Please be patient as the page loads
|
NAL14_HUMAN
|
||||||
| SwissProt Accessions | Q86W24, Q7RTR6 | Gene names | NALP14, NOD5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 14 (Nucleotide-binding oligomerization domain protein 5). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
NAL14_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q86W24, Q7RTR6 | Gene names | NALP14, NOD5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 14 (Nucleotide-binding oligomerization domain protein 5). | |||||
|
NALP4_HUMAN
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.983506 (rank : 2) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96MN2, Q86W87, Q96AY6 | Gene names | NALP4, PAN2, PYPAF4, RNH2 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 4 (PYRIN-containing APAF1-like protein 4) (PAAD and NACHT-containing protein 2) (PYRIN and NACHT- containing protein 2) (Ribonuclease inhibitor 2). | |||||
|
NALP5_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.977508 (rank : 5) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P59047, Q86W29 | Gene names | NALP5, MATER | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Mater protein homolog). | |||||
|
NALP5_MOUSE
|
||||||
| θ value | 9.13544e-178 (rank : 4) | NC score | 0.910126 (rank : 14) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
NAL12_HUMAN
|
||||||
| θ value | 4.53388e-177 (rank : 5) | NC score | 0.976793 (rank : 6) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P59046, Q8NEU4, Q9BY26 | Gene names | NALP12, PYPAF7, RNO | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 12 (PYRIN-containing APAF1-like protein 7) (Monarch-1) (Regulated by nitric oxide). | |||||
|
NAL13_HUMAN
|
||||||
| θ value | 3.36238e-172 (rank : 6) | NC score | 0.980319 (rank : 3) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q86W25, Q7RTR5 | Gene names | NALP13, NOD14 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 13 (Nucleotide-binding oligomerization domain protein 14). | |||||
|
NALP2_HUMAN
|
||||||
| θ value | 2.49359e-167 (rank : 7) | NC score | 0.978480 (rank : 4) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NX02, Q53FL5, Q59G09, Q8IXT0, Q9BVN5, Q9H6G6, Q9HAV9, Q9NWK3 | Gene names | NALP2, NBS1, PAN1, PYPAF2 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 2 (PYRIN domain and NACHT domain-containing protein 1) (PYRIN-containing APAF1-like protein 2) (Nucleotide-binding site protein 1). | |||||
|
CIAS1_HUMAN
|
||||||
| θ value | 6.35588e-163 (rank : 8) | NC score | 0.974196 (rank : 8) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96P20, O75434, Q59H68, Q5JQS8, Q5JQS9, Q6TG35, Q8TCW0, Q8TEU9, Q8WXH9 | Gene names | CIAS1, NALP3, PYPAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Cold autoinflammatory syndrome 1 protein (Cryopyrin) (NACHT-, LRR- and PYD-containing protein 3) (PYRIN-containing APAF1-like protein 1) (Angiotensin/vasopressin receptor AII/AVP-like). | |||||
|
CIAS1_MOUSE
|
||||||
| θ value | 7.52544e-156 (rank : 9) | NC score | 0.974484 (rank : 7) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8R4B8, Q6JEL0 | Gene names | Cias1, Mmig1, Nalp3, Pypaf1 | |||
|
Domain Architecture |

|
|||||
| Description | Cold autoinflammatory syndrome 1 protein homolog (PYRIN-containing APAF1-like protein 1) (Mast cell maturation-associated-inducible protein 1) (Cryopyrin). | |||||
|
NALP7_HUMAN
|
||||||
| θ value | 1.42253e-146 (rank : 10) | NC score | 0.970727 (rank : 9) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8WX94, Q7RTR1 | Gene names | NALP7, NOD12, PYPAF3 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 7 (PYRIN-containing APAF1-like protein 3). | |||||
|
NAL11_HUMAN
|
||||||
| θ value | 4.90052e-139 (rank : 11) | NC score | 0.964459 (rank : 10) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P59045, Q53ZZ0, Q8NBF5 | Gene names | NALP11, NOD17, PYPAF6 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 11 (PYRIN-containing APAF1-like protein 6) (Nucleotide-binding oligomerization domain protein 17). | |||||
|
NALP1_HUMAN
|
||||||
| θ value | 8.41738e-115 (rank : 12) | NC score | 0.923679 (rank : 13) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9C000, Q9BZZ8, Q9BZZ9, Q9HAV8, Q9UFT4, Q9Y2E0 | Gene names | NALP1, CARD7, DEFCAP, KIAA0926, NAC | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 1 (Death effector filament- forming ced-4-like apoptosis protein) (Nucleotide-binding domain and caspase recruitment domain) (Caspase recruitment domain protein 7). | |||||
|
NALP6_HUMAN
|
||||||
| θ value | 3.22841e-82 (rank : 13) | NC score | 0.932449 (rank : 11) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P59044 | Gene names | NALP6, PYPAF5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 6 (PYRIN-containing APAF1-like protein 5). | |||||
|
NALP6_MOUSE
|
||||||
| θ value | 4.82422e-78 (rank : 14) | NC score | 0.930973 (rank : 12) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q91WS2, Q8K0L4 | Gene names | Nalp6, Pypaf5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 6 (PYRIN-containing APAF1-like protein 5-like). | |||||
|
NAL10_HUMAN
|
||||||
| θ value | 2.84138e-62 (rank : 15) | NC score | 0.890562 (rank : 16) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86W26, Q6JGT0 | Gene names | NALP10, NOD8, PYNOD | |||
|
Domain Architecture |
|
|||||
| Description | NACHT, LRR and PYD-containing protein 10 (Nucleotide-binding oligomerization domain protein 8). | |||||
|
RINI_MOUSE
|
||||||
| θ value | 9.1404e-61 (rank : 16) | NC score | 0.890233 (rank : 17) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q91VI7, Q924P4 | Gene names | Rnh1, Rnh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribonuclease inhibitor (Ribonuclease/angiogenin inhibitor 1). | |||||
|
RINI_HUMAN
|
||||||
| θ value | 8.85319e-56 (rank : 17) | NC score | 0.892489 (rank : 15) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P13489 | Gene names | RNH1, PRI, RNH | |||
|
Domain Architecture |
|
|||||
| Description | Ribonuclease inhibitor (Ribonuclease/angiogenin inhibitor 1) (RAI) (Placental ribonuclease inhibitor) (RNase inhibitor) (RI). | |||||
