Please be patient as the page loads
|
K1024_MOUSE
|
||||||
| SwissProt Accessions | Q8K3V7, Q69ZS9 | Gene names | Kiaa1024 | |||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein KIAA1024 (Protein DD1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
K1024_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.986534 (rank : 2) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UPX6 | Gene names | KIAA1024 | |||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein KIAA1024. | |||||
|
K1024_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8K3V7, Q69ZS9 | Gene names | Kiaa1024 | |||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein KIAA1024 (Protein DD1). | |||||
|
U258_MOUSE
|
||||||
| θ value | 1.02238e-19 (rank : 3) | NC score | 0.703479 (rank : 4) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C4X7, Q6DI98 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein. | |||||
|
U258_HUMAN
|
||||||
| θ value | 1.38178e-16 (rank : 4) | NC score | 0.715893 (rank : 3) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P59773 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0258 protein. | |||||
|
KR122_HUMAN
|
||||||
| θ value | 0.21417 (rank : 5) | NC score | 0.048299 (rank : 5) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59991 | Gene names | KRTAP12-2, KAP12.2, KRTAP12.2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 12-2 (Keratin-associated protein 12.2) (High sulfur keratin-associated protein 12.2). | |||||
|
TRI15_HUMAN
|
||||||
| θ value | 0.279714 (rank : 6) | NC score | 0.026689 (rank : 17) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C019, O95604, Q8IUX9, Q9C018 | Gene names | TRIM15, RNF93, ZNF178, ZNFB7 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 15 (Zinc finger protein B7) (RING finger protein 93) (Zinc finger protein 178). | |||||
|
RABE2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 7) | NC score | 0.028530 (rank : 15) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 598 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q91WG2, Q99KN3 | Gene names | Rabep2, Rabpt5b | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 2 (Rabaptin-5beta). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 8) | NC score | 0.032380 (rank : 11) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
CHD9_MOUSE
|
||||||
| θ value | 0.813845 (rank : 9) | NC score | 0.025720 (rank : 18) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 10) | NC score | 0.031395 (rank : 12) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
K0241_HUMAN
|
||||||
| θ value | 1.06291 (rank : 11) | NC score | 0.038001 (rank : 6) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NBF6, Q92573 | Gene names | KIAA0241 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0241. | |||||
|
LASS2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 12) | NC score | 0.036021 (rank : 7) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q924Z4, Q3TXC5, Q9DCN6 | Gene names | Lass2, Trh3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LAG1 longevity assurance homolog 2 (Translocating chain-associating membrane protein homolog 3) (TRAM homolog 3). | |||||
|
PIWL3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 13) | NC score | 0.021033 (rank : 22) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z3Z3 | Gene names | PIWIL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 3. | |||||
|
TRIPC_HUMAN
|
||||||
| θ value | 1.38821 (rank : 14) | NC score | 0.033496 (rank : 9) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
PERI_HUMAN
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.012302 (rank : 39) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P41219, Q8N577 | Gene names | PRPH | |||
|
Domain Architecture |
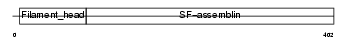
|
|||||
| Description | Peripherin. | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.034418 (rank : 8) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
ADH1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.014191 (rank : 34) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P00329 | Gene names | Adh1, Adh-1 | |||
|
Domain Architecture |
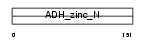
|
|||||
| Description | Alcohol dehydrogenase 1 (EC 1.1.1.1) (Alcohol dehydrogenase A subunit) (ADH-A2). | |||||
|
CCAR1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.029781 (rank : 13) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CH18, Q6AXC9, Q6PAR2, Q80XE4, Q8BJY0, Q8BVN2, Q8CGG1, Q9CSR5 | Gene names | Ccar1, Carp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle and apoptosis regulator protein 1 (Cell cycle and apoptosis regulatory protein 1) (CARP-1). | |||||
|
CHD1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.028096 (rank : 16) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
KI21B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 20) | NC score | 0.017552 (rank : 25) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
LIPA3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.015953 (rank : 29) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O75145, Q9H8B5, Q9UEW4 | Gene names | PPFIA3, KIAA0654 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
CAC1B_MOUSE
