Please be patient as the page loads
|
WBP4_MOUSE
|
||||||
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
WBP4_MOUSE
|
||||||
| θ value | 2.83983e-179 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 100 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
WBP4_HUMAN
|
||||||
| θ value | 2.52267e-127 (rank : 2) | NC score | 0.942255 (rank : 2) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75554 | Gene names | WBP4, FBP21, FNBP21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
PRP40_HUMAN
|
||||||
| θ value | 5.08577e-11 (rank : 3) | NC score | 0.421886 (rank : 4) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
PRP40_MOUSE
|
||||||
| θ value | 5.08577e-11 (rank : 4) | NC score | 0.422915 (rank : 3) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
PKCB1_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 5) | NC score | 0.104693 (rank : 27) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
TCRG1_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 6) | NC score | 0.184423 (rank : 8) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
TCRG1_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 7) | NC score | 0.183994 (rank : 9) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
K1688_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 8) | NC score | 0.128420 (rank : 15) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
SETD2_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 9) | NC score | 0.133392 (rank : 12) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
RU1C_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 10) | NC score | 0.219364 (rank : 5) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P09234 | Gene names | SNRPC | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
RU1C_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 11) | NC score | 0.218461 (rank : 6) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62241 | Gene names | Snrp1c, Snrpc | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
PIN1_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 12) | NC score | 0.185507 (rank : 7) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QUR7, Q543B3 | Gene names | Pin1 | |||
|
Domain Architecture |
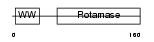
|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
IQGA1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 13) | NC score | 0.130393 (rank : 14) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P46940 | Gene names | IQGAP1, KIAA0051 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1 (p195). | |||||
|
TRIPC_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 14) | NC score | 0.103513 (rank : 28) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 15) | NC score | 0.100104 (rank : 33) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
POT15_HUMAN
|
||||||
| θ value | 0.125558 (rank : 16) | NC score | 0.045243 (rank : 89) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6S5H4, Q6NXN7, Q6S5H7 | Gene names | POTE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 15. | |||||
|
TEX14_MOUSE
|
||||||
| θ value | 0.125558 (rank : 17) | NC score | 0.027545 (rank : 108) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1333 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q7M6U3, Q3UT36, Q5NC10, Q5NC11, Q8CGK1, Q8CGK2, Q99MV8 | Gene names | Tex14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
RBP2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 18) | NC score | 0.030233 (rank : 105) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ERU9, Q61992, Q8C9K9 | Gene names | Ranbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2 (RanBP2). | |||||
|
IQGA1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 19) | NC score | 0.125201 (rank : 16) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JKF1 | Gene names | Iqgap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1. | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.065007 (rank : 52) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
ANR21_HUMAN
|
||||||
| θ value | 0.279714 (rank : 21) | NC score | 0.042667 (rank : 92) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q86YR6 | Gene names | ANKRD21, POTE | |||
|
Domain Architecture |
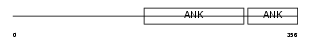
|
|||||
| Description | Ankyrin repeat domain-containing protein 21 (Protein POTE) (Prostate, ovary, testis-expressed protein). | |||||
|
DGC14_MOUSE
|
||||||
| θ value | 0.279714 (rank : 22) | NC score | 0.091523 (rank : 39) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O70279, Q91YX1 | Gene names | Dgcr14, Dgsi, Es2, Es2el | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DGCR14 protein (DiGeorge syndrome critical region 14 homolog) (ES2 protein) (Expressed sequence 2 embryonic lethal). | |||||
|
K1688_MOUSE
|
||||||
| θ value | 0.279714 (rank : 23) | NC score | 0.089112 (rank : 40) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P59281, Q69ZD4, Q6P9R6 | Gene names | Kiaa1688, D15Wsu169e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1688. | |||||
|
PIN1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 24) | NC score | 0.170952 (rank : 10) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13526 | Gene names | PIN1 | |||
|
Domain Architecture |
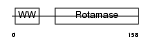
|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 0.279714 (rank : 25) | NC score | 0.016824 (rank : 121) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
WWOX_HUMAN
|
||||||
| θ value | 0.279714 (rank : 26) | NC score | 0.102602 (rank : 29) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NZC7, Q5MYT5, Q96KM3, Q96RF2, Q9BTT8, Q9NPC9, Q9NRF4, Q9NRF5, Q9NRF6, Q9NRK1, Q9NZC5 | Gene names | WWOX, FOR, WOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-) (Fragile site FRA16D oxidoreductase). | |||||
|
WWOX_MOUSE
|
||||||
| θ value | 0.279714 (rank : 27) | NC score | 0.101674 (rank : 30) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q91WL8, Q8C8J6, Q920Y2, Q9D2B3, Q9D339, Q9JLF5 | Gene names | Wwox, Wox1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-). | |||||
|
IQGA2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 28) | NC score | 0.116794 (rank : 24) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
ITCH_HUMAN
|
||||||
| θ value | 0.47712 (rank : 29) | NC score | 0.116914 (rank : 23) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
ITCH_MOUSE
|
||||||
