Please be patient as the page loads
|
PKP4_MOUSE
|
||||||
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CTND2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.951748 (rank : 4) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
CTND2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.964230 (rank : 3) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O35927 | Gene names | Ctnnd2, Catnd2, Nprap | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin). | |||||
|
PKP4_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.993729 (rank : 2) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
PKP4_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 129 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
ARVC_HUMAN
|
||||||
| θ value | 1.80368e-133 (rank : 5) | NC score | 0.947803 (rank : 6) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O00192 | Gene names | ARVCF | |||
|
Domain Architecture |

|
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome. | |||||
|
ARVC_MOUSE
|
||||||
| θ value | 5.80223e-132 (rank : 6) | NC score | 0.949704 (rank : 5) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
CTND1_MOUSE
|
||||||
| θ value | 5.26011e-125 (rank : 7) | NC score | 0.944051 (rank : 7) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P30999 | Gene names | Ctnnd1, Catns | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
CTND1_HUMAN
|
||||||
| θ value | 2.88631e-123 (rank : 8) | NC score | 0.941782 (rank : 8) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60716, O15088, O60713, O60714, O60715, O60935, Q6RBX8, Q9UP71, Q9UP72, Q9UP73 | Gene names | CTNND1, KIAA0384 | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
PKP1_MOUSE
|
||||||
| θ value | 1.7756e-72 (rank : 9) | NC score | 0.903597 (rank : 10) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P97350 | Gene names | Pkp1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1. | |||||
|
PKP2_HUMAN
|
||||||
| θ value | 5.71191e-71 (rank : 10) | NC score | 0.913060 (rank : 9) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99959, Q99960 | Gene names | PKP2 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-2. | |||||
|
PKP1_HUMAN
|
||||||
| θ value | 2.83479e-70 (rank : 11) | NC score | 0.902626 (rank : 11) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13835, O00645, Q15152 | Gene names | PKP1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1 (Band-6 protein) (B6P). | |||||
|
PKP3_HUMAN
|
||||||
| θ value | 2.93355e-67 (rank : 12) | NC score | 0.901271 (rank : 12) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP3_MOUSE
|
||||||
| θ value | 2.93355e-67 (rank : 13) | NC score | 0.900334 (rank : 13) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
ARMC4_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 14) | NC score | 0.378102 (rank : 14) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
IMA7_MOUSE
|
||||||
| θ value | 0.000121331 (rank : 15) | NC score | 0.188309 (rank : 21) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O35345, Q9CVP9 | Gene names | Kpna6, Kpna5 | |||
|
Domain Architecture |
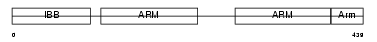
|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6 subunit) (Importin alpha S2). | |||||
|
IMA7_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 16) | NC score | 0.184223 (rank : 22) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |
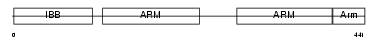
|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
APC_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 17) | NC score | 0.227933 (rank : 18) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
APC_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 18) | NC score | 0.221281 (rank : 19) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
CTNB1_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 19) | NC score | 0.275142 (rank : 16) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P35222, Q8NEW9, Q8NI94, Q9H391 | Gene names | CTNNB1, CTNNB | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
CTNB1_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 20) | NC score | 0.275209 (rank : 15) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q02248, Q922W1, Q9D335 | Gene names | Ctnnb1, Catnb | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
IMA2_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 21) | NC score | 0.169700 (rank : 26) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |
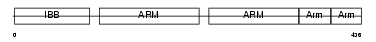
|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
IMA2_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 22) | NC score | 0.170658 (rank : 25) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52293, Q64292 | Gene names | Kpna2, Rch1 | |||
|
Domain Architecture |
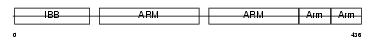
|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1) (Pendulin) (Pore targeting complex 58 kDa subunit) (PTAC58) (Importin alpha P1). | |||||
|
IMA4_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 23) | NC score | 0.162241 (rank : 28) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O00629, O00190 | Gene names | KPNA4, QIP1 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Qip1 protein). | |||||
|
IMA4_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 24) | NC score | 0.162253 (rank : 27) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O35343 | Gene names | Kpna4 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Importin alpha Q1). | |||||
|
K1718_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 25) | NC score | 0.030796 (rank : 59) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZMT4, Q6MZL8, Q9C0E5 | Gene names | KIAA1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
IMA5_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 26) | NC score | 0.177495 (rank : 23) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O15131 | Gene names | KPNA5 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-6 subunit (Karyopherin alpha-5 subunit). | |||||
|
PLAK_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 27) | NC score | 0.246601 (rank : 17) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q02257, Q8CGD3 | Gene names | Jup | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 28) | NC score | 0.048421 (rank : 48) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
CDKL5_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 29) | NC score | 0.006928 (rank : 112) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O76039, Q14198, Q5H985, Q9UJL6 | Gene names | CDKL5, STK9 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase-like 5 (EC 2.7.11.22) (Serine/threonine- protein kinase 9). | |||||
|
MAVS_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 30) | NC score | 0.051596 (rank : 44) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7Z434, Q3I0Y2, Q5T7I6, Q86VY7, Q9H1H3, Q9H4Y1, Q9H8D3, Q9ULE9 | Gene names | MAVS, IPS1, KIAA1271, VISA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif) (Putative NF-kappa-B- activating protein 031N). | |||||
|
PLAK_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 31) | NC score | 0.209943 (rank : 20) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P14923, Q9HCX9 | Gene names | JUP, DP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
SLK_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 32) | NC score | 0.001771 (rank : 128) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1798 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O54988, Q80U65, Q8CAU2, Q8CDW2, Q9WU41 | Gene names | Slk, Kiaa0204, Stk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | STE20-like serine/threonine-protein kinase (EC 2.7.11.1) (STE20-like kinase) (STE20-related serine/threonine-protein kinase) (STE20-related kinase) (mSLK) (Serine/threonine-protein kinase 2) (STE20-related kinase SMAK) (Etk4). | |||||
|
IMA1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 33) | NC score | 0.147508 (rank : 31) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52294, Q9BQ56 | Gene names | KPNA1, RCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (NPI-1). | |||||
|
IMA1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 34) | NC score | 0.147237 (rank : 32) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q60960, Q3TF32 | Gene names | Kpna1, Rch2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (Importin alpha S1). | |||||
|
IMA3_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 35) | NC score | 0.154850 (rank : 30) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00505, O00191, O43195, Q96AA7 | Gene names | KPNA3 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (SRP1-gamma). | |||||
|
IMA3_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 36) | NC score | 0.154986 (rank : 29) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O35344 | Gene names | Kpna3 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (Importin alpha Q2). | |||||
|
FILA_HUMAN
|
||||||
| θ value | 0.125558 (rank : 37) | NC score | 0.025564 (rank : 66) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
ARMX3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 38) | NC score | 0.063452 (rank : 37) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UH62, Q53HC6, Q7LCF5, Q9NPE4 | Gene names | ARMCX3, ALEX3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 3 (Protein ALEX3) (ARM protein lost in epithelial cancers on chromosome X 3). | |||||
|
ARMX3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 39) | NC score | 0.063463 (rank : 36) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BHS6, Q91VP8, Q9DC32 | Gene names | Armcx3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 3. | |||||
|
SMN_HUMAN
|
||||||
| θ value | 0.21417 (rank : 40) | NC score | 0.047338 (rank : 50) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16637, Q13119, Q96J51 | Gene names | SMN1, SMN, SMNT | |||
|
Domain Architecture |
|
|||||
| Description | Survival motor neuron protein (Component of gems 1) (Gemin-1). | |||||
|
EPCR_HUMAN
|
||||||
| θ value | 0.279714 (rank : 41) | NC score | 0.048496 (rank : 47) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UNN8, Q14218, Q6IB56, Q96CB3, Q9ULX1 | Gene names | PROCR, EPCR | |||
|
Domain Architecture |
|
|||||
| Description | Endothelial protein C receptor precursor (Endothelial cell protein C receptor) (Activated protein C receptor) (APC receptor) (CD201 antigen). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 42) | NC score | 0.063690 (rank : 35) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
NFRKB_MOUSE
|
||||||
| θ value | 0.365318 (rank : 43) | NC score | 0.040203 (rank : 55) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
SPTN5_HUMAN
|
||||||
| θ value | 0.365318 (rank : 44) | NC score | 0.013884 (rank : 93) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NRC6 | Gene names | SPTBN5, SPTBN4 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 4 (Spectrin, non-erythroid beta chain 4) (Beta-V spectrin) (BSPECV). | |||||
|
TAF9_HUMAN
|
||||||
| θ value | 0.365318 (rank : 45) | NC score | 0.032167 (rank : 57) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q16594, Q9BTS1 | Gene names | TAF9, TAF2G, TAFII31 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 9 (Transcription initiation factor TFIID 31 kDa subunit) (TAFII-31) (TAFII-32) (TAFII32) (STAF31/32). | |||||
|
TRI66_MOUSE
|
||||||
| θ value | 0.365318 (rank : 46) | NC score | 0.019764 (rank : 77) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
HEAT3_MOUSE
|
||||||
| θ value | 0.47712 (rank : 47) | NC score | 0.080227 (rank : 33) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BQM4, Q8C731 | Gene names | Heatr3 | |||
|
Domain Architecture |
|
|||||
| Description | HEAT repeat-containing protein 3. | |||||
|
AFF4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 48) | NC score | 0.026953 (rank : 64) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
AOC2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 49) | NC score | 0.015384 (rank : 88) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q812C9, Q80WP3 | Gene names | Aoc2 | |||
|
Domain Architecture |

