Please be patient as the page loads
|
STAT6_MOUSE
|
||||||
| SwissProt Accessions | P52633 | Gene names | Stat6 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and transcription activator 6. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
STAT6_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.986300 (rank : 2) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P42226 | Gene names | STAT6 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 6 (IL-4 Stat). | |||||
|
STAT6_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | P52633 | Gene names | Stat6 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and transcription activator 6. | |||||
|
STA5A_MOUSE
|
||||||
| θ value | 3.29472e-127 (rank : 3) | NC score | 0.944703 (rank : 4) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P42230 | Gene names | Stat5a, Mgf, Mpf | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5A (Mammary gland factor). | |||||
|
STA5B_MOUSE
|
||||||
| θ value | 3.64273e-126 (rank : 4) | NC score | 0.944470 (rank : 5) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P42232, Q541Q5, Q60804, Q8K3Q1, Q9JKM1 | Gene names | Stat5b | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5B. | |||||
|
STA5A_HUMAN
|
||||||
| θ value | 6.21357e-126 (rank : 5) | NC score | 0.942533 (rank : 6) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P42229 | Gene names | STAT5A, STAT5 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5A. | |||||
|
STA5B_HUMAN
|
||||||
| θ value | 1.05987e-125 (rank : 6) | NC score | 0.947292 (rank : 3) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P51692, Q8WWS8 | Gene names | STAT5B | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5B. | |||||
|
STAT4_HUMAN
|
||||||
| θ value | 3.48141e-52 (rank : 7) | NC score | 0.865966 (rank : 8) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q14765, Q96NZ6 | Gene names | STAT4 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 4. | |||||
|
STAT4_MOUSE
|
||||||
| θ value | 1.01294e-51 (rank : 8) | NC score | 0.871283 (rank : 7) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P42228 | Gene names | Stat4 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 4. | |||||
|
STAT3_HUMAN
|
||||||
| θ value | 6.35935e-46 (rank : 9) | NC score | 0.853533 (rank : 9) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P40763, O14916, Q9BW54 | Gene names | STAT3, APRF | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 3 (Acute-phase response factor). | |||||
|
STAT3_MOUSE
|
||||||
| θ value | 6.35935e-46 (rank : 10) | NC score | 0.853530 (rank : 10) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P42227 | Gene names | Stat3, Aprf | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 3 (Acute-phase response factor). | |||||
|
STAT1_HUMAN
|
||||||
| θ value | 2.26182e-43 (rank : 11) | NC score | 0.848087 (rank : 11) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P42224 | Gene names | STAT1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 1-alpha/beta (Transcription factor ISGF-3 components p91/p84). | |||||
|
STAT1_MOUSE
|
||||||
| θ value | 4.71619e-41 (rank : 12) | NC score | 0.844391 (rank : 13) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P42225 | Gene names | Stat1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 1. | |||||
|
STAT2_HUMAN
|
||||||
| θ value | 8.32485e-38 (rank : 13) | NC score | 0.847057 (rank : 12) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P52630, Q16430, Q16431 | Gene names | STAT2 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 2 (p113). | |||||
|
STAT2_MOUSE
|
||||||
| θ value | 7.54701e-31 (rank : 14) | NC score | 0.793713 (rank : 14) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9WVL2, Q64188, Q64189, Q64250 | Gene names | Stat2 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 2. | |||||
|
ARI1A_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 15) | NC score | 0.048422 (rank : 18) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
EP400_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 16) | NC score | 0.030929 (rank : 29) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.043267 (rank : 21) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
ARID2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.055381 (rank : 17) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q68CP9, Q5EB51, Q645I3, Q6ZRY5, Q7Z3I5, Q86T28, Q96SJ6, Q9HCL5 | Gene names | ARID2, KIAA1557 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 2 (ARID domain- containing protein 2) (BRG1-associated factor 200) (BAF200). | |||||
|
LMA2L_HUMAN
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.033081 (rank : 27) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0V9, Q53GV3, Q53S67, Q63HN6, Q8NBQ6, Q9BQ14 | Gene names | LMAN2L, VIPL | |||
