Please be patient as the page loads
|
DNJCD_HUMAN
|
||||||
| SwissProt Accessions | O75165, Q6PI82, Q6UJ77, Q6ZSW1, Q6ZUT5, Q86XG3, Q96DC1, Q9BWK9 | Gene names | DNAJC13, KIAA0678, RME8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 13 (Required for receptor-mediated endocytosis 8). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
DNJCD_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | O75165, Q6PI82, Q6UJ77, Q6ZSW1, Q6ZUT5, Q86XG3, Q96DC1, Q9BWK9 | Gene names | DNAJC13, KIAA0678, RME8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 13 (Required for receptor-mediated endocytosis 8). | |||||
|
DNJA2_HUMAN
|
||||||
| θ value | 5.81887e-07 (rank : 2) | NC score | 0.525688 (rank : 9) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O60884, O14711 | Gene names | DNAJA2, CPR3, HIRIP4 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily A member 2 (HIRA-interacting protein 4) (Cell cycle progression restoration gene 3 protein) (Dnj3) (NY-REN-14 antigen). | |||||
|
DNJA2_MOUSE
|
||||||
| θ value | 5.81887e-07 (rank : 3) | NC score | 0.525898 (rank : 8) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYJ0 | Gene names | Dnaja2 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily A member 2 (mDj3). | |||||
|
DNJA3_MOUSE
|
||||||
| θ value | 3.77169e-06 (rank : 4) | NC score | 0.527171 (rank : 5) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q99M87, Q8BSM0, Q99L09, Q99P71, Q99P76, Q9CT11, Q9DBJ7, Q9DC44 | Gene names | Dnaja3, Tid1 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily A member 3, mitochondrial precursor (Tumorous imaginal discs protein Tid56 homolog) (DnaJ protein Tid-1) (mTid-1). | |||||
|
DNJB7_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 5) | NC score | 0.527048 (rank : 6) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q7Z6W7, Q5H904, Q8WYJ7 | Gene names | DNAJB7, HSC3 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 7. | |||||
|
DNJB6_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 6) | NC score | 0.529033 (rank : 4) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O75190, O95806, Q9UIK6 | Gene names | DNAJB6, HSJ2, MSJ1 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 6 (Heat shock protein J2) (HSJ-2) (MSJ-1) (HHDJ1) (MRJ). | |||||
|
DNJB5_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 7) | NC score | 0.508352 (rank : 24) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O75953, Q8TDR7 | Gene names | DNAJB5, HSC40 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 5 (Heat shock protein Hsp40-3) (Heat shock protein cognate 40) (Hsc40) (Hsp40-2). | |||||
|
DNJB5_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 8) | NC score | 0.508640 (rank : 23) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O89114 | Gene names | Dnajb5, Hsc40 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 5 (Heat shock protein Hsp40-3) (Heat shock protein cognate 40) (Hsc40). | |||||
|
DNJA3_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 9) | NC score | 0.521725 (rank : 14) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96EY1, O75472, Q8WUJ6, Q8WXJ3, Q96D76, Q96IV1, Q9NYH8 | Gene names | DNAJA3, TID1 | |||
|
Domain Architecture |
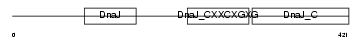
|
|||||
| Description | DnaJ homolog subfamily A member 3, mitochondrial precursor (Tumorous imaginal discs protein Tid56 homolog) (DnaJ protein Tid-1) (hTid-1). | |||||
|
DNJA4_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 10) | NC score | 0.518063 (rank : 17) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8WW22, Q8N7P2 | Gene names | DNAJA4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 4. | |||||
|
DNJA4_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 11) | NC score | 0.517706 (rank : 18) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9JMC3, Q543S9, Q9CTD6, Q9CUD4 | Gene names | Dnaja4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 4 (MmDjA4). | |||||
|
DNJBA_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 12) | NC score | 0.536380 (rank : 2) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYI5 | Gene names | Dnajb10 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 10 (mDJ8). | |||||
|
DNJB8_MOUSE
|
||||||
| θ value | 3.19293e-05 (rank : 13) | NC score | 0.526709 (rank : 7) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYI7 | Gene names | Dnajb8 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 8 (mDJ6). | |||||
|
DNJB3_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 14) | NC score | 0.525292 (rank : 11) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O35723, Q9DAN3, Q9DAN4 | Gene names | Dnajb3, Hsj3, Msj1 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 3 (DnaJ protein homolog 3) (Heat shock J3 protein) (HSJ-3) (MSJ-1). | |||||
|
DNJB6_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 15) | NC score | 0.525365 (rank : 10) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O54946, Q99LA5, Q9QYI9 | Gene names | Dnajb6, Hsj2, Mrj | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 6 (Heat shock protein J2) (HSJ-2) (MRJ) (mDj4). | |||||
|
DNJB7_MOUSE
|
||||||
| θ value | 5.44631e-05 (rank : 16) | NC score | 0.521888 (rank : 13) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYI8, Q9DA41 | Gene names | Dnajb7 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 7 (mDJ5). | |||||
|
DNJA1_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 17) | NC score | 0.511878 (rank : 21) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P63037, P54102 | Gene names | Dnaja1, Dnaj2, Hsj2, Hspf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily A member 1 (Heat shock 40 kDa protein 4) (DnaJ protein homolog 2) (HSJ-2). | |||||
|
DNJB8_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 18) | NC score | 0.524035 (rank : 12) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8NHS0 | Gene names | DNAJB8 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 8. | |||||
|
DNJA1_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 19) | NC score | 0.511622 (rank : 22) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P31689 | Gene names | DNAJA1, DNAJ2, HDJ2, HSJ2, HSPF4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 1 (Heat shock 40 kDa protein 4) (DnaJ protein homolog 2) (HSJ-2) (HSDJ). | |||||
|
DNJB2_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 20) | NC score | 0.529057 (rank : 3) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P25686, Q8IUK1, Q8IUK2, Q96F52 | Gene names | DNAJB2, HSJ1, HSPF3 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 2 (Heat shock 40 kDa protein 3) (DnaJ protein homolog 1) (HSJ-1). | |||||
|
DNJB9_MOUSE
|
||||||
| θ value | 0.000121331 (rank : 21) | NC score | 0.519059 (rank : 15) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9QYI6, Q9DAW1 | Gene names | Dnajb9 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 9 (mDJ7). | |||||
|
DNJB9_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 22) | NC score | 0.518440 (rank : 16) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UBS3 | Gene names | DNAJB9, MDG1 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 9 (Microvascular endothelial differentiation gene 1 protein) (Mdg-1). | |||||
|
DJC16_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 23) | NC score | 0.477866 (rank : 36) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9Y2G8, Q68D57, Q86X32, Q8N5P4 | Gene names | DNAJC16, KIAA0962 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 16 precursor. | |||||