|
|||||
| Description | Retina-specific copper amine oxidase precursor (EC 1.4.3.6) (RAO) (Amine oxidase [copper-containing]). | |||||
|
FLIP1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 50) | NC score | 0.004594 (rank : 119) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 51) | NC score | 0.023708 (rank : 70) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
PDE1C_MOUSE
|
||||||
| θ value | 0.62314 (rank : 52) | NC score | 0.009690 (rank : 103) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64338, Q62045, Q8BSV6 | Gene names | Pde1c | |||
|
Domain Architecture |
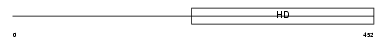
|
|||||
| Description | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C (EC 3.1.4.17) (Cam-PDE 1C). | |||||
|
PRC1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 53) | NC score | 0.029986 (rank : 61) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43663, Q9BSB6 | Gene names | PRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein regulator of cytokinesis 1. | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.813845 (rank : 54) | NC score | 0.021578 (rank : 76) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
COE1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 55) | NC score | 0.022402 (rank : 73) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UH73, Q8IW11 | Gene names | EBF1, COE1, EBF | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE1 (OE-1) (O/E-1) (Early B-cell factor). | |||||
|
COE1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 56) | NC score | 0.022402 (rank : 74) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q07802 | Gene names | Ebf1, Coe1, Ebf | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE1 (OE-1) (O/E-1) (Early B-cell factor). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 1.38821 (rank : 57) | NC score | 0.025344 (rank : 67) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
GAB1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.024656 (rank : 69) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QYY0, Q91VW7 | Gene names | Gab1 | |||
|
Domain Architecture |
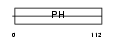
|
|||||
| Description | GRB2-associated-binding protein 1 (GRB2-associated binder 1). | |||||
|
GDS1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.176464 (rank : 24) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P52306, Q9NYM2, Q9NZA8 | Gene names | RAP1GDS1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-GDP dissociation stimulator 1 (SMG P21 stimulatory GDP/GTP exchange protein) (SMG GDS protein) (Exchange factor smgGDS). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.041581 (rank : 53) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
ZMYM1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.016802 (rank : 84) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5SVZ6, Q7Z3Q4 | Gene names | ZMYM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger MYM-type protein 1. | |||||
|
KLF12_HUMAN
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.001879 (rank : 126) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 717 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y4X4, Q5VZM7, Q9UHZ0 | Gene names | KLF12, AP2REP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 12 (Transcriptional repressor AP-2rep). | |||||
|
NCOR1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.022486 (rank : 72) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
TAF9_MOUSE
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.026093 (rank : 65) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VI33, Q32P09, Q80XS3, Q9CV61 | Gene names | Taf9, Taf2g | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 9 (Transcription initiation factor TFIID 31 kDa subunit) (TAFII-31) (TAFII-32) (TAFII32). | |||||
|
CCD27_MOUSE
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.019394 (rank : 78) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3V036 | Gene names | Ccdc27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 27. | |||||
|
CHD9_HUMAN
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.008893 (rank : 105) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
PDXL2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.029220 (rank : 62) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CAE9, Q8CFW3 | Gene names | Podxl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
RIMB1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.016352 (rank : 85) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
SFR12_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.015551 (rank : 87) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
SFR12_MOUSE
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.017277 (rank : 82) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
TLN2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.013783 (rank : 94) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q71LX4, Q8BWK0 | Gene names | Tln2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Talin-2. | |||||
|
TNR8_HUMAN
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.014762 (rank : 90) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P28908 | Gene names | TNFRSF8, CD30 | |||
|
Domain Architecture |
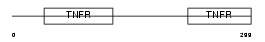
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 8 precursor (CD30L receptor) (Lymphocyte activation antigen CD30) (KI-1 antigen). | |||||
|
AKAP2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.030083 (rank : 60) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
ARMC6_MOUSE
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.053356 (rank : 42) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BNU0, Q8C4P5, Q8C7S1, Q8K2S7, Q99JQ2, Q9CWE4 | Gene names | Armc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 6. | |||||
|
ARMX1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 75) | NC score | 0.043127 (rank : 52) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9CX83 | Gene names | Armcx1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 1. | |||||
|
BAP1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.022243 (rank : 75) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92560, Q6LEM0, Q7Z5E8 | Gene names | BAP1, KIAA0272 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase BAP1 (EC 3.4.19.12) (BRCA1- associated protein 1) (Cerebral protein 6). | |||||
|
CHD9_MOUSE
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.017799 (rank : 80) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
BMP2K_MOUSE
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.003849 (rank : 121) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q91Z96, Q8C8L7 | Gene names | Bmp2k, Bike | |||
|
Domain Architecture |
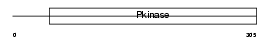
|
|||||
| Description | BMP-2-inducible protein kinase (EC 2.7.11.1) (BIKe). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 79) | NC score | 0.025235 (rank : 68) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
DOT1L_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | 0.041521 (rank : 54) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
MAML1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | 0.027306 (rank : 63) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6T264, Q505D8, Q5SUC2, Q6PDK3, Q6ZQG5, Q8BIU5, Q8R3T0 | Gene names | Maml1, Kiaa0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
MUCDL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.013641 (rank : 96) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8VHF2, Q8CEJ3, Q9D8I9 | Gene names | Mucdhl | |||
|
Domain Architecture |
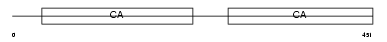
|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
RNF12_MOUSE
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | 0.007839 (rank : 108) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WTV7 | Gene names | Rnf12, Rlim | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 12 (LIM domain-interacting RING finger protein) (RING finger LIM domain-binding protein) (R-LIM). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.031128 (rank : 58) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 85) | NC score | 0.050616 (rank : 46) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
UBP47_MOUSE
|
||||||
| θ value | 4.03905 (rank : 86) | NC score | 0.007901 (rank : 107) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BY87, Q5EBP2, Q6KAR9, Q80V06, Q8BHU1, Q8BI15, Q8BI16, Q8BUW4, Q91X25 | Gene names | Usp47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
ZN692_MOUSE
|
||||||
| θ value | 4.03905 (rank : 87) | NC score | -0.001563 (rank : 135) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3U381 | Gene names | Znf692, Zfp692 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 692. | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | 0.010264 (rank : 102) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
HCFC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.011727 (rank : 100) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q61191, Q684R1, Q7TSB0, Q8C2D0, Q9QWH2 | Gene names | Hcfc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N- terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C-terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
HEAT3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 90) | NC score | 0.053229 (rank : 43) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7Z4Q2, Q8N525, Q8WV56, Q96CC9, Q9NWN7 | Gene names | HEATR3 | |||
|
Domain Architecture |
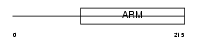
|
|||||
| Description | HEAT repeat-containing protein 3. | |||||
|
MTMR7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 91) | NC score | 0.012309 (rank : 98) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y216, Q68DX4 | Gene names | MTMR7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
NPHN_MOUSE
|
||||||
| θ value | 5.27518 (rank : 92) | NC score | 0.002785 (rank : 124) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QZS7 | Gene names | Nphs1, Nphn | |||
|
Domain Architecture |

