Please be patient as the page loads
|
EZH2_HUMAN
|
||||||
| SwissProt Accessions | Q15910, Q15755, Q92857 | Gene names | EZH2 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 2 (ENX-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
EZH1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.988594 (rank : 3) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q92800, O43287, Q14459 | Gene names | EZH1, KIAA0388 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 1 (ENX-2). | |||||
|
EZH1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.988470 (rank : 4) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P70351, Q9R089 | Gene names | Ezh1, Enx2 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 1 (ENX-2). | |||||
|
EZH2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q15910, Q15755, Q92857 | Gene names | EZH2 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 2 (ENX-1). | |||||
|
EZH2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999387 (rank : 2) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q61188, Q9R090 | Gene names | Ezh2, Enx1h | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 2 (ENX-1). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 3.17815e-21 (rank : 5) | NC score | 0.558268 (rank : 15) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
NSD1_MOUSE
|
||||||
| θ value | 2.06002e-20 (rank : 6) | NC score | 0.565590 (rank : 14) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
SUV91_HUMAN
|
||||||
| θ value | 2.06002e-20 (rank : 7) | NC score | 0.701618 (rank : 5) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O43463, Q53G60, Q6FHK6 | Gene names | SUV39H1, SUV39H | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1). | |||||
|
SUV91_MOUSE
|
||||||
| θ value | 2.06002e-20 (rank : 8) | NC score | 0.695477 (rank : 6) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O54864, Q9JLC7, Q9JLP8 | Gene names | Suv39h1, Suv39h | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1) (Position-effect variegation 3-9 homolog). | |||||
|
NSD1_HUMAN
|
||||||
| θ value | 2.69047e-20 (rank : 9) | NC score | 0.568112 (rank : 13) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
SETD2_HUMAN
|
||||||
| θ value | 2.69047e-20 (rank : 10) | NC score | 0.656445 (rank : 9) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
SET1A_HUMAN
|
||||||
| θ value | 3.51386e-20 (rank : 11) | NC score | 0.614956 (rank : 10) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
SUV92_MOUSE
|
||||||
| θ value | 2.97466e-19 (rank : 12) | NC score | 0.685381 (rank : 8) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9EQQ0, Q9CUK3, Q9JLP7 | Gene names | Suv39h2 | |||
|
Domain Architecture |
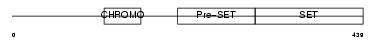
|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
SUV92_HUMAN
|
||||||
| θ value | 3.88503e-19 (rank : 13) | NC score | 0.691368 (rank : 7) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9H5I1 | Gene names | SUV39H2 | |||
|
Domain Architecture |
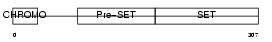
|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 5.07402e-19 (rank : 14) | NC score | 0.539904 (rank : 16) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 8.65492e-19 (rank : 15) | NC score | 0.415554 (rank : 22) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 8.65492e-19 (rank : 16) | NC score | 0.436981 (rank : 20) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 2.5182e-18 (rank : 17) | NC score | 0.424426 (rank : 21) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
EHMT2_MOUSE
|
||||||
| θ value | 7.32683e-18 (rank : 18) | NC score | 0.341998 (rank : 24) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
EHMT2_HUMAN
|
||||||
| θ value | 1.63225e-17 (rank : 19) | NC score | 0.344295 (rank : 23) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q96KQ7, Q14349, Q6PK06, Q96MH5, Q96QD0, Q9UQL8, Q9Y331 | Gene names | EHMT2, BAT8, G9A, NG36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
EHMT1_HUMAN
|
||||||
| θ value | 2.60593e-15 (rank : 20) | NC score | 0.333982 (rank : 25) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9H9B1, Q8TCN7, Q96F53, Q96JF1, Q96KH4 | Gene names | EHMT1, EUHMTASE1, KIAA1876 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 5 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 5) (H3-K9-HMTase 5) (Euchromatic histone-lysine N-methyltransferase 1) (Eu-HMTase1) (G9a- like protein 1) (GLP1). | |||||
|
SETD8_HUMAN
|
||||||
| θ value | 3.29651e-10 (rank : 21) | NC score | 0.612180 (rank : 11) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
SETD8_MOUSE
|
||||||
| θ value | 3.29651e-10 (rank : 22) | NC score | 0.611189 (rank : 12) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
SETB1_MOUSE
|
||||||
| θ value | 6.43352e-06 (rank : 23) | NC score | 0.469258 (rank : 18) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O88974, Q922K1 | Gene names | Setdb1, Eset | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
SETB1_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 24) | NC score | 0.464149 (rank : 19) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q15047, Q96GM9 | Gene names | SETDB1, KIAA0067 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
SETB2_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 25) | NC score | 0.504110 (rank : 17) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96T68, Q86UD6, Q96AI6 | Gene names | SETDB2, CLLD8 | |||
|
Domain Architecture |

|
|||||
| Description | Probable histone-lysine N-methyltransferase, H3 lysine-9 specific (EC 2.1.1.43) (Histone H3-K9 methyltransferase) (H3-K9-HMTase) (SET domain bifurcated 2) (Chronic lymphocytic leukemia deletion region gene 8 protein). | |||||
|
SETD7_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 26) | NC score | 0.196432 (rank : 26) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8VHL1 | Gene names | Setd7, Set7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7). | |||||
|
SETD7_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 27) | NC score | 0.190698 (rank : 27) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WTS6, Q9C0E6 | Gene names | SETD7, KIAA1717, SET7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7) (Set9) (SET7/9). | |||||
|
MCSP_MOUSE
|
||||||
| θ value | 0.125558 (rank : 28) | NC score | 0.160192 (rank : 28) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P15265, O70613, Q6P8N3 | Gene names | Smcp, Mcs, Mcsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm mitochondrial-associated cysteine-rich protein. | |||||
|
KRA58_HUMAN
|
||||||
| θ value | 0.21417 (rank : 29) | NC score | 0.093920 (rank : 31) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75690, Q6L8G7, Q6UTX6 | Gene names | KRTAP5-8, KAP5.8, KRTAP5.8, UHSKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-8 (Keratin-associated protein 5.8) (Ultrahigh sulfur keratin-associated protein 5.8) (Keratin, ultra high-sulfur matrix protein B) (UHS keratin B) (UHS KerB). | |||||
|
LR37A_HUMAN
|
||||||
| θ value | 0.21417 (rank : 30) | NC score | 0.049430 (rank : 79) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60309, Q49A01, Q49A80, Q8NB33 | Gene names | LRRC37A, KIAA0563 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37A. | |||||
|
NOTC2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 31) | NC score | 0.040558 (rank : 87) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q04721, Q99734, Q9H240 | Gene names | NOTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (hN2) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
CENB5_HUMAN
|
||||||