|
Domain Architecture |
|
|||||
| Description | VIP36-like protein precursor (Lectin, mannose-binding 2-like) (LMAN2- like protein). | |||||
|
LMA2L_MOUSE
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.033194 (rank : 26) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59481, Q3TZ91 | Gene names | Lman2l, Vipl | |||
|
Domain Architecture |
|
|||||
| Description | VIP36-like protein precursor (Lectin, mannose-binding 2-like) (LMAN2- like protein). | |||||
|
BAT3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.023743 (rank : 38) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46379, O95874, Q5SQ35, Q5SQ36, Q5SQ37, Q5SRP8, Q96SA6, Q9BCN4 | Gene names | BAT3, G3 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3) (Protein G3). | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.013528 (rank : 53) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
EP400_MOUSE
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.026941 (rank : 36) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
P85B_HUMAN
|
||||||
| θ value | 1.38821 (rank : 24) | NC score | 0.042583 (rank : 22) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O00459, Q9UPH9 | Gene names | PIK3R2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit beta (PI3-kinase p85- subunit beta) (PtdIns-3-kinase p85-beta). | |||||
|
P85B_MOUSE
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.043979 (rank : 20) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O08908 | Gene names | Pik3r2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit beta (PI3-kinase p85- subunit beta) (PtdIns-3-kinase p85-beta). | |||||
|
RERE_MOUSE
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.031545 (rank : 28) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
STRC_HUMAN
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.029319 (rank : 32) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7RTU9 | Gene names | STRC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stereocilin precursor. | |||||
|
TEKT2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.015329 (rank : 48) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q922G7, Q9WVR0 | Gene names | Tekt2 | |||
|
Domain Architecture |

|
|||||
| Description | Tektin-2 (Testicular tektin) (Tektin-t). | |||||
|
EGR4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.000384 (rank : 78) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q05215 | Gene names | EGR4 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 4 (EGR-4) (AT133). | |||||
|
EP300_HUMAN
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.027737 (rank : 34) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
PHF20_HUMAN
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.019918 (rank : 40) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
POLH_HUMAN
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.017178 (rank : 45) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y253, Q7L8E3, Q96BC4, Q9BX13 | Gene names | POLH, RAD30, RAD30A, XPV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase eta (EC 2.7.7.7) (RAD30 homolog A) (Xeroderma pigmentosum variant type protein). | |||||
|
ZO1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.014854 (rank : 49) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q07157, Q4ZGJ6 | Gene names | TJP1, ZO1 | |||
|
Domain Architecture |

|
|||||
| Description | Tight junction protein ZO-1 (Zonula occludens 1 protein) (Zona occludens 1 protein) (Tight junction protein 1). | |||||
|
ZO1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.014304 (rank : 50) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P39447 | Gene names | Tjp1, Zo1 | |||
|
Domain Architecture |

|
|||||
| Description | Tight junction protein ZO-1 (Zonula occludens 1 protein) (Zona occludens 1 protein) (Tight junction protein 1). | |||||
|
DOT1L_HUMAN
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.019599 (rank : 42) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
ENAH_MOUSE
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.010772 (rank : 60) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
MA1B1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.013759 (rank : 52) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UKM7, Q5VSG3, Q9BRS9, Q9Y5K7 | Gene names | MAN1B1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase (EC 3.2.1.113) (ER alpha-1,2-mannosidase) (Mannosidase alpha class 1B member 1) (Man9GlcNAc2-specific-processing alpha-mannosidase). | |||||
|
