Please be patient as the page loads
|
SACS_HUMAN
|
||||||
| SwissProt Accessions | Q9NZJ4, O94835 | Gene names | SACS, KIAA0730 | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SACS_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 128 | |
| SwissProt Accessions | Q9NZJ4, O94835 | Gene names | SACS, KIAA0730 | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
SACS_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.994673 (rank : 2) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9JLC8 | Gene names | Sacs | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
DNJB6_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 3) | NC score | 0.190155 (rank : 5) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O75190, O95806, Q9UIK6 | Gene names | DNAJB6, HSJ2, MSJ1 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 6 (Heat shock protein J2) (HSJ-2) (MSJ-1) (HHDJ1) (MRJ). | |||||
|
DNJB7_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 4) | NC score | 0.190880 (rank : 4) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYI8, Q9DA41 | Gene names | Dnajb7 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 7 (mDJ5). | |||||
|
DNJB8_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 5) | NC score | 0.185891 (rank : 7) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NHS0 | Gene names | DNAJB8 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 8. | |||||
|
DNJB8_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 6) | NC score | 0.184165 (rank : 9) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYI7 | Gene names | Dnajb8 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 8 (mDJ6). | |||||
|
DNJB2_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 7) | NC score | 0.188298 (rank : 6) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P25686, Q8IUK1, Q8IUK2, Q96F52 | Gene names | DNAJB2, HSJ1, HSPF3 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 2 (Heat shock 40 kDa protein 3) (DnaJ protein homolog 1) (HSJ-1). | |||||
|
DNJB6_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 8) | NC score | 0.183414 (rank : 11) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O54946, Q99LA5, Q9QYI9 | Gene names | Dnajb6, Hsj2, Mrj | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 6 (Heat shock protein J2) (HSJ-2) (MRJ) (mDj4). | |||||
|
DNJB7_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 9) | NC score | 0.184470 (rank : 8) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q7Z6W7, Q5H904, Q8WYJ7 | Gene names | DNAJB7, HSC3 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 7. | |||||
|
DNJB3_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 10) | NC score | 0.183673 (rank : 10) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35723, Q9DAN3, Q9DAN4 | Gene names | Dnajb3, Hsj3, Msj1 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 3 (DnaJ protein homolog 3) (Heat shock J3 protein) (HSJ-3) (MSJ-1). | |||||
|
DNJBA_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 11) | NC score | 0.194012 (rank : 3) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYI5 | Gene names | Dnajb10 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 10 (mDJ8). | |||||
|
DNJC5_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 12) | NC score | 0.167951 (rank : 12) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H3Z4, Q9H3Z5, Q9H7H2 | Gene names | DNAJC5, CSP | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 5 (Cysteine string protein) (CSP). | |||||
|
DNJC5_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 13) | NC score | 0.167923 (rank : 13) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P60904, P54101 | Gene names | Dnajc5, Csp | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 5 (Cysteine string protein) (CSP). | |||||
|
DNJ5B_HUMAN
|
||||||
| θ value | 0.163984 (rank : 14) | NC score | 0.156775 (rank : 14) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UF47, Q969Y8 | Gene names | DNAJC5B | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 5B (Beta cysteine string protein) (Beta-CSP). | |||||
|
BPAEA_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.029703 (rank : 95) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
DNJB5_MOUSE
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.124307 (rank : 21) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O89114 | Gene names | Dnajb5, Hsc40 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 5 (Heat shock protein Hsp40-3) (Heat shock protein cognate 40) (Hsc40). | |||||
|
DNJCD_HUMAN
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.138281 (rank : 17) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O75165, Q6PI82, Q6UJ77, Q6ZSW1, Q6ZUT5, Q86XG3, Q96DC1, Q9BWK9 | Gene names | DNAJC13, KIAA0678, RME8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 13 (Required for receptor-mediated endocytosis 8). | |||||
|
UTY_MOUSE
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.038096 (rank : 86) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P79457, O97979 | Gene names | Uty | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed Y chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein ON the Y chromosome) (Male- specific histocompatibility antigen H-YDB). | |||||
|
DNJB5_HUMAN
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.122995 (rank : 23) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O75953, Q8TDR7 | Gene names | DNAJB5, HSC40 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 5 (Heat shock protein Hsp40-3) (Heat shock protein cognate 40) (Hsc40) (Hsp40-2). | |||||
|
CENPQ_HUMAN
|
||||||
| θ value | 0.47712 (rank : 20) | NC score | 0.048813 (rank : 76) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7L2Z9, Q6IN61, Q9NVS5 | Gene names | CENPQ, C6orf139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein Q (CENP-Q). | |||||
|
DNJ5B_MOUSE
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.149078 (rank : 15) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9CQ94 | Gene names | Dnajc5b | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 5B (Beta cysteine string protein) (Beta-CSP). | |||||
|
DNJA4_MOUSE
|
||||||
| θ value | 0.47712 (rank : 22) | NC score | 0.126939 (rank : 18) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9JMC3, Q543S9, Q9CTD6, Q9CUD4 | Gene names | Dnaja4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 4 (MmDjA4). | |||||
|
SAPS1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 23) | NC score | 0.021544 (rank : 109) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
CF152_HUMAN
|
||||||
| θ value | 0.62314 (rank : 24) | NC score | 0.040938 (rank : 82) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q86VQ0, Q9BWX7 | Gene names | C6orf152 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152. | |||||
|
DNJA4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 25) | NC score | 0.125837 (rank : 19) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WW22, Q8N7P2 | Gene names | DNAJA4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 4. | |||||
|
HEAT1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 26) | NC score | 0.070930 (rank : 67) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H583, Q5T3Q8, Q9NW23 | Gene names | HEATR1, BAP28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HEAT repeat-containing protein 1 (Protein BAP28). | |||||
|
MAST4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.004850 (rank : 161) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
MOR2B_MOUSE
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.038150 (rank : 85) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C5W4, Q8CG24 | Gene names | Morc2b, Tce6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2B (TCE6). | |||||
|
CEP35_HUMAN
|
||||||
| θ value | 0.813845 (rank : 29) | NC score | 0.042182 (rank : 81) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
DNJA1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 30) | NC score | 0.120312 (rank : 25) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P31689 | Gene names | DNAJA1, DNAJ2, HDJ2, HSJ2, HSPF4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 1 (Heat shock 40 kDa protein 4) (DnaJ protein homolog 2) (HSJ-2) (HSDJ). | |||||
|
DNJA1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 31) | NC score | 0.120170 (rank : 26) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63037, P54102 | Gene names | Dnaja1, Dnaj2, Hsj2, Hspf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily A member 1 (Heat shock 40 kDa protein 4) (DnaJ protein homolog 2) (HSJ-2). | |||||
|
DNJBC_HUMAN
|
||||||
| θ value | 1.06291 (rank : 32) | NC score | 0.124397 (rank : 20) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NXW2, Q9H6H0 | Gene names | DNAJB12 | |||
|
Domain Architecture |
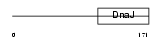
|
|||||
| Description | DnaJ homolog subfamily B member 12. | |||||
|
PERT_MOUSE
|
||||||
| θ value | 1.06291 (rank : 33) | NC score | 0.025603 (rank : 99) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35419, Q8C8B1 | Gene names | Tpo | |||
|
Domain Architecture |
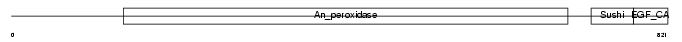
|
|||||
| Description | Thyroid peroxidase precursor (EC 1.11.1.8) (TPO). | |||||
|
RECQ4_MOUSE
|
||||||
| θ value | 1.06291 (rank : 34) | NC score | 0.020161 (rank : 112) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q75NR7, Q76MT1, Q99PV9 | Gene names | Recql4, Recq4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4). | |||||
|
TTBK2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 35) | NC score | 0.017331 (rank : 120) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3UVR3, Q3TSR6, Q3UFW0, Q571D1, Q8BKA4, Q924U8 | Gene names | Ttbk2, Kiaa0847, Ttbk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 2. | |||||
|
OTOAN_MOUSE
|
||||||
| θ value | 1.38821 (rank : 36) | NC score | 0.043482 (rank : 79) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K561 | Gene names | Otoa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Otoancorin precursor. | |||||
|
CF152_MOUSE
|
||||||
| θ value | 1.81305 (rank : 37) | NC score | 0.030111 (rank : 94) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80ST9, Q9CYM9, Q9D5J9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152 homolog. | |||||
|
DEN2A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 38) | NC score | 0.018848 (rank : 114) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULE3, Q1RMD5, Q86XY0 | Gene names | DENND2A, KIAA1277 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2A. | |||||
|
DJC17_HUMAN
|
||||||
| θ value | 1.81305 (rank : 39) | NC score | 0.120606 (rank : 24) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9NVM6 | Gene names | DNAJC17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 17. | |||||
|
DJC17_MOUSE
|
||||||
| θ value | 1.81305 (rank : 40) | NC score | 0.123580 (rank : 22) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q91WT4, Q8C5R1 | Gene names | Dnajc17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 17. | |||||
|
DNJA2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.115553 (rank : 28) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O60884, O14711 | Gene names | DNAJA2, CPR3, HIRIP4 | |||
|
Domain Architecture |
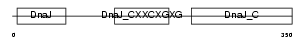
|
|||||
| Description | DnaJ homolog subfamily A member 2 (HIRA-interacting protein 4) (Cell cycle progression restoration gene 3 protein) (Dnj3) (NY-REN-14 antigen). | |||||
|
DNJA2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 42) | NC score | 0.115682 (rank : 27) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYJ0 | Gene names | Dnaja2 | |||
|
Domain Architecture |
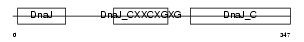
|
|||||
| Description | DnaJ homolog subfamily A member 2 (mDj3). | |||||
|
ENPL_HUMAN
|
||||||
| θ value | 1.81305 (rank : 43) | NC score | 0.039389 (rank : 83) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P14625, Q96A97 | Gene names | HSP90B1, TRA1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (gp96 homolog) (Tumor rejection antigen 1). | |||||
|
AKAP9_HUMAN
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.015330 (rank : 126) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
COMDA_MOUSE
|
||||||
| θ value | 2.36792 (rank : 45) | NC score | 0.071102 (rank : 66) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8JZY2 | Gene names | Commd10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | COMM domain-containing protein 10 (Down-regulated in W/WV mouse stomach 2) (mDRWMS2). | |||||
|
CUL4A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 46) | NC score | 0.022478 (rank : 105) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13619, O75834, Q589T6, Q5TC62, Q6UP08, Q9UP17 | Gene names | CUL4A | |||
|
Domain Architecture |
|
|||||
| Description | Cullin-4A (CUL-4A). | |||||
|
DNJ5G_HUMAN
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.149057 (rank : 16) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8N7S2, Q8IYQ4, Q96RJ8 | Gene names | DNAJC5G | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 5G (Gamma cysteine string protein) (Gamma-CSP). | |||||
|
ENPL_MOUSE
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.037542 (rank : 88) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P08113, P11427 | Gene names | Hsp90b1, Tra-1, Tra1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (ERP99) (Polymorphic tumor rejection antigen 1) (Tumor rejection antigen gp96). | |||||
|
FOXM1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.023448 (rank : 101) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q08050, O43258, O43259, O43260, Q4ZGG7, Q9BRL2 | Gene names | FOXM1, FKHL16, HFH11, MPP2, WIN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead-related protein FKHL16) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/fork-head homolog 11) (HFH-11) (Winged helix factor from INS-1 cells) (M-phase phosphoprotein 2) (MPM-2 reactive phosphoprotein 2) (Transcription factor Trident). | |||||
|
K0586_HUMAN
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.043719 (rank : 78) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BVV6, O60328 | Gene names | KIAA0586 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0586. | |||||
|
KIF5C_MOUSE
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.015325 (rank : 127) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P28738, Q9Z2F8 | Gene names | Kif5c, Nkhc2 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
POLK_MOUSE
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.034844 (rank : 89) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QUG2, Q7TPY7 | Gene names | Polk, Dinb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase kappa (EC 2.7.7.7) (DINB protein) (DINP). | |||||
|
ABCAD_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.014780 (rank : 129) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86UQ4, Q6ZTT7, Q86WI2, Q8N248 | Gene names | ABCA13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-binding cassette sub-family A member 13. | |||||
|
ANGL2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.008883 (rank : 152) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9R045 | Gene names | Angptl2, Arp2 | |||
|
Domain Architecture |

