Please be patient as the page loads
|
PHF22_MOUSE
|
||||||
| SwissProt Accessions | Q9D168, Q3U2Q5, Q921U2 | Gene names | Ints12, Phf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PHF22_MOUSE
|
||||||
| θ value | 1.08667e-154 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | Q9D168, Q3U2Q5, Q921U2 | Gene names | Ints12, Phf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
PHF22_HUMAN
|
||||||
| θ value | 6.17048e-150 (rank : 2) | NC score | 0.962905 (rank : 2) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96CB8, Q9HD71 | Gene names | INTS12, PHF22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
PHF1_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 3) | NC score | 0.420123 (rank : 3) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O43189, O60929, Q96KM7 | Gene names | PHF1 | |||
|
Domain Architecture |
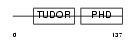
|
|||||
| Description | PHD finger protein 1 (Protein PHF1). | |||||
|
PHF1_MOUSE
|
||||||
| θ value | 1.99992e-07 (rank : 4) | NC score | 0.418163 (rank : 4) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Z1B8, O54808 | Gene names | Phf1, Tctex-3, Tctex3 | |||
|
Domain Architecture |
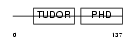
|
|||||
| Description | PHD finger protein 1 (Protein PHF1) (T-complex testis-expressed 3) (Polycomblike 1) (mPCl1). | |||||
|
MTF2_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 5) | NC score | 0.353631 (rank : 5) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9Y483, Q9UES9, Q9UP40 | Gene names | MTF2 | |||
|
Domain Architecture |
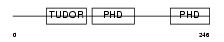
|
|||||
| Description | Metal-response element-binding transcription factor 2 (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
MTF2_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 6) | NC score | 0.351982 (rank : 6) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q02395, Q569Z8, Q6PG89, Q8BGP9, Q8BSJ7 | Gene names | Mtf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-response element-binding transcription factor 2 (Zinc-regulated factor 1) (ZiRF1) (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
PF21A_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 7) | NC score | 0.276305 (rank : 7) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
PF21A_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 8) | NC score | 0.269651 (rank : 8) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 9) | NC score | 0.200988 (rank : 13) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
SP110_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 10) | NC score | 0.236402 (rank : 9) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9HB58, Q14976, Q14977, Q8WUZ6, Q9HCT8 | Gene names | SP110 | |||
|
Domain Architecture |

|
|||||
| Description | Sp110 nuclear body protein (Speckled 110 kDa) (Transcriptional coactivator Sp110) (Interferon-induced protein 41/75). | |||||
|
TF3B_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 11) | NC score | 0.122704 (rank : 42) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFK2 | Gene names | Brf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor IIIB 90 kDa subunit (TFIIIB90) (hTFIIIB90) (B- related factor 1) (BRF-1). | |||||
|
TIF1A_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 12) | NC score | 0.195062 (rank : 16) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 13) | NC score | 0.164084 (rank : 23) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
PHF12_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 14) | NC score | 0.211075 (rank : 11) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
RAI1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 15) | NC score | 0.122977 (rank : 41) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q7Z5J4, Q8N3B4, Q8ND08, Q8WU64, Q96JK5, Q9H1C1, Q9H1C2, Q9UF69 | Gene names | RAI1, KIAA1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
TIF1A_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 16) | NC score | 0.198612 (rank : 14) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
LY10_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 17) | NC score | 0.217712 (rank : 10) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 18) | NC score | 0.153981 (rank : 27) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
DPF1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 19) | NC score | 0.155112 (rank : 26) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9QX66, Q9QX65, Q9QYA4 | Gene names | Dpf1, Neud4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
PHF12_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 20) | NC score | 0.208312 (rank : 12) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
TBX2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 21) | NC score | 0.027826 (rank : 121) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13207, Q16424, Q7Z647 | Gene names | TBX2 | |||
|
Domain Architecture |

|
|||||
| Description | T-box transcription factor TBX2 (T-box protein 2). | |||||
|
TIF1B_MOUSE
|
||||||
| θ value | 0.21417 (rank : 22) | NC score | 0.195157 (rank : 15) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q62318, P70391, Q8C283, Q99PN4 | Gene names | Trim28, Krip1, Tif1b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (KRAB-A-interacting protein) (KRIP-1). | |||||
|
KAISO_MOUSE
|
||||||
| θ value | 0.365318 (rank : 23) | NC score | 0.012749 (rank : 133) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BN78, Q8C226, Q9WTZ7 | Gene names | Zbtb33, Kaiso | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional regulator Kaiso (Zinc finger and BTB domain-containing protein 33). | |||||
|
RPGR_MOUSE
|
||||||
| θ value | 0.365318 (rank : 24) | NC score | 0.048608 (rank : 109) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
IF3X_HUMAN
|
||||||
| θ value | 0.47712 (rank : 25) | NC score | 0.070013 (rank : 82) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75153, Q6AHY2, Q6P3X7, Q6ZUG8, Q8N4U7, Q9BTA3, Q9H979 | Gene names | KIAA0664 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative eukaryotic translation initiation factor 3 subunit (eIF-3). | |||||
|
TIF1B_HUMAN
|
||||||
| θ value | 0.47712 (rank : 26) | NC score | 0.191602 (rank : 17) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q13263, O00677, Q7Z632, Q93040, Q96IM1 | Gene names | TRIM28, KAP1, RNF96, TIF1B | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (Nuclear corepressor KAP-1) (KRAB- associated protein 1) (KAP-1) (KRAB-interacting protein 1) (KRIP-1) (RING finger protein 96). | |||||
|
TRI66_MOUSE
|
||||||
| θ value | 0.47712 (rank : 27) | NC score | 0.185704 (rank : 18) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
ZN583_MOUSE
|
||||||
| θ value | 0.47712 (rank : 28) | NC score | 0.004677 (rank : 136) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3V080, Q3V3L4 | Gene names | Znf583, Zfp583 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 583. | |||||
|
CO002_HUMAN
|
||||||
| θ value | 0.62314 (rank : 29) | NC score | 0.057417 (rank : 95) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
JADE2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 30) | NC score | 0.144647 (rank : 34) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 31) | NC score | 0.163808 (rank : 24) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
