Please be patient as the page loads
|
CS021_HUMAN
|
||||||
| SwissProt Accessions | Q8IVT2 | Gene names | C19orf21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf21. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CS021_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 98 | |
| SwissProt Accessions | Q8IVT2 | Gene names | C19orf21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf21. | |||||
|
CS021_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.948204 (rank : 2) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D279, Q3UI04 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf21 homolog. | |||||
|
AKAP2_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 3) | NC score | 0.171412 (rank : 3) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
MYO15_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 4) | NC score | 0.035260 (rank : 26) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QZZ4, O70395, Q9QWL6 | Gene names | Myo15a, Myo15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
AKAP2_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 5) | NC score | 0.130171 (rank : 4) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
ROBO3_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 6) | NC score | 0.024732 (rank : 48) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96MS0 | Gene names | ROBO3 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Roundabout-like protein 3). | |||||
|
CHD4_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 7) | NC score | 0.034983 (rank : 27) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 0.365318 (rank : 8) | NC score | 0.058997 (rank : 5) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
LIMD1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 9) | NC score | 0.028332 (rank : 40) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UGP4, Q9BQQ9, Q9NQ47 | Gene names | LIMD1 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain-containing protein 1. | |||||
|
ACK1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 10) | NC score | 0.011117 (rank : 88) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1080 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O54967, Q8C2U0, Q8K0K4 | Gene names | Tnk2, Ack1 | |||
|
Domain Architecture |
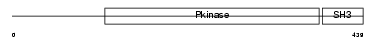
|
|||||
| Description | Activated CDC42 kinase 1 (EC 2.7.10.2) (ACK-1) (Non-receptor protein tyrosine kinase Ack) (Tyrosine kinase non-receptor protein 2). | |||||
|
MAGI2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 11) | NC score | 0.023918 (rank : 51) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 12) | NC score | 0.040646 (rank : 21) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.016136 (rank : 74) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
DIP2A_HUMAN
|
||||||
| θ value | 0.813845 (rank : 14) | NC score | 0.034624 (rank : 28) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14689, Q8IVA3, Q8N4S2, Q8TD89, Q96ML9 | Gene names | DIP2A, C21orf106, DIP2, KIAA0184 | |||
|
Domain Architecture |

|
|||||
| Description | Disco-interacting protein 2 homolog A. | |||||
|
AN13D_HUMAN
|
||||||
| θ value | 1.06291 (rank : 15) | NC score | 0.027828 (rank : 42) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZTN6, Q6ZVD0, Q86SU1 | Gene names | ANKRD13D | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 13D. | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.021965 (rank : 53) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
KKCC1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 17) | NC score | 0.010345 (rank : 93) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 866 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8VBY2, Q9R054 | Gene names | Camkk1, Camkk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium/calmodulin-dependent protein kinase kinase 1 (EC 2.7.11.17) (Calcium/calmodulin-dependent protein kinase kinase alpha) (CaM-kinase kinase alpha) (CaM-KK alpha) (CaMKK alpha) (CaMKK 1) (CaM-kinase IV kinase). | |||||
|
RAI1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.049413 (rank : 7) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7Z5J4, Q8N3B4, Q8ND08, Q8WU64, Q96JK5, Q9H1C1, Q9H1C2, Q9UF69 | Gene names | RAI1, KIAA1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
BAZ2A_MOUSE
|
||||||
| θ value | 1.38821 (rank : 19) | NC score | 0.041270 (rank : 20) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
ICAL_MOUSE
|
||||||
| θ value | 1.38821 (rank : 20) | NC score | 0.045568 (rank : 13) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P51125, Q9EQV4, Q9EQV5, Q9QXQ3, Q9QXQ4, Q9R0N1 | Gene names | Cast | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor). | |||||
|
MEGF9_MOUSE
|
||||||
| θ value | 1.38821 (rank : 21) | NC score | 0.024767 (rank : 47) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 22) | NC score | 0.047344 (rank : 10) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
T22D2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 23) | NC score | 0.041969 (rank : 18) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
CK024_MOUSE
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.049117 (rank : 8) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9D8N1, Q8VCP2 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 homolog precursor. | |||||
|