|
|||||
| Description | Angiopoietin-related protein 2 precursor (Angiopoietin-like 2). | |||||
|
F125B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.027846 (rank : 96) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H7P6, Q8N6S7 | Gene names | FAM125B, C9orf28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM125B. | |||||
|
GOGA1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.022555 (rank : 104) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 994 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9CW79, Q80YB0 | Gene names | Golga1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
PARN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.023337 (rank : 102) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95453 | Gene names | PARN, DAN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A)-specific ribonuclease PARN (EC 3.1.13.4) (Polyadenylate- specific ribonuclease) (Deadenylating nuclease) (Deadenylation nuclease). | |||||
|
PDZD6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.048096 (rank : 77) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULD6, Q4W5I8, Q86V55 | Gene names | PDZD6, KIAA1284, PDZK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 6. | |||||
|
SPAG5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.022177 (rank : 106) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96R06, O95213, Q9BWE8, Q9NT17, Q9UFE6 | Gene names | SPAG5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 5 (Astrin) (Mitotic spindle-associated protein p126) (MAP126) (Deepest). | |||||
|
SYMPK_MOUSE
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.042772 (rank : 80) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80X82 | Gene names | Sympk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Symplekin. | |||||
|
BCL7C_MOUSE
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.020602 (rank : 110) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O08664, Q99LS2 | Gene names | Bcl7c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell CLL/lymphoma 7 protein family member C (B-cell chronic lymphocytic leukemia/lymphoma 7C protein). | |||||
|
CCNB3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.014505 (rank : 131) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q810T2, Q810T3, Q8VDC8 | Gene names | Ccnb3, Cycb3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
CDC27_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.019035 (rank : 113) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30260, Q16349, Q96F35 | Gene names | CDC27 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 27 homolog (CDC27Hs) (H-NUC). | |||||
|
COMDA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.055063 (rank : 73) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6G5, Q9P077 | Gene names | COMMD10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | COMM domain-containing protein 10. | |||||
|
COVA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.022928 (rank : 103) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q16206, Q5VTJ1, Q5VTJ2, Q8WUX0, Q9NTP6, Q9UH82 | Gene names | COVA1 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor-associated hydroquinone oxidase (tNOX) (Cytosolic ovarian carcinoma antigen 1) (APK1 antigen) [Includes: Hydroquinone [NADH] oxidase (EC 1.-.-.-); Protein disulfide-thiol oxidoreductase (EC 1.-.-.-)]. | |||||
|
COVA1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.023457 (rank : 100) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8R0Z2, Q8BY59, Q8R372 | Gene names | Cova1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor-associated hydroquinone oxidase (tNOX) (Cytosolic ovarian carcinoma antigen 1) (APK1 antigen) [Includes: Hydroquinone [NADH] oxidase (EC 1.-.-.-); Protein disulfide-thiol oxidoreductase (EC 1.-.-.-)]. | |||||
|
DNJBC_MOUSE
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.114103 (rank : 29) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYI4 | Gene names | Dnajb12 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 12 (mDJ10). | |||||
|
GEMI2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.039366 (rank : 84) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CQQ4, Q9DAD7 | Gene names | Sip1, Gemin2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Survival of motor neuron protein-interacting protein 1 (SMN- interacting protein 1) (Component of gems 2) (Gemin-2). | |||||
|
HEM3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.033412 (rank : 91) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P22907 | Gene names | Hmbs, Uros1 | |||
|
Domain Architecture |
|
|||||
| Description | Porphobilinogen deaminase (EC 2.5.1.61) (Hydroxymethylbilane synthase) (HMBS) (Pre-uroporphyrinogen synthase) (PBG-D). | |||||
|
KIF1A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.010584 (rank : 144) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q12756 | Gene names | KIF1A, ATSV | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF1A (Axonal transporter of synaptic vesicles). | |||||
|
KIF1A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.010605 (rank : 143) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P33173, Q61770 | Gene names | Kif1a, Atsv, Kif1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF1A (Axonal transporter of synaptic vesicles). | |||||
|
KTN1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.017135 (rank : 121) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q86UP2, Q13999, Q14707, Q15387, Q86W57 | Gene names | KTN1, CG1, KIAA0004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin (Kinesin receptor) (CG-1 antigen). | |||||
|
MPP4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.011816 (rank : 141) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96JB8, Q6ZNH6, Q96Q43, Q96Q44 | Gene names | MPP4, ALS2CR5, DLG6 | |||
|
Domain Architecture |

|
|||||
| Description | MAGUK p55 subfamily member 4 (Discs large homolog 6) (Amyotrophic lateral sclerosis 2 chromosomal region candidate gene 5 protein). | |||||
|
MYH13_HUMAN
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.008379 (rank : 154) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
PYC_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.026516 (rank : 97) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P11498, Q16705 | Gene names | PC | |||
|
Domain Architecture |

|
|||||
| Description | Pyruvate carboxylase, mitochondrial precursor (EC 6.4.1.1) (Pyruvic carboxylase) (PCB). | |||||
|
PYC_MOUSE
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.026449 (rank : 98) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q05920 | Gene names | Pc, Pcx | |||
|
Domain Architecture |

|
|||||
| Description | Pyruvate carboxylase, mitochondrial precursor (EC 6.4.1.1) (Pyruvic carboxylase) (PCB). | |||||
|
SCN1A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.006306 (rank : 158) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35498, Q16172, Q96LA3, Q9C008 | Gene names | SCN1A, NAC1, SCN1 | |||
|
Domain Architecture |
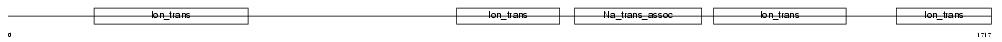
|
|||||
| Description | Sodium channel protein type 1 subunit alpha (Sodium channel protein type I subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.1) (Sodium channel protein, brain I subunit alpha). | |||||
|
SRXN1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.054278 (rank : 74) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D975, Q3TWN6, Q91VN5 | Gene names | Srxn1, Npn3, Srx | |||
|
Domain Architecture |
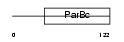
|
|||||
| Description | Sulfiredoxin-1 (EC 1.8.98.2) (Neoplastic progression protein 3). | |||||
|
TAF5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 79) | NC score | 0.006257 (rank : 159) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15542, Q86UZ7, Q9Y4K5 | Gene names | TAF5, TAF2D | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 5 (Transcription initiation factor TFIID 100 kDa subunit) (TAF(II)100) (TAFII-100) (TAFII100). | |||||
|
TAOK1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | 0.006999 (rank : 156) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1734 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7L7X3, Q96L75, Q9H2K7, Q9H7S5, Q9P2I6 | Gene names | TAOK1, KIAA1361, MAP3K16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1) (STE20-like kinase PSK2) (Kinase from chicken homolog B) (hKFC-B). | |||||
|
TAOK1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | 0.006946 (rank : 157) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1720 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5F2E8, Q69ZL2, Q8JZX2, Q91VG7 | Gene names | Taok1, Kiaa1361 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1). | |||||
|
TLR13_MOUSE
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.009852 (rank : 148) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6R5N8, Q3TDS2 | Gene names | Tlr13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Toll-like receptor 13 precursor. | |||||
|
TP53B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | 0.016032 (rank : 125) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 413 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q12888, Q2M1Z7, Q4LE46, Q5FWZ3, Q7Z3U4 | Gene names | TP53BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
AQR_HUMAN
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.014093 (rank : 132) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60306, Q2YDX9, Q6IRU8, Q6PIC8 | Gene names | AQR, KIAA0560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intron-binding protein aquarius (Intron-binding protein of 160 kDa) (IBP160). | |||||
|
CAPS2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.011821 (rank : 140) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BYR5, O08903, Q66JM7, Q6PCL7, Q76I88, Q7TMM6, Q80ZV8, Q8BL25, Q8BY04, Q8K3K6 | Gene names | Cadps2, Caps2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-dependent secretion activator 2 (Calcium-dependent activator protein for secretion 2) (CAPS-2). | |||||
|
IQGA2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | 0.018530 (rank : 117) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 87) | NC score | 0.018622 (rank : 115) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
STAP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | 0.030324 (rank : 93) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R0L1, Q8BWS2 | Gene names | Stap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-transducing adaptor protein 2 (STAP-2). | |||||
|
SYMPK_HUMAN
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.037879 (rank : 87) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92797, O00521, O00689, O00733, Q59GT5, Q8N2U5 | Gene names | SYMPK, SPK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Symplekin. | |||||
|
TTBK2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 90) | NC score | 0.010245 (rank : 146) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 822 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6IQ55, O94932, Q6ZN52, Q8IVV1 | Gene names | TTBK2, KIAA0847 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 2 (EC 2.7.11.1). | |||||
|
UACA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 91) | NC score | 0.011559 (rank : 142) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1622 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9BZF9, Q8N3B8, Q9HCL1, Q9NWC6 | Gene names | UACA, KIAA1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein. | |||||
|
ZO1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 92) | NC score | 0.007633 (rank : 155) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q07157, Q4ZGJ6 | Gene names | TJP1, ZO1 | |||
|
Domain Architecture |