JADE2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 32) | NC score | 0.150120 (rank : 30) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NQC1, Q6IE80, Q8TEK0, Q92513, Q96GQ6 | Gene names | PHF15, JADE2, KIAA0239 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 0.813845 (rank : 33) | NC score | 0.044233 (rank : 113) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
SIN3B_MOUSE
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.036868 (rank : 117) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62141, O54976, Q6A013, Q8VCB8, Q8VDZ5 | Gene names | Sin3b, Kiaa0700 | |||
|
Domain Architecture |
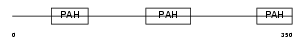
|
|||||
| Description | Paired amphipathic helix protein Sin3b (Transcriptional corepressor Sin3b) (Histone deacetylase complex subunit Sin3b). | |||||
|
TIF1G_HUMAN
|
||||||
| θ value | 0.813845 (rank : 35) | NC score | 0.170198 (rank : 21) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
TIF1G_MOUSE
|
||||||
| θ value | 0.813845 (rank : 36) | NC score | 0.168991 (rank : 22) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q99PP7, Q6SI71, Q6ZPX5 | Gene names | Trim33, Kiaa1113 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33). | |||||
|
UHRF1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 37) | NC score | 0.110627 (rank : 46) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8VDF2, Q8C6F1, Q8VIA1, Q9Z1H6 | Gene names | Uhrf1, Np95 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Nuclear zinc finger protein Np95) (Nuclear protein 95). | |||||
|
JADE3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 38) | NC score | 0.146200 (rank : 33) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q92613, Q6IE79 | Gene names | PHF16, JADE3, KIAA0215 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
JADE3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 39) | NC score | 0.146207 (rank : 32) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6IE82, Q6ZQG2 | Gene names | Phf16, Jade3, Kiaa0215 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
PF21B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.176130 (rank : 20) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PHF13_HUMAN
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.106550 (rank : 54) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86YI8, Q8N551, Q9UJP2 | Gene names | PHF13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
PHF13_MOUSE
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.106647 (rank : 53) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K2W6 | Gene names | Phf13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
CCNA2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 43) | NC score | 0.038520 (rank : 116) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20248 | Gene names | CCNA2, CCN1, CCNA | |||
|
Domain Architecture |
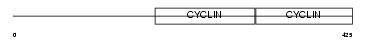
|
|||||
| Description | Cyclin-A2 (Cyclin-A). | |||||
|
DPF1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 44) | NC score | 0.149350 (rank : 31) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 45) | NC score | 0.140800 (rank : 35) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
PODXL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.047915 (rank : 110) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00592 | Gene names | PODXL, PCLP, PCLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Podocalyxin-like protein 1 precursor. | |||||
|
TRI66_HUMAN
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.176181 (rank : 19) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
VGFR1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 48) | NC score | 0.004766 (rank : 135) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P17948, O60722, P16057, Q12954 | Gene names | FLT1, FLT, FRT | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor receptor 1 precursor (EC 2.7.10.1) (VEGFR-1) (Vascular permeability factor receptor) (Tyrosine-protein kinase receptor FLT) (Flt-1) (Tyrosine-protein kinase FRT) (Fms-like tyrosine kinase 1). | |||||
|
ARI4A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.065693 (rank : 86) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
CIR_MOUSE
|
||||||
| θ value | 1.81305 (rank : 50) | NC score | 0.077056 (rank : 75) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
FXL19_HUMAN
|
||||||
| θ value | 1.81305 (rank : 51) | NC score | 0.065731 (rank : 85) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
JADE1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 52) | NC score | 0.139175 (rank : 36) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6IE81, Q4W5D5, Q6ZSL7, Q8NC41, Q96JL8, Q96SQ1, Q9H692 | Gene names | PHF17, JADE1, KIAA1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
JADE1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 53) | NC score | 0.138829 (rank : 37) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6ZPI0, Q6IE84, Q8C7S6, Q8CFM2, Q8R362 | Gene names | Phf17, Jade1, Kiaa1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
NCOA1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.020827 (rank : 123) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
ATRX_HUMAN
|
||||||
| θ value | 2.36792 (rank : 55) | NC score | 0.044252 (rank : 112) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
ATRX_MOUSE
|
||||||
| θ value | 2.36792 (rank : 56) | NC score | 0.043540 (rank : 114) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
BAZ2B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 57) | NC score | 0.110509 (rank : 47) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
BRPF1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 58) | NC score | 0.130330 (rank : 39) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
GON4L_HUMAN
|
||||||
| θ value | 2.36792 (rank : 59) | NC score | 0.028713 (rank : 120) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
JHD1B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 60) | NC score | 0.052264 (rank : 101) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
JHD1B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.051596 (rank : 103) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
LIMD1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.019820 (rank : 125) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UGP4, Q9BQQ9, Q9NQ47 | Gene names | LIMD1 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain-containing protein 1. | |||||
|
SPLC2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.039581 (rank : 115) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96DR5, Q9BQQ0 | Gene names | SPLUNC2, C20orf70 | |||
|
Domain Architecture |
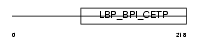
|
|||||
| Description | Short palate, lung and nasal epithelium carcinoma-associated protein 2 precursor (Parotid secretory protein) (PSP). | |||||
|
TAF3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.090411 (rank : 65) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
ZN583_HUMAN
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.002671 (rank : 137) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96ND8, O14850, Q2NKK3 | Gene names | ZNF583 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 583 (Zinc finger protein L3-5). | |||||
|
