Please be patient as the page loads
|
HTF4_MOUSE
|
||||||
| SwissProt Accessions | Q61286 | Gene names | Tcf12, Alf1, Meb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4) (Class A helix-loop-helix transcription factor ME1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HTF4_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.990652 (rank : 2) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 111 | |
| SwissProt Accessions | Q99081 | Gene names | TCF12, HEB, HTF4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4). | |||||
|
HTF4_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 153 | |
| SwissProt Accessions | Q61286 | Gene names | Tcf12, Alf1, Meb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4) (Class A helix-loop-helix transcription factor ME1). | |||||
|
ITF2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.964384 (rank : 4) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P15884, Q15439, Q15440 | Gene names | TCF4, ITF2, SEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (SL3-3 enhancer factor 2) (SEF-2). | |||||
|
ITF2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.965029 (rank : 3) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q60722, Q60721, Q62211, Q64072, Q80UE8 | Gene names | Tcf4, Itf2, Sef2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (MITF-2) (SL3-3 enhancer factor 2) (SEF-2) (Class A helix-loop-helix transcription factor ME2). | |||||
|
TFE2_HUMAN
|
||||||
| θ value | 2.68277e-145 (rank : 5) | NC score | 0.953977 (rank : 6) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 53 | |
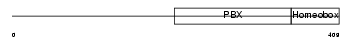
| SwissProt Accessions | P15923, P15883, Q14208, Q14635, Q14636, Q9UPI9 | Gene names | TCF3, E2A, ITF1 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Immunoglobulin transcription factor 1) (Transcription factor ITF-1) (Kappa-E2-binding factor). | |||||
|
TFE2_MOUSE
|
||||||
| θ value | 2.511e-143 (rank : 6) | NC score | 0.957399 (rank : 5) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P15806, Q3U153, Q8CAH9, Q8VCY4, Q922S2, Q99MB8, Q9CYJ4 | Gene names | Tcf3, Alf2, Me2, Tcfe2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Transcription factor A1). | |||||
|
MYF5_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 7) | NC score | 0.232154 (rank : 8) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 44 | |
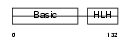
| SwissProt Accessions | P13349 | Gene names | MYF5 | |||
|
Domain Architecture |
|
|||||
| Description | Myogenic factor 5 (Myf-5). | |||||
|
MYF5_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 8) | NC score | 0.234931 (rank : 7) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 44 | |
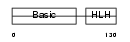
| SwissProt Accessions | P24699, Q543W7 | Gene names | Myf5, Myf-5 | |||
|
Domain Architecture |
|
|||||
| Description | Myogenic factor 5 (Myf-5). | |||||
|
ATOH1_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 9) | NC score | 0.224828 (rank : 9) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P48985 | Gene names | Atoh1, Ath1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein mATH-1) (MATH1). | |||||
|
NDF2_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 10) | NC score | 0.189217 (rank : 17) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q15784, Q8TBI7, Q9UQC6 | Gene names | NEUROD2, NDRF | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 2 (NeuroD2) (NeuroD-related factor) (NDRF). | |||||
|
NDF2_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 11) | NC score | 0.188949 (rank : 18) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q62414, Q61952, Q925V5 | Gene names | Neurod2, Ndrf | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 2 (NeuroD2) (NeuroD-related factor) (NDRF). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 12) | NC score | 0.049594 (rank : 88) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
NDF1_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 13) | NC score | 0.174083 (rank : 26) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 55 | |
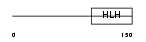
| SwissProt Accessions | Q13562, O00343, Q13340, Q5U095, Q96TH0, Q99455, Q9UEC8 | Gene names | NEUROD1, NEUROD | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenic differentiation factor 1 (NeuroD1) (NeuroD). | |||||
|
MYOD1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 14) | NC score | 0.211937 (rank : 11) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P15172 | Gene names | MYOD1, MYF3, MYOD | |||
|
Domain Architecture |
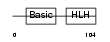
|
|||||
| Description | Myoblast determination protein 1 (Myogenic factor 3) (Myf-3). | |||||
|
MYOD1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 15) | NC score | 0.213523 (rank : 10) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P10085 | Gene names | Myod1, Myod | |||
|
Domain Architecture |
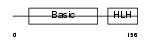
|
|||||
| Description | Myoblast determination protein 1. | |||||
|
MYOG_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 16) | NC score | 0.200549 (rank : 13) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P15173 | Gene names | MYOG, MYF4 | |||
|
Domain Architecture |
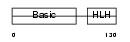
|
|||||
| Description | Myogenin (Myogenic factor 4) (Myf-4). | |||||
|
NDF1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 17) | NC score | 0.173779 (rank : 27) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q60867, Q545N9, Q60897 | Gene names | Neurod1, Neurod | |||
|
Domain Architecture |
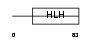
|
|||||
| Description | Neurogenic differentiation factor 1 (NeuroD1). | |||||
|
TWST2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 18) | NC score | 0.181154 (rank : 22) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8WVJ9 | Gene names | TWIST2, DERMO1 | |||
|
Domain Architecture |
|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
TWST2_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 19) | NC score | 0.181398 (rank : 21) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9D030 | Gene names | Twist2, Dermo1 | |||
|
Domain Architecture |
|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
ATOH1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 20) | NC score | 0.198770 (rank : 14) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q92858 | Gene names | ATOH1, ATH1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein hATH-1). | |||||
|
CROP_HUMAN
|
||||||
| θ value | 0.125558 (rank : 21) | NC score | 0.067393 (rank : 78) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95232, Q6PHR9, Q9NUY0, Q9P2S7 | Gene names | CROP, CREAP1, O48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein (cAMP regulatory element-associated protein 1) (CRE-associated protein 1) (CREAP-1) (Luc7A) (Okadaic acid-inducible phosphoprotein OA48-18). | |||||
|
CROP_MOUSE
|
||||||
| θ value | 0.125558 (rank : 22) | NC score | 0.068307 (rank : 76) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5SUF2, Q3U9D5, Q8BUJ5, Q921Z3, Q9CRS7, Q9CTY4 | Gene names | Crop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein. | |||||
|
SRGP2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 23) | NC score | 0.030618 (rank : 100) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O43295, Q8IX13, Q8IZV8 | Gene names | SRGAP3, ARHGAP14, KIAA0411, MEGAP, SRGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Mental disorder- associated GAP) (Rho-GTPase-activating protein 14). | |||||
|
NDF4_HUMAN
|
||||||
| θ value | 0.163984 (rank : 24) | NC score | 0.179756 (rank : 23) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9HD90 | Gene names | NEUROD4 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4). | |||||
|
NDF4_MOUSE
|
||||||
| θ value | 0.163984 (rank : 25) | NC score | 0.177074 (rank : 24) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O09105 | Gene names | Neurod4, Ath3, Atoh3 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4) (Atonal protein homolog 3) (Helix-loop-helix protein mATH-3) (MATH3). | |||||
|
MYOG_MOUSE
|
||||||
| θ value | 0.21417 (rank : 26) | NC score | 0.194579 (rank : 15) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P12979 | Gene names | Myog | |||
|
Domain Architecture |
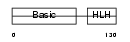
|
|||||
| Description | Myogenin (MYOD1-related protein). | |||||
|
NDF6_HUMAN
|
||||||
| θ value | 0.21417 (rank : 27) | NC score | 0.163430 (rank : 30) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96NK8, Q9H3H6 | Gene names | NEUROD6 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 6 (NeuroD6). | |||||
|
NDF6_MOUSE
|
||||||
| θ value | 0.21417 (rank : 28) | NC score | 0.163183 (rank : 31) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P48986 | Gene names | Neurod6, Ath2, Atoh2, Nex1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 6 (NeuroD6) (Atonal protein homolog 2) (Helix-loop-helix protein mATH-2) (MATH2) (NEX-1 protein). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 0.279714 (rank : 29) | NC score | 0.055694 (rank : 85) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
F10A1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 30) | NC score | 0.044191 (rank : 93) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P50502, O14999 | Gene names | ST13, FAM10A1, HIP, SNC6 | |||
|
Domain Architecture |
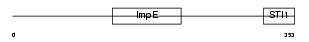
|
|||||
| Description | Hsc70-interacting protein (Hip) (Suppression of tumorigenicity protein 13) (Putative tumor suppressor ST13) (Protein FAM10A1) (Progesterone receptor-associated p48 protein) (NY-REN-33 antigen). | |||||
|
F10A5_HUMAN
|
||||||
| θ value | 0.279714 (rank : 31) | NC score | 0.044671 (rank : 91) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NFI4 | Gene names | FAM10A5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A5. | |||||
|
BSN_MOUSE
|
||||||
| θ value | 0.365318 (rank : 32) | NC score | 0.040579 (rank : 97) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CDSN_MOUSE
|
||||||
| θ value | 0.365318 (rank : 33) | NC score | 0.060383 (rank : 83) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7TPC1 | Gene names | Cdsn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor. | |||||
|
FIGLA_MOUSE
|
||||||
| θ value | 0.365318 (rank : 34) | NC score | 0.209031 (rank : 12) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O55208 | Gene names | Figla | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
ASCL2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 35) | NC score | 0.101284 (rank : 48) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q99929, Q9UM68 | Gene names | ASCL2 | |||
|
Domain Architecture |
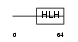
|
|||||
| Description | Achaete-scute homolog 2 (Mash2) (HASH2). | |||||
|
BHLH8_MOUSE
|
||||||
| θ value | 0.47712 (rank : 36) | NC score | 0.193259 (rank : 16) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9QYC3, Q9QYE4 | Gene names | Bhlhb8, Mist1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
FIBA_HUMAN
|
||||||
| θ value | 0.47712 (rank : 37) | NC score | 0.014049 (rank : 136) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P02671, Q9BX62, Q9UCH2 | Gene names | FGA | |||
|
Domain Architecture |