PHLA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.030212 (rank : 30) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WV24, Q15184, Q2TAN2, Q9NZ17 | Gene names | PHLDA1, PHRIP, TDAG51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Apoptosis-associated nuclear protein) (Proline- and histidine-rich protein) (Proline- and glutamine-rich protein) (PQ-rich protein). | |||||
|
R3HD2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.029745 (rank : 31) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
RIN3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.013915 (rank : 51) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P59729 | Gene names | Rin3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
WASF3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.028758 (rank : 33) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UPY6, O94974 | Gene names | WASF3, KIAA0900, SCAR3, WAVE3 | |||
|
Domain Architecture |

|
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3) (Verprolin homology domain-containing protein 3). | |||||
|
AMPH_HUMAN
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.011791 (rank : 57) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49418, O43538, Q75MJ8, Q75MK5, Q75MM3, Q8N4G0 | Gene names | AMPH, AMPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Amphiphysin. | |||||
|
HM20A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.012727 (rank : 55) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DC33, Q3LSF9, Q6NV87, Q8BSK1, Q8C3C1, Q8CAA0, Q9CYG2 | Gene names | Hmg20a, Ibraf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (iBRAF) (Inhibitor of BRAF35) (HMG domain protein HMGX1). | |||||
|
PYGO2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.019880 (rank : 41) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BRQ0, Q8WYZ4, Q96CY2 | Gene names | PYGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 2. | |||||
|
RERE_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.027208 (rank : 35) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
ZNFX1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.008955 (rank : 63) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
CECR2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.017204 (rank : 44) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9BXF3, Q658Z4, Q96P58, Q9C0C3 | Gene names | CECR2, KIAA1740 | |||
|
Domain Architecture |

|
|||||
| Description | Cat eye syndrome critical region protein 2. | |||||
|
FYCO1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.011200 (rank : 58) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 1149 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BQS8, Q3MJE6, Q86T41, Q86TB1, Q8TEF9, Q96IV5, Q9H8P9 | Gene names | FYCO1, ZFYVE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1 (Zinc finger FYVE domain-containing protein 7). | |||||
|
HM20A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.012091 (rank : 56) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NP66, Q53G31, Q9NSF6 | Gene names | HMG20A, HMGX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (HMG domain-containing protein HMGX1). | |||||
|
KIF1B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.004532 (rank : 74) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60333, Q96Q94, Q9BV80, Q9P280 | Gene names | KIF1B, KIAA0591, KIAA1448 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF1B (Klp). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.033687 (rank : 25) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.017428 (rank : 43) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
K1802_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.007310 (rank : 67) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
PHC3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.015823 (rank : 46) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CHP6, Q3TLV5 | Gene names | Phc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3. | |||||
|
PHF20_MOUSE
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.015464 (rank : 47) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
TCRG1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.006276 (rank : 70) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.024050 (rank : 37) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
CD6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.005807 (rank : 71) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61003, Q60679, Q61004 | Gene names | Cd6 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell differentiation antigen CD6 precursor. | |||||
|
HOMEZ_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.009351 (rank : 61) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IX15, Q6P049, Q86XB6, Q9P2A5 | Gene names | HOMEZ, KIAA1443 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox and leucine zipper protein Homez (Homeodomain leucine zipper- containing factor). | |||||
