R3X8 | Gene names | Stard8, Kiaa0189 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | StAR-related lipid transfer protein 8 (StARD8) (START domain- containing protein 8). | |||||
|
TCF20_HUMAN
|
||||||
| θ value | 8.99809 (rank : 156) | NC score | 0.032709 (rank : 83) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UGU0, O14528, Q13078, Q4V353, Q9H4M0 | Gene names | TCF20, KIAA0292, SPBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP) (AR1). | |||||
|
TPX2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 157) | NC score | 0.023501 (rank : 114) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULW0, Q9H1R4, Q9NRA3, Q9UFN9, Q9UL00, Q9Y2M1 | Gene names | TPX2, C20orf1, C20orf2, DIL2, HCA519 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Targeting protein for Xklp2 (Restricted expression proliferation- associated protein 100) (p100) (Differentially expressed in cancerous and noncancerous lung cells 2) (DIL-2) (Protein FLS353) (Hepatocellular carcinoma-associated antigen 519). | |||||
|
TTC16_HUMAN
|
||||||
| θ value | 8.99809 (rank : 158) | NC score | 0.036863 (rank : 71) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8NEE8, Q5JU66, Q96M72 | Gene names | TTC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.056859 (rank : 43) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
IWS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.063111 (rank : 27) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.056936 (rank : 42) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
NFH_MOUSE
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.057704 (rank : 40) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
RING1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.062450 (rank : 31) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q06587, Q5JP96, Q5SQW2, Q86V19 | Gene names | RING1, RNF1 | |||
|
Domain Architecture |
|
|||||
| Description | Polycomb complex protein RING1 (RING finger protein 1). | |||||
|
RING1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.061291 (rank : 35) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35730, Q3U242, Q3U333, Q4FK33, Q63ZX8, Q921Z8 | Gene names | Ring1, Ring1A, Rnf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycomb complex protein RING1 (RING finger protein 1) (Transcription repressor Ring1A). | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.057295 (rank : 41) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.071203 (rank : 17) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
SEN34_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 2.40964e-170 (rank : 1) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 158 | |
| SwissProt Accessions | Q8BMZ5, Q58EU2, Q99LV6, Q9CYW1 | Gene names | Tsen34, Leng5, Sen34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-splicing endonuclease subunit Sen34 (EC 3.1.27.9) (tRNA-intron endonuclease Sen34) (Leukocyte receptor cluster member 5 homolog). | |||||
|
SEN34_HUMAN
|
||||||
| NC score | 0.923826 (rank : 2) | θ value | 4.13891e-146 (rank : 2) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BSV6, Q9BVT1, Q9H6H5 | Gene names | TSEN34, LENG5, SEN34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-splicing endonuclease subunit Sen34 (EC 3.1.27.9) (tRNA-intron endonuclease Sen34) (HsSen34) (Leukocyte receptor cluster member 5). | |||||
|
RING2_MOUSE
|
||||||
| NC score | 0.133902 (rank : 3) | θ value | 0.00175202 (rank : 4) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CQJ4, O35699, O35729, Q4FJV5, Q8C1X8 | Gene names | Rnf2, DinG, Ring1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein RING2 (EC 6.3.2.-) (RING finger protein 2) (RING finger protein 1B) (RING1b). | |||||
|
RING2_HUMAN
|
||||||
| NC score | 0.133890 (rank : 4) | θ value | 0.00175202 (rank : 3) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99496, Q5TEN1, Q5TEN2 | Gene names | RNF2, BAP1, DING, HIPI3, RING1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein RING2 (EC 6.3.2.-) (RING finger protein 2) (RING finger protein 1B) (RING1b) (RING finger protein BAP-1) (DinG protein) (Huntingtin-interacting protein 2-interacting protein 3) (HIP2-interacting protein 3). | |||||
|
SEN2_HUMAN
|
||||||
| NC score | 0.126909 (rank : 5) | θ value | 0.125558 (rank : 14) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NCE0, Q8WTW7, Q9BPU7 | Gene names | TSEN2, SEN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-splicing endonuclease subunit Sen2 (EC 3.1.27.9) (tRNA-intron endonuclease Sen2) (HsSen2 protein). | |||||
|
SEN2_MOUSE
|
||||||
| NC score | 0.116362 (rank : 6) | θ value | 0.365318 (rank : 25) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P7W5, Q3UYG6 | Gene names | Tsen2, Sen2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-splicing endonuclease subunit Sen2 (EC 3.1.27.9) (tRNA-intron endonuclease Sen2). | |||||
|
MARCS_MOUSE
|
||||||
| NC score | 0.112301 (rank : 7) | θ value | 0.0961366 (rank : 10) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
MARCS_HUMAN
|
||||||
| NC score | 0.101044 (rank : 8) | θ value | 0.21417 (rank : 22) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
TR150_HUMAN
|
||||||
| NC score | 0.100099 (rank : 9) | θ value | 0.0330416 (rank : 6) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y2W1, Q5VTK6 | Gene names | THRAP3, TRAP150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
TR150_MOUSE
|
||||||
| NC score | 0.093600 (rank : 10) | θ value | 0.813845 (rank : 38) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
LYAR_MOUSE
|
||||||
| NC score | 0.091128 (rank : 11) | θ value | 0.0431538 (rank : 8) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q08288, Q9D9X2 | Gene names | Lyar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth-regulating nucleolar protein. | |||||
|
NASP_HUMAN
|
||||||
| NC score | 0.079961 (rank : 12) | θ value | 2.36792 (rank : 75) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
SCG1_MOUSE
|
||||||
| NC score | 0.074737 (rank : 13) | θ value | 0.125558 (rank : 13) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P16014 | Gene names | Chgb, Scg-1, Scg1 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-1 precursor (Secretogranin I) (SgI) (Chromogranin B) (CgB). | |||||
|
MRP_MOUSE
|
||||||
| NC score | 0.074458 (rank : 14) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P28667, Q3TEZ4, Q91W07 | Gene names | Marcksl1, Mlp, Mrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MARCKS-related protein (MARCKS-like protein 1) (Macrophage myristoylated alanine-rich C kinase substrate) (Mac-MARCKS) (MacMARCKS) (Brain protein F52). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.072606 (rank : 15) | θ value | 0.163984 (rank : 18) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
CA106_HUMAN
|
||||||
| NC score | 0.071484 (rank : 16) | θ value | 0.47712 (rank : 27) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q3KP66, Q9NV65, Q9NVI0 | Gene names | C1orf106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf106. | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.071203 (rank : 17) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
SRCA_HUMAN
|
||||||
| NC score | 0.070703 (rank : 18) | θ value | 0.0736092 (rank : 9) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
PTMS_HUMAN
|
||||||
| NC score | 0.070654 (rank : 19) | θ value | 1.81305 (rank : 61) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.069938 (rank : 20) | θ value | 0.21417 (rank : 21) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
LNP_MOUSE
|
||||||
| NC score | 0.068684 (rank : 21) | θ value | 0.47712 (rank : 29) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TQ95, Q69ZC5, Q6PAQ1, Q6PEN8 | Gene names | Lnp, Kiaa1715, Uln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein lunapark (Protein ulnaless). | |||||
|
CA106_MOUSE
|
||||||
| NC score | 0.068142 (rank : 22) | θ value | 0.813845 (rank : 35) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7TN12, Q7TN27, Q8BGK3 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf106 homolog. | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.065071 (rank : 23) | θ value | 8.99809 (rank : 147) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.064687 (rank : 24) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
TBP_HUMAN
|
||||||
| NC score | 0.064495 (rank : 25) | θ value | 0.163984 (rank : 19) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20226, Q16845, Q9UC02 | Gene names | TBP, TF2D, TFIID | |||
|
Domain Architecture |

