Please be patient as the page loads
|
TF3B_MOUSE
|
||||||
| SwissProt Accessions | Q8CFK2 | Gene names | Brf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor IIIB 90 kDa subunit (TFIIIB90) (hTFIIIB90) (B- related factor 1) (BRF-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TF3B_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.981784 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92994, Q13223, Q5PR24, Q6IQ02, Q9HCW6, Q9HCW7, Q9HCW8 | Gene names | BRF1, BRF | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor IIIB 90 kDa subunit (TFIIIB90) (hTFIIIB90) (B- related factor 1) (BRF-1) (hBRF) (TATA box-binding protein-associated factor, RNA polymerase III, subunit 2) (TAF3B2). | |||||
|
TF3B_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8CFK2 | Gene names | Brf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor IIIB 90 kDa subunit (TFIIIB90) (hTFIIIB90) (B- related factor 1) (BRF-1). | |||||
|
TF2B_HUMAN
|
||||||
| θ value | 3.18553e-13 (rank : 3) | NC score | 0.523639 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00403, Q5JS30 | Gene names | GTF2B, TF2B, TFIIB | |||
|
Domain Architecture |
|
|||||
| Description | Transcription initiation factor IIB (General transcription factor TFIIB) (S300-II). | |||||
|
TF2B_MOUSE
|
||||||
| θ value | 3.18553e-13 (rank : 4) | NC score | 0.523664 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P62915, P29053 | Gene names | Gtf2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor IIB (General transcription factor TFIIB) (RNA polymerase II alpha initiation factor). | |||||
|
CP250_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 5) | NC score | 0.048171 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
PHF22_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 6) | NC score | 0.122704 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D168, Q3U2Q5, Q921U2 | Gene names | Ints12, Phf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
PHF22_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 7) | NC score | 0.117535 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96CB8, Q9HD71 | Gene names | INTS12, PHF22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
GD1L1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 8) | NC score | 0.109974 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VE33 | Gene names | Gdap1l1, Gdap1l | |||
|
Domain Architecture |
|
|||||
| Description | Ganglioside-induced differentiation-associated protein 1-like 1 (GDAP1-L1). | |||||
|
GD1L1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 9) | NC score | 0.107485 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96MZ0, Q5TE60, Q68CW7, Q9BQJ7, Q9BQV4, Q9BWJ4, Q9H3Y2, Q9H4G5 | Gene names | GDAP1L1 | |||
|
Domain Architecture |
|
|||||
| Description | Ganglioside-induced differentiation-associated protein 1-like 1 (GDAP1-L1). | |||||
|
BRD3_MOUSE
|
||||||
| θ value | 0.279714 (rank : 10) | NC score | 0.048558 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K2F0, Q8C665, Q8CAX7, Q9JI25 | Gene names | Brd3, Fsrg2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (Bromodomain-containing FSH-like protein FSRG2). | |||||
|
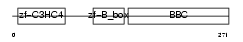
TRI55_HUMAN
|
||||||
| θ value | 0.279714 (rank : 11) | NC score | 0.039558 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BYV6, Q53XX3, Q8IUD9, Q8IUE4, Q96DV2, Q96DV3, Q9BYV5 | Gene names | TRIM55, MURF2, RNF29 | |||
|
Domain Architecture |
|
|||||
| Description | Tripartite motif-containing protein 55 (Muscle-specific RING finger protein 2) (MuRF2) (MURF-2) (RING finger protein 29). | |||||
|
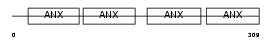
ANXA3_MOUSE
|
||||||
| θ value | 0.47712 (rank : 12) | NC score | 0.027453 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35639 | Gene names | Anxa3, Anx3 | |||
|
Domain Architecture |
|
|||||
| Description | Annexin A3 (Annexin III) (Lipocortin III) (Placental anticoagulant protein III) (PAP-III) (35-alpha calcimedin). | |||||
|
CUTL1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.043376 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
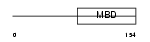
MECP2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 14) | NC score | 0.061000 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51608, O15233 | Gene names | MECP2 | |||
|
Domain Architecture |
|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
PHLB1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.050337 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q86UU1, O75133, Q4KMF8, Q8TEQ2 | Gene names | PHLDB1, DLNB07, KIAA0638, LL5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
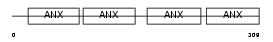
ANXA3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 16) | NC score | 0.026041 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P12429, Q6LET2 | Gene names | ANXA3, ANX3 | |||
|
Domain Architecture |
