Please be patient as the page loads
|
RTEL1_HUMAN
|
||||||
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
RTEL1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 116 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
DDX11_MOUSE
|
||||||
| θ value | 2.84138e-62 (rank : 2) | NC score | 0.842785 (rank : 2) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6AXC6 | Gene names | Ddx11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX11 (EC 3.6.1.-) (DEAD/H box protein 11). | |||||
|
FANCJ_MOUSE
|
||||||
| θ value | 2.7521e-57 (rank : 3) | NC score | 0.809450 (rank : 6) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5SXJ3, Q8BJQ8, Q8BKI6 | Gene names | Brip1, Bach1, Fancj | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group J protein homolog (EC 3.6.1.-) (ATP-dependent RNA helicase BRIP1) (Protein FACJ) (BRCA1-interacting protein C-terminal helicase 1) (BRCA1-interacting protein 1) (BRCA1-associated C-terminal helicase 1). | |||||
|
FANCJ_HUMAN
|
||||||
| θ value | 1.36585e-56 (rank : 4) | NC score | 0.820228 (rank : 3) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BX63, Q8NCI5 | Gene names | BRIP1, BACH1, FANCJ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group J protein (EC 3.6.1.-) (ATP-dependent RNA helicase BRIP1) (Protein FACJ) (BRCA1-interacting protein C-terminal helicase 1) (BRCA1-interacting protein 1) (BRCA1-associated C-terminal helicase 1). | |||||
|
ERCC2_MOUSE
|
||||||
| θ value | 3.1488e-53 (rank : 5) | NC score | 0.817762 (rank : 4) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08811, Q9DC01 | Gene names | Ercc2, Xpd | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase subunit (EC 3.6.1.-) (DNA-repair protein complementing XP-D cells) (Xeroderma pigmentosum group D-complementing protein) (CXPD) (DNA excision repair protein ERCC-2). | |||||
|
ERCC2_HUMAN
|
||||||
| θ value | 7.75577e-52 (rank : 6) | NC score | 0.815878 (rank : 5) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P18074, Q8N721 | Gene names | ERCC2, XPD, XPDC | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase subunit (EC 3.6.1.-) (DNA-repair protein complementing XP-D cells) (Xeroderma pigmentosum group D-complementing protein) (CXPD) (DNA excision repair protein ERCC-2). | |||||
|
DDX11_HUMAN
|
||||||
| θ value | 2.18568e-46 (rank : 7) | NC score | 0.802851 (rank : 7) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96FC9, Q13333, Q86VQ4, Q86W62, Q92498, Q92770, Q92998, Q92999 | Gene names | DDX11, CHL1, CHLR1, KRG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX11 (EC 3.6.1.-) (DEAD/H box protein 11) (CHL1 homolog) (Keratinocyte growth factor-regulated gene 2 protein) (KRG-2). | |||||
|
RPKL1_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 8) | NC score | 0.025916 (rank : 49) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y6S9, Q69YT9, Q6ZMQ6, Q96NR9, Q9BSU9 | Gene names | RPS6KL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal protein S6 kinase-like 1 (EC 2.7.11.1). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 9) | NC score | 0.042520 (rank : 16) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
AHTF1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 10) | NC score | 0.050543 (rank : 9) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CJF7, Q8BVJ5, Q8VD55 | Gene names | Ahctf1, Elys | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
TP53B_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 11) | NC score | 0.047866 (rank : 12) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 413 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12888, Q2M1Z7, Q4LE46, Q5FWZ3, Q7Z3U4 | Gene names | TP53BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
MOT3_MOUSE
|
||||||
| θ value | 0.125558 (rank : 12) | NC score | 0.031246 (rank : 33) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35308 | Gene names | Slc16a8, Mct3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Monocarboxylate transporter 3 (MCT 3) (Solute carrier family 16 member 8) (Proton-coupled monocarboxylate transporter 3). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 0.163984 (rank : 13) | NC score | 0.040427 (rank : 19) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
IQEC2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 14) | NC score | 0.041745 (rank : 17) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5JU85, O60275 | Gene names | IQSEC2, KIAA0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
IQEC2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 15) | NC score | 0.041314 (rank : 18) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
RBM28_HUMAN
|
||||||
| θ value | 0.163984 (rank : 16) | NC score | 0.028240 (rank : 44) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NW13, Q96CV3 | Gene names | RBM28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 28 (RNA-binding motif protein 28). | |||||
|
APOL5_HUMAN
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.058925 (rank : 8) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BWW9, Q5TFL9, Q9UGW5 | Gene names | APOL5 | |||
|
Domain Architecture |
|
|||||
| Description | Apolipoprotein-L5 (Apolipoprotein L-V) (ApoL-V). | |||||
|
IRX3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 18) | NC score | 0.029664 (rank : 38) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P81067 | Gene names | Irx3, Irxb1 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-3 (Iroquois homeobox protein 3) (Homeodomain protein IRXB1). | |||||
|
SREC2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 19) | NC score | 0.016650 (rank : 68) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 605 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P59222 | Gene names | Scarf2, Srec2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II). | |||||
|
DOK7_HUMAN
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.047951 (rank : 11) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q18PE1, Q6P6A6, Q86XG5, Q8N2J3, Q8NBC1 | Gene names | DOK7, C4orf25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Dok-7 (Downstream of tyrosine kinase 7). | |||||
|
SN1L2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 21) | NC score | 0.004939 (rank : 104) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CFH6 | Gene names | Snf1lk2, Sik2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Salt-inducible kinase 2). | |||||
|
CCNK_HUMAN
|
||||||
| θ value | 0.47712 (rank : 22) | NC score | 0.033847 (rank : 27) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75909, Q96B63, Q9NNY9 | Gene names | CCNK, CPR4 | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-K. | |||||
|
GP152_HUMAN
|
||||||
| θ value | 0.47712 (rank : 23) | NC score | 0.022442 (rank : 55) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
LYAR_MOUSE
|
||||||
| θ value | 0.62314 (rank : 24) | NC score | 0.042699 (rank : 15) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q08288, Q9D9X2 | Gene names | Lyar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth-regulating nucleolar protein. | |||||
|
SRCA_HUMAN
|
||||||
| θ value | 0.62314 (rank : 25) | NC score | 0.029154 (rank : 41) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
CIC_HUMAN
|
||||||
| θ value | 0.813845 (rank : 26) | NC score | 0.025213 (rank : 52) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
FA61A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 27) | NC score | 0.034800 (rank : 24) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K2F8, Q9CTG8 | Gene names | Fam61a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM61A. | |||||
|
K0226_HUMAN
|
||||||
| θ value | 0.813845 (rank : 28) | NC score | 0.048409 (rank : 10) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92622 | Gene names | KIAA0226 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0226. | |||||
