Please be patient as the page loads
|
SAM68_MOUSE
|
||||||
| SwissProt Accessions | Q60749, Q3U8T3, Q60735, Q99M33 | Gene names | Khdrbs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SAM68_MOUSE
|
||||||
| θ value | 1.08667e-154 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q60749, Q3U8T3, Q60735, Q99M33 | Gene names | Khdrbs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
SAM68_HUMAN
|
||||||
| θ value | 1.68038e-147 (rank : 2) | NC score | 0.989054 (rank : 2) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q07666, Q6PJX7, Q8NB97, Q99760 | Gene names | KHDRBS1, SAM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
KHDR3_HUMAN
|
||||||
| θ value | 3.68522e-86 (rank : 3) | NC score | 0.961898 (rank : 3) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75525, Q6NUL8, Q9UPA8 | Gene names | KHDRBS3, SALP, SLM2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 3 (Sam68-like mammalian protein 2) (SLM-2) (Sam68-like phosphotyrosine protein) (RNA-binding protein T-Star). | |||||
|
KHDR3_MOUSE
|
||||||
| θ value | 7.68416e-84 (rank : 4) | NC score | 0.958979 (rank : 4) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9R226, O88624 | Gene names | Khdrbs3, Salp, Slm2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 3 (Sam68-like mammalian protein 2) (SLM-2) (RNA-binding protein Etoile). | |||||
|
QKI_HUMAN
|
||||||
| θ value | 6.38894e-30 (rank : 5) | NC score | 0.821772 (rank : 5) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96PU8, Q2I375, Q5MJQ1, Q969L9, Q96EJ3, Q96KA3, Q96PU6, Q96PU7, Q9P0X6, Q9P0X7, Q9P0X8, Q9P0X9, Q9P0Y0, Q9P0Y1 | Gene names | QKI, HKQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Quaking protein (Hqk). | |||||
|
QKI_MOUSE
|
||||||
| θ value | 6.38894e-30 (rank : 6) | NC score | 0.821772 (rank : 6) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QYS9, O88972, Q61110, Q78ZE4, Q78ZE5, Q8K4X9, Q8K4Y0, Q9CW34, Q9QUH4, Q9R2A8, Q9Z246 | Gene names | Qki, Qk, Qk1, Qka1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Quaking protein (qkI). | |||||
|
SF01_HUMAN
|
||||||
| θ value | 5.08577e-11 (rank : 7) | NC score | 0.534409 (rank : 8) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q15637, Q14818, Q14819, Q15913, Q8IY00, Q92744, Q92745, Q969H7, Q9BW01, Q9UEI0 | Gene names | SF1, ZFM1, ZNF162 | |||
|
Domain Architecture |
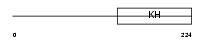
|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (Zinc finger gene in MEN1 locus) (Mammalian branch point-binding protein mBBP) (BBP). | |||||
|
SF01_MOUSE
|
||||||
| θ value | 8.67504e-11 (rank : 8) | NC score | 0.537540 (rank : 7) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q64213, O08817, P70167, Q61454, Q921Z4 | Gene names | Sf1, Zfm1, Zfp162 | |||
|
Domain Architecture |
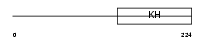
|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (mZFM) (Zinc finger gene in MEN1 locus) (Mammalian branch point- binding protein mBBP) (BBP) (CW17). | |||||
|
IF2B2_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 9) | NC score | 0.206324 (rank : 9) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5SF07, Q3TCU4, Q7TQF9 | Gene names | Igf2bp2, Imp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2). | |||||
|
FUBP3_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 10) | NC score | 0.185300 (rank : 10) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96I24, Q92946, Q9BVB6 | Gene names | FUBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 3 (FUSE-binding protein 3). | |||||
|
DCC_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 11) | NC score | 0.033626 (rank : 38) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
K0174_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 12) | NC score | 0.083017 (rank : 30) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P53990, Q9BQ81, Q9BWN2 | Gene names | KIAA0174 | |||
|
Domain Architecture |
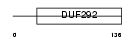
|
|||||
| Description | Protein KIAA0174 (Putative MAPK-activating protein PM28). | |||||
|
BIM_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 13) | NC score | 0.085536 (rank : 28) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43521, O43522 | Gene names | BCL2L11, BIM | |||
|
Domain Architecture |

|
|||||
| Description | Bcl-2-like protein 11 (Bcl2-interacting mediator of cell death). | |||||
|
NOVA2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 14) | NC score | 0.166275 (rank : 12) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UNW9, O43267, Q9UEA1 | Gene names | NOVA2, ANOVA, NOVA3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein Nova-2 (Neuro-oncological ventral antigen 2) (Astrocytic NOVA1-like RNA-binding protein). | |||||