|
HSH2D_MOUSE
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.060393 (rank : 16) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6VYH9, Q52KL3 | Gene names | Hsh2d, Alx | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hematopoietic SH2 domain-containing protein (Hematopoietic SH2 protein) (Adaptor in lymphocytes of unknown function X). | |||||
|
M3K14_MOUSE
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | -0.001351 (rank : 79) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9WUL6 | Gene names | Map3k14, Nik | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 14 (EC 2.7.11.25) (NF- kappa beta-inducing kinase) (Serine/threonine-protein kinase NIK). | |||||
|
MYST4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.006538 (rank : 69) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
PDC6I_MOUSE
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.008392 (rank : 64) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WU78, O88695, O89014, Q8BSL8, Q8R0H5, Q99LR3, Q9QZN8 | Gene names | Pdcd6ip, Aip1, Alix | |||
|
Domain Architecture |
|
|||||
| Description | Programmed cell death 6-interacting protein (ALG-2-interacting protein X) (ALG-2-interacting protein 1) (E2F1-inducible protein) (Eig2). | |||||
|
SMCA4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.011048 (rank : 59) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
2A5D_HUMAN
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.004515 (rank : 75) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14738, O00494, O00696, Q15171 | Gene names | PPP2R5D | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit delta isoform (PP2A, B subunit, B' delta isoform) (PP2A, B subunit, B56 delta isoform) (PP2A, B subunit, PR61 delta isoform) (PP2A, B subunit, R5 delta isoform). | |||||
|
AUTS2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.033997 (rank : 24) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8WXX7, Q9Y4F2 | Gene names | AUTS2, KIAA0442 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Autism susceptibility gene 2 protein. | |||||
|
AUXI_HUMAN
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.005767 (rank : 72) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75061, Q32M66, Q4G0K1, Q5T614, Q5T615 | Gene names | DNAJC6, KIAA0473 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative tyrosine-protein phosphatase auxilin (EC 3.1.3.48) (DnaJ homolog subfamily C member 6). | |||||
|
HNRL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.007378 (rank : 66) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VDM6, Q3U201, Q3UPB0, Q6AZA7, Q8BY45, Q8K365 | Gene names | Hnrpul1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U-like protein 1. | |||||
|
KI13B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.004554 (rank : 73) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQT8, O75134, Q9BYJ6 | Gene names | KIF13B, GAKIN, KIAA0639 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13B (Kinesin-like protein GAKIN). | |||||
|
LR37B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.004164 (rank : 76) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96QE4, Q5YKG6 | Gene names | LRRC37B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37B precursor (C66 SLIT-like testicular protein). | |||||
|
MAML1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.009078 (rank : 62) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92585, Q9NZ12 | Gene names | MAML1, KIAA0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.008061 (rank : 65) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
P55G_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.042386 (rank : 23) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92569, O60482 | Gene names | PIK3R3 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit gamma (PI3-kinase p85-subunit gamma) (PtdIns-3-kinase p85-gamma) (p55PIK). | |||||
|
P55G_MOUSE
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.047900 (rank : 19) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q64143 | Gene names | Pik3r3 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit gamma (PI3-kinase p85-subunit gamma) (PtdIns-3-kinase p85-gamma) (p55PIK). | |||||
|
POLS_MOUSE
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.007144 (rank : 68) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6PB75 | Gene names | Pols | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase sigma (EC 2.7.7.7). | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.021635 (rank : 39) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
WASF3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.013259 (rank : 54) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VHI6 | Gene names | Wasf3, Wave3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3). | |||||