CA103_MOUSE
|
||||||
| θ value | 3.0926 (rank : 66) | NC score | 0.049887 (rank : 108) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CDD9, Q8C893 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C1orf103 homolog. | |||||
|
CN037_HUMAN
|
||||||
| θ value | 3.0926 (rank : 67) | NC score | 0.033489 (rank : 118) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86TY3, Q6P5Q1, Q86TY1 | Gene names | C14orf37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf37 precursor. | |||||
|
DMN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 68) | NC score | 0.016114 (rank : 129) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15061 | Gene names | DMN, KIAA0353 | |||
|
Domain Architecture |
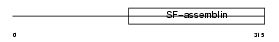
|
|||||
| Description | Desmuslin. | |||||
|
ITSN1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.010811 (rank : 134) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
REQU_HUMAN
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.136720 (rank : 38) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 523 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q92785 | Gene names | DPF2, REQ, UBID4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
YTHD2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.020666 (rank : 124) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y5A9, Q5VSZ9, Q8TDH0, Q9BUJ5 | Gene names | YTHDF2, HGRG8 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain family protein 2 (High-glucose-regulated protein 8) (NY- REN-2 antigen) (CLL-associated antigen KW-14). | |||||
|
DDX21_MOUSE
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.014004 (rank : 130) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.051638 (rank : 102) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.032224 (rank : 119) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
REQU_MOUSE
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.129832 (rank : 40) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q61103, Q3UNP5, Q60663, Q9QYA3 | Gene names | Dpf2, Req, Ubid4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
FXL19_MOUSE
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.056014 (rank : 97) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
HRH1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.001445 (rank : 138) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70174, Q91V75, Q91XN0, Q91XN1, Q91XN2, Q91XN3 | Gene names | Hrh1, Bphs | |||
|
Domain Architecture |
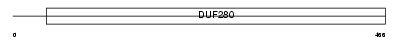
|
|||||
| Description | Histamine H1 receptor. | |||||
|
MAML2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.026902 (rank : 122) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IZL2, Q6AI23, Q6Y3A3, Q8IUL3, Q96JK6 | Gene names | MAML2, KIAA1819 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 2 (Mam-2). | |||||
|
MAP4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.045416 (rank : 111) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P27816, Q13082, Q96A76 | Gene names | MAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
MAST2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.001159 (rank : 139) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1189 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6P0Q8, O94899, Q5VT07, Q5VT08, Q7LGC4, Q8NDG1, Q96B94, Q9BYE8 | Gene names | MAST2, KIAA0807, MAST205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||
|
UTY_HUMAN
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.016828 (rank : 128) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O14607, O14608 | Gene names | UTY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed Y chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the Y chromosome). | |||||
|
B3GA2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.013891 (rank : 131) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NPZ5, Q5JS09, Q8TF38, Q96NK4 | Gene names | B3GAT2, GLCATS, KIAA1963 | |||
|
Domain Architecture |
|
|||||
| Description | Galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase 2 (EC 2.4.1.135) (Beta-1,3-glucuronyltransferase 2) (Glucuronosyltransferase-S) (GlcAT-S) (UDP-glucuronosyltransferase-S) (GlcAT-D). | |||||
|
BPTF_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.121333 (rank : 43) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
BRD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.115399 (rank : 44) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O95696 | Gene names | BRD1, BRL, BRPF2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 1 (BR140-like protein). | |||||
|
CDKL5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | -0.000153 (rank : 140) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O76039, Q14198, Q5H985, Q9UJL6 | Gene names | CDKL5, STK9 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase-like 5 (EC 2.7.11.22) (Serine/threonine- protein kinase 9). | |||||
|
CEP70_HUMAN
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.017147 (rank : 127) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NHQ1, Q96B31, Q9H2C3, Q9H2Z1, Q9H940 | Gene names | CEP70, BITE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 70 kDa (Cep70 protein) (p10-binding protein). | |||||
|
LPIN2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.013555 (rank : 132) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92539 | Gene names | LPIN2, KIAA0249 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipin-2. | |||||
|
TAF3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.080450 (rank : 70) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
SAFB2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.018345 (rank : 126) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14151, Q8TB13 | Gene names | SAFB2, KIAA0138 | |||
|
Domain Architecture |

|
|||||
| Description | Scaffold attachment factor B2. | |||||
|
AF10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.064563 (rank : 87) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P55197 | Gene names | MLLT10, AF10 | |||
|
Domain Architecture |
|
|||||
| Description | Protein AF-10. | |||||
|
AF10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.064503 (rank : 88) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O54826 | Gene names | Mllt10, Af10 | |||
|
Domain Architecture |
|
|||||
| Description | Protein AF-10. | |||||
|
AF17_HUMAN
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.066063 (rank : 84) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P55198, Q9UF49 | Gene names | MLLT6, AF17 | |||
|
Domain Architecture |
|
|||||
| Description | Protein AF-17. | |||||
|
AIRE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.161051 (rank : 25) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
AIRE_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.153180 (rank : 28) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
BAZ1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.099896 (rank : 57) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
BAZ1B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.092691 (rank : 63) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
BAZ1B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.097043 (rank : 60) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