|
DJC16_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 24) | NC score | 0.476152 (rank : 37) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q80TN4, Q811G1, Q8BHI2 | Gene names | Dnajc16, Kiaa0962 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 16 precursor. | |||||
|
DNJC5_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 25) | NC score | 0.512817 (rank : 19) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9H3Z4, Q9H3Z5, Q9H7H2 | Gene names | DNAJC5, CSP | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 5 (Cysteine string protein) (CSP). | |||||
|
DNJC5_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 26) | NC score | 0.512730 (rank : 20) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P60904, P54101 | Gene names | Dnajc5, Csp | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 5 (Cysteine string protein) (CSP). | |||||
|
DNJB4_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 27) | NC score | 0.495440 (rank : 30) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9D832, Q3TS92, Q9D9U2 | Gene names | Dnajb4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 4. | |||||
|
DNJBC_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 28) | NC score | 0.486088 (rank : 34) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NXW2, Q9H6H0 | Gene names | DNAJB12 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 12. | |||||
|
DNJB4_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 29) | NC score | 0.493739 (rank : 31) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UDY4, Q13431 | Gene names | DNAJB4, DNAJW, HLJ1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 4 (Heat shock 40 kDa protein 1 homolog) (Heat shock protein 40 homolog) (HSP40 homolog). | |||||
|
DNJ5B_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 30) | NC score | 0.504292 (rank : 25) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9UF47, Q969Y8 | Gene names | DNAJC5B | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 5B (Beta cysteine string protein) (Beta-CSP). | |||||
|
DNJB1_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 31) | NC score | 0.492390 (rank : 33) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P25685 | Gene names | DNAJB1, DNAJ1, HDJ1, HSPF1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 1 (Heat shock 40 kDa protein 1) (Heat shock protein 40) (HSP40) (DnaJ protein homolog 1) (HDJ-1). | |||||
|
DNJB1_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 32) | NC score | 0.492479 (rank : 32) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9QYJ3 | Gene names | Dnajb1, Hsp40, Hspf1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 1 (Heat shock 40 kDa protein 1) (Heat shock protein 40) (HSP40). | |||||
|
DNJ5B_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 33) | NC score | 0.501534 (rank : 26) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9CQ94 | Gene names | Dnajc5b | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 5B (Beta cysteine string protein) (Beta-CSP). | |||||
|
DNJBB_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 34) | NC score | 0.497364 (rank : 28) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UBS4, Q96JC6 | Gene names | DNAJB11, EDJ, ERJ3 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 11 precursor (ER-associated dnaJ protein 3) (ErJ3) (ER-associated Hsp40 co-chaperone) (hDj9) (PWP1- interacting protein 4). | |||||
|
DNJBB_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 35) | NC score | 0.497205 (rank : 29) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q99KV1, Q543I7 | Gene names | Dnajb11 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 11 precursor. | |||||
|
DNJBC_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 36) | NC score | 0.481722 (rank : 35) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYI4 | Gene names | Dnajb12 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily B member 12 (mDJ10). | |||||
|
SYMPK_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 37) | NC score | 0.119356 (rank : 74) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92797, O00521, O00689, O00733, Q59GT5, Q8N2U5 | Gene names | SYMPK, SPK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Symplekin. | |||||
|
SEC63_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 38) | NC score | 0.337309 (rank : 59) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 39) | NC score | 0.336687 (rank : 62) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
DJC18_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 40) | NC score | 0.465222 (rank : 41) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9H819 | Gene names | DNAJC18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 18. | |||||
|
SYMPK_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 41) | NC score | 0.116144 (rank : 75) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80X82 | Gene names | Sympk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Symplekin. | |||||
|
WBS18_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 42) | NC score | 0.473363 (rank : 39) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P59041 | Gene names | Wbscr18 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 18 protein homolog. | |||||
|
DNJ5G_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 43) | NC score | 0.500689 (rank : 27) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8N7S2, Q8IYQ4, Q96RJ8 | Gene names | DNAJC5G | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 5G (Gamma cysteine string protein) (Gamma-CSP). | |||||
|
DNJBD_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 44) | NC score | 0.474193 (rank : 38) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P59910, Q8IZW5 | Gene names | DNAJB13, TSARG3, TSARG6 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 13 (Testis spermatocyte apoptosis- related gene 6 protein) (Testis and spermatogenesis cell-related protein 6) (Testis spermatogenesis apoptosis-related gene 6 protein) (Testis spermatogenesis apoptosis-related gene 3 protein). | |||||
|
CU055_MOUSE
|
||||||
| θ value | 0.125558 (rank : 45) | NC score | 0.338111 (rank : 57) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8VCE1, Q99JX0 | Gene names | ORF28 | |||
|
Domain Architecture |
|
|||||
| Description | J domain-containing protein C21orf55 homolog. | |||||
|
DNJC8_HUMAN
|
||||||
| θ value | 0.125558 (rank : 46) | NC score | 0.338535 (rank : 56) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O75937, Q6IBA4, Q8N4Z5, Q9P051, Q9P067 | Gene names | DNAJC8, SPF31 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 8 (Splicing protein spf31). | |||||
|
DJC18_MOUSE
|
||||||
| θ value | 0.163984 (rank : 47) | NC score | 0.449633 (rank : 42) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9CZJ9, Q8R371 | Gene names | Dnajc18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 18. | |||||
|
SACS_HUMAN
|
||||||
| θ value | 0.279714 (rank : 48) | NC score | 0.138281 (rank : 72) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9NZJ4, O94835 | Gene names | SACS, KIAA0730 | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
DJC17_HUMAN
|
||||||
| θ value | 0.365318 (rank : 49) | NC score | 0.403505 (rank : 49) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9NVM6 | Gene names | DNAJC17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 17. | |||||
|
DJC17_MOUSE
|
||||||