|
|||||
| Description | TATA-box-binding protein (TATA-box factor) (TATA-binding factor) (TATA sequence-binding protein) (Transcription initiation factor TFIID TBP subunit). | |||||
|
TRI44_HUMAN
|
||||||
| NC score | 0.064446 (rank : 26) | θ value | 0.125558 (rank : 16) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96DX7, Q96QY2, Q9UGK0 | Gene names | TRIM44, DIPB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.063111 (rank : 27) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
CLSPN_HUMAN
|
||||||
| NC score | 0.062741 (rank : 28) | θ value | 0.125558 (rank : 11) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9HAW4, Q5VYG0, Q6P6H5, Q8IWI1 | Gene names | CLSPN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin (hClaspin) (Hu-Claspin). | |||||
|
TRI44_MOUSE
|
||||||
| NC score | 0.062612 (rank : 29) | θ value | 0.365318 (rank : 26) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QXA7, Q80SV9, Q9JHR8 | Gene names | Trim44, Dipb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB) (Protein Mc7). | |||||
|
MRP_HUMAN
|
||||||
| NC score | 0.062478 (rank : 30) | θ value | 6.88961 (rank : 128) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P49006, Q5TEE6, Q6NXS5 | Gene names | MARCKSL1, MLP, MRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MARCKS-related protein (MARCKS-like protein 1) (Macrophage myristoylated alanine-rich C kinase substrate) (Mac-MARCKS) (MacMARCKS). | |||||
|
RING1_HUMAN
|
||||||
| NC score | 0.062450 (rank : 31) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q06587, Q5JP96, Q5SQW2, Q86V19 | Gene names | RING1, RNF1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycomb complex protein RING1 (RING finger protein 1). | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.062202 (rank : 32) | θ value | 0.365318 (rank : 24) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
SNPC4_HUMAN
|
||||||
| NC score | 0.061605 (rank : 33) | θ value | 0.163984 (rank : 17) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5SXM2, Q9Y6P7 | Gene names | SNAPC4, SNAP190 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit) (Proximal sequence element-binding transcription factor subunit alpha) (PSE-binding factor subunit alpha) (PTF subunit alpha). | |||||
|
SRCA_MOUSE
|
||||||
| NC score | 0.061345 (rank : 34) | θ value | 1.06291 (rank : 45) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7TQ48, Q8BS43 | Gene names | Srl, Sar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
RING1_MOUSE
|
||||||
| NC score | 0.061291 (rank : 35) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35730, Q3U242, Q3U333, Q4FK33, Q63ZX8, Q921Z8 | Gene names | Ring1, Ring1A, Rnf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycomb complex protein RING1 (RING finger protein 1) (Transcription repressor Ring1A). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.061233 (rank : 36) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.058525 (rank : 37) | θ value | 3.0926 (rank : 85) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
TM16H_MOUSE
|
||||||
| NC score | 0.058202 (rank : 38) | θ value | 0.125558 (rank : 15) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PB70, Q69ZE4 | Gene names | Tmem16h, Kiaa1623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 16H. | |||||
|
SAM14_HUMAN
|
||||||
| NC score | 0.057952 (rank : 39) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IZD0, Q8N2X0 | Gene names | SAMD14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha motif domain-containing protein 14. | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.057704 (rank : 40) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.057295 (rank : 41) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.056936 (rank : 42) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.056859 (rank : 43) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
IPP2_MOUSE
|
||||||
| NC score | 0.056663 (rank : 44) | θ value | 0.62314 (rank : 32) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DCL8, Q3UD25, Q8C1A6 | Gene names | Ppp1r2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase inhibitor 2 (IPP-2). | |||||
|
SAM14_MOUSE
|
||||||
| NC score | 0.056379 (rank : 45) | θ value | 2.36792 (rank : 78) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K070, Q5SWB7, Q8BHE2, Q8C8N5 | Gene names | Samd14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha motif domain-containing protein 14. | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.055936 (rank : 46) | θ value | 2.36792 (rank : 74) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.055566 (rank : 47) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.055162 (rank : 48) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
ZCH10_HUMAN
|
||||||
| NC score | 0.054636 (rank : 49) | θ value | 2.36792 (rank : 83) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8TBK6, Q9NXR4 | Gene names | ZCCHC10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 10. | |||||
|
ARI1A_HUMAN
|
||||||
| NC score | 0.053736 (rank : 50) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
LNP_HUMAN
|
||||||
| NC score | 0.052721 (rank : 51) | θ value | 5.27518 (rank : 116) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9C0E8, Q2M2V8, Q2YD99, Q658W8, Q8N5V9, Q96MS5 | Gene names | LNP, KIAA1715 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein lunapark. | |||||
|
PDXL2_MOUSE
|
||||||
| NC score | 0.052152 (rank : 52) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8CAE9, Q8CFW3 | Gene names | Podxl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.048777 (rank : 53) | θ value | 1.81305 (rank : 60) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
NEUM_HUMAN
|
||||||
| NC score | 0.048243 (rank : 54) | θ value | 3.0926 (rank : 88) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P17677 | Gene names | GAP43 | |||
|
Domain Architecture |

