Please be patient as the page loads
|
NUMB_MOUSE
|
||||||
| SwissProt Accessions | Q9QZS3, P70422, Q8CIB1, Q9DC57, Q9QZR1, Q9QZS4 | Gene names | Numb | |||
|
Domain Architecture |
|
|||||
| Description | Protein numb homolog (m-Numb) (m-Nb). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
NUMB_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.983232 (rank : 2) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P49757, Q6NUQ7, Q86SY1, Q8WW73, Q9UBG1, Q9UEQ4, Q9UKE8, Q9UKE9, Q9UKF0, Q9UQJ4 | Gene names | NUMB | |||
|
Domain Architecture |
|
|||||
| Description | Protein numb homolog (h-Numb) (Protein S171). | |||||
|
NUMB_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9QZS3, P70422, Q8CIB1, Q9DC57, Q9QZR1, Q9QZS4 | Gene names | Numb | |||
|
Domain Architecture |

|
|||||
| Description | Protein numb homolog (m-Numb) (m-Nb). | |||||
|
NUMBL_HUMAN
|
||||||
| θ value | 7.28896e-151 (rank : 3) | NC score | 0.850304 (rank : 4) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y6R0, Q7Z4J9 | Gene names | NUMBL | |||
|
Domain Architecture |
|
|||||
| Description | Numb-like protein (Numb-R). | |||||
|
NUMBL_MOUSE
|
||||||
| θ value | 3.61748e-150 (rank : 4) | NC score | 0.945521 (rank : 3) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O08919, Q6NVG8 | Gene names | Numbl, Nbl | |||
|
Domain Architecture |
|
|||||
| Description | Numb-like protein. | |||||
|
ARH_HUMAN
|
||||||
| θ value | 1.58096e-12 (rank : 5) | NC score | 0.588644 (rank : 5) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5SW96, Q6TQS9, Q8N2Y0, Q9UFI9 | Gene names | LDLRAP1, ARH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Low density lipoprotein receptor adapter protein 1 (Autosomal recessive hypercholesterolemia protein). | |||||
|
ARH_MOUSE
|
||||||
| θ value | 1.33837e-11 (rank : 6) | NC score | 0.576983 (rank : 6) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8C142, Q6NWV6, Q8VDQ0 | Gene names | Ldlrap1, Arh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Low density lipoprotein receptor adapter protein 1 (Autosomal recessive hypercholesterolemia protein homolog). | |||||
|
DAB1_HUMAN
|
||||||
| θ value | 1.53129e-07 (rank : 7) | NC score | 0.389817 (rank : 8) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O75553, Q9NYA8 | Gene names | DAB1 | |||
|
Domain Architecture |
|
|||||
| Description | Disabled homolog 1. | |||||
|
DAB1_MOUSE
|
||||||
| θ value | 1.53129e-07 (rank : 8) | NC score | 0.399509 (rank : 7) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P97318, P97316, P97317 | Gene names | Dab1 | |||
|
Domain Architecture |
|
|||||
| Description | Disabled homolog 1. | |||||
|
ANKS1_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 9) | NC score | 0.159953 (rank : 18) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q92625, Q5JYI9, Q5SYR2, Q86WQ7 | Gene names | ANKS1A, ANKS1, KIAA0229 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 1A (Odin). | |||||
|
ANKS1_MOUSE
|
||||||
| θ value | 3.41135e-07 (rank : 10) | NC score | 0.167787 (rank : 17) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P59672, Q6ZQG0 | Gene names | Anks1a, Anks1, Kiaa0229 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 1A. | |||||
|
DAB2_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 11) | NC score | 0.339470 (rank : 11) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P98078, Q3U3K1, Q91W56, Q923E1 | Gene names | Dab2, Doc2 | |||
|
Domain Architecture |
|
|||||
| Description | Disabled homolog 2 (DOC-2) (Mitogen-responsive phosphoprotein). | |||||
|
DAB2_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 12) | NC score | 0.334008 (rank : 12) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P98082, Q13598, Q9BTY0, Q9UK04 | Gene names | DAB2, DOC2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (Differentially expressed protein 2) (DOC-2). | |||||
|
CAPON_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 13) | NC score | 0.344092 (rank : 10) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9D3A8, Q80TZ6 | Gene names | Nos1ap, Capon, Kiaa0464 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxyl-terminal PDZ ligand of neuronal nitric oxide synthase protein (C--terminal PDZ ligand of neuronal nitric oxide synthase protein) (Nitric oxide synthase 1 adaptor protein). | |||||
|
CAPON_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 14) | NC score | 0.355211 (rank : 9) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O75052, O43564 | Gene names | NOS1AP, CAPON, KIAA0464 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxyl-terminal PDZ ligand of neuronal nitric oxide synthase protein (C--terminal PDZ ligand of neuronal nitric oxide synthase protein) (Nitric oxide synthase 1 adaptor protein). | |||||
|
JIP1_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 15) | NC score | 0.212813 (rank : 13) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9WVI9, O35145, Q925J8, Q9R1H9, Q9R1Z1, Q9WVI7, Q9WVI8 | Gene names | Mapk8ip1, Ib1, Jip1, Mapk8ip, Prkm8ip | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 1 (JNK-interacting protein 1) (JIP-1) (JNK MAP kinase scaffold protein 1) (Islet-brain-1) (IB-1) (Mitogen-activated protein kinase 8-interacting protein 1). | |||||
|
JIP1_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 16) | NC score | 0.204153 (rank : 14) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UQF2, O43407 | Gene names | MAPK8IP1, IB1, JIP1 | |||
|
Domain Architecture |
|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 1 (JNK-interacting protein 1) (JIP-1) (JNK MAP kinase scaffold protein 1) (Islet-brain 1) (IB-1) (Mitogen-activated protein kinase 8-interacting protein 1). | |||||
|
NOTC2_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 17) | NC score | 0.032818 (rank : 75) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1069 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35516, Q06008, Q60941 | Gene names | Notch2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (Motch B) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
SHC2_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 18) | NC score | 0.111218 (rank : 20) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BMC3 | Gene names | Shc2, Sck, ShcB | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 2 (SH2 domain protein C2) (Src homology 2 domain-containing-transforming protein C2) (Protein Sck) (Protein Sli). | |||||
|
SHC2_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 19) | NC score | 0.108116 (rank : 21) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P98077, O60230, Q9NPL5, Q9UCX4 | Gene names | SHC2, SCK, SHCB | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 2 (SH2 domain protein C2) (Src homology 2 domain-containing-transforming protein C2) (Protein Sck). | |||||
|