|
|||||
| Description | Fibrinogen alpha chain precursor [Contains: Fibrinopeptide A]. | |||||
|
MUC13_MOUSE
|
||||||
| θ value | 0.47712 (rank : 38) | NC score | 0.044838 (rank : 90) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 0.62314 (rank : 39) | NC score | 0.015568 (rank : 133) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
MYCN_HUMAN
|
||||||
| θ value | 0.62314 (rank : 40) | NC score | 0.077577 (rank : 70) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYF6_MOUSE
|
||||||
| θ value | 0.62314 (rank : 41) | NC score | 0.174371 (rank : 25) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P15375 | Gene names | Myf6, Myf-6 | |||
|
Domain Architecture |
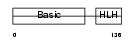
|
|||||
| Description | Myogenic factor 6 (Myf-6) (Herculin). | |||||
|
NUPL1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 42) | NC score | 0.057447 (rank : 84) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BVL2, O43160 | Gene names | NUPL1, KIAA0410 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin p58/p45 (Nucleoporin-like 1). | |||||
|
SRGP2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 43) | NC score | 0.023200 (rank : 111) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q812A2, Q80U09, Q8BKP4, Q8BLD0, Q925I2 | Gene names | Srgap3, Arhgap14, Kiaa0411, Srgap2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Rho-GTPase-activating protein 14). | |||||
|
TWST1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 44) | NC score | 0.181737 (rank : 19) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q15672, Q92487, Q99804 | Gene names | TWIST1, TWIST | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (H-twist). | |||||
|
TWST1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 45) | NC score | 0.181473 (rank : 20) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P26687 | Gene names | Twist1, Twist | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (M-twist). | |||||
|
ZKSC1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 46) | NC score | 0.003271 (rank : 172) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P17029, P52745, Q8TBW5, Q8TEK7 | Gene names | ZKSCAN1, KOX18, ZNF139, ZNF36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1 (Zinc finger protein 36) (Zinc finger protein KOX18). | |||||
|
AMOL1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 47) | NC score | 0.019287 (rank : 121) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
BAZ2A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 48) | NC score | 0.043850 (rank : 94) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
GPS2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 49) | NC score | 0.044548 (rank : 92) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13227 | Gene names | GPS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein pathway suppressor 2 (Protein GPS2). | |||||
|
HAND2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 50) | NC score | 0.148136 (rank : 37) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P61296, O95300, O95301, P97833 | Gene names | HAND2, DHAND | |||
|
Domain Architecture |
|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 2 (Deciduum, heart, autonomic nervous system and neural crest derivatives-expressed protein 2) (dHAND). | |||||
|
HAND2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 51) | NC score | 0.148136 (rank : 38) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q61039, Q61100 | Gene names | Hand2, Dhand, Hed, Thing2 | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 2 (Deciduum, heart, autonomic nervous system and neural crest derivatives-expressed protein 2) (dHAND) (Helix-loop-helix transcription factor expressed in embryo and deciduum) (Thing-2). | |||||
|
KI67_HUMAN
|
||||||
| θ value | 0.813845 (rank : 52) | NC score | 0.036396 (rank : 98) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
MYF6_HUMAN
|
||||||
| θ value | 0.813845 (rank : 53) | NC score | 0.173772 (rank : 28) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P23409, Q53X80, Q6FHI9 | Gene names | MYF6 | |||
|
Domain Architecture |
|
|||||
| Description | Myogenic factor 6 (Myf-6). | |||||
|
NCOA6_MOUSE
|
||||||
| θ value | 0.813845 (rank : 54) | NC score | 0.040680 (rank : 96) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
AHTF1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 55) | NC score | 0.024851 (rank : 105) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
BCAR1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 56) | NC score | 0.034695 (rank : 99) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
BHLH8_HUMAN
|
||||||
| θ value | 1.06291 (rank : 57) | NC score | 0.171797 (rank : 29) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q7RTS1 | Gene names | BHLHB8, MIST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
HEN2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 58) | NC score | 0.161638 (rank : 33) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q02577 | Gene names | NHLH2, HEN2 | |||
|
Domain Architecture |
|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
HEN2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 59) | NC score | 0.161482 (rank : 34) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q64221 | Gene names | Nhlh2, Hen2 | |||
|
Domain Architecture |
|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
KIF5A_HUMAN
|
||||||
| θ value | 1.06291 (rank : 60) | NC score | 0.012875 (rank : 143) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q12840, Q4LE26 | Gene names | KIF5A, NKHC1 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
ZBTB6_HUMAN
|
||||||
| θ value | 1.06291 (rank : 61) | NC score | 0.007318 (rank : 160) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 805 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q15916 | Gene names | ZBTB6, ZID, ZNF482 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 6 (Zinc finger protein 482) (Zinc finger protein with interaction domain). | |||||
|
DHX8_HUMAN
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.015234 (rank : 134) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14562 | Gene names | DHX8, DDX8 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DHX8 (EC 3.6.1.-) (DEAH box protein 8) (RNA helicase HRH1). | |||||
|
HEN1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 63) | NC score | 0.160601 (rank : 35) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q02575 | Gene names | NHLH1, HEN1 | |||
|
Domain Architecture |
|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
HEN1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 64) | NC score | 0.162928 (rank : 32) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q02576 | Gene names | Nhlh1, Hen1 | |||
|
Domain Architecture |
|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
INVS_HUMAN
|
||||||
| θ value | 1.38821 (rank : 65) | NC score | 0.015568 (rank : 132) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y283, Q5W0T6, Q8IVX8, Q9BRB9, Q9Y488, Q9Y498 | Gene names | INVS, INV, NPHP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning homolog) (Nephrocystin-2). | |||||
|
MYCN_MOUSE
|
||||||
| θ value | 1.38821 (rank : 66) | NC score | 0.070801 (rank : 74) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
OLIG2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 67) | NC score | 0.093398 (rank : 57) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9EQW6, Q9JKN4 | Gene names | Olig2 | |||
|
Domain Architecture |

|
|||||
| Description | Oligodendrocyte transcription factor 2 (Oligo2). | |||||
|
SI1L1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 68) | NC score | 0.013001 (rank : 142) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C0T5, Q6PDI8, Q80U02, Q8C026 | Gene names | Sipa1l1, Kiaa0440 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 1. | |||||
|
WT1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 69) | NC score | 0.005330 (rank : 167) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P19544, Q15881, Q16256, Q16575, Q8IYZ5 | Gene names | WT1 | |||
|
Domain Architecture |

|
|||||
| Description | Wilms' tumor protein (WT33). | |||||
|
BRE1A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 70) | NC score | 0.018453 (rank : 127) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1071 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5VTR2, Q69YL5, Q6P527, Q8N3J4, Q96JD3, Q9H9Y7, Q9HA51, Q9NUR4, Q9NWQ3, Q9NX83 | Gene names | RNF20, BRE1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1A (EC 6.3.2.-) (BRE1-A) (hBRE1) (RING finger protein 20). | |||||
|
EYA1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 71) | NC score | 0.020511 (rank : 118) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97767, O08818 | Gene names | Eya1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
EZRI_MOUSE
|
||||||
| θ value | 1.81305 (rank : 72) | NC score | 0.008242 (rank : 158) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P26040, Q80ZT8, Q9DCI1 | Gene names | Vil2 | |||
|
Domain Architecture |

|
|||||
| Description | Ezrin (p81) (Cytovillin) (Villin-2). | |||||
|
FBX24_MOUSE
|
||||||
| θ value | 1.81305 (rank : 73) | NC score | 0.045140 (rank : 89) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D417 | Gene names | Fbxo24, Fbx24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 24. | |||||
|
HORN_HUMAN
|
||||||
| θ value | 1.81305 (rank : 74) | NC score | 0.074054 (rank : 72) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86YZ3 | Gene names | HRNR, S100A18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hornerin. | |||||
|
K0157_MOUSE
|
||||||
| θ value | 1.81305 (rank : 75) | NC score | 0.029425 (rank : 101) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3TCJ1, Q3TB08, Q8K0R4 | Gene names | Kiaa0157 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0157. | |||||
|
ZBTB6_MOUSE
|
||||||
| θ value | 1.81305 (rank : 76) | NC score | 0.006680 (rank : 161) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K088, Q8BLM4 | Gene names | Zbtb6, Zfp482, Znf482 | |||
|
Domain Architecture |
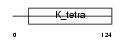
|
|||||
| Description | Zinc finger and BTB domain-containing protein 6 (Zinc finger protein 482). | |||||
|
ZN750_MOUSE
|
||||||
| θ value | 1.81305 (rank : 77) | NC score | 0.029284 (rank : 102) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BH05, Q66JP3, Q8C0L1 | Gene names | Znf750 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF750. | |||||
|
CLCN1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 78) | NC score | 0.010892 (rank : 153) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35523 | Gene names | CLCN1, CLC1 | |||
|
Domain Architecture |

|
|||||
| Description | Chloride channel protein, skeletal muscle (Chloride channel protein 1) (ClC-1). | |||||
|
MTR1L_HUMAN
|
||||||
| θ value | 2.36792 (rank : 79) | NC score | 0.002110 (rank : 175) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1180 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13585 | Gene names | GPR50 | |||
|
Domain Architecture |

|
|||||
| Description | Melatonin-related receptor (G protein-coupled receptor 50) (H9). | |||||
|
NU153_HUMAN
|
||||||
| θ value | 2.36792 (rank : 80) | NC score | 0.027062 (rank : 103) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
OLIG2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 81) | NC score | 0.090717 (rank : 58) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q13516, Q86X04, Q9NZ14 | Gene names | OLIG2, BHLHB1, PRKCBP2, RACK17 | |||
|
Domain Architecture |