R113A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.036355 (rank : 25) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15541 | Gene names | RNF113A, RNF113, ZNF183 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 113A (Zinc finger protein 183). | |||||
|
SSH2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.019262 (rank : 59) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q76I76, Q8TDB5, Q8WYL1, Q8WYL2, Q96F40, Q96H36, Q9C0D8 | Gene names | SSH2, KIAA1725, SSH2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 2 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-2L) (hSSH-2L). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.051044 (rank : 6) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
MN1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.045897 (rank : 12) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q10571 | Gene names | MN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable tumor suppressor protein MN1. | |||||
|
SREC2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.013893 (rank : 80) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96GP6, Q8IXF3, Q9BW74 | Gene names | SCARF2, SREC2, SREPCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II) (SRECRP-1). | |||||
|
TFE3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.030571 (rank : 35) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
TMAP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.046513 (rank : 11) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P84157 | Gene names | TMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane anchor protein 1. | |||||
|
TRI47_MOUSE
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.016902 (rank : 70) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C0E3, Q6P249, Q811J7, Q8BVZ8, Q8R1K0, Q8R3Y1 | Gene names | Trim47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 47. | |||||
|
COEA1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.016799 (rank : 72) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80X19, Q8C6X3, Q9WV05 | Gene names | Col14a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor. | |||||
|
CSKI1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.019058 (rank : 61) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
ENAH_MOUSE
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.030355 (rank : 36) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
LDB3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.019104 (rank : 60) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O75112, Q5K6N9, Q5K6P0, Q5K6P1, Q96FH2, Q9Y4Z3, Q9Y4Z4, Q9Y4Z5 | Gene names | LDB3, KIAA0613, ZASP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher). | |||||
|
MAP2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.029825 (rank : 37) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P20357 | Gene names | Map2, Mtap2 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 2 (MAP 2). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.044530 (rank : 16) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
NPHP4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.028234 (rank : 41) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O75161 | Gene names | NPHP4, KIAA0673 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nephrocystin-4 (Nephroretinin). | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.045245 (rank : 14) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
RAI1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.045114 (rank : 15) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61818, Q6ZPH7, Q8CEV1 | Gene names | Rai1, Kiaa1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
RUNX1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.021525 (rank : 54) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q01196, O60472, O60473, O76047, O76089, Q13081, Q13755, Q13756, Q13757, Q13758, Q13759, Q15341, Q15343, Q16122, Q16284, Q16285, Q16286, Q16346, Q16347, Q92479 | Gene names | RUNX1, AML1, CBFA2 | |||
|
Domain Architecture |

|
|||||
| Description | Runt-related transcription factor 1 (Core-binding factor, alpha 2 subunit) (CBF-alpha 2) (Acute myeloid leukemia 1 protein) (Oncogene AML-1) (Polyomavirus enhancer-binding protein 2 alpha B subunit) (PEBP2-alpha B) (PEA2-alpha B) (SL3-3 enhancer factor 1 alpha B subunit) (SL3/AKV core-binding factor alpha B subunit). | |||||
|
SAM14_HUMAN
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.029763 (rank : 38) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IZD0, Q8N2X0 | Gene names | SAMD14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha motif domain-containing protein 14. | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.038125 (rank : 24) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
ATBF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.013648 (rank : 82) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
BRD4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.032889 (rank : 31) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
CHD4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.025618 (rank : 45) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
DCJ14_MOUSE
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.018464 (rank : 64) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q921R4, Q3TX73, Q8BUU3, Q9CYB7 | Gene names | Dnajc14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 14. | |||||
|
H15_HUMAN
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.024256 (rank : 49) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P16401, Q14529 | Gene names | HIST1H1B, H1F5 | |||