|
|||||
| Description | Nephrin precursor (Renal glomerulus-specific cell adhesion receptor). | |||||
|
SNP29_HUMAN
|
||||||
| θ value | 5.27518 (rank : 93) | NC score | 0.012092 (rank : 99) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95721 | Gene names | SNAP29 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptosomal-associated protein 29 (SNAP-29) (Vesicle-membrane fusion protein SNAP-29) (Soluble 29 kDa NSF attachment protein). | |||||
|
SYNJ1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 94) | NC score | 0.009490 (rank : 104) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
TAB3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 95) | NC score | 0.018361 (rank : 79) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N5C8, Q6VQR0 | Gene names | MAP3K7IP3, TAB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 3 (TAK1-binding protein 3) (NF-kappa-B-activating protein 1). | |||||
|
TAB3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 96) | NC score | 0.017143 (rank : 83) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q571K4 | Gene names | Map3k7ip3, Kiaa4135, Tab3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 3 (TAK1-binding protein 3). | |||||
|
ARMX1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.048308 (rank : 49) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9P291, Q53HK2, Q9H2Q0 | Gene names | ARMCX1, ALEX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 1 (Protein ALEX1) (ARM protein lost in epithelial cancers on chromosome X 1). | |||||
|
ATX1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.014050 (rank : 92) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P54254 | Gene names | Atxn1, Sca1 | |||
|
Domain Architecture |