|
|||||
| Description | Neuromodulin (Axonal membrane protein GAP-43) (Growth-associated protein 43) (PP46) (Neural phosphoprotein B-50). | |||||
|
TEX2_HUMAN
|
||||||
| NC score | 0.048243 (rank : 55) | θ value | 2.36792 (rank : 81) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IWB9, Q6AHZ5, Q8N3L0, Q9C0C5 | Gene names | TEX2, KIAA1738 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 2 protein. | |||||
|
TEX2_MOUSE
|
||||||
| NC score | 0.048089 (rank : 56) | θ value | 2.36792 (rank : 82) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZPJ0 | Gene names | Tex2, Kiaa1738 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 2 protein. | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.046580 (rank : 57) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
PTMS_MOUSE
|
||||||
| NC score | 0.045774 (rank : 58) | θ value | 8.99809 (rank : 150) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
HPS4_HUMAN
|
||||||
| NC score | 0.045697 (rank : 59) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NQG7, Q5H8V6, Q96LX6, Q9BY93, Q9UH37, Q9UH38 | Gene names | HPS4, KIAA1667 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 4 protein (Light-ear protein homolog). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.045415 (rank : 60) | θ value | 6.88961 (rank : 130) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TAF10_HUMAN
|
||||||
| NC score | 0.044785 (rank : 61) | θ value | 2.36792 (rank : 80) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12962, O00703, Q13175 | Gene names | TAF10, TAF2H, TAFII30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 10 (Transcription initiation factor TFIID 30 kDa subunit) (TAF(II)30) (TAFII-30) (TAFII30) (STAF28). | |||||
|
RP1L1_MOUSE
|
||||||
| NC score | 0.042459 (rank : 62) | θ value | 1.06291 (rank : 44) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CGM2, Q8K0I4 | Gene names | Rp1l1, Rp1hl1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein (Retinitis pigmentosa 1-like protein 1). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.041828 (rank : 63) | θ value | 8.99809 (rank : 154) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
NFM_MOUSE
|
||||||
| NC score | 0.040821 (rank : 64) | θ value | 0.00665767 (rank : 5) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
FAM9A_HUMAN
|
||||||
| NC score | 0.039950 (rank : 65) | θ value | 3.0926 (rank : 84) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IZU1 | Gene names | FAM9A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM9A. | |||||
|
PO121_MOUSE
|
||||||
| NC score | 0.039920 (rank : 66) | θ value | 1.38821 (rank : 52) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
TTC28_HUMAN
|
||||||
| NC score | 0.039194 (rank : 67) | θ value | 0.62314 (rank : 33) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96AY4, O95928, O95929, Q9NTE4, Q9UG31, Q9UGG5, Q9UPV8, Q9Y3S5 | Gene names | TTC28, KIAA1043 | |||
|
Domain Architecture |

