Please be patient as the page loads
|
PCBP4_MOUSE
|
||||||
| SwissProt Accessions | P57724, Q91VR3 | Gene names | Pcbp4 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PCBP4_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.998411 (rank : 2) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P57723, Q96AH7 | Gene names | PCBP4 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
PCBP4_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P57724, Q91VR3 | Gene names | Pcbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
PCBP3_HUMAN
|
||||||
| θ value | 6.0463e-105 (rank : 3) | NC score | 0.984056 (rank : 3) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P57721, Q8N9K6, Q96EP6 | Gene names | PCBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
PCBP3_MOUSE
|
||||||
| θ value | 2.2976e-104 (rank : 4) | NC score | 0.983946 (rank : 4) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P57722, Q8BSB0, Q8C544 | Gene names | Pcbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
PCBP2_MOUSE
|
||||||
| θ value | 2.72037e-97 (rank : 5) | NC score | 0.979568 (rank : 5) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q61990, Q61383, Q62042 | Gene names | Pcbp2, Cbp, Hnrnpx, Hnrpx | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (Putative heterogeneous nuclear ribonucleoprotein X) (hnRNP X) (CTBP) (CBP). | |||||
|
PCBP2_HUMAN
|
||||||
| θ value | 1.49272e-95 (rank : 6) | NC score | 0.979357 (rank : 6) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q15366, Q6PKG5 | Gene names | PCBP2 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (hnRNP-E2). | |||||
|
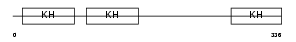
PCBP1_HUMAN
|
||||||
| θ value | 4.20666e-90 (rank : 7) | NC score | 0.979207 (rank : 7) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15365, Q13157, Q14975 | Gene names | PCBP1 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1) (Nucleic acid- binding protein SUB2.3). | |||||
|
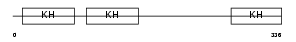
PCBP1_MOUSE
|
||||||
| θ value | 4.20666e-90 (rank : 8) | NC score | 0.979207 (rank : 8) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P60335 | Gene names | Pcbp1 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1). | |||||
|
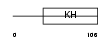
HNRPK_HUMAN
|
||||||
| θ value | 4.15078e-21 (rank : 9) | NC score | 0.824960 (rank : 9) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P61978, Q07244, Q15671, Q60577, Q922Y7, Q96J62 | Gene names | HNRPK, HNRNPK | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K (hnRNP K) (Transformation up-regulated nuclear protein) (TUNP). | |||||
|
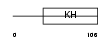
HNRPK_MOUSE
|
||||||
| θ value | 4.15078e-21 (rank : 10) | NC score | 0.824960 (rank : 10) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P61979, Q07244, Q15671, Q60577, Q8BGQ8, Q922Y7, Q96J62 | Gene names | Hnrpk, Hnrnpk | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K. | |||||
|
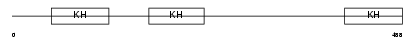
NOVA1_HUMAN
|
||||||
| θ value | 5.8054e-15 (rank : 11) | NC score | 0.769575 (rank : 11) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P51513 | Gene names | NOVA1 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding protein Nova-1 (Neuro-oncological ventral antigen 1) (Onconeural ventral antigen 1) (Paraneoplastic Ri antigen) (Ventral neuron-specific protein 1). | |||||
|
NOVA2_HUMAN
|
||||||
| θ value | 2.88119e-14 (rank : 12) | NC score | 0.769322 (rank : 12) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UNW9, O43267, Q9UEA1 | Gene names | NOVA2, ANOVA, NOVA3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein Nova-2 (Neuro-oncological ventral antigen 2) (Astrocytic NOVA1-like RNA-binding protein). | |||||
|
IF2B2_MOUSE
|
||||||
| θ value | 1.09485e-13 (rank : 13) | NC score | 0.607946 (rank : 18) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5SF07, Q3TCU4, Q7TQF9 | Gene names | Igf2bp2, Imp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2). | |||||
|
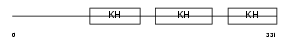
FUBP1_HUMAN
|
||||||
| θ value | 1.42992e-13 (rank : 14) | NC score | 0.617843 (rank : 16) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96AE4, Q12828 | Gene names | FUBP1 | |||
|
Domain Architecture |
|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP) (DNA helicase V) (HDH V). | |||||
|
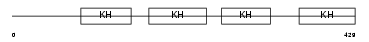
FUBP2_HUMAN
|
||||||