|
Domain Architecture |
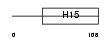
|
|||||
| Description | Histone H1.5 (Histone H1a). | |||||
|
IQEC3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.019384 (rank : 58) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UPP2, Q8TB43 | Gene names | IQSEC3, KIAA1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
MAML1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.027726 (rank : 43) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6T264, Q505D8, Q5SUC2, Q6PDK3, Q6ZQG5, Q8BIU5, Q8R3T0 | Gene names | Maml1, Kiaa0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
PHC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.025545 (rank : 46) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P78364, Q8WVM3, Q9BU63 | Gene names | PHC1, EDR1, PH1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (hPH1) (Early development regulatory protein 1). | |||||
|
PTTG1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.033341 (rank : 30) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95997 | Gene names | PTTG1, EAP1, PTTG, TUTR1 | |||
|
Domain Architecture |

|
|||||
| Description | Securin (Pituitary tumor-transforming protein 1) (Tumor-transforming protein 1) (Esp1-associated protein) (hPTTG). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.010963 (rank : 89) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
TAU_MOUSE
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.040074 (rank : 23) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P10637, P10638, Q60684, Q60685, Q60686, Q62286 | Gene names | Mapt, Mtapt, Tau | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
TTBK1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.010530 (rank : 91) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1058 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5TCY1, Q2L6C6, Q8N444, Q96JH2 | Gene names | TTBK1, BDTK, KIAA1855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 1 (EC 2.7.11.1) (Brain-derived tau kinase). | |||||
|
ATF7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.014377 (rank : 79) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17544, Q13814 | Gene names | ATF7, ATFA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.031625 (rank : 33) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.033341 (rank : 29) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CXA7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.013275 (rank : 84) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P36383 | Gene names | GJA7 | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-7 protein (Connexin-45) (Cx45). | |||||
|
CXA7_MOUSE
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.013361 (rank : 83) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28229 | Gene names | Gja7, Cxn-45 | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-7 protein (Connexin-45) (Cx45). | |||||
|
HCN2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.012388 (rank : 86) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O88703, O70506 | Gene names | Hcn2, Bcng2, Hac1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2) (Hyperpolarization-activated cation channel 1) (HAC-1). | |||||
|
IF3A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.010242 (rank : 94) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14152, O00653 | Gene names | EIF3S10, KIAA0139 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.030827 (rank : 34) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
ODO2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.023999 (rank : 50) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P36957, Q9BQ32 | Gene names | DLST, DLTS | |||
|
Domain Architecture |
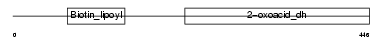
|
|||||
| Description | Dihydrolipoyllysine-residue succinyltransferase component of 2- oxoglutarate dehydrogenase complex, mitochondrial precursor (EC 2.3.1.61) (Dihydrolipoamide succinyltransferase component of 2- oxoglutarate dehydrogenase complex) (E2) (E2K). | |||||
|
RIF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.018320 (rank : 66) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 320 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5UIP0, Q5H9R3, Q5UIP2, Q66YK6, Q6PRU2, Q8TE94, Q99772, Q9H830, Q9H9B9, Q9NVP5, Q9Y4R4 | Gene names | RIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.016578 (rank : 73) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
TSSK2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.005156 (rank : 96) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 826 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96PF2, Q8IY55 | Gene names | TSSK2, DGSG, SPOGA2, STK22B | |||
|
Domain Architecture |
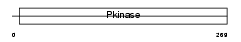
|
|||||
| Description | Testis-specific serine/threonine-protein kinase 2 (EC 2.7.11.1) (TSSK- 2) (Testis-specific kinase 2) (TSK-2) (Serine/threonine-protein kinase 22B) (DiGeorge syndrome protein G). | |||||
|
WASIP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.043434 (rank : 17) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
AFF1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.018182 (rank : 68) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
COEA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.015277 (rank : 76) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q05707, O00260, O00261, O00262, Q05708, Q5XJ18, Q96C67 | Gene names | COL14A1, UND | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor (Undulin). | |||||