|
AMOL1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 29) | NC score | 0.021166 (rank : 57) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
DEND_MOUSE
|
||||||
| θ value | 1.06291 (rank : 30) | NC score | 0.039541 (rank : 21) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80TS7 | Gene names | Ddn, Kiaa0749 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dendrin. | |||||
|
IER5_MOUSE
|
||||||
| θ value | 1.06291 (rank : 31) | NC score | 0.042783 (rank : 14) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O89113, Q8CGI0 | Gene names | Ier5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immediate early response gene 5 protein. | |||||
|
IF4G1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 32) | NC score | 0.035640 (rank : 22) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
KLF13_HUMAN
|
||||||
| θ value | 1.06291 (rank : 33) | NC score | 0.003859 (rank : 109) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2Y9, Q9Y356 | Gene names | KLF13, BTEB3, NSLP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 13 (Transcription factor BTEB3) (Basic transcription element-binding protein 3) (BTE-binding protein 3) (RANTES factor of late activated T-lymphocytes 1) (RFLAT-1) (Transcription factor NSLP1) (Novel Sp1-like zinc finger transcription factor 1). | |||||
|
PHLA1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 34) | NC score | 0.040037 (rank : 20) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WV24, Q15184, Q2TAN2, Q9NZ17 | Gene names | PHLDA1, PHRIP, TDAG51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Apoptosis-associated nuclear protein) (Proline- and histidine-rich protein) (Proline- and glutamine-rich protein) (PQ-rich protein). | |||||
|
APC_HUMAN
|
||||||
| θ value | 1.38821 (rank : 35) | NC score | 0.035241 (rank : 23) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 36) | NC score | 0.030029 (rank : 37) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
K0226_MOUSE
|
||||||
| θ value | 1.38821 (rank : 37) | NC score | 0.046371 (rank : 13) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80U62, Q3TAX3, Q3TDF7, Q3TDU2, Q6P9T7, Q6PG18, Q8BMP7, Q8BY22 | Gene names | Kiaa0226 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0226. | |||||
|
CT128_HUMAN
|
||||||
| θ value | 1.81305 (rank : 38) | NC score | 0.034184 (rank : 26) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BQN1, Q5JWN6, Q8N276 | Gene names | C20orf128 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf128. | |||||
|
FOXC2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 39) | NC score | 0.010544 (rank : 86) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99958 | Gene names | FOXC2, FKHL14, MFH1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
PO3F3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 40) | NC score | 0.017084 (rank : 67) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P20264, P78379 | Gene names | POU3F3, BRN1, OTF8 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 3 (Brain-specific homeobox/POU domain protein 1) (Brain-1) (Brn-1 protein). | |||||
|
PO3F3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.017086 (rank : 66) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P31361 | Gene names | Pou3f3, Brn-1, Brn1, Otf8 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 3 (Brain-specific homeobox/POU domain protein 1) (Brain-1) (Brn-1 protein). | |||||
|
PRGC2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 42) | NC score | 0.033064 (rank : 29) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86YN6, Q86YN3, Q86YN4, Q86YN5, Q8N1N9, Q8TDE4, Q8TDE5 | Gene names | PPARGC1B, PERC, PGC1, PGC1B, PPARGC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta) (PGC-1-related estrogen receptor alpha coactivator) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 43) | NC score | 0.006877 (rank : 95) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
ACK1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.003491 (rank : 111) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1140 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q07912, Q8N6U7, Q96H59 | Gene names | TNK2, ACK1 | |||
|
Domain Architecture |
|
|||||
| Description | Activated CDC42 kinase 1 (EC 2.7.10.2) (ACK-1) (Tyrosine kinase non- receptor protein 2). | |||||
|
ASPX_HUMAN
|
||||||
| θ value | 2.36792 (rank : 45) | NC score | 0.033503 (rank : 28) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P26436 | Gene names | ACRV1 | |||
|
Domain Architecture |

|
|||||
| Description | Acrosomal protein SP-10 precursor (Acrosomal vesicle protein 1). | |||||
|
BAI1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 46) | NC score | 0.009093 (rank : 90) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 340 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q3UHD1, Q3UH36, Q8CGM0 | Gene names | Bai1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain-specific angiogenesis inhibitor 1 precursor. | |||||
|
CKLF3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.030895 (rank : 35) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96MX0, Q8IUU8, Q8IWQ6, Q8IYE2, Q8IZ39, Q8IZ59 | Gene names | CMTM3, CKLFSF3 | |||
|
Domain Architecture |
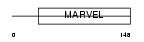
|
|||||
| Description | CKLF-like MARVEL transmembrane domain-containing protein 3 (Chemokine- like factor superfamily member 3). | |||||
|
COKA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.010688 (rank : 85) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P218, Q4VXQ4, Q8WUT2, Q96CY9, Q9BQU6, Q9BQU7 | Gene names | COL20A1, KIAA1510 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XX) chain precursor. | |||||
|
CPT1B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.013189 (rank : 78) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92523, Q13389, Q99655, Q9BY90 | Gene names | CPT1B, KIAA1670 | |||
|
Domain Architecture |

|
|||||
| Description | Carnitine O-palmitoyltransferase I, muscle isoform (EC 2.3.1.21) (CPT I) (CPTI-M) (Carnitine palmitoyltransferase 1B) (Carnitine palmitoyltransferase I-like protein). | |||||
|
DEN1A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.024985 (rank : 53) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TEH3, Q5VWF0, Q8IVD6, Q9H796 | Gene names | DENND1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
IF4G1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.031111 (rank : 34) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
MAVS_MOUSE
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.031929 (rank : 32) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VCF0 | Gene names | Mavs, Ips1, Visa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif). | |||||
|
MUCDL_MOUSE
|
||||||
| θ value | 2.36792 (rank : 53) | NC score | 0.020095 (rank : 58) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8VHF2, Q8CEJ3, Q9D8I9 | Gene names | Mucdhl | |||
|
Domain Architecture |
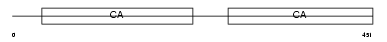
|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
PO6F2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 54) | NC score | 0.018100 (rank : 64) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P78424, P78425, Q75ME8, Q86UM6, Q9UDS7 | Gene names | POU6F2, RPF1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 2 (Retina-derived POU-domain factor 1) (RPF-1). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 55) | NC score | 0.026492 (rank : 47) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