|
NAL10_MOUSE
|
||||||
| θ value | 1.08223e-53 (rank : 18) | NC score | 0.887971 (rank : 18) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8CCN1 | Gene names | Nalp10, Pynod | |||
|
Domain Architecture |
|
|||||
| Description | NACHT, LRR and PYD-containing protein 10. | |||||
|
CARD4_HUMAN
|
||||||
| θ value | 4.28545e-26 (rank : 19) | NC score | 0.714100 (rank : 22) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Y239, Q8IWF5 | Gene names | CARD4, NOD1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 4 (Protein Nod1). | |||||
|
CARD4_MOUSE
|
||||||
| θ value | 5.59698e-26 (rank : 20) | NC score | 0.715031 (rank : 21) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BHB0, Q8BUT6 | Gene names | Card4 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 4. | |||||
|
CAR15_HUMAN
|
||||||
| θ value | 1.24688e-25 (rank : 21) | NC score | 0.732827 (rank : 20) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HC29, Q96RH5, Q96RH6, Q96RH8 | Gene names | CARD15, IBD1, NOD2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 15 (Protein Nod2) (Inflammatory bowel disease protein 1). | |||||
|
CAR15_MOUSE
|
||||||
| θ value | 4.01107e-24 (rank : 22) | NC score | 0.749751 (rank : 19) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8K3Z0 | Gene names | Card15, Nod2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 15 (Protein Nod2). | |||||
|
C2TA_MOUSE
|
||||||
| θ value | 2.98157e-11 (rank : 23) | NC score | 0.614188 (rank : 23) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P79621, O46787, O78036, O78109, Q31115, Q9TPP1 | Gene names | Ciita, C2ta, Mhc2ta | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II transactivator (CIITA). | |||||
|
LRC31_HUMAN
|
||||||
| θ value | 1.06045e-08 (rank : 24) | NC score | 0.529761 (rank : 27) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6UY01, Q8N848, Q9H5N5 | Gene names | LRRC31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 31. | |||||
|
LRC34_HUMAN
|
||||||
| θ value | 5.26297e-08 (rank : 25) | NC score | 0.564790 (rank : 25) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8IZ02 | Gene names | LRRC34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 34. | |||||
|
CF154_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 26) | NC score | 0.536974 (rank : 26) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5JTD7 | Gene names | C6orf154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf154. | |||||
|
C2TA_HUMAN
|
||||||
| θ value | 4.45536e-07 (rank : 27) | NC score | 0.592622 (rank : 24) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P33076 | Gene names | CIITA, MHC2TA | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II transactivator (CIITA). | |||||
|
LRC34_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 28) | NC score | 0.525908 (rank : 28) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9DAM1 | Gene names | Lrrc34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 34. | |||||
|
RGP1_HUMAN
|
||||||
| θ value | 3.77169e-06 (rank : 29) | NC score | 0.375808 (rank : 29) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P46060 | Gene names | RANGAP1 | |||
|
Domain Architecture |
|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
LRC45_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 30) | NC score | 0.183564 (rank : 40) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8CIM1, Q3U5Z2 | Gene names | Lrrc45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
RGP1_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 31) | NC score | 0.360312 (rank : 30) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P46061, Q60801 | Gene names | Rangap1, Fug1 | |||
|
Domain Architecture |
|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
BIR1A_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 32) | NC score | 0.225317 (rank : 34) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QWK5, Q9JIB5, Q9R017 | Gene names | Birc1a, Naip, Naip1 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1a (Neuronal apoptosis inhibitory protein 1). | |||||
|
MEFV_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 33) | NC score | 0.086618 (rank : 45) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15553, Q96PN4, Q96PN5 | Gene names | MEFV, MEF | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
BIRC1_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 34) | NC score | 0.242507 (rank : 33) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13075, O75857, Q13730, Q99796 | Gene names | BIRC1, NAIP | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1 (Neuronal apoptosis inhibitory protein). | |||||
|
BIR1F_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 35) | NC score | 0.219733 (rank : 35) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9JIB6, O09121, O09122, P81704 | Gene names | Birc1f, Naip-rs4, Naip6 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1f (Neuronal apoptosis inhibitory protein 6). | |||||
|
LRC45_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 36) | NC score | 0.172972 (rank : 41) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 933 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96CN5 | Gene names | LRRC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
BIR1B_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 37) | NC score | 0.205991 (rank : 38) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9QUK4, O09124, Q9R030 | Gene names | Birc1b, Naip-rs6, Naip2 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1b (Neuronal apoptosis inhibitory protein 2). | |||||
|
BIR1E_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 38) | NC score | 0.217456 (rank : 36) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9R016, O09121, O09122, P81703, Q9R029 | Gene names | Birc1e, Naip-rs3, Naip5 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1e (Neuronal apoptosis inhibitory protein 5). | |||||
|
BIR1G_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 39) | NC score | 0.211786 (rank : 37) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JIB3 | Gene names | Birc1g, Naip7 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1g (Neuronal apoptosis inhibitory protein 7). | |||||