|
G7D_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.026467 (rank : 44) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5SSQ6, Q9Y335 | Gene names | C6orf26, G7D, NG23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein G7d. | |||||
|
IRX1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.010444 (rank : 92) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78414, Q7Z2F8, Q8N312 | Gene names | IRX1, IRXA1 | |||
|
Domain Architecture |
|
|||||
| Description | Iroquois-class homeodomain protein IRX-1 (Iroquois homeobox protein 1) (Homeodomain protein IRXA1). | |||||
|
KBTBB_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.010574 (rank : 90) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O94819 | Gene names | KBTBD11, KIAA0711, KLHDC7C | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch repeat and BTB domain-containing protein 11 (Kelch domain- containing protein 7B). | |||||
|
KKCC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.007179 (rank : 95) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 858 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N5S9, Q9BQH3 | Gene names | CAMKK1, CAMKKA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium/calmodulin-dependent protein kinase kinase 1 (EC 2.7.11.17) (Calcium/calmodulin-dependent protein kinase kinase alpha) (CaM-kinase kinase alpha) (CaM-KK alpha) (CaMKK alpha) (CaMKK 1) (CaM-kinase IV kinase). | |||||
|
MARCS_HUMAN
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.028828 (rank : 39) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.048959 (rank : 9) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
RIN3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.041493 (rank : 19) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8TB24, Q8NF30, Q8TEE8, Q8WYP4, Q9H6A5, Q9HAG1 | Gene names | RIN3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
RTEL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.017480 (rank : 69) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
SPEG_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.013141 (rank : 85) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
TFEB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.022017 (rank : 52) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
WASIP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.040196 (rank : 22) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
ZIMP7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.014669 (rank : 77) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NF64, O94790, Q659A8, Q6JKL5, Q8WTX8, Q96Q01, Q9BQH7 | Gene names | ZIMP7, KIAA1886 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
ADA2B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.000024 (rank : 98) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 891 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P30545 | Gene names | Adra2b | |||
|
Domain Architecture |
|
|||||
| Description | Alpha-2B adrenergic receptor (Alpha-2B adrenoceptor) (Alpha-2B adrenoreceptor). | |||||
|
ATBF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.013845 (rank : 81) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
EPN3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.015756 (rank : 75) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H201, Q9BVN6, Q9NWK2 | Gene names | EPN3 | |||
|
Domain Architecture |
|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
FOXC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.004746 (rank : 97) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61850, P97948, Q63869, Q8C694 | Gene names | Foxc2, Fkh14, Fkhl14, Mfh1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
H14_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.019388 (rank : 57) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P10412 | Gene names | HIST1H1E, H1F4 | |||
|
Domain Architecture |
|
|||||
| Description | Histone H1.4 (Histone H1b). | |||||
|
H15_MOUSE
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.018657 (rank : 63) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P43276, Q9CRM8 | Gene names | Hist1h1b, H1f5 | |||
|
Domain Architecture |
|
|||||
| Description | Histone H1.5 (H1 VAR.5) (H1b). | |||||
|
IASPP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.018334 (rank : 65) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.019942 (rank : 56) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
MGAT1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.014647 (rank : 78) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P27808 | Gene names | Mgat1, Gnt1 | |||
|
Domain Architecture |
|
|||||
| Description | Alpha-1,3-mannosyl-glycoprotein 2-beta-N-acetylglucosaminyltransferase (EC 2.4.1.101) (N-glycosyl-oligosaccharide-glycoprotein N- acetylglucosaminyltransferase I) (GNT-I) (GlcNAc-T I). | |||||
|
ODP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.018313 (rank : 67) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10515, Q16783 | Gene names | DLAT, DLTA | |||
|
Domain Architecture |

|
|||||
| Description | Dihydrolipoyllysine-residue acetyltransferase component of pyruvate dehydrogenase complex, mitochondrial precursor (EC 2.3.1.12) (Pyruvate dehydrogenase complex E2 subunit) (PDCE2) (E2) (Dihydrolipoamide S- acetyltransferase component of pyruvate dehydrogenase complex) (PDC- E2) (70 kDa mitochondrial autoantigen of primary biliary cirrhosis) (PBC) (M2 antigen complex 70 kDa subunit). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.031717 (rank : 32) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
