Please be patient as the page loads
|
SAM68_HUMAN
|
||||||
| SwissProt Accessions | Q07666, Q6PJX7, Q8NB97, Q99760 | Gene names | KHDRBS1, SAM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SAM68_MOUSE
|
||||||
| θ value | 1.68038e-147 (rank : 1) | NC score | 0.989054 (rank : 2) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q60749, Q3U8T3, Q60735, Q99M33 | Gene names | Khdrbs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
SAM68_HUMAN
|
||||||
| θ value | 3.7435e-147 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q07666, Q6PJX7, Q8NB97, Q99760 | Gene names | KHDRBS1, SAM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
KHDR3_HUMAN
|
||||||
| θ value | 1.40038e-85 (rank : 3) | NC score | 0.963004 (rank : 3) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75525, Q6NUL8, Q9UPA8 | Gene names | KHDRBS3, SALP, SLM2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 3 (Sam68-like mammalian protein 2) (SLM-2) (Sam68-like phosphotyrosine protein) (RNA-binding protein T-Star). | |||||
|
KHDR3_MOUSE
|
||||||
| θ value | 4.98072e-83 (rank : 4) | NC score | 0.960190 (rank : 4) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R226, O88624 | Gene names | Khdrbs3, Salp, Slm2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 3 (Sam68-like mammalian protein 2) (SLM-2) (RNA-binding protein Etoile). | |||||
|
QKI_HUMAN
|
||||||
| θ value | 6.38894e-30 (rank : 5) | NC score | 0.823474 (rank : 5) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96PU8, Q2I375, Q5MJQ1, Q969L9, Q96EJ3, Q96KA3, Q96PU6, Q96PU7, Q9P0X6, Q9P0X7, Q9P0X8, Q9P0X9, Q9P0Y0, Q9P0Y1 | Gene names | QKI, HKQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Quaking protein (Hqk). | |||||
|
QKI_MOUSE
|
||||||
| θ value | 6.38894e-30 (rank : 6) | NC score | 0.823474 (rank : 6) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QYS9, O88972, Q61110, Q78ZE4, Q78ZE5, Q8K4X9, Q8K4Y0, Q9CW34, Q9QUH4, Q9R2A8, Q9Z246 | Gene names | Qki, Qk, Qk1, Qka1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Quaking protein (qkI). | |||||
|
SF01_HUMAN
|
||||||
| θ value | 5.08577e-11 (rank : 7) | NC score | 0.539670 (rank : 8) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15637, Q14818, Q14819, Q15913, Q8IY00, Q92744, Q92745, Q969H7, Q9BW01, Q9UEI0 | Gene names | SF1, ZFM1, ZNF162 | |||
|
Domain Architecture |
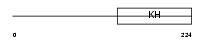
|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (Zinc finger gene in MEN1 locus) (Mammalian branch point-binding protein mBBP) (BBP). | |||||
|
SF01_MOUSE
|
||||||
| θ value | 8.67504e-11 (rank : 8) | NC score | 0.543049 (rank : 7) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q64213, O08817, P70167, Q61454, Q921Z4 | Gene names | Sf1, Zfm1, Zfp162 | |||
|
Domain Architecture |
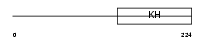
|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (mZFM) (Zinc finger gene in MEN1 locus) (Mammalian branch point- binding protein mBBP) (BBP) (CW17). | |||||
|
FUBP3_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 9) | NC score | 0.195068 (rank : 10) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96I24, Q92946, Q9BVB6 | Gene names | FUBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 3 (FUSE-binding protein 3). | |||||
|
FUBP2_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 10) | NC score | 0.179537 (rank : 11) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q92945, O00301, Q9UNT5, Q9UQH5 | Gene names | KHSRP, FUBP2 | |||
|
Domain Architecture |
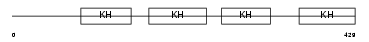
|
|||||
| Description | Far upstream element-binding protein 2 (FUSE-binding protein 2) (KH type-splicing regulatory protein) (KSRP) (p75). | |||||
|
IF2B2_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 11) | NC score | 0.204090 (rank : 9) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5SF07, Q3TCU4, Q7TQF9 | Gene names | Igf2bp2, Imp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2). | |||||
|
FUBP1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 12) | NC score | 0.169553 (rank : 14) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96AE4, Q12828 | Gene names | FUBP1 | |||
|
Domain Architecture |
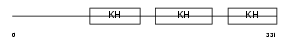
|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP) (DNA helicase V) (HDH V). | |||||
|
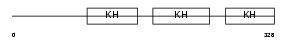
FUBP1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 13) | NC score | 0.169808 (rank : 13) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q91WJ8, Q8C0Y8 | Gene names | Fubp1, D3Ertd330e | |||
|
Domain Architecture |