| θ value | 0.47712 (rank : 30) | NC score | 0.117547 (rank : 22) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8C863, O54971 | Gene names | Itch | |||
|
Domain Architecture |

|
|||||
| Description | Itchy E3 ubiquitin protein ligase (EC 6.3.2.-). | |||||
|
NED4L_MOUSE
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.098798 (rank : 34) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8CFI0, Q8BRT9, Q8BS42, Q99PK2 | Gene names | Nedd4l, Kiaa0439, Nedd4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
OXR1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 32) | NC score | 0.067165 (rank : 47) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
CALD1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 33) | NC score | 0.050999 (rank : 81) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
NSD1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 34) | NC score | 0.060220 (rank : 58) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
WWP2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 35) | NC score | 0.118030 (rank : 21) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
WWP2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 36) | NC score | 0.118548 (rank : 20) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9DBH0, Q8BTG4, Q923F6 | Gene names | Wwp2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 37) | NC score | 0.094164 (rank : 36) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.813845 (rank : 38) | NC score | 0.040910 (rank : 94) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SPTN5_HUMAN
|
||||||
| θ value | 0.813845 (rank : 39) | NC score | 0.019667 (rank : 117) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NRC6 | Gene names | SPTBN5, SPTBN4 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 4 (Spectrin, non-erythroid beta chain 4) (Beta-V spectrin) (BSPECV). | |||||
|
IQGA3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.123995 (rank : 17) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86VI3 | Gene names | IQGAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP3. | |||||
|
JHD2C_HUMAN
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.030526 (rank : 103) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15652, Q5SQZ8, Q5SQZ9, Q5SR00, Q7Z3E7, Q8N3U0, Q96KB9, Q9P2G7 | Gene names | JMJD1C, JHDM2C, KIAA1380, TRIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C) (Thyroid receptor-interacting protein 8) (TRIP-8). | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.072749 (rank : 45) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
POTE8_HUMAN
|
||||||
| θ value | 1.06291 (rank : 43) | NC score | 0.036018 (rank : 97) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6S8J7, Q6S8J6 | Gene names | POTE8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 8. | |||||
|
PSIP1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 44) | NC score | 0.065446 (rank : 51) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q99JF8, Q3TEJ7, Q3TTD7, Q3UTA1, Q80WQ7, Q99JF7, Q99LR4, Q9CT03 | Gene names | Psip1, Ledgf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (mLEDGF). | |||||
|
RHG12_HUMAN
|
||||||
| θ value | 1.06291 (rank : 45) | NC score | 0.084388 (rank : 42) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8IWW6, Q86UB3, Q8IWW7, Q8N3L1, Q9NT76 | Gene names | ARHGAP12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
NED4L_HUMAN
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.100613 (rank : 31) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96PU5, O43165, Q7Z5F1, Q7Z5F2, Q7Z5N3, Q8N5A7, Q8WUU9, Q9BW58, Q9H2W4, Q9NT88 | Gene names | NEDD4L, KIAA0439, NEDL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
RHG12_MOUSE
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.094092 (rank : 37) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 1.38821 (rank : 48) | NC score | 0.067163 (rank : 48) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
2A5D_HUMAN
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.020485 (rank : 115) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14738, O00494, O00696, Q15171 | Gene names | PPP2R5D | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit delta isoform (PP2A, B subunit, B' delta isoform) (PP2A, B subunit, B56 delta isoform) (PP2A, B subunit, PR61 delta isoform) (PP2A, B subunit, R5 delta isoform). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 50) | NC score | 0.052218 (rank : 78) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
YAP1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 51) | NC score | 0.112659 (rank : 26) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
AFF2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.044678 (rank : 90) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
MPP8_HUMAN
|
||||||
| θ value | 2.36792 (rank : 53) | NC score | 0.042687 (rank : 91) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 54) | NC score | 0.064078 (rank : 53) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
SMUF2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 55) | NC score | 0.094245 (rank : 35) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9HAU4, Q9H260 | Gene names | SMURF2 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 2 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF2) (Smad-specific E3 ubiquitin ligase 2) (hSMURF2). | |||||
|
STXB4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 56) | NC score | 0.048806 (rank : 86) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 629 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WV89, Q5SUA6, Q5SUA8, Q5SUA9, Q8CFL1 | Gene names | Stxbp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
DGC14_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.063847 (rank : 55) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96DF8, Q9BTZ4 | Gene names | DGCR14, DGSI, ES2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DGCR14 protein (DiGeorge syndrome critical region 14) (ES2 protein). | |||||
|
HXC10_MOUSE
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.009032 (rank : 129) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P31257, P31312, Q7TMT7 | Gene names | Hoxc10, Hox-3.6, Hoxc-10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein Hox-C10 (Hox-3.6). | |||||
|
MNDA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.023686 (rank : 110) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P41218 | Gene names | MNDA | |||
|
Domain Architecture |
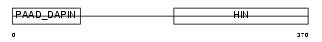
|
|||||
| Description | Myeloid cell nuclear differentiation antigen. | |||||