TSSC4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 56) | NC score | 0.034202 (rank : 25) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5U2, Q86VL2, Q9BRS6 | Gene names | TSSC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein TSSC4 (Tumor-suppressing subchromosomal transferable fragment candidate gene 4 protein) (Tumor-suppressing STF cDNA 4 protein). | |||||
|
UB7I4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 57) | NC score | 0.021900 (rank : 56) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q925F3, Q6A0B4 | Gene names | Rnf144, Kiaa0161, Ubce7ip4, Uip4 | |||
|
Domain Architecture |
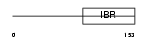
|
|||||
| Description | Ubiquitin-conjugating enzyme 7-interacting protein 4 (UbcM4- interacting protein 4) (RING finger protein 144). | |||||
|
CD2L7_HUMAN
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.003835 (rank : 110) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
COT1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.005490 (rank : 101) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60632, Q61438 | Gene names | Nr2f1, Erbal3, Tfcoup1 | |||
|
Domain Architecture |
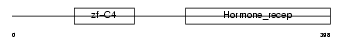
|
|||||
| Description | COUP transcription factor 1 (COUP-TF1) (COUP-TF I). | |||||
|
ERF_MOUSE
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.011863 (rank : 80) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
PK1L1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.025743 (rank : 50) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TDX9, Q6UWK1 | Gene names | PKD1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 1-like 1 protein (Polycystin-1L1). | |||||
|
WASIP_MOUSE
|
||||||
| θ value | 3.0926 (rank : 62) | NC score | 0.032390 (rank : 30) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
ASCL1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.016576 (rank : 70) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02067 | Gene names | Ascl1, Ash1, Mash-1, Mash1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 1 (Mash-1). | |||||
|
AT10A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.008470 (rank : 92) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
ATX2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.025621 (rank : 51) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O70305, P97421 | Gene names | Atxn2, Atx2, Sca2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein homolog). | |||||
|
CJ078_MOUSE
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.027795 (rank : 45) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 559 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BP27, Q3UIQ6, Q8R3W0, Q9CRT7, Q9D0D7, Q9D116, Q9D4W4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf78 homolog. | |||||
|
CNOT4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.029272 (rank : 40) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O95628, O95339, O95627, Q8IYM7, Q8NCL0, Q9NPQ1, Q9NZN6 | Gene names | CNOT4, NOT4 | |||
|
Domain Architecture |
|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
CNOT4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.029598 (rank : 39) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |
|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
FBXL6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.019214 (rank : 61) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N531, Q9H5W9, Q9UKC7 | Gene names | FBXL6, FBL6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 6 (F-box and leucine-rich repeat protein 6) (F-box protein FBL6) (FBL6A). | |||||
|
GGN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.026853 (rank : 46) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
HN1L_HUMAN
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.025917 (rank : 48) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H910, Q6EIC7 | Gene names | C16orf34, HN1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hematological and neurological expressed 1-like protein (HN1-like protein). | |||||
|
K1718_MOUSE
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.013657 (rank : 77) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
PRD15_HUMAN
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.000144 (rank : 116) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 805 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P57071, Q8N0X3, Q8NEX0, Q9NQV3 | Gene names | PRDM15, C21orf83, ZNF298 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 15 (PR domain-containing protein 15) (Zinc finger protein 298). | |||||
|
APC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.028338 (rank : 43) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
ATX2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.023359 (rank : 54) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99700, Q6ZQZ7, Q99493 | Gene names | ATXN2, ATX2, SCA2, TNRC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein) (Trinucleotide repeat-containing gene 13 protein). | |||||
|
BMP6_MOUSE
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.006416 (rank : 98) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20722 | Gene names | Bmp6, Bmp-6, Vgr1 | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein 6 precursor (BMP-6) (VG-1-related protein) (VGR-1). | |||||
|
CAD24_HUMAN
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.002976 (rank : 112) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86UP0, Q86UP1, Q9NT84 | Gene names | CDH24, CDH11L | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-24 precursor. | |||||
|
CSPG4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.011465 (rank : 81) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6UVK1, Q92675 | Gene names | CSPG4, MCSP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 4 precursor (Chondroitin sulfate proteoglycan NG2) (Melanoma chondroitin sulfate proteoglycan) (Melanoma-associated chondroitin sulfate proteoglycan). | |||||
|
CTGE2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.006105 (rank : 99) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96RT6 | Gene names | CTAGE1, CTAGE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein cTAGE-2. | |||||
|
DOCK1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.009055 (rank : 91) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14185 | Gene names | DOCK1 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 1 (180 kDa protein downstream of CRK) (DOCK180). | |||||
|
ICAL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.019841 (rank : 60) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51125, Q9EQV4, Q9EQV5, Q9QXQ3, Q9QXQ4, Q9R0N1 | Gene names | Cast | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor). | |||||
|
JPH3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.010808 (rank : 84) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WXH2, Q8N471, Q9HDC3, Q9HDC4 | Gene names | JPH3, JP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
LRP11_MOUSE
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | 0.015603 (rank : 72) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CB67, Q8C7Y7 | Gene names | Lrp11 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 11 precursor. | |||||
|
NOL3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.030487 (rank : 36) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D1X0 | Gene names | Nol3 | |||
|
Domain Architecture |
|
|||||
| Description | Nucleolar protein 3. | |||||
|
NOTC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.004389 (rank : 107) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 916 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q01705, Q06007, Q61905, Q99JC2, Q9QW58, Q9R0X7 | Gene names | Notch1, Motch | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 1 precursor (Notch 1) (Motch A) (mT14) (p300) [Contains: Notch 1 extracellular truncation; Notch 1 intracellular domain]. | |||||