|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP). | |||||
|
K1688_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 14) | NC score | 0.047325 (rank : 32) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
NOVA2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 15) | NC score | 0.168973 (rank : 15) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UNW9, O43267, Q9UEA1 | Gene names | NOVA2, ANOVA, NOVA3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein Nova-2 (Neuro-oncological ventral antigen 2) (Astrocytic NOVA1-like RNA-binding protein). | |||||
|
DCC_MOUSE
|
||||||
| θ value | 0.163984 (rank : 16) | NC score | 0.027339 (rank : 40) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
IF2B2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.179144 (rank : 12) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y6M1 | Gene names | IGF2BP2, IMP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2) (Hepatocellular carcinoma autoantigen p62). | |||||
|
PCBP3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 18) | NC score | 0.147908 (rank : 18) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P57721, Q8N9K6, Q96EP6 | Gene names | PCBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
PCBP3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 19) | NC score | 0.147980 (rank : 17) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P57722, Q8BSB0, Q8C544 | Gene names | Pcbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
RFIP1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.036894 (rank : 35) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D620, Q3UBC2, Q8BN24 | Gene names | Rab11fip1, Rcp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
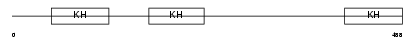
NOVA1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 21) | NC score | 0.161286 (rank : 16) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P51513 | Gene names | NOVA1 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding protein Nova-1 (Neuro-oncological ventral antigen 1) (Onconeural ventral antigen 1) (Paraneoplastic Ri antigen) (Ventral neuron-specific protein 1). | |||||
|
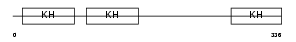
PCBP1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 22) | NC score | 0.143753 (rank : 19) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15365, Q13157, Q14975 | Gene names | PCBP1 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1) (Nucleic acid- binding protein SUB2.3). | |||||
|
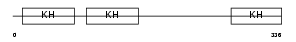
PCBP1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 23) | NC score | 0.143753 (rank : 20) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P60335 | Gene names | Pcbp1 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1). | |||||
|
BCAR1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 24) | NC score | 0.042873 (rank : 33) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
MYO15_HUMAN
|
||||||
| θ value | 0.62314 (rank : 25) | NC score | 0.017849 (rank : 56) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
NPAS4_MOUSE
|
||||||
| θ value | 0.62314 (rank : 26) | NC score | 0.038902 (rank : 34) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
PCBP2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.138829 (rank : 21) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q15366, Q6PKG5 | Gene names | PCBP2 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (hnRNP-E2). | |||||
|
PCBP2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.138046 (rank : 22) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61990, Q61383, Q62042 | Gene names | Pcbp2, Cbp, Hnrnpx, Hnrpx | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (Putative heterogeneous nuclear ribonucleoprotein X) (hnRNP X) (CTBP) (CBP). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 1.06291 (rank : 29) | NC score | 0.009830 (rank : 66) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
CABL2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 30) | NC score | 0.034598 (rank : 38) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BTV7, Q5JWL0, Q9BYK0 | Gene names | CABLES2, C20orf150 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 and ABL1 enzyme substrate 2 (Interactor with CDK3 2) (Ik3-2). | |||||
|
M3K11_HUMAN
|
||||||
| θ value | 1.38821 (rank : 31) | NC score | 0.005384 (rank : 71) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1136 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q16584, Q6P2G4 | Gene names | MAP3K11, MLK3, SPRK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 11 (EC 2.7.11.25) (Mixed lineage kinase 3) (Src-homology 3 domain-containing proline- rich kinase). | |||||