|
WDR33_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.002690 (rank : 77) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 638 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9C0J8 | Gene names | WDR33, WDC146 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 33 (WD repeat protein WDC146). | |||||
|
SH22A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.081846 (rank : 15) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NP31, O43817, Q9UPA7 | Gene names | SH2D2A, SCAP, TSAD, VRAP | |||
|
Domain Architecture |
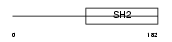
|
|||||
| Description | SH2 domain protein 2A (T cell-specific adapter protein) (TSAd) (VEGF receptor-associated protein) (SH2 domain-containing adapter protein). | |||||
|
STAT6_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | P52633 | Gene names | Stat6 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and transcription activator 6. | |||||
|
STAT6_HUMAN
|
||||||
| NC score | 0.986300 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P42226 | Gene names | STAT6 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 6 (IL-4 Stat). | |||||
|
STA5B_HUMAN
|
||||||
| NC score | 0.947292 (rank : 3) | θ value | 1.05987e-125 (rank : 6) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P51692, Q8WWS8 | Gene names | STAT5B | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5B. | |||||
|
STA5A_MOUSE
|
||||||
| NC score | 0.944703 (rank : 4) | θ value | 3.29472e-127 (rank : 3) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P42230 | Gene names | Stat5a, Mgf, Mpf | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5A (Mammary gland factor). | |||||
|
STA5B_MOUSE
|
||||||
| NC score | 0.944470 (rank : 5) | θ value | 3.64273e-126 (rank : 4) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P42232, Q541Q5, Q60804, Q8K3Q1, Q9JKM1 | Gene names | Stat5b | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5B. | |||||
|
STA5A_HUMAN
|
||||||
| NC score | 0.942533 (rank : 6) | θ value | 6.21357e-126 (rank : 5) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P42229 | Gene names | STAT5A, STAT5 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5A. | |||||
|
STAT4_MOUSE
|
||||||
| NC score | 0.871283 (rank : 7) | θ value | 1.01294e-51 (rank : 8) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P42228 | Gene names | Stat4 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 4. | |||||
|
STAT4_HUMAN
|
||||||
| NC score | 0.865966 (rank : 8) | θ value | 3.48141e-52 (rank : 7) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q14765, Q96NZ6 | Gene names | STAT4 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 4. | |||||
|
STAT3_HUMAN
|
||||||
| NC score | 0.853533 (rank : 9) | θ value | 6.35935e-46 (rank : 9) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P40763, O14916, Q9BW54 | Gene names | STAT3, APRF | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 3 (Acute-phase response factor). | |||||
|
STAT3_MOUSE
|
||||||
| NC score | 0.853530 (rank : 10) | θ value | 6.35935e-46 (rank : 10) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P42227 | Gene names | Stat3, Aprf | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 3 (Acute-phase response factor). | |||||
|
STAT1_HUMAN
|
||||||
| NC score | 0.848087 (rank : 11) | θ value | 2.26182e-43 (rank : 11) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P42224 | Gene names | STAT1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 1-alpha/beta (Transcription factor ISGF-3 components p91/p84). | |||||
|
STAT2_HUMAN
|
||||||
| NC score | 0.847057 (rank : 12) | θ value | 8.32485e-38 (rank : 13) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P52630, Q16430, Q16431 | Gene names | STAT2 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 2 (p113). | |||||
|
STAT1_MOUSE
|
||||||
| NC score | 0.844391 (rank : 13) | θ value | 4.71619e-41 (rank : 12) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P42225 | Gene names | Stat1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 1. | |||||
|
STAT2_MOUSE
|
||||||
| NC score | 0.793713 (rank : 14) | θ value | 7.54701e-31 (rank : 14) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9WVL2, Q64188, Q64189, Q64250 | Gene names | Stat2 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 2. | |||||
|
SH22A_HUMAN
|
||||||
| NC score | 0.081846 (rank : 15) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NP31, O43817, Q9UPA7 | Gene names | SH2D2A, SCAP, TSAD, VRAP | |||
|
Domain Architecture |

|
|||||
| Description | SH2 domain protein 2A (T cell-specific adapter protein) (TSAd) (VEGF receptor-associated protein) (SH2 domain-containing adapter protein). | |||||