|
UB7I4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | 0.018928 (rank : 62) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P50876 | Gene names | RNF144, KIAA0161, UBCE7IP4 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-conjugating enzyme 7-interacting protein 4 (UbcM4- interacting protein 4) (RING finger protein 144). | |||||
|
ZEP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 87) | NC score | 0.003929 (rank : 108) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3UHF7, O55140, Q3UVD4, Q3UVH5 | Gene names | Hivep2, Mibp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 homolog (Myc intron-binding protein 1) (MIBP-1). | |||||
|
CCD96_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.016616 (rank : 69) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q2M329, Q8N2I7 | Gene names | CCDC96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
CS021_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.017480 (rank : 65) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IVT2 | Gene names | C19orf21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf21. | |||||
|
CSPG3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.006723 (rank : 96) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14594, Q9UPK6 | Gene names | CSPG3, NCAN, NEUR | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
EMIL3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.010895 (rank : 83) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NT22, Q76KT4 | Gene names | EMILIN3, C20orf130, EMILIN5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EMILIN-3 precursor (EMILIN-5) (Elastin microfibril interface-located protein 5) (Elastin microfibril interfacer 5). | |||||
|
HXD3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.005604 (rank : 100) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P09027, Q3UUD3 | Gene names | Hoxd3, Hox-4.1, Hoxd-3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D3 (Hox-4.1) (Homeobox protein MH-19). | |||||
|
INVS_HUMAN
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.004768 (rank : 105) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y283, Q5W0T6, Q8IVX8, Q9BRB9, Q9Y488, Q9Y498 | Gene names | INVS, INV, NPHP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning homolog) (Nephrocystin-2). | |||||
|
PRPC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.032303 (rank : 31) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P02810, Q4VBP2, Q53XA2, Q6P2F6 | Gene names | PRH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Salivary acidic proline-rich phosphoprotein 1/2 precursor (PRP-1/PRP- 2) (Parotid proline-rich protein 1/2) (Pr1/Pr2) (Protein C) (Parotid acidic protein) (Pa) (Parotid isoelectric focusing variant protein) (PIF-S) (Parotid double-band protein) (Db-s) [Contains: Salivary acidic proline-rich phosphoprotein 1/2; Salivary acidic proline-rich phosphoprotein 3/4 (PRP-3/PRP-4) (Protein A) (PIF-F) (Db-F); Peptide P-C]. | |||||
|
S3TC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.014046 (rank : 76) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TE82 | Gene names | SH3TC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 domain and tetratricopeptide repeats-containing protein 1. | |||||
|
SFR15_HUMAN
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.020083 (rank : 59) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O95104, Q6P1M5, Q8N3I8, Q9UFM1, Q9ULP8 | Gene names | SFRS15, KIAA1172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 15 (CTD-binding SR-like protein RA4). | |||||
|
UBP8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.005303 (rank : 103) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
ANR11_HUMAN
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.006642 (rank : 97) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
B4GN3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.009551 (rank : 88) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6L9W6, Q6ZNC1, Q8N7T6 | Gene names | B4GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
BRD4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.008389 (rank : 93) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.028413 (rank : 42) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CEL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 102) | NC score | 0.014416 (rank : 74) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
CF118_HUMAN
|
||||||
| θ value | 8.99809 (rank : 103) | NC score | 0.016155 (rank : 71) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5T5N4, Q8TC11 | Gene names | C6orf118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf118. | |||||
|
DEN1A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.018142 (rank : 63) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K382 | Gene names | Dennd1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
EMID1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.011064 (rank : 82) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91VF5 | Gene names | Emid1, Emu1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EMI domain-containing protein 1 precursor (Protein Emu1) (Emilin and multimerin domain-containing protein 1). | |||||
|
FGRL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.002819 (rank : 113) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N441, Q6PJN1, Q9BXN7, Q9H4D7 | Gene names | FGFRL1, FGFR5, FHFR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor receptor-like 1 precursor (FGF receptor-like protein 1) (Fibroblast growth factor receptor 5) (FGFR-like protein) (FGF homologous factor receptor). | |||||
|
FOXE1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.006997 (rank : 94) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O00358, O75765 | Gene names | FOXE1, FKHL15, TITF2, TTF2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein E1 (Thyroid transcription factor 2) (TTF-2) (Forkhead-related protein FKHL15). | |||||
|
HSP74_HUMAN
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.004680 (rank : 106) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P34932, O95756 | Gene names | HSPA4 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4 (Heat shock 70-related protein APG-2) (HSP70RY). | |||||
|
JIP3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.010235 (rank : 87) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1039 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UPT6, Q96RY4, Q9H4I4, Q9H7P1, Q9NUG0 | Gene names | MAPK8IP3, JIP3, KIAA1066 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 3 (JNK-interacting protein 3) (JIP-3) (JNK MAP kinase scaffold protein 3) (Mitogen- activated protein kinase 8-interacting protein 3). | |||||
|
K2027_HUMAN
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.015400 (rank : 73) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6ZVF9, Q8IVE4 | Gene names | KIAA2027 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA2027. | |||||
|
KLF13_MOUSE
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.001724 (rank : 114) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 757 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JJZ6, Q9ESX3, Q9JHF8 | Gene names | Klf13, Bteb3, Fklf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 13 (Transcription factor BTEB3) (Basic transcription element-binding protein 3) (BTE-binding protein 3) (RANTES factor of late activated T-lymphocytes 1) (RFLAT-1) (Erythroid transcription factor FKLF-2). | |||||
|
KLKB1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.000969 (rank : 115) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P26262 | Gene names | Klkb1, Klk3, Pk | |||
|
Domain Architecture |

|
|||||