|
NPAS4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 32) | NC score | 0.035948 (rank : 37) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
PCBP4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.135409 (rank : 24) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P57723, Q96AH7 | Gene names | PCBP4 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
PCBP4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.135809 (rank : 23) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P57724, Q91VR3 | Gene names | Pcbp4 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
SPI1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 35) | NC score | 0.019208 (rank : 52) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P17947 | Gene names | SPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein). | |||||
|
CEL_HUMAN
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.023511 (rank : 44) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.024838 (rank : 42) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
RKHD1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.096537 (rank : 25) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q86XN8 | Gene names | RKHD1, KIAA2031 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and KH domain-containing protein 1. | |||||
|
ZN692_HUMAN
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.000850 (rank : 73) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BU19, Q5SRA5, Q5SRA6, Q9HBC9, Q9NW93, Q9NWY6, Q9UF97 | Gene names | ZNF692 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 692. | |||||
|
AKAP2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.017901 (rank : 55) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
CB025_MOUSE
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.024607 (rank : 43) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99LS1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C2orf25 homolog, mitochondrial precursor. | |||||
|
SEM6C_MOUSE
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.007899 (rank : 69) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
APC_HUMAN
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.021805 (rank : 46) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.020823 (rank : 49) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
RPTN_HUMAN
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.020924 (rank : 48) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
SPHK2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.017460 (rank : 58) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JIA7, Q91VA9, Q9DBH6 | Gene names | Sphk2 | |||
|
Domain Architecture |

|
|||||
| Description | Sphingosine kinase 2 (EC 2.7.1.-) (SK 2) (SPK 2). | |||||
|
ZCH14_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.021401 (rank : 47) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WYQ9, O60324, Q9UFP0 | Gene names | ZCCHC14, KIAA0579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 14 (BDG-29). | |||||
|
DCC_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.016777 (rank : 59) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 583 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P43146 | Gene names | DCC | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC) (Colorectal cancer suppressor). | |||||
|
EP15R_MOUSE
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.007073 (rank : 70) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60902, Q8CB60, Q8CB70, Q91WH8 | Gene names | Eps15l1, Eps15-rs, Eps15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R) (Epidermal growth factor receptor pathway substrate 15-related sequence) (Eps15-rs). | |||||
|
HNRPG_HUMAN
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.014286 (rank : 61) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P38159, Q5JQ67, Q969R3 | Gene names | RBMX, HNRPG, RBMXP1 | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein G (hnRNP G) (RNA-binding motif protein, X chromosome) (Glycoprotein p43). | |||||
|
PNPT1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.036291 (rank : 36) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TCS8, Q53SU0, Q68CN1, Q7Z7D1, Q8IWX1, Q96T05, Q9BRU3, Q9BVX0 | Gene names | PNPT1, PNPASE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyribonucleotide nucleotidyltransferase 1, mitochondrial precursor (EC 2.7.7.8) (PNPase 1) (Polynucleotide phosphorylase-like protein) (PNPase old-35) (3'-5' RNA exonuclease OLD35). | |||||