|
|||||
| Description | Oligodendrocyte transcription factor 2 (Oligo2) (Basic helix-loop- helix protein class B 1) (Protein kinase C-binding protein RACK17) (Protein kinase C-binding protein 2). | |||||
|
PGCA_HUMAN
|
||||||
| θ value | 2.36792 (rank : 82) | NC score | 0.022972 (rank : 113) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
RAD50_MOUSE
|
||||||
| θ value | 2.36792 (rank : 83) | NC score | 0.019978 (rank : 119) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
TAL1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 84) | NC score | 0.146657 (rank : 41) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P17542 | Gene names | TAL1, SCL, TCL5 | |||
|
Domain Architecture |
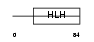
|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein (TAL-1 protein) (Stem cell protein) (T-cell leukemia/lymphoma-5 protein). | |||||
|
TAL1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 85) | NC score | 0.147244 (rank : 40) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P22091 | Gene names | Tal1, Scl, Tal-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein homolog (TAL-1 protein) (Stem cell protein). | |||||
|
TRIM2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 86) | NC score | 0.011782 (rank : 149) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9C040, O60272, Q9BSI9, Q9UFZ1 | Gene names | TRIM2, KIAA0517, RNF86 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 2 (RING finger protein 86). | |||||
|
TRIM2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 87) | NC score | 0.011824 (rank : 148) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ESN6, Q3UHP5, Q8C981 | Gene names | Trim2, Narf | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 2 (Neural activity-related RING finger protein). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 88) | NC score | 0.023803 (rank : 109) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 3.0926 (rank : 89) | NC score | 0.023907 (rank : 108) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
BRE1A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 90) | NC score | 0.019347 (rank : 120) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1025 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5DTM8, Q3UT10, Q3V350, Q7TT11, Q8BKA8, Q8BKN8, Q8BUF7, Q8BVU4 | Gene names | Rnf20, Bre1a, Kiaa4116 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1A (EC 6.3.2.-) (BRE1-A) (RING finger protein 20). | |||||
|
FIGLA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 91) | NC score | 0.150950 (rank : 36) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q6QHK4 | Gene names | FIGLA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
GLI1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 92) | NC score | 0.004359 (rank : 171) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P47806, Q9QYK1 | Gene names | Gli1, Gli | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI1 (Glioma-associated oncogene homolog). | |||||
|
HIRP3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 93) | NC score | 0.020595 (rank : 117) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BW71, O75707, O75708 | Gene names | HIRIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIRA-interacting protein 3. | |||||
|
IRS2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 94) | NC score | 0.021502 (rank : 116) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P81122 | Gene names | Irs2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin receptor substrate 2 (IRS-2) (4PS). | |||||
|
KIF5A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 95) | NC score | 0.011828 (rank : 147) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P33175, Q5DTP1, Q6PDY7, Q9Z2F9 | Gene names | Kif5a, Kiaa4086, Kif5, Nkhc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
LYL1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 96) | NC score | 0.147927 (rank : 39) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P12980, O76102 | Gene names | LYL1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
SPEG_HUMAN
|
||||||
| θ value | 3.0926 (rank : 97) | NC score | -0.000149 (rank : 179) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
TB182_HUMAN
|
||||||
| θ value | 3.0926 (rank : 98) | NC score | 0.054393 (rank : 86) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9C0C2 | Gene names | TNKS1BP1, KIAA1741, TAB182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 182 kDa tankyrase 1-binding protein. | |||||
|
TFE3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 99) | NC score | 0.051054 (rank : 87) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
XKR5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 100) | NC score | 0.022479 (rank : 115) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6UX68, Q5GH74 | Gene names | XKR5, XRG5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | XK-related protein 5. | |||||
|
DLG5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.008680 (rank : 157) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8TDM6, Q5T1H7, Q5T1H8, Q6DKG3, Q86WC0, Q8TDM7, Q9UE73, Q9Y4E3 | Gene names | DLG5, KIAA0583, PDLG | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 5 (Placenta and prostate DLG) (Discs large protein P-dlg). | |||||
|
DUS16_HUMAN
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.011650 (rank : 150) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BY84, Q9C0G3 | Gene names | DUSP16, KIAA1700, MKP7 | |||
|
Domain Architecture |
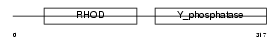
|
|||||
| Description | Dual specificity protein phosphatase 16 (EC 3.1.3.48) (EC 3.1.3.16) (Mitogen-activated protein kinase phosphatase 7) (MAP kinase phosphatase 7) (MKP-7). | |||||
|
GLP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 103) | NC score | 0.042722 (rank : 95) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P14220 | Gene names | Gypa | |||
|
Domain Architecture |

|
|||||
| Description | Glycophorin. | |||||
|
PPIG_HUMAN
|
||||||
| θ value | 4.03905 (rank : 104) | NC score | 0.013441 (rank : 141) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
SMRC1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 105) | NC score | 0.018804 (rank : 125) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P97496, Q7TS80, Q7TT29 | Gene names | Smarcc1, Baf155, Srg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155) (SWI3-related protein). | |||||
|
TARA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 106) | NC score | 0.023206 (rank : 110) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
WT1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 107) | NC score | 0.003023 (rank : 173) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P22561 | Gene names | Wt1, Wt-1 | |||
|
Domain Architecture |
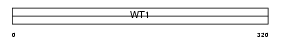
|
|||||
| Description | Wilms' tumor protein homolog. | |||||
|
ARHGA_MOUSE
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.007331 (rank : 159) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C033, Q5DU38, Q80VH8, Q8BW76, Q922S7 | Gene names | Arhgef10, Kiaa0294 | |||
|
Domain Architecture |
|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
COIA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 109) | NC score | 0.015002 (rank : 135) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P39060, Q9UK38, Q9Y6Q7, Q9Y6Q8 | Gene names | COL18A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
K1267_MOUSE
|
||||||
| θ value | 5.27518 (rank : 110) | NC score | 0.018967 (rank : 123) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TG1, Q3TT88, Q3U5D8, Q3V3N3, Q7TMU3, Q80XP7, Q8R3L6, Q9D9G0 | Gene names | Kiaa1267 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1267. | |||||
|
LYL1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 111) | NC score | 0.141385 (rank : 42) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P27792 | Gene names | Lyl1, Lyl-1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 5.27518 (rank : 112) | NC score | 0.016747 (rank : 131) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NCOA3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | 0.012378 (rank : 144) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NOP14_HUMAN
|
||||||
| θ value | 5.27518 (rank : 114) | NC score | 0.017392 (rank : 129) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
PRD16_HUMAN
|
||||||
| θ value | 5.27518 (rank : 115) | NC score | 0.001240 (rank : 176) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 832 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9HAZ2, Q8WYJ9, Q9C0I8 | Gene names | PRDM16, KIAA1675, MEL1, PFM13 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 16 (PR domain-containing protein 16) (Transcription factor MEL1). | |||||
|
PYGO1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 116) | NC score | 0.025106 (rank : 104) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y3Y4 | Gene names | PYGO1 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 1. | |||||
|
STX4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 117) | NC score | 0.011879 (rank : 146) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q12846, Q15525 | Gene names | STX4, STX4A | |||
|
Domain Architecture |
|
|||||
| Description | Syntaxin-4 (NY-REN-31 antigen). | |||||
|
CJ022_HUMAN
|
||||||
| θ value | 6.88961 (rank : 118) | NC score | 0.024495 (rank : 106) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96SZ5 | Gene names | C10orf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf22. | |||||
|
COHA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 119) | NC score | 0.022863 (rank : 114) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UMD9, Q02802, Q99018, Q9NQK9, Q9UC14 | Gene names | COL17A1, BP180, BPAG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
ENAH_HUMAN
|
||||||
| θ value | 6.88961 (rank : 120) | NC score | 0.009875 (rank : 155) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N8S7, Q502W5, Q5T5M7, Q5VTQ9, Q5VTR0, Q9NVF3, Q9UFB8 | Gene names | ENAH, MENA | |||
|
Domain Architecture |
|
|||||
| Description | Protein enabled homolog. | |||||
|
HXA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 121) | NC score | 0.001027 (rank : 177) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49639, O43363 | Gene names | HOXA1, HOX1F | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A1 (Hox-1F). | |||||
|
K1267_HUMAN
|
||||||
| θ value | 6.88961 (rank : 122) | NC score | 0.018019 (rank : 128) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z3B3, Q6AW85, Q8IYH1, Q9NTE7, Q9UFT0, Q9ULF3 | Gene names | KIAA1267 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1267. | |||||
|
RERE_MOUSE
|
||||||
| θ value | 6.88961 (rank : 123) | NC score | 0.024225 (rank : 107) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
RFX1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 124) | NC score | 0.011508 (rank : 152) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P48377 | Gene names | Rfx1 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein RFX1. | |||||
|
SAKS1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 125) | NC score | 0.010099 (rank : 154) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q922Y1, Q3UCP8 | Gene names | Saks1, D19Ertd721e | |||
|
Domain Architecture |

|
|||||
| Description | SAPK substrate protein 1 (UBA/UBX 33.3 kDa protein) (mY33K) (Protein 2B28). | |||||
|
SC24C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 126) | NC score | 0.018732 (rank : 126) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
SMRC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 127) | NC score | 0.013679 (rank : 139) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92922, Q6P172, Q8IWH2 | Gene names | SMARCC1, BAF155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155). | |||||
|
TTLL4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 128) | NC score | 0.012067 (rank : 145) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80UG8, Q5U4C4 | Gene names | Ttll4, Kiaa0173 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 4. | |||||
|
ZBT38_HUMAN
|
||||||
| θ value | 6.88961 (rank : 129) | NC score | 0.000861 (rank : 178) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 885 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NAP3 | Gene names | ZBTB38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 38. | |||||
|
ANS4B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 130) | NC score | 0.005894 (rank : 165) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K3X6, Q9D8A5 | Gene names | Anks4b, Harp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 4B (Harmonin- interacting ankyrin repeat-containing protein) (Harp). | |||||
|
ASH2L_MOUSE
|
||||||
| θ value | 8.99809 (rank : 131) | NC score | 0.009125 (rank : 156) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91X20, Q3UIF9, Q9Z2X4 | Gene names | Ash2l | |||
|
Domain Architecture |

|
|||||
| Description | Set1/Ash2 histone methyltransferase complex subunit ASH2 (ASH2-like protein). | |||||
|
BHLH4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 132) | NC score | 0.108836 (rank : 47) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8BGW3, Q71MB7 | Gene names | Bhlhb4, H20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 4 (bHLHB4). | |||||
|
CDKL5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.006423 (rank : 162) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O76039, Q14198, Q5H985, Q9UJL6 | Gene names | CDKL5, STK9 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase-like 5 (EC 2.7.11.22) (Serine/threonine- protein kinase 9). | |||||
|
CE152_HUMAN
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | 0.019253 (rank : 122) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O94986, Q6NTA0 | Gene names | CEP152, KIAA0912 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 152 kDa (Cep152 protein). | |||||
|
DCDC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 135) | NC score | 0.013484 (rank : 140) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
FXR2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.005947 (rank : 164) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WVR4, Q9WVR5 | Gene names | Fxr2, Fxr2h | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
KCNH1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.005207 (rank : 170) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95259, O76035 | Gene names | KCNH1, EAG, EAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (hEAG1) (h-eag). | |||||
|
KCNH1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.005214 (rank : 169) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60603 | Gene names | Kcnh1, Eag | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (EAG1) (m-eag). | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.017378 (rank : 130) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
MUSC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.112091 (rank : 46) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O60682, O75946, Q9BRE7 | Gene names | MSC, ABF1 | |||
|
Domain Architecture |

|
|||||
| Description | Musculin (Activated B-cell factor 1) (ABF-1). | |||||
|
MUSC_MOUSE
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.113191 (rank : 45) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O88940 | Gene names | Msc, Myor | |||
|
Domain Architecture |