|
LONM_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 40) | NC score | 0.052398 (rank : 49) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P36776, P36777, Q9UQ95 | Gene names | PRSS15 | |||
|
Domain Architecture |

|
|||||
| Description | Lon protease homolog, mitochondrial precursor (EC 3.4.21.-) (Lon protease-like protein) (LONP) (LONHs). | |||||
|
FXL16_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 41) | NC score | 0.048076 (rank : 51) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N461, Q2MHR2, Q96S14, Q9UJI0 | Gene names | FBXL16, C16orf22, FBL16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 16 (F-box and leucine-rich repeat protein 16). | |||||
|
MAPE_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 42) | NC score | 0.072320 (rank : 46) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P78395, O43481, Q8IXN8 | Gene names | PRAME, MAPE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma antigen preferentially expressed in tumors (Preferentially expressed antigen of melanoma) (OPA-interacting protein 4) (OIP4). | |||||
|
LRRK2_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 43) | NC score | 0.005000 (rank : 75) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1382 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5S007, Q6ZS50, Q8NCX9 | Gene names | LRRK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 2 (EC 2.7.11.1) (Dardarin). | |||||
|
TLR13_MOUSE
|
||||||
| θ value | 0.125558 (rank : 44) | NC score | 0.022058 (rank : 57) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6R5N8, Q3TDS2 | Gene names | Tlr13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Toll-like receptor 13 precursor. | |||||
|
AIM2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 45) | NC score | 0.053943 (rank : 48) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O14862 | Gene names | AIM2 | |||
|
Domain Architecture |
|
|||||
| Description | Interferon-inducible protein AIM2 (Absent in melanoma 2). | |||||
|
CAR12_HUMAN
|
||||||
| θ value | 0.21417 (rank : 46) | NC score | 0.331270 (rank : 31) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9NPP4, Q96J81, Q96J82, Q96J83 | Gene names | CARD12, CLAN, CLAN1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 12 (Ice protease- activating factor) (Ipaf) (CARD, LRR, and NACHT-containing protein) (Clan protein). | |||||
|
CP2A4_MOUSE
|
||||||
| θ value | 0.279714 (rank : 47) | NC score | 0.007555 (rank : 71) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15392 | Gene names | Cyp2a4, Cyp2a-4 | |||
|
Domain Architecture |
|
|||||
| Description | Cytochrome P450 2A4 (EC 1.14.14.1) (CYPIIA4) (Testosterone 15-alpha- hydroxylase) (P450-15-alpha) (P450-IIA3.1). | |||||
|
CP2A5_MOUSE
|
||||||
| θ value | 0.279714 (rank : 48) | NC score | 0.007533 (rank : 73) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20852 | Gene names | Cyp2a5, Cyp2a-5 | |||
|
Domain Architecture |
|
|||||
| Description | Cytochrome P450 2A5 (EC 1.14.14.1) (CYPIIA5) (Coumarin 7-hydroxylase) (P450-15-COH) (P450-IIA3.2). | |||||
|
FBXL5_HUMAN
|
||||||
| θ value | 0.47712 (rank : 49) | NC score | 0.020857 (rank : 60) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKA1, Q8NHP3, Q9NXN2, Q9P0I0, Q9P0X5, Q9UJT7, Q9UKC8 | Gene names | FBXL5, FBL4, FBL5, FLR1 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 5 (F-box and leucine-rich repeat protein 5) (F-box protein FBL4/FBL5) (p45SKP2-like protein). | |||||
|
LRC14_HUMAN
|
||||||
| θ value | 0.62314 (rank : 50) | NC score | 0.108135 (rank : 44) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15048 | Gene names | LRRC14, KIAA0014 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing protein 14. | |||||
|
BMS1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 51) | NC score | 0.027917 (rank : 55) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14692, Q86XJ9 | Gene names | BMS1L, KIAA0187 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome biogenesis protein BMS1 homolog. | |||||
|
PHLPL_HUMAN
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.021097 (rank : 59) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6ZVD8, Q9NV17, Q9Y2E3 | Gene names | PHLPPL, KIAA0931 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat protein phosphatase-like (EC 3.1.3.16). | |||||
|
PRAM1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.025958 (rank : 56) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O95521, Q9UQP2 | Gene names | PRAMEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PRAME family member 1. | |||||
|
DOCK2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 54) | NC score | 0.011502 (rank : 65) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92608, Q96AK7 | Gene names | DOCK2, KIAA0209 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 2. | |||||
|
FXL21_HUMAN
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.012527 (rank : 64) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UKT6 | Gene names | FBXL21, FBL21, FBL3, FBXL3B | |||
|
Domain Architecture |
|
|||||
| Description | F-box/LRR-repeat protein 21 (F-box and leucine-rich repeat protein 21) (F-box/LRR-repeat protein 3B) (F-box and leucine-rich repeat protein 3B). | |||||
|
LRC42_MOUSE
|
||||||
| θ value | 2.36792 (rank : 56) | NC score | 0.042354 (rank : 53) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8R2U7, Q3TE62, Q9CTP9 | Gene names | Lrrc42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 42. | |||||
|
OR5DD_HUMAN
|
||||||
| θ value | 2.36792 (rank : 57) | NC score | -0.003197 (rank : 80) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NGL4 | Gene names | OR5D13 | |||
|
Domain Architecture |
|
|||||
| Description | Olfactory receptor 5D13. | |||||
|
CACB4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.008246 (rank : 69) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R0S4, Q8BRN6, Q8CAJ9 | Gene names | Cacnb4, Cacnlb4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit beta-4 (CAB4) (Calcium channel voltage-dependent subunit beta 4). | |||||