| θ value | 0.47712 (rank : 32) | NC score | 0.035757 (rank : 92) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96P50, Q9BSR9, Q9C0E7 | Gene names | CENTB5, KIAA1716 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 5 (Cnt-b5). | |||||
|
KR511_HUMAN
|
||||||
| θ value | 0.47712 (rank : 33) | NC score | 0.088388 (rank : 32) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6L8G4 | Gene names | KRTAP5-11, KAP5.11, KRTAP5.11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-11 (Keratin-associated protein 5.11) (Ultrahigh sulfur keratin-associated protein 5.11). | |||||
|
KRA24_HUMAN
|
||||||
| θ value | 0.47712 (rank : 34) | NC score | 0.066491 (rank : 46) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9BYR9, Q495J2 | Gene names | KRTAP2-4, KAP2.4, KRTAP2.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 2-4 (Keratin-associated protein 2.4) (High sulfur keratin-associated protein 2.4). | |||||
|
ITB1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 35) | NC score | 0.036893 (rank : 90) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P05556, P78466, P78467, Q13089, Q13090, Q13091, Q13212, Q14622, Q14647 | Gene names | ITGB1, FNRB | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
KIF4A_HUMAN
|
||||||
| θ value | 0.62314 (rank : 36) | NC score | 0.017989 (rank : 109) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O95239, Q86TN3, Q86XX7, Q9NNY6, Q9NY24, Q9UMW3 | Gene names | KIF4A, KIF4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
KRA53_HUMAN
|
||||||
| θ value | 0.62314 (rank : 37) | NC score | 0.076953 (rank : 41) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6L8H2, Q6PL44, Q701N3 | Gene names | KRTAP5-3, KAP5-9, KAP5.3, KRTAP5-9, KRTAP5.3, KRTAP5.9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-3 (Keratin-associated protein 5.3) (Ultrahigh sulfur keratin-associated protein 5.3) (Keratin-associated protein 5-9) (Keratin-associated protein 5.9) (UHS KerB-like). | |||||
|
KRA57_HUMAN
|
||||||
| θ value | 0.62314 (rank : 38) | NC score | 0.067761 (rank : 45) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6L8G8, Q701N5 | Gene names | KRTAP5-7, KAP5-7, KAP5.3, KRTAP5.3, KRTAP5.7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-7 (Keratin-associated protein 5.7) (Ultrahigh sulfur keratin-associated protein 5.7) (Keratin-associated protein 5-3) (Keratin-associated protein 5.3). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 0.62314 (rank : 39) | NC score | 0.045007 (rank : 84) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
DYNA_MOUSE
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.013834 (rank : 114) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O08788 | Gene names | Dctn1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued). | |||||
|
IF4G3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.025900 (rank : 105) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80XI3, Q80Y69, Q8BJ68, Q8BQF3, Q8C6Q3, Q8CIH0 | Gene names | Eif4g3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
ITB1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.029234 (rank : 100) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P09055 | Gene names | Itgb1 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
NOTC2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 43) | NC score | 0.036583 (rank : 91) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1069 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O35516, Q06008, Q60941 | Gene names | Notch2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (Motch B) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
SSPO_MOUSE
|
||||||
| θ value | 1.38821 (rank : 44) | NC score | 0.031244 (rank : 95) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
STAB1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 45) | NC score | 0.020825 (rank : 106) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 546 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NY15, Q8IUH0, Q8IUH1, Q93072 | Gene names | STAB1, FEEL1, KIAA0246 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein) (MS-1 antigen). | |||||
|
ZNFX1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.047446 (rank : 82) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8R151, Q3U016, Q3UD71, Q3UEZ0 | Gene names | ZNFX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
ATS13_MOUSE
|
||||||
| θ value | 1.81305 (rank : 47) | NC score | 0.013518 (rank : 115) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q769J6, Q76LW1 | Gene names | Adamts13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-13 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 13) (ADAM-TS 13) (ADAM-TS13) (von Willebrand factor-cleaving protease) (vWF-cleaving protease) (vWF-CP). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 48) | NC score | 0.041615 (rank : 85) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
NOTC1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.031944 (rank : 94) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P46531 | Gene names | NOTCH1, TAN1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 1 precursor (Notch 1) (hN1) (Translocation-associated notch protein TAN-1) [Contains: Notch 1 extracellular truncation; Notch 1 intracellular domain]. | |||||
|
RCOR3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.030285 (rank : 99) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
ZNFX1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.045981 (rank : 83) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
MT2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.047640 (rank : 81) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P02798 | Gene names | Mt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-2 (MT-2) (Metallothionein-II) (MT-II). | |||||
|
ATL2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.011752 (rank : 116) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TSK7, Q3U0C8 | Gene names | Adamtsl2, Kiaa0605, Tcp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-like protein 2 precursor (ADAMTSL-2) (TSP1-repeats-containing protein 1) (TCP-1). | |||||
|
DYNA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.015647 (rank : 112) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1194 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14203, O95296, Q9BRM9, Q9UIU1, Q9UIU2 | Gene names | DCTN1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued) (p135). | |||||
|
KRA52_HUMAN
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.031094 (rank : 96) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q701N4 | Gene names | KRTAP5-2, KAP5-8, KAP5.2, KRTAP5-8, KRTAP5.2, KRTAP5.8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-2 (Keratin-associated protein 5.2) (Ultrahigh sulfur keratin-associated protein 5.2) (Keratin-associated protein 5-8) (Keratin-associated protein 5.8). | |||||
|
MEGF6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.019911 (rank : 107) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.047885 (rank : 80) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
OXRP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.017869 (rank : 110) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 544 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y4L1 | Gene names | HYOU1, ORP150 | |||
|
Domain Architecture |

|
|||||
| Description | 150 kDa oxygen-regulated protein precursor (Orp150) (Hypoxia up- regulated 1). | |||||
|
PGBM_MOUSE
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.007592 (rank : 120) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q05793 | Gene names | Hspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