| θ value | 2.43908e-13 (rank : 15) | NC score | 0.633423 (rank : 14) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q92945, O00301, Q9UNT5, Q9UQH5 | Gene names | KHSRP, FUBP2 | |||
|
Domain Architecture |
|
|||||
| Description | Far upstream element-binding protein 2 (FUSE-binding protein 2) (KH type-splicing regulatory protein) (KSRP) (p75). | |||||
|
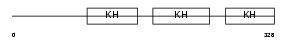
FUBP1_MOUSE
|
||||||
| θ value | 3.18553e-13 (rank : 16) | NC score | 0.618462 (rank : 15) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q91WJ8, Q8C0Y8 | Gene names | Fubp1, D3Ertd330e | |||
|
Domain Architecture |
|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP). | |||||
|
IF2B2_HUMAN
|
||||||
| θ value | 7.09661e-13 (rank : 17) | NC score | 0.608147 (rank : 17) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y6M1 | Gene names | IGF2BP2, IMP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2) (Hepatocellular carcinoma autoantigen p62). | |||||
|
FUBP3_HUMAN
|
||||||
| θ value | 1.58096e-12 (rank : 18) | NC score | 0.660505 (rank : 13) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96I24, Q92946, Q9BVB6 | Gene names | FUBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 3 (FUSE-binding protein 3). | |||||
|
TDRKH_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 19) | NC score | 0.448223 (rank : 19) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y2W6, Q8N582, Q9NYV5 | Gene names | TDRKH, TDRD2 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
TDRKH_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 20) | NC score | 0.445566 (rank : 20) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80VL1 | Gene names | Tdrkh | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
VIGLN_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 21) | NC score | 0.352093 (rank : 22) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q00341, Q9UCY3 | Gene names | HDLBP, HBP, VGL | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
VIGLN_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 22) | NC score | 0.352730 (rank : 21) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8VDJ3 | Gene names | Hdlbp | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
EFS_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 23) | NC score | 0.062296 (rank : 31) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43281, O43282 | Gene names | EFS | |||
|
Domain Architecture |
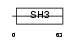
|
|||||
| Description | Embryonal Fyn-associated substrate (HEFS). | |||||
|
TF2AY_HUMAN
|
||||||
| θ value | 0.163984 (rank : 24) | NC score | 0.047302 (rank : 35) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UNN4 | Gene names | GTF2A1LF, ALF | |||
|
Domain Architecture |

|
|||||
| Description | TFIIA-alpha and beta-like factor (General transcription factor II A, 1-like factor). | |||||
|
TAF4_HUMAN
|
||||||
| θ value | 0.21417 (rank : 25) | NC score | 0.037818 (rank : 37) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
DDX53_HUMAN
|
||||||
| θ value | 0.279714 (rank : 26) | NC score | 0.058291 (rank : 32) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q86TM3, Q6NVV4 | Gene names | DDX53, CAGE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX53 (EC 3.6.1.-) (DEAD box protein 53) (DEAD box protein CAGE) (Cancer-associated gene protein). | |||||
|
AKAP1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 27) | NC score | 0.193701 (rank : 25) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O08715, O08714, P97488 | Gene names | Akap1, Akap | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein) (S-AKAP). | |||||
|
RKHD1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 28) | NC score | 0.235121 (rank : 23) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q86XN8 | Gene names | RKHD1, KIAA2031 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and KH domain-containing protein 1. | |||||
|
AKAP1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 29) | NC score | 0.205586 (rank : 24) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q92667, Q13320, Q9BW14 | Gene names | AKAP1, AKAP149 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (A-kinase anchor protein 149 kDa) (AKAP 149) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein 84) (S-AKAP84). | |||||
|
SAM68_HUMAN