|
NALP5_MOUSE
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.039321 (rank : 96) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
PINL_HUMAN
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.149324 (rank : 11) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15428 | Gene names | PIN1L | |||
|
Domain Architecture |
|
|||||
| Description | PIN1-like protein. | |||||
|
RHG09_HUMAN
|
||||||
| θ value | 3.0926 (rank : 62) | NC score | 0.040827 (rank : 95) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BRR9, Q8TCJ3, Q8WYR0, Q96EZ2, Q96S74 | Gene names | ARHGAP9 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 9. | |||||
|
ANKZ1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.031286 (rank : 102) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80UU1, Q9CZF5 | Gene names | Ankzf1, D1Ertd161e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and zinc finger domain-containing protein 1. | |||||
|
ARI4A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.048573 (rank : 87) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
PRG4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.034522 (rank : 98) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SEC63_HUMAN
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.062714 (rank : 57) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
ZC11A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.032516 (rank : 99) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
ACINU_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.053273 (rank : 76) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
AP3B1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.031952 (rank : 101) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
CAC1F_MOUSE
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.008314 (rank : 130) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JIS7 | Gene names | Cacna1f | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.018590 (rank : 119) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
GAS7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.122599 (rank : 18) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O60861, O43144 | Gene names | GAS7, KIAA0394 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
LBN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.017518 (rank : 120) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86UK5, Q86YT3, Q86YT4, Q8NG49 | Gene names | EVC2, LBN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Limbin. | |||||
|
NNP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.025604 (rank : 109) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P56183, O35712, Q9ERE1, Q9JI07, Q9JK67, Q9JKU2 | Gene names | Nnp1 | |||
|
Domain Architecture |
|
|||||
| Description | NNP-1 protein (Novel nuclear protein 1) (Nop52). | |||||
|
SMRC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.028450 (rank : 106) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
WWP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.119079 (rank : 19) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
ARMC4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.021419 (rank : 113) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
DCDC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.022772 (rank : 112) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
DDX21_MOUSE
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.015610 (rank : 123) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
GABPA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.009293 (rank : 127) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q06546, Q12939 | Gene names | GABPA, E4TF1A | |||
|
Domain Architecture |
|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha) (Transcription factor E4TF1-60) (Nuclear respiratory factor 2 subunit alpha). | |||||
|
GAS7_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.130706 (rank : 13) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q60780, Q9QY25, Q9QY26, Q9QY34 | Gene names | Gas7 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
NEDD4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.100148 (rank : 32) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
NFM_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.030465 (rank : 104) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
RNF25_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.020118 (rank : 116) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96BH1, Q9H874 | Gene names | RNF25 | |||
|
Domain Architecture |
|
|||||
| Description | RING finger protein 25 (EC 6.3.2.-). | |||||
|
TCF8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.005367 (rank : 131) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
WASF2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.015155 (rank : 124) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BH43 | Gene names | Wasf2, Wave2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2). | |||||
|
WWP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.114104 (rank : 25) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.010143 (rank : 126) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CENPB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.032009 (rank : 100) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P07199 | Gene names | CENPB | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
DC1I2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.009157 (rank : 128) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13409, Q96NG7, Q96S87, Q9BXZ5, Q9NT58 | Gene names | DYNC1I2, DNCI2, DNCIC2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 2 (Dynein intermediate chain 2, cytosolic) (DH IC-2) (Cytoplasmic dynein intermediate chain 2). | |||||
|
ENAH_MOUSE
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.019121 (rank : 118) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
ENAM_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.023357 (rank : 111) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O55196 | Gene names | Enam | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enamelin precursor. | |||||
|
K1024_MOUSE
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.016345 (rank : 122) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K3V7, Q69ZS9 | Gene names | Kiaa1024 | |||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein KIAA1024 (Protein DD1). | |||||
|