|
|||||
| Description | Ataxin-1 (Spinocerebellar ataxia type 1 protein homolog). | |||||
|
CCNL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.006913 (rank : 113) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96S94, Q5T2N5, Q5T2N6, Q6IQ12, Q7Z4Z8, Q8N3C9, Q8N3D5, Q8NHE3, Q8TEL0, Q96B00 | Gene names | CCNL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L2 (Paneth cell-enhanced expression protein). | |||||
|
CO002_HUMAN
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | 0.013672 (rank : 95) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
DAG1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 101) | NC score | 0.023680 (rank : 71) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14118 | Gene names | DAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Dystroglycan precursor (Dystrophin-associated glycoprotein 1) [Contains: Alpha-dystroglycan (Alpha-DG); Beta-dystroglycan (Beta- DG)]. | |||||
|
EGR1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.000119 (rank : 132) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P08046, Q61777 | Gene names | Egr1, Egr-1, Krox-24 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 1 (EGR-1) (Krox-24 protein) (Transcription factor Zif268) (Nerve growth factor-induced protein A) (NGFI-A). | |||||
|
FA5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.004417 (rank : 120) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P12259, Q14285, Q6UPU6 | Gene names | F5 | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor V precursor (Activated protein C cofactor) [Contains: Coagulation factor V heavy chain; Coagulation factor V light chain]. | |||||
|
GP125_HUMAN
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.003027 (rank : 123) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IWK6, Q6UXK9, Q86SQ5, Q8TC55 | Gene names | GPR125 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 125 precursor. | |||||
|
HIF1N_MOUSE
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.033571 (rank : 56) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BLR9, Q3U3G4 | Gene names | Hif1an | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha inhibitor (EC 1.14.11.16) (Hypoxia- inducible factor asparagine hydroxylase). | |||||
|
MAML3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.014811 (rank : 89) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96JK9 | Gene names | MAML3, KIAA1816 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 3 (Mam-3). | |||||
|
MTMRD_HUMAN
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.005514 (rank : 117) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86WG5, Q3MJF0, Q68DQ3, Q6P459, Q6PJD1, Q7Z325, Q7Z621, Q86VE2, Q96FE2, Q9C097 | Gene names | SBF2, CMT4B2, KIAA1766, MTMR13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 13 (SET-binding factor 2). | |||||
|
PHF2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.014729 (rank : 91) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PO2F3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.001791 (rank : 127) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P31362 | Gene names | Pou2f3, Epoc1, Oct11, Otf-11, Otf11 | |||
|
Domain Architecture |
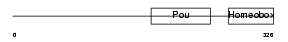
|
|||||
| Description | POU domain, class 2, transcription factor 3 (Octamer-binding transcription factor 11) (Oct-11) (Epoc-1). | |||||
|
SMAD2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.005953 (rank : 116) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15796 | Gene names | SMAD2, MADH2, MADR2 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 2 (SMAD 2) (Mothers against DPP homolog 2) (Mad-related protein 2) (hMAD-2) (JV18-1) (hSMAD2). | |||||
|
SMAD2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.005954 (rank : 115) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62432, Q9D8P6 | Gene names | Smad2, Madh2, Madr2 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 2 (SMAD 2) (Mothers against DPP homolog 2) (Mad-related protein 2) (mMad2). | |||||
|
TR150_HUMAN
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.012680 (rank : 97) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2W1, Q5VTK6 | Gene names | THRAP3, TRAP150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
WNK1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | 0.003077 (rank : 122) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1179 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P83741 | Gene names | Wnk1, Prkwnk1 | |||
|
Domain Architecture |
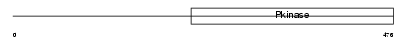
|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1). | |||||
|
ZCH10_HUMAN
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | 0.016124 (rank : 86) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TBK6, Q9NXR4 | Gene names | ZCCHC10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 10. | |||||
|
ZFAN3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | 0.010454 (rank : 101) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q497H0, Q8K230, Q8R2V7, Q8R3V1 | Gene names | Zfand3, Tex27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AN1-type zinc finger protein 3 (Testis-expressed sequence 27). | |||||
|
ABL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | -0.001502 (rank : 134) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1045 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P42684, Q5T0X6, Q6NZY6 | Gene names | ABL2, ABLL, ARG | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase ABL2 (EC 2.7.10.2) (Abelson murine leukemia viral oncogene homolog 2) (Tyrosine kinase ARG). | |||||
|
ANK3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.002664 (rank : 125) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 853 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q12955 | Gene names | ANK3 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-3 (ANK-3) (Ankyrin-G). | |||||
|
COA2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.007209 (rank : 111) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O00763, Q16852 | Gene names | ACACB, ACC2, ACCB | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 2 (EC 6.4.1.2) (ACC-beta) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.017455 (rank : 81) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.045527 (rank : 51) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
MYOTI_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.001388 (rank : 130) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UBF9 | Gene names | MYOT, TTID | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotilin (Titin immunoglobulin domain protein) (Myofibrillar titin- like Ig domains protein) (57 kDa cytoskeletal protein). | |||||
|
PAX3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.000347 (rank : 131) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P23760, Q16448 | Gene names | PAX3, HUP2 | |||
|
Domain Architecture |
|
|||||
| Description | Paired box protein Pax-3 (HUP2). | |||||
|
TENS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.007691 (rank : 110) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9HBL0 | Gene names | TNS1, TNS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-1. | |||||
|
TLN2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.007700 (rank : 109) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y4G6 | Gene names | TLN2, KIAA0320 | |||
|
Domain Architecture |

|
|||||
| Description | Talin-2. | |||||
|
UBP47_HUMAN
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.006483 (rank : 114) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96K76, Q658U0, Q86Y73, Q8TEP6, Q9BWI0, Q9NWN1 | Gene names | USP47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
WT1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | -0.000074 (rank : 133) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P19544, Q15881, Q16256, Q16575, Q8IYZ5 | Gene names | WT1 | |||
|
Domain Architecture |

|
|||||
| Description | Wilms' tumor protein (WT33). | |||||
|
ZBTB3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 127) | NC score | 0.001482 (rank : 129) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 910 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9H5J0 | Gene names | ZBTB3 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 3. | |||||
|
ZCH14_MOUSE
|
||||||
| θ value | 8.99809 (rank : 128) | NC score | 0.008063 (rank : 106) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VIG0, Q80TX3, Q91VU4 | Gene names | Zcchc14, Kiaa0579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 14 (BDG-29). | |||||
|
ZCHC6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 129) | NC score | 0.005370 (rank : 118) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5BLK4, Q8CIH3 | Gene names | Zcchc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 6. | |||||
|
CK024_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.056176 (rank : 39) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96F05, Q9H2K4 | Gene names | C11orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 precursor (Protein DM4E3). | |||||
|
MUC13_MOUSE
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.053837 (rank : 41) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
PARM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.054268 (rank : 40) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q923D3, Q3TTV0, Q3UFU4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PARM-1 precursor. | |||||
|
RTDR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.050655 (rank : 45) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UHP6 | Gene names | RTDR1 | |||
|
Domain Architecture |

|
|||||
| Description | Rhabdoid tumor deletion region protein 1. | |||||
|
SPAG6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.070126 (rank : 34) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O75602, Q5VUX5, Q5VUX6, Q5VUX7, Q6FI74, Q8NHQ6 | Gene names | SPAG6, PF16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Sperm flagellar protein) (Repro-SA-1). | |||||
|
SPAG6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.063158 (rank : 38) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JLI7, Q8K461 | Gene names | Spag6, Pf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Axoneme central apparatus protein). | |||||
|
PKP4_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 129 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
PKP4_HUMAN
|
||||||
| NC score | 0.993729 (rank : 2) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
CTND2_MOUSE
|
||||||
| NC score | 0.964230 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O35927 | Gene names | Ctnnd2, Catnd2, Nprap | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin). | |||||
|
CTND2_HUMAN
|
||||||
| NC score | 0.951748 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
ARVC_MOUSE
|
||||||
| NC score | 0.949704 (rank : 5) | θ value | 5.80223e-132 (rank : 6) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
ARVC_HUMAN
|
||||||
| NC score | 0.947803 (rank : 6) | θ value | 1.80368e-133 (rank : 5) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O00192 | Gene names | ARVCF | |||
|
Domain Architecture |