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.135809 (rank : 26) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q07666, Q6PJX7, Q8NB97, Q99760 | Gene names | KHDRBS1, SAM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
SAM68_MOUSE
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.133533 (rank : 27) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q60749, Q3U8T3, Q60735, Q99M33 | Gene names | Khdrbs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
FOXK1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.005440 (rank : 46) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P42128, O35939 | Gene names | Foxk1, Mnf | |||
|
Domain Architecture |
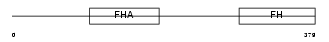
|
|||||
| Description | Forkhead box protein K1 (Myocyte nuclear factor) (MNF). | |||||
|
JHD2B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.011871 (rank : 42) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7LBC6, Q9BVH6, Q9BW93, Q9BZ52, Q9NYF4, Q9UPS0 | Gene names | JMJD1B, C5orf7, JHDM2B, KIAA1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B) (Nuclear protein 5qNCA). | |||||
|
FXR1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.088052 (rank : 28) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P51114, Q7Z450, Q8N6R8 | Gene names | FXR1 | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 1 (hFXR1P). | |||||
|
FXR1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.086110 (rank : 30) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61584, Q8VCU4, Q9R1E2, Q9R1E3, Q9R1E4, Q9R1E5, Q9WUA7, Q9WUA8, Q9WUA9 | Gene names | Fxr1, Fxr1h | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 1 (mFxr1p). | |||||
|
SC24C_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.009389 (rank : 44) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
UBE2T_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.012293 (rank : 40) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CQ37 | Gene names | Ube2t | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-conjugating enzyme E2 T (EC 6.3.2.19) (Ubiquitin-protein ligase T) (Ubiquitin carrier protein T). | |||||
|
DDX43_HUMAN
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.042897 (rank : 36) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NXZ2, Q6NXR1 | Gene names | DDX43, HAGE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX43 (EC 3.6.1.-) (DEAD box protein 43) (DEAD box protein HAGE) (Helical antigen). | |||||
|
NFAC2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.007270 (rank : 45) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60591, Q60984, Q60985 | Gene names | Nfatc2, Nfat1, Nfatp | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 2 (T cell transcription factor NFAT1) (NFAT pre-existing subunit) (NF-ATp). | |||||
|
UBE2T_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.011155 (rank : 43) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NPD8 | Gene names | UBE2T | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-conjugating enzyme E2 T (EC 6.3.2.19) (Ubiquitin-protein ligase T) (Ubiquitin carrier protein T). | |||||
|
HXD12_MOUSE
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.001096 (rank : 47) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23812 | Gene names | Hoxd12, Hox-4.7, Hoxd-12 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D12 (Hox-4.7) (Hox-5.6). | |||||
|
SON_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.012093 (rank : 41) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SPEG_HUMAN
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | -0.000037 (rank : 48) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | -0.000153 (rank : 49) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
TOM1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.012504 (rank : 39) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88746, Q3V4C6, Q9D120 | Gene names | Tom1 | |||
|
Domain Architecture |
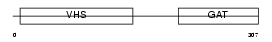
|
|||||
| Description | Target of Myb protein 1. | |||||
|
MISS_MOUSE
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.015874 (rank : 38) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D7G9, Q7M749 | Gene names | Mapk1ip1, Miss | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAPK-interacting and spindle-stabilizing protein (Mitogen-activated protein kinase 1-interacting protein 1). | |||||
|