|
|||||
| Description | Annexin A3 (Annexin III) (Lipocortin III) (Placental anticoagulant protein III) (PAP-III) (35-alpha calcimedin) (Inositol 1,2-cyclic phosphate 2-phosphohydrolase). | |||||
|
INT1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 17) | NC score | 0.056795 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N201, Q6NT70, Q6UX74, Q8WV40, Q96D36, Q9NTD1, Q9P2A8, Q9Y3W8 | Gene names | INTS1, KIAA1440 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 1 (Int1). | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.040715 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
MECP2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 19) | NC score | 0.058095 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z2D6 | Gene names | Mecp2 | |||
|
Domain Architecture |
|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
INT1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 20) | NC score | 0.053881 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P4S8, Q80UQ7, Q91Z01, Q9CTF7 | Gene names | Ints1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 1 (Int1). | |||||
|
PHF20_HUMAN
|
||||||
| θ value | 1.38821 (rank : 21) | NC score | 0.041925 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 22) | NC score | 0.031928 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
TP53B_HUMAN
|
||||||
| θ value | 1.38821 (rank : 23) | NC score | 0.041589 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 413 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12888, Q2M1Z7, Q4LE46, Q5FWZ3, Q7Z3U4 | Gene names | TP53BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
CYTSA_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.034996 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1138 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q69YQ0, O15081 | Gene names | CYTSA, KIAA0376 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytospin-A (NY-REN-22 antigen). | |||||
|
CYTSA_MOUSE
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.034330 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1104 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q2KN98, Q8CHF9 | Gene names | Cytsa, Kiaa0376 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytospin-A. | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.035002 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.031338 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
MNT_MOUSE
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.042849 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
SAS6_MOUSE
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.026167 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80UK7, Q9CYT4 | Gene names | Sass6, Sas6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spindle assembly abnormal protein 6 homolog. | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.029541 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
TRIO_HUMAN
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.006310 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1580 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O75962, Q13458 | Gene names | TRIO | |||
|
Domain Architecture |

|
|||||
| Description | Triple functional domain protein (PTPRF-interacting protein). | |||||
|
CAC1C_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.009222 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q01815, Q04476, Q61242, Q99242 | Gene names | Cacna1c, Cach2, Cacn2, Cacnl1a1, Cchl1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle) (Mouse brain class C) (MBC) (MELC-CC). | |||||
|
CK035_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.035945 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IXW0 | Gene names | C11orf35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf35. | |||||
|
DYHC_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.034145 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 733 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q14204, Q6DKQ7, Q8WU28, Q92814, Q9Y4G5 | Gene names | DYNC1H1, DNCH1, DNECL, KIAA0325 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dynein heavy chain, cytosolic (DYHC) (Cytoplasmic dynein heavy chain 1) (DHC1) (Dynein heavy chain 1, cytoplasmic 1). | |||||
|
ERC2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.027733 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
HIRA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.013864 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P54198, Q8IXN2 | Gene names | HIRA, DGCR1, HIR, TUPLE1 | |||
|
Domain Architecture |
|
|||||
| Description | HIRA protein (TUP1-like enhancer of split protein 1). | |||||
|
I2C2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.024815 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKV8, Q8TCZ5, Q8WV58, Q96ID1 | Gene names | EIF2C2, AGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 2 (eIF2C 2) (eIF-2C 2) (Argonaute-2) (Slicer protein) (PAZ Piwi domain protein) (PPD). | |||||