S23IP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.018692 (rank : 62) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6NZC7, Q8BXY6 | Gene names | Sec23ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SEC23-interacting protein. | |||||
|
TAF3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.020864 (rank : 55) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
TOP2A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.011297 (rank : 87) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q01320 | Gene names | Top2a, Top-2, Top2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
ZN318_MOUSE
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.016872 (rank : 71) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 642 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99PP2, Q3TZL5, Q9JJ01 | Gene names | Znf318, Tzf, Zfp318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Testicular zinc finger protein). | |||||
|
CS021_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 98 | |
| SwissProt Accessions | Q8IVT2 | Gene names | C19orf21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf21. | |||||
|
CS021_MOUSE
|
||||||
| NC score | 0.948204 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D279, Q3UI04 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf21 homolog. | |||||
|
AKAP2_HUMAN
|
||||||
| NC score | 0.171412 (rank : 3) | θ value | 0.00035302 (rank : 3) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
AKAP2_MOUSE
|
||||||
| NC score | 0.130171 (rank : 4) | θ value | 0.0252991 (rank : 5) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.058997 (rank : 5) | θ value | 0.365318 (rank : 8) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.051044 (rank : 6) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
RAI1_HUMAN
|
||||||
| NC score | 0.049413 (rank : 7) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7Z5J4, Q8N3B4, Q8ND08, Q8WU64, Q96JK5, Q9H1C1, Q9H1C2, Q9UF69 | Gene names | RAI1, KIAA1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
CK024_MOUSE
|
||||||
| NC score | 0.049117 (rank : 8) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9D8N1, Q8VCP2 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 homolog precursor. | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.048959 (rank : 9) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.047344 (rank : 10) | θ value | 1.38821 (rank : 22) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
TMAP1_HUMAN
|
||||||
| NC score | 0.046513 (rank : 11) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P84157 | Gene names | TMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane anchor protein 1. | |||||
|
MN1_HUMAN
|
||||||
| NC score | 0.045897 (rank : 12) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q10571 | Gene names | MN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable tumor suppressor protein MN1. | |||||
|
ICAL_MOUSE
|
||||||
| NC score | 0.045568 (rank : 13) | θ value | 1.38821 (rank : 20) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P51125, Q9EQV4, Q9EQV5, Q9QXQ3, Q9QXQ4, Q9R0N1 | Gene names | Cast | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor). | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.045245 (rank : 14) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
RAI1_MOUSE
|
||||||
| NC score | 0.045114 (rank : 15) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61818, Q6ZPH7, Q8CEV1 | Gene names | Rai1, Kiaa1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.044530 (rank : 16) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
WASIP_MOUSE
|
||||||
| NC score | 0.043434 (rank : 17) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
T22D2_HUMAN
|
||||||
| NC score | 0.041969 (rank : 18) | θ value | 1.38821 (rank : 23) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
RIN3_HUMAN
|
||||||
| NC score | 0.041493 (rank : 19) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8TB24, Q8NF30, Q8TEE8, Q8WYP4, Q9H6A5, Q9HAG1 | Gene names | RIN3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
BAZ2A_MOUSE
|
||||||
| NC score | 0.041270 (rank : 20) | θ value | 1.38821 (rank : 19) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.040646 (rank : 21) | θ value | 0.62314 (rank : 12) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.040196 (rank : 22) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
TAU_MOUSE
|
||||||
| NC score | 0.040074 (rank : 23) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P10637, P10638, Q60684, Q60685, Q60686, Q62286 | Gene names | Mapt, Mtapt, Tau | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.038125 (rank : 24) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
R113A_HUMAN
|
||||||
| NC score | 0.036355 (rank : 25) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15541 | Gene names | RNF113A, RNF113, ZNF183 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 113A (Zinc finger protein 183). | |||||
|
MYO15_MOUSE
|
||||||
| NC score | 0.035260 (rank : 26) | θ value | 0.00665767 (rank : 4) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QZZ4, O70395, Q9QWL6 | Gene names | Myo15a, Myo15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
CHD4_MOUSE
|
||||||
| NC score | 0.034983 (rank : 27) | θ value | 0.0431538 (rank : 7) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