|
|||||
| Description | Tight junction protein ZO-1 (Zonula occludens 1 protein) (Zona occludens 1 protein) (Tight junction protein 1). | |||||
|
AVEN_MOUSE
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.030972 (rank : 92) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D9K3 | Gene names | Aven | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell death regulator Aven. | |||||
|
CCD13_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.018483 (rank : 118) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8IYE1 | Gene names | CCDC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 13. | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.014809 (rank : 128) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
CN094_HUMAN
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.034101 (rank : 90) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H6D7, Q86T15, Q86T16, Q86U43, Q9NX59 | Gene names | C14orf94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf94. | |||||
|
DCDC2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.012077 (rank : 139) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UHG0, Q5VTR8, Q5VTR9, Q86W35, Q9UFD1, Q9ULR6 | Gene names | DCDC2, KIAA1154 | |||
|
Domain Architecture |
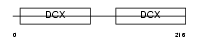
|
|||||
| Description | Doublecortin domain-containing protein 2 (RU2S protein). | |||||
|
DEN2A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.014555 (rank : 130) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C4S8, Q3TJU2, Q3TUG3, Q3UXA3 | Gene names | Dennd2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2A. | |||||
|
DJC16_HUMAN
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.102647 (rank : 31) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y2G8, Q68D57, Q86X32, Q8N5P4 | Gene names | DNAJC16, KIAA0962 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 16 precursor. | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | 0.013584 (rank : 134) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
IF4G1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 101) | NC score | 0.009224 (rank : 149) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
TPM3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.013350 (rank : 136) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 689 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P06753, P12324, Q969Q2, Q9NQH8 | Gene names | TPM3 | |||
|
Domain Architecture |

|
|||||
| Description | Tropomyosin alpha-3 chain (Tropomyosin-3) (Tropomyosin gamma) (hTM5). | |||||
|
TPM3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.013588 (rank : 133) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P21107, Q09021, Q60606, Q80SW6, Q9EPW3 | Gene names | Tpm3, Tpm-5, Tpm5 | |||
|
Domain Architecture |
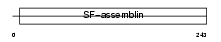
|
|||||
| Description | Tropomyosin alpha-3 chain (Tropomyosin-3) (Tropomyosin gamma). | |||||
|
A2MG_HUMAN
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.012522 (rank : 138) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P01023, Q13677, Q59F47, Q5QTS0, Q68DN2, Q6PIY3, Q6PN97 | Gene names | A2M | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-2-macroglobulin precursor (Alpha-2-M). | |||||
|
BRD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.006060 (rank : 160) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P25440, O00699, O00700, Q15310, Q5STC9, Q969U4 | Gene names | BRD2, KIAA9001, RING3 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 2 (Protein RING3) (O27.1.1). | |||||
|
DCJ14_HUMAN
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.088552 (rank : 52) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6Y2X3, Q66K17, Q96N59, Q96T63 | Gene names | DNAJC14, DRIP78 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 14 (Dopamine receptor-interacting protein of 78 kDa) (DRiP78). | |||||
|
DCJ14_MOUSE
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.086882 (rank : 54) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q921R4, Q3TX73, Q8BUU3, Q9CYB7 | Gene names | Dnajc14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 14. | |||||
|
DJC18_HUMAN
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.104636 (rank : 30) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H819 | Gene names | DNAJC18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 18. | |||||
|
DYH11_HUMAN
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.008666 (rank : 153) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96DT5, Q9UJ82 | Gene names | DNAH11 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 11 (Axonemal beta dynein heavy chain 11). | |||||
|
F125B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.020571 (rank : 111) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6KAU4, Q6PB42 | Gene names | Fam125b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM125B. | |||||
|
GRHL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.010361 (rank : 145) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ISB3, Q6NT03, Q9H8B8 | Gene names | GRHL2, BOM, TFCP2L3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Grainyhead-like protein 2 homolog (Brother of mammalian grainyhead) (Transcription factor CP2-like 3). | |||||
|
GRHL2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.010223 (rank : 147) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K5C0, Q80UZ5 | Gene names | Grhl2, Bom, Tcfcp2l3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Grainyhead-like protein 2 homolog (Brother of mammalian grainyhead) (Transcription factor CP2-like 3). | |||||
|
K1219_MOUSE
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.017657 (rank : 119) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BQZ4, Q80TH5, Q8BQN1 | Gene names | Kiaa1219 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1219. | |||||
|
MECP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.016351 (rank : 124) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51608, O15233 | Gene names | MECP2 | |||
|
Domain Architecture |
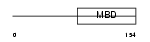
|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
MECP2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.016377 (rank : 123) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z2D6 | Gene names | Mecp2 | |||
|
Domain Architecture |
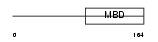
|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
MELPH_HUMAN
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.016736 (rank : 122) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BV36, Q9HA71 | Gene names | MLPH, SLAC2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanophilin (Exophilin-3) (Synaptotagmin-like protein 2a) (Slp homolog lacking C2 domains a). | |||||
|
MPP4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.013505 (rank : 135) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6P7F1, Q8BTT3, Q8BXM5, Q920P7, Q920P8 | Gene names | Mpp4, Dlg6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAGUK p55 subfamily member 4 (Discs large homolog 6) (MDLG6). | |||||
|
MYO1E_HUMAN
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.003672 (rank : 163) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q12965, Q14778 | Gene names | MYO1E, MYO1C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ie (Myosin Ic). | |||||
|
NVL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.009088 (rank : 150) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DBY8, Q3USC4, Q8BW27, Q8K2B5 | Gene names | Nvl | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear valosin-containing protein-like (Nuclear VCP-like protein) (NVLp). | |||||
|
P4HA2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.008912 (rank : 151) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15460, Q8WWN0 | Gene names | P4HA2 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-2 subunit precursor (EC 1.14.11.2) (4-PH alpha-2) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-2 subunit). | |||||
|
SGOL2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.012713 (rank : 137) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TSY8, Q811I4, Q8C9C4, Q8CGJ4, Q9CS55 | Gene names | Sgol2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2. | |||||
|
SNX13_MOUSE
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.018562 (rank : 116) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PHS6 | Gene names | Snx13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-13. | |||||
|
SPB5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.002906 (rank : 164) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70124 | Gene names | Serpinb5, Pi5, Spi5, Spi7 | |||
|
Domain Architecture |