|
I2C2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.024647 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CJG0, Q4VAB3, Q571J6 | Gene names | Eif2c2, Ago2, Kiaa4215 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 2 (eIF2C 2) (eIF-2C 2) (Argonaute-2) (Slicer protein) (Piwi/argonaute family protein meIF2C2). | |||||
|
CP26B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.017699 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NR63, Q53TW1, Q9NP41 | Gene names | CYP26B1, CYP26A2, P450RAI2 | |||
|
Domain Architecture |
|
|||||
| Description | Cytochrome P450 26B1 (EC 1.14.-.-) (P450 26A2) (P450 retinoic acid- inactivating 2) (P450RAI-2) (Retinoic acid-metabolizing cytochrome). | |||||
|
DIAP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.024484 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 839 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O08808 | Gene names | Diaph1, Diap1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1) (mDIA1) (p140mDIA). | |||||
|
DYHC_MOUSE
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.032911 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 729 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JHU4 | Gene names | Dync1h1, Dnch1, Dnchc1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynein heavy chain, cytosolic (DYHC) (Cytoplasmic dynein heavy chain 1) (DHC1) (Dynein heavy chain 1, cytoplasmic 1). | |||||
|
MRCKB_MOUSE
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.010288 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
TP53B_MOUSE
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.034059 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
CASP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.036644 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
DDX21_MOUSE
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.007256 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
ERC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.024189 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15083, Q86TK4 | Gene names | ERC2, KIAA0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2. | |||||
|
NCOA3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.012475 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
T53I2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.023720 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CFU8, Q711P5, Q8CHM4 | Gene names | Tp53inp2, Dor, Trp53inp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor protein p53-inducible nuclear protein 2 (Diabetes and obesity- regulated protein). | |||||
|
ZN746_MOUSE
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.001079 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3U133 | Gene names | Znf746, Zfp746 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 746. | |||||
|
BLNK_MOUSE
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.018142 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QUN3, O88504 | Gene names | Blnk, Bash, Ly57, Slp65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell linker protein (Cytoplasmic adapter protein) (B-cell adapter containing SH2 domain protein) (B-cell adapter containing Src homology 2 domain protein) (Src homology 2 domain-containing leukocyte protein of 65 kDa) (Slp-65) (Lymphocyte antigen 57). | |||||
|
BRD3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.025409 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15059, Q5T1R7, Q8N5M3, Q92645 | Gene names | BRD3, KIAA0043, RING3L | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (RING3-like protein). | |||||
|
CALD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 52) | NC score | 0.023641 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
DHX36_HUMAN
|
||||||
| θ value | 6.88961 (rank : 53) | NC score | 0.007177 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H2U1, Q70JU3, Q8IYE5, Q9P240 | Gene names | DHX36, DDX36, KIAA1488, MLEL1, RHAU | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DHX36 (EC 3.6.1.-) (DEAH box protein 36) (MLE-like protein 1) (RNA helicase associated with AU-rich element ARE). | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.023226 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
K1C10_HUMAN
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.007923 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P13645, Q14664, Q8N175 | Gene names | KRT10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10). | |||||
|
MK15_MOUSE
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.000808 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80Y86 | Gene names | Mapk15, Erk7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase 15 (EC 2.7.11.24) (Extracellular signal-regulated kinase 7). | |||||
|
MNT_HUMAN
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.032619 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