DIP2A_HUMAN
|
||||||
| NC score | 0.034624 (rank : 28) | θ value | 0.813845 (rank : 14) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14689, Q8IVA3, Q8N4S2, Q8TD89, Q96ML9 | Gene names | DIP2A, C21orf106, DIP2, KIAA0184 | |||
|
Domain Architecture |

|
|||||
| Description | Disco-interacting protein 2 homolog A. | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.033341 (rank : 29) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
PTTG1_HUMAN
|
||||||
| NC score | 0.033341 (rank : 30) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95997 | Gene names | PTTG1, EAP1, PTTG, TUTR1 | |||
|
Domain Architecture |

|
|||||
| Description | Securin (Pituitary tumor-transforming protein 1) (Tumor-transforming protein 1) (Esp1-associated protein) (hPTTG). | |||||
|
BRD4_MOUSE
|
||||||
| NC score | 0.032889 (rank : 31) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.031717 (rank : 32) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.031625 (rank : 33) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.030827 (rank : 34) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.030571 (rank : 35) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.030355 (rank : 36) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
MAP2_MOUSE
|
||||||
| NC score | 0.029825 (rank : 37) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P20357 | Gene names | Map2, Mtap2 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 2 (MAP 2). | |||||
|
SAM14_HUMAN
|
||||||
| NC score | 0.029763 (rank : 38) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IZD0, Q8N2X0 | Gene names | SAMD14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha motif domain-containing protein 14. | |||||
|
MARCS_HUMAN
|
||||||
| NC score | 0.028828 (rank : 39) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
LIMD1_HUMAN
|
||||||
| NC score | 0.028332 (rank : 40) | θ value | 0.47712 (rank : 9) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UGP4, Q9BQQ9, Q9NQ47 | Gene names | LIMD1 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain-containing protein 1. | |||||
|
NPHP4_HUMAN
|
||||||
| NC score | 0.028234 (rank : 41) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O75161 | Gene names | NPHP4, KIAA0673 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nephrocystin-4 (Nephroretinin). | |||||
|
AN13D_HUMAN
|
||||||
| NC score | 0.027828 (rank : 42) | θ value | 1.06291 (rank : 15) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZTN6, Q6ZVD0, Q86SU1 | Gene names | ANKRD13D | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 13D. | |||||
|
MAML1_MOUSE
|
||||||
| NC score | 0.027726 (rank : 43) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6T264, Q505D8, Q5SUC2, Q6PDK3, Q6ZQG5, Q8BIU5, Q8R3T0 | Gene names | Maml1, Kiaa0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
G7D_HUMAN
|
||||||
| NC score | 0.026467 (rank : 44) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5SSQ6, Q9Y335 | Gene names | C6orf26, G7D, NG23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein G7d. | |||||
|
CHD4_HUMAN
|
||||||
| NC score | 0.025618 (rank : 45) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
PHC1_HUMAN
|
||||||
| NC score | 0.025545 (rank : 46) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P78364, Q8WVM3, Q9BU63 | Gene names | PHC1, EDR1, PH1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (hPH1) (Early development regulatory protein 1). | |||||
|
MEGF9_MOUSE
|
||||||
| NC score | 0.024767 (rank : 47) | θ value | 1.38821 (rank : 21) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
ROBO3_HUMAN
|
||||||
| NC score | 0.024732 (rank : 48) | θ value | 0.0252991 (rank : 6) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96MS0 | Gene names | ROBO3 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Roundabout-like protein 3). | |||||
|
H15_HUMAN
|
||||||
| NC score | 0.024256 (rank : 49) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P16401, Q14529 | Gene names | HIST1H1B, H1F5 | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.5 (Histone H1a). | |||||
|
ODO2_HUMAN
|
||||||
| NC score | 0.023999 (rank : 50) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P36957, Q9BQ32 | Gene names | DLST, DLTS | |||
|
Domain Architecture |

|
|||||
| Description | Dihydrolipoyllysine-residue succinyltransferase component of 2- oxoglutarate dehydrogenase complex, mitochondrial precursor (EC 2.3.1.61) (Dihydrolipoamide succinyltransferase component of 2- oxoglutarate dehydrogenase complex) (E2) (E2K). | |||||
|
MAGI2_HUMAN
|
||||||
| NC score | 0.023918 (rank : 51) | θ value | 0.62314 (rank : 11) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.022017 (rank : 52) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.021965 (rank : 53) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
RUNX1_HUMAN
|
||||||
| NC score | 0.021525 (rank : 54) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q01196, O60472, O60473, O76047, O76089, Q13081, Q13755, Q13756, Q13757, Q13758, Q13759, Q15341, Q15343, Q16122, Q16284, Q16285, Q16286, Q16346, Q16347, Q92479 | Gene names | RUNX1, AML1, CBFA2 | |||
|
Domain Architecture |

|
|||||
| Description | Runt-related transcription factor 1 (Core-binding factor, alpha 2 subunit) (CBF-alpha 2) (Acute myeloid leukemia 1 protein) (Oncogene AML-1) (Polyomavirus enhancer-binding protein 2 alpha B subunit) (PEBP2-alpha B) (PEA2-alpha B) (SL3-3 enhancer factor 1 alpha B subunit) (SL3/AKV core-binding factor alpha B subunit). | |||||