BAZ2A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.107114 (rank : 51) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
BAZ2A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.104459 (rank : 55) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
BRPF3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.106934 (rank : 52) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||
|
CHD3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.076826 (rank : 77) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
CHD4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.080739 (rank : 69) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
CHD4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.079162 (rank : 73) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
CHD5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.080393 (rank : 72) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
CSR2B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.072620 (rank : 80) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H8E8, Q96GW6, Q96IH3, Q9HBF0, Q9UIY5 | Gene names | CSRP2BP | |||
|
Domain Architecture |
|
|||||
| Description | Cysteine-rich protein 2-binding protein (CSRP2-binding protein) (CRP2BP) (CRP2-binding partner). | |||||
|
CXCC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.058142 (rank : 94) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9P0U4, Q8N2W4, Q96BC8, Q9P2V7 | Gene names | CXXC1, CGBP, PCCX1, PHF18 | |||
|
Domain Architecture |
|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
CXCC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.056304 (rank : 96) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9CWW7 | Gene names | Cxxc1, Cgbp, Pccx1 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
IPR1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.053527 (rank : 99) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BVK9, Q3UCV7, Q80V00 | Gene names | Ipr1, Ifi75 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intracellular pathogen resistance protein 1. | |||||
|
JAD1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.077540 (rank : 74) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
JAD1C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.058520 (rank : 92) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
JAD1C_MOUSE
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.058853 (rank : 91) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P41230, O54995, Q9CVI4, Q9D0C3, Q9QVR8, Q9R039 | Gene names | Smcx, Jarid1c, Xe169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (SmcX protein) (Xe169 protein). | |||||
|
JAD1D_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.069289 (rank : 83) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9BY66, Q92509, Q92809, Q9HCU1 | Gene names | SMCY, HY, HYA, JARID1D, KIAA0234 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
JAD1D_MOUSE
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.062168 (rank : 90) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q62240, Q9QVR9, Q9R040 | Gene names | Smcy, Hya, Jarid1d | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
MYST3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.097930 (rank : 58) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
MYST3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.086855 (rank : 68) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
MYST4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.089803 (rank : 66) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
MYST4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.091355 (rank : 64) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
NSD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.095948 (rank : 62) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
NSD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.097365 (rank : 59) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
PF21B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.153094 (rank : 29) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PHF10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.111326 (rank : 45) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8WUB8, Q2YDA3, Q9BXD2, Q9NV26 | Gene names | PHF10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10 (XAP135). | |||||
|
PHF10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.109481 (rank : 48) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
PHF11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.063507 (rank : 89) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UIL8, Q9Y5A2 | Gene names | PHF11, BCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 11 (BRCA1 C-terminus-associated protein) (NY-REN-34 antigen). | |||||
|
PHF14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.108436 (rank : 49) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
PHF14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.107934 (rank : 50) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
PHF6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.053986 (rank : 98) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IWS0, Q96JK3, Q9BRU0 | Gene names | PHF6, KIAA1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6 (PHD-like zinc finger protein). | |||||
|
PHF6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.051078 (rank : 105) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D4J7, Q80T86, Q8BYT8, Q8C1B5 | Gene names | Phf6, Kiaa1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6. | |||||
|
PKCB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.096238 (rank : 61) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
RAI1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.076423 (rank : 79) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61818, Q6ZPH7, Q8CEV1 | Gene names | Rai1, Kiaa1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
RSF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.100241 (rank : 56) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
SET1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.058296 (rank : 93) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
TCF20_HUMAN
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.052919 (rank : 100) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UGU0, O14528, Q13078, Q4V353, Q9H4M0 | Gene names | TCF20, KIAA0292, SPBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP) (AR1). | |||||
|
TCF20_MOUSE
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.050406 (rank : 107) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
TRI45_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.051373 (rank : 104) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H8W5, Q5T2K4, Q5T2K5, Q8IYV6 | Gene names | TRIM45, RNF99 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 45 (RING finger protein 99). | |||||
|
TRI45_MOUSE
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.051039 (rank : 106) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6PFY8, Q8BVT5 | Gene names | Trim45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 45. | |||||
|
UHRF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.088331 (rank : 67) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96T88, Q8J022, Q9H6S6, Q9P115, Q9P1U7 | Gene names | UHRF1, ICBP90, NP95, RNF106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Inverted CCAAT box-binding protein of 90 kDa) (Transcription factor ICBP90) (Nuclear zinc finger protein Np95) (Nuclear protein 95) (HuNp95) (RING finger protein 106). | |||||