NSD1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.048110 (rank : 88) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
PEG3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.001760 (rank : 132) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
SCYBG_MOUSE
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.020536 (rank : 114) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BSU2, Q8VE25, Q9EPB3 | Gene names | Cxcl16, Srpsox | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small inducible cytokine B16 precursor (Transmembrane chemokine CXCL16) (SR-PSOX) (Scavenger receptor for phosphatidylserine and oxidized low density lipoprotein). | |||||
|
SMRC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.027994 (rank : 107) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
STXB4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.041019 (rank : 93) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 616 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6ZWJ1, Q8IVZ5 | Gene names | STXBP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
TACC3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.014193 (rank : 125) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y6A5, Q9UMQ1 | Gene names | TACC3, ERIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ERIC-1). | |||||
|
YAP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.085363 (rank : 41) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P46937 | Gene names | YAP1, YAP65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
BAG3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.051834 (rank : 80) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
CYLC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.053767 (rank : 74) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
DEK_MOUSE
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.050642 (rank : 84) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
HECD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.057813 (rank : 65) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5U5R9, Q5VZ97, Q5VZ99, Q8TCP5 | Gene names | HECTD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
HECD2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.058350 (rank : 62) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CDU6, Q8CBQ9 | Gene names | Hectd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
HUWE1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.065785 (rank : 50) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
HUWE1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.066293 (rank : 49) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
K0317_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.059930 (rank : 60) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O15033, Q7LDY1, Q8IYY9 | Gene names | KIAA0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
K0317_MOUSE
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.060116 (rank : 59) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CHG5, Q6P9Q1, Q80YC4, Q8C5W5 | Gene names | Kiaa0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
LEO1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.053675 (rank : 75) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.067220 (rank : 46) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
NEDD4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.092204 (rank : 38) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P46934 | Gene names | NEDD4, KIAA0093 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-). | |||||
|
NKTR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.057447 (rank : 68) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.050731 (rank : 82) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
RSF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.052093 (rank : 79) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
SAV1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.063903 (rank : 54) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9H4B6, Q6IA58, Q9H949, Q9HAK9 | Gene names | SAV1, WW45 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (hWW45). | |||||
|
SAV1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.062968 (rank : 56) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8VEB2, Q9D9N9, Q9ER46 | Gene names | Sav1, Ww45, Wwp3 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (mWW45). | |||||
|
SFRIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.053216 (rank : 77) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
SMUF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.075982 (rank : 44) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9HCE7, O75853, Q9UJT8 | Gene names | SMURF1, KIAA1625 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1) (hSMURF1). | |||||
|
SMUF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.078819 (rank : 43) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9CUN6, Q3U412, Q8K300 | Gene names | Smurf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1). | |||||
|
THOC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.050241 (rank : 85) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.050670 (rank : 83) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
TXND2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.054844 (rank : 73) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
UBE3A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.057909 (rank : 63) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q05086, P78355, Q93066, Q9UEP4, Q9UEP5, Q9UEP6, Q9UEP7, Q9UEP8, Q9UEP9 | Gene names | UBE3A, E6AP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (E6AP ubiquitin-protein ligase) (Oncogenic protein-associated protein E6-AP) (Human papillomavirus E6-associated protein) (NY-REN-54 antigen). | |||||
|
UBE3A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.056949 (rank : 69) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O08759, P97482 | Gene names | Ube3a | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (Oncogenic protein- associated protein E6-AP). | |||||
|
UBE3C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.056410 (rank : 71) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15386, Q8TC15, Q96CR4, Q9UDU3 | Gene names | UBE3C, KIAA0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
UBE3C_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.056519 (rank : 70) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80U95, Q8BQZ6, Q8C7W6, Q8CDJ1, Q8VDL5 | Gene names | Ube3c, Kiaa0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