|
BIM_MOUSE
|
||||||
| θ value | 0.163984 (rank : 15) | NC score | 0.083023 (rank : 29) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54918, O54919, O54920 | Gene names | Bcl2l11, Bim | |||
|
Domain Architecture |

|
|||||
| Description | Bcl-2-like protein 11 (Bcl2-interacting mediator of cell death). | |||||
|
FUBP1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 16) | NC score | 0.159580 (rank : 15) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96AE4, Q12828 | Gene names | FUBP1 | |||
|
Domain Architecture |
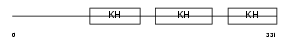
|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP) (DNA helicase V) (HDH V). | |||||
|
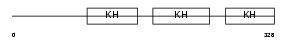
FUBP1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 17) | NC score | 0.159657 (rank : 14) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q91WJ8, Q8C0Y8 | Gene names | Fubp1, D3Ertd330e | |||
|
Domain Architecture |
|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP). | |||||
|
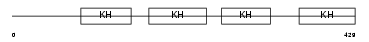
FUBP2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 18) | NC score | 0.165884 (rank : 13) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q92945, O00301, Q9UNT5, Q9UQH5 | Gene names | KHSRP, FUBP2 | |||
|
Domain Architecture |
|
|||||
| Description | Far upstream element-binding protein 2 (FUSE-binding protein 2) (KH type-splicing regulatory protein) (KSRP) (p75). | |||||
|
IF2B2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 19) | NC score | 0.180858 (rank : 11) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y6M1 | Gene names | IGF2BP2, IMP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2) (Hepatocellular carcinoma autoantigen p62). | |||||
|
PCBP3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.145850 (rank : 18) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P57721, Q8N9K6, Q96EP6 | Gene names | PCBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
PCBP3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 21) | NC score | 0.145919 (rank : 17) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P57722, Q8BSB0, Q8C544 | Gene names | Pcbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
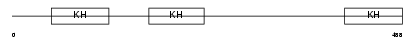
NOVA1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 22) | NC score | 0.158718 (rank : 16) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P51513 | Gene names | NOVA1 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding protein Nova-1 (Neuro-oncological ventral antigen 1) (Onconeural ventral antigen 1) (Paraneoplastic Ri antigen) (Ventral neuron-specific protein 1). | |||||
|
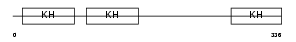
PCBP1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 23) | NC score | 0.141414 (rank : 19) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15365, Q13157, Q14975 | Gene names | PCBP1 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1) (Nucleic acid- binding protein SUB2.3). | |||||
|
PCBP1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 24) | NC score | 0.141414 (rank : 20) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P60335 | Gene names | Pcbp1 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 0.365318 (rank : 25) | NC score | 0.027722 (rank : 44) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
GLYG2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 26) | NC score | 0.036204 (rank : 37) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15488, O15485, O15486, O15487, O15489, O15490 | Gene names | GYG2 | |||
|
Domain Architecture |

|
|||||
| Description | Glycogenin-2 (EC 2.4.1.186) (GN-2) (GN2). | |||||
|
PCBP2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.136485 (rank : 21) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q15366, Q6PKG5 | Gene names | PCBP2 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (hnRNP-E2). | |||||
|
PCBP2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.136039 (rank : 22) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q61990, Q61383, Q62042 | Gene names | Pcbp2, Cbp, Hnrnpx, Hnrpx | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (Putative heterogeneous nuclear ribonucleoprotein X) (hnRNP X) (CTBP) (CBP). | |||||
|
K0174_MOUSE
|
||||||
| θ value | 0.813845 (rank : 29) | NC score | 0.070859 (rank : 31) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CX00, Q80U68 | Gene names | Kiaa0174 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0174. | |||||
|
M3K11_HUMAN
|
||||||