|
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome. | |||||
|
CTND1_MOUSE
|
||||||
| NC score | 0.944051 (rank : 7) | θ value | 5.26011e-125 (rank : 7) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P30999 | Gene names | Ctnnd1, Catns | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
CTND1_HUMAN
|
||||||
| NC score | 0.941782 (rank : 8) | θ value | 2.88631e-123 (rank : 8) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60716, O15088, O60713, O60714, O60715, O60935, Q6RBX8, Q9UP71, Q9UP72, Q9UP73 | Gene names | CTNND1, KIAA0384 | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
PKP2_HUMAN
|
||||||
| NC score | 0.913060 (rank : 9) | θ value | 5.71191e-71 (rank : 10) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99959, Q99960 | Gene names | PKP2 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-2. | |||||
|
PKP1_MOUSE
|
||||||
| NC score | 0.903597 (rank : 10) | θ value | 1.7756e-72 (rank : 9) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P97350 | Gene names | Pkp1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1. | |||||
|
PKP1_HUMAN
|
||||||
| NC score | 0.902626 (rank : 11) | θ value | 2.83479e-70 (rank : 11) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13835, O00645, Q15152 | Gene names | PKP1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1 (Band-6 protein) (B6P). | |||||
|
PKP3_HUMAN
|
||||||
| NC score | 0.901271 (rank : 12) | θ value | 2.93355e-67 (rank : 12) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP3_MOUSE
|
||||||
| NC score | 0.900334 (rank : 13) | θ value | 2.93355e-67 (rank : 13) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
ARMC4_HUMAN
|
||||||
| NC score | 0.378102 (rank : 14) | θ value | 4.1701e-05 (rank : 14) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
CTNB1_MOUSE
|
||||||
| NC score | 0.275209 (rank : 15) | θ value | 0.00175202 (rank : 20) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q02248, Q922W1, Q9D335 | Gene names | Ctnnb1, Catnb | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
CTNB1_HUMAN
|
||||||
| NC score | 0.275142 (rank : 16) | θ value | 0.00175202 (rank : 19) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P35222, Q8NEW9, Q8NI94, Q9H391 | Gene names | CTNNB1, CTNNB | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
PLAK_MOUSE
|
||||||
| NC score | 0.246601 (rank : 17) | θ value | 0.0193708 (rank : 27) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q02257, Q8CGD3 | Gene names | Jup | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
APC_MOUSE
|
||||||
| NC score | 0.227933 (rank : 18) | θ value | 0.000602161 (rank : 17) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
APC_HUMAN
|
||||||
| NC score | 0.221281 (rank : 19) | θ value | 0.000786445 (rank : 18) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
PLAK_HUMAN
|
||||||
| NC score | 0.209943 (rank : 20) | θ value | 0.0736092 (rank : 31) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P14923, Q9HCX9 | Gene names | JUP, DP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
IMA7_MOUSE
|
||||||
| NC score | 0.188309 (rank : 21) | θ value | 0.000121331 (rank : 15) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O35345, Q9CVP9 | Gene names | Kpna6, Kpna5 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6 subunit) (Importin alpha S2). | |||||
|
IMA7_HUMAN
|
||||||
| NC score | 0.184223 (rank : 22) | θ value | 0.00035302 (rank : 16) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
IMA5_HUMAN
|
||||||
| NC score | 0.177495 (rank : 23) | θ value | 0.0148317 (rank : 26) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O15131 | Gene names | KPNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-6 subunit (Karyopherin alpha-5 subunit). | |||||
|
GDS1_HUMAN
|
||||||
| NC score | 0.176464 (rank : 24) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P52306, Q9NYM2, Q9NZA8 | Gene names | RAP1GDS1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-GDP dissociation stimulator 1 (SMG P21 stimulatory GDP/GTP exchange protein) (SMG GDS protein) (Exchange factor smgGDS). | |||||
|
IMA2_MOUSE
|
||||||
| NC score | 0.170658 (rank : 25) | θ value | 0.00175202 (rank : 22) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52293, Q64292 | Gene names | Kpna2, Rch1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1) (Pendulin) (Pore targeting complex 58 kDa subunit) (PTAC58) (Importin alpha P1). | |||||
|
IMA2_HUMAN
|
||||||
| NC score | 0.169700 (rank : 26) | θ value | 0.00175202 (rank : 21) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
IMA4_MOUSE
|
||||||
| NC score | 0.162253 (rank : 27) | θ value | 0.00509761 (rank : 24) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O35343 | Gene names | Kpna4 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Importin alpha Q1). | |||||
|
IMA4_HUMAN
|
||||||
| NC score | 0.162241 (rank : 28) | θ value | 0.00509761 (rank : 23) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O00629, O00190 | Gene names | KPNA4, QIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Qip1 protein). | |||||
|
IMA3_MOUSE
|
||||||
| NC score | 0.154986 (rank : 29) | θ value | 0.0961366 (rank : 36) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O35344 | Gene names | Kpna3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (Importin alpha Q2). | |||||
|
IMA3_HUMAN
|
||||||
| NC score | 0.154850 (rank : 30) | θ value | 0.0961366 (rank : 35) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00505, O00191, O43195, Q96AA7 | Gene names | KPNA3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (SRP1-gamma). | |||||
|
IMA1_HUMAN
|
||||||
| NC score | 0.147508 (rank : 31) | θ value | 0.0961366 (rank : 33) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52294, Q9BQ56 | Gene names | KPNA1, RCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (NPI-1). | |||||
|
IMA1_MOUSE
|
||||||
| NC score | 0.147237 (rank : 32) | θ value | 0.0961366 (rank : 34) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q60960, Q3TF32 | Gene names | Kpna1, Rch2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (Importin alpha S1). | |||||
|
HEAT3_MOUSE
|
||||||
| NC score | 0.080227 (rank : 33) | θ value | 0.47712 (rank : 47) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BQM4, Q8C731 | Gene names | Heatr3 | |||
|
Domain Architecture |

|
|||||
| Description | HEAT repeat-containing protein 3. | |||||
|
SPAG6_HUMAN
|
||||||
| NC score | 0.070126 (rank : 34) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O75602, Q5VUX5, Q5VUX6, Q5VUX7, Q6FI74, Q8NHQ6 | Gene names | SPAG6, PF16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Sperm flagellar protein) (Repro-SA-1). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.063690 (rank : 35) | θ value | 0.365318 (rank : 42) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
ARMX3_MOUSE
|
||||||
| NC score | 0.063463 (rank : 36) | θ value | 0.21417 (rank : 39) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BHS6, Q91VP8, Q9DC32 | Gene names | Armcx3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 3. | |||||
|
ARMX3_HUMAN
|
||||||
| NC score | 0.063452 (rank : 37) | θ value | 0.21417 (rank : 38) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UH62, Q53HC6, Q7LCF5, Q9NPE4 | Gene names | ARMCX3, ALEX3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 3 (Protein ALEX3) (ARM protein lost in epithelial cancers on chromosome X 3). | |||||
|
SPAG6_MOUSE
|
||||||
| NC score | 0.063158 (rank : 38) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JLI7, Q8K461 | Gene names | Spag6, Pf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Axoneme central apparatus protein). | |||||
|
CK024_HUMAN
|
||||||
| NC score | 0.056176 (rank : 39) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96F05, Q9H2K4 | Gene names | C11orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 precursor (Protein DM4E3). | |||||
|
PARM1_MOUSE
|
||||||
| NC score | 0.054268 (rank : 40) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q923D3, Q3TTV0, Q3UFU4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PARM-1 precursor. | |||||
|
MUC13_MOUSE
|
||||||
| NC score | 0.053837 (rank : 41) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
ARMC6_MOUSE
|
||||||
| NC score | 0.053356 (rank : 42) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BNU0, Q8C4P5, Q8C7S1, Q8K2S7, Q99JQ2, Q9CWE4 | Gene names | Armc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 6. | |||||
|
HEAT3_HUMAN
|
||||||
| NC score | 0.053229 (rank : 43) | θ value | 5.27518 (rank : 90) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7Z4Q2, Q8N525, Q8WV56, Q96CC9, Q9NWN7 | Gene names | HEATR3 | |||
|
Domain Architecture |