WWC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.057659 (rank : 66) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8IX03, O94946, Q6MZX4, Q6Y2F8, Q7Z4G8, Q8WVM4, Q9BT29 | Gene names | WWC1, KIAA0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA) (HBeAg-binding protein 3). | |||||
|
WWC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.057486 (rank : 67) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5SXA9, Q571D0, Q8K1Y3, Q8VD17, Q922W3 | Gene names | Wwc1, Kiaa0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA). | |||||
|
WWTR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.057840 (rank : 64) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9GZV5, Q8N3P2, Q9Y3W6 | Gene names | WWTR1, TAZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
WWTR1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.056232 (rank : 72) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9EPK5, Q99KI4 | Gene names | Wwtr1, Taz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
YTDC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.058506 (rank : 61) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
WBP4_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 2.83983e-179 (rank : 1) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 100 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
WBP4_HUMAN
|
||||||
| NC score | 0.942255 (rank : 2) | θ value | 2.52267e-127 (rank : 2) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75554 | Gene names | WBP4, FBP21, FNBP21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
PRP40_MOUSE
|
||||||
| NC score | 0.422915 (rank : 3) | θ value | 5.08577e-11 (rank : 4) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
PRP40_HUMAN
|
||||||
| NC score | 0.421886 (rank : 4) | θ value | 5.08577e-11 (rank : 3) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
RU1C_HUMAN
|
||||||
| NC score | 0.219364 (rank : 5) | θ value | 0.0330416 (rank : 10) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P09234 | Gene names | SNRPC | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
RU1C_MOUSE
|
||||||
| NC score | 0.218461 (rank : 6) | θ value | 0.0330416 (rank : 11) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62241 | Gene names | Snrp1c, Snrpc | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
PIN1_MOUSE
|
||||||
| NC score | 0.185507 (rank : 7) | θ value | 0.0431538 (rank : 12) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QUR7, Q543B3 | Gene names | Pin1 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
TCRG1_HUMAN
|
||||||
| NC score | 0.184423 (rank : 8) | θ value | 0.00509761 (rank : 6) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
TCRG1_MOUSE
|
||||||
| NC score | 0.183994 (rank : 9) | θ value | 0.00509761 (rank : 7) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
PIN1_HUMAN
|
||||||
| NC score | 0.170952 (rank : 10) | θ value | 0.279714 (rank : 24) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13526 | Gene names | PIN1 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
PINL_HUMAN
|
||||||
| NC score | 0.149324 (rank : 11) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15428 | Gene names | PIN1L | |||
|
Domain Architecture |

|
|||||
| Description | PIN1-like protein. | |||||
|
SETD2_HUMAN
|
||||||
| NC score | 0.133392 (rank : 12) | θ value | 0.0148317 (rank : 9) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
GAS7_MOUSE
|
||||||
| NC score | 0.130706 (rank : 13) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q60780, Q9QY25, Q9QY26, Q9QY34 | Gene names | Gas7 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
IQGA1_HUMAN
|
||||||
| NC score | 0.130393 (rank : 14) | θ value | 0.0736092 (rank : 13) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P46940 | Gene names | IQGAP1, KIAA0051 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1 (p195). | |||||
|
K1688_HUMAN
|
||||||
| NC score | 0.128420 (rank : 15) | θ value | 0.0148317 (rank : 8) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
IQGA1_MOUSE
|
||||||
| NC score | 0.125201 (rank : 16) | θ value | 0.21417 (rank : 19) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JKF1 | Gene names | Iqgap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1. | |||||
|
IQGA3_HUMAN
|
||||||
| NC score | 0.123995 (rank : 17) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86VI3 | Gene names | IQGAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP3. | |||||
|
GAS7_HUMAN
|
||||||
| NC score | 0.122599 (rank : 18) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O60861, O43144 | Gene names | GAS7, KIAA0394 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
WWP1_MOUSE
|
||||||
| NC score | 0.119079 (rank : 19) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
WWP2_MOUSE
|
||||||
| NC score | 0.118548 (rank : 20) | θ value | 0.62314 (rank : 36) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9DBH0, Q8BTG4, Q923F6 | Gene names | Wwp2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2). | |||||
|
WWP2_HUMAN
|
||||||
| NC score | 0.118030 (rank : 21) | θ value | 0.62314 (rank : 35) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
ITCH_MOUSE
|
||||||
| NC score | 0.117547 (rank : 22) | θ value | 0.47712 (rank : 30) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8C863, O54971 | Gene names | Itch | |||
|
Domain Architecture |

|
|||||
| Description | Itchy E3 ubiquitin protein ligase (EC 6.3.2.-). | |||||
|
ITCH_HUMAN
|
||||||
| NC score | 0.116914 (rank : 23) | θ value | 0.47712 (rank : 29) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
IQGA2_HUMAN
|
||||||
| NC score | 0.116794 (rank : 24) | θ value | 0.47712 (rank : 28) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
WWP1_HUMAN
|
||||||
| NC score | 0.114104 (rank : 25) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
YAP1_MOUSE
|
||||||
| NC score | 0.112659 (rank : 26) | θ value | 1.81305 (rank : 51) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
PKCB1_HUMAN
|
||||||
| NC score | 0.104693 (rank : 27) | θ value | 0.00134147 (rank : 5) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
TRIPC_HUMAN
|
||||||
| NC score | 0.103513 (rank : 28) | θ value | 0.0961366 (rank : 14) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
WWOX_HUMAN
|
||||||
| NC score | 0.102602 (rank : 29) | θ value | 0.279714 (rank : 26) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NZC7, Q5MYT5, Q96KM3, Q96RF2, Q9BTT8, Q9NPC9, Q9NRF4, Q9NRF5, Q9NRF6, Q9NRK1, Q9NZC5 | Gene names | WWOX, FOR, WOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-) (Fragile site FRA16D oxidoreductase). | |||||