ZAN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.036993 (rank : 89) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
FGD4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.005645 (rank : 125) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96M96, Q6ULS2, Q8TCP6 | Gene names | FGD4, FRABP, ZFYVE6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein) (Zinc finger FYVE domain-containing protein 6). | |||||
|
KRA59_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.059906 (rank : 58) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
RCOR2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.032372 (rank : 93) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IZ40, Q96FP3 | Gene names | RCOR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2. | |||||
|
RCOR2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.037295 (rank : 88) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C796, Q9JMK4 | Gene names | Rcor2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2 (M-CoREST). | |||||
|
UBP25_MOUSE
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.007612 (rank : 119) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P57080, Q80ZT9 | Gene names | Usp25 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (mUSP25). | |||||
|
VEGFC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.027175 (rank : 104) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97953, Q543R6 | Gene names | Vegfc | |||
|
Domain Architecture |
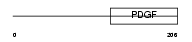
|
|||||
| Description | Vascular endothelial growth factor C precursor (VEGF-C) (Vascular endothelial growth factor-related protein) (VRP) (Flt4 ligand) (Flt4- L). | |||||
|
ALBU_MOUSE
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.007006 (rank : 121) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P07724, Q61802 | Gene names | Alb, Alb-1, Alb1 | |||
|
Domain Architecture |

|
|||||
| Description | Serum albumin precursor. | |||||
|
GPC6A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.006242 (rank : 123) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K4Z6, Q5DK51, Q5DK52 | Gene names | Gprc6a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor family C group 6 member A precursor. | |||||
|
KIF4A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.010392 (rank : 118) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
KR105_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.041224 (rank : 86) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P60370, Q70LJ3 | Gene names | KRTAP10-5, KAP10.5, KAP18-5, KRTAP10.5, KRTAP18-5, KRTAP18.5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 10-5 (Keratin-associated protein 10.5) (High sulfur keratin-associated protein 10.5) (Keratin-associated protein 18-5) (Keratin-associated protein 18.5). | |||||
|
MYSM1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.030674 (rank : 97) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5VVJ2, Q7Z3G8, Q96PX3 | Gene names | MYSM1, KIAA1915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MYSM1 (Myb-like, SWIRM and MPN domain-containing protein 1). | |||||
|
PKHA6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.017578 (rank : 111) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
SMYD3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.062266 (rank : 56) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H7B4, Q86TL8, Q8N5Z6, Q96AI5 | Gene names | SMYD3, ZMYND1, ZNFN3A1 | |||
|
Domain Architecture |
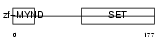
|
|||||
| Description | SET and MYND domain-containing protein 3 (EC 2.1.1.43) (Zinc finger MYND domain-containing protein 1). | |||||
|
TGON1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.019717 (rank : 108) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
UB7I4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.011299 (rank : 117) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q925F3, Q6A0B4 | Gene names | Rnf144, Kiaa0161, Ubce7ip4, Uip4 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin-conjugating enzyme 7-interacting protein 4 (UbcM4- interacting protein 4) (RING finger protein 144). | |||||
|
VWF_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.028645 (rank : 102) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8CIZ8, Q60863, Q6XUV6, Q8BIU9, Q8CGN0, Q9JK16 | Gene names | Vwf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | von Willebrand factor precursor (vWF) [Contains: von Willebrand antigen 2 (von Willebrand antigen II)]. | |||||
|
BAI3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.005793 (rank : 124) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60242, O60297 | Gene names | BAI3, KIAA0550 | |||
|
Domain Architecture |

|
|||||
| Description | Brain-specific angiogenesis inhibitor 3 precursor. | |||||
|
CJ012_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.029102 (rank : 101) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
CRIM1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.030425 (rank : 98) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NZV1, Q59GH0, Q7LCQ5, Q9H318 | Gene names | CRIM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich motor neuron 1 protein precursor (CRIM-1) (Cysteine-rich repeat-containing protein S52). | |||||
|
DLL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.015560 (rank : 113) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00548, Q9NU41, Q9UJV2 | Gene names | DLL1 | |||
|
Domain Architecture |
|
|||||
| Description | Delta-like protein 1 precursor (Drosophila Delta homolog 1) (Delta1) (H-Delta-1). | |||||
|
EGFR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | -0.001354 (rank : 127) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P00533, O00688, O00732, P06268, Q14225, Q92795, Q9BZS2, Q9GZX1, Q9H2C9, Q9H3C9, Q9UMD7, Q9UMD8, Q9UMG5 | Gene names | EGFR, ERBB1 | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor precursor (EC 2.7.10.1) (Receptor tyrosine-protein kinase ErbB-1). | |||||
|
GP116_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.006947 (rank : 122) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IZF2, O94858, Q6RGN2, Q86SP0, Q9Y3Z2 | Gene names | GPR116, KIAA0758 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 116 precursor. | |||||
|
MIB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.028404 (rank : 103) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86YT6, Q68D01, Q6YI51, Q8NBY0, Q8TCB5, Q8TCL7, Q9P2M3 | Gene names | MIB1, DIP1, KIAA1323, ZZANK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein MIB1 (EC 6.3.2.-) (Mind bomb homolog 1) (DAPK-interacting protein 1) (DIP-1) (Zinc finger ZZ type with ankyrin repeat domain protein 2). | |||||
|
TIE2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.000123 (rank : 126) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1204 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q02858 | Gene names | Tek, Hyk, Tie-2, Tie2 | |||
|
Domain Architecture |

|
|||||
| Description | Angiopoietin-1 receptor precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor TIE-2) (mTIE2) (Tyrosine-protein kinase receptor TEK) (p140 TEK) (Tunica interna endothelial cell kinase) (HYK) (STK1) (CD202b antigen). | |||||
|
AIRE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.064092 (rank : 55) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
AIRE_MOUSE
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.066341 (rank : 48) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