|
|||||
| Description | HEAT repeat-containing protein 3. | |||||
|
MAVS_HUMAN
|
||||||
| NC score | 0.051596 (rank : 44) | θ value | 0.0563607 (rank : 30) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7Z434, Q3I0Y2, Q5T7I6, Q86VY7, Q9H1H3, Q9H4Y1, Q9H8D3, Q9ULE9 | Gene names | MAVS, IPS1, KIAA1271, VISA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif) (Putative NF-kappa-B- activating protein 031N). | |||||
|
RTDR1_HUMAN
|
||||||
| NC score | 0.050655 (rank : 45) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UHP6 | Gene names | RTDR1 | |||
|
Domain Architecture |

|
|||||
| Description | Rhabdoid tumor deletion region protein 1. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.050616 (rank : 46) | θ value | 4.03905 (rank : 85) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
EPCR_HUMAN
|
||||||
| NC score | 0.048496 (rank : 47) | θ value | 0.279714 (rank : 41) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UNN8, Q14218, Q6IB56, Q96CB3, Q9ULX1 | Gene names | PROCR, EPCR | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial protein C receptor precursor (Endothelial cell protein C receptor) (Activated protein C receptor) (APC receptor) (CD201 antigen). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.048421 (rank : 48) | θ value | 0.0330416 (rank : 28) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
ARMX1_HUMAN
|
||||||
| NC score | 0.048308 (rank : 49) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9P291, Q53HK2, Q9H2Q0 | Gene names | ARMCX1, ALEX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 1 (Protein ALEX1) (ARM protein lost in epithelial cancers on chromosome X 1). | |||||
|
SMN_HUMAN
|
||||||
| NC score | 0.047338 (rank : 50) | θ value | 0.21417 (rank : 40) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16637, Q13119, Q96J51 | Gene names | SMN1, SMN, SMNT | |||
|
Domain Architecture |

|
|||||
| Description | Survival motor neuron protein (Component of gems 1) (Gemin-1). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.045527 (rank : 51) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
ARMX1_MOUSE
|
||||||
| NC score | 0.043127 (rank : 52) | θ value | 3.0926 (rank : 75) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9CX83 | Gene names | Armcx1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 1. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.041581 (rank : 53) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
DOT1L_HUMAN
|
||||||
| NC score | 0.041521 (rank : 54) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
NFRKB_MOUSE
|
||||||
| NC score | 0.040203 (rank : 55) | θ value | 0.365318 (rank : 43) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
HIF1N_MOUSE
|
||||||
| NC score | 0.033571 (rank : 56) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BLR9, Q3U3G4 | Gene names | Hif1an | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha inhibitor (EC 1.14.11.16) (Hypoxia- inducible factor asparagine hydroxylase). | |||||
|
TAF9_HUMAN
|
||||||
| NC score | 0.032167 (rank : 57) | θ value | 0.365318 (rank : 45) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q16594, Q9BTS1 | Gene names | TAF9, TAF2G, TAFII31 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 9 (Transcription initiation factor TFIID 31 kDa subunit) (TAFII-31) (TAFII-32) (TAFII32) (STAF31/32). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.031128 (rank : 58) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
K1718_HUMAN
|
||||||
| NC score | 0.030796 (rank : 59) | θ value | 0.0113563 (rank : 25) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZMT4, Q6MZL8, Q9C0E5 | Gene names | KIAA1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
AKAP2_MOUSE
|
||||||
| NC score | 0.030083 (rank : 60) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
PRC1_HUMAN
|
||||||
| NC score | 0.029986 (rank : 61) | θ value | 0.813845 (rank : 53) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43663, Q9BSB6 | Gene names | PRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein regulator of cytokinesis 1. | |||||
|
PDXL2_MOUSE
|
||||||
| NC score | 0.029220 (rank : 62) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CAE9, Q8CFW3 | Gene names | Podxl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
MAML1_MOUSE
|
||||||
| NC score | 0.027306 (rank : 63) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6T264, Q505D8, Q5SUC2, Q6PDK3, Q6ZQG5, Q8BIU5, Q8R3T0 | Gene names | Maml1, Kiaa0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.026953 (rank : 64) | θ value | 0.62314 (rank : 48) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
TAF9_MOUSE
|
||||||
| NC score | 0.026093 (rank : 65) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VI33, Q32P09, Q80XS3, Q9CV61 | Gene names | Taf9, Taf2g | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 9 (Transcription initiation factor TFIID 31 kDa subunit) (TAFII-31) (TAFII-32) (TAFII32). | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.025564 (rank : 66) | θ value | 0.125558 (rank : 37) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.025344 (rank : 67) | θ value | 1.38821 (rank : 57) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.025235 (rank : 68) | θ value | 4.03905 (rank : 79) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
GAB1_MOUSE
|
||||||
| NC score | 0.024656 (rank : 69) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QYY0, Q91VW7 | Gene names | Gab1 | |||
|
Domain Architecture |

|
|||||
| Description | GRB2-associated-binding protein 1 (GRB2-associated binder 1). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.023708 (rank : 70) | θ value | 0.62314 (rank : 51) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
DAG1_HUMAN
|
||||||
| NC score | 0.023680 (rank : 71) | θ value | 6.88961 (rank : 101) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14118 | Gene names | DAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Dystroglycan precursor (Dystrophin-associated glycoprotein 1) [Contains: Alpha-dystroglycan (Alpha-DG); Beta-dystroglycan (Beta- DG)]. | |||||
|
NCOR1_MOUSE
|
||||||
| NC score | 0.022486 (rank : 72) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
COE1_HUMAN
|
||||||
| NC score | 0.022402 (rank : 73) | θ value | 1.06291 (rank : 55) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UH73, Q8IW11 | Gene names | EBF1, COE1, EBF | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE1 (OE-1) (O/E-1) (Early B-cell factor). | |||||
|
COE1_MOUSE
|
||||||
| NC score | 0.022402 (rank : 74) | θ value | 1.06291 (rank : 56) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q07802 | Gene names | Ebf1, Coe1, Ebf | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE1 (OE-1) (O/E-1) (Early B-cell factor). | |||||
|
BAP1_HUMAN
|
||||||
| NC score | 0.022243 (rank : 75) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92560, Q6LEM0, Q7Z5E8 | Gene names | BAP1, KIAA0272 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase BAP1 (EC 3.4.19.12) (BRCA1- associated protein 1) (Cerebral protein 6). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.021578 (rank : 76) | θ value | 0.813845 (rank : 54) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
TRI66_MOUSE
|
||||||
| NC score | 0.019764 (rank : 77) | θ value | 0.365318 (rank : 46) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
CCD27_MOUSE
|
||||||
| NC score | 0.019394 (rank : 78) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3V036 | Gene names | Ccdc27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 27. | |||||
|
TAB3_HUMAN
|
||||||
| NC score | 0.018361 (rank : 79) | θ value | 5.27518 (rank : 95) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N5C8, Q6VQR0 | Gene names | MAP3K7IP3, TAB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 3 (TAK1-binding protein 3) (NF-kappa-B-activating protein 1). | |||||
|
CHD9_MOUSE
|
||||||
| NC score | 0.017799 (rank : 80) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.017455 (rank : 81) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
SFR12_MOUSE
|
||||||
| NC score | 0.017277 (rank : 82) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
TAB3_MOUSE
|
||||||
| NC score | 0.017143 (rank : 83) | θ value | 5.27518 (rank : 96) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q571K4 | Gene names | Map3k7ip3, Kiaa4135, Tab3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 3 (TAK1-binding protein 3). | |||||
|
ZMYM1_HUMAN
|
||||||
| NC score | 0.016802 (rank : 84) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5SVZ6, Q7Z3Q4 | Gene names | ZMYM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger MYM-type protein 1. | |||||
|
RIMB1_MOUSE
|
||||||
| NC score | 0.016352 (rank : 85) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
ZCH10_HUMAN
|
||||||
| NC score | 0.016124 (rank : 86) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TBK6, Q9NXR4 | Gene names | ZCCHC10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 10. | |||||
|
SFR12_HUMAN
|
||||||
| NC score | 0.015551 (rank : 87) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
AOC2_MOUSE
|
||||||
| NC score | 0.015384 (rank : 88) | θ value | 0.62314 (rank : 49) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q812C9, Q80WP3 | Gene names | Aoc2 | |||
|
Domain Architecture |