|
WWOX_MOUSE
|
||||||
| NC score | 0.101674 (rank : 30) | θ value | 0.279714 (rank : 27) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q91WL8, Q8C8J6, Q920Y2, Q9D2B3, Q9D339, Q9JLF5 | Gene names | Wwox, Wox1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-). | |||||
|
NED4L_HUMAN
|
||||||
| NC score | 0.100613 (rank : 31) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96PU5, O43165, Q7Z5F1, Q7Z5F2, Q7Z5N3, Q8N5A7, Q8WUU9, Q9BW58, Q9H2W4, Q9NT88 | Gene names | NEDD4L, KIAA0439, NEDL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
NEDD4_MOUSE
|
||||||
| NC score | 0.100148 (rank : 32) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.100104 (rank : 33) | θ value | 0.125558 (rank : 15) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
NED4L_MOUSE
|
||||||
| NC score | 0.098798 (rank : 34) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8CFI0, Q8BRT9, Q8BS42, Q99PK2 | Gene names | Nedd4l, Kiaa0439, Nedd4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
SMUF2_HUMAN
|
||||||
| NC score | 0.094245 (rank : 35) | θ value | 2.36792 (rank : 55) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9HAU4, Q9H260 | Gene names | SMURF2 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 2 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF2) (Smad-specific E3 ubiquitin ligase 2) (hSMURF2). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.094164 (rank : 36) | θ value | 0.813845 (rank : 37) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
RHG12_MOUSE
|
||||||
| NC score | 0.094092 (rank : 37) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
NEDD4_HUMAN
|
||||||
| NC score | 0.092204 (rank : 38) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P46934 | Gene names | NEDD4, KIAA0093 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-). | |||||
|
DGC14_MOUSE
|
||||||
| NC score | 0.091523 (rank : 39) | θ value | 0.279714 (rank : 22) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O70279, Q91YX1 | Gene names | Dgcr14, Dgsi, Es2, Es2el | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DGCR14 protein (DiGeorge syndrome critical region 14 homolog) (ES2 protein) (Expressed sequence 2 embryonic lethal). | |||||
|
K1688_MOUSE
|
||||||
| NC score | 0.089112 (rank : 40) | θ value | 0.279714 (rank : 23) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P59281, Q69ZD4, Q6P9R6 | Gene names | Kiaa1688, D15Wsu169e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1688. | |||||
|
YAP1_HUMAN
|
||||||
| NC score | 0.085363 (rank : 41) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P46937 | Gene names | YAP1, YAP65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
RHG12_HUMAN
|
||||||
| NC score | 0.084388 (rank : 42) | θ value | 1.06291 (rank : 45) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8IWW6, Q86UB3, Q8IWW7, Q8N3L1, Q9NT76 | Gene names | ARHGAP12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
SMUF1_MOUSE
|
||||||
| NC score | 0.078819 (rank : 43) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9CUN6, Q3U412, Q8K300 | Gene names | Smurf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1). | |||||
|
SMUF1_HUMAN
|
||||||
| NC score | 0.075982 (rank : 44) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9HCE7, O75853, Q9UJT8 | Gene names | SMURF1, KIAA1625 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1) (hSMURF1). | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.072749 (rank : 45) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.067220 (rank : 46) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
OXR1_MOUSE
|
||||||
| NC score | 0.067165 (rank : 47) | θ value | 0.47712 (rank : 32) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.067163 (rank : 48) | θ value | 1.38821 (rank : 48) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
HUWE1_MOUSE
|
||||||
| NC score | 0.066293 (rank : 49) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
HUWE1_HUMAN
|
||||||
| NC score | 0.065785 (rank : 50) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
PSIP1_MOUSE
|
||||||
| NC score | 0.065446 (rank : 51) | θ value | 1.06291 (rank : 44) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q99JF8, Q3TEJ7, Q3TTD7, Q3UTA1, Q80WQ7, Q99JF7, Q99LR4, Q9CT03 | Gene names | Psip1, Ledgf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (mLEDGF). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.065007 (rank : 52) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.064078 (rank : 53) | θ value | 2.36792 (rank : 54) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
SAV1_HUMAN
|
||||||
| NC score | 0.063903 (rank : 54) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9H4B6, Q6IA58, Q9H949, Q9HAK9 | Gene names | SAV1, WW45 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (hWW45). | |||||
|
DGC14_HUMAN
|
||||||
| NC score | 0.063847 (rank : 55) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96DF8, Q9BTZ4 | Gene names | DGCR14, DGSI, ES2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DGCR14 protein (DiGeorge syndrome critical region 14) (ES2 protein). | |||||
|
SAV1_MOUSE
|
||||||
| NC score | 0.062968 (rank : 56) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8VEB2, Q9D9N9, Q9ER46 | Gene names | Sav1, Ww45, Wwp3 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (mWW45). | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.062714 (rank : 57) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
NSD1_MOUSE
|
||||||
| NC score | 0.060220 (rank : 58) | θ value | 0.62314 (rank : 34) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
K0317_MOUSE
|
||||||
| NC score | 0.060116 (rank : 59) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CHG5, Q6P9Q1, Q80YC4, Q8C5W5 | Gene names | Kiaa0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
K0317_HUMAN
|
||||||
| NC score | 0.059930 (rank : 60) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O15033, Q7LDY1, Q8IYY9 | Gene names | KIAA0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
YTDC1_HUMAN
|
||||||
| NC score | 0.058506 (rank : 61) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
HECD2_MOUSE
|
||||||
| NC score | 0.058350 (rank : 62) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CDU6, Q8CBQ9 | Gene names | Hectd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
UBE3A_HUMAN
|
||||||