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.007963 (rank : 51) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O55017, Q60609 | Gene names | Cacna1b, Cach5, Cacnl1a5, Cchn1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
CAF1A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.018870 (rank : 24) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9QWF0 | Gene names | Chaf1a, Caip150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
CD2L7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 24) | NC score | 0.003289 (rank : 54) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
PERI_MOUSE
|
||||||
| θ value | 4.03905 (rank : 25) | NC score | 0.011148 (rank : 43) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P15331, O35688, O35689 | Gene names | Prph, Prph1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripherin. | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 4.03905 (rank : 26) | NC score | 0.003692 (rank : 53) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
ZN655_HUMAN
|
||||||
| θ value | 4.03905 (rank : 27) | NC score | 0.001589 (rank : 57) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N720, Q8IV00, Q8TA89, Q96EZ3 | Gene names | ZNF655, VIK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 655 (Vav-interacting Krueppel-like protein). | |||||
|
BRCA2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.021932 (rank : 20) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97929, O35922, P97383 | Gene names | Brca2, Fancd1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein homolog (Fanconi anemia group D1 protein homolog). | |||||
|
CHD9_HUMAN
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.021315 (rank : 21) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
E2AK2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.002486 (rank : 55) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P19525 | Gene names | EIF2AK2, PKR, PRKR | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced, double-stranded RNA-activated protein kinase (EC 2.7.11.1) (Interferon-inducible RNA-dependent protein kinase) (Protein kinase RNA-activated) (PKR) (p68 kinase) (P1/eIF-2A protein kinase). | |||||
|
GCC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.012993 (rank : 38) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
HSH2D_HUMAN
|
||||||
| θ value | 5.27518 (rank : 32) | NC score | 0.028732 (rank : 14) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96JZ2, Q6ZNG7 | Gene names | HSH2D, ALX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hematopoietic SH2 domain-containing protein (Hematopoietic SH2 protein) (Adaptor in lymphocytes of unknown function X). | |||||
|
KI21B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.015903 (rank : 30) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9QXL1, P97424 | Gene names | Kif21b, Kif6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B (Kinesin-like protein KIF6). | |||||
|
KRA59_HUMAN
|
||||||
| θ value | 5.27518 (rank : 34) | NC score | 0.009338 (rank : 48) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
MTMR7_MOUSE
|
||||||
| θ value | 5.27518 (rank : 35) | NC score | 0.016866 (rank : 27) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z2C9, Q8C4J6 | Gene names | Mtmr7 | |||
|
Domain Architecture |
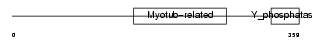
|
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
MYCB2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 36) | NC score | 0.014968 (rank : 32) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TPH6, Q6PCM8 | Gene names | Mycbp2, Pam, Phr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
RBP24_HUMAN
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.012997 (rank : 37) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
SRGP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.008395 (rank : 50) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z6B7, Q9H8A3, Q9P2P2 | Gene names | SRGAP1, ARHGAP13, KIAA1304 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 1 (srGAP1) (Rho-GTPase- activating protein 13). | |||||
|
AINX_MOUSE
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.009048 (rank : 49) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P46660, Q61958, Q8VCW5 | Gene names | Ina | |||
|
Domain Architecture |
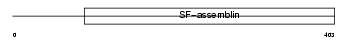
|
|||||
| Description | Alpha-internexin (Alpha-Inx) (66 kDa neurofilament protein) (Neurofilament-66) (NF-66). | |||||
|
B2L13_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.018956 (rank : 23) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BXK5, Q96B37, Q96IB7, Q9BY01, Q9HC05, Q9UFE0, Q9UKN3 | Gene names | BCL2L13, MIL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bcl-2-like 13 protein (Protein Mil1) (Bcl-rambo). | |||||
|
BCAR1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.009757 (rank : 46) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
CE152_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.013429 (rank : 35) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O94986, Q6NTA0 | Gene names | CEP152, KIAA0912 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 152 kDa (Cep152 protein). | |||||
|
CHD1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.032611 (rank : 10) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