PHLB1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.039532 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 678 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PDH0, Q80TV2 | Gene names | Phldb1, Kiaa0638, Ll5a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
RIOK2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.017092 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CQS5, Q91XF3 | Gene names | Riok2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase RIO2 (EC 2.7.11.1) (RIO kinase 2). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.022140 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
UTP18_MOUSE
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.016308 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5SSI6 | Gene names | Utp18, Wdr50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 18 homolog (WD repeat protein 50). | |||||
|
WDR55_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.010180 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CX97 | Gene names | Wdr55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 55. | |||||
|
ARMET_HUMAN
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.017614 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P55145, Q86U67, Q96IS4 | Gene names | ARMET, ARP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ARMET protein precursor (Arginine-rich protein). | |||||
|
CASP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.032180 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1152 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P70403, Q7TQJ4, Q91WN6 | Gene names | Cutl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CCD91_HUMAN
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.020906 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 868 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7Z6B0, Q68D43, Q6IA78, Q8NEN7, Q9NUW9 | Gene names | CCDC91, GGABP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 91 (GGA-binding partner) (p56 accessory protein). | |||||
|
KLF10_MOUSE
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.000447 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 741 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O89091 | Gene names | Klf10, Gdnfif, Tieg, Tieg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 10 (Transforming growth factor-beta-inducible early growth response protein 1) (TGFB-inducible early growth response protein 1) (TIEG-1) (GDNF-inducible factor) (Transcription factor GIF) (mGIF). | |||||
|
PCNT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.020448 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
PDZD6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.018151 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULD6, Q4W5I8, Q86V55 | Gene names | PDZD6, KIAA1284, PDZK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 6. | |||||
|
TDRD3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.013607 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91W18, Q8BZW6 | Gene names | Tdrd3 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 3. | |||||
|
TF3B_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8CFK2 | Gene names | Brf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor IIIB 90 kDa subunit (TFIIIB90) (hTFIIIB90) (B- related factor 1) (BRF-1). | |||||
|
TF3B_HUMAN
|
||||||
| NC score | 0.981784 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92994, Q13223, Q5PR24, Q6IQ02, Q9HCW6, Q9HCW7, Q9HCW8 | Gene names | BRF1, BRF | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor IIIB 90 kDa subunit (TFIIIB90) (hTFIIIB90) (B- related factor 1) (BRF-1) (hBRF) (TATA box-binding protein-associated factor, RNA polymerase III, subunit 2) (TAF3B2). | |||||
|
TF2B_MOUSE
|
||||||
| NC score | 0.523664 (rank : 3) | θ value | 3.18553e-13 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P62915, P29053 | Gene names | Gtf2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor IIB (General transcription factor TFIIB) (RNA polymerase II alpha initiation factor). | |||||
|
TF2B_HUMAN
|
||||||
| NC score | 0.523639 (rank : 4) | θ value | 3.18553e-13 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00403, Q5JS30 | Gene names | GTF2B, TF2B, TFIIB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIB (General transcription factor TFIIB) (S300-II). | |||||
|
PHF22_MOUSE
|
||||||
| NC score | 0.122704 (rank : 5) | θ value | 0.0148317 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D168, Q3U2Q5, Q921U2 | Gene names | Ints12, Phf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
PHF22_HUMAN
|
||||||
| NC score | 0.117535 (rank : 6) | θ value | 0.0330416 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96CB8, Q9HD71 | Gene names | INTS12, PHF22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
GD1L1_MOUSE
|
||||||
