Please be patient as the page loads
|
CJ006_MOUSE
|
||||||
| SwissProt Accessions | Q6P9P0, Q8C041, Q8C0J5 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf6 homolog. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CJ006_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.937182 (rank : 2) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IX21, Q5W0L8, Q9NPE8 | Gene names | C10orf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf6. | |||||
|
CJ006_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 141 | |
| SwissProt Accessions | Q6P9P0, Q8C041, Q8C0J5 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf6 homolog. | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 3) | NC score | 0.118565 (rank : 4) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 4) | NC score | 0.050442 (rank : 60) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
SFR17_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 5) | NC score | 0.136734 (rank : 3) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 6) | NC score | 0.093697 (rank : 6) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 7) | NC score | 0.033275 (rank : 94) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 8) | NC score | 0.104620 (rank : 5) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
CD248_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 9) | NC score | 0.028016 (rank : 106) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91V98, Q3UMV6, Q91ZV1 | Gene names | Cd248, Tem1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
DHX37_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 10) | NC score | 0.037819 (rank : 76) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IY37, Q9BUI7, Q9P211 | Gene names | DHX37, DDX37, KIAA1517 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DHX37 (EC 3.6.1.-) (DEAH box protein 37). | |||||
|
RB3GP_HUMAN
|
||||||
| θ value | 0.125558 (rank : 11) | NC score | 0.068823 (rank : 19) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15042, Q659F5, Q8TBB4 | Gene names | RAB3GAP1, KIAA0066, RAB3GAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab3 GTPase-activating protein catalytic subunit (RAB3 GTPase- activating protein 130 kDa subunit) (Rab3-GAP p130) (Rab3-GAP). | |||||
|
SMCE1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 12) | NC score | 0.088629 (rank : 9) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q969G3, O43539 | Gene names | SMARCE1, BAF57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
NBN_MOUSE
|
||||||
| θ value | 0.163984 (rank : 13) | NC score | 0.064571 (rank : 25) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9R207, O88981, Q3UY57, Q811I6, Q8CCY0, Q9R1X1 | Gene names | Nbn, Nbs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nibrin (Nijmegen breakage syndrome protein 1 homolog) (Cell cycle regulatory protein p95). | |||||
|
SMCE1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 14) | NC score | 0.083103 (rank : 12) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O54941, Q8BPD9 | Gene names | Smarce1, Baf57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
BRCA1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.043076 (rank : 71) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P48754, Q60957, Q60983 | Gene names | Brca1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein homolog. | |||||
|
MYH8_HUMAN
|
||||||
| θ value | 0.21417 (rank : 16) | NC score | 0.037450 (rank : 79) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
CDKL5_HUMAN
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.011084 (rank : 144) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O76039, Q14198, Q5H985, Q9UJL6 | Gene names | CDKL5, STK9 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase-like 5 (EC 2.7.11.22) (Serine/threonine- protein kinase 9). | |||||
|
GRIN1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.045666 (rank : 67) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
NIN_MOUSE
|
||||||
| θ value | 0.279714 (rank : 19) | NC score | 0.053849 (rank : 50) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||
|
SEMG1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 20) | NC score | 0.069362 (rank : 18) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P04279, Q6X4I9, Q6Y809, Q6Y822, Q6Y823, Q86U64, Q96QM3 | Gene names | SEMG1, SEMG | |||
|
Domain Architecture |

|
|||||
| Description | Semenogelin-1 precursor (Semenogelin I) (SGI) [Contains: Alpha- inhibin-92; Alpha-inhibin-31; Seminal basic protein]. | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 21) | NC score | 0.089088 (rank : 8) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | 0.365318 (rank : 22) | NC score | 0.078354 (rank : 14) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
ATRX_HUMAN
|
||||||
| θ value | 0.365318 (rank : 23) | NC score | 0.063132 (rank : 29) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 24) | NC score | 0.091866 (rank : 7) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 0.365318 (rank : 25) | NC score | 0.059024 (rank : 39) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
RBMX2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 26) | NC score | 0.061907 (rank : 32) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
FGD6_HUMAN
|
||||||
| θ value | 0.47712 (rank : 27) | NC score | 0.022523 (rank : 118) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZV73, Q6ZR53, Q7Z2Z7, Q96D44, Q9NUR8, Q9P2I5 | Gene names | FGD6, KIAA1362, ZFYVE24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6 (Zinc finger FYVE domain-containing protein 24). | |||||
|
NIN_HUMAN
|
||||||
| θ value | 0.47712 (rank : 28) | NC score | 0.051933 (rank : 56) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
RAB20_HUMAN
|
||||||