|
FXL13_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.037600 (rank : 54) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NEE6, Q6UVW7, Q6UVW8, Q75MN5, Q86UJ5, Q8N7Y4, Q8TCL2, Q8WUF9, Q8WUG0 | Gene names | FBXL13, FBL13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
LRC33_MOUSE
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.018985 (rank : 62) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BMT4, Q3TIA8, Q8BTT4, Q8BUI7, Q8BY16, Q8R063 | Gene names | Lrrc33 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 33 precursor. | |||||
|
PRA12_HUMAN
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.044942 (rank : 52) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95522 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PRAME family member DJ1198H6.2. | |||||
|
DOCK2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.009932 (rank : 67) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C3J5, Q99M79 | Gene names | Dock2 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 2 (Hch protein). | |||||
|
LRC33_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.021648 (rank : 58) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86YC3 | Gene names | LRRC33 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 33 precursor. | |||||
|
PHLPL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.019041 (rank : 61) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BXA7, Q8BX96 | Gene names | Phlppl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat protein phosphatase-like (EC 3.1.3.16). | |||||
|
LRC39_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.014197 (rank : 63) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96DD0, Q5VVK7 | Gene names | LRRC39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 39 (Densin hlg). | |||||
|
LRCH3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.008136 (rank : 70) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BVU0, Q3U222, Q3UZ74 | Gene names | Lrch3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 3 precursor. | |||||
|
TLR8_MOUSE
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.009729 (rank : 68) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P58682, Q91XI7 | Gene names | Tlr8 | |||
|
Domain Architecture |

|
|||||
| Description | Toll-like receptor 8 precursor. | |||||
|
CD14_MOUSE
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.063674 (rank : 47) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P10810 | Gene names | Cd14 | |||
|
Domain Architecture |
|
|||||
| Description | Monocyte differentiation antigen CD14 precursor (Myeloid cell-specific leucine-rich glycoprotein). | |||||
|
MOD5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.010782 (rank : 66) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H3H1, Q6IAC9, Q96FJ3, Q96L45, Q9NXT7 | Gene names | TRIT1, IPT, MOD5 | |||
|
Domain Architecture |
|
|||||
| Description | tRNA isopentenyltransferase, mitochondrial precursor (EC 2.5.1.8) (Isopentenyl-diphosphate:tRNA isopentenyltransferase) (IPP transferase) (IPTase) (IPPT) (hGRO1). | |||||
|
MYO9B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | -0.000519 (rank : 79) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
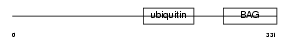
BAG1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.007534 (rank : 72) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60739 | Gene names | Bag1 | |||
|
Domain Architecture |
|
|||||
| Description | BAG family molecular chaperone regulator 1 (BCL-2-binding athanogene- 1) (BAG-1). | |||||
|
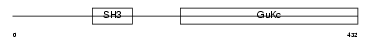
CACB4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.006215 (rank : 74) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O00305, O60515, Q96L40 | Gene names | CACNB4, CACNLB4 | |||
|
Domain Architecture |
|
|||||
| Description | Voltage-dependent L-type calcium channel subunit beta-4 (CAB4) (Calcium channel voltage-dependent subunit beta 4). | |||||
|
CCD46_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | -0.000467 (rank : 78) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
CDKL3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | -0.004245 (rank : 81) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 857 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BLF2, Q5M6W2, Q8BKR2, Q8BL49, Q8BVE0, Q8K134 | Gene names | Cdkl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase-like 3 (EC 2.7.11.22). | |||||
|
KLDC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.003766 (rank : 76) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80YG3 | Gene names | Klhdc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch domain-containing protein 1. | |||||
|
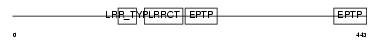
LGI3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.002703 (rank : 77) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8K406 | Gene names | Lgi3 | |||
|
Domain Architecture |
|
|||||
| Description | Leucine-rich repeat LGI family member 3 precursor (Leucine-rich glioma-inactivated protein 3) (Leubrin). | |||||
|
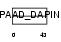
ASC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.203734 (rank : 39) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9ULZ3, Q96D12, Q9BSZ5, Q9HBD0, Q9NXJ8 | Gene names | PYCARD, ASC, CARD5, TMS1 | |||
|
Domain Architecture |
|
|||||
| Description | Apoptosis-associated speck-like protein containing a CARD (hASC) (PYD and CARD domain-containing protein) (Target of methylation-induced silencing 1) (Caspase recruitment domain-containing protein 5). | |||||
|
ASC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.258658 (rank : 32) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9EPB4, Q9D2W9 | Gene names | Pycard, Asc | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-associated speck-like protein containing a CARD (mASC) (PYD and CARD domain-containing protein). | |||||
|