| NC score | 0.109974 (rank : 7) | θ value | 0.0961366 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VE33 | Gene names | Gdap1l1, Gdap1l | |||
|
Domain Architecture |

|
|||||
| Description | Ganglioside-induced differentiation-associated protein 1-like 1 (GDAP1-L1). | |||||
|
GD1L1_HUMAN
|
||||||
| NC score | 0.107485 (rank : 8) | θ value | 0.125558 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96MZ0, Q5TE60, Q68CW7, Q9BQJ7, Q9BQV4, Q9BWJ4, Q9H3Y2, Q9H4G5 | Gene names | GDAP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Ganglioside-induced differentiation-associated protein 1-like 1 (GDAP1-L1). | |||||
|
MECP2_HUMAN
|
||||||
| NC score | 0.061000 (rank : 9) | θ value | 0.62314 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51608, O15233 | Gene names | MECP2 | |||
|
Domain Architecture |

|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
MECP2_MOUSE
|
||||||
| NC score | 0.058095 (rank : 10) | θ value | 1.06291 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z2D6 | Gene names | Mecp2 | |||
|
Domain Architecture |

|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
INT1_HUMAN
|
||||||
| NC score | 0.056795 (rank : 11) | θ value | 0.813845 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N201, Q6NT70, Q6UX74, Q8WV40, Q96D36, Q9NTD1, Q9P2A8, Q9Y3W8 | Gene names | INTS1, KIAA1440 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 1 (Int1). | |||||
|
INT1_MOUSE
|
||||||
| NC score | 0.053881 (rank : 12) | θ value | 1.38821 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P4S8, Q80UQ7, Q91Z01, Q9CTF7 | Gene names | Ints1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 1 (Int1). | |||||
|
PHLB1_HUMAN
|
||||||
| NC score | 0.050337 (rank : 13) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q86UU1, O75133, Q4KMF8, Q8TEQ2 | Gene names | PHLDB1, DLNB07, KIAA0638, LL5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
BRD3_MOUSE
|
||||||
| NC score | 0.048558 (rank : 14) | θ value | 0.279714 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K2F0, Q8C665, Q8CAX7, Q9JI25 | Gene names | Brd3, Fsrg2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (Bromodomain-containing FSH-like protein FSRG2). | |||||
|
CP250_MOUSE
|
||||||
| NC score | 0.048171 (rank : 15) | θ value | 0.0113563 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
CUTL1_HUMAN
|
||||||
| NC score | 0.043376 (rank : 16) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
MNT_MOUSE
|
||||||
| NC score | 0.042849 (rank : 17) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
PHF20_HUMAN
|
||||||
| NC score | 0.041925 (rank : 18) | θ value | 1.38821 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
TP53B_HUMAN
|
||||||
| NC score | 0.041589 (rank : 19) | θ value | 1.38821 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 413 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12888, Q2M1Z7, Q4LE46, Q5FWZ3, Q7Z3U4 | Gene names | TP53BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.040715 (rank : 20) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
TRI55_HUMAN
|
||||||
| NC score | 0.039558 (rank : 21) | θ value | 0.279714 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BYV6, Q53XX3, Q8IUD9, Q8IUE4, Q96DV2, Q96DV3, Q9BYV5 | Gene names | TRIM55, MURF2, RNF29 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 55 (Muscle-specific RING finger protein 2) (MuRF2) (MURF-2) (RING finger protein 29). | |||||
|
PHLB1_MOUSE
|
||||||
| NC score | 0.039532 (rank : 22) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 678 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PDH0, Q80TV2 | Gene names | Phldb1, Kiaa0638, Ll5a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
CASP_HUMAN
|
||||||
| NC score | 0.036644 (rank : 23) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CK035_HUMAN
|
||||||
| NC score | 0.035945 (rank : 24) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IXW0 | Gene names | C11orf35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf35. | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.035002 (rank : 25) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
CYTSA_HUMAN
|
||||||
| NC score | 0.034996 (rank : 26) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1138 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q69YQ0, O15081 | Gene names | CYTSA, KIAA0376 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytospin-A (NY-REN-22 antigen). | |||||
|
CYTSA_MOUSE
|
||||||
| NC score | 0.034330 (rank : 27) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1104 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q2KN98, Q8CHF9 | Gene names | Cytsa, Kiaa0376 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytospin-A. | |||||
|
DYHC_HUMAN
|
||||||
| NC score | 0.034145 (rank : 28) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 733 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q14204, Q6DKQ7, Q8WU28, Q92814, Q9Y4G5 | Gene names | DYNC1H1, DNCH1, DNECL, KIAA0325 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dynein heavy chain, cytosolic (DYHC) (Cytoplasmic dynein heavy chain 1) (DHC1) (Dynein heavy chain 1, cytoplasmic 1). | |||||