|
RPTN_MOUSE
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.009349 (rank : 67) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P97347 | Gene names | Rptn | |||
|
Domain Architecture |

|
|||||
| Description | Repetin. | |||||
|
ZN385_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.019371 (rank : 51) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
AXN2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.011022 (rank : 65) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
CIC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.017586 (rank : 57) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
DDX53_HUMAN
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.014313 (rank : 60) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q86TM3, Q6NVV4 | Gene names | DDX53, CAGE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX53 (EC 3.6.1.-) (DEAD box protein 53) (DEAD box protein CAGE) (Cancer-associated gene protein). | |||||
|
MTR1L_HUMAN
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | -0.000659 (rank : 74) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1180 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13585 | Gene names | GPR50 | |||
|
Domain Architecture |

|
|||||
| Description | Melatonin-related receptor (G protein-coupled receptor 50) (H9). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.025840 (rank : 41) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
SCAP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.017948 (rank : 54) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12770, Q8N2E0, Q8WUA1 | Gene names | SCAP, KIAA0199 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein cleavage-activating protein (SREBP cleavage-activating protein) (SCAP). | |||||
|
AMBN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.013303 (rank : 62) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O55189 | Gene names | Ambn | |||
|
Domain Architecture |
|
|||||
| Description | Ameloblastin precursor. | |||||
|
ATX2L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.018482 (rank : 53) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8WWM7, O95135, Q6NVJ8, Q6PJW6, Q8IU61, Q8IU95, Q8WWM3, Q8WWM4, Q8WWM5, Q8WWM6, Q99703 | Gene names | ATXN2L, A2D, A2LG, A2LP, A2RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein (Ataxin-2 domain protein) (Ataxin-2-related protein). | |||||
|
BCAR1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.022592 (rank : 45) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61140, Q60869 | Gene names | Bcar1, Cas, Crkas | |||
|
Domain Architecture |
|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
CTDP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.012709 (rank : 63) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
DPYL4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.004820 (rank : 72) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O14531, O00240, Q5T0Q7 | Gene names | DPYSL4, CRMP3, ULIP4 | |||
|
Domain Architecture |
|
|||||
| Description | Dihydropyrimidinase-related protein 4 (DRP-4) (Collapsin response mediator protein 3) (CRMP-3) (UNC33-like phosphoprotein 4) (ULIP4 protein). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.019475 (rank : 50) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
PNPT1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.030603 (rank : 39) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K1R3, Q3UEP9, Q810U7, Q812B3, Q8R2U3, Q9DC52 | Gene names | Pnpt1, Pnpase | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyribonucleotide nucleotidyltransferase 1, mitochondrial precursor (EC 2.7.7.8) (PNPase 1) (Polynucleotide phosphorylase-like protein) (PNPase old-35) (3'-5' RNA exonuclease OLD35). | |||||
|
TARA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.009090 (rank : 68) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TAU_HUMAN
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.012416 (rank : 64) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
HNRPK_HUMAN
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.093073 (rank : 26) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P61978, Q07244, Q15671, Q60577, Q922Y7, Q96J62 | Gene names | HNRPK, HNRNPK | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K (hnRNP K) (Transformation up-regulated nuclear protein) (TUNP). | |||||
|
HNRPK_MOUSE
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.093073 (rank : 27) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P61979, Q07244, Q15671, Q60577, Q8BGQ8, Q922Y7, Q96J62 | Gene names | Hnrpk, Hnrnpk | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K. | |||||