|
HSH2D_MOUSE
|
||||||
| NC score | 0.060393 (rank : 16) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6VYH9, Q52KL3 | Gene names | Hsh2d, Alx | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hematopoietic SH2 domain-containing protein (Hematopoietic SH2 protein) (Adaptor in lymphocytes of unknown function X). | |||||
|
ARID2_HUMAN
|
||||||
| NC score | 0.055381 (rank : 17) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q68CP9, Q5EB51, Q645I3, Q6ZRY5, Q7Z3I5, Q86T28, Q96SJ6, Q9HCL5 | Gene names | ARID2, KIAA1557 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 2 (ARID domain- containing protein 2) (BRG1-associated factor 200) (BAF200). | |||||
|
ARI1A_HUMAN
|
||||||
| NC score | 0.048422 (rank : 18) | θ value | 0.0252991 (rank : 15) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
P55G_MOUSE
|
||||||
| NC score | 0.047900 (rank : 19) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q64143 | Gene names | Pik3r3 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit gamma (PI3-kinase p85-subunit gamma) (PtdIns-3-kinase p85-gamma) (p55PIK). | |||||
|
P85B_MOUSE
|
||||||
| NC score | 0.043979 (rank : 20) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O08908 | Gene names | Pik3r2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit beta (PI3-kinase p85- subunit beta) (PtdIns-3-kinase p85-beta). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.043267 (rank : 21) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
P85B_HUMAN
|
||||||
| NC score | 0.042583 (rank : 22) | θ value | 1.38821 (rank : 24) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O00459, Q9UPH9 | Gene names | PIK3R2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit beta (PI3-kinase p85- subunit beta) (PtdIns-3-kinase p85-beta). | |||||
|
P55G_HUMAN
|
||||||
| NC score | 0.042386 (rank : 23) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92569, O60482 | Gene names | PIK3R3 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit gamma (PI3-kinase p85-subunit gamma) (PtdIns-3-kinase p85-gamma) (p55PIK). | |||||
|
AUTS2_HUMAN
|
||||||
| NC score | 0.033997 (rank : 24) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8WXX7, Q9Y4F2 | Gene names | AUTS2, KIAA0442 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Autism susceptibility gene 2 protein. | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.033687 (rank : 25) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
LMA2L_MOUSE
|
||||||
| NC score | 0.033194 (rank : 26) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59481, Q3TZ91 | Gene names | Lman2l, Vipl | |||
|
Domain Architecture |

|
|||||
| Description | VIP36-like protein precursor (Lectin, mannose-binding 2-like) (LMAN2- like protein). | |||||
|
LMA2L_HUMAN
|
||||||
| NC score | 0.033081 (rank : 27) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0V9, Q53GV3, Q53S67, Q63HN6, Q8NBQ6, Q9BQ14 | Gene names | LMAN2L, VIPL | |||
|
Domain Architecture |

|
|||||
| Description | VIP36-like protein precursor (Lectin, mannose-binding 2-like) (LMAN2- like protein). | |||||
|
RERE_MOUSE
|
||||||
| NC score | 0.031545 (rank : 28) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
EP400_HUMAN
|
||||||
| NC score | 0.030929 (rank : 29) | θ value | 0.0563607 (rank : 16) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
PHLA1_HUMAN
|
||||||
| NC score | 0.030212 (rank : 30) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WV24, Q15184, Q2TAN2, Q9NZ17 | Gene names | PHLDA1, PHRIP, TDAG51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Apoptosis-associated nuclear protein) (Proline- and histidine-rich protein) (Proline- and glutamine-rich protein) (PQ-rich protein). | |||||
|
R3HD2_MOUSE
|
||||||
| NC score | 0.029745 (rank : 31) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
STRC_HUMAN
|
||||||
| NC score | 0.029319 (rank : 32) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7RTU9 | Gene names | STRC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stereocilin precursor. | |||||
|
WASF3_HUMAN
|
||||||
| NC score | 0.028758 (rank : 33) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UPY6, O94974 | Gene names | WASF3, KIAA0900, SCAR3, WAVE3 | |||
|
Domain Architecture |

|
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3) (Verprolin homology domain-containing protein 3). | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.027737 (rank : 34) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.027208 (rank : 35) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
EP400_MOUSE
|
||||||
| NC score | 0.026941 (rank : 36) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.024050 (rank : 37) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
BAT3_HUMAN