CARD8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.119970 (rank : 43) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y2G2, Q6PGP8, Q96P82 | Gene names | CARD8, KIAA0955, NDPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 8 (Apoptotic protein NDPP1) (DACAR) (CARD-inhibitor of NF-kappa-B-activating ligand) (CARDINAL) (Tumor-up-regulated CARD-containing antagonist of CASP9) (TUCAN). | |||||
|
MEFV_MOUSE
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.143237 (rank : 42) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JJ26 | Gene names | Mefv | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
TMOD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.052199 (rank : 50) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NZR1 | Gene names | TMOD2, NTMOD | |||
|
Domain Architecture |

|
|||||
| Description | Tropomodulin-2 (Neuronal tropomodulin) (N-Tmod). | |||||
|
NAL14_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q86W24, Q7RTR6 | Gene names | NALP14, NOD5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 14 (Nucleotide-binding oligomerization domain protein 5). | |||||
|
NALP4_HUMAN
|
||||||
| NC score | 0.983506 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96MN2, Q86W87, Q96AY6 | Gene names | NALP4, PAN2, PYPAF4, RNH2 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 4 (PYRIN-containing APAF1-like protein 4) (PAAD and NACHT-containing protein 2) (PYRIN and NACHT- containing protein 2) (Ribonuclease inhibitor 2). | |||||
|
NAL13_HUMAN
|
||||||
| NC score | 0.980319 (rank : 3) | θ value | 3.36238e-172 (rank : 6) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q86W25, Q7RTR5 | Gene names | NALP13, NOD14 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 13 (Nucleotide-binding oligomerization domain protein 14). | |||||
|
NALP2_HUMAN
|
||||||
| NC score | 0.978480 (rank : 4) | θ value | 2.49359e-167 (rank : 7) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NX02, Q53FL5, Q59G09, Q8IXT0, Q9BVN5, Q9H6G6, Q9HAV9, Q9NWK3 | Gene names | NALP2, NBS1, PAN1, PYPAF2 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 2 (PYRIN domain and NACHT domain-containing protein 1) (PYRIN-containing APAF1-like protein 2) (Nucleotide-binding site protein 1). | |||||
|
NALP5_HUMAN
|
||||||
| NC score | 0.977508 (rank : 5) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P59047, Q86W29 | Gene names | NALP5, MATER | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Mater protein homolog). | |||||
|
NAL12_HUMAN
|
||||||
| NC score | 0.976793 (rank : 6) | θ value | 4.53388e-177 (rank : 5) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P59046, Q8NEU4, Q9BY26 | Gene names | NALP12, PYPAF7, RNO | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 12 (PYRIN-containing APAF1-like protein 7) (Monarch-1) (Regulated by nitric oxide). | |||||
|
CIAS1_MOUSE
|
||||||
| NC score | 0.974484 (rank : 7) | θ value | 7.52544e-156 (rank : 9) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8R4B8, Q6JEL0 | Gene names | Cias1, Mmig1, Nalp3, Pypaf1 | |||
|
Domain Architecture |

|
|||||
| Description | Cold autoinflammatory syndrome 1 protein homolog (PYRIN-containing APAF1-like protein 1) (Mast cell maturation-associated-inducible protein 1) (Cryopyrin). | |||||
|
CIAS1_HUMAN
|
||||||
| NC score | 0.974196 (rank : 8) | θ value | 6.35588e-163 (rank : 8) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96P20, O75434, Q59H68, Q5JQS8, Q5JQS9, Q6TG35, Q8TCW0, Q8TEU9, Q8WXH9 | Gene names | CIAS1, NALP3, PYPAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Cold autoinflammatory syndrome 1 protein (Cryopyrin) (NACHT-, LRR- and PYD-containing protein 3) (PYRIN-containing APAF1-like protein 1) (Angiotensin/vasopressin receptor AII/AVP-like). | |||||
|
NALP7_HUMAN
|
||||||
| NC score | 0.970727 (rank : 9) | θ value | 1.42253e-146 (rank : 10) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8WX94, Q7RTR1 | Gene names | NALP7, NOD12, PYPAF3 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 7 (PYRIN-containing APAF1-like protein 3). | |||||
|
NAL11_HUMAN
|
||||||
| NC score | 0.964459 (rank : 10) | θ value | 4.90052e-139 (rank : 11) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P59045, Q53ZZ0, Q8NBF5 | Gene names | NALP11, NOD17, PYPAF6 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 11 (PYRIN-containing APAF1-like protein 6) (Nucleotide-binding oligomerization domain protein 17). | |||||
|
NALP6_HUMAN
|
||||||
| NC score | 0.932449 (rank : 11) | θ value | 3.22841e-82 (rank : 13) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P59044 | Gene names | NALP6, PYPAF5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 6 (PYRIN-containing APAF1-like protein 5). | |||||
|
NALP6_MOUSE
|
||||||
| NC score | 0.930973 (rank : 12) | θ value | 4.82422e-78 (rank : 14) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q91WS2, Q8K0L4 | Gene names | Nalp6, Pypaf5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 6 (PYRIN-containing APAF1-like protein 5-like). | |||||
|
NALP1_HUMAN
|
||||||
| NC score | 0.923679 (rank : 13) | θ value | 8.41738e-115 (rank : 12) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9C000, Q9BZZ8, Q9BZZ9, Q9HAV8, Q9UFT4, Q9Y2E0 | Gene names | NALP1, CARD7, DEFCAP, KIAA0926, NAC | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 1 (Death effector filament- forming ced-4-like apoptosis protein) (Nucleotide-binding domain and caspase recruitment domain) (Caspase recruitment domain protein 7). | |||||
|
NALP5_MOUSE
|
||||||
| NC score | 0.910126 (rank : 14) | θ value | 9.13544e-178 (rank : 4) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
RINI_HUMAN
|
||||||
| NC score | 0.892489 (rank : 15) | θ value | 8.85319e-56 (rank : 17) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P13489 | Gene names | RNH1, PRI, RNH | |||
|
Domain Architecture |

|
|||||
| Description | Ribonuclease inhibitor (Ribonuclease/angiogenin inhibitor 1) (RAI) (Placental ribonuclease inhibitor) (RNase inhibitor) (RI). | |||||
|
NAL10_HUMAN
|
||||||