| θ value | 0.365318 (rank : 50) | NC score | 0.417803 (rank : 44) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q91WT4, Q8C5R1 | Gene names | Dnajc17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 17. | |||||
|
DNJBD_MOUSE
|
||||||
| θ value | 0.365318 (rank : 51) | NC score | 0.468725 (rank : 40) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q80Y75, Q8CJA2 | Gene names | Dnajb13, Tsarg, Tsarg3, Tsarg6 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 13 (Testis spermatocyte apoptosis- related gene 6 protein) (Testis and spermatogenesis cell-related protein 6) (Testis spermatogenesis apoptosis-related gene 6 protein) (Testis spermatogenesis apoptosis-related gene 3 protein). | |||||
|
DNJC8_MOUSE
|
||||||
| θ value | 0.365318 (rank : 52) | NC score | 0.330984 (rank : 64) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q6NZB0, Q6PG72, Q8C2M6 | Gene names | Dnajc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 8. | |||||
|
SACS_MOUSE
|
||||||
| θ value | 0.365318 (rank : 53) | NC score | 0.137978 (rank : 73) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9JLC8 | Gene names | Sacs | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
CU055_HUMAN
|
||||||
| θ value | 0.62314 (rank : 54) | NC score | 0.301489 (rank : 69) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9NX36 | Gene names | C21orf55 | |||
|
Domain Architecture |
|
|||||
| Description | J domain-containing protein C21orf55. | |||||
|
METH_HUMAN
|
||||||
| θ value | 0.62314 (rank : 55) | NC score | 0.061528 (rank : 79) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99707, Q99713, Q99723 | Gene names | MTR | |||
|
Domain Architecture |

|
|||||
| Description | Methionine synthase (EC 2.1.1.13) (5-methyltetrahydrofolate-- homocysteine methyltransferase) (Methionine synthase, vitamin-B12 dependent) (MS). | |||||
|
WBS18_HUMAN
|
||||||
| θ value | 0.62314 (rank : 56) | NC score | 0.448002 (rank : 43) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96LL9, Q9BSG8 | Gene names | WBSCR18 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 18 protein. | |||||
|
DCJ11_HUMAN
|
||||||
| θ value | 0.813845 (rank : 57) | NC score | 0.408649 (rank : 48) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9NVH1, Q4VWF5, Q5VZN0, Q6PK20, Q6PK70, Q8NDM2, Q96CL7 | Gene names | DNAJC11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 11. | |||||
|
DCJ11_MOUSE
|
||||||
| θ value | 0.813845 (rank : 58) | NC score | 0.408661 (rank : 47) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q5U458, Q8BP83, Q8C1Z4, Q8C6U5 | Gene names | Dnajc11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 11. | |||||
|
ZRF1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 59) | NC score | 0.253361 (rank : 70) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q99543 | Gene names | ZRF1, DNAJC2, MPHOSPH11, MPP11 | |||
|
Domain Architecture |

|
|||||
| Description | Zuotin-related factor 1 (M-phase phosphoprotein 11). | |||||
|
ZRF1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 60) | NC score | 0.246909 (rank : 71) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P54103 | Gene names | Zrf1, Dnajc2 | |||
|
Domain Architecture |
|
|||||
| Description | Zuotin-related factor 1. | |||||
|
DNJC4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.415790 (rank : 45) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9D844, O70278 | Gene names | Dnajc4, Mcg18 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 4 (Multiple endocrine neoplasia type 1 candidate protein number 18 homolog). | |||||
|
AP3D1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.033670 (rank : 86) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14617, O00202, O75262, Q59HF5, Q96G11, Q9H3C6 | Gene names | AP3D1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit delta-1 (Adapter-related protein complex 3 subunit delta-1) (Delta-adaptin 3) (AP-3 complex subunit delta) (Delta-adaptin). | |||||
|
AP3D1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.034815 (rank : 84) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O54774 | Gene names | Ap3d1, Ap3d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit delta-1 (Adapter-related protein complex 3 subunit delta-1) (Delta-adaptin 3) (AP-3 complex subunit delta) (Delta-adaptin) (mBLVR1). | |||||
|
CO029_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.040088 (rank : 83) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H079, Q2TAC0, Q9H670 | Gene names | C15orf29 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf29. | |||||
|
COG3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.033959 (rank : 85) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CI04 | Gene names | Cog3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Conserved oligomeric Golgi complex component 3. | |||||
|
MELK_MOUSE
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.002960 (rank : 98) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61846, Q61804, Q6ZQH6 | Gene names | Melk, Kiaa0175, Pk38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Maternal embryonic leucine zipper kinase (EC 2.7.11.1) (Protein kinase PK38) (mPK38). | |||||
|
NDUAA_MOUSE
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.040155 (rank : 82) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99LC3, Q8BL57, Q9CW21 | Gene names | Ndufa10 | |||
|
Domain Architecture |
|
|||||
| Description | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 10, mitochondrial precursor (EC 1.6.5.3) (EC 1.6.99.3) (NADH-ubiquinone oxidoreductase 42 kDa subunit) (Complex I-42kD) (CI-42kD). | |||||
|
SRPK2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 68) | NC score | 0.017831 (rank : 91) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 664 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78362, O75220, O75221, Q6NUL0, Q6V1X2, Q8IYQ3 | Gene names | SRPK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK2 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 2) (SR-protein-specific kinase 2) (SFRS protein kinase 2). | |||||
|
SRPK2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 69) | NC score | 0.018064 (rank : 90) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O54781, Q8VCD9 | Gene names | Srpk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK2 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 2) (SR-protein-specific kinase 2) (SFRS protein kinase 2). | |||||
|
MELK_HUMAN
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.002839 (rank : 99) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q14680, Q7L3C3 | Gene names | MELK, KIAA0175 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Maternal embryonic leucine zipper kinase (EC 2.7.11.1) (hMELK) (Protein kinase PK38) (hPK38). | |||||
|
MET_HUMAN
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.002318 (rank : 100) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P08581, O60366, Q12875, Q9UDX7, Q9UPL8 | Gene names | MET | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor receptor precursor (EC 2.7.10.1) (HGF receptor) (Scatter factor receptor) (SF receptor) (HGF/SF receptor) (Met proto-oncogene tyrosine kinase) (c-Met). | |||||
|
ARLY_MOUSE
|
||||||
| θ value | 3.0926 (rank : 72) | NC score | 0.030912 (rank : 87) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91YI0, Q3UFA2 | Gene names | Asl | |||
|
Domain Architecture |
|
|||||
| Description | Argininosuccinate lyase (EC 4.3.2.1) (Arginosuccinase) (ASAL). | |||||
|