| Description | Plasma kallikrein precursor (EC 3.4.21.34) (Plasma prekallikrein) (Kininogenin) (Fletcher factor) [Contains: Plasma kallikrein heavy chain; Plasma kallikrein light chain]. | |||||
|
LTBP4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.005436 (rank : 102) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K4G1, Q8K4G0 | Gene names | Ltbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 4 precursor (LTBP-4). | |||||
|
MAML1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.012896 (rank : 79) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q6T264, Q505D8, Q5SUC2, Q6PDK3, Q6ZQG5, Q8BIU5, Q8R3T0 | Gene names | Maml1, Kiaa0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.009188 (rank : 89) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SNIP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.014251 (rank : 75) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QWI6, O70298 | Gene names | P140, Kiaa1684 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p130Cas-associated protein (p140Cap) (SNAP-25-interacting protein) (SNIP). | |||||
|
RTEL1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 116 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
DDX11_MOUSE
|
||||||
| NC score | 0.842785 (rank : 2) | θ value | 2.84138e-62 (rank : 2) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6AXC6 | Gene names | Ddx11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX11 (EC 3.6.1.-) (DEAD/H box protein 11). | |||||
|
FANCJ_HUMAN
|
||||||
| NC score | 0.820228 (rank : 3) | θ value | 1.36585e-56 (rank : 4) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BX63, Q8NCI5 | Gene names | BRIP1, BACH1, FANCJ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group J protein (EC 3.6.1.-) (ATP-dependent RNA helicase BRIP1) (Protein FACJ) (BRCA1-interacting protein C-terminal helicase 1) (BRCA1-interacting protein 1) (BRCA1-associated C-terminal helicase 1). | |||||
|
ERCC2_MOUSE
|
||||||
| NC score | 0.817762 (rank : 4) | θ value | 3.1488e-53 (rank : 5) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08811, Q9DC01 | Gene names | Ercc2, Xpd | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase subunit (EC 3.6.1.-) (DNA-repair protein complementing XP-D cells) (Xeroderma pigmentosum group D-complementing protein) (CXPD) (DNA excision repair protein ERCC-2). | |||||
|
ERCC2_HUMAN
|
||||||
| NC score | 0.815878 (rank : 5) | θ value | 7.75577e-52 (rank : 6) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P18074, Q8N721 | Gene names | ERCC2, XPD, XPDC | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase subunit (EC 3.6.1.-) (DNA-repair protein complementing XP-D cells) (Xeroderma pigmentosum group D-complementing protein) (CXPD) (DNA excision repair protein ERCC-2). | |||||
|
FANCJ_MOUSE
|
||||||
| NC score | 0.809450 (rank : 6) | θ value | 2.7521e-57 (rank : 3) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5SXJ3, Q8BJQ8, Q8BKI6 | Gene names | Brip1, Bach1, Fancj | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group J protein homolog (EC 3.6.1.-) (ATP-dependent RNA helicase BRIP1) (Protein FACJ) (BRCA1-interacting protein C-terminal helicase 1) (BRCA1-interacting protein 1) (BRCA1-associated C-terminal helicase 1). | |||||
|
DDX11_HUMAN
|
||||||
| NC score | 0.802851 (rank : 7) | θ value | 2.18568e-46 (rank : 7) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96FC9, Q13333, Q86VQ4, Q86W62, Q92498, Q92770, Q92998, Q92999 | Gene names | DDX11, CHL1, CHLR1, KRG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX11 (EC 3.6.1.-) (DEAD/H box protein 11) (CHL1 homolog) (Keratinocyte growth factor-regulated gene 2 protein) (KRG-2). | |||||
|
APOL5_HUMAN
|
||||||
| NC score | 0.058925 (rank : 8) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BWW9, Q5TFL9, Q9UGW5 | Gene names | APOL5 | |||
|
Domain Architecture |

|
|||||
| Description | Apolipoprotein-L5 (Apolipoprotein L-V) (ApoL-V). | |||||
|
AHTF1_MOUSE
|
||||||
| NC score | 0.050543 (rank : 9) | θ value | 0.0961366 (rank : 10) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CJF7, Q8BVJ5, Q8VD55 | Gene names | Ahctf1, Elys | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
K0226_HUMAN
|
||||||
| NC score | 0.048409 (rank : 10) | θ value | 0.813845 (rank : 28) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92622 | Gene names | KIAA0226 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0226. | |||||
|
DOK7_HUMAN
|
||||||
| NC score | 0.047951 (rank : 11) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q18PE1, Q6P6A6, Q86XG5, Q8N2J3, Q8NBC1 | Gene names | DOK7, C4orf25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Dok-7 (Downstream of tyrosine kinase 7). | |||||
|
TP53B_HUMAN
|
||||||
| NC score | 0.047866 (rank : 12) | θ value | 0.0961366 (rank : 11) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 413 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12888, Q2M1Z7, Q4LE46, Q5FWZ3, Q7Z3U4 | Gene names | TP53BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
K0226_MOUSE
|
||||||
| NC score | 0.046371 (rank : 13) | θ value | 1.38821 (rank : 37) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80U62, Q3TAX3, Q3TDF7, Q3TDU2, Q6P9T7, Q6PG18, Q8BMP7, Q8BY22 | Gene names | Kiaa0226 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0226. | |||||
|
IER5_MOUSE
|
||||||
| NC score | 0.042783 (rank : 14) | θ value | 1.06291 (rank : 31) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O89113, Q8CGI0 | Gene names | Ier5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immediate early response gene 5 protein. | |||||
|
LYAR_MOUSE
|
||||||
| NC score | 0.042699 (rank : 15) | θ value | 0.62314 (rank : 24) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q08288, Q9D9X2 | Gene names | Lyar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth-regulating nucleolar protein. | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.042520 (rank : 16) | θ value | 0.0736092 (rank : 9) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
IQEC2_HUMAN
|
||||||
| NC score | 0.041745 (rank : 17) | θ value | 0.163984 (rank : 14) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5JU85, O60275 | Gene names | IQSEC2, KIAA0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
IQEC2_MOUSE
|
||||||
| NC score | 0.041314 (rank : 18) | θ value | 0.163984 (rank : 15) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.040427 (rank : 19) | θ value | 0.163984 (rank : 13) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
PHLA1_HUMAN
|
||||||
| NC score | 0.040037 (rank : 20) | θ value | 1.06291 (rank : 34) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WV24, Q15184, Q2TAN2, Q9NZ17 | Gene names | PHLDA1, PHRIP, TDAG51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Apoptosis-associated nuclear protein) (Proline- and histidine-rich protein) (Proline- and glutamine-rich protein) (PQ-rich protein). | |||||
|
DEND_MOUSE
|
||||||
| NC score | 0.039541 (rank : 21) | θ value | 1.06291 (rank : 30) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80TS7 | Gene names | Ddn, Kiaa0749 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dendrin. | |||||
|
IF4G1_MOUSE
|
||||||
| NC score | 0.035640 (rank : 22) | θ value | 1.06291 (rank : 32) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
APC_HUMAN
|
||||||
| NC score | 0.035241 (rank : 23) | θ value | 1.38821 (rank : 35) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
FA61A_MOUSE
|
||||||
| NC score | 0.034800 (rank : 24) | θ value | 0.813845 (rank : 27) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K2F8, Q9CTG8 | Gene names | Fam61a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM61A. | |||||
|
TSSC4_HUMAN
|
||||||