|
|||||
| Description | Retina-specific copper amine oxidase precursor (EC 1.4.3.6) (RAO) (Amine oxidase [copper-containing]). | |||||
|
MAML3_HUMAN
|
||||||
| NC score | 0.014811 (rank : 89) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96JK9 | Gene names | MAML3, KIAA1816 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 3 (Mam-3). | |||||
|
TNR8_HUMAN
|
||||||
| NC score | 0.014762 (rank : 90) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P28908 | Gene names | TNFRSF8, CD30 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 8 precursor (CD30L receptor) (Lymphocyte activation antigen CD30) (KI-1 antigen). | |||||
|
PHF2_MOUSE
|
||||||
| NC score | 0.014729 (rank : 91) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
ATX1_MOUSE
|
||||||
| NC score | 0.014050 (rank : 92) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P54254 | Gene names | Atxn1, Sca1 | |||
|
Domain Architecture |

|
|||||
| Description | Ataxin-1 (Spinocerebellar ataxia type 1 protein homolog). | |||||
|
SPTN5_HUMAN
|
||||||
| NC score | 0.013884 (rank : 93) | θ value | 0.365318 (rank : 44) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NRC6 | Gene names | SPTBN5, SPTBN4 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 4 (Spectrin, non-erythroid beta chain 4) (Beta-V spectrin) (BSPECV). | |||||
|
TLN2_MOUSE
|
||||||
| NC score | 0.013783 (rank : 94) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q71LX4, Q8BWK0 | Gene names | Tln2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Talin-2. | |||||
|
CO002_HUMAN
|
||||||
| NC score | 0.013672 (rank : 95) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
MUCDL_MOUSE
|
||||||
| NC score | 0.013641 (rank : 96) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8VHF2, Q8CEJ3, Q9D8I9 | Gene names | Mucdhl | |||
|
Domain Architecture |

|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
TR150_HUMAN
|
||||||
| NC score | 0.012680 (rank : 97) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2W1, Q5VTK6 | Gene names | THRAP3, TRAP150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
MTMR7_HUMAN
|
||||||
| NC score | 0.012309 (rank : 98) | θ value | 5.27518 (rank : 91) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y216, Q68DX4 | Gene names | MTMR7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
SNP29_HUMAN
|
||||||
| NC score | 0.012092 (rank : 99) | θ value | 5.27518 (rank : 93) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95721 | Gene names | SNAP29 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptosomal-associated protein 29 (SNAP-29) (Vesicle-membrane fusion protein SNAP-29) (Soluble 29 kDa NSF attachment protein). | |||||
|
HCFC1_MOUSE
|
||||||
| NC score | 0.011727 (rank : 100) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q61191, Q684R1, Q7TSB0, Q8C2D0, Q9QWH2 | Gene names | Hcfc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N- terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C-terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
ZFAN3_MOUSE
|
||||||
| NC score | 0.010454 (rank : 101) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q497H0, Q8K230, Q8R2V7, Q8R3V1 | Gene names | Zfand3, Tex27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AN1-type zinc finger protein 3 (Testis-expressed sequence 27). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.010264 (rank : 102) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
PDE1C_MOUSE
|
||||||
| NC score | 0.009690 (rank : 103) | θ value | 0.62314 (rank : 52) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64338, Q62045, Q8BSV6 | Gene names | Pde1c | |||
|
Domain Architecture |

|
|||||
| Description | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C (EC 3.1.4.17) (Cam-PDE 1C). | |||||
|
SYNJ1_MOUSE
|
||||||
| NC score | 0.009490 (rank : 104) | θ value | 5.27518 (rank : 94) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
CHD9_HUMAN
|
||||||
| NC score | 0.008893 (rank : 105) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
ZCH14_MOUSE
|
||||||
| NC score | 0.008063 (rank : 106) | θ value | 8.99809 (rank : 128) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VIG0, Q80TX3, Q91VU4 | Gene names | Zcchc14, Kiaa0579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 14 (BDG-29). | |||||
|
UBP47_MOUSE
|
||||||
| NC score | 0.007901 (rank : 107) | θ value | 4.03905 (rank : 86) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BY87, Q5EBP2, Q6KAR9, Q80V06, Q8BHU1, Q8BI15, Q8BI16, Q8BUW4, Q91X25 | Gene names | Usp47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
RNF12_MOUSE
|
||||||
| NC score | 0.007839 (rank : 108) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WTV7 | Gene names | Rnf12, Rlim | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 12 (LIM domain-interacting RING finger protein) (RING finger LIM domain-binding protein) (R-LIM). | |||||
|
TLN2_HUMAN
|
||||||
| NC score | 0.007700 (rank : 109) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y4G6 | Gene names | TLN2, KIAA0320 | |||
|
Domain Architecture |