APBB2_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 20) | NC score | 0.123184 (rank : 19) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92870, Q8IUI6 | Gene names | APBB2, FE65L, FE65L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2 (Fe65-like protein). | |||||
|
NOTC2_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 21) | NC score | 0.030286 (rank : 76) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q04721, Q99734, Q9H240 | Gene names | NOTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (hN2) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
SALL3_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 22) | NC score | 0.014944 (rank : 91) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1066 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9BXA9, Q9UGH1 | Gene names | SALL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 3 (Zinc finger protein SALL3) (hSALL3). | |||||
|
FA43A_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 23) | NC score | 0.201292 (rank : 15) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N2R8, Q8IXP4, Q8WZ07 | Gene names | FAM43A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM43A. | |||||
|
FA43A_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 24) | NC score | 0.201193 (rank : 16) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BUP8, Q8R231 | Gene names | Fam43a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM43A. | |||||
|
CD2L6_MOUSE
|
||||||
| θ value | 0.125558 (rank : 25) | NC score | 0.008927 (rank : 101) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BWD8, Q80TM1 | Gene names | Cdc2l6, Kiaa1028 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 6 (EC 2.7.11.22) (CDC2- related protein kinase 6). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 0.163984 (rank : 26) | NC score | 0.049853 (rank : 55) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
MAML2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 27) | NC score | 0.049626 (rank : 56) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IZL2, Q6AI23, Q6Y3A3, Q8IUL3, Q96JK6 | Gene names | MAML2, KIAA1819 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 2 (Mam-2). | |||||
|
ANX11_MOUSE
|
||||||
| θ value | 0.279714 (rank : 28) | NC score | 0.038546 (rank : 68) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97384 | Gene names | Anxa11, Anx11 | |||
|
Domain Architecture |
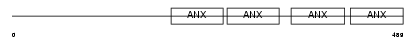
|
|||||
| Description | Annexin A11 (Annexin XI) (Calcyclin-associated annexin 50) (CAP-50). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 0.365318 (rank : 29) | NC score | 0.041859 (rank : 66) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
SN1L1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 30) | NC score | 0.007240 (rank : 105) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P57059, Q5R2V5, Q86YJ2 | Gene names | SNF1LK | |||
|
Domain Architecture |
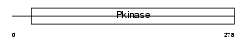
|
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 1 (EC 2.7.11.1) (Serine/threonine-protein kinase SNF1LK). | |||||
|
PER2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.049930 (rank : 54) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 0.62314 (rank : 32) | NC score | 0.054568 (rank : 41) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CJ012_HUMAN
|
||||||
| θ value | 0.62314 (rank : 33) | NC score | 0.055947 (rank : 38) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 34) | NC score | 0.037506 (rank : 72) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 35) | NC score | 0.057750 (rank : 35) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
ADA29_HUMAN
|
||||||
| θ value | 0.813845 (rank : 36) | NC score | 0.017366 (rank : 88) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UKF5, Q9UHP1, Q9UKF3, Q9UKF4 | Gene names | ADAM29 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 29 precursor (A disintegrin and metalloproteinase domain 29). | |||||
|
APBB1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 37) | NC score | 0.071167 (rank : 26) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QXJ1, O08642, Q3TPU0, Q8BNF4, Q8BSR9 | Gene names | Apbb1, Fe65 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1 (Fe65 protein). | |||||
|
ARI1B_HUMAN
|
||||||
| θ value | 0.813845 (rank : 38) | NC score | 0.044358 (rank : 64) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 0.813845 (rank : 39) | NC score | 0.051703 (rank : 50) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
FUBP2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.038464 (rank : 70) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92945, O00301, Q9UNT5, Q9UQH5 | Gene names | KHSRP, FUBP2 | |||
|
Domain Architecture |
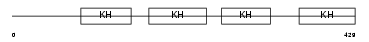
|
|||||
| Description | Far upstream element-binding protein 2 (FUSE-binding protein 2) (KH type-splicing regulatory protein) (KSRP) (p75). | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.043034 (rank : 65) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
APBB2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 42) | NC score | 0.098448 (rank : 22) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9DBR4, Q6DFX8 | Gene names | Apbb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2. | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 1.81305 (rank : 43) | NC score | 0.007769 (rank : 104) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
SEM6C_MOUSE
|
||||||
| θ value | 1.81305 (rank : 44) | NC score | 0.011094 (rank : 93) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
SON_HUMAN
|
||||||
| θ value | 1.81305 (rank : 45) | NC score | 0.044514 (rank : 63) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
APBB1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 46) | NC score | 0.065082 (rank : 30) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O00213, Q96A93 | Gene names | APBB1, FE65 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1 (Fe65 protein). | |||||
|
EMSY_MOUSE
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.030092 (rank : 77) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BMB0, Q5FWK5, Q80XU1, Q8VDW9 | Gene names | Emsy | |||
|
Domain Architecture |
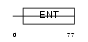
|
|||||
| Description | Protein EMSY. | |||||
|
MUC5B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.029515 (rank : 78) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
NU214_HUMAN
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.048514 (rank : 57) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.045580 (rank : 62) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 51) | NC score | 0.045964 (rank : 59) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
MYCL1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.016583 (rank : 90) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
NUFP1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.027409 (rank : 80) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UHK0, Q8WVM5, Q96SG1 | Gene names | NUFIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 1 (Nuclear FMRP-interacting protein 1). | |||||