|
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
TCF20_MOUSE
|
||||||
| NC score | 0.038677 (rank : 68) | θ value | 5.27518 (rank : 121) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
NFM_HUMAN
|
||||||
| NC score | 0.038596 (rank : 69) | θ value | 0.125558 (rank : 12) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
UBF1_MOUSE
|
||||||
| NC score | 0.038139 (rank : 70) | θ value | 1.06291 (rank : 47) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P25976 | Gene names | Ubtf, Tcfubf, Ubf-1, Ubf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1). | |||||
|
TTC16_HUMAN
|
||||||
| NC score | 0.036863 (rank : 71) | θ value | 8.99809 (rank : 158) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8NEE8, Q5JU66, Q96M72 | Gene names | TTC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.036346 (rank : 72) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
DCDC2_HUMAN
|
||||||
| NC score | 0.035743 (rank : 73) | θ value | 4.03905 (rank : 98) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UHG0, Q5VTR8, Q5VTR9, Q86W35, Q9UFD1, Q9ULR6 | Gene names | DCDC2, KIAA1154 | |||
|
Domain Architecture |

|
|||||
| Description | Doublecortin domain-containing protein 2 (RU2S protein). | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.035607 (rank : 74) | θ value | 1.06291 (rank : 43) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
MCRS1_MOUSE
|
||||||
| NC score | 0.034884 (rank : 75) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99L90, O35255, Q32P11 | Gene names | Mcrs1, Msp58 | |||
|
Domain Architecture |

|
|||||
| Description | Microspherule protein 1 (58 kDa microspherule protein). | |||||
|
ACRBP_MOUSE
|
||||||
| NC score | 0.034723 (rank : 76) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3V140, Q62253, Q62254, Q8C621, Q91VQ1 | Gene names | Acrbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acrosin-binding protein precursor (Proacrosin-binding protein sp32). | |||||
|
TNC6B_HUMAN
|
||||||
| NC score | 0.034010 (rank : 77) | θ value | 4.03905 (rank : 105) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
MARH4_MOUSE
|
||||||
| NC score | 0.033584 (rank : 78) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80TE3 | Gene names | March4, Kiaa1399 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated RING finger protein 4 (Membrane-associated RING-CH protein IV) (MARCH-IV). | |||||
|
HTSF1_MOUSE
|
||||||
| NC score | 0.033231 (rank : 79) | θ value | 6.88961 (rank : 126) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.033162 (rank : 80) | θ value | 5.27518 (rank : 118) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
JHD2B_HUMAN
|
||||||
| NC score | 0.033153 (rank : 81) | θ value | 0.47712 (rank : 28) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7LBC6, Q9BVH6, Q9BW93, Q9BZ52, Q9NYF4, Q9UPS0 | Gene names | JMJD1B, C5orf7, JHDM2B, KIAA1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B) (Nuclear protein 5qNCA). | |||||
|
CMTA2_MOUSE
|
||||||
| NC score | 0.032869 (rank : 82) | θ value | 1.38821 (rank : 48) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80Y50, Q3TFK0, Q5SX68, Q80TP1, Q8R0D9, Q8R2N5 | Gene names | Camta2, Kiaa0909 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-binding transcription activator 2. | |||||
|
TCF20_HUMAN
|
||||||
| NC score | 0.032709 (rank : 83) | θ value | 8.99809 (rank : 156) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UGU0, O14528, Q13078, Q4V353, Q9H4M0 | Gene names | TCF20, KIAA0292, SPBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP) (AR1). | |||||
|
MYRIP_HUMAN
|
||||||
| NC score | 0.032254 (rank : 84) | θ value | 4.03905 (rank : 100) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8NFW9, Q8IUF5, Q9Y3V4 | Gene names | MYRIP, SLAC2C | |||
|
Domain Architecture |

|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
BL1S3_HUMAN
|
||||||
| NC score | 0.031931 (rank : 85) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6QNY0 | Gene names | BLOC1S3, BLOS3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Biogenesis of lysosome-related organelles complex-1 subunit 3 (BLOC subunit 3). | |||||
|
RALY_MOUSE
|
||||||
| NC score | 0.031602 (rank : 86) | θ value | 1.38821 (rank : 53) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q64012, Q99K76, Q9CXH8, Q9QZX6 | Gene names | Raly, Merc | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein Raly (hnRNP associated with lethal yellow protein) (Maternally expressed hnRNP C-related protein). | |||||
|
SHAN2_MOUSE
|
||||||
| NC score | 0.030976 (rank : 87) | θ value | 1.38821 (rank : 54) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 446 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80Z38, Q3UTK4, Q5DU07 | Gene names | Shank2, Kiaa1022 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2) (Cortactin- binding protein 1) (CortBP1). | |||||
|
SFR17_HUMAN
|
||||||
| NC score | 0.030896 (rank : 88) | θ value | 8.99809 (rank : 153) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
CTDP1_HUMAN
|
||||||
| NC score | 0.030660 (rank : 89) | θ value | 5.27518 (rank : 111) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
NFL_HUMAN
|
||||||
| NC score | 0.029518 (rank : 90) | θ value | 0.813845 (rank : 37) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 666 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P07196, Q16154, Q8IU72 | Gene names | NEFL, NF68, NFL | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet L protein (68 kDa neurofilament protein) (Neurofilament light polypeptide) (NF-L). | |||||
|
PDC6I_HUMAN
|
||||||
| NC score | 0.029062 (rank : 91) | θ value | 2.36792 (rank : 76) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WUM4, Q9BX86, Q9NUN0, Q9P2H2, Q9UKL5 | Gene names | PDCD6IP, AIP1, ALIX, KIAA1375 | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death 6-interacting protein (PDCD6-interacting protein) (ALG-2-interacting protein 1) (Hp95). | |||||
|
BMP6_MOUSE
|
||||||
| NC score | 0.028898 (rank : 92) | θ value | 0.0431538 (rank : 7) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P20722 | Gene names | Bmp6, Bmp-6, Vgr1 | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein 6 precursor (BMP-6) (VG-1-related protein) (VGR-1). | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.028344 (rank : 93) | θ value | 3.0926 (rank : 87) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
NPHP1_MOUSE
|
||||||
| NC score | 0.027974 (rank : 94) | θ value | 1.38821 (rank : 51) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QY53, Q9D7G8, Q9QZP0, Q9WUZ2 | Gene names | Nphp1, Nph1 | |||
|
Domain Architecture |