|
UHRF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.076956 (rank : 76) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96PU4, Q5VYR1, Q5VYR3, Q659C8, Q8TAG7 | Gene names | UHRF2, NIRF, RNF107 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95/ICBP90-like RING finger protein) (Np95-like RING finger protein) (Nuclear zinc finger protein Np97) (RING finger protein 107). | |||||
|
UHRF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.080421 (rank : 71) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q7TMI3, Q8BG56, Q8BJP6, Q8BY30, Q8K1I5 | Gene names | Uhrf2, Nirf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95-like ring finger protein) (Nuclear zinc finger protein Np97) (NIRF). | |||||
|
ZMY11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.071184 (rank : 81) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q15326 | Gene names | ZMYND11, BS69 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11 (Adenovirus 5 E1A- binding protein) (BS69 protein). | |||||
|
ZMY11_MOUSE
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.076512 (rank : 78) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8R5C8, Q8C155 | Gene names | Zmynd11 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11. | |||||
|
PHF22_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 1.08667e-154 (rank : 1) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | Q9D168, Q3U2Q5, Q921U2 | Gene names | Ints12, Phf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
PHF22_HUMAN
|
||||||
| NC score | 0.962905 (rank : 2) | θ value | 6.17048e-150 (rank : 2) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96CB8, Q9HD71 | Gene names | INTS12, PHF22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
PHF1_HUMAN
|
||||||
| NC score | 0.420123 (rank : 3) | θ value | 1.99992e-07 (rank : 3) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O43189, O60929, Q96KM7 | Gene names | PHF1 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 1 (Protein PHF1). | |||||
|
PHF1_MOUSE
|
||||||
| NC score | 0.418163 (rank : 4) | θ value | 1.99992e-07 (rank : 4) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Z1B8, O54808 | Gene names | Phf1, Tctex-3, Tctex3 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 1 (Protein PHF1) (T-complex testis-expressed 3) (Polycomblike 1) (mPCl1). | |||||
|
MTF2_HUMAN
|
||||||
| NC score | 0.353631 (rank : 5) | θ value | 0.00020696 (rank : 5) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9Y483, Q9UES9, Q9UP40 | Gene names | MTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Metal-response element-binding transcription factor 2 (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
MTF2_MOUSE
|
||||||
| NC score | 0.351982 (rank : 6) | θ value | 0.00020696 (rank : 6) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q02395, Q569Z8, Q6PG89, Q8BGP9, Q8BSJ7 | Gene names | Mtf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-response element-binding transcription factor 2 (Zinc-regulated factor 1) (ZiRF1) (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
PF21A_MOUSE
|
||||||
| NC score | 0.276305 (rank : 7) | θ value | 0.00020696 (rank : 7) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
PF21A_HUMAN
|
||||||
| NC score | 0.269651 (rank : 8) | θ value | 0.000270298 (rank : 8) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
SP110_HUMAN
|
||||||
| NC score | 0.236402 (rank : 9) | θ value | 0.00228821 (rank : 10) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9HB58, Q14976, Q14977, Q8WUZ6, Q9HCT8 | Gene names | SP110 | |||
|
Domain Architecture |

|
|||||
| Description | Sp110 nuclear body protein (Speckled 110 kDa) (Transcriptional coactivator Sp110) (Interferon-induced protein 41/75). | |||||
|
LY10_HUMAN
|
||||||
| NC score | 0.217712 (rank : 10) | θ value | 0.0736092 (rank : 17) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
PHF12_MOUSE
|
||||||
| NC score | 0.211075 (rank : 11) | θ value | 0.0431538 (rank : 14) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF12_HUMAN
|
||||||
| NC score | 0.208312 (rank : 12) | θ value | 0.0961366 (rank : 20) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.200988 (rank : 13) | θ value | 0.000786445 (rank : 9) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
TIF1A_MOUSE
|
||||||
| NC score | 0.198612 (rank : 14) | θ value | 0.0431538 (rank : 16) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
TIF1B_MOUSE
|
||||||
| NC score | 0.195157 (rank : 15) | θ value | 0.21417 (rank : 22) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q62318, P70391, Q8C283, Q99PN4 | Gene names | Trim28, Krip1, Tif1b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (KRAB-A-interacting protein) (KRIP-1). | |||||
|
TIF1A_HUMAN
|
||||||
| NC score | 0.195062 (rank : 16) | θ value | 0.0252991 (rank : 12) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
TIF1B_HUMAN
|
||||||
| NC score | 0.191602 (rank : 17) | θ value | 0.47712 (rank : 26) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q13263, O00677, Q7Z632, Q93040, Q96IM1 | Gene names | TRIM28, KAP1, RNF96, TIF1B | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (Nuclear corepressor KAP-1) (KRAB- associated protein 1) (KAP-1) (KRAB-interacting protein 1) (KRIP-1) (RING finger protein 96). | |||||
|
TRI66_MOUSE
|
||||||
| NC score | 0.185704 (rank : 18) | θ value | 0.47712 (rank : 27) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
TRI66_HUMAN
|
||||||
| NC score | 0.176181 (rank : 19) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
PF21B_HUMAN
|
||||||
| NC score | 0.176130 (rank : 20) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
TIF1G_HUMAN
|
||||||
| NC score | 0.170198 (rank : 21) | θ value | 0.813845 (rank : 35) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
TIF1G_MOUSE
|
||||||
| NC score | 0.168991 (rank : 22) | θ value | 0.813845 (rank : 36) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q99PP7, Q6SI71, Q6ZPX5 | Gene names | Trim33, Kiaa1113 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.164084 (rank : 23) | θ value | 0.0431538 (rank : 13) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.163808 (rank : 24) | θ value | 0.62314 (rank : 31) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
AIRE_HUMAN
|
||||||
| NC score | 0.161051 (rank : 25) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
DPF1_MOUSE
|
||||||
| NC score | 0.155112 (rank : 26) | θ value | 0.0961366 (rank : 19) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9QX66, Q9QX65, Q9QYA4 | Gene names | Dpf1, Neud4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.153981 (rank : 27) | θ value | 0.0736092 (rank : 18) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
AIRE_MOUSE
|
||||||
| NC score | 0.153180 (rank : 28) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
PF21B_MOUSE
|
||||||
| NC score | 0.153094 (rank : 29) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
JADE2_HUMAN
|
||||||
| NC score | 0.150120 (rank : 30) | θ value | 0.813845 (rank : 32) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NQC1, Q6IE80, Q8TEK0, Q92513, Q96GQ6 | Gene names | PHF15, JADE2, KIAA0239 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