|
TP53B_MOUSE
|
||||||
| NC score | 0.034059 (rank : 29) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
DYHC_MOUSE
|
||||||
| NC score | 0.032911 (rank : 30) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 729 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JHU4 | Gene names | Dync1h1, Dnch1, Dnchc1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynein heavy chain, cytosolic (DYHC) (Cytoplasmic dynein heavy chain 1) (DHC1) (Dynein heavy chain 1, cytoplasmic 1). | |||||
|
MNT_HUMAN
|
||||||
| NC score | 0.032619 (rank : 31) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
CASP_MOUSE
|
||||||
| NC score | 0.032180 (rank : 32) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1152 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P70403, Q7TQJ4, Q91WN6 | Gene names | Cutl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.031928 (rank : 33) | θ value | 1.38821 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.031338 (rank : 34) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.029541 (rank : 35) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
ERC2_MOUSE
|
||||||
| NC score | 0.027733 (rank : 36) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
ANXA3_MOUSE
|
||||||
| NC score | 0.027453 (rank : 37) | θ value | 0.47712 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35639 | Gene names | Anxa3, Anx3 | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A3 (Annexin III) (Lipocortin III) (Placental anticoagulant protein III) (PAP-III) (35-alpha calcimedin). | |||||
|
SAS6_MOUSE
|
||||||
| NC score | 0.026167 (rank : 38) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80UK7, Q9CYT4 | Gene names | Sass6, Sas6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spindle assembly abnormal protein 6 homolog. | |||||
|
ANXA3_HUMAN
|
||||||
| NC score | 0.026041 (rank : 39) | θ value | 0.813845 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P12429, Q6LET2 | Gene names | ANXA3, ANX3 | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A3 (Annexin III) (Lipocortin III) (Placental anticoagulant protein III) (PAP-III) (35-alpha calcimedin) (Inositol 1,2-cyclic phosphate 2-phosphohydrolase). | |||||
|
BRD3_HUMAN
|
||||||
| NC score | 0.025409 (rank : 40) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15059, Q5T1R7, Q8N5M3, Q92645 | Gene names | BRD3, KIAA0043, RING3L | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (RING3-like protein). | |||||
|
I2C2_HUMAN
|
||||||
| NC score | 0.024815 (rank : 41) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKV8, Q8TCZ5, Q8WV58, Q96ID1 | Gene names | EIF2C2, AGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 2 (eIF2C 2) (eIF-2C 2) (Argonaute-2) (Slicer protein) (PAZ Piwi domain protein) (PPD). | |||||
|
I2C2_MOUSE
|
||||||
| NC score | 0.024647 (rank : 42) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CJG0, Q4VAB3, Q571J6 | Gene names | Eif2c2, Ago2, Kiaa4215 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 2 (eIF2C 2) (eIF-2C 2) (Argonaute-2) (Slicer protein) (Piwi/argonaute family protein meIF2C2). | |||||
|
DIAP1_MOUSE
|
||||||
| NC score | 0.024484 (rank : 43) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 839 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O08808 | Gene names | Diaph1, Diap1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1) (mDIA1) (p140mDIA). | |||||
|
ERC2_HUMAN
|
||||||
| NC score | 0.024189 (rank : 44) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15083, Q86TK4 | Gene names | ERC2, KIAA0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2. | |||||
|
T53I2_MOUSE
|
||||||
| NC score | 0.023720 (rank : 45) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CFU8, Q711P5, Q8CHM4 | Gene names | Tp53inp2, Dor, Trp53inp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor protein p53-inducible nuclear protein 2 (Diabetes and obesity- regulated protein). | |||||
|
CALD1_HUMAN
|
||||||
| NC score | 0.023641 (rank : 46) | θ value | 6.88961 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.023226 (rank : 47) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.022140 (rank : 48) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
CCD91_HUMAN
|
||||||
| NC score | 0.020906 (rank : 49) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 868 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7Z6B0, Q68D43, Q6IA78, Q8NEN7, Q9NUW9 | Gene names | CCDC91, GGABP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 91 (GGA-binding partner) (p56 accessory protein). | |||||
|
PCNT_HUMAN
|
||||||
| NC score | 0.020448 (rank : 50) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
PDZD6_HUMAN
|
||||||