CCNB3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.015926 (rank : 93) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WWL7, Q96SB5, Q96SB6, Q96SB7, Q9NT38 | Gene names | CCNB3, CYCB3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
DNJC1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.392347 (rank : 50) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q61712 | Gene names | Dnajc1, Dnajl1, Mtj1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1 (DnaJ protein homolog MTJ1). | |||||
|
IFIT2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 75) | NC score | 0.020341 (rank : 89) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09913 | Gene names | IFIT2, G10P2, IFI54 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced protein with tetratricopeptide repeats 2 (IFIT-2) (Interferon-induced 54 kDa protein) (IFI-54K) (ISG-54 K). | |||||
|
IQGA2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.016676 (rank : 92) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
KIFA3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.025046 (rank : 88) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92845, Q8NHU7, Q9H416 | Gene names | KIFAP3, SMAP | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-associated protein 3 (Smg GDS-associated protein). | |||||
|
DJC12_MOUSE
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.337105 (rank : 61) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9R022 | Gene names | Dnajc12, Jdp1 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 12 (J domain-containing protein 1). | |||||
|
DNJC4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.413342 (rank : 46) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9NNZ3, O14716 | Gene names | DNAJC4, HSPF2, MCG18 | |||
|
Domain Architecture |
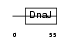
|
|||||
| Description | DnaJ homolog subfamily C member 4 (Multiple endocrine neoplasia type 1 candidate protein number 18) (DnaJ-like protein HSPF2). | |||||
|
DCJ14_MOUSE
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.344657 (rank : 55) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q921R4, Q3TX73, Q8BUU3, Q9CYB7 | Gene names | Dnajc14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 14. | |||||
|
DNJC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.386439 (rank : 51) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96KC8 | Gene names | DNAJC1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1. | |||||
|
AP1G2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.015074 (rank : 94) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88512 | Gene names | Ap1g2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-2 (Adapter-related protein complex 1 gamma- 2 subunit) (Gamma2-adaptin) (Adaptor protein complex AP-1 gamma-2 subunit) (G2ad). | |||||
|
BBS10_HUMAN
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.014253 (rank : 95) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TAM1, Q96CW2, Q9H5D2 | Gene names | BBS10, C12orf58 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bardet-Biedl syndrome 10 protein. | |||||
|
DNJC7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.315538 (rank : 67) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q99615 | Gene names | DNAJC7, TPR2, TTC2 | |||
|
Domain Architecture |
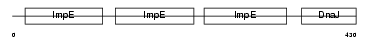
|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2). | |||||
|
DNJC7_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.312093 (rank : 68) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9QYI3, Q3TKR1, Q8BPG3, Q8CIL2, Q9CT29, Q9D026 | Gene names | Dnajc7, Ttc2 | |||
|
Domain Architecture |
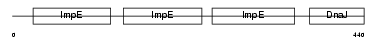
|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2) (MDj11). | |||||
|
MTSS1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.011205 (rank : 96) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R1S4, Q8BMM3, Q99LB3 | Gene names | Mtss1, Mim | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein). | |||||
|
RAG1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.009994 (rank : 97) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15918, Q8NER2 | Gene names | RAG1, RNF74 | |||
|
Domain Architecture |

|
|||||
| Description | V(D)J recombination-activating protein 1 (RAG-1) (RING finger protein 74). | |||||
|
AUXI_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.053262 (rank : 81) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O75061, Q32M66, Q4G0K1, Q5T614, Q5T615 | Gene names | DNAJC6, KIAA0473 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative tyrosine-protein phosphatase auxilin (EC 3.1.3.48) (DnaJ homolog subfamily C member 6). | |||||
|
DCJ14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.337881 (rank : 58) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q6Y2X3, Q66K17, Q96N59, Q96T63 | Gene names | DNAJC14, DRIP78 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 14 (Dopamine receptor-interacting protein of 78 kDa) (DRiP78). | |||||
|
DCJ15_HUMAN
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.067156 (rank : 78) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y5T4, Q5T219, Q6X963 | Gene names | DNAJC15, DNAJD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 15 (Methylation-controlled J protein) (MCJ). | |||||
|
DCJ15_MOUSE
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.056944 (rank : 80) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q78YY6 | Gene names | Dnajc15, Dnajd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 15. | |||||
|
DJC12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.337304 (rank : 60) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9UKB3, Q9UKB2 | Gene names | DNAJC12, JDP1 | |||
|
Domain Architecture |
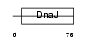
|
|||||
| Description | DnaJ homolog subfamily C member 12 (J domain-containing protein 1). | |||||
|
DNJC3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.317355 (rank : 65) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q13217, Q86WT9, Q8N4N2 | Gene names | DNAJC3, P58IPK, PRKRI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 3 (Interferon-induced, double-stranded RNA-activated protein kinase inhibitor) (Protein kinase inhibitor p58) (Protein kinase inhibitor of 58 kDa). | |||||
|
DNJC3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.316119 (rank : 66) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q91YW3, Q60873 | Gene names | Dnajc3, P58ipk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 3 (Interferon-induced, double-stranded RNA-activated protein kinase inhibitor) (Protein kinase inhibitor p58) (Protein kinase inhibitor of 58 kDa). | |||||
|
DNJC9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.375021 (rank : 52) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q8WXX5 | Gene names | DNAJC9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 9 (DnaJ protein SB73). | |||||
|
DNJC9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.368056 (rank : 53) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q91WN1, Q3TSG3, Q8R0E3 | Gene names | Dnajc9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 9. | |||||
|
TIM14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.072504 (rank : 76) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96DA6 | Gene names | DNAJC19, TIM14, TIMM14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial import inner membrane translocase subunit TIM14 (DnaJ homolog subfamily C member 19). | |||||