| NC score | 0.057909 (rank : 63) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q05086, P78355, Q93066, Q9UEP4, Q9UEP5, Q9UEP6, Q9UEP7, Q9UEP8, Q9UEP9 | Gene names | UBE3A, E6AP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (E6AP ubiquitin-protein ligase) (Oncogenic protein-associated protein E6-AP) (Human papillomavirus E6-associated protein) (NY-REN-54 antigen). | |||||
|
WWTR1_HUMAN
|
||||||
| NC score | 0.057840 (rank : 64) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9GZV5, Q8N3P2, Q9Y3W6 | Gene names | WWTR1, TAZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
HECD2_HUMAN
|
||||||
| NC score | 0.057813 (rank : 65) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5U5R9, Q5VZ97, Q5VZ99, Q8TCP5 | Gene names | HECTD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
WWC1_HUMAN
|
||||||
| NC score | 0.057659 (rank : 66) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8IX03, O94946, Q6MZX4, Q6Y2F8, Q7Z4G8, Q8WVM4, Q9BT29 | Gene names | WWC1, KIAA0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA) (HBeAg-binding protein 3). | |||||
|
WWC1_MOUSE
|
||||||
| NC score | 0.057486 (rank : 67) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5SXA9, Q571D0, Q8K1Y3, Q8VD17, Q922W3 | Gene names | Wwc1, Kiaa0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA). | |||||
|
NKTR_HUMAN
|
||||||
| NC score | 0.057447 (rank : 68) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
UBE3A_MOUSE
|
||||||
| NC score | 0.056949 (rank : 69) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O08759, P97482 | Gene names | Ube3a | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (Oncogenic protein- associated protein E6-AP). | |||||
|
UBE3C_MOUSE
|
||||||
| NC score | 0.056519 (rank : 70) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80U95, Q8BQZ6, Q8C7W6, Q8CDJ1, Q8VDL5 | Gene names | Ube3c, Kiaa0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
UBE3C_HUMAN
|
||||||
| NC score | 0.056410 (rank : 71) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15386, Q8TC15, Q96CR4, Q9UDU3 | Gene names | UBE3C, KIAA0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
WWTR1_MOUSE
|
||||||
| NC score | 0.056232 (rank : 72) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9EPK5, Q99KI4 | Gene names | Wwtr1, Taz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.054844 (rank : 73) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
CYLC2_HUMAN
|
||||||
| NC score | 0.053767 (rank : 74) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.053675 (rank : 75) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
ACINU_HUMAN
|
||||||
| NC score | 0.053273 (rank : 76) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
SFRIP_HUMAN
|
||||||
| NC score | 0.053216 (rank : 77) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.052218 (rank : 78) | θ value | 1.81305 (rank : 50) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
RSF1_HUMAN
|
||||||
| NC score | 0.052093 (rank : 79) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
BAG3_MOUSE
|
||||||
| NC score | 0.051834 (rank : 80) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
CALD1_HUMAN
|
||||||
| NC score | 0.050999 (rank : 81) | θ value | 0.62314 (rank : 33) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.050731 (rank : 82) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.050670 (rank : 83) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
DEK_MOUSE
|
||||||
| NC score | 0.050642 (rank : 84) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
THOC2_HUMAN
|
||||||
| NC score | 0.050241 (rank : 85) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
STXB4_MOUSE
|
||||||
| NC score | 0.048806 (rank : 86) | θ value | 2.36792 (rank : 56) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 629 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WV89, Q5SUA6, Q5SUA8, Q5SUA9, Q8CFL1 | Gene names | Stxbp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
ARI4A_HUMAN
|
||||||
| NC score | 0.048573 (rank : 87) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.048110 (rank : 88) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
POT15_HUMAN
|
||||||
| NC score | 0.045243 (rank : 89) | θ value | 0.125558 (rank : 16) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6S5H4, Q6NXN7, Q6S5H7 | Gene names | POTE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 15. | |||||
|
AFF2_MOUSE
|
||||||
| NC score | 0.044678 (rank : 90) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
MPP8_HUMAN
|
||||||
| NC score | 0.042687 (rank : 91) | θ value | 2.36792 (rank : 53) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
ANR21_HUMAN
|
||||||
| NC score | 0.042667 (rank : 92) | θ value | 0.279714 (rank : 21) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q86YR6 | Gene names | ANKRD21, POTE | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat domain-containing protein 21 (Protein POTE) (Prostate, ovary, testis-expressed protein). | |||||
|
STXB4_HUMAN
|
||||||
| NC score | 0.041019 (rank : 93) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 616 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6ZWJ1, Q8IVZ5 | Gene names | STXBP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.040910 (rank : 94) | θ value | 0.813845 (rank : 38) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
RHG09_HUMAN
|
||||||
| NC score | 0.040827 (rank : 95) | θ value | 3.0926 (rank : 62) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BRR9, Q8TCJ3, Q8WYR0, Q96EZ2, Q96S74 | Gene names | ARHGAP9 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 9. | |||||
|
NALP5_MOUSE
|
||||||
| NC score | 0.039321 (rank : 96) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
POTE8_HUMAN
|
||||||
| NC score | 0.036018 (rank : 97) | θ value | 1.06291 (rank : 43) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6S8J7, Q6S8J6 | Gene names | POTE8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 8. | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.034522 (rank : 98) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
ZC11A_MOUSE
|
||||||
| NC score | 0.032516 (rank : 99) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
CENPB_HUMAN
|
||||||
| NC score | 0.032009 (rank : 100) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P07199 | Gene names | CENPB | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
AP3B1_HUMAN
|
||||||
| NC score | 0.031952 (rank : 101) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
ANKZ1_MOUSE
|
||||||