| θ value | 1.06291 (rank : 30) | NC score | 0.004941 (rank : 69) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1136 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q16584, Q6P2G4 | Gene names | MAP3K11, MLK3, SPRK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 11 (EC 2.7.11.25) (Mixed lineage kinase 3) (Src-homology 3 domain-containing proline- rich kinase). | |||||
|
DCC_HUMAN
|
||||||
| θ value | 1.38821 (rank : 31) | NC score | 0.022842 (rank : 48) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 583 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P43146 | Gene names | DCC | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC) (Colorectal cancer suppressor). | |||||
|
PCBP4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.133084 (rank : 24) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P57723, Q96AH7 | Gene names | PCBP4 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
PCBP4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.133533 (rank : 23) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P57724, Q91VR3 | Gene names | Pcbp4 | |||
|
Domain Architecture |
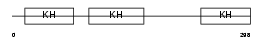
|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.010170 (rank : 62) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
CO039_MOUSE
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.024776 (rank : 47) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
DZIP1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.017666 (rank : 56) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BMD2, Q6ZQ10, Q9CRK5 | Gene names | Dzip1, Kiaa0996 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein DZIP1 (DAZ-interacting protein 1 homolog). | |||||
|
HSF1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.018294 (rank : 54) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P38532, O70462 | Gene names | Hsf1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 1 (HSF 1) (Heat shock transcription factor 1) (HSTF 1). | |||||
|
RKHD1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.096207 (rank : 25) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q86XN8 | Gene names | RKHD1, KIAA2031 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and KH domain-containing protein 1. | |||||
|
RPTN_HUMAN
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.017877 (rank : 55) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
CABL2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.030598 (rank : 41) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BTV7, Q5JWL0, Q9BYK0 | Gene names | CABLES2, C20orf150 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 and ABL1 enzyme substrate 2 (Interactor with CDK3 2) (Ik3-2). | |||||
|
EP15R_MOUSE
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.008664 (rank : 66) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60902, Q8CB60, Q8CB70, Q91WH8 | Gene names | Eps15l1, Eps15-rs, Eps15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R) (Epidermal growth factor receptor pathway substrate 15-related sequence) (Eps15-rs). | |||||
|
EVPL_MOUSE
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.006732 (rank : 67) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
K0423_HUMAN
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.020309 (rank : 50) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y4F4, Q68D66, Q6PG27 | Gene names | KIAA0423 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0423. | |||||
|
SON_HUMAN
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.019183 (rank : 53) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
UBP36_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.013152 (rank : 60) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9P275, Q8NDM8, Q9NVC8 | Gene names | USP36, KIAA1453 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 36 (EC 3.1.2.15) (Ubiquitin thioesterase 36) (Ubiquitin-specific-processing protease 36) (Deubiquitinating enzyme 36). | |||||
|
K1688_HUMAN
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.030955 (rank : 40) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.024925 (rank : 46) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
ZN385_HUMAN
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.019518 (rank : 51) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
BCAR1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.031442 (rank : 39) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
C1QR1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.004000 (rank : 70) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NPY3, O00274 | Gene names | CD93, C1QR1, MXRA4 | |||
|
Domain Architecture |
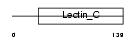
|