|
TIM14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.071959 (rank : 77) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9CQV7, Q8R1N1, Q9D896 | Gene names | Dnajc19, Tim14, Timm14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial import inner membrane translocase subunit TIM14 (DnaJ homolog subfamily C member 19). | |||||
|
ZCSL3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.353860 (rank : 54) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q6P3W2 | Gene names | ZCSL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CSL-type zinc finger-containing protein 3. | |||||
|
ZCSL3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.335853 (rank : 63) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q91ZF0, Q9D9S7 | Gene names | Zcsl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CSL-type zinc finger-containing protein 3 (J-domain protein DjC7). | |||||
|
DNJCD_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | O75165, Q6PI82, Q6UJ77, Q6ZSW1, Q6ZUT5, Q86XG3, Q96DC1, Q9BWK9 | Gene names | DNAJC13, KIAA0678, RME8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 13 (Required for receptor-mediated endocytosis 8). | |||||
|
DNJBA_MOUSE
|
||||||
| NC score | 0.536380 (rank : 2) | θ value | 1.87187e-05 (rank : 12) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYI5 | Gene names | Dnajb10 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 10 (mDJ8). | |||||
|
DNJB2_HUMAN
|
||||||
| NC score | 0.529057 (rank : 3) | θ value | 9.29e-05 (rank : 20) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P25686, Q8IUK1, Q8IUK2, Q96F52 | Gene names | DNAJB2, HSJ1, HSPF3 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 2 (Heat shock 40 kDa protein 3) (DnaJ protein homolog 1) (HSJ-1). | |||||
|
DNJB6_HUMAN
|
||||||
| NC score | 0.529033 (rank : 4) | θ value | 8.40245e-06 (rank : 6) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O75190, O95806, Q9UIK6 | Gene names | DNAJB6, HSJ2, MSJ1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 6 (Heat shock protein J2) (HSJ-2) (MSJ-1) (HHDJ1) (MRJ). | |||||
|
DNJA3_MOUSE
|
||||||
| NC score | 0.527171 (rank : 5) | θ value | 3.77169e-06 (rank : 4) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q99M87, Q8BSM0, Q99L09, Q99P71, Q99P76, Q9CT11, Q9DBJ7, Q9DC44 | Gene names | Dnaja3, Tid1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 3, mitochondrial precursor (Tumorous imaginal discs protein Tid56 homolog) (DnaJ protein Tid-1) (mTid-1). | |||||
|
DNJB7_HUMAN
|
||||||
| NC score | 0.527048 (rank : 6) | θ value | 6.43352e-06 (rank : 5) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q7Z6W7, Q5H904, Q8WYJ7 | Gene names | DNAJB7, HSC3 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 7. | |||||
|
DNJB8_MOUSE
|
||||||
| NC score | 0.526709 (rank : 7) | θ value | 3.19293e-05 (rank : 13) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYI7 | Gene names | Dnajb8 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 8 (mDJ6). | |||||
|
DNJA2_MOUSE
|
||||||
| NC score | 0.525898 (rank : 8) | θ value | 5.81887e-07 (rank : 3) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYJ0 | Gene names | Dnaja2 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily A member 2 (mDj3). | |||||
|
DNJA2_HUMAN
|
||||||
| NC score | 0.525688 (rank : 9) | θ value | 5.81887e-07 (rank : 2) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O60884, O14711 | Gene names | DNAJA2, CPR3, HIRIP4 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily A member 2 (HIRA-interacting protein 4) (Cell cycle progression restoration gene 3 protein) (Dnj3) (NY-REN-14 antigen). | |||||
|
DNJB6_MOUSE
|
||||||
| NC score | 0.525365 (rank : 10) | θ value | 4.1701e-05 (rank : 15) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O54946, Q99LA5, Q9QYI9 | Gene names | Dnajb6, Hsj2, Mrj | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 6 (Heat shock protein J2) (HSJ-2) (MRJ) (mDj4). | |||||
|
DNJB3_MOUSE
|
||||||
| NC score | 0.525292 (rank : 11) | θ value | 4.1701e-05 (rank : 14) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O35723, Q9DAN3, Q9DAN4 | Gene names | Dnajb3, Hsj3, Msj1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 3 (DnaJ protein homolog 3) (Heat shock J3 protein) (HSJ-3) (MSJ-1). | |||||
|
DNJB8_HUMAN
|
||||||
| NC score | 0.524035 (rank : 12) | θ value | 7.1131e-05 (rank : 18) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8NHS0 | Gene names | DNAJB8 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 8. | |||||
|
DNJB7_MOUSE
|
||||||
| NC score | 0.521888 (rank : 13) | θ value | 5.44631e-05 (rank : 16) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYI8, Q9DA41 | Gene names | Dnajb7 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 7 (mDJ5). | |||||
|
DNJA3_HUMAN
|
||||||
| NC score | 0.521725 (rank : 14) | θ value | 1.43324e-05 (rank : 9) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96EY1, O75472, Q8WUJ6, Q8WXJ3, Q96D76, Q96IV1, Q9NYH8 | Gene names | DNAJA3, TID1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 3, mitochondrial precursor (Tumorous imaginal discs protein Tid56 homolog) (DnaJ protein Tid-1) (hTid-1). | |||||
|
DNJB9_MOUSE
|
||||||
| NC score | 0.519059 (rank : 15) | θ value | 0.000121331 (rank : 21) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9QYI6, Q9DAW1 | Gene names | Dnajb9 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 9 (mDJ7). | |||||
|
DNJB9_HUMAN
|
||||||
| NC score | 0.518440 (rank : 16) | θ value | 0.000158464 (rank : 22) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UBS3 | Gene names | DNAJB9, MDG1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 9 (Microvascular endothelial differentiation gene 1 protein) (Mdg-1). | |||||
|
DNJA4_HUMAN
|
||||||
| NC score | 0.518063 (rank : 17) | θ value | 1.43324e-05 (rank : 10) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8WW22, Q8N7P2 | Gene names | DNAJA4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 4. | |||||
|
DNJA4_MOUSE
|
||||||
| NC score | 0.517706 (rank : 18) | θ value | 1.43324e-05 (rank : 11) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9JMC3, Q543S9, Q9CTD6, Q9CUD4 | Gene names | Dnaja4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 4 (MmDjA4). | |||||
|
DNJC5_HUMAN
|
||||||
| NC score | 0.512817 (rank : 19) | θ value | 0.00035302 (rank : 25) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9H3Z4, Q9H3Z5, Q9H7H2 | Gene names | DNAJC5, CSP | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 5 (Cysteine string protein) (CSP). | |||||
|
DNJC5_MOUSE
|
||||||
| NC score | 0.512730 (rank : 20) | θ value | 0.00035302 (rank : 26) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P60904, P54101 | Gene names | Dnajc5, Csp | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 5 (Cysteine string protein) (CSP). | |||||
|
DNJA1_MOUSE
|
||||||
| NC score | 0.511878 (rank : 21) | θ value | 7.1131e-05 (rank : 17) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P63037, P54102 | Gene names | Dnaja1, Dnaj2, Hsj2, Hspf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily A member 1 (Heat shock 40 kDa protein 4) (DnaJ protein homolog 2) (HSJ-2). | |||||
|
DNJA1_HUMAN
|
||||||
| NC score | 0.511622 (rank : 22) | θ value | 9.29e-05 (rank : 19) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P31689 | Gene names | DNAJA1, DNAJ2, HDJ2, HSJ2, HSPF4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 1 (Heat shock 40 kDa protein 4) (DnaJ protein homolog 2) (HSJ-2) (HSDJ). | |||||