|
|||||
| Description | Musculin (Myogenic repressor). | |||||
|
NUPL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.011529 (rank : 151) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K2K6, O70448, Q8BQL5, Q8CDK9 | Gene names | Hrb, Rip | |||
|
Domain Architecture |
|
|||||
| Description | Nucleoporin-like protein RIP (HIV-1 Rev-binding protein homolog). | |||||
|
OLIG1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.090019 (rank : 59) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8TAK6, Q7RTS0 | Gene names | OLIG1 | |||
|
Domain Architecture |
|
|||||
| Description | Oligodendrocyte transcription factor 1 (Oligo1). | |||||
|
PTF1A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.115596 (rank : 43) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q7RTS3, Q9HC25 | Gene names | PTF1A, PTF1P48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pancreas transcription factor 1 subunit alpha (Pancreas-specific transcription factor 1a) (bHLH transcription factor p48) (p48 DNA- binding subunit of transcription factor PTF1) (PTF1-p48). | |||||
|
PTF1A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.114588 (rank : 44) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9QX98, Q9QYF5, Q9QYF6 | Gene names | Ptf1a, Ptf1p48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pancreas transcription factor 1 subunit alpha (Pancreas-specific transcription factor 1a) (bHLH transcription factor p48) (p48 DNA- binding subunit of transcription factor PTF1) (PTF1-p48). | |||||
|
PTPRA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | 0.003017 (rank : 174) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P18433, Q14513, Q7Z2I2, Q96TD9 | Gene names | PTPRA, PTPA | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase alpha precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase alpha) (R-PTP-alpha). | |||||
|
REN3B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 147) | NC score | 0.018942 (rank : 124) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BZI7, Q9H1J0 | Gene names | UPF3B, RENT3B, UPF3X | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 3B (Nonsense mRNA reducing factor 3B) (Up-frameshift suppressor 3 homolog B) (hUpf3B) (hUpf3p-X). | |||||
|
RERE_HUMAN
|
||||||
| θ value | 8.99809 (rank : 148) | NC score | 0.023027 (rank : 112) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
SFR12_MOUSE
|
||||||
| θ value | 8.99809 (rank : 149) | NC score | 0.013827 (rank : 138) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
SOX9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 150) | NC score | 0.005318 (rank : 168) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q04887, Q8C7L2, Q91ZK2, Q99KQ0 | Gene names | Sox9, Sox-9 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-9. | |||||
|
SSH1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 151) | NC score | 0.005847 (rank : 166) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WYL5, Q6P6C0, Q8N9A7, Q8WYL3, Q8WYL4, Q9P2P8 | Gene names | SSH1, KIAA1298, SSH1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 1 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-1L) (hSSH-1L). | |||||
|
ST18_HUMAN
|
||||||
| θ value | 8.99809 (rank : 152) | NC score | 0.006018 (rank : 163) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O60284 | Gene names | ST18, KIAA0535, ZNF387 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18 (Zinc finger protein 387). | |||||
|
TP53B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 153) | NC score | 0.014010 (rank : 137) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
ASCL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.076677 (rank : 71) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P50553, Q9BQ30 | Gene names | ASCL1, ASH1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 1 (HASH1). | |||||
|
ASCL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.077582 (rank : 69) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q02067 | Gene names | Ascl1, Ash1, Mash-1, Mash1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 1 (Mash-1). | |||||
|
ASCL2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.080182 (rank : 67) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O35885, Q9WUJ7 | Gene names | Ascl2, Mash2 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 2 (Mash-2). | |||||
|
ASCL3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.080593 (rank : 66) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9NQ33, Q8WYQ6 | Gene names | ASCL3, HASH3, SGN1 | |||
|
Domain Architecture |
|
|||||
| Description | Achaete-scute homolog 3 (bHLH transcriptional regulator Sgn-1). | |||||
|
ASCL3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.083664 (rank : 63) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JJR7 | Gene names | Ascl3, Mash3, Sgn1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 3 (bHLH transcriptional regulator Sgn-1) (Mash- 3). | |||||
|
BHLH4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.097290 (rank : 51) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NDY6 | Gene names | BHLHB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 4 (bHLHB4). | |||||
|
HAND1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.083041 (rank : 64) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O96004 | Gene names | HAND1, EHAND | |||
|
Domain Architecture |
|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND). | |||||
|
HAND1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.084359 (rank : 62) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q64279, Q61099 | Gene names | Hand1, Ehand, Hxt, Thing1 | |||
|
Domain Architecture |
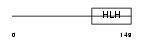
|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND) (Helix-loop-helix transcription factor expressed in extraembryonic mesoderm and trophoblast) (Thing-1) (Th1). | |||||
|
HORN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.064650 (rank : 82) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
NGN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.097705 (rank : 50) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q92886, Q96HE1 | Gene names | NEUROG1, NEUROD3, NGN, NGN1 | |||
|
Domain Architecture |
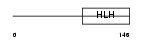
|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein). | |||||
|
NGN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.100478 (rank : 49) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P70660 | Gene names | Neurog1, Ath4c, Neurod3, Ngn, Ngn1 | |||
|
Domain Architecture |
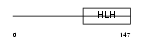
|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein) (Helix-loop-helix protein mATH-4C) (MATH4C). | |||||
|
NGN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.093798 (rank : 55) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9H2A3, Q8N416 | Gene names | NEUROG2, NGN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurogenin 2. | |||||
|
NGN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.093568 (rank : 56) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P70447, P70237 | Gene names | Neurog2, Ath4a, Atoh4, Ngn2 | |||
|
Domain Architecture |
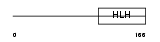
|
|||||
| Description | Neurogenin 2 (Atonal protein homolog 4) (Helix-loop-helix protein mATH-4A) (MATH4A). | |||||
|
NGN3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.094587 (rank : 53) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y4Z2, Q9BY24 | Gene names | NEUROG3, NGN3 | |||
|
Domain Architecture |
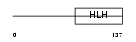
|
|||||
| Description | Neurogenin 3. | |||||
|
NGN3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.095315 (rank : 52) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P70661 | Gene names | Neurog3, Ath4b, Atoh5, Ngn3 | |||
|
Domain Architecture |
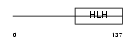
|
|||||
| Description | Neurogenin 3 (Atonal protein homolog 5) (Helix-loop-helix protein mATH-4B) (MATH4B). | |||||
|
OLIG1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.066376 (rank : 80) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JKN5 | Gene names | Olig1 | |||
|
Domain Architecture |
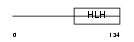
|
|||||
| Description | Oligodendrocyte transcription factor 1 (Oligo1) (Oligodendrocyte- specific bHLH transcription factor 1). | |||||
|
OLIG3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.067303 (rank : 79) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q7RTU3, Q8N8Q0 | Gene names | OLIG3, BHLHB7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oligodendrocyte transcription factor 3 (Oligo3) (Basic helix-loop- helix domain-containing class B protein 7). | |||||
|
OLIG3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.068253 (rank : 77) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6PFG8, Q9EQW5 | Gene names | Olig3, Bhlhb7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oligodendrocyte transcription factor 3 (Oligo3) (Oligodendrocyte- specific bHLH transcription factor 3) (Basic helix-loop-helix domain- containing class B protein 7). | |||||
|
SCX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.094419 (rank : 54) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q64124 | Gene names | Scx | |||
|
Domain Architecture |
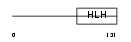
|
|||||
| Description | Basic helix-loop-helix transcription factor scleraxis. | |||||
|
TAL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.088770 (rank : 61) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q16559 | Gene names | TAL2 | |||
|
Domain Architecture |
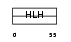
|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein (TAL-2 protein). | |||||
|
TAL2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.089888 (rank : 60) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q62282 | Gene names | Tal2 | |||
|
Domain Architecture |
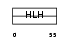
|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein homolog (TAL-2 protein). | |||||
|
TCF15_HUMAN
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.078536 (rank : 68) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q12870, Q9NQQ1 | Gene names | TCF15, BHLHEC2 | |||
|
Domain Architecture |
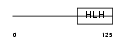
|
|||||
| Description | Transcription factor 15 (bHLH-EC2 protein) (Paraxis). | |||||
|
TCF15_MOUSE
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.082910 (rank : 65) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q60756, Q60788 | Gene names | Tcf15, Bhlhec2 | |||
|
Domain Architecture |
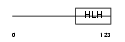
|
|||||
| Description | Transcription factor 15 (bHLH-EC2 protein) (Paraxis) (Meso1). | |||||
|
TCF21_HUMAN
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.070825 (rank : 73) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O43680, O43545, Q9BZ14 | Gene names | TCF21, POD1 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor 21 (Podocyte-expressed 1) (Pod-1) (Epicardin) (Capsulin). | |||||
|
TCF21_MOUSE
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.070449 (rank : 75) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O35437, Q3U023 | Gene names | Tcf21, Pod1 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor 21 (Podocyte-expressed 1) (Pod-1) (Epicardin) (Capsulin). | |||||
|
TFAP4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.065694 (rank : 81) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
HTF4_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 153 | |
| SwissProt Accessions | Q61286 | Gene names | Tcf12, Alf1, Meb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4) (Class A helix-loop-helix transcription factor ME1). | |||||
|
HTF4_HUMAN
|
||||||
| NC score | 0.990652 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 111 | |
| SwissProt Accessions | Q99081 | Gene names | TCF12, HEB, HTF4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4). | |||||
|
ITF2_MOUSE
|
||||||
| NC score | 0.965029 (rank : 3) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q60722, Q60721, Q62211, Q64072, Q80UE8 | Gene names | Tcf4, Itf2, Sef2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (MITF-2) (SL3-3 enhancer factor 2) (SEF-2) (Class A helix-loop-helix transcription factor ME2). | |||||
|
ITF2_HUMAN
|
||||||
| NC score | 0.964384 (rank : 4) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P15884, Q15439, Q15440 | Gene names | TCF4, ITF2, SEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (SL3-3 enhancer factor 2) (SEF-2). | |||||
|
TFE2_MOUSE
|
||||||
| NC score | 0.957399 (rank : 5) | θ value | 2.511e-143 (rank : 6) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P15806, Q3U153, Q8CAH9, Q8VCY4, Q922S2, Q99MB8, Q9CYJ4 | Gene names | Tcf3, Alf2, Me2, Tcfe2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Transcription factor A1). | |||||
|
TFE2_HUMAN
|
||||||
| NC score | 0.953977 (rank : 6) | θ value | 2.68277e-145 (rank : 5) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P15923, P15883, Q14208, Q14635, Q14636, Q9UPI9 | Gene names | TCF3, E2A, ITF1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Immunoglobulin transcription factor 1) (Transcription factor ITF-1) (Kappa-E2-binding factor). | |||||
|
MYF5_MOUSE
|
||||||
| NC score | 0.234931 (rank : 7) | θ value | 0.00869519 (rank : 8) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P24699, Q543W7 | Gene names | Myf5, Myf-5 | |||
|
Domain Architecture |