| NC score | 0.034202 (rank : 25) | θ value | 2.36792 (rank : 56) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5U2, Q86VL2, Q9BRS6 | Gene names | TSSC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein TSSC4 (Tumor-suppressing subchromosomal transferable fragment candidate gene 4 protein) (Tumor-suppressing STF cDNA 4 protein). | |||||
|
CT128_HUMAN
|
||||||
| NC score | 0.034184 (rank : 26) | θ value | 1.81305 (rank : 38) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BQN1, Q5JWN6, Q8N276 | Gene names | C20orf128 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf128. | |||||
|
CCNK_HUMAN
|
||||||
| NC score | 0.033847 (rank : 27) | θ value | 0.47712 (rank : 22) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75909, Q96B63, Q9NNY9 | Gene names | CCNK, CPR4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-K. | |||||
|
ASPX_HUMAN
|
||||||
| NC score | 0.033503 (rank : 28) | θ value | 2.36792 (rank : 45) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P26436 | Gene names | ACRV1 | |||
|
Domain Architecture |

|
|||||
| Description | Acrosomal protein SP-10 precursor (Acrosomal vesicle protein 1). | |||||
|
PRGC2_HUMAN
|
||||||
| NC score | 0.033064 (rank : 29) | θ value | 1.81305 (rank : 42) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86YN6, Q86YN3, Q86YN4, Q86YN5, Q8N1N9, Q8TDE4, Q8TDE5 | Gene names | PPARGC1B, PERC, PGC1, PGC1B, PPARGC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta) (PGC-1-related estrogen receptor alpha coactivator) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
WASIP_MOUSE
|
||||||
| NC score | 0.032390 (rank : 30) | θ value | 3.0926 (rank : 62) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
PRPC_HUMAN
|
||||||
| NC score | 0.032303 (rank : 31) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P02810, Q4VBP2, Q53XA2, Q6P2F6 | Gene names | PRH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Salivary acidic proline-rich phosphoprotein 1/2 precursor (PRP-1/PRP- 2) (Parotid proline-rich protein 1/2) (Pr1/Pr2) (Protein C) (Parotid acidic protein) (Pa) (Parotid isoelectric focusing variant protein) (PIF-S) (Parotid double-band protein) (Db-s) [Contains: Salivary acidic proline-rich phosphoprotein 1/2; Salivary acidic proline-rich phosphoprotein 3/4 (PRP-3/PRP-4) (Protein A) (PIF-F) (Db-F); Peptide P-C]. | |||||
|
MAVS_MOUSE
|
||||||
| NC score | 0.031929 (rank : 32) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VCF0 | Gene names | Mavs, Ips1, Visa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif). | |||||
|
MOT3_MOUSE
|
||||||
| NC score | 0.031246 (rank : 33) | θ value | 0.125558 (rank : 12) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35308 | Gene names | Slc16a8, Mct3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Monocarboxylate transporter 3 (MCT 3) (Solute carrier family 16 member 8) (Proton-coupled monocarboxylate transporter 3). | |||||
|
IF4G1_HUMAN
|
||||||
| NC score | 0.031111 (rank : 34) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
CKLF3_HUMAN
|
||||||
| NC score | 0.030895 (rank : 35) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96MX0, Q8IUU8, Q8IWQ6, Q8IYE2, Q8IZ39, Q8IZ59 | Gene names | CMTM3, CKLFSF3 | |||
|
Domain Architecture |

|
|||||
| Description | CKLF-like MARVEL transmembrane domain-containing protein 3 (Chemokine- like factor superfamily member 3). | |||||
|
NOL3_MOUSE
|
||||||
| NC score | 0.030487 (rank : 36) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D1X0 | Gene names | Nol3 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar protein 3. | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.030029 (rank : 37) | θ value | 1.38821 (rank : 36) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
IRX3_MOUSE
|
||||||
| NC score | 0.029664 (rank : 38) | θ value | 0.21417 (rank : 18) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P81067 | Gene names | Irx3, Irxb1 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-3 (Iroquois homeobox protein 3) (Homeodomain protein IRXB1). | |||||
|
CNOT4_MOUSE
|
||||||
| NC score | 0.029598 (rank : 39) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
CNOT4_HUMAN
|
||||||
| NC score | 0.029272 (rank : 40) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O95628, O95339, O95627, Q8IYM7, Q8NCL0, Q9NPQ1, Q9NZN6 | Gene names | CNOT4, NOT4 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
SRCA_HUMAN
|
||||||
| NC score | 0.029154 (rank : 41) | θ value | 0.62314 (rank : 25) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.028413 (rank : 42) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
APC_MOUSE
|
||||||
| NC score | 0.028338 (rank : 43) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
RBM28_HUMAN
|
||||||
| NC score | 0.028240 (rank : 44) | θ value | 0.163984 (rank : 16) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NW13, Q96CV3 | Gene names | RBM28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 28 (RNA-binding motif protein 28). | |||||
|
CJ078_MOUSE
|
||||||
| NC score | 0.027795 (rank : 45) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 559 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BP27, Q3UIQ6, Q8R3W0, Q9CRT7, Q9D0D7, Q9D116, Q9D4W4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf78 homolog. | |||||
|
GGN_MOUSE
|
||||||
| NC score | 0.026853 (rank : 46) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.026492 (rank : 47) | θ value | 2.36792 (rank : 55) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
HN1L_HUMAN
|
||||||
| NC score | 0.025917 (rank : 48) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H910, Q6EIC7 | Gene names | C16orf34, HN1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hematological and neurological expressed 1-like protein (HN1-like protein). | |||||
|
RPKL1_HUMAN
|
||||||
| NC score | 0.025916 (rank : 49) | θ value | 0.00390308 (rank : 8) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y6S9, Q69YT9, Q6ZMQ6, Q96NR9, Q9BSU9 | Gene names | RPS6KL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal protein S6 kinase-like 1 (EC 2.7.11.1). | |||||
|
PK1L1_HUMAN
|
||||||
| NC score | 0.025743 (rank : 50) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TDX9, Q6UWK1 | Gene names | PKD1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 1-like 1 protein (Polycystin-1L1). | |||||
|
ATX2_MOUSE
|
||||||
| NC score | 0.025621 (rank : 51) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O70305, P97421 | Gene names | Atxn2, Atx2, Sca2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein homolog). | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.025213 (rank : 52) | θ value | 0.813845 (rank : 26) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
DEN1A_HUMAN
|
||||||
| NC score | 0.024985 (rank : 53) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TEH3, Q5VWF0, Q8IVD6, Q9H796 | Gene names | DENND1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
ATX2_HUMAN
|
||||||
| NC score | 0.023359 (rank : 54) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99700, Q6ZQZ7, Q99493 | Gene names | ATXN2, ATX2, SCA2, TNRC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein) (Trinucleotide repeat-containing gene 13 protein). | |||||
|
GP152_HUMAN
|
||||||
| NC score | 0.022442 (rank : 55) | θ value | 0.47712 (rank : 23) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
UB7I4_MOUSE
|
||||||
| NC score | 0.021900 (rank : 56) | θ value | 2.36792 (rank : 57) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q925F3, Q6A0B4 | Gene names | Rnf144, Kiaa0161, Ubce7ip4, Uip4 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-conjugating enzyme 7-interacting protein 4 (UbcM4- interacting protein 4) (RING finger protein 144). | |||||
|
AMOL1_MOUSE
|
||||||
| NC score | 0.021166 (rank : 57) | θ value | 1.06291 (rank : 29) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