DPF1_HUMAN
|
||||||
| NC score | 0.149350 (rank : 31) | θ value | 1.38821 (rank : 44) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
JADE3_MOUSE
|
||||||
| NC score | 0.146207 (rank : 32) | θ value | 1.06291 (rank : 39) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6IE82, Q6ZQG2 | Gene names | Phf16, Jade3, Kiaa0215 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
JADE3_HUMAN
|
||||||
| NC score | 0.146200 (rank : 33) | θ value | 1.06291 (rank : 38) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q92613, Q6IE79 | Gene names | PHF16, JADE3, KIAA0215 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
JADE2_MOUSE
|
||||||
| NC score | 0.144647 (rank : 34) | θ value | 0.62314 (rank : 30) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.140800 (rank : 35) | θ value | 1.38821 (rank : 45) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
JADE1_HUMAN
|
||||||
| NC score | 0.139175 (rank : 36) | θ value | 1.81305 (rank : 52) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6IE81, Q4W5D5, Q6ZSL7, Q8NC41, Q96JL8, Q96SQ1, Q9H692 | Gene names | PHF17, JADE1, KIAA1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
JADE1_MOUSE
|
||||||
| NC score | 0.138829 (rank : 37) | θ value | 1.81305 (rank : 53) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6ZPI0, Q6IE84, Q8C7S6, Q8CFM2, Q8R362 | Gene names | Phf17, Jade1, Kiaa1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
REQU_HUMAN
|
||||||
| NC score | 0.136720 (rank : 38) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 523 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q92785 | Gene names | DPF2, REQ, UBID4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
BRPF1_HUMAN
|
||||||
| NC score | 0.130330 (rank : 39) | θ value | 2.36792 (rank : 58) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
REQU_MOUSE
|
||||||
| NC score | 0.129832 (rank : 40) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q61103, Q3UNP5, Q60663, Q9QYA3 | Gene names | Dpf2, Req, Ubid4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
RAI1_HUMAN
|
||||||
| NC score | 0.122977 (rank : 41) | θ value | 0.0431538 (rank : 15) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q7Z5J4, Q8N3B4, Q8ND08, Q8WU64, Q96JK5, Q9H1C1, Q9H1C2, Q9UF69 | Gene names | RAI1, KIAA1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
TF3B_MOUSE
|
||||||
| NC score | 0.122704 (rank : 42) | θ value | 0.0148317 (rank : 11) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFK2 | Gene names | Brf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor IIIB 90 kDa subunit (TFIIIB90) (hTFIIIB90) (B- related factor 1) (BRF-1). | |||||
|
BPTF_HUMAN
|
||||||
| NC score | 0.121333 (rank : 43) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
BRD1_HUMAN
|
||||||
| NC score | 0.115399 (rank : 44) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O95696 | Gene names | BRD1, BRL, BRPF2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 1 (BR140-like protein). | |||||
|
PHF10_HUMAN
|
||||||
| NC score | 0.111326 (rank : 45) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8WUB8, Q2YDA3, Q9BXD2, Q9NV26 | Gene names | PHF10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10 (XAP135). | |||||
|
UHRF1_MOUSE
|
||||||
| NC score | 0.110627 (rank : 46) | θ value | 0.813845 (rank : 37) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8VDF2, Q8C6F1, Q8VIA1, Q9Z1H6 | Gene names | Uhrf1, Np95 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Nuclear zinc finger protein Np95) (Nuclear protein 95). | |||||
|
BAZ2B_HUMAN
|
||||||
| NC score | 0.110509 (rank : 47) | θ value | 2.36792 (rank : 57) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
PHF10_MOUSE
|
||||||
| NC score | 0.109481 (rank : 48) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
PHF14_HUMAN
|
||||||
| NC score | 0.108436 (rank : 49) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
PHF14_MOUSE
|
||||||
| NC score | 0.107934 (rank : 50) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
BAZ2A_HUMAN
|
||||||
| NC score | 0.107114 (rank : 51) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
BRPF3_HUMAN
|
||||||
| NC score | 0.106934 (rank : 52) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||
|
PHF13_MOUSE
|
||||||
| NC score | 0.106647 (rank : 53) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K2W6 | Gene names | Phf13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
PHF13_HUMAN
|
||||||
| NC score | 0.106550 (rank : 54) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86YI8, Q8N551, Q9UJP2 | Gene names | PHF13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
BAZ2A_MOUSE
|
||||||
| NC score | 0.104459 (rank : 55) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
RSF1_HUMAN
|
||||||
| NC score | 0.100241 (rank : 56) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
BAZ1A_HUMAN
|
||||||
| NC score | 0.099896 (rank : 57) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.097930 (rank : 58) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
NSD1_MOUSE
|
||||||
| NC score | 0.097365 (rank : 59) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
BAZ1B_MOUSE
|
||||||
| NC score | 0.097043 (rank : 60) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
PKCB1_HUMAN
|
||||||
| NC score | 0.096238 (rank : 61) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.095948 (rank : 62) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
BAZ1B_HUMAN
|
||||||
| NC score | 0.092691 (rank : 63) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
MYST4_MOUSE
|
||||||
| NC score | 0.091355 (rank : 64) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
TAF3_MOUSE
|
||||||
| NC score | 0.090411 (rank : 65) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
MYST4_HUMAN
|
||||||
| NC score | 0.089803 (rank : 66) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
UHRF1_HUMAN
|
||||||
| NC score | 0.088331 (rank : 67) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96T88, Q8J022, Q9H6S6, Q9P115, Q9P1U7 | Gene names | UHRF1, ICBP90, NP95, RNF106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Inverted CCAAT box-binding protein of 90 kDa) (Transcription factor ICBP90) (Nuclear zinc finger protein Np95) (Nuclear protein 95) (HuNp95) (RING finger protein 106). | |||||
|
MYST3_MOUSE
|
||||||
| NC score | 0.086855 (rank : 68) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
CHD4_HUMAN
|
||||||
| NC score | 0.080739 (rank : 69) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
TAF3_HUMAN
|
||||||
| NC score | 0.080450 (rank : 70) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
UHRF2_MOUSE
|
||||||
| NC score | 0.080421 (rank : 71) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q7TMI3, Q8BG56, Q8BJP6, Q8BY30, Q8K1I5 | Gene names | Uhrf2, Nirf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95-like ring finger protein) (Nuclear zinc finger protein Np97) (NIRF). | |||||
|
CHD5_HUMAN
|
||||||
| NC score | 0.080393 (rank : 72) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
CHD4_MOUSE
|
||||||
| NC score | 0.079162 (rank : 73) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
JAD1A_HUMAN
|
||||||
| NC score | 0.077540 (rank : 74) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
CIR_MOUSE
|
||||||