| NC score | 0.018151 (rank : 51) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULD6, Q4W5I8, Q86V55 | Gene names | PDZD6, KIAA1284, PDZK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 6. | |||||
|
BLNK_MOUSE
|
||||||
| NC score | 0.018142 (rank : 52) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QUN3, O88504 | Gene names | Blnk, Bash, Ly57, Slp65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell linker protein (Cytoplasmic adapter protein) (B-cell adapter containing SH2 domain protein) (B-cell adapter containing Src homology 2 domain protein) (Src homology 2 domain-containing leukocyte protein of 65 kDa) (Slp-65) (Lymphocyte antigen 57). | |||||
|
CP26B_HUMAN
|
||||||
| NC score | 0.017699 (rank : 53) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NR63, Q53TW1, Q9NP41 | Gene names | CYP26B1, CYP26A2, P450RAI2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome P450 26B1 (EC 1.14.-.-) (P450 26A2) (P450 retinoic acid- inactivating 2) (P450RAI-2) (Retinoic acid-metabolizing cytochrome). | |||||
|
ARMET_HUMAN
|
||||||
| NC score | 0.017614 (rank : 54) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P55145, Q86U67, Q96IS4 | Gene names | ARMET, ARP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ARMET protein precursor (Arginine-rich protein). | |||||
|
RIOK2_MOUSE
|
||||||
| NC score | 0.017092 (rank : 55) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CQS5, Q91XF3 | Gene names | Riok2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase RIO2 (EC 2.7.11.1) (RIO kinase 2). | |||||
|
UTP18_MOUSE
|
||||||
| NC score | 0.016308 (rank : 56) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5SSI6 | Gene names | Utp18, Wdr50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 18 homolog (WD repeat protein 50). | |||||
|
HIRA_HUMAN
|
||||||
| NC score | 0.013864 (rank : 57) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P54198, Q8IXN2 | Gene names | HIRA, DGCR1, HIR, TUPLE1 | |||
|
Domain Architecture |

|
|||||
| Description | HIRA protein (TUP1-like enhancer of split protein 1). | |||||
|
TDRD3_MOUSE
|
||||||
| NC score | 0.013607 (rank : 58) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91W18, Q8BZW6 | Gene names | Tdrd3 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 3. | |||||
|
NCOA3_MOUSE
|
||||||
| NC score | 0.012475 (rank : 59) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
MRCKB_MOUSE
|
||||||
| NC score | 0.010288 (rank : 60) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
WDR55_MOUSE
|
||||||
| NC score | 0.010180 (rank : 61) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CX97 | Gene names | Wdr55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 55. | |||||
|
CAC1C_MOUSE
|
||||||
| NC score | 0.009222 (rank : 62) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q01815, Q04476, Q61242, Q99242 | Gene names | Cacna1c, Cach2, Cacn2, Cacnl1a1, Cchl1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle) (Mouse brain class C) (MBC) (MELC-CC). | |||||
|
K1C10_HUMAN
|
||||||
| NC score | 0.007923 (rank : 63) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P13645, Q14664, Q8N175 | Gene names | KRT10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10). | |||||
|
DDX21_MOUSE
|
||||||
| NC score | 0.007256 (rank : 64) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
DHX36_HUMAN
|
||||||
| NC score | 0.007177 (rank : 65) | θ value | 6.88961 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H2U1, Q70JU3, Q8IYE5, Q9P240 | Gene names | DHX36, DDX36, KIAA1488, MLEL1, RHAU | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DHX36 (EC 3.6.1.-) (DEAH box protein 36) (MLE-like protein 1) (RNA helicase associated with AU-rich element ARE). | |||||
|
TRIO_HUMAN
|
||||||
| NC score | 0.006310 (rank : 66) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1580 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O75962, Q13458 | Gene names | TRIO | |||
|
Domain Architecture |

|
|||||
| Description | Triple functional domain protein (PTPRF-interacting protein). | |||||
|
ZN746_MOUSE
|
||||||
| NC score | 0.001079 (rank : 67) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3U133 | Gene names | Znf746, Zfp746 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 746. | |||||
|
MK15_MOUSE
|
||||||
| NC score | 0.000808 (rank : 68) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80Y86 | Gene names | Mapk15, Erk7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase 15 (EC 2.7.11.24) (Extracellular signal-regulated kinase 7). | |||||
|
KLF10_MOUSE
|
||||||
| NC score | 0.000447 (rank : 69) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 741 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O89091 | Gene names | Klf10, Gdnfif, Tieg, Tieg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 10 (Transforming growth factor-beta-inducible early growth response protein 1) (TGFB-inducible early growth response protein 1) (TIEG-1) (GDNF-inducible factor) (Transcription factor GIF) (mGIF). | |||||