CBX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.057709 (rank : 64) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P83916, P23197 | Gene names | CBX1, CBX | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25) (HP1Hsbeta) (p25beta). | |||||
|
CBX1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.057709 (rank : 65) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P83917, P23197 | Gene names | Cbx1, Cbx | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25). | |||||
|
CBX3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.059222 (rank : 63) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13185, Q96CD7, Q99409, Q9BVS3, Q9P0Z6 | Gene names | CBX3 | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (HECH). | |||||
|
CBX3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.059807 (rank : 59) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23198, Q921C4 | Gene names | Cbx3 | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (M32). | |||||
|
CBX5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.054797 (rank : 71) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P45973 | Gene names | CBX5, HP1A | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha) (Antigen p25). | |||||
|
CBX5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.054493 (rank : 74) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61686, Q9CS35, Q9CXD1 | Gene names | Cbx5, Hp1a | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha). | |||||
|
DPF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.074748 (rank : 42) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
DPF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.066159 (rank : 49) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QX66, Q9QX65, Q9QYA4 | Gene names | Dpf1, Neud4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
K1333_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.065162 (rank : 53) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7L622, Q9BVR2, Q9H9E9, Q9NXC0, Q9P2L3 | Gene names | KIAA1333 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1333 (EC 6.3.2.-). | |||||
|
K1333_MOUSE
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.065221 (rank : 52) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5RJY2, Q6ZPT7, Q8BNA4 | Gene names | Kiaa1333 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1333 (EC 6.3.2.-). | |||||
|
LCE1D_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.055227 (rank : 69) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5T752 | Gene names | LCE1D, LEP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Late cornified envelope protein 1D (Late envelope protein 4). | |||||
|
LCE1E_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.054141 (rank : 75) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5T753 | Gene names | LCE1E, LEP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Late cornified envelope protein 1E (Late envelope protein 5). | |||||
|
MYST3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.055510 (rank : 68) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
MYST3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.057004 (rank : 66) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
MYST4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.065687 (rank : 50) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
MYST4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.059233 (rank : 62) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
PF21A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.059593 (rank : 60) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
PF21A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.059281 (rank : 61) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
PF21B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.050131 (rank : 78) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PF21B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.055847 (rank : 67) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PHF10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.084335 (rank : 36) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WUB8, Q2YDA3, Q9BXD2, Q9NV26 | Gene names | PHF10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10 (XAP135). | |||||
|
PHF10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.082816 (rank : 37) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
PHF11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.114498 (rank : 30) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UIL8, Q9Y5A2 | Gene names | PHF11, BCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 11 (BRCA1 C-terminus-associated protein) (NY-REN-34 antigen). | |||||
|
PHF12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.054778 (rank : 72) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.054828 (rank : 70) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.087525 (rank : 33) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IWS0, Q96JK3, Q9BRU0 | Gene names | PHF6, KIAA1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6 (PHD-like zinc finger protein). | |||||
|
PHF6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.084980 (rank : 35) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D4J7, Q80T86, Q8BYT8, Q8C1B5 | Gene names | Phf6, Kiaa1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6. | |||||
|
PHF7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.071178 (rank : 44) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BWX1 | Gene names | PHF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 7 (Testis development protein NYD-SP6). | |||||
|
PHF7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.066462 (rank : 47) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DAG9, Q6PG81 | Gene names | Phf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 7 (Testis development protein NYD-SP6). | |||||
|
PTMA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.114519 (rank : 29) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
PTMA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.080314 (rank : 38) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
PYGO2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.050458 (rank : 77) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BRQ0, Q8WYZ4, Q96CY2 | Gene names | PYGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 2. | |||||
|
RAI1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.078618 (rank : 40) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z5J4, Q8N3B4, Q8ND08, Q8WU64, Q96JK5, Q9H1C1, Q9H1C2, Q9UF69 | Gene names | RAI1, KIAA1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
RAI1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.085103 (rank : 34) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61818, Q6ZPH7, Q8CEV1 | Gene names | Rai1, Kiaa1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
REQU_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.065408 (rank : 51) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 523 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92785 | Gene names | DPF2, REQ, UBID4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
REQU_MOUSE
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.064933 (rank : 54) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61103, Q3UNP5, Q60663, Q9QYA3 | Gene names | Dpf2, Req, Ubid4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
SMYD3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.060042 (rank : 57) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9CWR2, Q6P7V6, Q8BG90 | Gene names | Smyd3, Zmynd1 | |||