|
TAF3_HUMAN
|
||||||
| NC score | 0.020864 (rank : 55) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.019942 (rank : 56) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
H14_HUMAN
|
||||||
| NC score | 0.019388 (rank : 57) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P10412 | Gene names | HIST1H1E, H1F4 | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.4 (Histone H1b). | |||||
|
IQEC3_HUMAN
|
||||||
| NC score | 0.019384 (rank : 58) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UPP2, Q8TB43 | Gene names | IQSEC3, KIAA1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
SSH2_HUMAN
|
||||||
| NC score | 0.019262 (rank : 59) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q76I76, Q8TDB5, Q8WYL1, Q8WYL2, Q96F40, Q96H36, Q9C0D8 | Gene names | SSH2, KIAA1725, SSH2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 2 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-2L) (hSSH-2L). | |||||
|
LDB3_HUMAN
|
||||||
| NC score | 0.019104 (rank : 60) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O75112, Q5K6N9, Q5K6P0, Q5K6P1, Q96FH2, Q9Y4Z3, Q9Y4Z4, Q9Y4Z5 | Gene names | LDB3, KIAA0613, ZASP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher). | |||||
|
CSKI1_MOUSE
|
||||||
| NC score | 0.019058 (rank : 61) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
S23IP_MOUSE
|
||||||
| NC score | 0.018692 (rank : 62) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6NZC7, Q8BXY6 | Gene names | Sec23ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SEC23-interacting protein. | |||||
|
H15_MOUSE
|
||||||
| NC score | 0.018657 (rank : 63) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P43276, Q9CRM8 | Gene names | Hist1h1b, H1f5 | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.5 (H1 VAR.5) (H1b). | |||||
|
DCJ14_MOUSE
|
||||||
| NC score | 0.018464 (rank : 64) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q921R4, Q3TX73, Q8BUU3, Q9CYB7 | Gene names | Dnajc14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 14. | |||||
|
IASPP_MOUSE
|
||||||
| NC score | 0.018334 (rank : 65) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
RIF1_HUMAN
|
||||||
| NC score | 0.018320 (rank : 66) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 320 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5UIP0, Q5H9R3, Q5UIP2, Q66YK6, Q6PRU2, Q8TE94, Q99772, Q9H830, Q9H9B9, Q9NVP5, Q9Y4R4 | Gene names | RIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog). | |||||
|
ODP2_HUMAN
|
||||||
| NC score | 0.018313 (rank : 67) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10515, Q16783 | Gene names | DLAT, DLTA | |||
|
Domain Architecture |

|
|||||
| Description | Dihydrolipoyllysine-residue acetyltransferase component of pyruvate dehydrogenase complex, mitochondrial precursor (EC 2.3.1.12) (Pyruvate dehydrogenase complex E2 subunit) (PDCE2) (E2) (Dihydrolipoamide S- acetyltransferase component of pyruvate dehydrogenase complex) (PDC- E2) (70 kDa mitochondrial autoantigen of primary biliary cirrhosis) (PBC) (M2 antigen complex 70 kDa subunit). | |||||
|
AFF1_MOUSE
|
||||||
| NC score | 0.018182 (rank : 68) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
RTEL1_HUMAN
|
||||||
| NC score | 0.017480 (rank : 69) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
TRI47_MOUSE
|
||||||
| NC score | 0.016902 (rank : 70) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C0E3, Q6P249, Q811J7, Q8BVZ8, Q8R1K0, Q8R3Y1 | Gene names | Trim47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 47. | |||||
|
ZN318_MOUSE
|
||||||
| NC score | 0.016872 (rank : 71) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 642 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99PP2, Q3TZL5, Q9JJ01 | Gene names | Znf318, Tzf, Zfp318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Testicular zinc finger protein). | |||||
|
COEA1_MOUSE
|
||||||
| NC score | 0.016799 (rank : 72) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80X19, Q8C6X3, Q9WV05 | Gene names | Col14a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor. | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.016578 (rank : 73) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
WNK2_HUMAN
|
||||||
| NC score | 0.016136 (rank : 74) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
EPN3_HUMAN
|
||||||
| NC score | 0.015756 (rank : 75) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H201, Q9BVN6, Q9NWK2 | Gene names | EPN3 | |||
|
Domain Architecture |

|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
COEA1_HUMAN
|
||||||
| NC score | 0.015277 (rank : 76) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q05707, O00260, O00261, O00262, Q05708, Q5XJ18, Q96C67 | Gene names | COL14A1, UND | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor (Undulin). | |||||
|
ZIMP7_HUMAN
|
||||||
| NC score | 0.014669 (rank : 77) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NF64, O94790, Q659A8, Q6JKL5, Q8WTX8, Q96Q01, Q9BQH7 | Gene names | ZIMP7, KIAA1886 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
MGAT1_MOUSE
|
||||||
| NC score | 0.014647 (rank : 78) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P27808 | Gene names | Mgat1, Gnt1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-1,3-mannosyl-glycoprotein 2-beta-N-acetylglucosaminyltransferase (EC 2.4.1.101) (N-glycosyl-oligosaccharide-glycoprotein N- acetylglucosaminyltransferase I) (GNT-I) (GlcNAc-T I). | |||||