|
|||||
| Description | Nephrocystin-1. | |||||
|
TBCD2_HUMAN
|
||||||
| NC score | 0.027549 (rank : 95) | θ value | 5.27518 (rank : 120) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BYX2 | Gene names | TBC1D2, PARIS1 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 2 (Prostate antigen recognized and identified by SEREX) (PARIS-1). | |||||
|
ELL3_MOUSE
|
||||||
| NC score | 0.027528 (rank : 96) | θ value | 4.03905 (rank : 99) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80VR2 | Gene names | Ell3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL3. | |||||
|
ZN414_HUMAN
|
||||||
| NC score | 0.026716 (rank : 97) | θ value | 1.38821 (rank : 56) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96IQ9 | Gene names | ZNF414 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 414. | |||||
|
ARS2_MOUSE
|
||||||
| NC score | 0.026134 (rank : 98) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99MR6, Q99MR4, Q99MR5, Q99MR7 | Gene names | Ars2, Asr2 | |||
|
Domain Architecture |

|
|||||
| Description | Arsenite-resistance protein 2. | |||||
|
IF4G3_MOUSE
|
||||||
| NC score | 0.025794 (rank : 99) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80XI3, Q80Y69, Q8BJ68, Q8BQF3, Q8C6Q3, Q8CIH0 | Gene names | Eif4g3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
STK23_MOUSE
|
||||||
| NC score | 0.025740 (rank : 100) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 572 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Z0G2 | Gene names | Stk23, Mssk1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase 23 (EC 2.7.11.1) (Muscle-specific serine kinase 1) (MSSK-1). | |||||
|
CENPB_MOUSE
|
||||||
| NC score | 0.025454 (rank : 101) | θ value | 5.27518 (rank : 109) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P27790 | Gene names | Cenpb, Cenp-b | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
MYT1_MOUSE
|
||||||
| NC score | 0.025438 (rank : 102) | θ value | 1.38821 (rank : 50) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
IF2P_HUMAN
|
||||||
| NC score | 0.025209 (rank : 103) | θ value | 6.88961 (rank : 127) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
BAZ2A_HUMAN
|
||||||
| NC score | 0.024911 (rank : 104) | θ value | 4.03905 (rank : 94) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
BSPRY_MOUSE
|
||||||
| NC score | 0.024809 (rank : 105) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80YW5, Q3TU74, Q8BZF0, Q99KV7, Q9ER70 | Gene names | Bspry | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B box and SPRY domain-containing protein. | |||||
|
CDSN_MOUSE
|
||||||
| NC score | 0.024787 (rank : 106) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TPC1 | Gene names | Cdsn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor. | |||||
|
CS007_HUMAN
|
||||||
| NC score | 0.024594 (rank : 107) | θ value | 5.27518 (rank : 110) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
ASTL_HUMAN
|
||||||
| NC score | 0.024572 (rank : 108) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6HA08 | Gene names | ASTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Astacin-like metalloendopeptidase precursor (EC 3.4.-.-) (Oocyte astacin) (Ovastacin). | |||||
|
RIN3_HUMAN
|
||||||
| NC score | 0.024017 (rank : 109) | θ value | 4.03905 (rank : 103) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TB24, Q8NF30, Q8TEE8, Q8WYP4, Q9H6A5, Q9HAG1 | Gene names | RIN3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
DAXX_MOUSE
|
||||||
| NC score | 0.023994 (rank : 110) | θ value | 5.27518 (rank : 112) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O35613, Q9QWT8, Q9QWV3 | Gene names | Daxx | |||
|
Domain Architecture |

|
|||||
| Description | Death domain-associated protein 6 (Daxx). | |||||
|
DACH1_MOUSE
|
||||||
| NC score | 0.023882 (rank : 111) | θ value | 4.03905 (rank : 97) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QYB2, O88716, Q8BPQ0, Q8C8D7, Q9QY47, Q9R218, Q9Z0Y5 | Gene names | Dach1, Dach | |||
|
Domain Architecture |

|
|||||
| Description | Dachshund homolog 1 (Dach1). | |||||
|
RGP1_MOUSE
|
||||||
| NC score | 0.023811 (rank : 112) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46061, Q60801 | Gene names | Rangap1, Fug1 | |||
|
Domain Architecture |