|||||
| Description | Complement component C1q receptor precursor (Complement component 1 q subcomponent receptor 1) (C1qR) (C1qRp) (C1qR(p)) (C1q/MBL/SPA receptor) (CD93 antigen) (CDw93). | |||||
|
IRX5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.009297 (rank : 64) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JKQ4, Q80WV4, Q9JLL5 | Gene names | Irx5, Irxb2 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-5 (Iroquois homeobox protein 5) (Homeodomain protein IRXB2). | |||||
|
PNPT1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.036353 (rank : 36) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TCS8, Q53SU0, Q68CN1, Q7Z7D1, Q8IWX1, Q96T05, Q9BRU3, Q9BVX0 | Gene names | PNPT1, PNPASE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyribonucleotide nucleotidyltransferase 1, mitochondrial precursor (EC 2.7.7.8) (PNPase 1) (Polynucleotide phosphorylase-like protein) (PNPase old-35) (3'-5' RNA exonuclease OLD35). | |||||
|
REST_MOUSE
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.002292 (rank : 71) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
UBP6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.009581 (rank : 63) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35125, Q15634, Q86WP6, Q8IWT4 | Gene names | USP6, TRE2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 6 (EC 3.1.2.15) (Ubiquitin thioesterase 6) (Ubiquitin-specific-processing protease 6) (Deubiquitinating enzyme 6) (Proto-oncogene TRE-2). | |||||
|
APC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.020618 (rank : 49) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.029211 (rank : 43) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
BCL9_MOUSE
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.027562 (rank : 45) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
DDX53_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.013912 (rank : 59) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q86TM3, Q6NVV4 | Gene names | DDX53, CAGE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX53 (EC 3.6.1.-) (DEAD box protein 53) (DEAD box protein CAGE) (Cancer-associated gene protein). | |||||
|
DC1L2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.009099 (rank : 65) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PDL0 | Gene names | Dync1li2, Dncli2, Dnclic2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic dynein 1 light intermediate chain 2 (Dynein light intermediate chain 2, cytosolic). | |||||
|
MYO15_HUMAN
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.012710 (rank : 61) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
PADI4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.005630 (rank : 68) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UM07, Q5VTZ8, Q70SX4 | Gene names | PADI4, PADI5, PDI5 | |||
|
Domain Architecture |

|
|||||
| Description | Protein-arginine deiminase type-4 (EC 3.5.3.15) (Protein-arginine deiminase type IV) (Peptidylarginine deiminase IV) (HL-60 PAD). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.019470 (rank : 52) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PNPT1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.030466 (rank : 42) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K1R3, Q3UEP9, Q810U7, Q812B3, Q8R2U3, Q9DC52 | Gene names | Pnpt1, Pnpase | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyribonucleotide nucleotidyltransferase 1, mitochondrial precursor (EC 2.7.7.8) (PNPase 1) (Polynucleotide phosphorylase-like protein) (PNPase old-35) (3'-5' RNA exonuclease OLD35). | |||||
|
SCAP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.016000 (rank : 58) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q12770, Q8N2E0, Q8WUA1 | Gene names | SCAP, KIAA0199 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein cleavage-activating protein (SREBP cleavage-activating protein) (SCAP). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.017107 (rank : 57) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
HNRPK_HUMAN
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.090444 (rank : 26) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P61978, Q07244, Q15671, Q60577, Q922Y7, Q96J62 | Gene names | HNRPK, HNRNPK | |||
|
Domain Architecture |
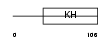
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K (hnRNP K) (Transformation up-regulated nuclear protein) (TUNP). | |||||
|
HNRPK_MOUSE
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.090444 (rank : 27) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P61979, Q07244, Q15671, Q60577, Q8BGQ8, Q922Y7, Q96J62 | Gene names | Hnrpk, Hnrnpk | |||