| NC score | 0.077056 (rank : 75) | θ value | 1.81305 (rank : 50) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
UHRF2_HUMAN
|
||||||
| NC score | 0.076956 (rank : 76) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96PU4, Q5VYR1, Q5VYR3, Q659C8, Q8TAG7 | Gene names | UHRF2, NIRF, RNF107 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95/ICBP90-like RING finger protein) (Np95-like RING finger protein) (Nuclear zinc finger protein Np97) (RING finger protein 107). | |||||
|
CHD3_HUMAN
|
||||||
| NC score | 0.076826 (rank : 77) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
ZMY11_MOUSE
|
||||||
| NC score | 0.076512 (rank : 78) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8R5C8, Q8C155 | Gene names | Zmynd11 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11. | |||||
|
RAI1_MOUSE
|
||||||
| NC score | 0.076423 (rank : 79) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61818, Q6ZPH7, Q8CEV1 | Gene names | Rai1, Kiaa1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
CSR2B_HUMAN
|
||||||
| NC score | 0.072620 (rank : 80) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H8E8, Q96GW6, Q96IH3, Q9HBF0, Q9UIY5 | Gene names | CSRP2BP | |||
|
Domain Architecture |

|
|||||
| Description | Cysteine-rich protein 2-binding protein (CSRP2-binding protein) (CRP2BP) (CRP2-binding partner). | |||||
|
ZMY11_HUMAN
|
||||||
| NC score | 0.071184 (rank : 81) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q15326 | Gene names | ZMYND11, BS69 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11 (Adenovirus 5 E1A- binding protein) (BS69 protein). | |||||
|
IF3X_HUMAN
|
||||||
| NC score | 0.070013 (rank : 82) | θ value | 0.47712 (rank : 25) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75153, Q6AHY2, Q6P3X7, Q6ZUG8, Q8N4U7, Q9BTA3, Q9H979 | Gene names | KIAA0664 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative eukaryotic translation initiation factor 3 subunit (eIF-3). | |||||
|
JAD1D_HUMAN
|
||||||
| NC score | 0.069289 (rank : 83) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9BY66, Q92509, Q92809, Q9HCU1 | Gene names | SMCY, HY, HYA, JARID1D, KIAA0234 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
AF17_HUMAN
|
||||||
| NC score | 0.066063 (rank : 84) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P55198, Q9UF49 | Gene names | MLLT6, AF17 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-17. | |||||
|
FXL19_HUMAN
|
||||||
| NC score | 0.065731 (rank : 85) | θ value | 1.81305 (rank : 51) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
ARI4A_HUMAN
|
||||||
| NC score | 0.065693 (rank : 86) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
AF10_HUMAN
|
||||||
| NC score | 0.064563 (rank : 87) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P55197 | Gene names | MLLT10, AF10 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-10. | |||||
|
AF10_MOUSE
|
||||||
| NC score | 0.064503 (rank : 88) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O54826 | Gene names | Mllt10, Af10 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-10. | |||||
|
PHF11_HUMAN
|
||||||
| NC score | 0.063507 (rank : 89) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UIL8, Q9Y5A2 | Gene names | PHF11, BCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 11 (BRCA1 C-terminus-associated protein) (NY-REN-34 antigen). | |||||
|
JAD1D_MOUSE
|
||||||
| NC score | 0.062168 (rank : 90) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q62240, Q9QVR9, Q9R040 | Gene names | Smcy, Hya, Jarid1d | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
JAD1C_MOUSE
|
||||||
| NC score | 0.058853 (rank : 91) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P41230, O54995, Q9CVI4, Q9D0C3, Q9QVR8, Q9R039 | Gene names | Smcx, Jarid1c, Xe169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (SmcX protein) (Xe169 protein). | |||||
|
JAD1C_HUMAN
|
||||||
| NC score | 0.058520 (rank : 92) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.058296 (rank : 93) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
CXCC1_HUMAN
|
||||||
| NC score | 0.058142 (rank : 94) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9P0U4, Q8N2W4, Q96BC8, Q9P2V7 | Gene names | CXXC1, CGBP, PCCX1, PHF18 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
CO002_HUMAN
|
||||||
| NC score | 0.057417 (rank : 95) | θ value | 0.62314 (rank : 29) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
CXCC1_MOUSE
|
||||||
| NC score | 0.056304 (rank : 96) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9CWW7 | Gene names | Cxxc1, Cgbp, Pccx1 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
FXL19_MOUSE
|
||||||
| NC score | 0.056014 (rank : 97) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
PHF6_HUMAN
|
||||||
| NC score | 0.053986 (rank : 98) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IWS0, Q96JK3, Q9BRU0 | Gene names | PHF6, KIAA1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6 (PHD-like zinc finger protein). | |||||
|
IPR1_MOUSE
|
||||||
| NC score | 0.053527 (rank : 99) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BVK9, Q3UCV7, Q80V00 | Gene names | Ipr1, Ifi75 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intracellular pathogen resistance protein 1. | |||||
|
TCF20_HUMAN
|
||||||
| NC score | 0.052919 (rank : 100) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UGU0, O14528, Q13078, Q4V353, Q9H4M0 | Gene names | TCF20, KIAA0292, SPBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP) (AR1). | |||||
|
JHD1B_HUMAN
|
||||||
| NC score | 0.052264 (rank : 101) | θ value | 2.36792 (rank : 60) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.051638 (rank : 102) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
JHD1B_MOUSE
|
||||||
| NC score | 0.051596 (rank : 103) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
TRI45_HUMAN
|
||||||
| NC score | 0.051373 (rank : 104) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H8W5, Q5T2K4, Q5T2K5, Q8IYV6 | Gene names | TRIM45, RNF99 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 45 (RING finger protein 99). | |||||
|
PHF6_MOUSE
|
||||||
| NC score | 0.051078 (rank : 105) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D4J7, Q80T86, Q8BYT8, Q8C1B5 | Gene names | Phf6, Kiaa1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6. | |||||
|
TRI45_MOUSE
|
||||||
| NC score | 0.051039 (rank : 106) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6PFY8, Q8BVT5 | Gene names | Trim45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 45. | |||||
|
TCF20_MOUSE
|
||||||
| NC score | 0.050406 (rank : 107) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
CA103_MOUSE
|
||||||
| NC score | 0.049887 (rank : 108) | θ value | 3.0926 (rank : 66) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CDD9, Q8C893 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C1orf103 homolog. | |||||
|
RPGR_MOUSE
|
||||||
| NC score | 0.048608 (rank : 109) | θ value | 0.365318 (rank : 24) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
PODXL_HUMAN
|
||||||
| NC score | 0.047915 (rank : 110) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00592 | Gene names | PODXL, PCLP, PCLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Podocalyxin-like protein 1 precursor. | |||||
|
MAP4_HUMAN
|
||||||
| NC score | 0.045416 (rank : 111) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P27816, Q13082, Q96A76 | Gene names | MAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