|
RERE_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.045722 (rank : 61) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
BRD4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.033214 (rank : 73) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.048095 (rank : 58) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
EP400_MOUSE
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.017930 (rank : 87) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
MAML3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.039765 (rank : 67) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96JK9 | Gene names | MAML3, KIAA1816 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 3 (Mam-3). | |||||
|
MCM3A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.023125 (rank : 83) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60318, Q9UMT4 | Gene names | MCM3AP, GANP, KIAA0572, MAP80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
MYO15_MOUSE
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.009834 (rank : 97) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9QZZ4, O70395, Q9QWL6 | Gene names | Myo15a, Myo15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
ANXA7_MOUSE
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.020361 (rank : 85) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q07076 | Gene names | Anxa7, Anx7 | |||
|
Domain Architecture |
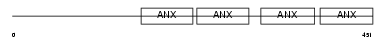
|
|||||
| Description | Annexin A7 (Annexin VII) (Synexin). | |||||
|
CD2L6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.005695 (rank : 110) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BWU1, Q5JQZ7, Q5JR00, Q8TC78, Q9UPX2 | Gene names | CDC2L6, CDK11, KIAA1028 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 6 (EC 2.7.11.22) (CDC2- related protein kinase 6) (Death-preventing kinase) (Cyclin-dependent kinase 11). | |||||
|
CNOT3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.028839 (rank : 79) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75175, Q9NZN7, Q9UF76 | Gene names | CNOT3, KIAA0691, NOT3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
DOCK7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.009016 (rank : 99) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96N67, Q6ZV32, Q8TB82, Q96NG6, Q9C092 | Gene names | DOCK7, KIAA1771 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dedicator of cytokinesis protein 7. | |||||
|
PRKN2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.019668 (rank : 86) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60260, Q5TFV8, Q6Q2I6, Q8NI41, Q8NI43, Q8NI44, Q8WW07 | Gene names | PARK2, PRKN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parkin (EC 6.3.2.-) (Ubiquitin E3 ligase PRKN) (Parkinson juvenile disease protein 2) (Parkinson disease protein 2). | |||||
|
SGIP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.038502 (rank : 69) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8VD37, Q3UFU3, Q3UGA0, Q8BXX4, Q8C034, Q9CXT2 | Gene names | Sgip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3-containing GRB2-like protein 3-interacting protein 1 (Endophilin- 3-interacting protein). | |||||
|
TM108_MOUSE
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.050640 (rank : 52) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BHE4, Q80WR9 | Gene names | Tmem108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 108 precursor. | |||||
|
WNK1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.007777 (rank : 103) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1158 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9H4A3, O15052, Q86WL5, Q8N673, Q9P1S9 | Gene names | WNK1, KDP, KIAA0344, PRKWNK1 | |||
|
Domain Architecture |
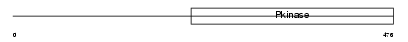
|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1) (Kinase deficient protein). | |||||
|
ANGL3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.004655 (rank : 112) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5C1 | Gene names | ANGPTL3, ANGPT5 | |||
|
Domain Architecture |
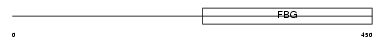
|
|||||
| Description | Angiopoietin-related protein 3 precursor (Angiopoietin-like 3) (Angiopoietin-5) (ANG-5). | |||||
|
ATX2L_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.023410 (rank : 82) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8WWM7, O95135, Q6NVJ8, Q6PJW6, Q8IU61, Q8IU95, Q8WWM3, Q8WWM4, Q8WWM5, Q8WWM6, Q99703 | Gene names | ATXN2L, A2D, A2LG, A2LP, A2RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein (Ataxin-2 domain protein) (Ataxin-2-related protein). | |||||
|
EPHAA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.002440 (rank : 115) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5JZY3, Q6NW42 | Gene names | EPHA10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ephrin type-A receptor 10 precursor (EC 2.7.10.1). | |||||
|
ITSN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.005910 (rank : 109) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
LYST_MOUSE
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.006639 (rank : 106) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97412, Q62403, Q8VBS6 | Gene names | Lyst, Bg, Chs1 | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal-trafficking regulator (Beige protein) (CHS1 homolog). | |||||
|
MEF2A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.010937 (rank : 94) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q02078, O43814, Q14223, Q14224, Q96D14 | Gene names | MEF2A, MEF2 | |||
|
Domain Architecture |
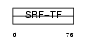
|
|||||
| Description | Myocyte-specific enhancer factor 2A (Serum response factor-like protein 1). | |||||
|
NEUM_MOUSE
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.021073 (rank : 84) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P06837 | Gene names | Gap43, Basp2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuromodulin (Axonal membrane protein GAP-43) (Growth-associated protein 43) (Calmodulin-binding protein P-57). | |||||
|
PI4KB_HUMAN
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.017307 (rank : 89) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBF8, O15096, P78405, Q9BWR6 | Gene names | PIK4CB, PI4KB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase beta (EC 2.7.1.67) (PtdIns 4-kinase beta) (PI4Kbeta) (PI4K-beta) (NPIK) (PI4K92). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.033084 (rank : 74) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SSH1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.006234 (rank : 107) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q76I79, Q3TDG3, Q69ZM4, Q811E5 | Gene names | Ssh1, Kiaa1298, Ssh1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 1 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-1L) (mSSH-1L). | |||||
|