|
Domain Architecture |
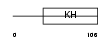
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K. | |||||
|
TDRKH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.067877 (rank : 32) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2W6, Q8N582, Q9NYV5 | Gene names | TDRKH, TDRD2 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
TDRKH_MOUSE
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.067794 (rank : 33) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q80VL1 | Gene names | Tdrkh | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
VIGLN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.050791 (rank : 35) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q00341, Q9UCY3 | Gene names | HDLBP, HBP, VGL | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
VIGLN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.050894 (rank : 34) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8VDJ3 | Gene names | Hdlbp | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
SAM68_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 1.08667e-154 (rank : 1) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q60749, Q3U8T3, Q60735, Q99M33 | Gene names | Khdrbs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
SAM68_HUMAN
|
||||||
| NC score | 0.989054 (rank : 2) | θ value | 1.68038e-147 (rank : 2) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q07666, Q6PJX7, Q8NB97, Q99760 | Gene names | KHDRBS1, SAM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
KHDR3_HUMAN
|
||||||
| NC score | 0.961898 (rank : 3) | θ value | 3.68522e-86 (rank : 3) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75525, Q6NUL8, Q9UPA8 | Gene names | KHDRBS3, SALP, SLM2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 3 (Sam68-like mammalian protein 2) (SLM-2) (Sam68-like phosphotyrosine protein) (RNA-binding protein T-Star). | |||||
|
KHDR3_MOUSE
|
||||||
| NC score | 0.958979 (rank : 4) | θ value | 7.68416e-84 (rank : 4) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9R226, O88624 | Gene names | Khdrbs3, Salp, Slm2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 3 (Sam68-like mammalian protein 2) (SLM-2) (RNA-binding protein Etoile). | |||||
|
QKI_HUMAN
|
||||||
| NC score | 0.821772 (rank : 5) | θ value | 6.38894e-30 (rank : 5) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96PU8, Q2I375, Q5MJQ1, Q969L9, Q96EJ3, Q96KA3, Q96PU6, Q96PU7, Q9P0X6, Q9P0X7, Q9P0X8, Q9P0X9, Q9P0Y0, Q9P0Y1 | Gene names | QKI, HKQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Quaking protein (Hqk). | |||||
|
QKI_MOUSE
|
||||||
| NC score | 0.821772 (rank : 6) | θ value | 6.38894e-30 (rank : 6) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QYS9, O88972, Q61110, Q78ZE4, Q78ZE5, Q8K4X9, Q8K4Y0, Q9CW34, Q9QUH4, Q9R2A8, Q9Z246 | Gene names | Qki, Qk, Qk1, Qka1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Quaking protein (qkI). | |||||
|
SF01_MOUSE
|
||||||
| NC score | 0.537540 (rank : 7) | θ value | 8.67504e-11 (rank : 8) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q64213, O08817, P70167, Q61454, Q921Z4 | Gene names | Sf1, Zfm1, Zfp162 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (mZFM) (Zinc finger gene in MEN1 locus) (Mammalian branch point- binding protein mBBP) (BBP) (CW17). | |||||
|
SF01_HUMAN
|
||||||
| NC score | 0.534409 (rank : 8) | θ value | 5.08577e-11 (rank : 7) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q15637, Q14818, Q14819, Q15913, Q8IY00, Q92744, Q92745, Q969H7, Q9BW01, Q9UEI0 | Gene names | SF1, ZFM1, ZNF162 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (Zinc finger gene in MEN1 locus) (Mammalian branch point-binding protein mBBP) (BBP). | |||||
|
IF2B2_MOUSE
|
||||||
| NC score | 0.206324 (rank : 9) | θ value | 0.00390308 (rank : 9) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5SF07, Q3TCU4, Q7TQF9 | Gene names | Igf2bp2, Imp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2). | |||||
|
FUBP3_HUMAN
|
||||||
| NC score | 0.185300 (rank : 10) | θ value | 0.00509761 (rank : 10) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96I24, Q92946, Q9BVB6 | Gene names | FUBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 3 (FUSE-binding protein 3). | |||||
|
IF2B2_HUMAN
|
||||||
| NC score | 0.180858 (rank : 11) | θ value | 0.21417 (rank : 19) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y6M1 | Gene names | IGF2BP2, IMP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2) (Hepatocellular carcinoma autoantigen p62). | |||||
|
NOVA2_HUMAN
|
||||||