|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
RBM28_MOUSE
|
||||||
| NC score | 0.023716 (rank : 113) | θ value | 3.0926 (rank : 89) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8CGC6, Q8BG25, Q8VEJ8, Q9CS22, Q9CSE6 | Gene names | Rbm28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 28 (RNA-binding motif protein 28). | |||||
|
TPX2_HUMAN
|
||||||
| NC score | 0.023501 (rank : 114) | θ value | 8.99809 (rank : 157) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULW0, Q9H1R4, Q9NRA3, Q9UFN9, Q9UL00, Q9Y2M1 | Gene names | TPX2, C20orf1, C20orf2, DIL2, HCA519 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Targeting protein for Xklp2 (Restricted expression proliferation- associated protein 100) (p100) (Differentially expressed in cancerous and noncancerous lung cells 2) (DIL-2) (Protein FLS353) (Hepatocellular carcinoma-associated antigen 519). | |||||
|
CD2AP_MOUSE
|
||||||
| NC score | 0.023366 (rank : 115) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9JLQ0, O88903, Q8K4Z1, Q8VCI9 | Gene names | Cd2ap, Mets1 | |||
|
Domain Architecture |

|
|||||
| Description | CD2-associated protein (Mesenchyme-to-epithelium transition protein with SH3 domains 1) (METS-1). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.023292 (rank : 116) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
UN119_MOUSE
|
||||||
| NC score | 0.023022 (rank : 117) | θ value | 6.88961 (rank : 132) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z2R6 | Gene names | Unc119, Unc119h | |||
|
Domain Architecture |

|
|||||
| Description | Unc-119 protein homolog (Retinal protein 4) (MRG4). | |||||
|
HEMGN_MOUSE
|
||||||
| NC score | 0.022973 (rank : 118) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ERZ0, Q3TAE6, Q80YT2, Q8CF29 | Gene names | Hemgn, Ndr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemogen (Hemopoietic gene protein) (Negative differentiation regulator protein) (mNDR). | |||||
|
TIF1G_HUMAN
|
||||||
| NC score | 0.022896 (rank : 119) | θ value | 1.06291 (rank : 46) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
DDX51_HUMAN
|
||||||
| NC score | 0.022847 (rank : 120) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
UBP42_HUMAN
|
||||||
| NC score | 0.022531 (rank : 121) | θ value | 3.0926 (rank : 91) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H9J4, Q3C166, Q6P9B4 | Gene names | USP42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 42 (EC 3.1.2.15) (Ubiquitin thioesterase 42) (Ubiquitin-specific-processing protease 42) (Deubiquitinating enzyme 42). | |||||
|
CAC1C_HUMAN
|
||||||
| NC score | 0.021782 (rank : 122) | θ value | 1.06291 (rank : 39) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
TMC2_HUMAN
|
||||||
| NC score | 0.021749 (rank : 123) | θ value | 3.0926 (rank : 90) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8TDI7, Q5JXT0, Q5JXT1, Q6UWW4, Q6ZS41, Q8N9F3, Q9BYN2, Q9BYN3, Q9BYN4, Q9BYN5 | Gene names | TMC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane cochlear-expressed protein 2. | |||||
|
HNRLL_MOUSE
|
||||||
| NC score | 0.021515 (rank : 124) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q921F4, Q8BIP6, Q99J40 | Gene names | Hnrpll | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L-like. | |||||
|
CEP35_HUMAN
|
||||||
| NC score | 0.021390 (rank : 125) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
BCOR_MOUSE
|
||||||
| NC score | 0.021223 (rank : 126) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CGN4, Q6PDK5, Q6ZPM3, Q8BKF5, Q8CGN1, Q8CGN2, Q8CGN3 | Gene names | Bcor, Kiaa1575 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BCoR protein (BCL-6 corepressor). | |||||
|
HNRPU_HUMAN
|
||||||
| NC score | 0.019750 (rank : 127) | θ value | 6.88961 (rank : 125) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q00839, O75507, Q96HY9 | Gene names | HNRPU, SAFA, U21.1 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U (hnRNP U) (Scaffold attachment factor A) (SAF-A) (p120) (pp120). | |||||
|
SMAL1_MOUSE
|
||||||
| NC score | 0.019590 (rank : 128) | θ value | 2.36792 (rank : 79) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BJL0, Q8BVK7, Q8BXW4, Q8K309, Q9EQK8, Q9QYC4 | Gene names | Smarcal1, Harp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A-like protein 1 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 1) (HepA-related protein) (mharp). | |||||
|
MAGB6_HUMAN
|
||||||
| NC score | 0.019009 (rank : 129) | θ value | 5.27518 (rank : 117) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8N7X4, Q6GS19, Q9H219 | Gene names | MAGEB6 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen B6 (MAGE-B6 antigen). | |||||
|
NCOA5_MOUSE
|
||||||
| NC score | 0.018939 (rank : 130) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91W39 | Gene names | Ncoa5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 5 (NCoA-5) (Coactivator independent of AF-2) (CIA). | |||||
|
DMRT2_HUMAN
|
||||||
| NC score | 0.018477 (rank : 131) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y5R5 | Gene names | DMRT2, DSXL2 | |||
|
Domain Architecture |