MUCDL_MOUSE
|
||||||
| NC score | 0.020095 (rank : 58) | θ value | 2.36792 (rank : 53) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8VHF2, Q8CEJ3, Q9D8I9 | Gene names | Mucdhl | |||
|
Domain Architecture |

|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
SFR15_HUMAN
|
||||||
| NC score | 0.020083 (rank : 59) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O95104, Q6P1M5, Q8N3I8, Q9UFM1, Q9ULP8 | Gene names | SFRS15, KIAA1172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 15 (CTD-binding SR-like protein RA4). | |||||
|
ICAL_MOUSE
|
||||||
| NC score | 0.019841 (rank : 60) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51125, Q9EQV4, Q9EQV5, Q9QXQ3, Q9QXQ4, Q9R0N1 | Gene names | Cast | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor). | |||||
|
FBXL6_HUMAN
|
||||||
| NC score | 0.019214 (rank : 61) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N531, Q9H5W9, Q9UKC7 | Gene names | FBXL6, FBL6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 6 (F-box and leucine-rich repeat protein 6) (F-box protein FBL6) (FBL6A). | |||||
|
UB7I4_HUMAN
|
||||||
| NC score | 0.018928 (rank : 62) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P50876 | Gene names | RNF144, KIAA0161, UBCE7IP4 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-conjugating enzyme 7-interacting protein 4 (UbcM4- interacting protein 4) (RING finger protein 144). | |||||
|
DEN1A_MOUSE
|
||||||
| NC score | 0.018142 (rank : 63) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K382 | Gene names | Dennd1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
PO6F2_HUMAN
|
||||||
| NC score | 0.018100 (rank : 64) | θ value | 2.36792 (rank : 54) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P78424, P78425, Q75ME8, Q86UM6, Q9UDS7 | Gene names | POU6F2, RPF1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 2 (Retina-derived POU-domain factor 1) (RPF-1). | |||||
|
CS021_HUMAN
|
||||||
| NC score | 0.017480 (rank : 65) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IVT2 | Gene names | C19orf21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf21. | |||||
|
PO3F3_MOUSE
|
||||||
| NC score | 0.017086 (rank : 66) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P31361 | Gene names | Pou3f3, Brn-1, Brn1, Otf8 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 3 (Brain-specific homeobox/POU domain protein 1) (Brain-1) (Brn-1 protein). | |||||
|
PO3F3_HUMAN
|
||||||
| NC score | 0.017084 (rank : 67) | θ value | 1.81305 (rank : 40) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P20264, P78379 | Gene names | POU3F3, BRN1, OTF8 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 3 (Brain-specific homeobox/POU domain protein 1) (Brain-1) (Brn-1 protein). | |||||
|
SREC2_MOUSE
|
||||||
| NC score | 0.016650 (rank : 68) | θ value | 0.279714 (rank : 19) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 605 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P59222 | Gene names | Scarf2, Srec2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II). | |||||
|
CCD96_HUMAN
|
||||||
| NC score | 0.016616 (rank : 69) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q2M329, Q8N2I7 | Gene names | CCDC96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
ASCL1_MOUSE
|
||||||
| NC score | 0.016576 (rank : 70) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02067 | Gene names | Ascl1, Ash1, Mash-1, Mash1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 1 (Mash-1). | |||||
|
CF118_HUMAN
|
||||||
| NC score | 0.016155 (rank : 71) | θ value | 8.99809 (rank : 103) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5T5N4, Q8TC11 | Gene names | C6orf118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf118. | |||||
|
LRP11_MOUSE
|
||||||
| NC score | 0.015603 (rank : 72) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CB67, Q8C7Y7 | Gene names | Lrp11 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 11 precursor. | |||||
|
K2027_HUMAN
|
||||||
| NC score | 0.015400 (rank : 73) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6ZVF9, Q8IVE4 | Gene names | KIAA2027 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA2027. | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.014416 (rank : 74) | θ value | 8.99809 (rank : 102) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
SNIP_MOUSE
|
||||||
| NC score | 0.014251 (rank : 75) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QWI6, O70298 | Gene names | P140, Kiaa1684 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p130Cas-associated protein (p140Cap) (SNAP-25-interacting protein) (SNIP). | |||||
|
S3TC1_HUMAN
|
||||||
| NC score | 0.014046 (rank : 76) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TE82 | Gene names | SH3TC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 domain and tetratricopeptide repeats-containing protein 1. | |||||
|
K1718_MOUSE
|
||||||
| NC score | 0.013657 (rank : 77) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
CPT1B_HUMAN
|
||||||
| NC score | 0.013189 (rank : 78) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92523, Q13389, Q99655, Q9BY90 | Gene names | CPT1B, KIAA1670 | |||
|
Domain Architecture |

|
|||||
| Description | Carnitine O-palmitoyltransferase I, muscle isoform (EC 2.3.1.21) (CPT I) (CPTI-M) (Carnitine palmitoyltransferase 1B) (Carnitine palmitoyltransferase I-like protein). | |||||
|
MAML1_MOUSE
|
||||||
| NC score | 0.012896 (rank : 79) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q6T264, Q505D8, Q5SUC2, Q6PDK3, Q6ZQG5, Q8BIU5, Q8R3T0 | Gene names | Maml1, Kiaa0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
ERF_MOUSE
|
||||||
| NC score | 0.011863 (rank : 80) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
CSPG4_HUMAN
|
||||||
| NC score | 0.011465 (rank : 81) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6UVK1, Q92675 | Gene names | CSPG4, MCSP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 4 precursor (Chondroitin sulfate proteoglycan NG2) (Melanoma chondroitin sulfate proteoglycan) (Melanoma-associated chondroitin sulfate proteoglycan). | |||||
|
EMID1_MOUSE
|
||||||
| NC score | 0.011064 (rank : 82) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91VF5 | Gene names | Emid1, Emu1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EMI domain-containing protein 1 precursor (Protein Emu1) (Emilin and multimerin domain-containing protein 1). | |||||
|
EMIL3_HUMAN
|
||||||
| NC score | 0.010895 (rank : 83) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NT22, Q76KT4 | Gene names | EMILIN3, C20orf130, EMILIN5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EMILIN-3 precursor (EMILIN-5) (Elastin microfibril interface-located protein 5) (Elastin microfibril interfacer 5). | |||||
|
JPH3_HUMAN
|
||||||
| NC score | 0.010808 (rank : 84) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WXH2, Q8N471, Q9HDC3, Q9HDC4 | Gene names | JPH3, JP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
COKA1_HUMAN
|
||||||
| NC score | 0.010688 (rank : 85) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P218, Q4VXQ4, Q8WUT2, Q96CY9, Q9BQU6, Q9BQU7 | Gene names | COL20A1, KIAA1510 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XX) chain precursor. | |||||
|
FOXC2_HUMAN
|
||||||
| NC score | 0.010544 (rank : 86) | θ value | 1.81305 (rank : 39) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99958 | Gene names | FOXC2, FKHL14, MFH1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
JIP3_HUMAN
|
||||||
| NC score | 0.010235 (rank : 87) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1039 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UPT6, Q96RY4, Q9H4I4, Q9H7P1, Q9NUG0 | Gene names | MAPK8IP3, JIP3, KIAA1066 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 3 (JNK-interacting protein 3) (JIP-3) (JNK MAP kinase scaffold protein 3) (Mitogen- activated protein kinase 8-interacting protein 3). | |||||