ATRX_HUMAN
|
||||||
| NC score | 0.044252 (rank : 112) | θ value | 2.36792 (rank : 55) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.044233 (rank : 113) | θ value | 0.813845 (rank : 33) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
ATRX_MOUSE
|
||||||
| NC score | 0.043540 (rank : 114) | θ value | 2.36792 (rank : 56) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
SPLC2_HUMAN
|
||||||
| NC score | 0.039581 (rank : 115) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96DR5, Q9BQQ0 | Gene names | SPLUNC2, C20orf70 | |||
|
Domain Architecture |

|
|||||
| Description | Short palate, lung and nasal epithelium carcinoma-associated protein 2 precursor (Parotid secretory protein) (PSP). | |||||
|
CCNA2_HUMAN
|
||||||
| NC score | 0.038520 (rank : 116) | θ value | 1.38821 (rank : 43) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20248 | Gene names | CCNA2, CCN1, CCNA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-A2 (Cyclin-A). | |||||
|
SIN3B_MOUSE
|
||||||
| NC score | 0.036868 (rank : 117) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62141, O54976, Q6A013, Q8VCB8, Q8VDZ5 | Gene names | Sin3b, Kiaa0700 | |||
|
Domain Architecture |

|
|||||
| Description | Paired amphipathic helix protein Sin3b (Transcriptional corepressor Sin3b) (Histone deacetylase complex subunit Sin3b). | |||||
|
CN037_HUMAN
|
||||||
| NC score | 0.033489 (rank : 118) | θ value | 3.0926 (rank : 67) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86TY3, Q6P5Q1, Q86TY1 | Gene names | C14orf37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf37 precursor. | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.032224 (rank : 119) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
GON4L_HUMAN
|
||||||
| NC score | 0.028713 (rank : 120) | θ value | 2.36792 (rank : 59) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
TBX2_HUMAN
|
||||||
| NC score | 0.027826 (rank : 121) | θ value | 0.0961366 (rank : 21) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13207, Q16424, Q7Z647 | Gene names | TBX2 | |||
|
Domain Architecture |

|
|||||
| Description | T-box transcription factor TBX2 (T-box protein 2). | |||||
|
MAML2_HUMAN
|
||||||
| NC score | 0.026902 (rank : 122) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IZL2, Q6AI23, Q6Y3A3, Q8IUL3, Q96JK6 | Gene names | MAML2, KIAA1819 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 2 (Mam-2). | |||||
|
NCOA1_HUMAN
|
||||||
| NC score | 0.020827 (rank : 123) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
YTHD2_HUMAN
|
||||||
| NC score | 0.020666 (rank : 124) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y5A9, Q5VSZ9, Q8TDH0, Q9BUJ5 | Gene names | YTHDF2, HGRG8 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain family protein 2 (High-glucose-regulated protein 8) (NY- REN-2 antigen) (CLL-associated antigen KW-14). | |||||
|
LIMD1_HUMAN
|
||||||
| NC score | 0.019820 (rank : 125) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UGP4, Q9BQQ9, Q9NQ47 | Gene names | LIMD1 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain-containing protein 1. | |||||
|
SAFB2_HUMAN
|
||||||
| NC score | 0.018345 (rank : 126) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14151, Q8TB13 | Gene names | SAFB2, KIAA0138 | |||
|
Domain Architecture |

|
|||||
| Description | Scaffold attachment factor B2. | |||||
|
CEP70_HUMAN
|
||||||
| NC score | 0.017147 (rank : 127) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NHQ1, Q96B31, Q9H2C3, Q9H2Z1, Q9H940 | Gene names | CEP70, BITE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 70 kDa (Cep70 protein) (p10-binding protein). | |||||
|
UTY_HUMAN
|
||||||
| NC score | 0.016828 (rank : 128) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O14607, O14608 | Gene names | UTY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed Y chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the Y chromosome). | |||||
|
DMN_HUMAN
|
||||||
| NC score | 0.016114 (rank : 129) | θ value | 3.0926 (rank : 68) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15061 | Gene names | DMN, KIAA0353 | |||
|
Domain Architecture |

|
|||||
| Description | Desmuslin. | |||||
|
DDX21_MOUSE
|
||||||
| NC score | 0.014004 (rank : 130) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
B3GA2_HUMAN
|
||||||
| NC score | 0.013891 (rank : 131) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NPZ5, Q5JS09, Q8TF38, Q96NK4 | Gene names | B3GAT2, GLCATS, KIAA1963 | |||
|
Domain Architecture |

|
|||||
| Description | Galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase 2 (EC 2.4.1.135) (Beta-1,3-glucuronyltransferase 2) (Glucuronosyltransferase-S) (GlcAT-S) (UDP-glucuronosyltransferase-S) (GlcAT-D). | |||||
|
LPIN2_HUMAN
|
||||||
| NC score | 0.013555 (rank : 132) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92539 | Gene names | LPIN2, KIAA0249 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipin-2. | |||||
|
KAISO_MOUSE
|
||||||
| NC score | 0.012749 (rank : 133) | θ value | 0.365318 (rank : 23) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BN78, Q8C226, Q9WTZ7 | Gene names | Zbtb33, Kaiso | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional regulator Kaiso (Zinc finger and BTB domain-containing protein 33). | |||||
|
ITSN1_MOUSE
|
||||||
| NC score | 0.010811 (rank : 134) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
VGFR1_HUMAN
|
||||||
| NC score | 0.004766 (rank : 135) | θ value | 1.38821 (rank : 48) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P17948, O60722, P16057, Q12954 | Gene names | FLT1, FLT, FRT | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor receptor 1 precursor (EC 2.7.10.1) (VEGFR-1) (Vascular permeability factor receptor) (Tyrosine-protein kinase receptor FLT) (Flt-1) (Tyrosine-protein kinase FRT) (Fms-like tyrosine kinase 1). | |||||
|
ZN583_MOUSE
|
||||||
| NC score | 0.004677 (rank : 136) | θ value | 0.47712 (rank : 28) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3V080, Q3V3L4 | Gene names | Znf583, Zfp583 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 583. | |||||
|
ZN583_HUMAN
|
||||||
| NC score | 0.002671 (rank : 137) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96ND8, O14850, Q2NKK3 | Gene names | ZNF583 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 583 (Zinc finger protein L3-5). | |||||
|
HRH1_MOUSE
|
||||||
| NC score | 0.001445 (rank : 138) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70174, Q91V75, Q91XN0, Q91XN1, Q91XN2, Q91XN3 | Gene names | Hrh1, Bphs | |||
|
Domain Architecture |

|
|||||
| Description | Histamine H1 receptor. | |||||
|
MAST2_HUMAN
|
||||||
| NC score | 0.001159 (rank : 139) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1189 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6P0Q8, O94899, Q5VT07, Q5VT08, Q7LGC4, Q8NDG1, Q96B94, Q9BYE8 | Gene names | MAST2, KIAA0807, MAST205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||
|
CDKL5_HUMAN
|
||||||
| NC score | -0.000153 (rank : 140) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O76039, Q14198, Q5H985, Q9UJL6 | Gene names | CDKL5, STK9 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase-like 5 (EC 2.7.11.22) (Serine/threonine- protein kinase 9). | |||||