|
TDRKH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.070419 (rank : 28) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2W6, Q8N582, Q9NYV5 | Gene names | TDRKH, TDRD2 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
TDRKH_MOUSE
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.070407 (rank : 29) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q80VL1 | Gene names | Tdrkh | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
VIGLN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.053103 (rank : 31) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q00341, Q9UCY3 | Gene names | HDLBP, HBP, VGL | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
VIGLN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.053233 (rank : 30) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8VDJ3 | Gene names | Hdlbp | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
SAM68_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 3.7435e-147 (rank : 2) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q07666, Q6PJX7, Q8NB97, Q99760 | Gene names | KHDRBS1, SAM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
SAM68_MOUSE
|
||||||
| NC score | 0.989054 (rank : 2) | θ value | 1.68038e-147 (rank : 1) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q60749, Q3U8T3, Q60735, Q99M33 | Gene names | Khdrbs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
KHDR3_HUMAN
|
||||||
| NC score | 0.963004 (rank : 3) | θ value | 1.40038e-85 (rank : 3) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75525, Q6NUL8, Q9UPA8 | Gene names | KHDRBS3, SALP, SLM2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 3 (Sam68-like mammalian protein 2) (SLM-2) (Sam68-like phosphotyrosine protein) (RNA-binding protein T-Star). | |||||
|
KHDR3_MOUSE
|
||||||
| NC score | 0.960190 (rank : 4) | θ value | 4.98072e-83 (rank : 4) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R226, O88624 | Gene names | Khdrbs3, Salp, Slm2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 3 (Sam68-like mammalian protein 2) (SLM-2) (RNA-binding protein Etoile). | |||||
|
QKI_HUMAN
|
||||||
| NC score | 0.823474 (rank : 5) | θ value | 6.38894e-30 (rank : 5) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96PU8, Q2I375, Q5MJQ1, Q969L9, Q96EJ3, Q96KA3, Q96PU6, Q96PU7, Q9P0X6, Q9P0X7, Q9P0X8, Q9P0X9, Q9P0Y0, Q9P0Y1 | Gene names | QKI, HKQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Quaking protein (Hqk). | |||||
|
QKI_MOUSE
|
||||||
| NC score | 0.823474 (rank : 6) | θ value | 6.38894e-30 (rank : 6) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QYS9, O88972, Q61110, Q78ZE4, Q78ZE5, Q8K4X9, Q8K4Y0, Q9CW34, Q9QUH4, Q9R2A8, Q9Z246 | Gene names | Qki, Qk, Qk1, Qka1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Quaking protein (qkI). | |||||
|
SF01_MOUSE
|
||||||
| NC score | 0.543049 (rank : 7) | θ value | 8.67504e-11 (rank : 8) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q64213, O08817, P70167, Q61454, Q921Z4 | Gene names | Sf1, Zfm1, Zfp162 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (mZFM) (Zinc finger gene in MEN1 locus) (Mammalian branch point- binding protein mBBP) (BBP) (CW17). | |||||
|
SF01_HUMAN
|
||||||
| NC score | 0.539670 (rank : 8) | θ value | 5.08577e-11 (rank : 7) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15637, Q14818, Q14819, Q15913, Q8IY00, Q92744, Q92745, Q969H7, Q9BW01, Q9UEI0 | Gene names | SF1, ZFM1, ZNF162 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (Zinc finger gene in MEN1 locus) (Mammalian branch point-binding protein mBBP) (BBP). | |||||
|
IF2B2_MOUSE
|
||||||
| NC score | 0.204090 (rank : 9) | θ value | 0.0330416 (rank : 11) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5SF07, Q3TCU4, Q7TQF9 | Gene names | Igf2bp2, Imp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2). | |||||
|
FUBP3_HUMAN
|
||||||
| NC score | 0.195068 (rank : 10) | θ value | 0.00390308 (rank : 9) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96I24, Q92946, Q9BVB6 | Gene names | FUBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 3 (FUSE-binding protein 3). | |||||
|
FUBP2_HUMAN
|
||||||
| NC score | 0.179537 (rank : 11) | θ value | 0.0193708 (rank : 10) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q92945, O00301, Q9UNT5, Q9UQH5 | Gene names | KHSRP, FUBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 2 (FUSE-binding protein 2) (KH type-splicing regulatory protein) (KSRP) (p75). | |||||
|
IF2B2_HUMAN
|
||||||