ZEP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.003679 (rank : 114) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P31629, Q02646, Q5THT5, Q9NS05 | Gene names | HIVEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 (HIV- EP2) (MHC-binding protein 2) (MBP-2). | |||||
|
BAT3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.038189 (rank : 71) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Z1R2, Q8SNA3 | Gene names | Bat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3). | |||||
|
BNC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.005394 (rank : 111) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35914, O54886 | Gene names | Bnc1, Bnc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein basonuclin-1. | |||||
|
CCD45_MOUSE
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.008979 (rank : 100) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 663 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BVV7 | Gene names | Ccdc45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 45. | |||||
|
DEN1A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.012991 (rank : 92) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8K382 | Gene names | Dennd1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
GSCR1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.045938 (rank : 60) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
PGBM_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.008095 (rank : 102) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q05793 | Gene names | Hspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
RC3H1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.010527 (rank : 95) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
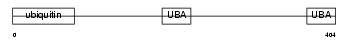
RD23B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.026927 (rank : 81) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P54728 | Gene names | Rad23b, Mhr23b | |||
|
Domain Architecture |
|
|||||
| Description | UV excision repair protein RAD23 homolog B (mHR23B) (XP-C repair- complementing complex 58 kDa protein) (p58). | |||||
|
RPGF2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.009124 (rank : 98) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y4G8 | Gene names | RAPGEF2, KIAA0313, NRAPGEP, PDZGEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rap guanine nucleotide exchange factor 2 (Neural RAP guanine nucleotide exchange protein) (nRap GEP) (PDZ domain-containing guanine nucleotide exchange factor 1) (PDZ-GEF1) (RA-GEF). | |||||
|
SKI_MOUSE
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.006008 (rank : 108) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q60698, Q8VIL5 | Gene names | Ski | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ski oncogene (C-ski). | |||||
|
WNK3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.003912 (rank : 113) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BYP7, Q9HCK6 | Gene names | WNK3, KIAA1566, PRKWNK3 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK3 (EC 2.7.11.1) (Protein kinase with no lysine 3) (Protein kinase, lysine-deficient 3). | |||||
|
XLKD1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.010150 (rank : 96) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5Y7, Q8TC18, Q9UNF4 | Gene names | XLKD1, CRSBP1, HAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lymphatic vessel endothelial hyaluronic acid receptor 1 precursor (LYVE-1) (Cell surface retention sequence-binding protein 1) (CRSBP-1) (Hyaluronic acid receptor) (Extracellular link domain-containing protein 1). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.050426 (rank : 53) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
CCNK_MOUSE
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.054257 (rank : 42) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
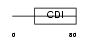
CDN1C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.081974 (rank : 24) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P49918 | Gene names | CDKN1C, KIP2 | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
CN032_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.066431 (rank : 28) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
CN032_MOUSE
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.063265 (rank : 33) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BH93, Q3TF12, Q3TGL0, Q8CC90 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32 homolog. | |||||
|
FA43B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.065828 (rank : 29) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZT52 | Gene names | FAM43B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM43B. | |||||
|
FMN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.051040 (rank : 51) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
FMN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.055969 (rank : 37) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JL04 | Gene names | Fmn2 | |||
|
Domain Architecture |

|
|||||
| Description | Formin-2. | |||||
|
GGN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.052437 (rank : 48) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
JIP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.052572 (rank : 47) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13387, Q96G62, Q99771, Q9NZ59, Q9UKQ4 | Gene names | MAPK8IP2, IB2, JIP2 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 2 (JNK-interacting protein 2) (JIP-2) (JNK MAP kinase scaffold protein 2) (Islet-brain-2) (IB-2) (Mitogen-activated protein kinase 8-interacting protein 2). | |||||
|
JIP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.053315 (rank : 45) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ERE9, Q9CXI4 | Gene names | Mapk8ip2, Ib2, Jip2 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 2 (JNK-interacting protein 2) (JIP-2) (JNK MAP kinase scaffold protein 2) (Islet-brain-2) (IB-2) (Mitogen-activated protein kinase 8-interacting protein 2). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.053963 (rank : 43) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.057322 (rank : 36) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
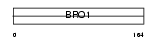
PDC6I_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.053757 (rank : 44) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9WU78, O88695, O89014, Q8BSL8, Q8R0H5, Q99LR3, Q9QZN8 | Gene names | Pdcd6ip, Aip1, Alix | |||
|
Domain Architecture |
|
|||||
| Description | Programmed cell death 6-interacting protein (ALG-2-interacting protein X) (ALG-2-interacting protein 1) (E2F1-inducible protein) (Eig2). | |||||
|
PRR12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.063665 (rank : 32) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