|
|||||
| Description | Serpin B5 precursor (Maspin) (Protease inhibitor 5). | |||||
|
STK4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.001752 (rank : 165) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13043, Q15802, Q4G156, Q5H982, Q6PD60, Q9BR32, Q9NTZ4 | Gene names | STK4, MST1 | |||
|
Domain Architecture |
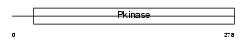
|
|||||
| Description | Serine/threonine-protein kinase 4 (EC 2.7.11.1) (STE20-like kinase MST1) (MST-1) (Mammalian STE20-like protein kinase 1) (Serine/threonine-protein kinase Krs-2). | |||||
|
STK4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.001724 (rank : 166) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 873 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JI11 | Gene names | Stk4, Mst1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase 4 (EC 2.7.11.1) (STE20-like kinase MST1) (MST-1) (Mammalian STE20-like protein kinase 1). | |||||
|
TAF5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.004665 (rank : 162) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C092, Q8BTR2 | Gene names | Taf5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 5 (Transcription initiation factor TFIID 100 kDa subunit) (TAF(II)100) (TAFII-100) (TAFII100). | |||||
|
TTLL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 127) | NC score | 0.022138 (rank : 108) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95922, Q9BR27, Q9NRS9, Q9UMU0 | Gene names | TTLL1, C22orf7 | |||
|
Domain Architecture |
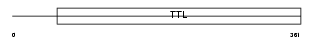
|
|||||
| Description | Probable tubulin polyglutamylase (EC 6.-.-.-) (Tubulin polyglutamylase complex subunit 3) (PGs3) (Tubulin--tyrosine ligase-like protein 1). | |||||
|
TTLL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 128) | NC score | 0.022174 (rank : 107) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91V51, Q3TGC8, Q543S4, Q8C0A2, Q91ZG1 | Gene names | Ttll1 | |||
|
Domain Architecture |
|
|||||
| Description | Probable tubulin polyglutamylase (EC 6.-.-.-) (Tubulin polyglutamylase complex subunit 3) (PGs3) (Tubulin--tyrosine ligase-like protein 1) (p49). | |||||
|
CU055_HUMAN
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.050183 (rank : 75) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NX36 | Gene names | C21orf55 | |||
|
Domain Architecture |
|
|||||
| Description | J domain-containing protein C21orf55. | |||||
|
CU055_MOUSE
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.056256 (rank : 72) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8VCE1, Q99JX0 | Gene names | ORF28 | |||
|
Domain Architecture |
|
|||||
| Description | J domain-containing protein C21orf55 homolog. | |||||
|
DCJ11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.089332 (rank : 50) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9NVH1, Q4VWF5, Q5VZN0, Q6PK20, Q6PK70, Q8NDM2, Q96CL7 | Gene names | DNAJC11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 11. | |||||
|
DCJ11_MOUSE
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.088648 (rank : 51) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5U458, Q8BP83, Q8C1Z4, Q8C6U5 | Gene names | Dnajc11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 11. | |||||
|
DJC12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.080710 (rank : 59) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UKB3, Q9UKB2 | Gene names | DNAJC12, JDP1 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 12 (J domain-containing protein 1). | |||||
|
DJC12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.081340 (rank : 58) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9R022 | Gene names | Dnajc12, Jdp1 | |||
|
Domain Architecture |
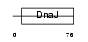
|
|||||
| Description | DnaJ homolog subfamily C member 12 (J domain-containing protein 1). | |||||
|
DJC16_MOUSE
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.090017 (rank : 47) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80TN4, Q811G1, Q8BHI2 | Gene names | Dnajc16, Kiaa0962 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 16 precursor. | |||||
|
DJC18_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.091071 (rank : 46) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9CZJ9, Q8R371 | Gene names | Dnajc18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 18. | |||||
|
DNJA3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.097423 (rank : 37) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96EY1, O75472, Q8WUJ6, Q8WXJ3, Q96D76, Q96IV1, Q9NYH8 | Gene names | DNAJA3, TID1 | |||
|
Domain Architecture |
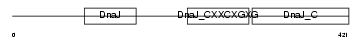
|
|||||
| Description | DnaJ homolog subfamily A member 3, mitochondrial precursor (Tumorous imaginal discs protein Tid56 homolog) (DnaJ protein Tid-1) (hTid-1). | |||||
|
DNJA3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.098279 (rank : 34) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q99M87, Q8BSM0, Q99L09, Q99P71, Q99P76, Q9CT11, Q9DBJ7, Q9DC44 | Gene names | Dnaja3, Tid1 | |||
|
Domain Architecture |
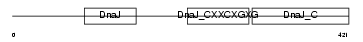
|
|||||
| Description | DnaJ homolog subfamily A member 3, mitochondrial precursor (Tumorous imaginal discs protein Tid56 homolog) (DnaJ protein Tid-1) (mTid-1). | |||||
|
DNJB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.096635 (rank : 43) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P25685 | Gene names | DNAJB1, DNAJ1, HDJ1, HSPF1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 1 (Heat shock 40 kDa protein 1) (Heat shock protein 40) (HSP40) (DnaJ protein homolog 1) (HDJ-1). | |||||
|
DNJB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.096868 (rank : 41) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9QYJ3 | Gene names | Dnajb1, Hsp40, Hspf1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 1 (Heat shock 40 kDa protein 1) (Heat shock protein 40) (HSP40). | |||||
|
DNJB4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.097260 (rank : 40) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UDY4, Q13431 | Gene names | DNAJB4, DNAJW, HLJ1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 4 (Heat shock 40 kDa protein 1 homolog) (Heat shock protein 40 homolog) (HSP40 homolog). | |||||
|
DNJB4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.097320 (rank : 39) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9D832, Q3TS92, Q9D9U2 | Gene names | Dnajb4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 4. | |||||
|
DNJB9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.099613 (rank : 33) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UBS3 | Gene names | DNAJB9, MDG1 | |||
|
Domain Architecture |
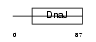
|
|||||
| Description | DnaJ homolog subfamily B member 9 (Microvascular endothelial differentiation gene 1 protein) (Mdg-1). | |||||
|
DNJB9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.100008 (rank : 32) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9QYI6, Q9DAW1 | Gene names | Dnajb9 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 9 (mDJ7). | |||||
|
DNJBB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.097818 (rank : 36) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UBS4, Q96JC6 | Gene names | DNAJB11, EDJ, ERJ3 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 11 precursor (ER-associated dnaJ protein 3) (ErJ3) (ER-associated Hsp40 co-chaperone) (hDj9) (PWP1- interacting protein 4). | |||||
|
DNJBB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.097955 (rank : 35) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q99KV1, Q543I7 | Gene names | Dnajb11 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 11 precursor. | |||||
|
DNJBD_HUMAN
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.097353 (rank : 38) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P59910, Q8IZW5 | Gene names | DNAJB13, TSARG3, TSARG6 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 13 (Testis spermatocyte apoptosis- related gene 6 protein) (Testis and spermatogenesis cell-related protein 6) (Testis spermatogenesis apoptosis-related gene 6 protein) (Testis spermatogenesis apoptosis-related gene 3 protein). | |||||
|
DNJBD_MOUSE
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.096763 (rank : 42) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80Y75, Q8CJA2 | Gene names | Dnajb13, Tsarg, Tsarg3, Tsarg6 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 13 (Testis spermatocyte apoptosis- related gene 6 protein) (Testis and spermatogenesis cell-related protein 6) (Testis spermatogenesis apoptosis-related gene 6 protein) (Testis spermatogenesis apoptosis-related gene 3 protein). | |||||
|
DNJC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.084461 (rank : 57) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96KC8 | Gene names | DNAJC1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1. | |||||
|
DNJC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.085732 (rank : 55) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q61712 | Gene names | Dnajc1, Dnajl1, Mtj1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1 (DnaJ protein homolog MTJ1). | |||||
|
DNJC3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.073706 (rank : 61) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q13217, Q86WT9, Q8N4N2 | Gene names | DNAJC3, P58IPK, PRKRI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 3 (Interferon-induced, double-stranded RNA-activated protein kinase inhibitor) (Protein kinase inhibitor p58) (Protein kinase inhibitor of 58 kDa). | |||||
|
DNJC3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.073473 (rank : 62) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q91YW3, Q60873 | Gene names | Dnajc3, P58ipk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 3 (Interferon-induced, double-stranded RNA-activated protein kinase inhibitor) (Protein kinase inhibitor p58) (Protein kinase inhibitor of 58 kDa). | |||||
|
DNJC4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.088330 (rank : 53) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9NNZ3, O14716 | Gene names | DNAJC4, HSPF2, MCG18 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 4 (Multiple endocrine neoplasia type 1 candidate protein number 18) (DnaJ-like protein HSPF2). | |||||
|
DNJC4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.085505 (rank : 56) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9D844, O70278 | Gene names | Dnajc4, Mcg18 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 4 (Multiple endocrine neoplasia type 1 candidate protein number 18 homolog). | |||||
|
DNJC7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.071406 (rank : 65) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99615 | Gene names | DNAJC7, TPR2, TTC2 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2). | |||||
|
DNJC7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.070415 (rank : 69) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYI3, Q3TKR1, Q8BPG3, Q8CIL2, Q9CT29, Q9D026 | Gene names | Dnajc7, Ttc2 | |||
|
Domain Architecture |
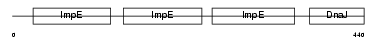
|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2) (MDj11). | |||||
|
DNJC8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.071530 (rank : 64) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75937, Q6IBA4, Q8N4Z5, Q9P051, Q9P067 | Gene names | DNAJC8, SPF31 | |||
|
Domain Architecture |
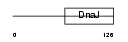
|
|||||
| Description | DnaJ homolog subfamily C member 8 (Splicing protein spf31). | |||||
|
DNJC8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.070923 (rank : 68) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6NZB0, Q6PG72, Q8C2M6 | Gene names | Dnajc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 8. | |||||
|
DNJC9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.089711 (rank : 49) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8WXX5 | Gene names | DNAJC9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 9 (DnaJ protein SB73). | |||||
|
DNJC9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.089952 (rank : 48) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91WN1, Q3TSG3, Q8R0E3 | Gene names | Dnajc9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 9. | |||||
|
SEC63_HUMAN
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.058672 (rank : 71) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SEC63_MOUSE
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.058944 (rank : 70) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
WBS18_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.091392 (rank : 45) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96LL9, Q9BSG8 | Gene names | WBSCR18 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 18 protein. | |||||
|
WBS18_MOUSE
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.096187 (rank : 44) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P59041 | Gene names | Wbscr18 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 18 protein homolog. | |||||
|
ZCSL3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.078731 (rank : 60) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6P3W2 | Gene names | ZCSL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CSL-type zinc finger-containing protein 3. | |||||
|
ZCSL3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.072319 (rank : 63) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q91ZF0, Q9D9S7 | Gene names | Zcsl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CSL-type zinc finger-containing protein 3 (J-domain protein DjC7). | |||||
|
SACS_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 128 | |
| SwissProt Accessions | Q9NZJ4, O94835 | Gene names | SACS, KIAA0730 | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
SACS_MOUSE
|
||||||
| NC score | 0.994673 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9JLC8 | Gene names | Sacs | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
DNJBA_MOUSE
|
||||||
| NC score | 0.194012 (rank : 3) | θ value | 0.0113563 (rank : 11) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYI5 | Gene names | Dnajb10 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 10 (mDJ8). | |||||
|
DNJB7_MOUSE
|
||||||
| NC score | 0.190880 (rank : 4) | θ value | 0.00134147 (rank : 4) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYI8, Q9DA41 | Gene names | Dnajb7 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 7 (mDJ5). | |||||
|
DNJB6_HUMAN
|
||||||
| NC score | 0.190155 (rank : 5) | θ value | 0.00102713 (rank : 3) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O75190, O95806, Q9UIK6 | Gene names | DNAJB6, HSJ2, MSJ1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 6 (Heat shock protein J2) (HSJ-2) (MSJ-1) (HHDJ1) (MRJ). | |||||
|
DNJB2_HUMAN
|
||||||
| NC score | 0.188298 (rank : 6) | θ value | 0.00509761 (rank : 7) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P25686, Q8IUK1, Q8IUK2, Q96F52 | Gene names | DNAJB2, HSJ1, HSPF3 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 2 (Heat shock 40 kDa protein 3) (DnaJ protein homolog 1) (HSJ-1). | |||||
|
DNJB8_HUMAN
|
||||||
| NC score | 0.185891 (rank : 7) | θ value | 0.00298849 (rank : 5) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NHS0 | Gene names | DNAJB8 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 8. | |||||
|
DNJB7_HUMAN
|
||||||
| NC score | 0.184470 (rank : 8) | θ value | 0.00509761 (rank : 9) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q7Z6W7, Q5H904, Q8WYJ7 | Gene names | DNAJB7, HSC3 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 7. | |||||
|
DNJB8_MOUSE
|
||||||
| NC score | 0.184165 (rank : 9) | θ value | 0.00390308 (rank : 6) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYI7 | Gene names | Dnajb8 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 8 (mDJ6). | |||||
|
DNJB3_MOUSE
|
||||||
| NC score | 0.183673 (rank : 10) | θ value | 0.0113563 (rank : 10) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35723, Q9DAN3, Q9DAN4 | Gene names | Dnajb3, Hsj3, Msj1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 3 (DnaJ protein homolog 3) (Heat shock J3 protein) (HSJ-3) (MSJ-1). | |||||
|
DNJB6_MOUSE
|
||||||
| NC score | 0.183414 (rank : 11) | θ value | 0.00509761 (rank : 8) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O54946, Q99LA5, Q9QYI9 | Gene names | Dnajb6, Hsj2, Mrj | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 6 (Heat shock protein J2) (HSJ-2) (MRJ) (mDj4). | |||||