|
|||||
| Description | Myogenic factor 5 (Myf-5). | |||||
|
MYF5_HUMAN
|
||||||
| NC score | 0.232154 (rank : 8) | θ value | 0.00869519 (rank : 7) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P13349 | Gene names | MYF5 | |||
|
Domain Architecture |

|
|||||
| Description | Myogenic factor 5 (Myf-5). | |||||
|
ATOH1_MOUSE
|
||||||
| NC score | 0.224828 (rank : 9) | θ value | 0.0148317 (rank : 9) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P48985 | Gene names | Atoh1, Ath1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein mATH-1) (MATH1). | |||||
|
MYOD1_MOUSE
|
||||||
| NC score | 0.213523 (rank : 10) | θ value | 0.0736092 (rank : 15) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P10085 | Gene names | Myod1, Myod | |||
|
Domain Architecture |

|
|||||
| Description | Myoblast determination protein 1. | |||||
|
MYOD1_HUMAN
|
||||||
| NC score | 0.211937 (rank : 11) | θ value | 0.0736092 (rank : 14) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P15172 | Gene names | MYOD1, MYF3, MYOD | |||
|
Domain Architecture |

|
|||||
| Description | Myoblast determination protein 1 (Myogenic factor 3) (Myf-3). | |||||
|
FIGLA_MOUSE
|
||||||
| NC score | 0.209031 (rank : 12) | θ value | 0.365318 (rank : 34) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O55208 | Gene names | Figla | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
MYOG_HUMAN
|
||||||
| NC score | 0.200549 (rank : 13) | θ value | 0.0736092 (rank : 16) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P15173 | Gene names | MYOG, MYF4 | |||
|
Domain Architecture |

|
|||||
| Description | Myogenin (Myogenic factor 4) (Myf-4). | |||||
|
ATOH1_HUMAN
|
||||||
| NC score | 0.198770 (rank : 14) | θ value | 0.125558 (rank : 20) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q92858 | Gene names | ATOH1, ATH1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein hATH-1). | |||||
|
MYOG_MOUSE
|
||||||
| NC score | 0.194579 (rank : 15) | θ value | 0.21417 (rank : 26) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P12979 | Gene names | Myog | |||
|
Domain Architecture |

|
|||||
| Description | Myogenin (MYOD1-related protein). | |||||
|
BHLH8_MOUSE
|
||||||
| NC score | 0.193259 (rank : 16) | θ value | 0.47712 (rank : 36) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9QYC3, Q9QYE4 | Gene names | Bhlhb8, Mist1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
NDF2_HUMAN
|
||||||
| NC score | 0.189217 (rank : 17) | θ value | 0.0431538 (rank : 10) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q15784, Q8TBI7, Q9UQC6 | Gene names | NEUROD2, NDRF | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 2 (NeuroD2) (NeuroD-related factor) (NDRF). | |||||
|
NDF2_MOUSE
|
||||||
| NC score | 0.188949 (rank : 18) | θ value | 0.0431538 (rank : 11) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q62414, Q61952, Q925V5 | Gene names | Neurod2, Ndrf | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 2 (NeuroD2) (NeuroD-related factor) (NDRF). | |||||
|
TWST1_HUMAN
|
||||||
| NC score | 0.181737 (rank : 19) | θ value | 0.62314 (rank : 44) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q15672, Q92487, Q99804 | Gene names | TWIST1, TWIST | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (H-twist). | |||||
|
TWST1_MOUSE
|
||||||
| NC score | 0.181473 (rank : 20) | θ value | 0.62314 (rank : 45) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P26687 | Gene names | Twist1, Twist | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (M-twist). | |||||
|
TWST2_MOUSE
|
||||||
| NC score | 0.181398 (rank : 21) | θ value | 0.0961366 (rank : 19) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9D030 | Gene names | Twist2, Dermo1 | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
TWST2_HUMAN
|
||||||
| NC score | 0.181154 (rank : 22) | θ value | 0.0961366 (rank : 18) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8WVJ9 | Gene names | TWIST2, DERMO1 | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
NDF4_HUMAN
|
||||||
| NC score | 0.179756 (rank : 23) | θ value | 0.163984 (rank : 24) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9HD90 | Gene names | NEUROD4 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4). | |||||
|
NDF4_MOUSE
|
||||||
| NC score | 0.177074 (rank : 24) | θ value | 0.163984 (rank : 25) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O09105 | Gene names | Neurod4, Ath3, Atoh3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4) (Atonal protein homolog 3) (Helix-loop-helix protein mATH-3) (MATH3). | |||||
|
MYF6_MOUSE
|
||||||
| NC score | 0.174371 (rank : 25) | θ value | 0.62314 (rank : 41) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P15375 | Gene names | Myf6, Myf-6 | |||
|
Domain Architecture |

|
|||||
| Description | Myogenic factor 6 (Myf-6) (Herculin). | |||||
|
NDF1_HUMAN
|
||||||
| NC score | 0.174083 (rank : 26) | θ value | 0.0563607 (rank : 13) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q13562, O00343, Q13340, Q5U095, Q96TH0, Q99455, Q9UEC8 | Gene names | NEUROD1, NEUROD | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 1 (NeuroD1) (NeuroD). | |||||
|
NDF1_MOUSE
|
||||||
| NC score | 0.173779 (rank : 27) | θ value | 0.0736092 (rank : 17) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q60867, Q545N9, Q60897 | Gene names | Neurod1, Neurod | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 1 (NeuroD1). | |||||
|
MYF6_HUMAN
|
||||||
| NC score | 0.173772 (rank : 28) | θ value | 0.813845 (rank : 53) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P23409, Q53X80, Q6FHI9 | Gene names | MYF6 | |||
|
Domain Architecture |

|
|||||
| Description | Myogenic factor 6 (Myf-6). | |||||
|
BHLH8_HUMAN
|
||||||
| NC score | 0.171797 (rank : 29) | θ value | 1.06291 (rank : 57) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q7RTS1 | Gene names | BHLHB8, MIST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
NDF6_HUMAN
|
||||||
| NC score | 0.163430 (rank : 30) | θ value | 0.21417 (rank : 27) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96NK8, Q9H3H6 | Gene names | NEUROD6 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 6 (NeuroD6). | |||||
|
NDF6_MOUSE
|
||||||
| NC score | 0.163183 (rank : 31) | θ value | 0.21417 (rank : 28) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P48986 | Gene names | Neurod6, Ath2, Atoh2, Nex1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 6 (NeuroD6) (Atonal protein homolog 2) (Helix-loop-helix protein mATH-2) (MATH2) (NEX-1 protein). | |||||
|
HEN1_MOUSE
|
||||||
| NC score | 0.162928 (rank : 32) | θ value | 1.38821 (rank : 64) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q02576 | Gene names | Nhlh1, Hen1 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
HEN2_HUMAN
|
||||||
| NC score | 0.161638 (rank : 33) | θ value | 1.06291 (rank : 58) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q02577 | Gene names | NHLH2, HEN2 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
HEN2_MOUSE
|
||||||
| NC score | 0.161482 (rank : 34) | θ value | 1.06291 (rank : 59) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q64221 | Gene names | Nhlh2, Hen2 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
HEN1_HUMAN
|
||||||
| NC score | 0.160601 (rank : 35) | θ value | 1.38821 (rank : 63) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q02575 | Gene names | NHLH1, HEN1 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
FIGLA_HUMAN
|
||||||
| NC score | 0.150950 (rank : 36) | θ value | 3.0926 (rank : 91) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q6QHK4 | Gene names | FIGLA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
HAND2_HUMAN
|
||||||
| NC score | 0.148136 (rank : 37) | θ value | 0.813845 (rank : 50) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P61296, O95300, O95301, P97833 | Gene names | HAND2, DHAND | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 2 (Deciduum, heart, autonomic nervous system and neural crest derivatives-expressed protein 2) (dHAND). | |||||
|
HAND2_MOUSE
|
||||||
| NC score | 0.148136 (rank : 38) | θ value | 0.813845 (rank : 51) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q61039, Q61100 | Gene names | Hand2, Dhand, Hed, Thing2 | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 2 (Deciduum, heart, autonomic nervous system and neural crest derivatives-expressed protein 2) (dHAND) (Helix-loop-helix transcription factor expressed in embryo and deciduum) (Thing-2). | |||||
|
LYL1_HUMAN
|
||||||
| NC score | 0.147927 (rank : 39) | θ value | 3.0926 (rank : 96) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P12980, O76102 | Gene names | LYL1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
TAL1_MOUSE
|
||||||
| NC score | 0.147244 (rank : 40) | θ value | 2.36792 (rank : 85) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P22091 | Gene names | Tal1, Scl, Tal-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein homolog (TAL-1 protein) (Stem cell protein). | |||||
|
TAL1_HUMAN
|
||||||
| NC score | 0.146657 (rank : 41) | θ value | 2.36792 (rank : 84) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P17542 | Gene names | TAL1, SCL, TCL5 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein (TAL-1 protein) (Stem cell protein) (T-cell leukemia/lymphoma-5 protein). | |||||
|
LYL1_MOUSE
|
||||||
| NC score | 0.141385 (rank : 42) | θ value | 5.27518 (rank : 111) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P27792 | Gene names | Lyl1, Lyl-1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
PTF1A_HUMAN
|
||||||
| NC score | 0.115596 (rank : 43) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q7RTS3, Q9HC25 | Gene names | PTF1A, PTF1P48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pancreas transcription factor 1 subunit alpha (Pancreas-specific transcription factor 1a) (bHLH transcription factor p48) (p48 DNA- binding subunit of transcription factor PTF1) (PTF1-p48). | |||||
|
PTF1A_MOUSE
|
||||||
| NC score | 0.114588 (rank : 44) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9QX98, Q9QYF5, Q9QYF6 | Gene names | Ptf1a, Ptf1p48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pancreas transcription factor 1 subunit alpha (Pancreas-specific transcription factor 1a) (bHLH transcription factor p48) (p48 DNA- binding subunit of transcription factor PTF1) (PTF1-p48). | |||||
|
MUSC_MOUSE
|
||||||
| NC score | 0.113191 (rank : 45) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O88940 | Gene names | Msc, Myor | |||
|
Domain Architecture |

|
|||||
| Description | Musculin (Myogenic repressor). | |||||
|
MUSC_HUMAN
|
||||||
| NC score | 0.112091 (rank : 46) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O60682, O75946, Q9BRE7 | Gene names | MSC, ABF1 | |||
|
Domain Architecture |