| NC score | 0.166275 (rank : 12) | θ value | 0.125558 (rank : 14) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UNW9, O43267, Q9UEA1 | Gene names | NOVA2, ANOVA, NOVA3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein Nova-2 (Neuro-oncological ventral antigen 2) (Astrocytic NOVA1-like RNA-binding protein). | |||||
|
FUBP2_HUMAN
|
||||||
| NC score | 0.165884 (rank : 13) | θ value | 0.21417 (rank : 18) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q92945, O00301, Q9UNT5, Q9UQH5 | Gene names | KHSRP, FUBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 2 (FUSE-binding protein 2) (KH type-splicing regulatory protein) (KSRP) (p75). | |||||
|
FUBP1_MOUSE
|
||||||
| NC score | 0.159657 (rank : 14) | θ value | 0.163984 (rank : 17) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q91WJ8, Q8C0Y8 | Gene names | Fubp1, D3Ertd330e | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP). | |||||
|
FUBP1_HUMAN
|
||||||
| NC score | 0.159580 (rank : 15) | θ value | 0.163984 (rank : 16) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96AE4, Q12828 | Gene names | FUBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP) (DNA helicase V) (HDH V). | |||||
|
NOVA1_HUMAN
|
||||||
| NC score | 0.158718 (rank : 16) | θ value | 0.279714 (rank : 22) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P51513 | Gene names | NOVA1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein Nova-1 (Neuro-oncological ventral antigen 1) (Onconeural ventral antigen 1) (Paraneoplastic Ri antigen) (Ventral neuron-specific protein 1). | |||||
|
PCBP3_MOUSE
|
||||||
| NC score | 0.145919 (rank : 17) | θ value | 0.21417 (rank : 21) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P57722, Q8BSB0, Q8C544 | Gene names | Pcbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
PCBP3_HUMAN
|
||||||
| NC score | 0.145850 (rank : 18) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P57721, Q8N9K6, Q96EP6 | Gene names | PCBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
PCBP1_HUMAN
|
||||||
| NC score | 0.141414 (rank : 19) | θ value | 0.279714 (rank : 23) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15365, Q13157, Q14975 | Gene names | PCBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1) (Nucleic acid- binding protein SUB2.3). | |||||
|
PCBP1_MOUSE
|
||||||
| NC score | 0.141414 (rank : 20) | θ value | 0.279714 (rank : 24) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P60335 | Gene names | Pcbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1). | |||||
|
PCBP2_HUMAN
|
||||||
| NC score | 0.136485 (rank : 21) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q15366, Q6PKG5 | Gene names | PCBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (hnRNP-E2). | |||||
|
PCBP2_MOUSE
|
||||||
| NC score | 0.136039 (rank : 22) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q61990, Q61383, Q62042 | Gene names | Pcbp2, Cbp, Hnrnpx, Hnrpx | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (Putative heterogeneous nuclear ribonucleoprotein X) (hnRNP X) (CTBP) (CBP). | |||||
|
PCBP4_MOUSE
|
||||||
| NC score | 0.133533 (rank : 23) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P57724, Q91VR3 | Gene names | Pcbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
PCBP4_HUMAN
|
||||||
| NC score | 0.133084 (rank : 24) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P57723, Q96AH7 | Gene names | PCBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
RKHD1_HUMAN
|
||||||
| NC score | 0.096207 (rank : 25) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q86XN8 | Gene names | RKHD1, KIAA2031 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and KH domain-containing protein 1. | |||||
|
HNRPK_HUMAN
|
||||||
| NC score | 0.090444 (rank : 26) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P61978, Q07244, Q15671, Q60577, Q922Y7, Q96J62 | Gene names | HNRPK, HNRNPK | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K (hnRNP K) (Transformation up-regulated nuclear protein) (TUNP). | |||||
|
HNRPK_MOUSE
|
||||||
| NC score | 0.090444 (rank : 27) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P61979, Q07244, Q15671, Q60577, Q8BGQ8, Q922Y7, Q96J62 | Gene names | Hnrpk, Hnrnpk | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K. | |||||
|
BIM_HUMAN
|
||||||
| NC score | 0.085536 (rank : 28) | θ value | 0.0736092 (rank : 13) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43521, O43522 | Gene names | BCL2L11, BIM | |||
|
Domain Architecture |

|
|||||
| Description | Bcl-2-like protein 11 (Bcl2-interacting mediator of cell death). | |||||
|
BIM_MOUSE
|
||||||
| NC score | 0.083023 (rank : 29) | θ value | 0.163984 (rank : 15) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54918, O54919, O54920 | Gene names | Bcl2l11, Bim | |||