PRR13_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.055104 (rank : 40) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZ81, Q6FIG7, Q6MZP8, Q6NXQ6, Q6PKF9 | Gene names | PRR13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 13. | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.066657 (rank : 27) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
SELV_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.052617 (rank : 46) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P59797 | Gene names | SELV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Selenoprotein V. | |||||
|
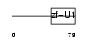
SF3A2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.060580 (rank : 34) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
SF3A2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.055461 (rank : 39) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |
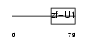
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
TAF4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.064638 (rank : 31) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
WASL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.052035 (rank : 49) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00401, Q7Z746 | Gene names | WASL | |||
|
Domain Architecture |
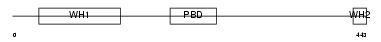
|
|||||
| Description | Neural Wiskott-Aldrich syndrome protein (N-WASP). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.078674 (rank : 25) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
ZN207_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.096401 (rank : 23) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
NUMB_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9QZS3, P70422, Q8CIB1, Q9DC57, Q9QZR1, Q9QZS4 | Gene names | Numb | |||
|
Domain Architecture |

|
|||||
| Description | Protein numb homolog (m-Numb) (m-Nb). | |||||
|
NUMB_HUMAN
|
||||||
| NC score | 0.983232 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P49757, Q6NUQ7, Q86SY1, Q8WW73, Q9UBG1, Q9UEQ4, Q9UKE8, Q9UKE9, Q9UKF0, Q9UQJ4 | Gene names | NUMB | |||
|
Domain Architecture |

|
|||||
| Description | Protein numb homolog (h-Numb) (Protein S171). | |||||
|
NUMBL_MOUSE
|
||||||
| NC score | 0.945521 (rank : 3) | θ value | 3.61748e-150 (rank : 4) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O08919, Q6NVG8 | Gene names | Numbl, Nbl | |||
|
Domain Architecture |

|
|||||
| Description | Numb-like protein. | |||||
|
NUMBL_HUMAN
|
||||||
| NC score | 0.850304 (rank : 4) | θ value | 7.28896e-151 (rank : 3) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y6R0, Q7Z4J9 | Gene names | NUMBL | |||
|
Domain Architecture |

|
|||||
| Description | Numb-like protein (Numb-R). | |||||
|
ARH_HUMAN
|
||||||
| NC score | 0.588644 (rank : 5) | θ value | 1.58096e-12 (rank : 5) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5SW96, Q6TQS9, Q8N2Y0, Q9UFI9 | Gene names | LDLRAP1, ARH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Low density lipoprotein receptor adapter protein 1 (Autosomal recessive hypercholesterolemia protein). | |||||
|
ARH_MOUSE
|
||||||
| NC score | 0.576983 (rank : 6) | θ value | 1.33837e-11 (rank : 6) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8C142, Q6NWV6, Q8VDQ0 | Gene names | Ldlrap1, Arh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Low density lipoprotein receptor adapter protein 1 (Autosomal recessive hypercholesterolemia protein homolog). | |||||
|
DAB1_MOUSE
|
||||||
| NC score | 0.399509 (rank : 7) | θ value | 1.53129e-07 (rank : 8) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P97318, P97316, P97317 | Gene names | Dab1 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 1. | |||||
|
DAB1_HUMAN
|
||||||
| NC score | 0.389817 (rank : 8) | θ value | 1.53129e-07 (rank : 7) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O75553, Q9NYA8 | Gene names | DAB1 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 1. | |||||
|
CAPON_HUMAN
|
||||||
| NC score | 0.355211 (rank : 9) | θ value | 0.000121331 (rank : 14) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O75052, O43564 | Gene names | NOS1AP, CAPON, KIAA0464 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxyl-terminal PDZ ligand of neuronal nitric oxide synthase protein (C--terminal PDZ ligand of neuronal nitric oxide synthase protein) (Nitric oxide synthase 1 adaptor protein). | |||||
|
CAPON_MOUSE
|
||||||
| NC score | 0.344092 (rank : 10) | θ value | 7.1131e-05 (rank : 13) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9D3A8, Q80TZ6 | Gene names | Nos1ap, Capon, Kiaa0464 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxyl-terminal PDZ ligand of neuronal nitric oxide synthase protein (C--terminal PDZ ligand of neuronal nitric oxide synthase protein) (Nitric oxide synthase 1 adaptor protein). | |||||
|
DAB2_MOUSE
|
||||||
| NC score | 0.339470 (rank : 11) | θ value | 4.1701e-05 (rank : 11) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P98078, Q3U3K1, Q91W56, Q923E1 | Gene names | Dab2, Doc2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (DOC-2) (Mitogen-responsive phosphoprotein). | |||||
|
DAB2_HUMAN
|
||||||
| NC score | 0.334008 (rank : 12) | θ value | 5.44631e-05 (rank : 12) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P98082, Q13598, Q9BTY0, Q9UK04 | Gene names | DAB2, DOC2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (Differentially expressed protein 2) (DOC-2). | |||||
|
JIP1_MOUSE
|
||||||
| NC score | 0.212813 (rank : 13) | θ value | 0.00298849 (rank : 15) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9WVI9, O35145, Q925J8, Q9R1H9, Q9R1Z1, Q9WVI7, Q9WVI8 | Gene names | Mapk8ip1, Ib1, Jip1, Mapk8ip, Prkm8ip | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 1 (JNK-interacting protein 1) (JIP-1) (JNK MAP kinase scaffold protein 1) (Islet-brain-1) (IB-1) (Mitogen-activated protein kinase 8-interacting protein 1). | |||||
|
JIP1_HUMAN
|
||||||
| NC score | 0.204153 (rank : 14) | θ value | 0.00665767 (rank : 16) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UQF2, O43407 | Gene names | MAPK8IP1, IB1, JIP1 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 1 (JNK-interacting protein 1) (JIP-1) (JNK MAP kinase scaffold protein 1) (Islet-brain 1) (IB-1) (Mitogen-activated protein kinase 8-interacting protein 1). | |||||
|
FA43A_HUMAN
|
||||||
| NC score | 0.201292 (rank : 15) | θ value | 0.0961366 (rank : 23) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N2R8, Q8IXP4, Q8WZ07 | Gene names | FAM43A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM43A. | |||||
|
FA43A_MOUSE
|
||||||
| NC score | 0.201193 (rank : 16) | θ value | 0.0961366 (rank : 24) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BUP8, Q8R231 | Gene names | Fam43a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM43A. | |||||
|
ANKS1_MOUSE
|
||||||
| NC score | 0.167787 (rank : 17) | θ value | 3.41135e-07 (rank : 10) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P59672, Q6ZQG0 | Gene names | Anks1a, Anks1, Kiaa0229 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 1A. | |||||