| NC score | 0.031286 (rank : 102) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80UU1, Q9CZF5 | Gene names | Ankzf1, D1Ertd161e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and zinc finger domain-containing protein 1. | |||||
|
JHD2C_HUMAN
|
||||||
| NC score | 0.030526 (rank : 103) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15652, Q5SQZ8, Q5SQZ9, Q5SR00, Q7Z3E7, Q8N3U0, Q96KB9, Q9P2G7 | Gene names | JMJD1C, JHDM2C, KIAA1380, TRIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C) (Thyroid receptor-interacting protein 8) (TRIP-8). | |||||
|
NFM_HUMAN
|
||||||
| NC score | 0.030465 (rank : 104) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
RBP2_MOUSE
|
||||||
| NC score | 0.030233 (rank : 105) | θ value | 0.163984 (rank : 18) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ERU9, Q61992, Q8C9K9 | Gene names | Ranbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2 (RanBP2). | |||||
|
SMRC2_HUMAN
|
||||||
| NC score | 0.028450 (rank : 106) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
SMRC2_MOUSE
|
||||||
| NC score | 0.027994 (rank : 107) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
TEX14_MOUSE
|
||||||
| NC score | 0.027545 (rank : 108) | θ value | 0.125558 (rank : 17) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1333 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q7M6U3, Q3UT36, Q5NC10, Q5NC11, Q8CGK1, Q8CGK2, Q99MV8 | Gene names | Tex14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
NNP1_MOUSE
|
||||||
| NC score | 0.025604 (rank : 109) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P56183, O35712, Q9ERE1, Q9JI07, Q9JK67, Q9JKU2 | Gene names | Nnp1 | |||
|
Domain Architecture |

|
|||||
| Description | NNP-1 protein (Novel nuclear protein 1) (Nop52). | |||||
|
MNDA_HUMAN
|
||||||
| NC score | 0.023686 (rank : 110) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P41218 | Gene names | MNDA | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid cell nuclear differentiation antigen. | |||||
|
ENAM_MOUSE
|
||||||
| NC score | 0.023357 (rank : 111) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O55196 | Gene names | Enam | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enamelin precursor. | |||||
|
DCDC2_MOUSE
|
||||||
| NC score | 0.022772 (rank : 112) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
ARMC4_HUMAN
|
||||||
| NC score | 0.021419 (rank : 113) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
SCYBG_MOUSE
|
||||||
| NC score | 0.020536 (rank : 114) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BSU2, Q8VE25, Q9EPB3 | Gene names | Cxcl16, Srpsox | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small inducible cytokine B16 precursor (Transmembrane chemokine CXCL16) (SR-PSOX) (Scavenger receptor for phosphatidylserine and oxidized low density lipoprotein). | |||||
|
2A5D_HUMAN
|
||||||
| NC score | 0.020485 (rank : 115) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14738, O00494, O00696, Q15171 | Gene names | PPP2R5D | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit delta isoform (PP2A, B subunit, B' delta isoform) (PP2A, B subunit, B56 delta isoform) (PP2A, B subunit, PR61 delta isoform) (PP2A, B subunit, R5 delta isoform). | |||||
|
RNF25_HUMAN
|
||||||
| NC score | 0.020118 (rank : 116) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96BH1, Q9H874 | Gene names | RNF25 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 25 (EC 6.3.2.-). | |||||
|
SPTN5_HUMAN
|
||||||
| NC score | 0.019667 (rank : 117) | θ value | 0.813845 (rank : 39) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NRC6 | Gene names | SPTBN5, SPTBN4 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 4 (Spectrin, non-erythroid beta chain 4) (Beta-V spectrin) (BSPECV). | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.019121 (rank : 118) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.018590 (rank : 119) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
LBN_HUMAN
|
||||||
| NC score | 0.017518 (rank : 120) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86UK5, Q86YT3, Q86YT4, Q8NG49 | Gene names | EVC2, LBN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Limbin. | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.016824 (rank : 121) | θ value | 0.279714 (rank : 25) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
K1024_MOUSE
|
||||||
| NC score | 0.016345 (rank : 122) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K3V7, Q69ZS9 | Gene names | Kiaa1024 | |||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein KIAA1024 (Protein DD1). | |||||
|
DDX21_MOUSE
|
||||||
| NC score | 0.015610 (rank : 123) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
WASF2_MOUSE
|
||||||
| NC score | 0.015155 (rank : 124) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BH43 | Gene names | Wasf2, Wave2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2). | |||||
|
TACC3_HUMAN
|
||||||
| NC score | 0.014193 (rank : 125) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y6A5, Q9UMQ1 | Gene names | TACC3, ERIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ERIC-1). | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.010143 (rank : 126) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
GABPA_HUMAN
|
||||||
| NC score | 0.009293 (rank : 127) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q06546, Q12939 | Gene names | GABPA, E4TF1A | |||
|
Domain Architecture |

|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha) (Transcription factor E4TF1-60) (Nuclear respiratory factor 2 subunit alpha). | |||||
|
DC1I2_HUMAN
|
||||||
| NC score | 0.009157 (rank : 128) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13409, Q96NG7, Q96S87, Q9BXZ5, Q9NT58 | Gene names | DYNC1I2, DNCI2, DNCIC2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 2 (Dynein intermediate chain 2, cytosolic) (DH IC-2) (Cytoplasmic dynein intermediate chain 2). | |||||
|
HXC10_MOUSE
|
||||||
| NC score | 0.009032 (rank : 129) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P31257, P31312, Q7TMT7 | Gene names | Hoxc10, Hox-3.6, Hoxc-10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein Hox-C10 (Hox-3.6). | |||||
|
CAC1F_MOUSE
|
||||||
| NC score | 0.008314 (rank : 130) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JIS7 | Gene names | Cacna1f | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
TCF8_HUMAN
|
||||||
| NC score | 0.005367 (rank : 131) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
PEG3_MOUSE
|
||||||
| NC score | 0.001760 (rank : 132) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||