|
Domain Architecture |

|
|||||
| Description | Bcl-2-like protein 11 (Bcl2-interacting mediator of cell death). | |||||
|
K0174_HUMAN
|
||||||
| NC score | 0.083017 (rank : 30) | θ value | 0.0563607 (rank : 12) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P53990, Q9BQ81, Q9BWN2 | Gene names | KIAA0174 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA0174 (Putative MAPK-activating protein PM28). | |||||
|
K0174_MOUSE
|
||||||
| NC score | 0.070859 (rank : 31) | θ value | 0.813845 (rank : 29) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CX00, Q80U68 | Gene names | Kiaa0174 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0174. | |||||
|
TDRKH_HUMAN
|
||||||
| NC score | 0.067877 (rank : 32) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2W6, Q8N582, Q9NYV5 | Gene names | TDRKH, TDRD2 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
TDRKH_MOUSE
|
||||||
| NC score | 0.067794 (rank : 33) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q80VL1 | Gene names | Tdrkh | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
VIGLN_MOUSE
|
||||||
| NC score | 0.050894 (rank : 34) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8VDJ3 | Gene names | Hdlbp | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
VIGLN_HUMAN
|
||||||
| NC score | 0.050791 (rank : 35) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q00341, Q9UCY3 | Gene names | HDLBP, HBP, VGL | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
PNPT1_HUMAN
|
||||||
| NC score | 0.036353 (rank : 36) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TCS8, Q53SU0, Q68CN1, Q7Z7D1, Q8IWX1, Q96T05, Q9BRU3, Q9BVX0 | Gene names | PNPT1, PNPASE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyribonucleotide nucleotidyltransferase 1, mitochondrial precursor (EC 2.7.7.8) (PNPase 1) (Polynucleotide phosphorylase-like protein) (PNPase old-35) (3'-5' RNA exonuclease OLD35). | |||||
|
GLYG2_HUMAN
|
||||||
| NC score | 0.036204 (rank : 37) | θ value | 0.62314 (rank : 26) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15488, O15485, O15486, O15487, O15489, O15490 | Gene names | GYG2 | |||
|
Domain Architecture |

|
|||||
| Description | Glycogenin-2 (EC 2.4.1.186) (GN-2) (GN2). | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.033626 (rank : 38) | θ value | 0.00869519 (rank : 11) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
BCAR1_HUMAN
|
||||||
| NC score | 0.031442 (rank : 39) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
K1688_HUMAN
|
||||||
| NC score | 0.030955 (rank : 40) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
CABL2_HUMAN
|
||||||
| NC score | 0.030598 (rank : 41) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BTV7, Q5JWL0, Q9BYK0 | Gene names | CABLES2, C20orf150 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 and ABL1 enzyme substrate 2 (Interactor with CDK3 2) (Ik3-2). | |||||
|
PNPT1_MOUSE
|
||||||
| NC score | 0.030466 (rank : 42) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K1R3, Q3UEP9, Q810U7, Q812B3, Q8R2U3, Q9DC52 | Gene names | Pnpt1, Pnpase | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyribonucleotide nucleotidyltransferase 1, mitochondrial precursor (EC 2.7.7.8) (PNPase 1) (Polynucleotide phosphorylase-like protein) (PNPase old-35) (3'-5' RNA exonuclease OLD35). | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.029211 (rank : 43) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.027722 (rank : 44) | θ value | 0.365318 (rank : 25) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
BCL9_MOUSE
|
||||||
| NC score | 0.027562 (rank : 45) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.024925 (rank : 46) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
CO039_MOUSE
|
||||||
| NC score | 0.024776 (rank : 47) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
DCC_HUMAN
|
||||||
| NC score | 0.022842 (rank : 48) | θ value | 1.38821 (rank : 31) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 583 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P43146 | Gene names | DCC | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC) (Colorectal cancer suppressor). | |||||
|
APC_HUMAN
|
||||||
| NC score | 0.020618 (rank : 49) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
K0423_HUMAN
|
||||||
| NC score | 0.020309 (rank : 50) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y4F4, Q68D66, Q6PG27 | Gene names | KIAA0423 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0423. | |||||
|
ZN385_HUMAN
|
||||||