|
DNJC5_HUMAN
|
||||||
| NC score | 0.167951 (rank : 12) | θ value | 0.0330416 (rank : 12) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H3Z4, Q9H3Z5, Q9H7H2 | Gene names | DNAJC5, CSP | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 5 (Cysteine string protein) (CSP). | |||||
|
DNJC5_MOUSE
|
||||||
| NC score | 0.167923 (rank : 13) | θ value | 0.0330416 (rank : 13) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P60904, P54101 | Gene names | Dnajc5, Csp | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 5 (Cysteine string protein) (CSP). | |||||
|
DNJ5B_HUMAN
|
||||||
| NC score | 0.156775 (rank : 14) | θ value | 0.163984 (rank : 14) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UF47, Q969Y8 | Gene names | DNAJC5B | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 5B (Beta cysteine string protein) (Beta-CSP). | |||||
|
DNJ5B_MOUSE
|
||||||
| NC score | 0.149078 (rank : 15) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9CQ94 | Gene names | Dnajc5b | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 5B (Beta cysteine string protein) (Beta-CSP). | |||||
|
DNJ5G_HUMAN
|
||||||
| NC score | 0.149057 (rank : 16) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8N7S2, Q8IYQ4, Q96RJ8 | Gene names | DNAJC5G | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 5G (Gamma cysteine string protein) (Gamma-CSP). | |||||
|
DNJCD_HUMAN
|
||||||
| NC score | 0.138281 (rank : 17) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O75165, Q6PI82, Q6UJ77, Q6ZSW1, Q6ZUT5, Q86XG3, Q96DC1, Q9BWK9 | Gene names | DNAJC13, KIAA0678, RME8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 13 (Required for receptor-mediated endocytosis 8). | |||||
|
DNJA4_MOUSE
|
||||||
| NC score | 0.126939 (rank : 18) | θ value | 0.47712 (rank : 22) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9JMC3, Q543S9, Q9CTD6, Q9CUD4 | Gene names | Dnaja4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 4 (MmDjA4). | |||||
|
DNJA4_HUMAN
|
||||||
| NC score | 0.125837 (rank : 19) | θ value | 0.62314 (rank : 25) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WW22, Q8N7P2 | Gene names | DNAJA4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 4. | |||||
|
DNJBC_HUMAN
|
||||||
| NC score | 0.124397 (rank : 20) | θ value | 1.06291 (rank : 32) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NXW2, Q9H6H0 | Gene names | DNAJB12 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 12. | |||||
|
DNJB5_MOUSE
|
||||||
| NC score | 0.124307 (rank : 21) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O89114 | Gene names | Dnajb5, Hsc40 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 5 (Heat shock protein Hsp40-3) (Heat shock protein cognate 40) (Hsc40). | |||||
|
DJC17_MOUSE
|
||||||
| NC score | 0.123580 (rank : 22) | θ value | 1.81305 (rank : 40) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q91WT4, Q8C5R1 | Gene names | Dnajc17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 17. | |||||
|
DNJB5_HUMAN
|
||||||
| NC score | 0.122995 (rank : 23) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O75953, Q8TDR7 | Gene names | DNAJB5, HSC40 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 5 (Heat shock protein Hsp40-3) (Heat shock protein cognate 40) (Hsc40) (Hsp40-2). | |||||
|
DJC17_HUMAN
|
||||||
| NC score | 0.120606 (rank : 24) | θ value | 1.81305 (rank : 39) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9NVM6 | Gene names | DNAJC17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 17. | |||||
|
DNJA1_HUMAN
|
||||||
| NC score | 0.120312 (rank : 25) | θ value | 1.06291 (rank : 30) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P31689 | Gene names | DNAJA1, DNAJ2, HDJ2, HSJ2, HSPF4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 1 (Heat shock 40 kDa protein 4) (DnaJ protein homolog 2) (HSJ-2) (HSDJ). | |||||
|
DNJA1_MOUSE
|
||||||
| NC score | 0.120170 (rank : 26) | θ value | 1.06291 (rank : 31) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63037, P54102 | Gene names | Dnaja1, Dnaj2, Hsj2, Hspf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily A member 1 (Heat shock 40 kDa protein 4) (DnaJ protein homolog 2) (HSJ-2). | |||||
|
DNJA2_MOUSE
|
||||||
| NC score | 0.115682 (rank : 27) | θ value | 1.81305 (rank : 42) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYJ0 | Gene names | Dnaja2 | |||
|
Domain Architecture |
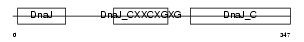
|
|||||
| Description | DnaJ homolog subfamily A member 2 (mDj3). | |||||
|
DNJA2_HUMAN
|
||||||
| NC score | 0.115553 (rank : 28) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O60884, O14711 | Gene names | DNAJA2, CPR3, HIRIP4 | |||
|
Domain Architecture |
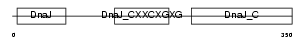
|
|||||
| Description | DnaJ homolog subfamily A member 2 (HIRA-interacting protein 4) (Cell cycle progression restoration gene 3 protein) (Dnj3) (NY-REN-14 antigen). | |||||
|
DNJBC_MOUSE
|
||||||
| NC score | 0.114103 (rank : 29) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYI4 | Gene names | Dnajb12 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 12 (mDJ10). | |||||
|
DJC18_HUMAN
|
||||||
| NC score | 0.104636 (rank : 30) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H819 | Gene names | DNAJC18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 18. | |||||
|
DJC16_HUMAN
|
||||||
| NC score | 0.102647 (rank : 31) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y2G8, Q68D57, Q86X32, Q8N5P4 | Gene names | DNAJC16, KIAA0962 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 16 precursor. | |||||
|
DNJB9_MOUSE
|
||||||
| NC score | 0.100008 (rank : 32) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9QYI6, Q9DAW1 | Gene names | Dnajb9 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 9 (mDJ7). | |||||
|
DNJB9_HUMAN
|
||||||
| NC score | 0.099613 (rank : 33) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UBS3 | Gene names | DNAJB9, MDG1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 9 (Microvascular endothelial differentiation gene 1 protein) (Mdg-1). | |||||
|
DNJA3_MOUSE
|
||||||
| NC score | 0.098279 (rank : 34) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q99M87, Q8BSM0, Q99L09, Q99P71, Q99P76, Q9CT11, Q9DBJ7, Q9DC44 | Gene names | Dnaja3, Tid1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 3, mitochondrial precursor (Tumorous imaginal discs protein Tid56 homolog) (DnaJ protein Tid-1) (mTid-1). | |||||
|
DNJBB_MOUSE
|
||||||
| NC score | 0.097955 (rank : 35) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q99KV1, Q543I7 | Gene names | Dnajb11 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 11 precursor. | |||||
|
DNJBB_HUMAN
|
||||||
| NC score | 0.097818 (rank : 36) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UBS4, Q96JC6 | Gene names | DNAJB11, EDJ, ERJ3 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 11 precursor (ER-associated dnaJ protein 3) (ErJ3) (ER-associated Hsp40 co-chaperone) (hDj9) (PWP1- interacting protein 4). | |||||
|
DNJA3_HUMAN
|
||||||
| NC score | 0.097423 (rank : 37) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96EY1, O75472, Q8WUJ6, Q8WXJ3, Q96D76, Q96IV1, Q9NYH8 | Gene names | DNAJA3, TID1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 3, mitochondrial precursor (Tumorous imaginal discs protein Tid56 homolog) (DnaJ protein Tid-1) (hTid-1). | |||||
|
DNJBD_HUMAN
|
||||||
| NC score | 0.097353 (rank : 38) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P59910, Q8IZW5 | Gene names | DNAJB13, TSARG3, TSARG6 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 13 (Testis spermatocyte apoptosis- related gene 6 protein) (Testis and spermatogenesis cell-related protein 6) (Testis spermatogenesis apoptosis-related gene 6 protein) (Testis spermatogenesis apoptosis-related gene 3 protein). | |||||
|
DNJB4_MOUSE
|
||||||
| NC score | 0.097320 (rank : 39) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9D832, Q3TS92, Q9D9U2 | Gene names | Dnajb4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 4. | |||||
|
DNJB4_HUMAN
|
||||||
| NC score | 0.097260 (rank : 40) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UDY4, Q13431 | Gene names | DNAJB4, DNAJW, HLJ1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 4 (Heat shock 40 kDa protein 1 homolog) (Heat shock protein 40 homolog) (HSP40 homolog). | |||||
|
DNJB1_MOUSE
|
||||||
| NC score | 0.096868 (rank : 41) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9QYJ3 | Gene names | Dnajb1, Hsp40, Hspf1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 1 (Heat shock 40 kDa protein 1) (Heat shock protein 40) (HSP40). | |||||
|
DNJBD_MOUSE
|
||||||
| NC score | 0.096763 (rank : 42) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80Y75, Q8CJA2 | Gene names | Dnajb13, Tsarg, Tsarg3, Tsarg6 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 13 (Testis spermatocyte apoptosis- related gene 6 protein) (Testis and spermatogenesis cell-related protein 6) (Testis spermatogenesis apoptosis-related gene 6 protein) (Testis spermatogenesis apoptosis-related gene 3 protein). | |||||
|
DNJB1_HUMAN
|
||||||
| NC score | 0.096635 (rank : 43) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P25685 | Gene names | DNAJB1, DNAJ1, HDJ1, HSPF1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 1 (Heat shock 40 kDa protein 1) (Heat shock protein 40) (HSP40) (DnaJ protein homolog 1) (HDJ-1). | |||||
|
WBS18_MOUSE
|
||||||
| NC score | 0.096187 (rank : 44) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P59041 | Gene names | Wbscr18 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 18 protein homolog. | |||||
|
WBS18_HUMAN
|
||||||
| NC score | 0.091392 (rank : 45) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96LL9, Q9BSG8 | Gene names | WBSCR18 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 18 protein. | |||||
|
DJC18_MOUSE
|
||||||
| NC score | 0.091071 (rank : 46) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9CZJ9, Q8R371 | Gene names | Dnajc18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 18. | |||||
|
DJC16_MOUSE
|
||||||
| NC score | 0.090017 (rank : 47) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80TN4, Q811G1, Q8BHI2 | Gene names | Dnajc16, Kiaa0962 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 16 precursor. | |||||
|
DNJC9_MOUSE
|
||||||
| NC score | 0.089952 (rank : 48) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91WN1, Q3TSG3, Q8R0E3 | Gene names | Dnajc9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 9. | |||||
|
DNJC9_HUMAN
|
||||||
| NC score | 0.089711 (rank : 49) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8WXX5 | Gene names | DNAJC9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 9 (DnaJ protein SB73). | |||||
|
DCJ11_HUMAN
|
||||||
| NC score | 0.089332 (rank : 50) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9NVH1, Q4VWF5, Q5VZN0, Q6PK20, Q6PK70, Q8NDM2, Q96CL7 | Gene names | DNAJC11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 11. | |||||
|
DCJ11_MOUSE
|
||||||
| NC score | 0.088648 (rank : 51) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5U458, Q8BP83, Q8C1Z4, Q8C6U5 | Gene names | Dnajc11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 11. | |||||
|
DCJ14_HUMAN
|
||||||
| NC score | 0.088552 (rank : 52) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6Y2X3, Q66K17, Q96N59, Q96T63 | Gene names | DNAJC14, DRIP78 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 14 (Dopamine receptor-interacting protein of 78 kDa) (DRiP78). | |||||
|
DNJC4_HUMAN
|
||||||
| NC score | 0.088330 (rank : 53) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9NNZ3, O14716 | Gene names | DNAJC4, HSPF2, MCG18 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 4 (Multiple endocrine neoplasia type 1 candidate protein number 18) (DnaJ-like protein HSPF2). | |||||
|
DCJ14_MOUSE
|
||||||
| NC score | 0.086882 (rank : 54) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q921R4, Q3TX73, Q8BUU3, Q9CYB7 | Gene names | Dnajc14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 14. | |||||
|
DNJC1_MOUSE
|
||||||
| NC score | 0.085732 (rank : 55) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q61712 | Gene names | Dnajc1, Dnajl1, Mtj1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1 (DnaJ protein homolog MTJ1). | |||||
|
DNJC4_MOUSE
|
||||||
| NC score | 0.085505 (rank : 56) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9D844, O70278 | Gene names | Dnajc4, Mcg18 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 4 (Multiple endocrine neoplasia type 1 candidate protein number 18 homolog). | |||||
|
DNJC1_HUMAN
|
||||||
| NC score | 0.084461 (rank : 57) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96KC8 | Gene names | DNAJC1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1. | |||||
|
DJC12_MOUSE
|
||||||
| NC score | 0.081340 (rank : 58) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9R022 | Gene names | Dnajc12, Jdp1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 12 (J domain-containing protein 1). | |||||
|
DJC12_HUMAN
|
||||||
| NC score | 0.080710 (rank : 59) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UKB3, Q9UKB2 | Gene names | DNAJC12, JDP1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 12 (J domain-containing protein 1). | |||||
|
ZCSL3_HUMAN
|
||||||
| NC score | 0.078731 (rank : 60) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6P3W2 | Gene names | ZCSL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CSL-type zinc finger-containing protein 3. | |||||
|
DNJC3_HUMAN
|
||||||
| NC score | 0.073706 (rank : 61) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q13217, Q86WT9, Q8N4N2 | Gene names | DNAJC3, P58IPK, PRKRI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 3 (Interferon-induced, double-stranded RNA-activated protein kinase inhibitor) (Protein kinase inhibitor p58) (Protein kinase inhibitor of 58 kDa). | |||||
|
DNJC3_MOUSE
|
||||||
| NC score | 0.073473 (rank : 62) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q91YW3, Q60873 | Gene names | Dnajc3, P58ipk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 3 (Interferon-induced, double-stranded RNA-activated protein kinase inhibitor) (Protein kinase inhibitor p58) (Protein kinase inhibitor of 58 kDa). | |||||
|
ZCSL3_MOUSE
|
||||||
| NC score | 0.072319 (rank : 63) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q91ZF0, Q9D9S7 | Gene names | Zcsl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CSL-type zinc finger-containing protein 3 (J-domain protein DjC7). | |||||
|
DNJC8_HUMAN
|
||||||
| NC score | 0.071530 (rank : 64) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75937, Q6IBA4, Q8N4Z5, Q9P051, Q9P067 | Gene names | DNAJC8, SPF31 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 8 (Splicing protein spf31). | |||||
|
DNJC7_HUMAN
|
||||||
| NC score | 0.071406 (rank : 65) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99615 | Gene names | DNAJC7, TPR2, TTC2 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2). | |||||
|
COMDA_MOUSE
|
||||||
| NC score | 0.071102 (rank : 66) | θ value | 2.36792 (rank : 45) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8JZY2 | Gene names | Commd10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | COMM domain-containing protein 10 (Down-regulated in W/WV mouse stomach 2) (mDRWMS2). | |||||
|
HEAT1_HUMAN
|
||||||
| NC score | 0.070930 (rank : 67) | θ value | 0.62314 (rank : 26) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H583, Q5T3Q8, Q9NW23 | Gene names | HEATR1, BAP28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HEAT repeat-containing protein 1 (Protein BAP28). | |||||
|
DNJC8_MOUSE
|
||||||
| NC score | 0.070923 (rank : 68) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6NZB0, Q6PG72, Q8C2M6 | Gene names | Dnajc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 8. | |||||
|
DNJC7_MOUSE
|
||||||
| NC score | 0.070415 (rank : 69) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYI3, Q3TKR1, Q8BPG3, Q8CIL2, Q9CT29, Q9D026 | Gene names | Dnajc7, Ttc2 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2) (MDj11). | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.058944 (rank : 70) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.058672 (rank : 71) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
CU055_MOUSE
|
||||||
| NC score | 0.056256 (rank : 72) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8VCE1, Q99JX0 | Gene names | ORF28 | |||
|
Domain Architecture |