|
ANKS1_HUMAN
|
||||||
| NC score | 0.159953 (rank : 18) | θ value | 3.41135e-07 (rank : 9) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q92625, Q5JYI9, Q5SYR2, Q86WQ7 | Gene names | ANKS1A, ANKS1, KIAA0229 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 1A (Odin). | |||||
|
APBB2_HUMAN
|
||||||
| NC score | 0.123184 (rank : 19) | θ value | 0.0431538 (rank : 20) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92870, Q8IUI6 | Gene names | APBB2, FE65L, FE65L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2 (Fe65-like protein). | |||||
|
SHC2_MOUSE
|
||||||
| NC score | 0.111218 (rank : 20) | θ value | 0.00869519 (rank : 18) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BMC3 | Gene names | Shc2, Sck, ShcB | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 2 (SH2 domain protein C2) (Src homology 2 domain-containing-transforming protein C2) (Protein Sck) (Protein Sli). | |||||
|
SHC2_HUMAN
|
||||||
| NC score | 0.108116 (rank : 21) | θ value | 0.0113563 (rank : 19) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P98077, O60230, Q9NPL5, Q9UCX4 | Gene names | SHC2, SCK, SHCB | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 2 (SH2 domain protein C2) (Src homology 2 domain-containing-transforming protein C2) (Protein Sck). | |||||
|
APBB2_MOUSE
|
||||||
| NC score | 0.098448 (rank : 22) | θ value | 1.81305 (rank : 42) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9DBR4, Q6DFX8 | Gene names | Apbb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2. | |||||
|
ZN207_HUMAN
|
||||||
| NC score | 0.096401 (rank : 23) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
CDN1C_HUMAN
|
||||||
| NC score | 0.081974 (rank : 24) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P49918 | Gene names | CDKN1C, KIP2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.078674 (rank : 25) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
APBB1_MOUSE
|
||||||
| NC score | 0.071167 (rank : 26) | θ value | 0.813845 (rank : 37) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QXJ1, O08642, Q3TPU0, Q8BNF4, Q8BSR9 | Gene names | Apbb1, Fe65 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1 (Fe65 protein). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.066657 (rank : 27) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
CN032_HUMAN
|
||||||
| NC score | 0.066431 (rank : 28) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
FA43B_HUMAN
|
||||||
| NC score | 0.065828 (rank : 29) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZT52 | Gene names | FAM43B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM43B. | |||||
|
APBB1_HUMAN
|
||||||
| NC score | 0.065082 (rank : 30) | θ value | 2.36792 (rank : 46) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O00213, Q96A93 | Gene names | APBB1, FE65 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1 (Fe65 protein). | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.064638 (rank : 31) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.063665 (rank : 32) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
CN032_MOUSE
|
||||||
| NC score | 0.063265 (rank : 33) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BH93, Q3TF12, Q3TGL0, Q8CC90 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32 homolog. | |||||
|
SF3A2_HUMAN
|
||||||
| NC score | 0.060580 (rank : 34) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.057750 (rank : 35) | θ value | 0.62314 (rank : 35) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.057322 (rank : 36) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
FMN2_MOUSE
|
||||||
| NC score | 0.055969 (rank : 37) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JL04 | Gene names | Fmn2 | |||
|
Domain Architecture |

|
|||||
| Description | Formin-2. | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.055947 (rank : 38) | θ value | 0.62314 (rank : 33) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
SF3A2_MOUSE
|
||||||
| NC score | 0.055461 (rank : 39) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
PRR13_HUMAN
|
||||||
| NC score | 0.055104 (rank : 40) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZ81, Q6FIG7, Q6MZP8, Q6NXQ6, Q6PKF9 | Gene names | PRR13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 13. | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.054568 (rank : 41) | θ value | 0.62314 (rank : 32) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CCNK_MOUSE
|
||||||
| NC score | 0.054257 (rank : 42) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.053963 (rank : 43) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
PDC6I_MOUSE
|
||||||
| NC score | 0.053757 (rank : 44) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9WU78, O88695, O89014, Q8BSL8, Q8R0H5, Q99LR3, Q9QZN8 | Gene names | Pdcd6ip, Aip1, Alix | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death 6-interacting protein (ALG-2-interacting protein X) (ALG-2-interacting protein 1) (E2F1-inducible protein) (Eig2). | |||||
|
JIP2_MOUSE
|
||||||
| NC score | 0.053315 (rank : 45) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ERE9, Q9CXI4 | Gene names | Mapk8ip2, Ib2, Jip2 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 2 (JNK-interacting protein 2) (JIP-2) (JNK MAP kinase scaffold protein 2) (Islet-brain-2) (IB-2) (Mitogen-activated protein kinase 8-interacting protein 2). | |||||
|
SELV_HUMAN
|
||||||
| NC score | 0.052617 (rank : 46) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P59797 | Gene names | SELV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Selenoprotein V. | |||||
|
JIP2_HUMAN
|
||||||
| NC score | 0.052572 (rank : 47) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13387, Q96G62, Q99771, Q9NZ59, Q9UKQ4 | Gene names | MAPK8IP2, IB2, JIP2 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 2 (JNK-interacting protein 2) (JIP-2) (JNK MAP kinase scaffold protein 2) (Islet-brain-2) (IB-2) (Mitogen-activated protein kinase 8-interacting protein 2). | |||||
|
GGN_MOUSE
|
||||||
| NC score | 0.052437 (rank : 48) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
WASL_HUMAN
|
||||||
| NC score | 0.052035 (rank : 49) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00401, Q7Z746 | Gene names | WASL | |||
|
Domain Architecture |

|
|||||
| Description | Neural Wiskott-Aldrich syndrome protein (N-WASP). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.051703 (rank : 50) | θ value | 0.813845 (rank : 39) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
FMN2_HUMAN
|
||||||
| NC score | 0.051040 (rank : 51) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
TM108_MOUSE
|
||||||
| NC score | 0.050640 (rank : 52) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BHE4, Q80WR9 | Gene names | Tmem108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 108 precursor. | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.050426 (rank : 53) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