DPOLZ_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.013066 (rank : 36) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61493, Q9JMD6, Q9QWX6 | Gene names | Rev3l, Polz, Sez4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (Seizure-related protein 4). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.017340 (rank : 26) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
JHD2C_MOUSE
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.011684 (rank : 41) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q69ZK6, Q6NV48, Q8BUF5, Q8C4I5, Q8C5Q9 | Gene names | Jmjd1c, Jhdm2c, Kiaa1380 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C). | |||||
|
LUZP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.014414 (rank : 33) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
MY18B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.012274 (rank : 40) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.023021 (rank : 19) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
ST18_HUMAN
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.011144 (rank : 44) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60284 | Gene names | ST18, KIAA0535, ZNF387 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18 (Zinc finger protein 387). | |||||
|
ANR26_MOUSE
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.007780 (rank : 52) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
AP3D1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.015111 (rank : 31) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O54774 | Gene names | Ap3d1, Ap3d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit delta-1 (Adapter-related protein complex 3 subunit delta-1) (Delta-adaptin 3) (AP-3 complex subunit delta) (Delta-adaptin) (mBLVR1). | |||||
|
KI21A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.011477 (rank : 42) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1543 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7Z4S6, Q6UKL9, Q7Z668, Q86WZ5, Q8IVZ8, Q9C0F5, Q9NXU4, Q9Y590 | Gene names | KIF21A, KIAA1708, KIF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21A (Kinesin-like protein KIF2) (NY-REN-62 antigen). | |||||
|
NF2L2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.010757 (rank : 45) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16236, Q96F71 | Gene names | NFE2L2, NRF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor erythroid 2-related factor 2 (NF-E2-related factor 2) (NFE2-related factor 2) (Nuclear factor, erythroid derived 2, like 2) (HEBP1). | |||||
|
NGAP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.009423 (rank : 47) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UJF2, O95174, Q5TFU9 | Gene names | RASAL2, NGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein nGAP (RAS protein activator-like 1). | |||||
|
WBP4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.016345 (rank : 28) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
ZN579_MOUSE
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.002403 (rank : 56) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 719 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80VM4 | Gene names | Znf579, Zfp579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 579. | |||||
|
K1024_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8K3V7, Q69ZS9 | Gene names | Kiaa1024 | |||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein KIAA1024 (Protein DD1). | |||||
|
K1024_HUMAN
|
||||||
| NC score | 0.986534 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UPX6 | Gene names | KIAA1024 | |||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein KIAA1024. | |||||
|
U258_HUMAN
|
||||||
| NC score | 0.715893 (rank : 3) | θ value | 1.38178e-16 (rank : 4) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P59773 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0258 protein. | |||||
|
U258_MOUSE
|
||||||
| NC score | 0.703479 (rank : 4) | θ value | 1.02238e-19 (rank : 3) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C4X7, Q6DI98 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein. | |||||
|
KR122_HUMAN
|
||||||
| NC score | 0.048299 (rank : 5) | θ value | 0.21417 (rank : 5) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59991 | Gene names | KRTAP12-2, KAP12.2, KRTAP12.2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 12-2 (Keratin-associated protein 12.2) (High sulfur keratin-associated protein 12.2). | |||||
|
K0241_HUMAN
|
||||||
| NC score | 0.038001 (rank : 6) | θ value | 1.06291 (rank : 11) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NBF6, Q92573 | Gene names | KIAA0241 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0241. | |||||
|
LASS2_MOUSE
|
||||||
| NC score | 0.036021 (rank : 7) | θ value | 1.06291 (rank : 12) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q924Z4, Q3TXC5, Q9DCN6 | Gene names | Lass2, Trh3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LAG1 longevity assurance homolog 2 (Translocating chain-associating membrane protein homolog 3) (TRAM homolog 3). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.034418 (rank : 8) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TRIPC_HUMAN
|
||||||