|
||||||
| NC score | 0.023743 (rank : 38) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46379, O95874, Q5SQ35, Q5SQ36, Q5SQ37, Q5SRP8, Q96SA6, Q9BCN4 | Gene names | BAT3, G3 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3) (Protein G3). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.021635 (rank : 39) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
PHF20_HUMAN
|
||||||
| NC score | 0.019918 (rank : 40) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
PYGO2_HUMAN
|
||||||
| NC score | 0.019880 (rank : 41) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BRQ0, Q8WYZ4, Q96CY2 | Gene names | PYGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 2. | |||||
|
DOT1L_HUMAN
|
||||||
| NC score | 0.019599 (rank : 42) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.017428 (rank : 43) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CECR2_HUMAN
|
||||||
| NC score | 0.017204 (rank : 44) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9BXF3, Q658Z4, Q96P58, Q9C0C3 | Gene names | CECR2, KIAA1740 | |||
|
Domain Architecture |

|
|||||
| Description | Cat eye syndrome critical region protein 2. | |||||
|
POLH_HUMAN
|
||||||
| NC score | 0.017178 (rank : 45) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y253, Q7L8E3, Q96BC4, Q9BX13 | Gene names | POLH, RAD30, RAD30A, XPV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase eta (EC 2.7.7.7) (RAD30 homolog A) (Xeroderma pigmentosum variant type protein). | |||||
|
PHC3_MOUSE
|
||||||
| NC score | 0.015823 (rank : 46) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CHP6, Q3TLV5 | Gene names | Phc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3. | |||||
|
PHF20_MOUSE
|
||||||
| NC score | 0.015464 (rank : 47) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
TEKT2_MOUSE
|
||||||
| NC score | 0.015329 (rank : 48) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q922G7, Q9WVR0 | Gene names | Tekt2 | |||
|
Domain Architecture |

|
|||||
| Description | Tektin-2 (Testicular tektin) (Tektin-t). | |||||
|
ZO1_HUMAN
|
||||||
| NC score | 0.014854 (rank : 49) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q07157, Q4ZGJ6 | Gene names | TJP1, ZO1 | |||
|
Domain Architecture |

|
|||||
| Description | Tight junction protein ZO-1 (Zonula occludens 1 protein) (Zona occludens 1 protein) (Tight junction protein 1). | |||||
|
ZO1_MOUSE
|
||||||
| NC score | 0.014304 (rank : 50) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P39447 | Gene names | Tjp1, Zo1 | |||
|
Domain Architecture |

|
|||||
| Description | Tight junction protein ZO-1 (Zonula occludens 1 protein) (Zona occludens 1 protein) (Tight junction protein 1). | |||||
|
RIN3_MOUSE
|
||||||
| NC score | 0.013915 (rank : 51) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P59729 | Gene names | Rin3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
MA1B1_HUMAN
|
||||||
| NC score | 0.013759 (rank : 52) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UKM7, Q5VSG3, Q9BRS9, Q9Y5K7 | Gene names | MAN1B1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase (EC 3.2.1.113) (ER alpha-1,2-mannosidase) (Mannosidase alpha class 1B member 1) (Man9GlcNAc2-specific-processing alpha-mannosidase). | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.013528 (rank : 53) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
WASF3_MOUSE
|
||||||
| NC score | 0.013259 (rank : 54) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VHI6 | Gene names | Wasf3, Wave3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3). | |||||
|
HM20A_MOUSE
|
||||||
| NC score | 0.012727 (rank : 55) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DC33, Q3LSF9, Q6NV87, Q8BSK1, Q8C3C1, Q8CAA0, Q9CYG2 | Gene names | Hmg20a, Ibraf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (iBRAF) (Inhibitor of BRAF35) (HMG domain protein HMGX1). | |||||
|
HM20A_HUMAN
|
||||||
| NC score | 0.012091 (rank : 56) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NP66, Q53G31, Q9NSF6 | Gene names | HMG20A, HMGX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (HMG domain-containing protein HMGX1). | |||||
|
AMPH_HUMAN
|
||||||
| NC score | 0.011791 (rank : 57) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49418, O43538, Q75MJ8, Q75MK5, Q75MM3, Q8N4G0 | Gene names | AMPH, AMPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Amphiphysin. | |||||
|
FYCO1_HUMAN
|
||||||
| NC score | 0.011200 (rank : 58) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 1149 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BQS8, Q3MJE6, Q86T41, Q86TB1, Q8TEF9, Q96IV5, Q9H8P9 | Gene names | FYCO1, ZFYVE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1 (Zinc finger FYVE domain-containing protein 7). | |||||