|
Domain Architecture |

|
|||||
| Description | SET and MYND domain-containing protein 3 (EC 2.1.1.43) (Zinc finger MYND domain-containing protein 1). | |||||
|
TCF20_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.078690 (rank : 39) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UGU0, O14528, Q13078, Q4V353, Q9H4M0 | Gene names | TCF20, KIAA0292, SPBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP) (AR1). | |||||
|
TCF20_MOUSE
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.074307 (rank : 43) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
UHRF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.051107 (rank : 76) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96T88, Q8J022, Q9H6S6, Q9P115, Q9P1U7 | Gene names | UHRF1, ICBP90, NP95, RNF106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Inverted CCAAT box-binding protein of 90 kDa) (Transcription factor ICBP90) (Nuclear zinc finger protein Np95) (Nuclear protein 95) (HuNp95) (RING finger protein 106). | |||||
|
UHRF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.054607 (rank : 73) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VDF2, Q8C6F1, Q8VIA1, Q9Z1H6 | Gene names | Uhrf1, Np95 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Nuclear zinc finger protein Np95) (Nuclear protein 95). | |||||
|
EZH2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q15910, Q15755, Q92857 | Gene names | EZH2 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 2 (ENX-1). | |||||
|
EZH2_MOUSE
|
||||||
| NC score | 0.999387 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q61188, Q9R090 | Gene names | Ezh2, Enx1h | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 2 (ENX-1). | |||||
|
EZH1_HUMAN
|
||||||
| NC score | 0.988594 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q92800, O43287, Q14459 | Gene names | EZH1, KIAA0388 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 1 (ENX-2). | |||||
|
EZH1_MOUSE
|
||||||
| NC score | 0.988470 (rank : 4) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P70351, Q9R089 | Gene names | Ezh1, Enx2 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 1 (ENX-2). | |||||
|
SUV91_HUMAN
|
||||||
| NC score | 0.701618 (rank : 5) | θ value | 2.06002e-20 (rank : 7) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O43463, Q53G60, Q6FHK6 | Gene names | SUV39H1, SUV39H | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1). | |||||
|
SUV91_MOUSE
|
||||||
| NC score | 0.695477 (rank : 6) | θ value | 2.06002e-20 (rank : 8) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O54864, Q9JLC7, Q9JLP8 | Gene names | Suv39h1, Suv39h | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1) (Position-effect variegation 3-9 homolog). | |||||
|
SUV92_HUMAN
|
||||||
| NC score | 0.691368 (rank : 7) | θ value | 3.88503e-19 (rank : 13) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9H5I1 | Gene names | SUV39H2 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
SUV92_MOUSE
|
||||||
| NC score | 0.685381 (rank : 8) | θ value | 2.97466e-19 (rank : 12) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9EQQ0, Q9CUK3, Q9JLP7 | Gene names | Suv39h2 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
SETD2_HUMAN
|
||||||
| NC score | 0.656445 (rank : 9) | θ value | 2.69047e-20 (rank : 10) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.614956 (rank : 10) | θ value | 3.51386e-20 (rank : 11) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
SETD8_HUMAN
|
||||||
| NC score | 0.612180 (rank : 11) | θ value | 3.29651e-10 (rank : 21) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
SETD8_MOUSE
|
||||||
| NC score | 0.611189 (rank : 12) | θ value | 3.29651e-10 (rank : 22) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.568112 (rank : 13) | θ value | 2.69047e-20 (rank : 9) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
NSD1_MOUSE
|
||||||
| NC score | 0.565590 (rank : 14) | θ value | 2.06002e-20 (rank : 6) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.558268 (rank : 15) | θ value | 3.17815e-21 (rank : 5) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.539904 (rank : 16) | θ value | 5.07402e-19 (rank : 14) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
SETB2_HUMAN
|
||||||
| NC score | 0.504110 (rank : 17) | θ value | 5.44631e-05 (rank : 25) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96T68, Q86UD6, Q96AI6 | Gene names | SETDB2, CLLD8 | |||
|
Domain Architecture |

|
|||||
| Description | Probable histone-lysine N-methyltransferase, H3 lysine-9 specific (EC 2.1.1.43) (Histone H3-K9 methyltransferase) (H3-K9-HMTase) (SET domain bifurcated 2) (Chronic lymphocytic leukemia deletion region gene 8 protein). | |||||
|
SETB1_MOUSE
|
||||||
| NC score | 0.469258 (rank : 18) | θ value | 6.43352e-06 (rank : 23) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O88974, Q922K1 | Gene names | Setdb1, Eset | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
SETB1_HUMAN
|
||||||
| NC score | 0.464149 (rank : 19) | θ value | 1.09739e-05 (rank : 24) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q15047, Q96GM9 | Gene names | SETDB1, KIAA0067 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.436981 (rank : 20) | θ value | 8.65492e-19 (rank : 16) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.424426 (rank : 21) | θ value | 2.5182e-18 (rank : 17) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.415554 (rank : 22) | θ value | 8.65492e-19 (rank : 15) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
EHMT2_HUMAN
|
||||||
| NC score | 0.344295 (rank : 23) | θ value | 1.63225e-17 (rank : 19) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q96KQ7, Q14349, Q6PK06, Q96MH5, Q96QD0, Q9UQL8, Q9Y331 | Gene names | EHMT2, BAT8, G9A, NG36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
EHMT2_MOUSE
|
||||||
| NC score | 0.341998 (rank : 24) | θ value | 7.32683e-18 (rank : 18) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
EHMT1_HUMAN
|
||||||
| NC score | 0.333982 (rank : 25) | θ value | 2.60593e-15 (rank : 20) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9H9B1, Q8TCN7, Q96F53, Q96JF1, Q96KH4 | Gene names | EHMT1, EUHMTASE1, KIAA1876 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 5 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 5) (H3-K9-HMTase 5) (Euchromatic histone-lysine N-methyltransferase 1) (Eu-HMTase1) (G9a- like protein 1) (GLP1). | |||||
|
SETD7_MOUSE
|
||||||
| NC score | 0.196432 (rank : 26) | θ value | 0.0113563 (rank : 26) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8VHL1 | Gene names | Setd7, Set7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7). | |||||
|
SETD7_HUMAN
|
||||||
| NC score | 0.190698 (rank : 27) | θ value | 0.0252991 (rank : 27) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WTS6, Q9C0E6 | Gene names | SETD7, KIAA1717, SET7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7) (Set9) (SET7/9). | |||||
|
MCSP_MOUSE
|
||||||
| NC score | 0.160192 (rank : 28) | θ value | 0.125558 (rank : 28) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P15265, O70613, Q6P8N3 | Gene names | Smcp, Mcs, Mcsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm mitochondrial-associated cysteine-rich protein. | |||||
|
PTMA_HUMAN
|
||||||
| NC score | 0.114519 (rank : 29) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
PHF11_HUMAN
|
||||||
| NC score | 0.114498 (rank : 30) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UIL8, Q9Y5A2 | Gene names | PHF11, BCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 11 (BRCA1 C-terminus-associated protein) (NY-REN-34 antigen). | |||||