|
|||||
| Description | Musculin (Activated B-cell factor 1) (ABF-1). | |||||
|
BHLH4_MOUSE
|
||||||
| NC score | 0.108836 (rank : 47) | θ value | 8.99809 (rank : 132) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8BGW3, Q71MB7 | Gene names | Bhlhb4, H20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 4 (bHLHB4). | |||||
|
ASCL2_HUMAN
|
||||||
| NC score | 0.101284 (rank : 48) | θ value | 0.47712 (rank : 35) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q99929, Q9UM68 | Gene names | ASCL2 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 2 (Mash2) (HASH2). | |||||
|
NGN1_MOUSE
|
||||||
| NC score | 0.100478 (rank : 49) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P70660 | Gene names | Neurog1, Ath4c, Neurod3, Ngn, Ngn1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein) (Helix-loop-helix protein mATH-4C) (MATH4C). | |||||
|
NGN1_HUMAN
|
||||||
| NC score | 0.097705 (rank : 50) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q92886, Q96HE1 | Gene names | NEUROG1, NEUROD3, NGN, NGN1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein). | |||||
|
BHLH4_HUMAN
|
||||||
| NC score | 0.097290 (rank : 51) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NDY6 | Gene names | BHLHB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 4 (bHLHB4). | |||||
|
NGN3_MOUSE
|
||||||
| NC score | 0.095315 (rank : 52) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P70661 | Gene names | Neurog3, Ath4b, Atoh5, Ngn3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 3 (Atonal protein homolog 5) (Helix-loop-helix protein mATH-4B) (MATH4B). | |||||
|
NGN3_HUMAN
|
||||||
| NC score | 0.094587 (rank : 53) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y4Z2, Q9BY24 | Gene names | NEUROG3, NGN3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 3. | |||||
|
SCX_MOUSE
|
||||||
| NC score | 0.094419 (rank : 54) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q64124 | Gene names | Scx | |||
|
Domain Architecture |

|
|||||
| Description | Basic helix-loop-helix transcription factor scleraxis. | |||||
|
NGN2_HUMAN
|
||||||
| NC score | 0.093798 (rank : 55) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9H2A3, Q8N416 | Gene names | NEUROG2, NGN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurogenin 2. | |||||
|
NGN2_MOUSE
|
||||||
| NC score | 0.093568 (rank : 56) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P70447, P70237 | Gene names | Neurog2, Ath4a, Atoh4, Ngn2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 2 (Atonal protein homolog 4) (Helix-loop-helix protein mATH-4A) (MATH4A). | |||||
|
OLIG2_MOUSE
|
||||||
| NC score | 0.093398 (rank : 57) | θ value | 1.38821 (rank : 67) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9EQW6, Q9JKN4 | Gene names | Olig2 | |||
|
Domain Architecture |

|
|||||
| Description | Oligodendrocyte transcription factor 2 (Oligo2). | |||||
|
OLIG2_HUMAN
|
||||||
| NC score | 0.090717 (rank : 58) | θ value | 2.36792 (rank : 81) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q13516, Q86X04, Q9NZ14 | Gene names | OLIG2, BHLHB1, PRKCBP2, RACK17 | |||
|
Domain Architecture |

|
|||||
| Description | Oligodendrocyte transcription factor 2 (Oligo2) (Basic helix-loop- helix protein class B 1) (Protein kinase C-binding protein RACK17) (Protein kinase C-binding protein 2). | |||||
|
OLIG1_HUMAN
|
||||||
| NC score | 0.090019 (rank : 59) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8TAK6, Q7RTS0 | Gene names | OLIG1 | |||
|
Domain Architecture |

|
|||||
| Description | Oligodendrocyte transcription factor 1 (Oligo1). | |||||
|
TAL2_MOUSE
|
||||||
| NC score | 0.089888 (rank : 60) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q62282 | Gene names | Tal2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein homolog (TAL-2 protein). | |||||
|
TAL2_HUMAN
|
||||||
| NC score | 0.088770 (rank : 61) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q16559 | Gene names | TAL2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein (TAL-2 protein). | |||||
|
HAND1_MOUSE
|
||||||
| NC score | 0.084359 (rank : 62) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q64279, Q61099 | Gene names | Hand1, Ehand, Hxt, Thing1 | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND) (Helix-loop-helix transcription factor expressed in extraembryonic mesoderm and trophoblast) (Thing-1) (Th1). | |||||
|
ASCL3_MOUSE
|
||||||
| NC score | 0.083664 (rank : 63) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JJR7 | Gene names | Ascl3, Mash3, Sgn1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 3 (bHLH transcriptional regulator Sgn-1) (Mash- 3). | |||||
|
HAND1_HUMAN
|
||||||
| NC score | 0.083041 (rank : 64) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O96004 | Gene names | HAND1, EHAND | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND). | |||||
|
TCF15_MOUSE
|
||||||
| NC score | 0.082910 (rank : 65) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q60756, Q60788 | Gene names | Tcf15, Bhlhec2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 15 (bHLH-EC2 protein) (Paraxis) (Meso1). | |||||
|
ASCL3_HUMAN
|
||||||
| NC score | 0.080593 (rank : 66) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9NQ33, Q8WYQ6 | Gene names | ASCL3, HASH3, SGN1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 3 (bHLH transcriptional regulator Sgn-1). | |||||
|
ASCL2_MOUSE
|
||||||
| NC score | 0.080182 (rank : 67) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O35885, Q9WUJ7 | Gene names | Ascl2, Mash2 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 2 (Mash-2). | |||||
|
TCF15_HUMAN
|
||||||
| NC score | 0.078536 (rank : 68) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q12870, Q9NQQ1 | Gene names | TCF15, BHLHEC2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 15 (bHLH-EC2 protein) (Paraxis). | |||||
|
ASCL1_MOUSE
|
||||||
| NC score | 0.077582 (rank : 69) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q02067 | Gene names | Ascl1, Ash1, Mash-1, Mash1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 1 (Mash-1). | |||||
|
MYCN_HUMAN
|
||||||
| NC score | 0.077577 (rank : 70) | θ value | 0.62314 (rank : 40) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
ASCL1_HUMAN
|
||||||
| NC score | 0.076677 (rank : 71) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P50553, Q9BQ30 | Gene names | ASCL1, ASH1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 1 (HASH1). | |||||
|
HORN_HUMAN
|
||||||
| NC score | 0.074054 (rank : 72) | θ value | 1.81305 (rank : 74) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86YZ3 | Gene names | HRNR, S100A18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hornerin. | |||||
|
TCF21_HUMAN
|
||||||
| NC score | 0.070825 (rank : 73) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O43680, O43545, Q9BZ14 | Gene names | TCF21, POD1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 21 (Podocyte-expressed 1) (Pod-1) (Epicardin) (Capsulin). | |||||
|
MYCN_MOUSE
|
||||||
| NC score | 0.070801 (rank : 74) | θ value | 1.38821 (rank : 66) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
TCF21_MOUSE
|
||||||
| NC score | 0.070449 (rank : 75) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O35437, Q3U023 | Gene names | Tcf21, Pod1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 21 (Podocyte-expressed 1) (Pod-1) (Epicardin) (Capsulin). | |||||
|
CROP_MOUSE
|
||||||
| NC score | 0.068307 (rank : 76) | θ value | 0.125558 (rank : 22) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5SUF2, Q3U9D5, Q8BUJ5, Q921Z3, Q9CRS7, Q9CTY4 | Gene names | Crop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein. | |||||
|
OLIG3_MOUSE
|
||||||
| NC score | 0.068253 (rank : 77) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6PFG8, Q9EQW5 | Gene names | Olig3, Bhlhb7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oligodendrocyte transcription factor 3 (Oligo3) (Oligodendrocyte- specific bHLH transcription factor 3) (Basic helix-loop-helix domain- containing class B protein 7). | |||||
|
CROP_HUMAN
|
||||||
| NC score | 0.067393 (rank : 78) | θ value | 0.125558 (rank : 21) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95232, Q6PHR9, Q9NUY0, Q9P2S7 | Gene names | CROP, CREAP1, O48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein (cAMP regulatory element-associated protein 1) (CRE-associated protein 1) (CREAP-1) (Luc7A) (Okadaic acid-inducible phosphoprotein OA48-18). | |||||
|
OLIG3_HUMAN
|
||||||
| NC score | 0.067303 (rank : 79) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q7RTU3, Q8N8Q0 | Gene names | OLIG3, BHLHB7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oligodendrocyte transcription factor 3 (Oligo3) (Basic helix-loop- helix domain-containing class B protein 7). | |||||
|
OLIG1_MOUSE
|
||||||
| NC score | 0.066376 (rank : 80) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JKN5 | Gene names | Olig1 | |||
|
Domain Architecture |

|
|||||
| Description | Oligodendrocyte transcription factor 1 (Oligo1) (Oligodendrocyte- specific bHLH transcription factor 1). | |||||
|
TFAP4_HUMAN
|
||||||
| NC score | 0.065694 (rank : 81) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
HORN_MOUSE
|
||||||
| NC score | 0.064650 (rank : 82) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
CDSN_MOUSE
|
||||||
| NC score | 0.060383 (rank : 83) | θ value | 0.365318 (rank : 33) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7TPC1 | Gene names | Cdsn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor. | |||||
|
NUPL1_HUMAN
|
||||||
| NC score | 0.057447 (rank : 84) | θ value | 0.62314 (rank : 42) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BVL2, O43160 | Gene names | NUPL1, KIAA0410 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin p58/p45 (Nucleoporin-like 1). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.055694 (rank : 85) | θ value | 0.279714 (rank : 29) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
TB182_HUMAN
|
||||||
| NC score | 0.054393 (rank : 86) | θ value | 3.0926 (rank : 98) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9C0C2 | Gene names | TNKS1BP1, KIAA1741, TAB182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 182 kDa tankyrase 1-binding protein. | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.051054 (rank : 87) | θ value | 3.0926 (rank : 99) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.049594 (rank : 88) | θ value | 0.0431538 (rank : 12) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
FBX24_MOUSE
|
||||||
| NC score | 0.045140 (rank : 89) | θ value | 1.81305 (rank : 73) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D417 | Gene names | Fbxo24, Fbx24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 24. | |||||
|
MUC13_MOUSE
|
||||||
| NC score | 0.044838 (rank : 90) | θ value | 0.47712 (rank : 38) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
F10A5_HUMAN
|
||||||
| NC score | 0.044671 (rank : 91) | θ value | 0.279714 (rank : 31) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NFI4 | Gene names | FAM10A5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A5. | |||||
|
GPS2_HUMAN
|
||||||
| NC score | 0.044548 (rank : 92) | θ value | 0.813845 (rank : 49) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13227 | Gene names | GPS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein pathway suppressor 2 (Protein GPS2). | |||||
|
F10A1_HUMAN
|
||||||
| NC score | 0.044191 (rank : 93) | θ value | 0.279714 (rank : 30) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P50502, O14999 | Gene names | ST13, FAM10A1, HIP, SNC6 | |||
|
Domain Architecture |