| NC score | 0.179144 (rank : 12) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y6M1 | Gene names | IGF2BP2, IMP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2) (Hepatocellular carcinoma autoantigen p62). | |||||
|
FUBP1_MOUSE
|
||||||
| NC score | 0.169808 (rank : 13) | θ value | 0.0736092 (rank : 13) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q91WJ8, Q8C0Y8 | Gene names | Fubp1, D3Ertd330e | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP). | |||||
|
FUBP1_HUMAN
|
||||||
| NC score | 0.169553 (rank : 14) | θ value | 0.0736092 (rank : 12) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96AE4, Q12828 | Gene names | FUBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP) (DNA helicase V) (HDH V). | |||||
|
NOVA2_HUMAN
|
||||||
| NC score | 0.168973 (rank : 15) | θ value | 0.125558 (rank : 15) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UNW9, O43267, Q9UEA1 | Gene names | NOVA2, ANOVA, NOVA3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein Nova-2 (Neuro-oncological ventral antigen 2) (Astrocytic NOVA1-like RNA-binding protein). | |||||
|
NOVA1_HUMAN
|
||||||
| NC score | 0.161286 (rank : 16) | θ value | 0.279714 (rank : 21) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P51513 | Gene names | NOVA1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein Nova-1 (Neuro-oncological ventral antigen 1) (Onconeural ventral antigen 1) (Paraneoplastic Ri antigen) (Ventral neuron-specific protein 1). | |||||
|
PCBP3_MOUSE
|
||||||
| NC score | 0.147980 (rank : 17) | θ value | 0.21417 (rank : 19) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P57722, Q8BSB0, Q8C544 | Gene names | Pcbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
PCBP3_HUMAN
|
||||||
| NC score | 0.147908 (rank : 18) | θ value | 0.21417 (rank : 18) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P57721, Q8N9K6, Q96EP6 | Gene names | PCBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 3 (Alpha-CP3). | |||||
|
PCBP1_HUMAN
|
||||||
| NC score | 0.143753 (rank : 19) | θ value | 0.279714 (rank : 22) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15365, Q13157, Q14975 | Gene names | PCBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1) (Nucleic acid- binding protein SUB2.3). | |||||
|
PCBP1_MOUSE
|
||||||
| NC score | 0.143753 (rank : 20) | θ value | 0.279714 (rank : 23) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P60335 | Gene names | Pcbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 1 (Alpha-CP1) (hnRNP-E1). | |||||
|
PCBP2_HUMAN
|
||||||
| NC score | 0.138829 (rank : 21) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q15366, Q6PKG5 | Gene names | PCBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (hnRNP-E2). | |||||
|
PCBP2_MOUSE
|
||||||
| NC score | 0.138046 (rank : 22) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61990, Q61383, Q62042 | Gene names | Pcbp2, Cbp, Hnrnpx, Hnrpx | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 2 (Alpha-CP2) (Putative heterogeneous nuclear ribonucleoprotein X) (hnRNP X) (CTBP) (CBP). | |||||
|
PCBP4_MOUSE
|
||||||
| NC score | 0.135809 (rank : 23) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P57724, Q91VR3 | Gene names | Pcbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
PCBP4_HUMAN
|
||||||
| NC score | 0.135409 (rank : 24) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P57723, Q96AH7 | Gene names | PCBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
RKHD1_HUMAN
|
||||||
| NC score | 0.096537 (rank : 25) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q86XN8 | Gene names | RKHD1, KIAA2031 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and KH domain-containing protein 1. | |||||
|
HNRPK_HUMAN
|
||||||
| NC score | 0.093073 (rank : 26) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P61978, Q07244, Q15671, Q60577, Q922Y7, Q96J62 | Gene names | HNRPK, HNRNPK | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K (hnRNP K) (Transformation up-regulated nuclear protein) (TUNP). | |||||
|
HNRPK_MOUSE
|
||||||
| NC score | 0.093073 (rank : 27) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P61979, Q07244, Q15671, Q60577, Q8BGQ8, Q922Y7, Q96J62 | Gene names | Hnrpk, Hnrnpk | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein K. | |||||
|
TDRKH_HUMAN
|
||||||
| NC score | 0.070419 (rank : 28) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2W6, Q8N582, Q9NYV5 | Gene names | TDRKH, TDRD2 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
TDRKH_MOUSE
|
||||||
| NC score | 0.070407 (rank : 29) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q80VL1 | Gene names | Tdrkh | |||