| NC score | 0.019518 (rank : 51) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.019470 (rank : 52) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.019183 (rank : 53) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
HSF1_MOUSE
|
||||||
| NC score | 0.018294 (rank : 54) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P38532, O70462 | Gene names | Hsf1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 1 (HSF 1) (Heat shock transcription factor 1) (HSTF 1). | |||||
|
RPTN_HUMAN
|
||||||
| NC score | 0.017877 (rank : 55) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
DZIP1_MOUSE
|
||||||
| NC score | 0.017666 (rank : 56) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BMD2, Q6ZQ10, Q9CRK5 | Gene names | Dzip1, Kiaa0996 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein DZIP1 (DAZ-interacting protein 1 homolog). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.017107 (rank : 57) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SCAP_HUMAN
|
||||||
| NC score | 0.016000 (rank : 58) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q12770, Q8N2E0, Q8WUA1 | Gene names | SCAP, KIAA0199 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein cleavage-activating protein (SREBP cleavage-activating protein) (SCAP). | |||||
|
DDX53_HUMAN
|
||||||
| NC score | 0.013912 (rank : 59) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q86TM3, Q6NVV4 | Gene names | DDX53, CAGE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX53 (EC 3.6.1.-) (DEAD box protein 53) (DEAD box protein CAGE) (Cancer-associated gene protein). | |||||
|
UBP36_HUMAN
|
||||||
| NC score | 0.013152 (rank : 60) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9P275, Q8NDM8, Q9NVC8 | Gene names | USP36, KIAA1453 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 36 (EC 3.1.2.15) (Ubiquitin thioesterase 36) (Ubiquitin-specific-processing protease 36) (Deubiquitinating enzyme 36). | |||||
|
MYO15_HUMAN
|
||||||
| NC score | 0.012710 (rank : 61) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.010170 (rank : 62) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
UBP6_HUMAN
|
||||||
| NC score | 0.009581 (rank : 63) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35125, Q15634, Q86WP6, Q8IWT4 | Gene names | USP6, TRE2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 6 (EC 3.1.2.15) (Ubiquitin thioesterase 6) (Ubiquitin-specific-processing protease 6) (Deubiquitinating enzyme 6) (Proto-oncogene TRE-2). | |||||
|
IRX5_MOUSE
|
||||||
| NC score | 0.009297 (rank : 64) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JKQ4, Q80WV4, Q9JLL5 | Gene names | Irx5, Irxb2 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-5 (Iroquois homeobox protein 5) (Homeodomain protein IRXB2). | |||||
|
DC1L2_MOUSE
|
||||||
| NC score | 0.009099 (rank : 65) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PDL0 | Gene names | Dync1li2, Dncli2, Dnclic2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic dynein 1 light intermediate chain 2 (Dynein light intermediate chain 2, cytosolic). | |||||
|
EP15R_MOUSE
|
||||||
| NC score | 0.008664 (rank : 66) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60902, Q8CB60, Q8CB70, Q91WH8 | Gene names | Eps15l1, Eps15-rs, Eps15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R) (Epidermal growth factor receptor pathway substrate 15-related sequence) (Eps15-rs). | |||||
|
EVPL_MOUSE
|
||||||
| NC score | 0.006732 (rank : 67) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
PADI4_HUMAN
|
||||||
| NC score | 0.005630 (rank : 68) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UM07, Q5VTZ8, Q70SX4 | Gene names | PADI4, PADI5, PDI5 | |||
|
Domain Architecture |

|
|||||
| Description | Protein-arginine deiminase type-4 (EC 3.5.3.15) (Protein-arginine deiminase type IV) (Peptidylarginine deiminase IV) (HL-60 PAD). | |||||
|
M3K11_HUMAN
|
||||||
| NC score | 0.004941 (rank : 69) | θ value | 1.06291 (rank : 30) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1136 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q16584, Q6P2G4 | Gene names | MAP3K11, MLK3, SPRK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 11 (EC 2.7.11.25) (Mixed lineage kinase 3) (Src-homology 3 domain-containing proline- rich kinase). | |||||
|
C1QR1_HUMAN
|
||||||
| NC score | 0.004000 (rank : 70) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NPY3, O00274 | Gene names | CD93, C1QR1, MXRA4 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C1q receptor precursor (Complement component 1 q subcomponent receptor 1) (C1qR) (C1qRp) (C1qR(p)) (C1q/MBL/SPA receptor) (CD93 antigen) (CDw93). | |||||
|
REST_MOUSE
|
||||||
| NC score | 0.002292 (rank : 71) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||