|
|||||
| Description | Hsc70-interacting protein (Hip) (Suppression of tumorigenicity protein 13) (Putative tumor suppressor ST13) (Protein FAM10A1) (Progesterone receptor-associated p48 protein) (NY-REN-33 antigen). | |||||
|
BAZ2A_MOUSE
|
||||||
| NC score | 0.043850 (rank : 94) | θ value | 0.813845 (rank : 48) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
GLP_MOUSE
|
||||||
| NC score | 0.042722 (rank : 95) | θ value | 4.03905 (rank : 103) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P14220 | Gene names | Gypa | |||
|
Domain Architecture |

|
|||||
| Description | Glycophorin. | |||||
|
NCOA6_MOUSE
|
||||||
| NC score | 0.040680 (rank : 96) | θ value | 0.813845 (rank : 54) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.040579 (rank : 97) | θ value | 0.365318 (rank : 32) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.036396 (rank : 98) | θ value | 0.813845 (rank : 52) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
BCAR1_HUMAN
|
||||||
| NC score | 0.034695 (rank : 99) | θ value | 1.06291 (rank : 56) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
SRGP2_HUMAN
|
||||||
| NC score | 0.030618 (rank : 100) | θ value | 0.125558 (rank : 23) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O43295, Q8IX13, Q8IZV8 | Gene names | SRGAP3, ARHGAP14, KIAA0411, MEGAP, SRGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Mental disorder- associated GAP) (Rho-GTPase-activating protein 14). | |||||
|
K0157_MOUSE
|
||||||
| NC score | 0.029425 (rank : 101) | θ value | 1.81305 (rank : 75) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3TCJ1, Q3TB08, Q8K0R4 | Gene names | Kiaa0157 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0157. | |||||
|
ZN750_MOUSE
|
||||||
| NC score | 0.029284 (rank : 102) | θ value | 1.81305 (rank : 77) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BH05, Q66JP3, Q8C0L1 | Gene names | Znf750 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF750. | |||||
|
NU153_HUMAN
|
||||||
| NC score | 0.027062 (rank : 103) | θ value | 2.36792 (rank : 80) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
PYGO1_HUMAN
|
||||||
| NC score | 0.025106 (rank : 104) | θ value | 5.27518 (rank : 116) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y3Y4 | Gene names | PYGO1 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 1. | |||||
|
AHTF1_HUMAN
|
||||||
| NC score | 0.024851 (rank : 105) | θ value | 1.06291 (rank : 55) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
CJ022_HUMAN
|
||||||
| NC score | 0.024495 (rank : 106) | θ value | 6.88961 (rank : 118) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96SZ5 | Gene names | C10orf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf22. | |||||
|
RERE_MOUSE
|
||||||
| NC score | 0.024225 (rank : 107) | θ value | 6.88961 (rank : 123) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.023907 (rank : 108) | θ value | 3.0926 (rank : 89) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.023803 (rank : 109) | θ value | 3.0926 (rank : 88) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.023206 (rank : 110) | θ value | 4.03905 (rank : 106) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
SRGP2_MOUSE
|
||||||
| NC score | 0.023200 (rank : 111) | θ value | 0.62314 (rank : 43) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q812A2, Q80U09, Q8BKP4, Q8BLD0, Q925I2 | Gene names | Srgap3, Arhgap14, Kiaa0411, Srgap2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Rho-GTPase-activating protein 14). | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.023027 (rank : 112) | θ value | 8.99809 (rank : 148) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
PGCA_HUMAN
|
||||||
| NC score | 0.022972 (rank : 113) | θ value | 2.36792 (rank : 82) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
COHA1_HUMAN
|
||||||
| NC score | 0.022863 (rank : 114) | θ value | 6.88961 (rank : 119) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UMD9, Q02802, Q99018, Q9NQK9, Q9UC14 | Gene names | COL17A1, BP180, BPAG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
XKR5_HUMAN
|
||||||
| NC score | 0.022479 (rank : 115) | θ value | 3.0926 (rank : 100) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6UX68, Q5GH74 | Gene names | XKR5, XRG5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | XK-related protein 5. | |||||
|
IRS2_MOUSE
|
||||||
| NC score | 0.021502 (rank : 116) | θ value | 3.0926 (rank : 94) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P81122 | Gene names | Irs2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin receptor substrate 2 (IRS-2) (4PS). | |||||
|
HIRP3_HUMAN
|
||||||
| NC score | 0.020595 (rank : 117) | θ value | 3.0926 (rank : 93) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BW71, O75707, O75708 | Gene names | HIRIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIRA-interacting protein 3. | |||||
|
EYA1_MOUSE
|
||||||
| NC score | 0.020511 (rank : 118) | θ value | 1.81305 (rank : 71) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97767, O08818 | Gene names | Eya1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
RAD50_MOUSE
|
||||||
| NC score | 0.019978 (rank : 119) | θ value | 2.36792 (rank : 83) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
BRE1A_MOUSE
|
||||||
| NC score | 0.019347 (rank : 120) | θ value | 3.0926 (rank : 90) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1025 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5DTM8, Q3UT10, Q3V350, Q7TT11, Q8BKA8, Q8BKN8, Q8BUF7, Q8BVU4 | Gene names | Rnf20, Bre1a, Kiaa4116 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1A (EC 6.3.2.-) (BRE1-A) (RING finger protein 20). | |||||
|
AMOL1_MOUSE
|
||||||
| NC score | 0.019287 (rank : 121) | θ value | 0.813845 (rank : 47) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
CE152_HUMAN
|
||||||
| NC score | 0.019253 (rank : 122) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O94986, Q6NTA0 | Gene names | CEP152, KIAA0912 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 152 kDa (Cep152 protein). | |||||
|
K1267_MOUSE
|
||||||
| NC score | 0.018967 (rank : 123) | θ value | 5.27518 (rank : 110) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TG1, Q3TT88, Q3U5D8, Q3V3N3, Q7TMU3, Q80XP7, Q8R3L6, Q9D9G0 | Gene names | Kiaa1267 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1267. | |||||
|
REN3B_HUMAN
|
||||||
| NC score | 0.018942 (rank : 124) | θ value | 8.99809 (rank : 147) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BZI7, Q9H1J0 | Gene names | UPF3B, RENT3B, UPF3X | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 3B (Nonsense mRNA reducing factor 3B) (Up-frameshift suppressor 3 homolog B) (hUpf3B) (hUpf3p-X). | |||||
|
SMRC1_MOUSE
|
||||||
| NC score | 0.018804 (rank : 125) | θ value | 4.03905 (rank : 105) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P97496, Q7TS80, Q7TT29 | Gene names | Smarcc1, Baf155, Srg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155) (SWI3-related protein). | |||||
|
SC24C_HUMAN
|
||||||
| NC score | 0.018732 (rank : 126) | θ value | 6.88961 (rank : 126) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
BRE1A_HUMAN
|
||||||
| NC score | 0.018453 (rank : 127) | θ value | 1.81305 (rank : 70) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1071 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5VTR2, Q69YL5, Q6P527, Q8N3J4, Q96JD3, Q9H9Y7, Q9HA51, Q9NUR4, Q9NWQ3, Q9NX83 | Gene names | RNF20, BRE1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1A (EC 6.3.2.-) (BRE1-A) (hBRE1) (RING finger protein 20). | |||||
|
K1267_HUMAN
|
||||||
| NC score | 0.018019 (rank : 128) | θ value | 6.88961 (rank : 122) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z3B3, Q6AW85, Q8IYH1, Q9NTE7, Q9UFT0, Q9ULF3 | Gene names | KIAA1267 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1267. | |||||
|
NOP14_HUMAN
|
||||||
| NC score | 0.017392 (rank : 129) | θ value | 5.27518 (rank : 114) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.017378 (rank : 130) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.016747 (rank : 131) | θ value | 5.27518 (rank : 112) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
INVS_HUMAN
|
||||||
| NC score | 0.015568 (rank : 132) | θ value | 1.38821 (rank : 65) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y283, Q5W0T6, Q8IVX8, Q9BRB9, Q9Y488, Q9Y498 | Gene names | INVS, INV, NPHP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning homolog) (Nephrocystin-2). | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.015568 (rank : 133) | θ value | 0.62314 (rank : 39) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
DHX8_HUMAN
|
||||||
| NC score | 0.015234 (rank : 134) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14562 | Gene names | DHX8, DDX8 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DHX8 (EC 3.6.1.-) (DEAH box protein 8) (RNA helicase HRH1). | |||||
|
COIA1_HUMAN
|
||||||
| NC score | 0.015002 (rank : 135) | θ value | 5.27518 (rank : 109) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P39060, Q9UK38, Q9Y6Q7, Q9Y6Q8 | Gene names | COL18A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
FIBA_HUMAN
|
||||||
| NC score | 0.014049 (rank : 136) | θ value | 0.47712 (rank : 37) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P02671, Q9BX62, Q9UCH2 | Gene names | FGA | |||
|
Domain Architecture |