|
|||||
| Description | J domain-containing protein C21orf55 homolog. | |||||
|
COMDA_HUMAN
|
||||||
| NC score | 0.055063 (rank : 73) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6G5, Q9P077 | Gene names | COMMD10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | COMM domain-containing protein 10. | |||||
|
SRXN1_MOUSE
|
||||||
| NC score | 0.054278 (rank : 74) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D975, Q3TWN6, Q91VN5 | Gene names | Srxn1, Npn3, Srx | |||
|
Domain Architecture |

|
|||||
| Description | Sulfiredoxin-1 (EC 1.8.98.2) (Neoplastic progression protein 3). | |||||
|
CU055_HUMAN
|
||||||
| NC score | 0.050183 (rank : 75) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NX36 | Gene names | C21orf55 | |||
|
Domain Architecture |

|
|||||
| Description | J domain-containing protein C21orf55. | |||||
|
CENPQ_HUMAN
|
||||||
| NC score | 0.048813 (rank : 76) | θ value | 0.47712 (rank : 20) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7L2Z9, Q6IN61, Q9NVS5 | Gene names | CENPQ, C6orf139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein Q (CENP-Q). | |||||
|
PDZD6_HUMAN
|
||||||
| NC score | 0.048096 (rank : 77) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULD6, Q4W5I8, Q86V55 | Gene names | PDZD6, KIAA1284, PDZK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 6. | |||||
|
K0586_HUMAN
|
||||||
| NC score | 0.043719 (rank : 78) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BVV6, O60328 | Gene names | KIAA0586 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0586. | |||||
|
OTOAN_MOUSE
|
||||||
| NC score | 0.043482 (rank : 79) | θ value | 1.38821 (rank : 36) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K561 | Gene names | Otoa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Otoancorin precursor. | |||||
|
SYMPK_MOUSE
|
||||||
| NC score | 0.042772 (rank : 80) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80X82 | Gene names | Sympk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Symplekin. | |||||
|
CEP35_HUMAN
|
||||||
| NC score | 0.042182 (rank : 81) | θ value | 0.813845 (rank : 29) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
CF152_HUMAN
|
||||||
| NC score | 0.040938 (rank : 82) | θ value | 0.62314 (rank : 24) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q86VQ0, Q9BWX7 | Gene names | C6orf152 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152. | |||||
|
ENPL_HUMAN
|
||||||
| NC score | 0.039389 (rank : 83) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P14625, Q96A97 | Gene names | HSP90B1, TRA1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (gp96 homolog) (Tumor rejection antigen 1). | |||||
|
GEMI2_MOUSE
|
||||||
| NC score | 0.039366 (rank : 84) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CQQ4, Q9DAD7 | Gene names | Sip1, Gemin2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Survival of motor neuron protein-interacting protein 1 (SMN- interacting protein 1) (Component of gems 2) (Gemin-2). | |||||
|
MOR2B_MOUSE
|
||||||
| NC score | 0.038150 (rank : 85) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C5W4, Q8CG24 | Gene names | Morc2b, Tce6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2B (TCE6). | |||||
|
UTY_MOUSE
|
||||||
| NC score | 0.038096 (rank : 86) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P79457, O97979 | Gene names | Uty | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed Y chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein ON the Y chromosome) (Male- specific histocompatibility antigen H-YDB). | |||||
|
SYMPK_HUMAN
|
||||||
| NC score | 0.037879 (rank : 87) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92797, O00521, O00689, O00733, Q59GT5, Q8N2U5 | Gene names | SYMPK, SPK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Symplekin. | |||||
|
ENPL_MOUSE
|
||||||
| NC score | 0.037542 (rank : 88) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P08113, P11427 | Gene names | Hsp90b1, Tra-1, Tra1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (ERP99) (Polymorphic tumor rejection antigen 1) (Tumor rejection antigen gp96). | |||||
|
POLK_MOUSE
|
||||||
| NC score | 0.034844 (rank : 89) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QUG2, Q7TPY7 | Gene names | Polk, Dinb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase kappa (EC 2.7.7.7) (DINB protein) (DINP). | |||||
|
CN094_HUMAN
|
||||||
| NC score | 0.034101 (rank : 90) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H6D7, Q86T15, Q86T16, Q86U43, Q9NX59 | Gene names | C14orf94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf94. | |||||
|
HEM3_MOUSE
|
||||||
| NC score | 0.033412 (rank : 91) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P22907 | Gene names | Hmbs, Uros1 | |||
|
Domain Architecture |