FMR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.054328 (rank : 33) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q06787, Q16578, Q99054 | Gene names | FMR1 | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation 1 protein (Protein FMR-1) (FMRP). | |||||
|
FMR1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.052887 (rank : 34) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P35922 | Gene names | Fmr1, Fmr-1 | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation protein 1 homolog (Protein FMR-1) (FMRP) (mFmr1p). | |||||
|
FXR2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.087888 (rank : 29) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51116, Q8WUM2 | Gene names | FXR2 | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
PCBP4_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P57724, Q91VR3 | Gene names | Pcbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
PCBP4_HUMAN
|
||||||
| NC score | 0.998411 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P57723, Q96AH7 | Gene names | PCBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
PCBP3_HUMAN
|
||||||
| NC score | 0.984056 (rank : 3) | θ value | 6.0463e-105 (rank : 3) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P57721, Q8N9K6, Q96EP6 | Gene names | PCBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
PCBP3_MOUSE
|
||||||
| NC score | 0.983946 (rank : 4) | θ value | 2.2976e-104 (rank : 4) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P57722, Q8BSB0, Q8C544 | Gene names | Pcbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
PCBP2_MOUSE
|
||||||
| NC score | 0.979568 (rank : 5) | θ value | 2.72037e-97 (rank : 5) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q61990, Q61383, Q62042 | Gene names | Pcbp2, Cbp, Hnrnpx, Hnrpx | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (Putative heterogeneous nuclear ribonucleoprotein X) (hnRNP X) (CTBP) (CBP). | |||||
|
PCBP2_HUMAN
|
||||||
| NC score | 0.979357 (rank : 6) | θ value | 1.49272e-95 (rank : 6) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q15366, Q6PKG5 | Gene names | PCBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (hnRNP-E2). | |||||
|
PCBP1_HUMAN
|
||||||
| NC score | 0.979207 (rank : 7) | θ value | 4.20666e-90 (rank : 7) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15365, Q13157, Q14975 | Gene names | PCBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1) (Nucleic acid- binding protein SUB2.3). | |||||
|
PCBP1_MOUSE
|
||||||
| NC score | 0.979207 (rank : 8) | θ value | 4.20666e-90 (rank : 8) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P60335 | Gene names | Pcbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1). | |||||
|
HNRPK_HUMAN
|
||||||
| NC score | 0.824960 (rank : 9) | θ value | 4.15078e-21 (rank : 9) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P61978, Q07244, Q15671, Q60577, Q922Y7, Q96J62 | Gene names | HNRPK, HNRNPK | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K (hnRNP K) (Transformation up-regulated nuclear protein) (TUNP). | |||||
|
HNRPK_MOUSE
|
||||||
| NC score | 0.824960 (rank : 10) | θ value | 4.15078e-21 (rank : 10) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P61979, Q07244, Q15671, Q60577, Q8BGQ8, Q922Y7, Q96J62 | Gene names | Hnrpk, Hnrnpk | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K. | |||||
|
NOVA1_HUMAN
|
||||||
| NC score | 0.769575 (rank : 11) | θ value | 5.8054e-15 (rank : 11) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P51513 | Gene names | NOVA1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein Nova-1 (Neuro-oncological ventral antigen 1) (Onconeural ventral antigen 1) (Paraneoplastic Ri antigen) (Ventral neuron-specific protein 1). | |||||
|
NOVA2_HUMAN
|
||||||
| NC score | 0.769322 (rank : 12) | θ value | 2.88119e-14 (rank : 12) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UNW9, O43267, Q9UEA1 | Gene names | NOVA2, ANOVA, NOVA3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein Nova-2 (Neuro-oncological ventral antigen 2) (Astrocytic NOVA1-like RNA-binding protein). | |||||
|
FUBP3_HUMAN
|
||||||
| NC score | 0.660505 (rank : 13) | θ value | 1.58096e-12 (rank : 18) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96I24, Q92946, Q9BVB6 | Gene names | FUBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 3 (FUSE-binding protein 3). | |||||
|
FUBP2_HUMAN
|
||||||