| NC score | 0.890562 (rank : 16) | θ value | 2.84138e-62 (rank : 15) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86W26, Q6JGT0 | Gene names | NALP10, NOD8, PYNOD | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 10 (Nucleotide-binding oligomerization domain protein 8). | |||||
|
RINI_MOUSE
|
||||||
| NC score | 0.890233 (rank : 17) | θ value | 9.1404e-61 (rank : 16) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q91VI7, Q924P4 | Gene names | Rnh1, Rnh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribonuclease inhibitor (Ribonuclease/angiogenin inhibitor 1). | |||||
|
NAL10_MOUSE
|
||||||
| NC score | 0.887971 (rank : 18) | θ value | 1.08223e-53 (rank : 18) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8CCN1 | Gene names | Nalp10, Pynod | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 10. | |||||
|
CAR15_MOUSE
|
||||||
| NC score | 0.749751 (rank : 19) | θ value | 4.01107e-24 (rank : 22) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8K3Z0 | Gene names | Card15, Nod2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 15 (Protein Nod2). | |||||
|
CAR15_HUMAN
|
||||||
| NC score | 0.732827 (rank : 20) | θ value | 1.24688e-25 (rank : 21) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HC29, Q96RH5, Q96RH6, Q96RH8 | Gene names | CARD15, IBD1, NOD2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 15 (Protein Nod2) (Inflammatory bowel disease protein 1). | |||||
|
CARD4_MOUSE
|
||||||
| NC score | 0.715031 (rank : 21) | θ value | 5.59698e-26 (rank : 20) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BHB0, Q8BUT6 | Gene names | Card4 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 4. | |||||
|
CARD4_HUMAN
|
||||||
| NC score | 0.714100 (rank : 22) | θ value | 4.28545e-26 (rank : 19) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Y239, Q8IWF5 | Gene names | CARD4, NOD1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 4 (Protein Nod1). | |||||
|
C2TA_MOUSE
|
||||||
| NC score | 0.614188 (rank : 23) | θ value | 2.98157e-11 (rank : 23) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P79621, O46787, O78036, O78109, Q31115, Q9TPP1 | Gene names | Ciita, C2ta, Mhc2ta | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II transactivator (CIITA). | |||||
|
C2TA_HUMAN
|
||||||
| NC score | 0.592622 (rank : 24) | θ value | 4.45536e-07 (rank : 27) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P33076 | Gene names | CIITA, MHC2TA | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II transactivator (CIITA). | |||||
|
LRC34_HUMAN
|
||||||
| NC score | 0.564790 (rank : 25) | θ value | 5.26297e-08 (rank : 25) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8IZ02 | Gene names | LRRC34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 34. | |||||
|
CF154_HUMAN
|
||||||
| NC score | 0.536974 (rank : 26) | θ value | 3.41135e-07 (rank : 26) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5JTD7 | Gene names | C6orf154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf154. | |||||
|
LRC31_HUMAN
|
||||||
| NC score | 0.529761 (rank : 27) | θ value | 1.06045e-08 (rank : 24) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6UY01, Q8N848, Q9H5N5 | Gene names | LRRC31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 31. | |||||
|
LRC34_MOUSE
|
||||||
| NC score | 0.525908 (rank : 28) | θ value | 2.21117e-06 (rank : 28) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9DAM1 | Gene names | Lrrc34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 34. | |||||
|
RGP1_HUMAN
|
||||||
| NC score | 0.375808 (rank : 29) | θ value | 3.77169e-06 (rank : 29) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P46060 | Gene names | RANGAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
RGP1_MOUSE
|
||||||
| NC score | 0.360312 (rank : 30) | θ value | 7.1131e-05 (rank : 31) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P46061, Q60801 | Gene names | Rangap1, Fug1 | |||
|
Domain Architecture |

|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
CAR12_HUMAN
|
||||||
| NC score | 0.331270 (rank : 31) | θ value | 0.21417 (rank : 46) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9NPP4, Q96J81, Q96J82, Q96J83 | Gene names | CARD12, CLAN, CLAN1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 12 (Ice protease- activating factor) (Ipaf) (CARD, LRR, and NACHT-containing protein) (Clan protein). | |||||
|
ASC_MOUSE
|
||||||
| NC score | 0.258658 (rank : 32) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9EPB4, Q9D2W9 | Gene names | Pycard, Asc | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-associated speck-like protein containing a CARD (mASC) (PYD and CARD domain-containing protein). | |||||
|
BIRC1_HUMAN
|
||||||
| NC score | 0.242507 (rank : 33) | θ value | 0.00020696 (rank : 34) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13075, O75857, Q13730, Q99796 | Gene names | BIRC1, NAIP | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1 (Neuronal apoptosis inhibitory protein). | |||||
|
BIR1A_MOUSE
|
||||||
| NC score | 0.225317 (rank : 34) | θ value | 9.29e-05 (rank : 32) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QWK5, Q9JIB5, Q9R017 | Gene names | Birc1a, Naip, Naip1 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1a (Neuronal apoptosis inhibitory protein 1). | |||||
|
BIR1F_MOUSE
|
||||||
| NC score | 0.219733 (rank : 35) | θ value | 0.00035302 (rank : 35) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9JIB6, O09121, O09122, P81704 | Gene names | Birc1f, Naip-rs4, Naip6 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1f (Neuronal apoptosis inhibitory protein 6). | |||||
|
BIR1E_MOUSE
|
||||||
| NC score | 0.217456 (rank : 36) | θ value | 0.00134147 (rank : 38) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9R016, O09121, O09122, P81703, Q9R029 | Gene names | Birc1e, Naip-rs3, Naip5 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1e (Neuronal apoptosis inhibitory protein 5). | |||||
|
BIR1G_MOUSE
|
||||||
| NC score | 0.211786 (rank : 37) | θ value | 0.00175202 (rank : 39) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JIB3 | Gene names | Birc1g, Naip7 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1g (Neuronal apoptosis inhibitory protein 7). | |||||