|
|||||
| Description | Fibrinogen alpha chain precursor [Contains: Fibrinopeptide A]. | |||||
|
TP53B_MOUSE
|
||||||
| NC score | 0.014010 (rank : 137) | θ value | 8.99809 (rank : 153) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
SFR12_MOUSE
|
||||||
| NC score | 0.013827 (rank : 138) | θ value | 8.99809 (rank : 149) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
SMRC1_HUMAN
|
||||||
| NC score | 0.013679 (rank : 139) | θ value | 6.88961 (rank : 127) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92922, Q6P172, Q8IWH2 | Gene names | SMARCC1, BAF155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155). | |||||
|
DCDC2_MOUSE
|
||||||
| NC score | 0.013484 (rank : 140) | θ value | 8.99809 (rank : 135) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
PPIG_HUMAN
|
||||||
| NC score | 0.013441 (rank : 141) | θ value | 4.03905 (rank : 104) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
SI1L1_MOUSE
|
||||||
| NC score | 0.013001 (rank : 142) | θ value | 1.38821 (rank : 68) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C0T5, Q6PDI8, Q80U02, Q8C026 | Gene names | Sipa1l1, Kiaa0440 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 1. | |||||
|
KIF5A_HUMAN
|
||||||
| NC score | 0.012875 (rank : 143) | θ value | 1.06291 (rank : 60) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q12840, Q4LE26 | Gene names | KIF5A, NKHC1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
NCOA3_HUMAN
|
||||||
| NC score | 0.012378 (rank : 144) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
TTLL4_MOUSE
|
||||||
| NC score | 0.012067 (rank : 145) | θ value | 6.88961 (rank : 128) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80UG8, Q5U4C4 | Gene names | Ttll4, Kiaa0173 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 4. | |||||
|
STX4_HUMAN
|
||||||
| NC score | 0.011879 (rank : 146) | θ value | 5.27518 (rank : 117) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q12846, Q15525 | Gene names | STX4, STX4A | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-4 (NY-REN-31 antigen). | |||||
|
KIF5A_MOUSE
|
||||||
| NC score | 0.011828 (rank : 147) | θ value | 3.0926 (rank : 95) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P33175, Q5DTP1, Q6PDY7, Q9Z2F9 | Gene names | Kif5a, Kiaa4086, Kif5, Nkhc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
TRIM2_MOUSE
|
||||||
| NC score | 0.011824 (rank : 148) | θ value | 2.36792 (rank : 87) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ESN6, Q3UHP5, Q8C981 | Gene names | Trim2, Narf | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 2 (Neural activity-related RING finger protein). | |||||
|
TRIM2_HUMAN
|
||||||
| NC score | 0.011782 (rank : 149) | θ value | 2.36792 (rank : 86) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9C040, O60272, Q9BSI9, Q9UFZ1 | Gene names | TRIM2, KIAA0517, RNF86 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 2 (RING finger protein 86). | |||||
|
DUS16_HUMAN
|
||||||
| NC score | 0.011650 (rank : 150) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BY84, Q9C0G3 | Gene names | DUSP16, KIAA1700, MKP7 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein phosphatase 16 (EC 3.1.3.48) (EC 3.1.3.16) (Mitogen-activated protein kinase phosphatase 7) (MAP kinase phosphatase 7) (MKP-7). | |||||
|
NUPL_MOUSE
|
||||||
| NC score | 0.011529 (rank : 151) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K2K6, O70448, Q8BQL5, Q8CDK9 | Gene names | Hrb, Rip | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoporin-like protein RIP (HIV-1 Rev-binding protein homolog). | |||||
|
RFX1_MOUSE
|
||||||
| NC score | 0.011508 (rank : 152) | θ value | 6.88961 (rank : 124) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P48377 | Gene names | Rfx1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein RFX1. | |||||
|
CLCN1_HUMAN
|
||||||
| NC score | 0.010892 (rank : 153) | θ value | 2.36792 (rank : 78) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35523 | Gene names | CLCN1, CLC1 | |||
|
Domain Architecture |

|
|||||
| Description | Chloride channel protein, skeletal muscle (Chloride channel protein 1) (ClC-1). | |||||
|
SAKS1_MOUSE
|
||||||
| NC score | 0.010099 (rank : 154) | θ value | 6.88961 (rank : 125) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q922Y1, Q3UCP8 | Gene names | Saks1, D19Ertd721e | |||
|
Domain Architecture |

|
|||||
| Description | SAPK substrate protein 1 (UBA/UBX 33.3 kDa protein) (mY33K) (Protein 2B28). | |||||
|
ENAH_HUMAN
|
||||||
| NC score | 0.009875 (rank : 155) | θ value | 6.88961 (rank : 120) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N8S7, Q502W5, Q5T5M7, Q5VTQ9, Q5VTR0, Q9NVF3, Q9UFB8 | Gene names | ENAH, MENA | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog. | |||||
|
ASH2L_MOUSE
|
||||||
| NC score | 0.009125 (rank : 156) | θ value | 8.99809 (rank : 131) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91X20, Q3UIF9, Q9Z2X4 | Gene names | Ash2l | |||
|
Domain Architecture |

|
|||||
| Description | Set1/Ash2 histone methyltransferase complex subunit ASH2 (ASH2-like protein). | |||||
|
DLG5_HUMAN
|
||||||
| NC score | 0.008680 (rank : 157) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8TDM6, Q5T1H7, Q5T1H8, Q6DKG3, Q86WC0, Q8TDM7, Q9UE73, Q9Y4E3 | Gene names | DLG5, KIAA0583, PDLG | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 5 (Placenta and prostate DLG) (Discs large protein P-dlg). | |||||
|
EZRI_MOUSE
|
||||||
| NC score | 0.008242 (rank : 158) | θ value | 1.81305 (rank : 72) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P26040, Q80ZT8, Q9DCI1 | Gene names | Vil2 | |||
|
Domain Architecture |

|
|||||
| Description | Ezrin (p81) (Cytovillin) (Villin-2). | |||||
|
ARHGA_MOUSE
|
||||||
| NC score | 0.007331 (rank : 159) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C033, Q5DU38, Q80VH8, Q8BW76, Q922S7 | Gene names | Arhgef10, Kiaa0294 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
ZBTB6_HUMAN
|
||||||
| NC score | 0.007318 (rank : 160) | θ value | 1.06291 (rank : 61) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 805 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q15916 | Gene names | ZBTB6, ZID, ZNF482 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 6 (Zinc finger protein 482) (Zinc finger protein with interaction domain). | |||||
|
ZBTB6_MOUSE
|
||||||
| NC score | 0.006680 (rank : 161) | θ value | 1.81305 (rank : 76) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K088, Q8BLM4 | Gene names | Zbtb6, Zfp482, Znf482 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 6 (Zinc finger protein 482). | |||||
|
CDKL5_HUMAN
|
||||||
| NC score | 0.006423 (rank : 162) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O76039, Q14198, Q5H985, Q9UJL6 | Gene names | CDKL5, STK9 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase-like 5 (EC 2.7.11.22) (Serine/threonine- protein kinase 9). | |||||
|
ST18_HUMAN
|
||||||
| NC score | 0.006018 (rank : 163) | θ value | 8.99809 (rank : 152) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O60284 | Gene names | ST18, KIAA0535, ZNF387 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18 (Zinc finger protein 387). | |||||
|
FXR2_MOUSE
|
||||||
| NC score | 0.005947 (rank : 164) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WVR4, Q9WVR5 | Gene names | Fxr2, Fxr2h | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
ANS4B_MOUSE
|
||||||
| NC score | 0.005894 (rank : 165) | θ value | 8.99809 (rank : 130) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K3X6, Q9D8A5 | Gene names | Anks4b, Harp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 4B (Harmonin- interacting ankyrin repeat-containing protein) (Harp). | |||||
|
SSH1_HUMAN
|
||||||
| NC score | 0.005847 (rank : 166) | θ value | 8.99809 (rank : 151) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WYL5, Q6P6C0, Q8N9A7, Q8WYL3, Q8WYL4, Q9P2P8 | Gene names | SSH1, KIAA1298, SSH1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 1 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-1L) (hSSH-1L). | |||||
|
WT1_HUMAN
|
||||||
| NC score | 0.005330 (rank : 167) | θ value | 1.38821 (rank : 69) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P19544, Q15881, Q16256, Q16575, Q8IYZ5 | Gene names | WT1 | |||
|
Domain Architecture |

|
|||||
| Description | Wilms' tumor protein (WT33). | |||||
|
SOX9_MOUSE
|
||||||
| NC score | 0.005318 (rank : 168) | θ value | 8.99809 (rank : 150) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q04887, Q8C7L2, Q91ZK2, Q99KQ0 | Gene names | Sox9, Sox-9 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-9. | |||||
|
KCNH1_MOUSE
|
||||||
| NC score | 0.005214 (rank : 169) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60603 | Gene names | Kcnh1, Eag | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (EAG1) (m-eag). | |||||
|
KCNH1_HUMAN
|
||||||
| NC score | 0.005207 (rank : 170) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95259, O76035 | Gene names | KCNH1, EAG, EAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 1 (Voltage-gated potassium channel subunit Kv10.1) (Ether-a-go-go potassium channel 1) (hEAG1) (h-eag). | |||||
|
GLI1_MOUSE
|
||||||
| NC score | 0.004359 (rank : 171) | θ value | 3.0926 (rank : 92) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P47806, Q9QYK1 | Gene names | Gli1, Gli | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI1 (Glioma-associated oncogene homolog). | |||||
|
ZKSC1_HUMAN
|
||||||
| NC score | 0.003271 (rank : 172) | θ value | 0.62314 (rank : 46) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P17029, P52745, Q8TBW5, Q8TEK7 | Gene names | ZKSCAN1, KOX18, ZNF139, ZNF36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1 (Zinc finger protein 36) (Zinc finger protein KOX18). | |||||
|
WT1_MOUSE
|
||||||
| NC score | 0.003023 (rank : 173) | θ value | 4.03905 (rank : 107) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P22561 | Gene names | Wt1, Wt-1 | |||
|
Domain Architecture |

|
|||||
| Description | Wilms' tumor protein homolog. | |||||
|
PTPRA_HUMAN
|
||||||
| NC score | 0.003017 (rank : 174) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P18433, Q14513, Q7Z2I2, Q96TD9 | Gene names | PTPRA, PTPA | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase alpha precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase alpha) (R-PTP-alpha). | |||||
|
MTR1L_HUMAN
|
||||||
| NC score | 0.002110 (rank : 175) | θ value | 2.36792 (rank : 79) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1180 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13585 | Gene names | GPR50 | |||
|
Domain Architecture |

|
|||||
| Description | Melatonin-related receptor (G protein-coupled receptor 50) (H9). | |||||
|
PRD16_HUMAN
|
||||||
| NC score | 0.001240 (rank : 176) | θ value | 5.27518 (rank : 115) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 832 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9HAZ2, Q8WYJ9, Q9C0I8 | Gene names | PRDM16, KIAA1675, MEL1, PFM13 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 16 (PR domain-containing protein 16) (Transcription factor MEL1). | |||||
|
HXA1_HUMAN
|
||||||
| NC score | 0.001027 (rank : 177) | θ value | 6.88961 (rank : 121) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49639, O43363 | Gene names | HOXA1, HOX1F | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A1 (Hox-1F). | |||||
|
ZBT38_HUMAN
|
||||||
| NC score | 0.000861 (rank : 178) | θ value | 6.88961 (rank : 129) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 885 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NAP3 | Gene names | ZBTB38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 38. | |||||
|
SPEG_HUMAN
|
||||||
| NC score | -0.000149 (rank : 179) | θ value | 3.0926 (rank : 97) | |||
| Query Neighborhood Hits | 153 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||