|
|||||
| Description | Talin-2. | |||||
|
TENS1_HUMAN
|
||||||
| NC score | 0.007691 (rank : 110) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9HBL0 | Gene names | TNS1, TNS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-1. | |||||
|
COA2_HUMAN
|
||||||
| NC score | 0.007209 (rank : 111) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O00763, Q16852 | Gene names | ACACB, ACC2, ACCB | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 2 (EC 6.4.1.2) (ACC-beta) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
CDKL5_HUMAN
|
||||||
| NC score | 0.006928 (rank : 112) | θ value | 0.0431538 (rank : 29) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O76039, Q14198, Q5H985, Q9UJL6 | Gene names | CDKL5, STK9 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase-like 5 (EC 2.7.11.22) (Serine/threonine- protein kinase 9). | |||||
|
CCNL2_HUMAN
|
||||||
| NC score | 0.006913 (rank : 113) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96S94, Q5T2N5, Q5T2N6, Q6IQ12, Q7Z4Z8, Q8N3C9, Q8N3D5, Q8NHE3, Q8TEL0, Q96B00 | Gene names | CCNL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L2 (Paneth cell-enhanced expression protein). | |||||
|
UBP47_HUMAN
|
||||||
| NC score | 0.006483 (rank : 114) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96K76, Q658U0, Q86Y73, Q8TEP6, Q9BWI0, Q9NWN1 | Gene names | USP47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
SMAD2_MOUSE
|
||||||
| NC score | 0.005954 (rank : 115) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62432, Q9D8P6 | Gene names | Smad2, Madh2, Madr2 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 2 (SMAD 2) (Mothers against DPP homolog 2) (Mad-related protein 2) (mMad2). | |||||
|
SMAD2_HUMAN
|
||||||
| NC score | 0.005953 (rank : 116) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15796 | Gene names | SMAD2, MADH2, MADR2 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 2 (SMAD 2) (Mothers against DPP homolog 2) (Mad-related protein 2) (hMAD-2) (JV18-1) (hSMAD2). | |||||
|
MTMRD_HUMAN
|
||||||
| NC score | 0.005514 (rank : 117) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86WG5, Q3MJF0, Q68DQ3, Q6P459, Q6PJD1, Q7Z325, Q7Z621, Q86VE2, Q96FE2, Q9C097 | Gene names | SBF2, CMT4B2, KIAA1766, MTMR13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 13 (SET-binding factor 2). | |||||
|
ZCHC6_MOUSE
|
||||||
| NC score | 0.005370 (rank : 118) | θ value | 8.99809 (rank : 129) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5BLK4, Q8CIH3 | Gene names | Zcchc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 6. | |||||
|
FLIP1_HUMAN
|
||||||
| NC score | 0.004594 (rank : 119) | θ value | 0.62314 (rank : 50) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
FA5_HUMAN
|
||||||
| NC score | 0.004417 (rank : 120) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P12259, Q14285, Q6UPU6 | Gene names | F5 | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor V precursor (Activated protein C cofactor) [Contains: Coagulation factor V heavy chain; Coagulation factor V light chain]. | |||||
|
BMP2K_MOUSE
|
||||||
| NC score | 0.003849 (rank : 121) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q91Z96, Q8C8L7 | Gene names | Bmp2k, Bike | |||
|
Domain Architecture |

|
|||||
| Description | BMP-2-inducible protein kinase (EC 2.7.11.1) (BIKe). | |||||
|
WNK1_MOUSE
|
||||||
| NC score | 0.003077 (rank : 122) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1179 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P83741 | Gene names | Wnk1, Prkwnk1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1). | |||||
|
GP125_HUMAN
|
||||||
| NC score | 0.003027 (rank : 123) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IWK6, Q6UXK9, Q86SQ5, Q8TC55 | Gene names | GPR125 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 125 precursor. | |||||
|
NPHN_MOUSE
|
||||||
| NC score | 0.002785 (rank : 124) | θ value | 5.27518 (rank : 92) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QZS7 | Gene names | Nphs1, Nphn | |||
|
Domain Architecture |

|
|||||
| Description | Nephrin precursor (Renal glomerulus-specific cell adhesion receptor). | |||||
|
ANK3_HUMAN
|
||||||
| NC score | 0.002664 (rank : 125) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 853 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q12955 | Gene names | ANK3 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-3 (ANK-3) (Ankyrin-G). | |||||
|
KLF12_HUMAN
|
||||||
| NC score | 0.001879 (rank : 126) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 717 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y4X4, Q5VZM7, Q9UHZ0 | Gene names | KLF12, AP2REP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 12 (Transcriptional repressor AP-2rep). | |||||
|
PO2F3_MOUSE
|
||||||
| NC score | 0.001791 (rank : 127) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P31362 | Gene names | Pou2f3, Epoc1, Oct11, Otf-11, Otf11 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 2, transcription factor 3 (Octamer-binding transcription factor 11) (Oct-11) (Epoc-1). | |||||
|
SLK_MOUSE
|
||||||
| NC score | 0.001771 (rank : 128) | θ value | 0.0736092 (rank : 32) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1798 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O54988, Q80U65, Q8CAU2, Q8CDW2, Q9WU41 | Gene names | Slk, Kiaa0204, Stk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | STE20-like serine/threonine-protein kinase (EC 2.7.11.1) (STE20-like kinase) (STE20-related serine/threonine-protein kinase) (STE20-related kinase) (mSLK) (Serine/threonine-protein kinase 2) (STE20-related kinase SMAK) (Etk4). | |||||
|
ZBTB3_HUMAN
|
||||||
| NC score | 0.001482 (rank : 129) | θ value | 8.99809 (rank : 127) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 910 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9H5J0 | Gene names | ZBTB3 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 3. | |||||
|
MYOTI_HUMAN
|
||||||
| NC score | 0.001388 (rank : 130) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UBF9 | Gene names | MYOT, TTID | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotilin (Titin immunoglobulin domain protein) (Myofibrillar titin- like Ig domains protein) (57 kDa cytoskeletal protein). | |||||
|
PAX3_HUMAN
|
||||||
| NC score | 0.000347 (rank : 131) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P23760, Q16448 | Gene names | PAX3, HUP2 | |||
|
Domain Architecture |

|
|||||
| Description | Paired box protein Pax-3 (HUP2). | |||||
|
EGR1_MOUSE
|
||||||
| NC score | 0.000119 (rank : 132) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P08046, Q61777 | Gene names | Egr1, Egr-1, Krox-24 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 1 (EGR-1) (Krox-24 protein) (Transcription factor Zif268) (Nerve growth factor-induced protein A) (NGFI-A). | |||||
|
WT1_HUMAN
|
||||||
| NC score | -0.000074 (rank : 133) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P19544, Q15881, Q16256, Q16575, Q8IYZ5 | Gene names | WT1 | |||
|
Domain Architecture |

|
|||||
| Description | Wilms' tumor protein (WT33). | |||||
|
ABL2_HUMAN
|
||||||
| NC score | -0.001502 (rank : 134) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 1045 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P42684, Q5T0X6, Q6NZY6 | Gene names | ABL2, ABLL, ARG | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase ABL2 (EC 2.7.10.2) (Abelson murine leukemia viral oncogene homolog 2) (Tyrosine kinase ARG). | |||||
|
ZN692_MOUSE
|
||||||
| NC score | -0.001563 (rank : 135) | θ value | 4.03905 (rank : 87) | |||
| Query Neighborhood Hits | 129 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3U381 | Gene names | Znf692, Zfp692 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 692. | |||||