|
BIR1B_MOUSE
|
||||||
| NC score | 0.205991 (rank : 38) | θ value | 0.000602161 (rank : 37) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9QUK4, O09124, Q9R030 | Gene names | Birc1b, Naip-rs6, Naip2 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1b (Neuronal apoptosis inhibitory protein 2). | |||||
|
ASC_HUMAN
|
||||||
| NC score | 0.203734 (rank : 39) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9ULZ3, Q96D12, Q9BSZ5, Q9HBD0, Q9NXJ8 | Gene names | PYCARD, ASC, CARD5, TMS1 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-associated speck-like protein containing a CARD (hASC) (PYD and CARD domain-containing protein) (Target of methylation-induced silencing 1) (Caspase recruitment domain-containing protein 5). | |||||
|
LRC45_MOUSE
|
||||||
| NC score | 0.183564 (rank : 40) | θ value | 1.87187e-05 (rank : 30) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8CIM1, Q3U5Z2 | Gene names | Lrrc45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
LRC45_HUMAN
|
||||||
| NC score | 0.172972 (rank : 41) | θ value | 0.000461057 (rank : 36) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 933 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96CN5 | Gene names | LRRC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
MEFV_MOUSE
|
||||||
| NC score | 0.143237 (rank : 42) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JJ26 | Gene names | Mefv | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
CARD8_HUMAN
|
||||||
| NC score | 0.119970 (rank : 43) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y2G2, Q6PGP8, Q96P82 | Gene names | CARD8, KIAA0955, NDPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 8 (Apoptotic protein NDPP1) (DACAR) (CARD-inhibitor of NF-kappa-B-activating ligand) (CARDINAL) (Tumor-up-regulated CARD-containing antagonist of CASP9) (TUCAN). | |||||
|
LRC14_HUMAN
|
||||||
| NC score | 0.108135 (rank : 44) | θ value | 0.62314 (rank : 50) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15048 | Gene names | LRRC14, KIAA0014 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing protein 14. | |||||
|
MEFV_HUMAN
|
||||||
| NC score | 0.086618 (rank : 45) | θ value | 9.29e-05 (rank : 33) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15553, Q96PN4, Q96PN5 | Gene names | MEFV, MEF | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
MAPE_HUMAN
|
||||||
| NC score | 0.072320 (rank : 46) | θ value | 0.0563607 (rank : 42) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P78395, O43481, Q8IXN8 | Gene names | PRAME, MAPE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma antigen preferentially expressed in tumors (Preferentially expressed antigen of melanoma) (OPA-interacting protein 4) (OIP4). | |||||
|
CD14_MOUSE
|
||||||
| NC score | 0.063674 (rank : 47) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P10810 | Gene names | Cd14 | |||
|
Domain Architecture |

|
|||||
| Description | Monocyte differentiation antigen CD14 precursor (Myeloid cell-specific leucine-rich glycoprotein). | |||||
|
AIM2_HUMAN
|
||||||
| NC score | 0.053943 (rank : 48) | θ value | 0.163984 (rank : 45) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O14862 | Gene names | AIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-inducible protein AIM2 (Absent in melanoma 2). | |||||
|
LONM_HUMAN
|
||||||
| NC score | 0.052398 (rank : 49) | θ value | 0.0193708 (rank : 40) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P36776, P36777, Q9UQ95 | Gene names | PRSS15 | |||
|
Domain Architecture |

|
|||||
| Description | Lon protease homolog, mitochondrial precursor (EC 3.4.21.-) (Lon protease-like protein) (LONP) (LONHs). | |||||
|
TMOD2_HUMAN
|
||||||
| NC score | 0.052199 (rank : 50) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NZR1 | Gene names | TMOD2, NTMOD | |||
|
Domain Architecture |

|
|||||
| Description | Tropomodulin-2 (Neuronal tropomodulin) (N-Tmod). | |||||
|
FXL16_HUMAN
|
||||||
| NC score | 0.048076 (rank : 51) | θ value | 0.0330416 (rank : 41) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N461, Q2MHR2, Q96S14, Q9UJI0 | Gene names | FBXL16, C16orf22, FBL16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 16 (F-box and leucine-rich repeat protein 16). | |||||
|
PRA12_HUMAN
|
||||||
| NC score | 0.044942 (rank : 52) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95522 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PRAME family member DJ1198H6.2. | |||||
|
LRC42_MOUSE
|
||||||
| NC score | 0.042354 (rank : 53) | θ value | 2.36792 (rank : 56) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8R2U7, Q3TE62, Q9CTP9 | Gene names | Lrrc42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 42. | |||||
|
FXL13_HUMAN
|
||||||
| NC score | 0.037600 (rank : 54) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NEE6, Q6UVW7, Q6UVW8, Q75MN5, Q86UJ5, Q8N7Y4, Q8TCL2, Q8WUF9, Q8WUG0 | Gene names | FBXL13, FBL13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
BMS1_HUMAN
|
||||||
| NC score | 0.027917 (rank : 55) | θ value | 0.813845 (rank : 51) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14692, Q86XJ9 | Gene names | BMS1L, KIAA0187 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome biogenesis protein BMS1 homolog. | |||||
|
PRAM1_HUMAN
|
||||||
| NC score | 0.025958 (rank : 56) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O95521, Q9UQP2 | Gene names | PRAMEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PRAME family member 1. | |||||
|
TLR13_MOUSE
|
||||||
| NC score | 0.022058 (rank : 57) | θ value | 0.125558 (rank : 44) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6R5N8, Q3TDS2 | Gene names | Tlr13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Toll-like receptor 13 precursor. | |||||
|
LRC33_HUMAN
|
||||||
| NC score | 0.021648 (rank : 58) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86YC3 | Gene names | LRRC33 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 33 precursor. | |||||
|
PHLPL_HUMAN
|
||||||