| θ value | 0.47712 (rank : 29) | NC score | 0.016008 (rank : 129) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NX57, Q9NX49 | Gene names | RAB20 | |||
|
Domain Architecture |
|
|||||
| Description | Ras-related protein Rab-20. | |||||
|
SFR12_MOUSE
|
||||||
| θ value | 0.47712 (rank : 30) | NC score | 0.079491 (rank : 13) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
ZC313_HUMAN
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.086736 (rank : 10) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q5T200, O94936, Q5T1Z9, Q7Z7J3, Q8NDT6, Q9H0L6 | Gene names | ZC3H13, KIAA0853 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 13. | |||||
|
ZFP37_MOUSE
|
||||||
| θ value | 0.47712 (rank : 32) | NC score | 0.005943 (rank : 154) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P17141, Q62514 | Gene names | Zfp37, Zfp-37 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 37 (Zfp-37) (Male germ cell-specific zinc finger protein). | |||||
|
TARA_HUMAN
|
||||||
| θ value | 0.62314 (rank : 33) | NC score | 0.062157 (rank : 30) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
A4_HUMAN
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.034108 (rank : 90) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P05067, P09000, P78438, Q13764, Q13778, Q13793, Q16011, Q16014, Q16019, Q16020, Q9BT38, Q9UCA9, Q9UCB6, Q9UCC8, Q9UCD1, Q9UQ58 | Gene names | APP, A4, AD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein) (Cerebral vascular amyloid peptide) (CVAP) (Protease nexin-II) (PN-II) (APPI) (PreA4) [Contains: Soluble APP-alpha (S-APP- alpha); Soluble APP-beta (S-APP-beta); C99; Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)); Gamma-CTF(57) (Gamma- secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
MARK2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 35) | NC score | 0.005856 (rank : 155) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||
|
RTP3_MOUSE
|
||||||
| θ value | 0.813845 (rank : 36) | NC score | 0.043238 (rank : 70) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5QGU6, Q8VDC2 | Gene names | Rtp3, Tmem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-transporting protein 3 (Transmembrane protein 7). | |||||
|
ZN467_MOUSE
|
||||||
| θ value | 0.813845 (rank : 37) | NC score | 0.003014 (rank : 158) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 794 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8JZL0, Q9JJ98 | Gene names | Znf467, Ezi, Zfp467 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 467 (Endothelial cell-derived zinc finger protein) (EZI). | |||||
|
CENPJ_HUMAN
|
||||||
| θ value | 1.06291 (rank : 38) | NC score | 0.056674 (rank : 44) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9HC77, Q5T6R5, Q96KS5, Q9C067 | Gene names | CENPJ, CPAP, LAP, LIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein J (CENP-J) (Centrosomal P4.1-associated protein) (LAG-3-associated protein) (LYST-interacting protein 1). | |||||
|
PCX1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 39) | NC score | 0.048587 (rank : 64) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96RV3, O94897, Q96AI7, Q9Y2J9 | Gene names | PCNX, KIAA0805, KIAA0995, PCNXL1 | |||
|
Domain Architecture |

|
|||||
| Description | Pecanex-like protein 1 (Pecanex homolog). | |||||
|
PER2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.022644 (rank : 117) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
REST_MOUSE
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.036907 (rank : 82) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
TOP2A_MOUSE
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.035486 (rank : 85) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q01320 | Gene names | Top2a, Top-2, Top2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 43) | NC score | 0.068593 (rank : 20) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
EST31_MOUSE
|
||||||
| θ value | 1.38821 (rank : 44) | NC score | 0.015927 (rank : 131) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q63880 | Gene names | Es31 | |||
|
Domain Architecture |

|
|||||
| Description | Liver carboxylesterase 31 precursor (EC 3.1.1.1) (ES-Male) (Esterase- 31). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 45) | NC score | 0.059019 (rank : 40) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
MYH3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.035939 (rank : 83) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
NKTR_HUMAN
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.057567 (rank : 43) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 1.38821 (rank : 48) | NC score | 0.083833 (rank : 11) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
PPIG_HUMAN
|
||||||
| θ value | 1.38821 (rank : 49) | NC score | 0.061840 (rank : 33) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
RBMX2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 50) | NC score | 0.048777 (rank : 63) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8R0F5, Q3TI42 | Gene names | Rbmx2 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
ZAN_HUMAN
|
||||||
| θ value | 1.38821 (rank : 51) | NC score | 0.026315 (rank : 112) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y493, O00218, Q96L85, Q96L86, Q96L87, Q96L88, Q96L89, Q96L90, Q9BXN9, Q9BZ83, Q9BZ84, Q9BZ85, Q9BZ86, Q9BZ87, Q9BZ88 | Gene names | ZAN | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
MYH6_MOUSE
|
||||||
| θ value | 1.81305 (rank : 52) | NC score | 0.034470 (rank : 89) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 1.81305 (rank : 53) | NC score | 0.039022 (rank : 75) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
RNH2C_MOUSE