|
Domain Architecture |

|
|||||
| Description | Tudor and KH domain-containing protein. | |||||
|
VIGLN_MOUSE
|
||||||
| NC score | 0.053233 (rank : 30) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8VDJ3 | Gene names | Hdlbp | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
VIGLN_HUMAN
|
||||||
| NC score | 0.053103 (rank : 31) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q00341, Q9UCY3 | Gene names | HDLBP, HBP, VGL | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
K1688_HUMAN
|
||||||
| NC score | 0.047325 (rank : 32) | θ value | 0.0961366 (rank : 14) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
BCAR1_HUMAN
|
||||||
| NC score | 0.042873 (rank : 33) | θ value | 0.62314 (rank : 24) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
NPAS4_MOUSE
|
||||||
| NC score | 0.038902 (rank : 34) | θ value | 0.62314 (rank : 26) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
RFIP1_MOUSE
|
||||||
| NC score | 0.036894 (rank : 35) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D620, Q3UBC2, Q8BN24 | Gene names | Rab11fip1, Rcp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
PNPT1_HUMAN
|
||||||
| NC score | 0.036291 (rank : 36) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TCS8, Q53SU0, Q68CN1, Q7Z7D1, Q8IWX1, Q96T05, Q9BRU3, Q9BVX0 | Gene names | PNPT1, PNPASE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyribonucleotide nucleotidyltransferase 1, mitochondrial precursor (EC 2.7.7.8) (PNPase 1) (Polynucleotide phosphorylase-like protein) (PNPase old-35) (3'-5' RNA exonuclease OLD35). | |||||
|
NPAS4_HUMAN
|
||||||
| NC score | 0.035948 (rank : 37) | θ value | 1.38821 (rank : 32) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
CABL2_HUMAN
|
||||||
| NC score | 0.034598 (rank : 38) | θ value | 1.38821 (rank : 30) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BTV7, Q5JWL0, Q9BYK0 | Gene names | CABLES2, C20orf150 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 and ABL1 enzyme substrate 2 (Interactor with CDK3 2) (Ik3-2). | |||||
|
PNPT1_MOUSE
|
||||||
| NC score | 0.030603 (rank : 39) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K1R3, Q3UEP9, Q810U7, Q812B3, Q8R2U3, Q9DC52 | Gene names | Pnpt1, Pnpase | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyribonucleotide nucleotidyltransferase 1, mitochondrial precursor (EC 2.7.7.8) (PNPase 1) (Polynucleotide phosphorylase-like protein) (PNPase old-35) (3'-5' RNA exonuclease OLD35). | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.027339 (rank : 40) | θ value | 0.163984 (rank : 16) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.025840 (rank : 41) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.024838 (rank : 42) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
CB025_MOUSE
|
||||||
| NC score | 0.024607 (rank : 43) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99LS1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C2orf25 homolog, mitochondrial precursor. | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.023511 (rank : 44) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
BCAR1_MOUSE
|
||||||
| NC score | 0.022592 (rank : 45) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61140, Q60869 | Gene names | Bcar1, Cas, Crkas | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
APC_HUMAN
|
||||||
| NC score | 0.021805 (rank : 46) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
ZCH14_HUMAN
|
||||||
| NC score | 0.021401 (rank : 47) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WYQ9, O60324, Q9UFP0 | Gene names | ZCCHC14, KIAA0579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 14 (BDG-29). | |||||
|
RPTN_HUMAN
|
||||||
| NC score | 0.020924 (rank : 48) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.020823 (rank : 49) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.019475 (rank : 50) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
ZN385_HUMAN
|
||||||
| NC score | 0.019371 (rank : 51) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
SPI1_HUMAN
|
||||||
| NC score | 0.019208 (rank : 52) | θ value | 1.81305 (rank : 35) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P17947 | Gene names | SPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein). | |||||
|
ATX2L_HUMAN
|
||||||
| NC score | 0.018482 (rank : 53) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8WWM7, O95135, Q6NVJ8, Q6PJW6, Q8IU61, Q8IU95, Q8WWM3, Q8WWM4, Q8WWM5, Q8WWM6, Q99703 | Gene names | ATXN2L, A2D, A2LG, A2LP, A2RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein (Ataxin-2 domain protein) (Ataxin-2-related protein). | |||||