| NC score | 0.021097 (rank : 59) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6ZVD8, Q9NV17, Q9Y2E3 | Gene names | PHLPPL, KIAA0931 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat protein phosphatase-like (EC 3.1.3.16). | |||||
|
FBXL5_HUMAN
|
||||||
| NC score | 0.020857 (rank : 60) | θ value | 0.47712 (rank : 49) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKA1, Q8NHP3, Q9NXN2, Q9P0I0, Q9P0X5, Q9UJT7, Q9UKC8 | Gene names | FBXL5, FBL4, FBL5, FLR1 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 5 (F-box and leucine-rich repeat protein 5) (F-box protein FBL4/FBL5) (p45SKP2-like protein). | |||||
|
PHLPL_MOUSE
|
||||||
| NC score | 0.019041 (rank : 61) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BXA7, Q8BX96 | Gene names | Phlppl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat protein phosphatase-like (EC 3.1.3.16). | |||||
|
LRC33_MOUSE
|
||||||
| NC score | 0.018985 (rank : 62) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BMT4, Q3TIA8, Q8BTT4, Q8BUI7, Q8BY16, Q8R063 | Gene names | Lrrc33 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 33 precursor. | |||||
|
LRC39_HUMAN
|
||||||
| NC score | 0.014197 (rank : 63) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96DD0, Q5VVK7 | Gene names | LRRC39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 39 (Densin hlg). | |||||
|
FXL21_HUMAN
|
||||||
| NC score | 0.012527 (rank : 64) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UKT6 | Gene names | FBXL21, FBL21, FBL3, FBXL3B | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 21 (F-box and leucine-rich repeat protein 21) (F-box/LRR-repeat protein 3B) (F-box and leucine-rich repeat protein 3B). | |||||
|
DOCK2_HUMAN
|
||||||
| NC score | 0.011502 (rank : 65) | θ value | 1.38821 (rank : 54) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92608, Q96AK7 | Gene names | DOCK2, KIAA0209 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 2. | |||||
|
MOD5_HUMAN
|
||||||
| NC score | 0.010782 (rank : 66) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H3H1, Q6IAC9, Q96FJ3, Q96L45, Q9NXT7 | Gene names | TRIT1, IPT, MOD5 | |||
|
Domain Architecture |

|
|||||
| Description | tRNA isopentenyltransferase, mitochondrial precursor (EC 2.5.1.8) (Isopentenyl-diphosphate:tRNA isopentenyltransferase) (IPP transferase) (IPTase) (IPPT) (hGRO1). | |||||
|
DOCK2_MOUSE
|
||||||
| NC score | 0.009932 (rank : 67) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C3J5, Q99M79 | Gene names | Dock2 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 2 (Hch protein). | |||||
|
TLR8_MOUSE
|
||||||
| NC score | 0.009729 (rank : 68) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P58682, Q91XI7 | Gene names | Tlr8 | |||
|
Domain Architecture |

|
|||||
| Description | Toll-like receptor 8 precursor. | |||||
|
CACB4_MOUSE
|
||||||
| NC score | 0.008246 (rank : 69) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R0S4, Q8BRN6, Q8CAJ9 | Gene names | Cacnb4, Cacnlb4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit beta-4 (CAB4) (Calcium channel voltage-dependent subunit beta 4). | |||||
|
LRCH3_MOUSE
|
||||||
| NC score | 0.008136 (rank : 70) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BVU0, Q3U222, Q3UZ74 | Gene names | Lrch3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 3 precursor. | |||||
|
CP2A4_MOUSE
|
||||||
| NC score | 0.007555 (rank : 71) | θ value | 0.279714 (rank : 47) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15392 | Gene names | Cyp2a4, Cyp2a-4 | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome P450 2A4 (EC 1.14.14.1) (CYPIIA4) (Testosterone 15-alpha- hydroxylase) (P450-15-alpha) (P450-IIA3.1). | |||||
|
BAG1_MOUSE
|
||||||
| NC score | 0.007534 (rank : 72) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60739 | Gene names | Bag1 | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 1 (BCL-2-binding athanogene- 1) (BAG-1). | |||||
|
CP2A5_MOUSE
|
||||||
| NC score | 0.007533 (rank : 73) | θ value | 0.279714 (rank : 48) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20852 | Gene names | Cyp2a5, Cyp2a-5 | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome P450 2A5 (EC 1.14.14.1) (CYPIIA5) (Coumarin 7-hydroxylase) (P450-15-COH) (P450-IIA3.2). | |||||
|
CACB4_HUMAN
|
||||||
| NC score | 0.006215 (rank : 74) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O00305, O60515, Q96L40 | Gene names | CACNB4, CACNLB4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit beta-4 (CAB4) (Calcium channel voltage-dependent subunit beta 4). | |||||
|
LRRK2_HUMAN
|
||||||
| NC score | 0.005000 (rank : 75) | θ value | 0.0736092 (rank : 43) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1382 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5S007, Q6ZS50, Q8NCX9 | Gene names | LRRK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 2 (EC 2.7.11.1) (Dardarin). | |||||
|
KLDC1_MOUSE
|
||||||
| NC score | 0.003766 (rank : 76) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80YG3 | Gene names | Klhdc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch domain-containing protein 1. | |||||
|
LGI3_MOUSE
|
||||||
| NC score | 0.002703 (rank : 77) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8K406 | Gene names | Lgi3 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat LGI family member 3 precursor (Leucine-rich glioma-inactivated protein 3) (Leubrin). | |||||
|
CCD46_HUMAN
|
||||||
| NC score | -0.000467 (rank : 78) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
MYO9B_HUMAN
|
||||||
| NC score | -0.000519 (rank : 79) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
OR5DD_HUMAN
|
||||||
| NC score | -0.003197 (rank : 80) | θ value | 2.36792 (rank : 57) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NGL4 | Gene names | OR5D13 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 5D13. | |||||
|
CDKL3_MOUSE
|
||||||
| NC score | -0.004245 (rank : 81) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 857 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BLF2, Q5M6W2, Q8BKR2, Q8BL49, Q8BVE0, Q8K134 | Gene names | Cdkl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase-like 3 (EC 2.7.11.22). | |||||