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.045726 (rank : 66) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CQ18, Q8R1N2 | Gene names | Rnaseh2c, Ayp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribonuclease H2 subunit C (RNase H2 subunit C) (Ribonuclease HI subunit C). | |||||
|
SFRS4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.067897 (rank : 21) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
TAGAP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.021905 (rank : 119) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N103, Q2NKM8, Q8NI40, Q96KZ2, Q96QA2 | Gene names | TAGAP, TAGAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell activation Rho GTPase-activating protein (T-cell activation GTPase-activating protein). | |||||
|
ZEP1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.012053 (rank : 139) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
ANR26_MOUSE
|
||||||
| θ value | 2.36792 (rank : 58) | NC score | 0.032435 (rank : 97) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
CENPC_HUMAN
|
||||||
| θ value | 2.36792 (rank : 59) | NC score | 0.060335 (rank : 35) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q03188, Q9P0M5 | Gene names | CENPC1, CENPC, ICEN7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C) (Interphase centromere complex protein 7). | |||||
|
CG034_HUMAN
|
||||||
| θ value | 2.36792 (rank : 60) | NC score | 0.058535 (rank : 41) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 2 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96L11, Q8TDV6 | Gene names | C7orf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C7orf34 precursor (MSSP-binding protein CTM-1). | |||||
|
CROCC_MOUSE
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.039161 (rank : 74) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
CROP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.061600 (rank : 34) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O95232, Q6PHR9, Q9NUY0, Q9P2S7 | Gene names | CROP, CREAP1, O48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein (cAMP regulatory element-associated protein 1) (CRE-associated protein 1) (CREAP-1) (Luc7A) (Okadaic acid-inducible phosphoprotein OA48-18). | |||||
|
GNL3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.029040 (rank : 103) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CI11, Q811R9, Q811S8, Q8BK21 | Gene names | Gnl3, Ns | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein-like 3 (Nucleolar GTP-binding protein 3) (Nucleostemin). | |||||
|
MAK_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.004342 (rank : 156) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 857 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20794, Q9NUH7 | Gene names | MAK | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MAK (EC 2.7.11.22) (Male germ cell- associated kinase). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.043959 (rank : 69) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
P52K_HUMAN
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.028967 (rank : 104) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43422, Q8WTW1, Q9Y3Z4 | Gene names | PRKRIR, DAP4, P52RIPK, THAP0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (Death-associated protein 4) (THAP domain-containing protein 0). | |||||
|
PIWL4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.016649 (rank : 128) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CGT6, Q8CC75 | Gene names | Piwil4, Miwi2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 4 (mAgo5). | |||||
|
YTDC1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.071380 (rank : 16) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
AP3B1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.031434 (rank : 98) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
CT2NL_MOUSE
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.042741 (rank : 72) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
GP179_HUMAN
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.035508 (rank : 84) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
LSP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 72) | NC score | 0.037281 (rank : 81) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19973, P97339, Q04950, Q62022, Q62023, Q62024, Q8CD28, Q99L65 | Gene names | Lsp1, Pp52, S37, Wp34 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte-specific protein 1 (Protein pp52) (52 kDa phosphoprotein) (Lymphocyte-specific antigen WP34) (S37 protein). | |||||
|
MYH13_HUMAN
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.034692 (rank : 88) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
NEK1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.006993 (rank : 151) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P51954 | Gene names | Nek1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek1 (EC 2.7.11.1) (NimA-related protein kinase 1). | |||||
|
SIX2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 75) | NC score | 0.011211 (rank : 142) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62232, P70179 | Gene names | Six2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein SIX2 (Sine oculis homeobox homolog 2). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.065994 (rank : 24) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
TSH1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.017947 (rank : 125) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5DTH5, Q6PE60, Q8VD19, Q9JLD5 | Gene names | Tshz1, Kiaa4206, Sdccag33, Tsh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 1 (Serologically defined colon cancer antigen 3 homolog). | |||||
|
WDHD1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 78) | NC score | 0.015960 (rank : 130) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75717 | Gene names | WDHD1, AND1 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
CDC37_MOUSE