|
|||||
| Description | Porphobilinogen deaminase (EC 2.5.1.61) (Hydroxymethylbilane synthase) (HMBS) (Pre-uroporphyrinogen synthase) (PBG-D). | |||||
|
AVEN_MOUSE
|
||||||
| NC score | 0.030972 (rank : 92) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D9K3 | Gene names | Aven | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell death regulator Aven. | |||||
|
STAP2_MOUSE
|
||||||
| NC score | 0.030324 (rank : 93) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R0L1, Q8BWS2 | Gene names | Stap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-transducing adaptor protein 2 (STAP-2). | |||||
|
CF152_MOUSE
|
||||||
| NC score | 0.030111 (rank : 94) | θ value | 1.81305 (rank : 37) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80ST9, Q9CYM9, Q9D5J9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152 homolog. | |||||
|
BPAEA_HUMAN
|
||||||
| NC score | 0.029703 (rank : 95) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
F125B_HUMAN
|
||||||
| NC score | 0.027846 (rank : 96) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H7P6, Q8N6S7 | Gene names | FAM125B, C9orf28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM125B. | |||||
|
PYC_HUMAN
|
||||||
| NC score | 0.026516 (rank : 97) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P11498, Q16705 | Gene names | PC | |||
|
Domain Architecture |

|
|||||
| Description | Pyruvate carboxylase, mitochondrial precursor (EC 6.4.1.1) (Pyruvic carboxylase) (PCB). | |||||
|
PYC_MOUSE
|
||||||
| NC score | 0.026449 (rank : 98) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q05920 | Gene names | Pc, Pcx | |||
|
Domain Architecture |

|
|||||
| Description | Pyruvate carboxylase, mitochondrial precursor (EC 6.4.1.1) (Pyruvic carboxylase) (PCB). | |||||
|
PERT_MOUSE
|
||||||
| NC score | 0.025603 (rank : 99) | θ value | 1.06291 (rank : 33) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35419, Q8C8B1 | Gene names | Tpo | |||
|
Domain Architecture |
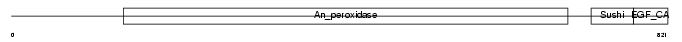
|
|||||
| Description | Thyroid peroxidase precursor (EC 1.11.1.8) (TPO). | |||||
|
COVA1_MOUSE
|
||||||
| NC score | 0.023457 (rank : 100) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8R0Z2, Q8BY59, Q8R372 | Gene names | Cova1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor-associated hydroquinone oxidase (tNOX) (Cytosolic ovarian carcinoma antigen 1) (APK1 antigen) [Includes: Hydroquinone [NADH] oxidase (EC 1.-.-.-); Protein disulfide-thiol oxidoreductase (EC 1.-.-.-)]. | |||||
|
FOXM1_HUMAN
|
||||||
| NC score | 0.023448 (rank : 101) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q08050, O43258, O43259, O43260, Q4ZGG7, Q9BRL2 | Gene names | FOXM1, FKHL16, HFH11, MPP2, WIN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead-related protein FKHL16) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/fork-head homolog 11) (HFH-11) (Winged helix factor from INS-1 cells) (M-phase phosphoprotein 2) (MPM-2 reactive phosphoprotein 2) (Transcription factor Trident). | |||||
|
PARN_HUMAN
|
||||||
| NC score | 0.023337 (rank : 102) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95453 | Gene names | PARN, DAN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A)-specific ribonuclease PARN (EC 3.1.13.4) (Polyadenylate- specific ribonuclease) (Deadenylating nuclease) (Deadenylation nuclease). | |||||
|
COVA1_HUMAN
|
||||||
| NC score | 0.022928 (rank : 103) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q16206, Q5VTJ1, Q5VTJ2, Q8WUX0, Q9NTP6, Q9UH82 | Gene names | COVA1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor-associated hydroquinone oxidase (tNOX) (Cytosolic ovarian carcinoma antigen 1) (APK1 antigen) [Includes: Hydroquinone [NADH] oxidase (EC 1.-.-.-); Protein disulfide-thiol oxidoreductase (EC 1.-.-.-)]. | |||||
|
GOGA1_MOUSE
|
||||||
| NC score | 0.022555 (rank : 104) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 994 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9CW79, Q80YB0 | Gene names | Golga1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
CUL4A_HUMAN
|
||||||
| NC score | 0.022478 (rank : 105) | θ value | 2.36792 (rank : 46) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13619, O75834, Q589T6, Q5TC62, Q6UP08, Q9UP17 | Gene names | CUL4A | |||
|
Domain Architecture |

|
|||||
| Description | Cullin-4A (CUL-4A). | |||||
|
SPAG5_HUMAN
|
||||||
| NC score | 0.022177 (rank : 106) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96R06, O95213, Q9BWE8, Q9NT17, Q9UFE6 | Gene names | SPAG5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 5 (Astrin) (Mitotic spindle-associated protein p126) (MAP126) (Deepest). | |||||
|
TTLL1_MOUSE
|
||||||
| NC score | 0.022174 (rank : 107) | θ value | 8.99809 (rank : 128) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91V51, Q3TGC8, Q543S4, Q8C0A2, Q91ZG1 | Gene names | Ttll1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable tubulin polyglutamylase (EC 6.-.-.-) (Tubulin polyglutamylase complex subunit 3) (PGs3) (Tubulin--tyrosine ligase-like protein 1) (p49). | |||||
|
TTLL1_HUMAN
|
||||||
| NC score | 0.022138 (rank : 108) | θ value | 8.99809 (rank : 127) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95922, Q9BR27, Q9NRS9, Q9UMU0 | Gene names | TTLL1, C22orf7 | |||
|
Domain Architecture |

|
|||||
| Description | Probable tubulin polyglutamylase (EC 6.-.-.-) (Tubulin polyglutamylase complex subunit 3) (PGs3) (Tubulin--tyrosine ligase-like protein 1). | |||||
|
SAPS1_HUMAN
|
||||||
| NC score | 0.021544 (rank : 109) | θ value | 0.47712 (rank : 23) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
BCL7C_MOUSE
|
||||||
| NC score | 0.020602 (rank : 110) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O08664, Q99LS2 | Gene names | Bcl7c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell CLL/lymphoma 7 protein family member C (B-cell chronic lymphocytic leukemia/lymphoma 7C protein). | |||||
|
F125B_MOUSE
|
||||||
| NC score | 0.020571 (rank : 111) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6KAU4, Q6PB42 | Gene names | Fam125b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM125B. | |||||
|
RECQ4_MOUSE
|
||||||
| NC score | 0.020161 (rank : 112) | θ value | 1.06291 (rank : 34) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q75NR7, Q76MT1, Q99PV9 | Gene names | Recql4, Recq4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4). | |||||
|
CDC27_HUMAN
|
||||||
| NC score | 0.019035 (rank : 113) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30260, Q16349, Q96F35 | Gene names | CDC27 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 27 homolog (CDC27Hs) (H-NUC). | |||||
|
DEN2A_HUMAN
|
||||||
| NC score | 0.018848 (rank : 114) | θ value | 1.81305 (rank : 38) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULE3, Q1RMD5, Q86XY0 | Gene names | DENND2A, KIAA1277 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2A. | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.018622 (rank : 115) | θ value | 5.27518 (rank : 87) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
SNX13_MOUSE
|
||||||
| NC score | 0.018562 (rank : 116) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PHS6 | Gene names | Snx13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-13. | |||||
|
IQGA2_HUMAN
|
||||||
| NC score | 0.018530 (rank : 117) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
CCD13_HUMAN
|
||||||
| NC score | 0.018483 (rank : 118) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8IYE1 | Gene names | CCDC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 13. | |||||
|
K1219_MOUSE
|
||||||
| NC score | 0.017657 (rank : 119) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BQZ4, Q80TH5, Q8BQN1 | Gene names | Kiaa1219 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1219. | |||||
|
TTBK2_MOUSE
|
||||||
| NC score | 0.017331 (rank : 120) | θ value | 1.06291 (rank : 35) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3UVR3, Q3TSR6, Q3UFW0, Q571D1, Q8BKA4, Q924U8 | Gene names | Ttbk2, Kiaa0847, Ttbk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 2. | |||||
|
KTN1_HUMAN
|
||||||
| NC score | 0.017135 (rank : 121) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q86UP2, Q13999, Q14707, Q15387, Q86W57 | Gene names | KTN1, CG1, KIAA0004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin (Kinesin receptor) (CG-1 antigen). | |||||
|
MELPH_HUMAN
|
||||||
| NC score | 0.016736 (rank : 122) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BV36, Q9HA71 | Gene names | MLPH, SLAC2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanophilin (Exophilin-3) (Synaptotagmin-like protein 2a) (Slp homolog lacking C2 domains a). | |||||
|
MECP2_MOUSE
|
||||||
| NC score | 0.016377 (rank : 123) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z2D6 | Gene names | Mecp2 | |||
|
Domain Architecture |

|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
MECP2_HUMAN
|
||||||
| NC score | 0.016351 (rank : 124) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51608, O15233 | Gene names | MECP2 | |||
|
Domain Architecture |

|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
TP53B_HUMAN
|
||||||
| NC score | 0.016032 (rank : 125) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 413 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q12888, Q2M1Z7, Q4LE46, Q5FWZ3, Q7Z3U4 | Gene names | TP53BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
AKAP9_HUMAN
|
||||||
| NC score | 0.015330 (rank : 126) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
KIF5C_MOUSE
|
||||||
| NC score | 0.015325 (rank : 127) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P28738, Q9Z2F8 | Gene names | Kif5c, Nkhc2 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.014809 (rank : 128) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
ABCAD_HUMAN
|
||||||
| NC score | 0.014780 (rank : 129) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86UQ4, Q6ZTT7, Q86WI2, Q8N248 | Gene names | ABCA13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-binding cassette sub-family A member 13. | |||||
|
DEN2A_MOUSE
|
||||||
| NC score | 0.014555 (rank : 130) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C4S8, Q3TJU2, Q3TUG3, Q3UXA3 | Gene names | Dennd2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2A. | |||||
|
CCNB3_MOUSE
|
||||||
| NC score | 0.014505 (rank : 131) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q810T2, Q810T3, Q8VDC8 | Gene names | Ccnb3, Cycb3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
AQR_HUMAN
|
||||||
| NC score | 0.014093 (rank : 132) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60306, Q2YDX9, Q6IRU8, Q6PIC8 | Gene names | AQR, KIAA0560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intron-binding protein aquarius (Intron-binding protein of 160 kDa) (IBP160). | |||||
|
TPM3_MOUSE
|
||||||
| NC score | 0.013588 (rank : 133) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P21107, Q09021, Q60606, Q80SW6, Q9EPW3 | Gene names | Tpm3, Tpm-5, Tpm5 | |||
|
Domain Architecture |