| NC score | 0.633423 (rank : 14) | θ value | 2.43908e-13 (rank : 15) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q92945, O00301, Q9UNT5, Q9UQH5 | Gene names | KHSRP, FUBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 2 (FUSE-binding protein 2) (KH type-splicing regulatory protein) (KSRP) (p75). | |||||
|
FUBP1_MOUSE
|
||||||
| NC score | 0.618462 (rank : 15) | θ value | 3.18553e-13 (rank : 16) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q91WJ8, Q8C0Y8 | Gene names | Fubp1, D3Ertd330e | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP). | |||||
|
FUBP1_HUMAN
|
||||||
| NC score | 0.617843 (rank : 16) | θ value | 1.42992e-13 (rank : 14) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96AE4, Q12828 | Gene names | FUBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP) (DNA helicase V) (HDH V). | |||||
|
IF2B2_HUMAN
|
||||||
| NC score | 0.608147 (rank : 17) | θ value | 7.09661e-13 (rank : 17) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y6M1 | Gene names | IGF2BP2, IMP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2) (Hepatocellular carcinoma autoantigen p62). | |||||
|
IF2B2_MOUSE
|
||||||
| NC score | 0.607946 (rank : 18) | θ value | 1.09485e-13 (rank : 13) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5SF07, Q3TCU4, Q7TQF9 | Gene names | Igf2bp2, Imp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2). | |||||
|
TDRKH_HUMAN
|
||||||
| NC score | 0.448223 (rank : 19) | θ value | 8.40245e-06 (rank : 19) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y2W6, Q8N582, Q9NYV5 | Gene names | TDRKH, TDRD2 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
TDRKH_MOUSE
|
||||||
| NC score | 0.445566 (rank : 20) | θ value | 1.87187e-05 (rank : 20) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80VL1 | Gene names | Tdrkh | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
VIGLN_MOUSE
|
||||||
| NC score | 0.352730 (rank : 21) | θ value | 0.00390308 (rank : 22) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8VDJ3 | Gene names | Hdlbp | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
VIGLN_HUMAN
|
||||||
| NC score | 0.352093 (rank : 22) | θ value | 0.00390308 (rank : 21) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q00341, Q9UCY3 | Gene names | HDLBP, HBP, VGL | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
RKHD1_HUMAN
|
||||||
| NC score | 0.235121 (rank : 23) | θ value | 0.47712 (rank : 28) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q86XN8 | Gene names | RKHD1, KIAA2031 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and KH domain-containing protein 1. | |||||
|
AKAP1_HUMAN
|
||||||
| NC score | 0.205586 (rank : 24) | θ value | 0.813845 (rank : 29) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q92667, Q13320, Q9BW14 | Gene names | AKAP1, AKAP149 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (A-kinase anchor protein 149 kDa) (AKAP 149) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein 84) (S-AKAP84). | |||||
|
AKAP1_MOUSE
|
||||||
| NC score | 0.193701 (rank : 25) | θ value | 0.365318 (rank : 27) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O08715, O08714, P97488 | Gene names | Akap1, Akap | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein) (S-AKAP). | |||||
|
SAM68_HUMAN
|
||||||
| NC score | 0.135809 (rank : 26) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q07666, Q6PJX7, Q8NB97, Q99760 | Gene names | KHDRBS1, SAM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
SAM68_MOUSE
|
||||||
| NC score | 0.133533 (rank : 27) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q60749, Q3U8T3, Q60735, Q99M33 | Gene names | Khdrbs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
FXR1_HUMAN
|
||||||
| NC score | 0.088052 (rank : 28) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P51114, Q7Z450, Q8N6R8 | Gene names | FXR1 | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 1 (hFXR1P). | |||||
|
FXR2_HUMAN
|
||||||
| NC score | 0.087888 (rank : 29) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51116, Q8WUM2 | Gene names | FXR2 | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
FXR1_MOUSE
|
||||||
| NC score | 0.086110 (rank : 30) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61584, Q8VCU4, Q9R1E2, Q9R1E3, Q9R1E4, Q9R1E5, Q9WUA7, Q9WUA8, Q9WUA9 | Gene names | Fxr1, Fxr1h | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 1 (mFxr1p). | |||||
|
EFS_HUMAN
|
||||||
| NC score | 0.062296 (rank : 31) | θ value | 0.00665767 (rank : 23) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43281, O43282 | Gene names | EFS | |||