|
B4GN3_HUMAN
|
||||||
| NC score | 0.009551 (rank : 88) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6L9W6, Q6ZNC1, Q8N7T6 | Gene names | B4GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.009188 (rank : 89) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
BAI1_MOUSE
|
||||||
| NC score | 0.009093 (rank : 90) | θ value | 2.36792 (rank : 46) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 340 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q3UHD1, Q3UH36, Q8CGM0 | Gene names | Bai1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain-specific angiogenesis inhibitor 1 precursor. | |||||
|
DOCK1_HUMAN
|
||||||
| NC score | 0.009055 (rank : 91) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14185 | Gene names | DOCK1 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 1 (180 kDa protein downstream of CRK) (DOCK180). | |||||
|
AT10A_HUMAN
|
||||||
| NC score | 0.008470 (rank : 92) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
BRD4_MOUSE
|
||||||
| NC score | 0.008389 (rank : 93) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
FOXE1_HUMAN
|
||||||
| NC score | 0.006997 (rank : 94) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O00358, O75765 | Gene names | FOXE1, FKHL15, TITF2, TTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein E1 (Thyroid transcription factor 2) (TTF-2) (Forkhead-related protein FKHL15). | |||||
|
WNK2_HUMAN
|
||||||
| NC score | 0.006877 (rank : 95) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
CSPG3_HUMAN
|
||||||
| NC score | 0.006723 (rank : 96) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14594, Q9UPK6 | Gene names | CSPG3, NCAN, NEUR | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
ANR11_HUMAN
|
||||||
| NC score | 0.006642 (rank : 97) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
BMP6_MOUSE
|
||||||
| NC score | 0.006416 (rank : 98) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20722 | Gene names | Bmp6, Bmp-6, Vgr1 | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein 6 precursor (BMP-6) (VG-1-related protein) (VGR-1). | |||||
|
CTGE2_HUMAN
|
||||||
| NC score | 0.006105 (rank : 99) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96RT6 | Gene names | CTAGE1, CTAGE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein cTAGE-2. | |||||
|
HXD3_MOUSE
|
||||||
| NC score | 0.005604 (rank : 100) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P09027, Q3UUD3 | Gene names | Hoxd3, Hox-4.1, Hoxd-3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D3 (Hox-4.1) (Homeobox protein MH-19). | |||||
|
COT1_MOUSE
|
||||||
| NC score | 0.005490 (rank : 101) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60632, Q61438 | Gene names | Nr2f1, Erbal3, Tfcoup1 | |||
|
Domain Architecture |

|
|||||
| Description | COUP transcription factor 1 (COUP-TF1) (COUP-TF I). | |||||
|
LTBP4_MOUSE
|
||||||
| NC score | 0.005436 (rank : 102) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K4G1, Q8K4G0 | Gene names | Ltbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 4 precursor (LTBP-4). | |||||
|
UBP8_HUMAN
|
||||||
| NC score | 0.005303 (rank : 103) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
SN1L2_MOUSE
|
||||||
| NC score | 0.004939 (rank : 104) | θ value | 0.365318 (rank : 21) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CFH6 | Gene names | Snf1lk2, Sik2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Salt-inducible kinase 2). | |||||
|
INVS_HUMAN
|
||||||
| NC score | 0.004768 (rank : 105) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y283, Q5W0T6, Q8IVX8, Q9BRB9, Q9Y488, Q9Y498 | Gene names | INVS, INV, NPHP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning homolog) (Nephrocystin-2). | |||||
|
HSP74_HUMAN
|
||||||
| NC score | 0.004680 (rank : 106) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P34932, O95756 | Gene names | HSPA4 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4 (Heat shock 70-related protein APG-2) (HSP70RY). | |||||
|
NOTC1_MOUSE
|
||||||
| NC score | 0.004389 (rank : 107) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 916 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q01705, Q06007, Q61905, Q99JC2, Q9QW58, Q9R0X7 | Gene names | Notch1, Motch | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 1 precursor (Notch 1) (Motch A) (mT14) (p300) [Contains: Notch 1 extracellular truncation; Notch 1 intracellular domain]. | |||||
|
ZEP2_MOUSE
|
||||||
| NC score | 0.003929 (rank : 108) | θ value | 5.27518 (rank : 87) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3UHF7, O55140, Q3UVD4, Q3UVH5 | Gene names | Hivep2, Mibp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 homolog (Myc intron-binding protein 1) (MIBP-1). | |||||
|
KLF13_HUMAN
|
||||||
| NC score | 0.003859 (rank : 109) | θ value | 1.06291 (rank : 33) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2Y9, Q9Y356 | Gene names | KLF13, BTEB3, NSLP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 13 (Transcription factor BTEB3) (Basic transcription element-binding protein 3) (BTE-binding protein 3) (RANTES factor of late activated T-lymphocytes 1) (RFLAT-1) (Transcription factor NSLP1) (Novel Sp1-like zinc finger transcription factor 1). | |||||
|
CD2L7_HUMAN
|
||||||
| NC score | 0.003835 (rank : 110) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
ACK1_HUMAN
|
||||||
| NC score | 0.003491 (rank : 111) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1140 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q07912, Q8N6U7, Q96H59 | Gene names | TNK2, ACK1 | |||
|
Domain Architecture |

|
|||||
| Description | Activated CDC42 kinase 1 (EC 2.7.10.2) (ACK-1) (Tyrosine kinase non- receptor protein 2). | |||||
|
CAD24_HUMAN
|
||||||
| NC score | 0.002976 (rank : 112) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86UP0, Q86UP1, Q9NT84 | Gene names | CDH24, CDH11L | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-24 precursor. | |||||
|
FGRL1_HUMAN
|
||||||
| NC score | 0.002819 (rank : 113) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N441, Q6PJN1, Q9BXN7, Q9H4D7 | Gene names | FGFRL1, FGFR5, FHFR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor receptor-like 1 precursor (FGF receptor-like protein 1) (Fibroblast growth factor receptor 5) (FGFR-like protein) (FGF homologous factor receptor). | |||||
|
KLF13_MOUSE
|
||||||
| NC score | 0.001724 (rank : 114) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 757 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JJZ6, Q9ESX3, Q9JHF8 | Gene names | Klf13, Bteb3, Fklf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 13 (Transcription factor BTEB3) (Basic transcription element-binding protein 3) (BTE-binding protein 3) (RANTES factor of late activated T-lymphocytes 1) (RFLAT-1) (Erythroid transcription factor FKLF-2). | |||||
|
KLKB1_MOUSE
|
||||||
| NC score | 0.000969 (rank : 115) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P26262 | Gene names | Klkb1, Klk3, Pk | |||
|
Domain Architecture |

|
|||||
| Description | Plasma kallikrein precursor (EC 3.4.21.34) (Plasma prekallikrein) (Kininogenin) (Fletcher factor) [Contains: Plasma kallikrein heavy chain; Plasma kallikrein light chain]. | |||||
|
PRD15_HUMAN
|
||||||
| NC score | 0.000144 (rank : 116) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 805 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P57071, Q8N0X3, Q8NEX0, Q9NQV3 | Gene names | PRDM15, C21orf83, ZNF298 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 15 (PR domain-containing protein 15) (Zinc finger protein 298). | |||||