|
||||||
| θ value | 4.03905 (rank : 79) | NC score | 0.042128 (rank : 73) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61081, Q3TGP0 | Gene names | Cdc37 | |||
|
Domain Architecture |
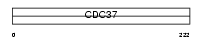
|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
ELL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | 0.020385 (rank : 120) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O08856 | Gene names | Ell | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL (Eleven-nineteen lysine-rich leukemia protein). | |||||
|
EVI1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | 0.003118 (rank : 157) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 826 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P14404 | Gene names | Evi1, Evi-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ecotropic virus integration 1 site protein. | |||||
|
FA21C_HUMAN
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.044208 (rank : 68) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y4E1, Q5SQU4, Q5SQU5, Q7L521, Q9UG79 | Gene names | FAM21C, KIAA0592 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM21C. | |||||
|
FA76B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | 0.026318 (rank : 111) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5HYJ3, Q6PIU3, Q8TC53 | Gene names | FAM76B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM76B. | |||||
|
MECP2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.033257 (rank : 95) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z2D6 | Gene names | Mecp2 | |||
|
Domain Architecture |
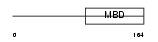
|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
MYH6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 85) | NC score | 0.033786 (rank : 91) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH8_MOUSE
|
||||||
| θ value | 4.03905 (rank : 86) | NC score | 0.033214 (rank : 96) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1600 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P13542, Q5SX36 | Gene names | Myh8, Myhsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
PHC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 87) | NC score | 0.018638 (rank : 124) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P78364, Q8WVM3, Q9BU63 | Gene names | PHC1, EDR1, PH1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (hPH1) (Early development regulatory protein 1). | |||||
|
PYGO1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 88) | NC score | 0.024579 (rank : 113) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y3Y4 | Gene names | PYGO1 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 1. | |||||
|
RB3GP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.050586 (rank : 59) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80UJ7, Q6A0D7, Q8C4Y0, Q8C679, Q8C6A5, Q8K324 | Gene names | Rab3gap1, Kiaa0066, Rab3gap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab3 GTPase-activating protein catalytic subunit (RAB3 GTPase- activating protein 130 kDa subunit) (Rab3-GAP p130) (Rab3-GAP). | |||||
|
SCN2A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.007851 (rank : 149) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99250, Q14472, Q9BZC9, Q9BZD0 | Gene names | SCN2A, NAC2, SCN2A2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 2 subunit alpha (Sodium channel protein type II subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.2) (Sodium channel protein, brain II subunit alpha) (HBSC II). | |||||
|
SNTA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.011228 (rank : 141) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13424, Q16438 | Gene names | SNTA1, SNT1 | |||
|
Domain Architecture |
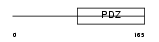
|
|||||
| Description | Alpha-1-syntrophin (59 kDa dystrophin-associated protein A1 acidic component 1) (Pro-TGF-alpha cytoplasmic domain-interacting protein 1) (TACIP1) (Syntrophin 1). | |||||
|
TOPRS_HUMAN
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.026478 (rank : 109) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NS56, O43273, Q6P987, Q9NS55, Q9UNR9 | Gene names | TOPORS, LUN, TP53BPL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
WWP2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.010707 (rank : 145) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
AF10_HUMAN
|
||||||
| θ value | 5.27518 (rank : 94) | NC score | 0.015757 (rank : 132) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P55197 | Gene names | MLLT10, AF10 | |||
|
Domain Architecture |
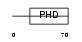
|
|||||
| Description | Protein AF-10. | |||||
|
CE110_HUMAN
|
||||||
| θ value | 5.27518 (rank : 95) | NC score | 0.023284 (rank : 116) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O43303, O43335, Q68DV9, Q8NE13 | Gene names | CEP110, CP110, KIAA0419 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 110 kDa (Cep110 protein). | |||||
|
CIR_MOUSE
|
||||||
| θ value | 5.27518 (rank : 96) | NC score | 0.066096 (rank : 23) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
DEK_MOUSE
|
||||||
| θ value | 5.27518 (rank : 97) | NC score | 0.064397 (rank : 26) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
LC7L2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.037360 (rank : 80) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7TNC4, Q99LM5, Q99PC3 | Gene names | Luc7l2 | |||
|
Domain Architecture |
|
|||||
| Description | Putative RNA-binding protein Luc7-like 2 (CGI-74 homolog). | |||||
|
LIPA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.033409 (rank : 92) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
RED_HUMAN