|
KRA58_HUMAN
|
||||||
| NC score | 0.093920 (rank : 31) | θ value | 0.21417 (rank : 29) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75690, Q6L8G7, Q6UTX6 | Gene names | KRTAP5-8, KAP5.8, KRTAP5.8, UHSKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-8 (Keratin-associated protein 5.8) (Ultrahigh sulfur keratin-associated protein 5.8) (Keratin, ultra high-sulfur matrix protein B) (UHS keratin B) (UHS KerB). | |||||
|
KR511_HUMAN
|
||||||
| NC score | 0.088388 (rank : 32) | θ value | 0.47712 (rank : 33) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6L8G4 | Gene names | KRTAP5-11, KAP5.11, KRTAP5.11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-11 (Keratin-associated protein 5.11) (Ultrahigh sulfur keratin-associated protein 5.11). | |||||
|
PHF6_HUMAN
|
||||||
| NC score | 0.087525 (rank : 33) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IWS0, Q96JK3, Q9BRU0 | Gene names | PHF6, KIAA1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6 (PHD-like zinc finger protein). | |||||
|
RAI1_MOUSE
|
||||||
| NC score | 0.085103 (rank : 34) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61818, Q6ZPH7, Q8CEV1 | Gene names | Rai1, Kiaa1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
PHF6_MOUSE
|
||||||
| NC score | 0.084980 (rank : 35) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D4J7, Q80T86, Q8BYT8, Q8C1B5 | Gene names | Phf6, Kiaa1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6. | |||||
|
PHF10_HUMAN
|
||||||
| NC score | 0.084335 (rank : 36) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WUB8, Q2YDA3, Q9BXD2, Q9NV26 | Gene names | PHF10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10 (XAP135). | |||||
|
PHF10_MOUSE
|
||||||
| NC score | 0.082816 (rank : 37) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
PTMA_MOUSE
|
||||||
| NC score | 0.080314 (rank : 38) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
TCF20_HUMAN
|
||||||
| NC score | 0.078690 (rank : 39) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UGU0, O14528, Q13078, Q4V353, Q9H4M0 | Gene names | TCF20, KIAA0292, SPBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP) (AR1). | |||||
|
RAI1_HUMAN
|
||||||
| NC score | 0.078618 (rank : 40) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z5J4, Q8N3B4, Q8ND08, Q8WU64, Q96JK5, Q9H1C1, Q9H1C2, Q9UF69 | Gene names | RAI1, KIAA1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
KRA53_HUMAN
|
||||||
| NC score | 0.076953 (rank : 41) | θ value | 0.62314 (rank : 37) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6L8H2, Q6PL44, Q701N3 | Gene names | KRTAP5-3, KAP5-9, KAP5.3, KRTAP5-9, KRTAP5.3, KRTAP5.9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-3 (Keratin-associated protein 5.3) (Ultrahigh sulfur keratin-associated protein 5.3) (Keratin-associated protein 5-9) (Keratin-associated protein 5.9) (UHS KerB-like). | |||||
|
DPF1_HUMAN
|
||||||
| NC score | 0.074748 (rank : 42) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
TCF20_MOUSE
|
||||||
| NC score | 0.074307 (rank : 43) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
PHF7_HUMAN
|
||||||
| NC score | 0.071178 (rank : 44) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BWX1 | Gene names | PHF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 7 (Testis development protein NYD-SP6). | |||||
|
KRA57_HUMAN
|
||||||
| NC score | 0.067761 (rank : 45) | θ value | 0.62314 (rank : 38) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6L8G8, Q701N5 | Gene names | KRTAP5-7, KAP5-7, KAP5.3, KRTAP5.3, KRTAP5.7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-7 (Keratin-associated protein 5.7) (Ultrahigh sulfur keratin-associated protein 5.7) (Keratin-associated protein 5-3) (Keratin-associated protein 5.3). | |||||
|
KRA24_HUMAN
|
||||||
| NC score | 0.066491 (rank : 46) | θ value | 0.47712 (rank : 34) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9BYR9, Q495J2 | Gene names | KRTAP2-4, KAP2.4, KRTAP2.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 2-4 (Keratin-associated protein 2.4) (High sulfur keratin-associated protein 2.4). | |||||
|
PHF7_MOUSE
|
||||||
| NC score | 0.066462 (rank : 47) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DAG9, Q6PG81 | Gene names | Phf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 7 (Testis development protein NYD-SP6). | |||||
|
AIRE_MOUSE
|
||||||
| NC score | 0.066341 (rank : 48) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
DPF1_MOUSE
|
||||||
| NC score | 0.066159 (rank : 49) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QX66, Q9QX65, Q9QYA4 | Gene names | Dpf1, Neud4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
MYST4_HUMAN
|
||||||
| NC score | 0.065687 (rank : 50) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
REQU_HUMAN
|
||||||
| NC score | 0.065408 (rank : 51) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 523 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92785 | Gene names | DPF2, REQ, UBID4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
K1333_MOUSE
|
||||||
| NC score | 0.065221 (rank : 52) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5RJY2, Q6ZPT7, Q8BNA4 | Gene names | Kiaa1333 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1333 (EC 6.3.2.-). | |||||
|
K1333_HUMAN
|
||||||
| NC score | 0.065162 (rank : 53) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7L622, Q9BVR2, Q9H9E9, Q9NXC0, Q9P2L3 | Gene names | KIAA1333 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1333 (EC 6.3.2.-). | |||||
|
REQU_MOUSE
|
||||||
| NC score | 0.064933 (rank : 54) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61103, Q3UNP5, Q60663, Q9QYA3 | Gene names | Dpf2, Req, Ubid4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
AIRE_HUMAN
|
||||||
| NC score | 0.064092 (rank : 55) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
SMYD3_HUMAN
|
||||||
| NC score | 0.062266 (rank : 56) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H7B4, Q86TL8, Q8N5Z6, Q96AI5 | Gene names | SMYD3, ZMYND1, ZNFN3A1 | |||
|
Domain Architecture |

|
|||||
| Description | SET and MYND domain-containing protein 3 (EC 2.1.1.43) (Zinc finger MYND domain-containing protein 1). | |||||
|
SMYD3_MOUSE
|
||||||
| NC score | 0.060042 (rank : 57) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9CWR2, Q6P7V6, Q8BG90 | Gene names | Smyd3, Zmynd1 | |||
|
Domain Architecture |

|
|||||
| Description | SET and MYND domain-containing protein 3 (EC 2.1.1.43) (Zinc finger MYND domain-containing protein 1). | |||||
|
KRA59_HUMAN
|
||||||
| NC score | 0.059906 (rank : 58) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
CBX3_MOUSE
|
||||||
| NC score | 0.059807 (rank : 59) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23198, Q921C4 | Gene names | Cbx3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (M32). | |||||
|
PF21A_HUMAN
|
||||||
| NC score | 0.059593 (rank : 60) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
PF21A_MOUSE
|
||||||
| NC score | 0.059281 (rank : 61) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
MYST4_MOUSE
|
||||||
| NC score | 0.059233 (rank : 62) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
CBX3_HUMAN
|
||||||
| NC score | 0.059222 (rank : 63) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13185, Q96CD7, Q99409, Q9BVS3, Q9P0Z6 | Gene names | CBX3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (HECH). | |||||
|
CBX1_HUMAN
|
||||||