| NC score | 0.033496 (rank : 9) | θ value | 1.38821 (rank : 14) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
CHD1_MOUSE
|
||||||
| NC score | 0.032611 (rank : 10) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.032380 (rank : 11) | θ value | 0.62314 (rank : 8) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.031395 (rank : 12) | θ value | 0.813845 (rank : 10) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
CCAR1_MOUSE
|
||||||
| NC score | 0.029781 (rank : 13) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CH18, Q6AXC9, Q6PAR2, Q80XE4, Q8BJY0, Q8BVN2, Q8CGG1, Q9CSR5 | Gene names | Ccar1, Carp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle and apoptosis regulator protein 1 (Cell cycle and apoptosis regulatory protein 1) (CARP-1). | |||||
|
HSH2D_HUMAN
|
||||||
| NC score | 0.028732 (rank : 14) | θ value | 5.27518 (rank : 32) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96JZ2, Q6ZNG7 | Gene names | HSH2D, ALX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hematopoietic SH2 domain-containing protein (Hematopoietic SH2 protein) (Adaptor in lymphocytes of unknown function X). | |||||
|
RABE2_MOUSE
|
||||||
| NC score | 0.028530 (rank : 15) | θ value | 0.47712 (rank : 7) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 598 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q91WG2, Q99KN3 | Gene names | Rabep2, Rabpt5b | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 2 (Rabaptin-5beta). | |||||
|
CHD1_HUMAN
|
||||||
| NC score | 0.028096 (rank : 16) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
TRI15_HUMAN
|
||||||
| NC score | 0.026689 (rank : 17) | θ value | 0.279714 (rank : 6) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C019, O95604, Q8IUX9, Q9C018 | Gene names | TRIM15, RNF93, ZNF178, ZNFB7 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 15 (Zinc finger protein B7) (RING finger protein 93) (Zinc finger protein 178). | |||||
|
CHD9_MOUSE
|
||||||
| NC score | 0.025720 (rank : 18) | θ value | 0.813845 (rank : 9) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.023021 (rank : 19) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
BRCA2_MOUSE
|
||||||
| NC score | 0.021932 (rank : 20) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97929, O35922, P97383 | Gene names | Brca2, Fancd1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein homolog (Fanconi anemia group D1 protein homolog). | |||||
|
CHD9_HUMAN
|
||||||
| NC score | 0.021315 (rank : 21) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
PIWL3_HUMAN
|
||||||
| NC score | 0.021033 (rank : 22) | θ value | 1.06291 (rank : 13) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z3Z3 | Gene names | PIWIL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 3. | |||||
|
B2L13_HUMAN
|
||||||
| NC score | 0.018956 (rank : 23) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BXK5, Q96B37, Q96IB7, Q9BY01, Q9HC05, Q9UFE0, Q9UKN3 | Gene names | BCL2L13, MIL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bcl-2-like 13 protein (Protein Mil1) (Bcl-rambo). | |||||
|
CAF1A_MOUSE
|
||||||
| NC score | 0.018870 (rank : 24) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9QWF0 | Gene names | Chaf1a, Caip150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
KI21B_HUMAN
|
||||||
| NC score | 0.017552 (rank : 25) | θ value | 3.0926 (rank : 20) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.017340 (rank : 26) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
MTMR7_MOUSE
|
||||||
| NC score | 0.016866 (rank : 27) | θ value | 5.27518 (rank : 35) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z2C9, Q8C4J6 | Gene names | Mtmr7 | |||
|
Domain Architecture |

|
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
WBP4_MOUSE
|
||||||
| NC score | 0.016345 (rank : 28) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
LIPA3_HUMAN
|
||||||
| NC score | 0.015953 (rank : 29) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O75145, Q9H8B5, Q9UEW4 | Gene names | PPFIA3, KIAA0654 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
KI21B_MOUSE
|
||||||
| NC score | 0.015903 (rank : 30) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9QXL1, P97424 | Gene names | Kif21b, Kif6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B (Kinesin-like protein KIF6). | |||||
|
AP3D1_MOUSE
|
||||||
| NC score | 0.015111 (rank : 31) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O54774 | Gene names | Ap3d1, Ap3d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit delta-1 (Adapter-related protein complex 3 subunit delta-1) (Delta-adaptin 3) (AP-3 complex subunit delta) (Delta-adaptin) (mBLVR1). | |||||
|
MYCB2_MOUSE
|
||||||
| NC score | 0.014968 (rank : 32) | θ value | 5.27518 (rank : 36) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TPH6, Q6PCM8 | Gene names | Mycbp2, Pam, Phr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
LUZP1_MOUSE
|
||||||
| NC score | 0.014414 (rank : 33) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
ADH1_MOUSE
|
||||||
| NC score | 0.014191 (rank : 34) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P00329 | Gene names | Adh1, Adh-1 | |||
|
Domain Architecture |

|
|||||
| Description | Alcohol dehydrogenase 1 (EC 1.1.1.1) (Alcohol dehydrogenase A subunit) (ADH-A2). | |||||