|
|||||
| Description | Doublesex- and mab-3-related transcription factor 2 (Doublesex-like 2 protein) (DSXL-2). | |||||
|
ENAH_HUMAN
|
||||||
| NC score | 0.018372 (rank : 132) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8N8S7, Q502W5, Q5T5M7, Q5VTQ9, Q5VTR0, Q9NVF3, Q9UFB8 | Gene names | ENAH, MENA | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog. | |||||
|
BAZ1A_HUMAN
|
||||||
| NC score | 0.018147 (rank : 133) | θ value | 8.99809 (rank : 135) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
CTR9_MOUSE
|
||||||
| NC score | 0.017991 (rank : 134) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62018, Q3UFF5, Q3UY40, Q66JX4, Q7TPS6, Q8BND9, Q8BRD1, Q8C9W7, Q8C9Y3, Q8CHI1 | Gene names | Ctr9, Kiaa0155, Sh2bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein CTR9 homolog (SH2 domain-binding protein 1) (Tetratricopeptide repeat-containing, SH2-binding phosphoprotein of 150 kDa) (TPR-containing, SH2-binding phosphoprotein of 150 kDa) (p150TSP). | |||||
|
CAMKV_HUMAN
|
||||||
| NC score | 0.016965 (rank : 135) | θ value | 0.365318 (rank : 23) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8NCB2, Q6FIB8, Q8NBS8, Q8NC85, Q8NDU4, Q8WTT8, Q9BQC9, Q9H0Q5 | Gene names | CAMKV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CaM kinase-like vesicle-associated protein. | |||||
|
CAC1C_MOUSE
|
||||||
| NC score | 0.016198 (rank : 136) | θ value | 6.88961 (rank : 123) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q01815, Q04476, Q61242, Q99242 | Gene names | Cacna1c, Cach2, Cacn2, Cacnl1a1, Cchl1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle) (Mouse brain class C) (MBC) (MELC-CC). | |||||
|
IQEC3_MOUSE
|
||||||
| NC score | 0.015374 (rank : 137) | θ value | 5.27518 (rank : 115) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
SOX12_HUMAN
|
||||||
| NC score | 0.015207 (rank : 138) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15370, Q9NUD4 | Gene names | SOX12, SOX22 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-12 protein (SOX-22 protein). | |||||
|
OFD1_HUMAN
|
||||||
| NC score | 0.015167 (rank : 139) | θ value | 8.99809 (rank : 148) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75665, O75666 | Gene names | OFD1, CXorf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oral-facial-digital syndrome 1 protein (Protein 71-7A). | |||||
|
JIP1_HUMAN
|
||||||
| NC score | 0.014752 (rank : 140) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UQF2, O43407 | Gene names | MAPK8IP1, IB1, JIP1 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 1 (JNK-interacting protein 1) (JIP-1) (JNK MAP kinase scaffold protein 1) (Islet-brain 1) (IB-1) (Mitogen-activated protein kinase 8-interacting protein 1). | |||||
|
SPEG_HUMAN
|
||||||
| NC score | 0.014194 (rank : 141) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
PROX1_HUMAN
|
||||||
| NC score | 0.013465 (rank : 142) | θ value | 8.99809 (rank : 149) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92786, Q5SW76, Q8TB91 | Gene names | PROX1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox prospero-like protein PROX1 (PROX 1). | |||||
|
HMGB1_MOUSE
|
||||||
| NC score | 0.012850 (rank : 143) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P63158, P07155, P27109, P27428 | Gene names | Hmgb1, Hmg-1, Hmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein B1 (High mobility group protein 1) (HMG- 1). | |||||
|
JPH2_MOUSE
|
||||||
| NC score | 0.012549 (rank : 144) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
LHX3_MOUSE
|
||||||
| NC score | 0.012488 (rank : 145) | θ value | 0.813845 (rank : 36) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50481, Q61800, Q61801 | Gene names | Lhx3, Lim-3, Lim3, Plim | |||
|
Domain Architecture |

|
|||||
| Description | LIM/homeobox protein Lhx3 (Homeobox protein LIM-3) (Homeobox protein P-LIM) (LIM homeobox protein 3). | |||||
|
CAF1B_MOUSE
|
||||||
| NC score | 0.012288 (rank : 146) | θ value | 4.03905 (rank : 95) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D0N7 | Gene names | Chaf1b | |||
|
Domain Architecture |

|
|||||
| Description | Chromatin assembly factor 1 subunit B (CAF-1 subunit B) (Chromatin assembly factor I p60 subunit) (CAF-I 60 kDa subunit) (CAF-Ip60). | |||||
|
MYLK2_HUMAN
|
||||||
| NC score | 0.012140 (rank : 147) | θ value | 0.47712 (rank : 30) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H1R3, Q96I84 | Gene names | MYLK2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin light chain kinase 2, skeletal/cardiac muscle (EC 2.7.11.18) (MLCK2). | |||||
|
CENG1_MOUSE
|
||||||
| NC score | 0.010944 (rank : 148) | θ value | 4.03905 (rank : 96) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3UHD9, Q5DU45 | Gene names | Centg1, Agap2, Kiaa0167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-gamma 1 (ARF-GAP with GTP-binding protein-like, ankyrin repeat and pleckstrin homology domains 2) (AGAP-2) (Phosphatidylinositol-3-kinase enhancer) (PIKE). | |||||
|
ANDR_HUMAN
|
||||||
| NC score | 0.010864 (rank : 149) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P10275 | Gene names | AR, DHTR, NR3C4 | |||
|
Domain Architecture |