| NC score | 0.057709 (rank : 64) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P83916, P23197 | Gene names | CBX1, CBX | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25) (HP1Hsbeta) (p25beta). | |||||
|
CBX1_MOUSE
|
||||||
| NC score | 0.057709 (rank : 65) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P83917, P23197 | Gene names | Cbx1, Cbx | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25). | |||||
|
MYST3_MOUSE
|
||||||
| NC score | 0.057004 (rank : 66) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
PF21B_MOUSE
|
||||||
| NC score | 0.055847 (rank : 67) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.055510 (rank : 68) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
LCE1D_HUMAN
|
||||||
| NC score | 0.055227 (rank : 69) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5T752 | Gene names | LCE1D, LEP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Late cornified envelope protein 1D (Late envelope protein 4). | |||||
|
PHF12_MOUSE
|
||||||
| NC score | 0.054828 (rank : 70) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
CBX5_HUMAN
|
||||||
| NC score | 0.054797 (rank : 71) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P45973 | Gene names | CBX5, HP1A | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha) (Antigen p25). | |||||
|
PHF12_HUMAN
|
||||||
| NC score | 0.054778 (rank : 72) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
UHRF1_MOUSE
|
||||||
| NC score | 0.054607 (rank : 73) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VDF2, Q8C6F1, Q8VIA1, Q9Z1H6 | Gene names | Uhrf1, Np95 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Nuclear zinc finger protein Np95) (Nuclear protein 95). | |||||
|
CBX5_MOUSE
|
||||||
| NC score | 0.054493 (rank : 74) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61686, Q9CS35, Q9CXD1 | Gene names | Cbx5, Hp1a | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha). | |||||
|
LCE1E_HUMAN
|
||||||
| NC score | 0.054141 (rank : 75) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5T753 | Gene names | LCE1E, LEP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Late cornified envelope protein 1E (Late envelope protein 5). | |||||
|
UHRF1_HUMAN
|
||||||
| NC score | 0.051107 (rank : 76) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96T88, Q8J022, Q9H6S6, Q9P115, Q9P1U7 | Gene names | UHRF1, ICBP90, NP95, RNF106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Inverted CCAAT box-binding protein of 90 kDa) (Transcription factor ICBP90) (Nuclear zinc finger protein Np95) (Nuclear protein 95) (HuNp95) (RING finger protein 106). | |||||
|
PYGO2_HUMAN
|
||||||
| NC score | 0.050458 (rank : 77) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BRQ0, Q8WYZ4, Q96CY2 | Gene names | PYGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 2. | |||||
|
PF21B_HUMAN
|
||||||
| NC score | 0.050131 (rank : 78) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
LR37A_HUMAN
|
||||||
| NC score | 0.049430 (rank : 79) | θ value | 0.21417 (rank : 30) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60309, Q49A01, Q49A80, Q8NB33 | Gene names | LRRC37A, KIAA0563 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37A. | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.047885 (rank : 80) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
MT2_MOUSE
|
||||||
| NC score | 0.047640 (rank : 81) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P02798 | Gene names | Mt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-2 (MT-2) (Metallothionein-II) (MT-II). | |||||
|
ZNFX1_MOUSE
|
||||||
| NC score | 0.047446 (rank : 82) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8R151, Q3U016, Q3UD71, Q3UEZ0 | Gene names | ZNFX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
ZNFX1_HUMAN
|
||||||
| NC score | 0.045981 (rank : 83) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.045007 (rank : 84) | θ value | 0.62314 (rank : 39) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.041615 (rank : 85) | θ value | 1.81305 (rank : 48) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
KR105_HUMAN
|
||||||
| NC score | 0.041224 (rank : 86) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P60370, Q70LJ3 | Gene names | KRTAP10-5, KAP10.5, KAP18-5, KRTAP10.5, KRTAP18-5, KRTAP18.5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 10-5 (Keratin-associated protein 10.5) (High sulfur keratin-associated protein 10.5) (Keratin-associated protein 18-5) (Keratin-associated protein 18.5). | |||||
|
NOTC2_HUMAN
|
||||||
| NC score | 0.040558 (rank : 87) | θ value | 0.365318 (rank : 31) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q04721, Q99734, Q9H240 | Gene names | NOTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (hN2) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
RCOR2_MOUSE
|
||||||
| NC score | 0.037295 (rank : 88) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C796, Q9JMK4 | Gene names | Rcor2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2 (M-CoREST). | |||||
|
ZAN_MOUSE
|
||||||
| NC score | 0.036993 (rank : 89) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
ITB1_HUMAN
|
||||||
| NC score | 0.036893 (rank : 90) | θ value | 0.62314 (rank : 35) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P05556, P78466, P78467, Q13089, Q13090, Q13091, Q13212, Q14622, Q14647 | Gene names | ITGB1, FNRB | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
NOTC2_MOUSE
|
||||||
| NC score | 0.036583 (rank : 91) | θ value | 1.06291 (rank : 43) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1069 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O35516, Q06008, Q60941 | Gene names | Notch2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (Motch B) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
CENB5_HUMAN
|
||||||
| NC score | 0.035757 (rank : 92) | θ value | 0.47712 (rank : 32) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96P50, Q9BSR9, Q9C0E7 | Gene names | CENTB5, KIAA1716 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 5 (Cnt-b5). | |||||
|
RCOR2_HUMAN
|
||||||
| NC score | 0.032372 (rank : 93) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IZ40, Q96FP3 | Gene names | RCOR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2. | |||||
|
NOTC1_HUMAN
|
||||||
| NC score | 0.031944 (rank : 94) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P46531 | Gene names | NOTCH1, TAN1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 1 precursor (Notch 1) (hN1) (Translocation-associated notch protein TAN-1) [Contains: Notch 1 extracellular truncation; Notch 1 intracellular domain]. | |||||
|
SSPO_MOUSE
|
||||||
| NC score | 0.031244 (rank : 95) | θ value | 1.38821 (rank : 44) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
KRA52_HUMAN
|
||||||
| NC score | 0.031094 (rank : 96) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q701N4 | Gene names | KRTAP5-2, KAP5-8, KAP5.2, KRTAP5-8, KRTAP5.2, KRTAP5.8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-2 (Keratin-associated protein 5.2) (Ultrahigh sulfur keratin-associated protein 5.2) (Keratin-associated protein 5-8) (Keratin-associated protein 5.8). | |||||
|
MYSM1_HUMAN
|
||||||
| NC score | 0.030674 (rank : 97) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5VVJ2, Q7Z3G8, Q96PX3 | Gene names | MYSM1, KIAA1915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MYSM1 (Myb-like, SWIRM and MPN domain-containing protein 1). | |||||
|
CRIM1_HUMAN
|
||||||
| NC score | 0.030425 (rank : 98) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NZV1, Q59GH0, Q7LCQ5, Q9H318 | Gene names | CRIM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich motor neuron 1 protein precursor (CRIM-1) (Cysteine-rich repeat-containing protein S52). | |||||