PER2_MOUSE
|
||||||
| NC score | 0.049930 (rank : 54) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.049853 (rank : 55) | θ value | 0.163984 (rank : 26) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
MAML2_HUMAN
|
||||||
| NC score | 0.049626 (rank : 56) | θ value | 0.21417 (rank : 27) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IZL2, Q6AI23, Q6Y3A3, Q8IUL3, Q96JK6 | Gene names | MAML2, KIAA1819 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 2 (Mam-2). | |||||
|
NU214_HUMAN
|
||||||
| NC score | 0.048514 (rank : 57) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.048095 (rank : 58) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.045964 (rank : 59) | θ value | 3.0926 (rank : 51) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
GSCR1_HUMAN
|
||||||
| NC score | 0.045938 (rank : 60) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.045722 (rank : 61) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.045580 (rank : 62) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.044514 (rank : 63) | θ value | 1.81305 (rank : 45) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
ARI1B_HUMAN
|
||||||
| NC score | 0.044358 (rank : 64) | θ value | 0.813845 (rank : 38) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.043034 (rank : 65) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.041859 (rank : 66) | θ value | 0.365318 (rank : 29) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
MAML3_HUMAN
|
||||||
| NC score | 0.039765 (rank : 67) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96JK9 | Gene names | MAML3, KIAA1816 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 3 (Mam-3). | |||||
|
ANX11_MOUSE
|
||||||
| NC score | 0.038546 (rank : 68) | θ value | 0.279714 (rank : 28) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97384 | Gene names | Anxa11, Anx11 | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A11 (Annexin XI) (Calcyclin-associated annexin 50) (CAP-50). | |||||
|
SGIP1_MOUSE
|
||||||
| NC score | 0.038502 (rank : 69) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8VD37, Q3UFU3, Q3UGA0, Q8BXX4, Q8C034, Q9CXT2 | Gene names | Sgip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3-containing GRB2-like protein 3-interacting protein 1 (Endophilin- 3-interacting protein). | |||||
|
FUBP2_HUMAN
|
||||||
| NC score | 0.038464 (rank : 70) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92945, O00301, Q9UNT5, Q9UQH5 | Gene names | KHSRP, FUBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 2 (FUSE-binding protein 2) (KH type-splicing regulatory protein) (KSRP) (p75). | |||||
|
BAT3_MOUSE
|
||||||
| NC score | 0.038189 (rank : 71) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Z1R2, Q8SNA3 | Gene names | Bat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.037506 (rank : 72) | θ value | 0.62314 (rank : 34) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
BRD4_MOUSE
|
||||||
| NC score | 0.033214 (rank : 73) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.033084 (rank : 74) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
NOTC2_MOUSE
|
||||||
| NC score | 0.032818 (rank : 75) | θ value | 0.00869519 (rank : 17) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1069 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35516, Q06008, Q60941 | Gene names | Notch2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (Motch B) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
NOTC2_HUMAN
|
||||||
| NC score | 0.030286 (rank : 76) | θ value | 0.0431538 (rank : 21) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q04721, Q99734, Q9H240 | Gene names | NOTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (hN2) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
EMSY_MOUSE
|
||||||
| NC score | 0.030092 (rank : 77) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BMB0, Q5FWK5, Q80XU1, Q8VDW9 | Gene names | Emsy | |||
|
Domain Architecture |

|
|||||
| Description | Protein EMSY. | |||||
|
MUC5B_HUMAN
|
||||||
| NC score | 0.029515 (rank : 78) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
CNOT3_HUMAN
|
||||||
| NC score | 0.028839 (rank : 79) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75175, Q9NZN7, Q9UF76 | Gene names | CNOT3, KIAA0691, NOT3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
NUFP1_HUMAN
|
||||||
| NC score | 0.027409 (rank : 80) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UHK0, Q8WVM5, Q96SG1 | Gene names | NUFIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 1 (Nuclear FMRP-interacting protein 1). | |||||
|
RD23B_MOUSE
|
||||||
| NC score | 0.026927 (rank : 81) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P54728 | Gene names | Rad23b, Mhr23b | |||
|
Domain Architecture |

|
|||||
| Description | UV excision repair protein RAD23 homolog B (mHR23B) (XP-C repair- complementing complex 58 kDa protein) (p58). | |||||
|
ATX2L_HUMAN
|
||||||
| NC score | 0.023410 (rank : 82) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8WWM7, O95135, Q6NVJ8, Q6PJW6, Q8IU61, Q8IU95, Q8WWM3, Q8WWM4, Q8WWM5, Q8WWM6, Q99703 | Gene names | ATXN2L, A2D, A2LG, A2LP, A2RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein (Ataxin-2 domain protein) (Ataxin-2-related protein). | |||||
|
MCM3A_HUMAN
|
||||||
| NC score | 0.023125 (rank : 83) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60318, Q9UMT4 | Gene names | MCM3AP, GANP, KIAA0572, MAP80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
NEUM_MOUSE
|
||||||
| NC score | 0.021073 (rank : 84) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P06837 | Gene names | Gap43, Basp2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuromodulin (Axonal membrane protein GAP-43) (Growth-associated protein 43) (Calmodulin-binding protein P-57). | |||||
|
ANXA7_MOUSE
|
||||||
| NC score | 0.020361 (rank : 85) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q07076 | Gene names | Anxa7, Anx7 | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A7 (Annexin VII) (Synexin). | |||||
|
PRKN2_HUMAN
|
||||||
| NC score | 0.019668 (rank : 86) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60260, Q5TFV8, Q6Q2I6, Q8NI41, Q8NI43, Q8NI44, Q8WW07 | Gene names | PARK2, PRKN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parkin (EC 6.3.2.-) (Ubiquitin E3 ligase PRKN) (Parkinson juvenile disease protein 2) (Parkinson disease protein 2). | |||||
|
EP400_MOUSE
|
||||||
| NC score | 0.017930 (rank : 87) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
ADA29_HUMAN
|
||||||
| NC score | 0.017366 (rank : 88) | θ value | 0.813845 (rank : 36) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UKF5, Q9UHP1, Q9UKF3, Q9UKF4 | Gene names | ADAM29 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 29 precursor (A disintegrin and metalloproteinase domain 29). | |||||
|
PI4KB_HUMAN
|
||||||