|
ATF7_HUMAN
|
||||||
| NC score | 0.014377 (rank : 79) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17544, Q13814 | Gene names | ATF7, ATFA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
SREC2_HUMAN
|
||||||
| NC score | 0.013893 (rank : 80) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96GP6, Q8IXF3, Q9BW74 | Gene names | SCARF2, SREC2, SREPCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II) (SRECRP-1). | |||||
|
ATBF1_HUMAN
|
||||||
| NC score | 0.013845 (rank : 81) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
ATBF1_MOUSE
|
||||||
| NC score | 0.013648 (rank : 82) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
CXA7_MOUSE
|
||||||
| NC score | 0.013361 (rank : 83) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28229 | Gene names | Gja7, Cxn-45 | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-7 protein (Connexin-45) (Cx45). | |||||
|
CXA7_HUMAN
|
||||||
| NC score | 0.013275 (rank : 84) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P36383 | Gene names | GJA7 | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-7 protein (Connexin-45) (Cx45). | |||||
|
SPEG_HUMAN
|
||||||
| NC score | 0.013141 (rank : 85) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
HCN2_MOUSE
|
||||||
| NC score | 0.012388 (rank : 86) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O88703, O70506 | Gene names | Hcn2, Bcng2, Hac1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2) (Hyperpolarization-activated cation channel 1) (HAC-1). | |||||
|
TOP2A_MOUSE
|
||||||
| NC score | 0.011297 (rank : 87) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q01320 | Gene names | Top2a, Top-2, Top2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
ACK1_MOUSE
|
||||||
| NC score | 0.011117 (rank : 88) | θ value | 0.62314 (rank : 10) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1080 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O54967, Q8C2U0, Q8K0K4 | Gene names | Tnk2, Ack1 | |||
|
Domain Architecture |

|
|||||
| Description | Activated CDC42 kinase 1 (EC 2.7.10.2) (ACK-1) (Non-receptor protein tyrosine kinase Ack) (Tyrosine kinase non-receptor protein 2). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.010963 (rank : 89) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
KBTBB_HUMAN
|
||||||
| NC score | 0.010574 (rank : 90) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O94819 | Gene names | KBTBD11, KIAA0711, KLHDC7C | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch repeat and BTB domain-containing protein 11 (Kelch domain- containing protein 7B). | |||||
|
TTBK1_HUMAN
|
||||||
| NC score | 0.010530 (rank : 91) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1058 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5TCY1, Q2L6C6, Q8N444, Q96JH2 | Gene names | TTBK1, BDTK, KIAA1855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 1 (EC 2.7.11.1) (Brain-derived tau kinase). | |||||
|
IRX1_HUMAN
|
||||||
| NC score | 0.010444 (rank : 92) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78414, Q7Z2F8, Q8N312 | Gene names | IRX1, IRXA1 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-1 (Iroquois homeobox protein 1) (Homeodomain protein IRXA1). | |||||
|
KKCC1_MOUSE
|
||||||
| NC score | 0.010345 (rank : 93) | θ value | 1.06291 (rank : 17) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 866 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8VBY2, Q9R054 | Gene names | Camkk1, Camkk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium/calmodulin-dependent protein kinase kinase 1 (EC 2.7.11.17) (Calcium/calmodulin-dependent protein kinase kinase alpha) (CaM-kinase kinase alpha) (CaM-KK alpha) (CaMKK alpha) (CaMKK 1) (CaM-kinase IV kinase). | |||||
|
IF3A_HUMAN
|
||||||
| NC score | 0.010242 (rank : 94) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14152, O00653 | Gene names | EIF3S10, KIAA0139 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a). | |||||
|
KKCC1_HUMAN
|
||||||
| NC score | 0.007179 (rank : 95) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 858 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N5S9, Q9BQH3 | Gene names | CAMKK1, CAMKKA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium/calmodulin-dependent protein kinase kinase 1 (EC 2.7.11.17) (Calcium/calmodulin-dependent protein kinase kinase alpha) (CaM-kinase kinase alpha) (CaM-KK alpha) (CaMKK alpha) (CaMKK 1) (CaM-kinase IV kinase). | |||||
|
TSSK2_HUMAN
|
||||||
| NC score | 0.005156 (rank : 96) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 826 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96PF2, Q8IY55 | Gene names | TSSK2, DGSG, SPOGA2, STK22B | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific serine/threonine-protein kinase 2 (EC 2.7.11.1) (TSSK- 2) (Testis-specific kinase 2) (TSK-2) (Serine/threonine-protein kinase 22B) (DiGeorge syndrome protein G). | |||||
|
FOXC2_MOUSE
|
||||||
| NC score | 0.004746 (rank : 97) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61850, P97948, Q63869, Q8C694 | Gene names | Foxc2, Fkh14, Fkhl14, Mfh1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
ADA2B_MOUSE
|
||||||
| NC score | 0.000024 (rank : 98) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 891 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P30545 | Gene names | Adra2b | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-2B adrenergic receptor (Alpha-2B adrenoceptor) (Alpha-2B adrenoreceptor). | |||||