|
Domain Architecture |

|
|||||
| Description | Embryonal Fyn-associated substrate (HEFS). | |||||
|
DDX53_HUMAN
|
||||||
| NC score | 0.058291 (rank : 32) | θ value | 0.279714 (rank : 26) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q86TM3, Q6NVV4 | Gene names | DDX53, CAGE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX53 (EC 3.6.1.-) (DEAD box protein 53) (DEAD box protein CAGE) (Cancer-associated gene protein). | |||||
|
FMR1_HUMAN
|
||||||
| NC score | 0.054328 (rank : 33) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q06787, Q16578, Q99054 | Gene names | FMR1 | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation 1 protein (Protein FMR-1) (FMRP). | |||||
|
FMR1_MOUSE
|
||||||
| NC score | 0.052887 (rank : 34) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P35922 | Gene names | Fmr1, Fmr-1 | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation protein 1 homolog (Protein FMR-1) (FMRP) (mFmr1p). | |||||
|
TF2AY_HUMAN
|
||||||
| NC score | 0.047302 (rank : 35) | θ value | 0.163984 (rank : 24) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UNN4 | Gene names | GTF2A1LF, ALF | |||
|
Domain Architecture |

|
|||||
| Description | TFIIA-alpha and beta-like factor (General transcription factor II A, 1-like factor). | |||||
|
DDX43_HUMAN
|
||||||
| NC score | 0.042897 (rank : 36) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NXZ2, Q6NXR1 | Gene names | DDX43, HAGE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX43 (EC 3.6.1.-) (DEAD box protein 43) (DEAD box protein HAGE) (Helical antigen). | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.037818 (rank : 37) | θ value | 0.21417 (rank : 25) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
MISS_MOUSE
|
||||||
| NC score | 0.015874 (rank : 38) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D7G9, Q7M749 | Gene names | Mapk1ip1, Miss | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAPK-interacting and spindle-stabilizing protein (Mitogen-activated protein kinase 1-interacting protein 1). | |||||
|
TOM1_MOUSE
|
||||||
| NC score | 0.012504 (rank : 39) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88746, Q3V4C6, Q9D120 | Gene names | Tom1 | |||
|
Domain Architecture |

|
|||||
| Description | Target of Myb protein 1. | |||||
|
UBE2T_MOUSE
|
||||||
| NC score | 0.012293 (rank : 40) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CQ37 | Gene names | Ube2t | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-conjugating enzyme E2 T (EC 6.3.2.19) (Ubiquitin-protein ligase T) (Ubiquitin carrier protein T). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.012093 (rank : 41) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
JHD2B_HUMAN
|
||||||
| NC score | 0.011871 (rank : 42) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7LBC6, Q9BVH6, Q9BW93, Q9BZ52, Q9NYF4, Q9UPS0 | Gene names | JMJD1B, C5orf7, JHDM2B, KIAA1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B) (Nuclear protein 5qNCA). | |||||
|
UBE2T_HUMAN
|
||||||
| NC score | 0.011155 (rank : 43) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NPD8 | Gene names | UBE2T | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-conjugating enzyme E2 T (EC 6.3.2.19) (Ubiquitin-protein ligase T) (Ubiquitin carrier protein T). | |||||
|
SC24C_HUMAN
|
||||||
| NC score | 0.009389 (rank : 44) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
NFAC2_MOUSE
|
||||||
| NC score | 0.007270 (rank : 45) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60591, Q60984, Q60985 | Gene names | Nfatc2, Nfat1, Nfatp | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 2 (T cell transcription factor NFAT1) (NFAT pre-existing subunit) (NF-ATp). | |||||
|
FOXK1_MOUSE
|
||||||
| NC score | 0.005440 (rank : 46) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P42128, O35939 | Gene names | Foxk1, Mnf | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein K1 (Myocyte nuclear factor) (MNF). | |||||
|
HXD12_MOUSE
|
||||||
| NC score | 0.001096 (rank : 47) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23812 | Gene names | Hoxd12, Hox-4.7, Hoxd-12 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D12 (Hox-4.7) (Hox-5.6). | |||||
|
SPEG_HUMAN
|
||||||
| NC score | -0.000037 (rank : 48) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | -0.000153 (rank : 49) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||