|
DNJB5_MOUSE
|
||||||
| NC score | 0.508640 (rank : 23) | θ value | 1.09739e-05 (rank : 8) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O89114 | Gene names | Dnajb5, Hsc40 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 5 (Heat shock protein Hsp40-3) (Heat shock protein cognate 40) (Hsc40). | |||||
|
DNJB5_HUMAN
|
||||||
| NC score | 0.508352 (rank : 24) | θ value | 1.09739e-05 (rank : 7) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O75953, Q8TDR7 | Gene names | DNAJB5, HSC40 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 5 (Heat shock protein Hsp40-3) (Heat shock protein cognate 40) (Hsc40) (Hsp40-2). | |||||
|
DNJ5B_HUMAN
|
||||||
| NC score | 0.504292 (rank : 25) | θ value | 0.00134147 (rank : 30) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9UF47, Q969Y8 | Gene names | DNAJC5B | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 5B (Beta cysteine string protein) (Beta-CSP). | |||||
|
DNJ5B_MOUSE
|
||||||
| NC score | 0.501534 (rank : 26) | θ value | 0.00298849 (rank : 33) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9CQ94 | Gene names | Dnajc5b | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 5B (Beta cysteine string protein) (Beta-CSP). | |||||
|
DNJ5G_HUMAN
|
||||||
| NC score | 0.500689 (rank : 27) | θ value | 0.0330416 (rank : 43) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8N7S2, Q8IYQ4, Q96RJ8 | Gene names | DNAJC5G | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 5G (Gamma cysteine string protein) (Gamma-CSP). | |||||
|
DNJBB_HUMAN
|
||||||
| NC score | 0.497364 (rank : 28) | θ value | 0.00390308 (rank : 34) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UBS4, Q96JC6 | Gene names | DNAJB11, EDJ, ERJ3 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 11 precursor (ER-associated dnaJ protein 3) (ErJ3) (ER-associated Hsp40 co-chaperone) (hDj9) (PWP1- interacting protein 4). | |||||
|
DNJBB_MOUSE
|
||||||
| NC score | 0.497205 (rank : 29) | θ value | 0.00390308 (rank : 35) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q99KV1, Q543I7 | Gene names | Dnajb11 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 11 precursor. | |||||
|
DNJB4_MOUSE
|
||||||
| NC score | 0.495440 (rank : 30) | θ value | 0.000602161 (rank : 27) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9D832, Q3TS92, Q9D9U2 | Gene names | Dnajb4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 4. | |||||
|
DNJB4_HUMAN
|
||||||
| NC score | 0.493739 (rank : 31) | θ value | 0.000786445 (rank : 29) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UDY4, Q13431 | Gene names | DNAJB4, DNAJW, HLJ1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 4 (Heat shock 40 kDa protein 1 homolog) (Heat shock protein 40 homolog) (HSP40 homolog). | |||||
|
DNJB1_MOUSE
|
||||||
| NC score | 0.492479 (rank : 32) | θ value | 0.00228821 (rank : 32) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9QYJ3 | Gene names | Dnajb1, Hsp40, Hspf1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 1 (Heat shock 40 kDa protein 1) (Heat shock protein 40) (HSP40). | |||||
|
DNJB1_HUMAN
|
||||||
| NC score | 0.492390 (rank : 33) | θ value | 0.00228821 (rank : 31) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P25685 | Gene names | DNAJB1, DNAJ1, HDJ1, HSPF1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 1 (Heat shock 40 kDa protein 1) (Heat shock protein 40) (HSP40) (DnaJ protein homolog 1) (HDJ-1). | |||||
|
DNJBC_HUMAN
|
||||||
| NC score | 0.486088 (rank : 34) | θ value | 0.000602161 (rank : 28) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NXW2, Q9H6H0 | Gene names | DNAJB12 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 12. | |||||
|
DNJBC_MOUSE
|
||||||
| NC score | 0.481722 (rank : 35) | θ value | 0.00665767 (rank : 36) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYI4 | Gene names | Dnajb12 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 12 (mDJ10). | |||||
|
DJC16_HUMAN
|
||||||
| NC score | 0.477866 (rank : 36) | θ value | 0.00035302 (rank : 23) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9Y2G8, Q68D57, Q86X32, Q8N5P4 | Gene names | DNAJC16, KIAA0962 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 16 precursor. | |||||
|
DJC16_MOUSE
|
||||||
| NC score | 0.476152 (rank : 37) | θ value | 0.00035302 (rank : 24) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q80TN4, Q811G1, Q8BHI2 | Gene names | Dnajc16, Kiaa0962 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 16 precursor. | |||||
|
DNJBD_HUMAN
|
||||||
| NC score | 0.474193 (rank : 38) | θ value | 0.0563607 (rank : 44) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P59910, Q8IZW5 | Gene names | DNAJB13, TSARG3, TSARG6 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 13 (Testis spermatocyte apoptosis- related gene 6 protein) (Testis and spermatogenesis cell-related protein 6) (Testis spermatogenesis apoptosis-related gene 6 protein) (Testis spermatogenesis apoptosis-related gene 3 protein). | |||||
|
WBS18_MOUSE
|
||||||
| NC score | 0.473363 (rank : 39) | θ value | 0.0252991 (rank : 42) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P59041 | Gene names | Wbscr18 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 18 protein homolog. | |||||
|
DNJBD_MOUSE
|
||||||
| NC score | 0.468725 (rank : 40) | θ value | 0.365318 (rank : 51) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q80Y75, Q8CJA2 | Gene names | Dnajb13, Tsarg, Tsarg3, Tsarg6 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily B member 13 (Testis spermatocyte apoptosis- related gene 6 protein) (Testis and spermatogenesis cell-related protein 6) (Testis spermatogenesis apoptosis-related gene 6 protein) (Testis spermatogenesis apoptosis-related gene 3 protein). | |||||
|
DJC18_HUMAN
|
||||||
| NC score | 0.465222 (rank : 41) | θ value | 0.0252991 (rank : 40) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9H819 | Gene names | DNAJC18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 18. | |||||
|
DJC18_MOUSE
|
||||||
| NC score | 0.449633 (rank : 42) | θ value | 0.163984 (rank : 47) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9CZJ9, Q8R371 | Gene names | Dnajc18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 18. | |||||
|
WBS18_HUMAN
|
||||||
| NC score | 0.448002 (rank : 43) | θ value | 0.62314 (rank : 56) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96LL9, Q9BSG8 | Gene names | WBSCR18 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 18 protein. | |||||
|
DJC17_MOUSE
|
||||||
| NC score | 0.417803 (rank : 44) | θ value | 0.365318 (rank : 50) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q91WT4, Q8C5R1 | Gene names | Dnajc17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 17. | |||||
|
DNJC4_MOUSE
|
||||||
| NC score | 0.415790 (rank : 45) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9D844, O70278 | Gene names | Dnajc4, Mcg18 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 4 (Multiple endocrine neoplasia type 1 candidate protein number 18 homolog). | |||||
|
DNJC4_HUMAN
|
||||||
| NC score | 0.413342 (rank : 46) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9NNZ3, O14716 | Gene names | DNAJC4, HSPF2, MCG18 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 4 (Multiple endocrine neoplasia type 1 candidate protein number 18) (DnaJ-like protein HSPF2). | |||||
|
DCJ11_MOUSE
|
||||||