|
RCOR3_HUMAN
|
||||||
| NC score | 0.030285 (rank : 99) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
ITB1_MOUSE
|
||||||
| NC score | 0.029234 (rank : 100) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P09055 | Gene names | Itgb1 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.029102 (rank : 101) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
VWF_MOUSE
|
||||||
| NC score | 0.028645 (rank : 102) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8CIZ8, Q60863, Q6XUV6, Q8BIU9, Q8CGN0, Q9JK16 | Gene names | Vwf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | von Willebrand factor precursor (vWF) [Contains: von Willebrand antigen 2 (von Willebrand antigen II)]. | |||||
|
MIB1_HUMAN
|
||||||
| NC score | 0.028404 (rank : 103) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86YT6, Q68D01, Q6YI51, Q8NBY0, Q8TCB5, Q8TCL7, Q9P2M3 | Gene names | MIB1, DIP1, KIAA1323, ZZANK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein MIB1 (EC 6.3.2.-) (Mind bomb homolog 1) (DAPK-interacting protein 1) (DIP-1) (Zinc finger ZZ type with ankyrin repeat domain protein 2). | |||||
|
VEGFC_MOUSE
|
||||||
| NC score | 0.027175 (rank : 104) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97953, Q543R6 | Gene names | Vegfc | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor C precursor (VEGF-C) (Vascular endothelial growth factor-related protein) (VRP) (Flt4 ligand) (Flt4- L). | |||||
|
IF4G3_MOUSE
|
||||||
| NC score | 0.025900 (rank : 105) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80XI3, Q80Y69, Q8BJ68, Q8BQF3, Q8C6Q3, Q8CIH0 | Gene names | Eif4g3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
STAB1_HUMAN
|
||||||
| NC score | 0.020825 (rank : 106) | θ value | 1.38821 (rank : 45) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 546 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NY15, Q8IUH0, Q8IUH1, Q93072 | Gene names | STAB1, FEEL1, KIAA0246 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein) (MS-1 antigen). | |||||
|
MEGF6_HUMAN
|
||||||
| NC score | 0.019911 (rank : 107) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
TGON1_MOUSE
|
||||||
| NC score | 0.019717 (rank : 108) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
KIF4A_HUMAN
|
||||||
| NC score | 0.017989 (rank : 109) | θ value | 0.62314 (rank : 36) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O95239, Q86TN3, Q86XX7, Q9NNY6, Q9NY24, Q9UMW3 | Gene names | KIF4A, KIF4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
OXRP_HUMAN
|
||||||
| NC score | 0.017869 (rank : 110) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 544 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y4L1 | Gene names | HYOU1, ORP150 | |||
|
Domain Architecture |

|
|||||
| Description | 150 kDa oxygen-regulated protein precursor (Orp150) (Hypoxia up- regulated 1). | |||||
|
PKHA6_HUMAN
|
||||||
| NC score | 0.017578 (rank : 111) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
DYNA_HUMAN
|
||||||
| NC score | 0.015647 (rank : 112) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1194 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14203, O95296, Q9BRM9, Q9UIU1, Q9UIU2 | Gene names | DCTN1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued) (p135). | |||||
|
DLL1_HUMAN
|
||||||
| NC score | 0.015560 (rank : 113) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00548, Q9NU41, Q9UJV2 | Gene names | DLL1 | |||
|
Domain Architecture |

|
|||||
| Description | Delta-like protein 1 precursor (Drosophila Delta homolog 1) (Delta1) (H-Delta-1). | |||||
|
DYNA_MOUSE
|
||||||
| NC score | 0.013834 (rank : 114) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O08788 | Gene names | Dctn1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued). | |||||
|
ATS13_MOUSE
|
||||||
| NC score | 0.013518 (rank : 115) | θ value | 1.81305 (rank : 47) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q769J6, Q76LW1 | Gene names | Adamts13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-13 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 13) (ADAM-TS 13) (ADAM-TS13) (von Willebrand factor-cleaving protease) (vWF-cleaving protease) (vWF-CP). | |||||
|
ATL2_MOUSE
|
||||||
| NC score | 0.011752 (rank : 116) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TSK7, Q3U0C8 | Gene names | Adamtsl2, Kiaa0605, Tcp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-like protein 2 precursor (ADAMTSL-2) (TSP1-repeats-containing protein 1) (TCP-1). | |||||
|
UB7I4_MOUSE
|
||||||
| NC score | 0.011299 (rank : 117) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q925F3, Q6A0B4 | Gene names | Rnf144, Kiaa0161, Ubce7ip4, Uip4 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-conjugating enzyme 7-interacting protein 4 (UbcM4- interacting protein 4) (RING finger protein 144). | |||||
|
KIF4A_MOUSE
|
||||||
| NC score | 0.010392 (rank : 118) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
UBP25_MOUSE
|
||||||
| NC score | 0.007612 (rank : 119) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P57080, Q80ZT9 | Gene names | Usp25 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (mUSP25). | |||||
|
PGBM_MOUSE
|
||||||
| NC score | 0.007592 (rank : 120) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q05793 | Gene names | Hspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
ALBU_MOUSE
|
||||||
| NC score | 0.007006 (rank : 121) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P07724, Q61802 | Gene names | Alb, Alb-1, Alb1 | |||
|
Domain Architecture |

|
|||||
| Description | Serum albumin precursor. | |||||
|
GP116_HUMAN
|
||||||
| NC score | 0.006947 (rank : 122) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IZF2, O94858, Q6RGN2, Q86SP0, Q9Y3Z2 | Gene names | GPR116, KIAA0758 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 116 precursor. | |||||
|
GPC6A_MOUSE
|
||||||
| NC score | 0.006242 (rank : 123) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K4Z6, Q5DK51, Q5DK52 | Gene names | Gprc6a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor family C group 6 member A precursor. | |||||
|
BAI3_HUMAN
|
||||||
| NC score | 0.005793 (rank : 124) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60242, O60297 | Gene names | BAI3, KIAA0550 | |||
|
Domain Architecture |

|
|||||
| Description | Brain-specific angiogenesis inhibitor 3 precursor. | |||||
|
FGD4_HUMAN
|
||||||
| NC score | 0.005645 (rank : 125) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96M96, Q6ULS2, Q8TCP6 | Gene names | FGD4, FRABP, ZFYVE6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein) (Zinc finger FYVE domain-containing protein 6). | |||||
|
TIE2_MOUSE
|
||||||
| NC score | 0.000123 (rank : 126) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1204 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q02858 | Gene names | Tek, Hyk, Tie-2, Tie2 | |||
|
Domain Architecture |

|
|||||
| Description | Angiopoietin-1 receptor precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor TIE-2) (mTIE2) (Tyrosine-protein kinase receptor TEK) (p140 TEK) (Tunica interna endothelial cell kinase) (HYK) (STK1) (CD202b antigen). | |||||
|
EGFR_HUMAN
|
||||||
| NC score | -0.001354 (rank : 127) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P00533, O00688, O00732, P06268, Q14225, Q92795, Q9BZS2, Q9GZX1, Q9H2C9, Q9H3C9, Q9UMD7, Q9UMD8, Q9UMG5 | Gene names | EGFR, ERBB1 | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor precursor (EC 2.7.10.1) (Receptor tyrosine-protein kinase ErbB-1). | |||||