|
|||||
| Description | Androgen receptor (Dihydrotestosterone receptor). | |||||
|
RHG06_HUMAN
|
||||||
| NC score | 0.010413 (rank : 150) | θ value | 8.99809 (rank : 152) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43182, O43437, Q9P1B3, Q9UK81, Q9UK82 | Gene names | ARHGAP6, RHOGAP6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
RBM19_MOUSE
|
||||||
| NC score | 0.009987 (rank : 151) | θ value | 8.99809 (rank : 151) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R3C6, Q8BHR0, Q9CW63 | Gene names | Rbm19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 19 (RNA-binding motif protein 19). | |||||
|
M4K4_MOUSE
|
||||||
| NC score | 0.009430 (rank : 152) | θ value | 1.38821 (rank : 49) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1560 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P97820 | Gene names | Map4k4, Nik | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase kinase 4 (EC 2.7.11.1) (MAPK/ERK kinase kinase kinase 4) (MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase HGK) (Nck-interacting kinase). | |||||
|
DHX37_HUMAN
|
||||||
| NC score | 0.008924 (rank : 153) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IY37, Q9BUI7, Q9P211 | Gene names | DHX37, DDX37, KIAA1517 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DHX37 (EC 3.6.1.-) (DEAH box protein 37). | |||||
|
B3A3_HUMAN
|
||||||
| NC score | 0.008789 (rank : 154) | θ value | 6.88961 (rank : 122) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P48751 | Gene names | SLC4A3, AE3 | |||
|
Domain Architecture |

|
|||||
| Description | Anion exchange protein 3 (Neuronal band 3-like protein) (Solute carrier family 4 member 3) (Cardiac/brain band 3-like protein) (CAE3/BAE3). | |||||
|
ZN447_HUMAN
|
||||||
| NC score | 0.008574 (rank : 155) | θ value | 4.03905 (rank : 106) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TBC5, Q9BRK7, Q9H9A0 | Gene names | ZNF447 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 447. | |||||
|
STAR8_MOUSE
|
||||||
| NC score | 0.008427 (rank : 156) | θ value | 8.99809 (rank : 155) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K031, Q3UZC7, Q6A0A8, Q6P5E0, Q8R3X8 | Gene names | Stard8, Kiaa0189 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | StAR-related lipid transfer protein 8 (StARD8) (START domain- containing protein 8). | |||||
|
MK07_HUMAN
|
||||||
| NC score | 0.007913 (rank : 157) | θ value | 3.0926 (rank : 86) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1087 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13164, Q16634 | Gene names | MAPK7, ERK4, ERK5, PRKM7 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (ERK4) (BMK1 kinase). | |||||
|
PLK2_MOUSE
|
||||||
| NC score | 0.007528 (rank : 158) | θ value | 2.36792 (rank : 77) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P53351 | Gene names | Plk2, Snk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase PLK2 (EC 2.7.11.21) (Polo-like kinase 1) (PLK-2) (Serine/threonine-protein kinase SNK) (Serum-inducible kinase). | |||||
|
SHOC2_HUMAN
|
||||||
| NC score | 0.007393 (rank : 159) | θ value | 5.27518 (rank : 119) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UQ13, O76063 | Gene names | SHOC2, KIAA0862 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat protein SHOC-2 (Ras-binding protein Sur-8). | |||||
|
HNF3A_HUMAN
|
||||||
| NC score | 0.007010 (rank : 160) | θ value | 5.27518 (rank : 114) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P55317, Q9H2A0 | Gene names | FOXA1, HNF3A, TCF3A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-alpha (HNF-3A) (Forkhead box protein A1). | |||||
|
SHOC2_MOUSE
|
||||||
| NC score | 0.006892 (rank : 161) | θ value | 6.88961 (rank : 129) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88520, Q3UJH6, Q8BVL0 | Gene names | Shoc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat protein SHOC-2 (Ras-binding protein Sur-8). | |||||
|
FARP2_HUMAN
|
||||||
| NC score | 0.006880 (rank : 162) | θ value | 6.88961 (rank : 124) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O94887, Q53QM5, Q8WU27, Q9UFE7 | Gene names | FARP2, KIAA0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
CD2L2_HUMAN
|
||||||
| NC score | 0.006865 (rank : 163) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 1439 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UQ88, O95227, O95228, O96012, Q12821, Q12853, Q12854, Q5QPR0, Q5QPR1, Q5QPR2, Q9UBC4, Q9UBI3, Q9UEI1, Q9UEI2, Q9UP53, Q9UP54, Q9UP55, Q9UP56, Q9UQ86, Q9UQ87, Q9UQ89 | Gene names | CDC2L2, PITSLREB | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L2 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 2) (CDK11). | |||||
|
SNRK_HUMAN
|
||||||
| NC score | 0.006781 (rank : 164) | θ value | 4.03905 (rank : 104) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 855 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NRH2, Q14706, Q68D15, Q6IQ46, Q9NXI7 | Gene names | SNRK, KIAA0096, SNFRK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SNF-related serine/threonine-protein kinase (EC 2.7.11.1) (SNF1- related kinase). | |||||
|
TAF5_HUMAN
|
||||||
| NC score | 0.003860 (rank : 165) | θ value | 6.88961 (rank : 131) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15542, Q86UZ7, Q9Y4K5 | Gene names | TAF5, TAF2D | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 5 (Transcription initiation factor TFIID 100 kDa subunit) (TAF(II)100) (TAFII-100) (TAFII100). | |||||
|
ZN696_HUMAN
|
||||||
| NC score | -0.001814 (rank : 166) | θ value | 3.0926 (rank : 92) | |||
| Query Neighborhood Hits | 158 | Target Neighborhood Hits | 743 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H7X3 | Gene names | ZNF696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 696. | |||||