| NC score | 0.017307 (rank : 89) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBF8, O15096, P78405, Q9BWR6 | Gene names | PIK4CB, PI4KB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase beta (EC 2.7.1.67) (PtdIns 4-kinase beta) (PI4Kbeta) (PI4K-beta) (NPIK) (PI4K92). | |||||
|
MYCL1_MOUSE
|
||||||
| NC score | 0.016583 (rank : 90) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
SALL3_HUMAN
|
||||||
| NC score | 0.014944 (rank : 91) | θ value | 0.0736092 (rank : 22) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1066 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9BXA9, Q9UGH1 | Gene names | SALL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 3 (Zinc finger protein SALL3) (hSALL3). | |||||
|
DEN1A_MOUSE
|
||||||
| NC score | 0.012991 (rank : 92) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8K382 | Gene names | Dennd1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
SEM6C_MOUSE
|
||||||
| NC score | 0.011094 (rank : 93) | θ value | 1.81305 (rank : 44) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
MEF2A_HUMAN
|
||||||
| NC score | 0.010937 (rank : 94) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q02078, O43814, Q14223, Q14224, Q96D14 | Gene names | MEF2A, MEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2A (Serum response factor-like protein 1). | |||||
|
RC3H1_HUMAN
|
||||||
| NC score | 0.010527 (rank : 95) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
XLKD1_HUMAN
|
||||||
| NC score | 0.010150 (rank : 96) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5Y7, Q8TC18, Q9UNF4 | Gene names | XLKD1, CRSBP1, HAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lymphatic vessel endothelial hyaluronic acid receptor 1 precursor (LYVE-1) (Cell surface retention sequence-binding protein 1) (CRSBP-1) (Hyaluronic acid receptor) (Extracellular link domain-containing protein 1). | |||||
|
MYO15_MOUSE
|
||||||
| NC score | 0.009834 (rank : 97) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9QZZ4, O70395, Q9QWL6 | Gene names | Myo15a, Myo15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
RPGF2_HUMAN
|
||||||
| NC score | 0.009124 (rank : 98) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y4G8 | Gene names | RAPGEF2, KIAA0313, NRAPGEP, PDZGEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rap guanine nucleotide exchange factor 2 (Neural RAP guanine nucleotide exchange protein) (nRap GEP) (PDZ domain-containing guanine nucleotide exchange factor 1) (PDZ-GEF1) (RA-GEF). | |||||
|
DOCK7_HUMAN
|
||||||
| NC score | 0.009016 (rank : 99) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96N67, Q6ZV32, Q8TB82, Q96NG6, Q9C092 | Gene names | DOCK7, KIAA1771 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dedicator of cytokinesis protein 7. | |||||
|
CCD45_MOUSE
|
||||||
| NC score | 0.008979 (rank : 100) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 663 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BVV7 | Gene names | Ccdc45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 45. | |||||
|
CD2L6_MOUSE
|
||||||
| NC score | 0.008927 (rank : 101) | θ value | 0.125558 (rank : 25) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BWD8, Q80TM1 | Gene names | Cdc2l6, Kiaa1028 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 6 (EC 2.7.11.22) (CDC2- related protein kinase 6). | |||||
|
PGBM_MOUSE
|
||||||
| NC score | 0.008095 (rank : 102) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q05793 | Gene names | Hspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
WNK1_HUMAN
|
||||||
| NC score | 0.007777 (rank : 103) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1158 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9H4A3, O15052, Q86WL5, Q8N673, Q9P1S9 | Gene names | WNK1, KDP, KIAA0344, PRKWNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1) (Kinase deficient protein). | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | 0.007769 (rank : 104) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
SN1L1_HUMAN
|
||||||
| NC score | 0.007240 (rank : 105) | θ value | 0.365318 (rank : 30) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P57059, Q5R2V5, Q86YJ2 | Gene names | SNF1LK | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 1 (EC 2.7.11.1) (Serine/threonine-protein kinase SNF1LK). | |||||
|
LYST_MOUSE
|
||||||
| NC score | 0.006639 (rank : 106) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97412, Q62403, Q8VBS6 | Gene names | Lyst, Bg, Chs1 | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal-trafficking regulator (Beige protein) (CHS1 homolog). | |||||
|
SSH1_MOUSE
|
||||||
| NC score | 0.006234 (rank : 107) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q76I79, Q3TDG3, Q69ZM4, Q811E5 | Gene names | Ssh1, Kiaa1298, Ssh1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 1 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-1L) (mSSH-1L). | |||||
|
SKI_MOUSE
|
||||||
| NC score | 0.006008 (rank : 108) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q60698, Q8VIL5 | Gene names | Ski | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ski oncogene (C-ski). | |||||
|
ITSN1_HUMAN
|
||||||
| NC score | 0.005910 (rank : 109) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
CD2L6_HUMAN
|
||||||
| NC score | 0.005695 (rank : 110) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BWU1, Q5JQZ7, Q5JR00, Q8TC78, Q9UPX2 | Gene names | CDC2L6, CDK11, KIAA1028 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 6 (EC 2.7.11.22) (CDC2- related protein kinase 6) (Death-preventing kinase) (Cyclin-dependent kinase 11). | |||||
|
BNC1_MOUSE
|
||||||
| NC score | 0.005394 (rank : 111) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35914, O54886 | Gene names | Bnc1, Bnc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein basonuclin-1. | |||||
|
ANGL3_HUMAN
|
||||||
| NC score | 0.004655 (rank : 112) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5C1 | Gene names | ANGPTL3, ANGPT5 | |||
|
Domain Architecture |

|
|||||
| Description | Angiopoietin-related protein 3 precursor (Angiopoietin-like 3) (Angiopoietin-5) (ANG-5). | |||||
|
WNK3_HUMAN
|
||||||
| NC score | 0.003912 (rank : 113) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BYP7, Q9HCK6 | Gene names | WNK3, KIAA1566, PRKWNK3 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK3 (EC 2.7.11.1) (Protein kinase with no lysine 3) (Protein kinase, lysine-deficient 3). | |||||
|
ZEP2_HUMAN
|
||||||
| NC score | 0.003679 (rank : 114) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P31629, Q02646, Q5THT5, Q9NS05 | Gene names | HIVEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 (HIV- EP2) (MHC-binding protein 2) (MBP-2). | |||||
|
EPHAA_HUMAN
|
||||||
| NC score | 0.002440 (rank : 115) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5JZY3, Q6NW42 | Gene names | EPHA10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ephrin type-A receptor 10 precursor (EC 2.7.10.1). | |||||