|
SMCA4_HUMAN
|
||||||
| NC score | 0.011048 (rank : 59) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.010772 (rank : 60) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
HOMEZ_HUMAN
|
||||||
| NC score | 0.009351 (rank : 61) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IX15, Q6P049, Q86XB6, Q9P2A5 | Gene names | HOMEZ, KIAA1443 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox and leucine zipper protein Homez (Homeodomain leucine zipper- containing factor). | |||||
|
MAML1_HUMAN
|
||||||
| NC score | 0.009078 (rank : 62) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92585, Q9NZ12 | Gene names | MAML1, KIAA0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
ZNFX1_HUMAN
|
||||||
| NC score | 0.008955 (rank : 63) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
PDC6I_MOUSE
|
||||||
| NC score | 0.008392 (rank : 64) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WU78, O88695, O89014, Q8BSL8, Q8R0H5, Q99LR3, Q9QZN8 | Gene names | Pdcd6ip, Aip1, Alix | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death 6-interacting protein (ALG-2-interacting protein X) (ALG-2-interacting protein 1) (E2F1-inducible protein) (Eig2). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.008061 (rank : 65) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
HNRL1_MOUSE
|
||||||
| NC score | 0.007378 (rank : 66) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VDM6, Q3U201, Q3UPB0, Q6AZA7, Q8BY45, Q8K365 | Gene names | Hnrpul1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U-like protein 1. | |||||
|
K1802_HUMAN
|
||||||
| NC score | 0.007310 (rank : 67) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
POLS_MOUSE
|
||||||
| NC score | 0.007144 (rank : 68) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6PB75 | Gene names | Pols | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase sigma (EC 2.7.7.7). | |||||
|
MYST4_MOUSE
|
||||||
| NC score | 0.006538 (rank : 69) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
TCRG1_HUMAN
|
||||||
| NC score | 0.006276 (rank : 70) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
CD6_MOUSE
|
||||||
| NC score | 0.005807 (rank : 71) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61003, Q60679, Q61004 | Gene names | Cd6 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell differentiation antigen CD6 precursor. | |||||
|
AUXI_HUMAN
|
||||||
| NC score | 0.005767 (rank : 72) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75061, Q32M66, Q4G0K1, Q5T614, Q5T615 | Gene names | DNAJC6, KIAA0473 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative tyrosine-protein phosphatase auxilin (EC 3.1.3.48) (DnaJ homolog subfamily C member 6). | |||||
|
KI13B_HUMAN
|
||||||
| NC score | 0.004554 (rank : 73) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQT8, O75134, Q9BYJ6 | Gene names | KIF13B, GAKIN, KIAA0639 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13B (Kinesin-like protein GAKIN). | |||||
|
KIF1B_HUMAN
|
||||||
| NC score | 0.004532 (rank : 74) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60333, Q96Q94, Q9BV80, Q9P280 | Gene names | KIF1B, KIAA0591, KIAA1448 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF1B (Klp). | |||||
|
2A5D_HUMAN
|
||||||
| NC score | 0.004515 (rank : 75) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14738, O00494, O00696, Q15171 | Gene names | PPP2R5D | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit delta isoform (PP2A, B subunit, B' delta isoform) (PP2A, B subunit, B56 delta isoform) (PP2A, B subunit, PR61 delta isoform) (PP2A, B subunit, R5 delta isoform). | |||||
|
LR37B_HUMAN
|
||||||
| NC score | 0.004164 (rank : 76) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96QE4, Q5YKG6 | Gene names | LRRC37B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37B precursor (C66 SLIT-like testicular protein). | |||||
|
WDR33_HUMAN
|
||||||
| NC score | 0.002690 (rank : 77) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 638 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9C0J8 | Gene names | WDR33, WDC146 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 33 (WD repeat protein WDC146). | |||||
|
EGR4_HUMAN
|
||||||
| NC score | 0.000384 (rank : 78) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q05215 | Gene names | EGR4 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 4 (EGR-4) (AT133). | |||||
|
M3K14_MOUSE
|
||||||
| NC score | -0.001351 (rank : 79) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 78 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9WUL6 | Gene names | Map3k14, Nik | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 14 (EC 2.7.11.25) (NF- kappa beta-inducing kinase) (Serine/threonine-protein kinase NIK). | |||||