| NC score | 0.408661 (rank : 47) | θ value | 0.813845 (rank : 58) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q5U458, Q8BP83, Q8C1Z4, Q8C6U5 | Gene names | Dnajc11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 11. | |||||
|
DCJ11_HUMAN
|
||||||
| NC score | 0.408649 (rank : 48) | θ value | 0.813845 (rank : 57) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9NVH1, Q4VWF5, Q5VZN0, Q6PK20, Q6PK70, Q8NDM2, Q96CL7 | Gene names | DNAJC11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 11. | |||||
|
DJC17_HUMAN
|
||||||
| NC score | 0.403505 (rank : 49) | θ value | 0.365318 (rank : 49) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9NVM6 | Gene names | DNAJC17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 17. | |||||
|
DNJC1_MOUSE
|
||||||
| NC score | 0.392347 (rank : 50) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q61712 | Gene names | Dnajc1, Dnajl1, Mtj1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1 (DnaJ protein homolog MTJ1). | |||||
|
DNJC1_HUMAN
|
||||||
| NC score | 0.386439 (rank : 51) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96KC8 | Gene names | DNAJC1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1. | |||||
|
DNJC9_HUMAN
|
||||||
| NC score | 0.375021 (rank : 52) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q8WXX5 | Gene names | DNAJC9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 9 (DnaJ protein SB73). | |||||
|
DNJC9_MOUSE
|
||||||
| NC score | 0.368056 (rank : 53) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q91WN1, Q3TSG3, Q8R0E3 | Gene names | Dnajc9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 9. | |||||
|
ZCSL3_HUMAN
|
||||||
| NC score | 0.353860 (rank : 54) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q6P3W2 | Gene names | ZCSL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CSL-type zinc finger-containing protein 3. | |||||
|
DCJ14_MOUSE
|
||||||
| NC score | 0.344657 (rank : 55) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q921R4, Q3TX73, Q8BUU3, Q9CYB7 | Gene names | Dnajc14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 14. | |||||
|
DNJC8_HUMAN
|
||||||
| NC score | 0.338535 (rank : 56) | θ value | 0.125558 (rank : 46) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O75937, Q6IBA4, Q8N4Z5, Q9P051, Q9P067 | Gene names | DNAJC8, SPF31 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 8 (Splicing protein spf31). | |||||
|
CU055_MOUSE
|
||||||
| NC score | 0.338111 (rank : 57) | θ value | 0.125558 (rank : 45) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8VCE1, Q99JX0 | Gene names | ORF28 | |||
|
Domain Architecture |

|
|||||
| Description | J domain-containing protein C21orf55 homolog. | |||||
|
DCJ14_HUMAN
|
||||||
| NC score | 0.337881 (rank : 58) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q6Y2X3, Q66K17, Q96N59, Q96T63 | Gene names | DNAJC14, DRIP78 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 14 (Dopamine receptor-interacting protein of 78 kDa) (DRiP78). | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.337309 (rank : 59) | θ value | 0.0148317 (rank : 38) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
DJC12_HUMAN
|
||||||
| NC score | 0.337304 (rank : 60) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9UKB3, Q9UKB2 | Gene names | DNAJC12, JDP1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 12 (J domain-containing protein 1). | |||||
|
DJC12_MOUSE
|
||||||
| NC score | 0.337105 (rank : 61) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9R022 | Gene names | Dnajc12, Jdp1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 12 (J domain-containing protein 1). | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.336687 (rank : 62) | θ value | 0.0148317 (rank : 39) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
ZCSL3_MOUSE
|
||||||
| NC score | 0.335853 (rank : 63) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q91ZF0, Q9D9S7 | Gene names | Zcsl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CSL-type zinc finger-containing protein 3 (J-domain protein DjC7). | |||||
|
DNJC8_MOUSE
|
||||||
| NC score | 0.330984 (rank : 64) | θ value | 0.365318 (rank : 52) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q6NZB0, Q6PG72, Q8C2M6 | Gene names | Dnajc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 8. | |||||
|
DNJC3_HUMAN
|
||||||
| NC score | 0.317355 (rank : 65) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q13217, Q86WT9, Q8N4N2 | Gene names | DNAJC3, P58IPK, PRKRI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 3 (Interferon-induced, double-stranded RNA-activated protein kinase inhibitor) (Protein kinase inhibitor p58) (Protein kinase inhibitor of 58 kDa). | |||||
|
DNJC3_MOUSE
|
||||||
| NC score | 0.316119 (rank : 66) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q91YW3, Q60873 | Gene names | Dnajc3, P58ipk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 3 (Interferon-induced, double-stranded RNA-activated protein kinase inhibitor) (Protein kinase inhibitor p58) (Protein kinase inhibitor of 58 kDa). | |||||
|
DNJC7_HUMAN
|
||||||
| NC score | 0.315538 (rank : 67) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q99615 | Gene names | DNAJC7, TPR2, TTC2 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2). | |||||
|
DNJC7_MOUSE
|
||||||
| NC score | 0.312093 (rank : 68) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9QYI3, Q3TKR1, Q8BPG3, Q8CIL2, Q9CT29, Q9D026 | Gene names | Dnajc7, Ttc2 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2) (MDj11). | |||||
|
CU055_HUMAN
|
||||||
| NC score | 0.301489 (rank : 69) | θ value | 0.62314 (rank : 54) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9NX36 | Gene names | C21orf55 | |||
|
Domain Architecture |

|
|||||
| Description | J domain-containing protein C21orf55. | |||||
|
ZRF1_HUMAN
|
||||||
| NC score | 0.253361 (rank : 70) | θ value | 1.06291 (rank : 59) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q99543 | Gene names | ZRF1, DNAJC2, MPHOSPH11, MPP11 | |||
|
Domain Architecture |

|
|||||
| Description | Zuotin-related factor 1 (M-phase phosphoprotein 11). | |||||
|
ZRF1_MOUSE
|
||||||
| NC score | 0.246909 (rank : 71) | θ value | 1.06291 (rank : 60) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P54103 | Gene names | Zrf1, Dnajc2 | |||
|
Domain Architecture |

|
|||||
| Description | Zuotin-related factor 1. | |||||
|
SACS_HUMAN
|
||||||
| NC score | 0.138281 (rank : 72) | θ value | 0.279714 (rank : 48) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9NZJ4, O94835 | Gene names | SACS, KIAA0730 | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
SACS_MOUSE
|
||||||
| NC score | 0.137978 (rank : 73) | θ value | 0.365318 (rank : 53) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9JLC8 | Gene names | Sacs | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
SYMPK_HUMAN
|
||||||
| NC score | 0.119356 (rank : 74) | θ value | 0.0113563 (rank : 37) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92797, O00521, O00689, O00733, Q59GT5, Q8N2U5 | Gene names | SYMPK, SPK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Symplekin. | |||||
|
SYMPK_MOUSE
|
||||||
| NC score | 0.116144 (rank : 75) | θ value | 0.0252991 (rank : 41) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80X82 | Gene names | Sympk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Symplekin. | |||||
|
TIM14_HUMAN
|
||||||