|
CE152_HUMAN
|
||||||
| NC score | 0.013429 (rank : 35) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O94986, Q6NTA0 | Gene names | CEP152, KIAA0912 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 152 kDa (Cep152 protein). | |||||
|
DPOLZ_MOUSE
|
||||||
| NC score | 0.013066 (rank : 36) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61493, Q9JMD6, Q9QWX6 | Gene names | Rev3l, Polz, Sez4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (Seizure-related protein 4). | |||||
|
RBP24_HUMAN
|
||||||
| NC score | 0.012997 (rank : 37) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
GCC2_HUMAN
|
||||||
| NC score | 0.012993 (rank : 38) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
PERI_HUMAN
|
||||||
| NC score | 0.012302 (rank : 39) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P41219, Q8N577 | Gene names | PRPH | |||
|
Domain Architecture |

|
|||||
| Description | Peripherin. | |||||
|
MY18B_HUMAN
|
||||||
| NC score | 0.012274 (rank : 40) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
JHD2C_MOUSE
|
||||||
| NC score | 0.011684 (rank : 41) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q69ZK6, Q6NV48, Q8BUF5, Q8C4I5, Q8C5Q9 | Gene names | Jmjd1c, Jhdm2c, Kiaa1380 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C). | |||||
|
KI21A_HUMAN
|
||||||
| NC score | 0.011477 (rank : 42) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1543 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7Z4S6, Q6UKL9, Q7Z668, Q86WZ5, Q8IVZ8, Q9C0F5, Q9NXU4, Q9Y590 | Gene names | KIF21A, KIAA1708, KIF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21A (Kinesin-like protein KIF2) (NY-REN-62 antigen). | |||||
|
PERI_MOUSE
|
||||||
| NC score | 0.011148 (rank : 43) | θ value | 4.03905 (rank : 25) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P15331, O35688, O35689 | Gene names | Prph, Prph1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripherin. | |||||
|
ST18_HUMAN
|
||||||
| NC score | 0.011144 (rank : 44) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60284 | Gene names | ST18, KIAA0535, ZNF387 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18 (Zinc finger protein 387). | |||||
|
NF2L2_HUMAN
|
||||||
| NC score | 0.010757 (rank : 45) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16236, Q96F71 | Gene names | NFE2L2, NRF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor erythroid 2-related factor 2 (NF-E2-related factor 2) (NFE2-related factor 2) (Nuclear factor, erythroid derived 2, like 2) (HEBP1). | |||||
|
BCAR1_HUMAN
|
||||||
| NC score | 0.009757 (rank : 46) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
NGAP_HUMAN
|
||||||
| NC score | 0.009423 (rank : 47) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UJF2, O95174, Q5TFU9 | Gene names | RASAL2, NGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein nGAP (RAS protein activator-like 1). | |||||
|
KRA59_HUMAN
|
||||||
| NC score | 0.009338 (rank : 48) | θ value | 5.27518 (rank : 34) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
AINX_MOUSE
|
||||||
| NC score | 0.009048 (rank : 49) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P46660, Q61958, Q8VCW5 | Gene names | Ina | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-internexin (Alpha-Inx) (66 kDa neurofilament protein) (Neurofilament-66) (NF-66). | |||||
|
SRGP1_HUMAN
|
||||||
| NC score | 0.008395 (rank : 50) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z6B7, Q9H8A3, Q9P2P2 | Gene names | SRGAP1, ARHGAP13, KIAA1304 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 1 (srGAP1) (Rho-GTPase- activating protein 13). | |||||
|
CAC1B_MOUSE
|
||||||
| NC score | 0.007963 (rank : 51) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O55017, Q60609 | Gene names | Cacna1b, Cach5, Cacnl1a5, Cchn1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
ANR26_MOUSE
|
||||||
| NC score | 0.007780 (rank : 52) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.003692 (rank : 53) | θ value | 4.03905 (rank : 26) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
CD2L7_HUMAN
|
||||||
| NC score | 0.003289 (rank : 54) | θ value | 4.03905 (rank : 24) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
E2AK2_HUMAN
|
||||||
| NC score | 0.002486 (rank : 55) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P19525 | Gene names | EIF2AK2, PKR, PRKR | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced, double-stranded RNA-activated protein kinase (EC 2.7.11.1) (Interferon-inducible RNA-dependent protein kinase) (Protein kinase RNA-activated) (PKR) (p68 kinase) (P1/eIF-2A protein kinase). | |||||
|
ZN579_MOUSE
|
||||||
| NC score | 0.002403 (rank : 56) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 719 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80VM4 | Gene names | Znf579, Zfp579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 579. | |||||
|
ZN655_HUMAN
|
||||||
| NC score | 0.001589 (rank : 57) | θ value | 4.03905 (rank : 27) | |||
| Query Neighborhood Hits | 57 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N720, Q8IV00, Q8TA89, Q96EZ3 | Gene names | ZNF655, VIK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 655 (Vav-interacting Krueppel-like protein). | |||||