|
|||||
| Description | Tropomyosin alpha-3 chain (Tropomyosin-3) (Tropomyosin gamma). | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.013584 (rank : 134) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
MPP4_MOUSE
|
||||||
| NC score | 0.013505 (rank : 135) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6P7F1, Q8BTT3, Q8BXM5, Q920P7, Q920P8 | Gene names | Mpp4, Dlg6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAGUK p55 subfamily member 4 (Discs large homolog 6) (MDLG6). | |||||
|
TPM3_HUMAN
|
||||||
| NC score | 0.013350 (rank : 136) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 689 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P06753, P12324, Q969Q2, Q9NQH8 | Gene names | TPM3 | |||
|
Domain Architecture |

|
|||||
| Description | Tropomyosin alpha-3 chain (Tropomyosin-3) (Tropomyosin gamma) (hTM5). | |||||
|
SGOL2_MOUSE
|
||||||
| NC score | 0.012713 (rank : 137) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TSY8, Q811I4, Q8C9C4, Q8CGJ4, Q9CS55 | Gene names | Sgol2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2. | |||||
|
A2MG_HUMAN
|
||||||
| NC score | 0.012522 (rank : 138) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P01023, Q13677, Q59F47, Q5QTS0, Q68DN2, Q6PIY3, Q6PN97 | Gene names | A2M | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-2-macroglobulin precursor (Alpha-2-M). | |||||
|
DCDC2_HUMAN
|
||||||
| NC score | 0.012077 (rank : 139) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UHG0, Q5VTR8, Q5VTR9, Q86W35, Q9UFD1, Q9ULR6 | Gene names | DCDC2, KIAA1154 | |||
|
Domain Architecture |

|
|||||
| Description | Doublecortin domain-containing protein 2 (RU2S protein). | |||||
|
CAPS2_MOUSE
|
||||||
| NC score | 0.011821 (rank : 140) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BYR5, O08903, Q66JM7, Q6PCL7, Q76I88, Q7TMM6, Q80ZV8, Q8BL25, Q8BY04, Q8K3K6 | Gene names | Cadps2, Caps2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-dependent secretion activator 2 (Calcium-dependent activator protein for secretion 2) (CAPS-2). | |||||
|
MPP4_HUMAN
|
||||||
| NC score | 0.011816 (rank : 141) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96JB8, Q6ZNH6, Q96Q43, Q96Q44 | Gene names | MPP4, ALS2CR5, DLG6 | |||
|
Domain Architecture |

|
|||||
| Description | MAGUK p55 subfamily member 4 (Discs large homolog 6) (Amyotrophic lateral sclerosis 2 chromosomal region candidate gene 5 protein). | |||||
|
UACA_HUMAN
|
||||||
| NC score | 0.011559 (rank : 142) | θ value | 5.27518 (rank : 91) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1622 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9BZF9, Q8N3B8, Q9HCL1, Q9NWC6 | Gene names | UACA, KIAA1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein. | |||||
|
KIF1A_MOUSE
|
||||||
| NC score | 0.010605 (rank : 143) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P33173, Q61770 | Gene names | Kif1a, Atsv, Kif1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF1A (Axonal transporter of synaptic vesicles). | |||||
|
KIF1A_HUMAN
|
||||||
| NC score | 0.010584 (rank : 144) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q12756 | Gene names | KIF1A, ATSV | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF1A (Axonal transporter of synaptic vesicles). | |||||
|
GRHL2_HUMAN
|
||||||
| NC score | 0.010361 (rank : 145) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ISB3, Q6NT03, Q9H8B8 | Gene names | GRHL2, BOM, TFCP2L3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Grainyhead-like protein 2 homolog (Brother of mammalian grainyhead) (Transcription factor CP2-like 3). | |||||
|
TTBK2_HUMAN
|
||||||
| NC score | 0.010245 (rank : 146) | θ value | 5.27518 (rank : 90) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 822 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6IQ55, O94932, Q6ZN52, Q8IVV1 | Gene names | TTBK2, KIAA0847 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 2 (EC 2.7.11.1). | |||||
|
GRHL2_MOUSE
|
||||||
| NC score | 0.010223 (rank : 147) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K5C0, Q80UZ5 | Gene names | Grhl2, Bom, Tcfcp2l3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Grainyhead-like protein 2 homolog (Brother of mammalian grainyhead) (Transcription factor CP2-like 3). | |||||
|
TLR13_MOUSE
|
||||||
| NC score | 0.009852 (rank : 148) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6R5N8, Q3TDS2 | Gene names | Tlr13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Toll-like receptor 13 precursor. | |||||
|
IF4G1_HUMAN
|
||||||
| NC score | 0.009224 (rank : 149) | θ value | 6.88961 (rank : 101) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
NVL_MOUSE
|
||||||
| NC score | 0.009088 (rank : 150) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DBY8, Q3USC4, Q8BW27, Q8K2B5 | Gene names | Nvl | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear valosin-containing protein-like (Nuclear VCP-like protein) (NVLp). | |||||
|
P4HA2_HUMAN
|
||||||
| NC score | 0.008912 (rank : 151) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15460, Q8WWN0 | Gene names | P4HA2 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-2 subunit precursor (EC 1.14.11.2) (4-PH alpha-2) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-2 subunit). | |||||
|
ANGL2_MOUSE
|
||||||
| NC score | 0.008883 (rank : 152) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9R045 | Gene names | Angptl2, Arp2 | |||
|
Domain Architecture |

|
|||||
| Description | Angiopoietin-related protein 2 precursor (Angiopoietin-like 2). | |||||
|
DYH11_HUMAN
|
||||||
| NC score | 0.008666 (rank : 153) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96DT5, Q9UJ82 | Gene names | DNAH11 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 11 (Axonemal beta dynein heavy chain 11). | |||||
|
MYH13_HUMAN
|
||||||
| NC score | 0.008379 (rank : 154) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
ZO1_HUMAN
|
||||||
| NC score | 0.007633 (rank : 155) | θ value | 5.27518 (rank : 92) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q07157, Q4ZGJ6 | Gene names | TJP1, ZO1 | |||
|
Domain Architecture |

|
|||||
| Description | Tight junction protein ZO-1 (Zonula occludens 1 protein) (Zona occludens 1 protein) (Tight junction protein 1). | |||||
|
TAOK1_HUMAN
|
||||||
| NC score | 0.006999 (rank : 156) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1734 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7L7X3, Q96L75, Q9H2K7, Q9H7S5, Q9P2I6 | Gene names | TAOK1, KIAA1361, MAP3K16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1) (STE20-like kinase PSK2) (Kinase from chicken homolog B) (hKFC-B). | |||||
|
TAOK1_MOUSE
|
||||||
| NC score | 0.006946 (rank : 157) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1720 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5F2E8, Q69ZL2, Q8JZX2, Q91VG7 | Gene names | Taok1, Kiaa1361 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1). | |||||
|
SCN1A_HUMAN
|
||||||
| NC score | 0.006306 (rank : 158) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35498, Q16172, Q96LA3, Q9C008 | Gene names | SCN1A, NAC1, SCN1 | |||
|
Domain Architecture |
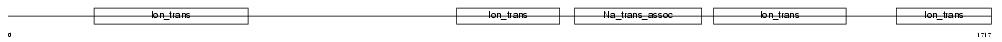
|
|||||
| Description | Sodium channel protein type 1 subunit alpha (Sodium channel protein type I subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.1) (Sodium channel protein, brain I subunit alpha). | |||||
|
TAF5_HUMAN
|
||||||
| NC score | 0.006257 (rank : 159) | θ value | 4.03905 (rank : 79) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15542, Q86UZ7, Q9Y4K5 | Gene names | TAF5, TAF2D | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 5 (Transcription initiation factor TFIID 100 kDa subunit) (TAF(II)100) (TAFII-100) (TAFII100). | |||||
|
BRD2_HUMAN
|
||||||
| NC score | 0.006060 (rank : 160) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P25440, O00699, O00700, Q15310, Q5STC9, Q969U4 | Gene names | BRD2, KIAA9001, RING3 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 2 (Protein RING3) (O27.1.1). | |||||
|
MAST4_HUMAN
|
||||||
| NC score | 0.004850 (rank : 161) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
TAF5_MOUSE
|
||||||
| NC score | 0.004665 (rank : 162) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C092, Q8BTR2 | Gene names | Taf5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 5 (Transcription initiation factor TFIID 100 kDa subunit) (TAF(II)100) (TAFII-100) (TAFII100). | |||||
|
MYO1E_HUMAN
|
||||||
| NC score | 0.003672 (rank : 163) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q12965, Q14778 | Gene names | MYO1E, MYO1C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ie (Myosin Ic). | |||||
|
SPB5_MOUSE
|
||||||
| NC score | 0.002906 (rank : 164) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70124 | Gene names | Serpinb5, Pi5, Spi5, Spi7 | |||
|
Domain Architecture |

|
|||||
| Description | Serpin B5 precursor (Maspin) (Protease inhibitor 5). | |||||
|
STK4_HUMAN
|
||||||
| NC score | 0.001752 (rank : 165) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13043, Q15802, Q4G156, Q5H982, Q6PD60, Q9BR32, Q9NTZ4 | Gene names | STK4, MST1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase 4 (EC 2.7.11.1) (STE20-like kinase MST1) (MST-1) (Mammalian STE20-like protein kinase 1) (Serine/threonine-protein kinase Krs-2). | |||||
|
STK4_MOUSE
|
||||||
| NC score | 0.001724 (rank : 166) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 873 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JI11 | Gene names | Stk4, Mst1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase 4 (EC 2.7.11.1) (STE20-like kinase MST1) (MST-1) (Mammalian STE20-like protein kinase 1). | |||||