| NC score | 0.072504 (rank : 76) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96DA6 | Gene names | DNAJC19, TIM14, TIMM14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial import inner membrane translocase subunit TIM14 (DnaJ homolog subfamily C member 19). | |||||
|
TIM14_MOUSE
|
||||||
| NC score | 0.071959 (rank : 77) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9CQV7, Q8R1N1, Q9D896 | Gene names | Dnajc19, Tim14, Timm14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial import inner membrane translocase subunit TIM14 (DnaJ homolog subfamily C member 19). | |||||
|
DCJ15_HUMAN
|
||||||
| NC score | 0.067156 (rank : 78) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y5T4, Q5T219, Q6X963 | Gene names | DNAJC15, DNAJD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 15 (Methylation-controlled J protein) (MCJ). | |||||
|
METH_HUMAN
|
||||||
| NC score | 0.061528 (rank : 79) | θ value | 0.62314 (rank : 55) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99707, Q99713, Q99723 | Gene names | MTR | |||
|
Domain Architecture |

|
|||||
| Description | Methionine synthase (EC 2.1.1.13) (5-methyltetrahydrofolate-- homocysteine methyltransferase) (Methionine synthase, vitamin-B12 dependent) (MS). | |||||
|
DCJ15_MOUSE
|
||||||
| NC score | 0.056944 (rank : 80) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q78YY6 | Gene names | Dnajc15, Dnajd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 15. | |||||
|
AUXI_HUMAN
|
||||||
| NC score | 0.053262 (rank : 81) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O75061, Q32M66, Q4G0K1, Q5T614, Q5T615 | Gene names | DNAJC6, KIAA0473 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative tyrosine-protein phosphatase auxilin (EC 3.1.3.48) (DnaJ homolog subfamily C member 6). | |||||
|
NDUAA_MOUSE
|
||||||
| NC score | 0.040155 (rank : 82) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99LC3, Q8BL57, Q9CW21 | Gene names | Ndufa10 | |||
|
Domain Architecture |

|
|||||
| Description | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 10, mitochondrial precursor (EC 1.6.5.3) (EC 1.6.99.3) (NADH-ubiquinone oxidoreductase 42 kDa subunit) (Complex I-42kD) (CI-42kD). | |||||
|
CO029_HUMAN
|
||||||
| NC score | 0.040088 (rank : 83) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H079, Q2TAC0, Q9H670 | Gene names | C15orf29 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf29. | |||||
|
AP3D1_MOUSE
|
||||||
| NC score | 0.034815 (rank : 84) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O54774 | Gene names | Ap3d1, Ap3d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit delta-1 (Adapter-related protein complex 3 subunit delta-1) (Delta-adaptin 3) (AP-3 complex subunit delta) (Delta-adaptin) (mBLVR1). | |||||
|
COG3_MOUSE
|
||||||
| NC score | 0.033959 (rank : 85) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CI04 | Gene names | Cog3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Conserved oligomeric Golgi complex component 3. | |||||
|
AP3D1_HUMAN
|
||||||
| NC score | 0.033670 (rank : 86) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14617, O00202, O75262, Q59HF5, Q96G11, Q9H3C6 | Gene names | AP3D1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit delta-1 (Adapter-related protein complex 3 subunit delta-1) (Delta-adaptin 3) (AP-3 complex subunit delta) (Delta-adaptin). | |||||
|
ARLY_MOUSE
|
||||||
| NC score | 0.030912 (rank : 87) | θ value | 3.0926 (rank : 72) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91YI0, Q3UFA2 | Gene names | Asl | |||
|
Domain Architecture |

|
|||||
| Description | Argininosuccinate lyase (EC 4.3.2.1) (Arginosuccinase) (ASAL). | |||||
|
KIFA3_HUMAN
|
||||||
| NC score | 0.025046 (rank : 88) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92845, Q8NHU7, Q9H416 | Gene names | KIFAP3, SMAP | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-associated protein 3 (Smg GDS-associated protein). | |||||
|
IFIT2_HUMAN
|
||||||
| NC score | 0.020341 (rank : 89) | θ value | 3.0926 (rank : 75) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09913 | Gene names | IFIT2, G10P2, IFI54 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced protein with tetratricopeptide repeats 2 (IFIT-2) (Interferon-induced 54 kDa protein) (IFI-54K) (ISG-54 K). | |||||
|
SRPK2_MOUSE
|
||||||
| NC score | 0.018064 (rank : 90) | θ value | 1.81305 (rank : 69) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O54781, Q8VCD9 | Gene names | Srpk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK2 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 2) (SR-protein-specific kinase 2) (SFRS protein kinase 2). | |||||
|
SRPK2_HUMAN
|
||||||
| NC score | 0.017831 (rank : 91) | θ value | 1.81305 (rank : 68) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 664 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78362, O75220, O75221, Q6NUL0, Q6V1X2, Q8IYQ3 | Gene names | SRPK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK2 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 2) (SR-protein-specific kinase 2) (SFRS protein kinase 2). | |||||
|
IQGA2_HUMAN
|
||||||
| NC score | 0.016676 (rank : 92) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
CCNB3_HUMAN
|
||||||
| NC score | 0.015926 (rank : 93) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WWL7, Q96SB5, Q96SB6, Q96SB7, Q9NT38 | Gene names | CCNB3, CYCB3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
AP1G2_MOUSE
|
||||||
| NC score | 0.015074 (rank : 94) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88512 | Gene names | Ap1g2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-2 (Adapter-related protein complex 1 gamma- 2 subunit) (Gamma2-adaptin) (Adaptor protein complex AP-1 gamma-2 subunit) (G2ad). | |||||
|
BBS10_HUMAN
|
||||||
| NC score | 0.014253 (rank : 95) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TAM1, Q96CW2, Q9H5D2 | Gene names | BBS10, C12orf58 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bardet-Biedl syndrome 10 protein. | |||||
|
MTSS1_MOUSE
|
||||||
| NC score | 0.011205 (rank : 96) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R1S4, Q8BMM3, Q99LB3 | Gene names | Mtss1, Mim | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein). | |||||
|
RAG1_HUMAN
|
||||||
| NC score | 0.009994 (rank : 97) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15918, Q8NER2 | Gene names | RAG1, RNF74 | |||
|
Domain Architecture |

|
|||||
| Description | V(D)J recombination-activating protein 1 (RAG-1) (RING finger protein 74). | |||||
|
MELK_MOUSE
|
||||||
| NC score | 0.002960 (rank : 98) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61846, Q61804, Q6ZQH6 | Gene names | Melk, Kiaa0175, Pk38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Maternal embryonic leucine zipper kinase (EC 2.7.11.1) (Protein kinase PK38) (mPK38). | |||||
|
MELK_HUMAN
|
||||||
| NC score | 0.002839 (rank : 99) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q14680, Q7L3C3 | Gene names | MELK, KIAA0175 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Maternal embryonic leucine zipper kinase (EC 2.7.11.1) (hMELK) (Protein kinase PK38) (hPK38). | |||||
|
MET_HUMAN
|
||||||
| NC score | 0.002318 (rank : 100) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 87 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P08581, O60366, Q12875, Q9UDX7, Q9UPL8 | Gene names | MET | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor receptor precursor (EC 2.7.10.1) (HGF receptor) (Scatter factor receptor) (SF receptor) (HGF/SF receptor) (Met proto-oncogene tyrosine kinase) (c-Met). | |||||