|
SCAP_HUMAN
|
||||||
| NC score | 0.017948 (rank : 54) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12770, Q8N2E0, Q8WUA1 | Gene names | SCAP, KIAA0199 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein cleavage-activating protein (SREBP cleavage-activating protein) (SCAP). | |||||
|
AKAP2_HUMAN
|
||||||
| NC score | 0.017901 (rank : 55) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
MYO15_HUMAN
|
||||||
| NC score | 0.017849 (rank : 56) | θ value | 0.62314 (rank : 25) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.017586 (rank : 57) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
SPHK2_MOUSE
|
||||||
| NC score | 0.017460 (rank : 58) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JIA7, Q91VA9, Q9DBH6 | Gene names | Sphk2 | |||
|
Domain Architecture |

|
|||||
| Description | Sphingosine kinase 2 (EC 2.7.1.-) (SK 2) (SPK 2). | |||||
|
DCC_HUMAN
|
||||||
| NC score | 0.016777 (rank : 59) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 583 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P43146 | Gene names | DCC | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC) (Colorectal cancer suppressor). | |||||
|
DDX53_HUMAN
|
||||||
| NC score | 0.014313 (rank : 60) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q86TM3, Q6NVV4 | Gene names | DDX53, CAGE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX53 (EC 3.6.1.-) (DEAD box protein 53) (DEAD box protein CAGE) (Cancer-associated gene protein). | |||||
|
HNRPG_HUMAN
|
||||||
| NC score | 0.014286 (rank : 61) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P38159, Q5JQ67, Q969R3 | Gene names | RBMX, HNRPG, RBMXP1 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein G (hnRNP G) (RNA-binding motif protein, X chromosome) (Glycoprotein p43). | |||||
|
AMBN_MOUSE
|
||||||
| NC score | 0.013303 (rank : 62) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O55189 | Gene names | Ambn | |||
|
Domain Architecture |

|
|||||
| Description | Ameloblastin precursor. | |||||
|
CTDP1_HUMAN
|
||||||
| NC score | 0.012709 (rank : 63) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
TAU_HUMAN
|
||||||
| NC score | 0.012416 (rank : 64) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
AXN2_MOUSE
|
||||||
| NC score | 0.011022 (rank : 65) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.009830 (rank : 66) | θ value | 1.06291 (rank : 29) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
RPTN_MOUSE
|
||||||
| NC score | 0.009349 (rank : 67) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P97347 | Gene names | Rptn | |||
|
Domain Architecture |

|
|||||
| Description | Repetin. | |||||
|
TARA_MOUSE
|
||||||
| NC score | 0.009090 (rank : 68) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
SEM6C_MOUSE
|
||||||
| NC score | 0.007899 (rank : 69) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
EP15R_MOUSE
|
||||||
| NC score | 0.007073 (rank : 70) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60902, Q8CB60, Q8CB70, Q91WH8 | Gene names | Eps15l1, Eps15-rs, Eps15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R) (Epidermal growth factor receptor pathway substrate 15-related sequence) (Eps15-rs). | |||||
|
M3K11_HUMAN
|
||||||
| NC score | 0.005384 (rank : 71) | θ value | 1.38821 (rank : 31) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1136 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q16584, Q6P2G4 | Gene names | MAP3K11, MLK3, SPRK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 11 (EC 2.7.11.25) (Mixed lineage kinase 3) (Src-homology 3 domain-containing proline- rich kinase). | |||||
|
DPYL4_HUMAN
|
||||||
| NC score | 0.004820 (rank : 72) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O14531, O00240, Q5T0Q7 | Gene names | DPYSL4, CRMP3, ULIP4 | |||
|
Domain Architecture |

|
|||||
| Description | Dihydropyrimidinase-related protein 4 (DRP-4) (Collapsin response mediator protein 3) (CRMP-3) (UNC33-like phosphoprotein 4) (ULIP4 protein). | |||||
|
ZN692_HUMAN
|
||||||
| NC score | 0.000850 (rank : 73) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BU19, Q5SRA5, Q5SRA6, Q9HBC9, Q9NW93, Q9NWY6, Q9UF97 | Gene names | ZNF692 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 692. | |||||
|
MTR1L_HUMAN
|
||||||
| NC score | -0.000659 (rank : 74) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1180 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13585 | Gene names | GPR50 | |||
|
Domain Architecture |

|
|||||
| Description | Melatonin-related receptor (G protein-coupled receptor 50) (H9). | |||||