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.030793 (rank : 100) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13123 | Gene names | IK, RED, RER | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
RED_MOUSE
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.031041 (rank : 99) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z1M8 | Gene names | Ik, Red, Rer | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
RENT2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | 0.054920 (rank : 47) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9HAU5, Q8N8U1, Q9H1J2, Q9NWL1, Q9P2D9, Q9Y4M9 | Gene names | UPF2, KIAA1408, RENT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 2 (Nonsense mRNA reducing factor 2) (Up-frameshift suppressor 2 homolog) (hUpf2). | |||||
|
SPTA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 103) | NC score | 0.023894 (rank : 114) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P02549, Q15514, Q6LDY5 | Gene names | SPTA1, SPTA | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin alpha chain, erythrocyte (Erythroid alpha-spectrin). | |||||
|
TRHY_HUMAN
|
||||||
| θ value | 5.27518 (rank : 104) | NC score | 0.037739 (rank : 77) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
CASR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.006066 (rank : 153) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P41180, Q13912, Q16108, Q16109, Q16110, Q16379, Q4PJ19 | Gene names | CASR, GPRC2A, PCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Extracellular calcium-sensing receptor precursor (CaSR) (Parathyroid Cell calcium-sensing receptor). | |||||
|
CCNB3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.037588 (rank : 78) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WWL7, Q96SB5, Q96SB6, Q96SB7, Q9NT38 | Gene names | CCNB3, CYCB3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
DACT1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.026912 (rank : 108) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NYF0, Q86TY0 | Gene names | DACT1, DPR1, HNG3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dapper homolog 1 (hDPR1) (Heptacellular carcinoma novel gene 3 protein). | |||||
|
FGD3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.011100 (rank : 143) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.034991 (rank : 87) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
IFNW1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.007499 (rank : 150) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P05000 | Gene names | IFNW1 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon omega-1 precursor (Interferon alpha-II-1). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.047049 (rank : 65) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
MO4L2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.015643 (rank : 134) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15014, Q567V0, Q8J026 | Gene names | MORF4L2, KIAA0026, MRGX | |||
|
Domain Architecture |
|
|||||
| Description | Mortality factor 4-like protein 2 (MORF-related gene X protein) (Transcription factor-like protein MRGX) (MSL3-2 protein). | |||||
|
ANR12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.035058 (rank : 86) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1081 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q6UB98, O94951, Q658K1, Q6QMF7, Q9H231, Q9H784 | Gene names | ANKRD12, ANCO2, KIAA0874 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 12 (Ankyrin repeat-containing cofactor 2) (GAC-1 protein). | |||||
|
ARI4A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.053951 (rank : 49) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
CA026_HUMAN
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.019042 (rank : 123) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5T5J6, Q8NEK9, Q9BZQ7, Q9NXQ0 | Gene names | C1orf26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf26. | |||||
|
CDC37_HUMAN
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.033295 (rank : 93) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q16543 | Gene names | CDC37 | |||
|
Domain Architecture |
|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
CNGA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.008726 (rank : 147) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P29973, Q16279, Q16485 | Gene names | CNGA1, CNCG, CNCG1 | |||
|
Domain Architecture |
|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
CP135_HUMAN
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.030544 (rank : 101) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
CTGE2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.026469 (rank : 110) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96RT6 | Gene names | CTAGE1, CTAGE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein cTAGE-2. | |||||
|
EP15_HUMAN
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.023713 (rank : 115) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1021 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P42566 | Gene names | EPS15, AF1P | |||
|
Domain Architecture |
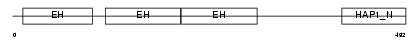
|
|||||
| Description | Epidermal growth factor receptor substrate 15 (Protein Eps15) (AF-1p protein). | |||||
|
FSBP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.016737 (rank : 127) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95073, Q8N4S5 | Gene names | FSBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibrinogen silencer-binding protein. | |||||
|
ITF2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.015698 (rank : 133) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P15884, Q15439, Q15440 | Gene names | TCF4, ITF2, SEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (SL3-3 enhancer factor 2) (SEF-2). | |||||
|
ITF2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.017120 (rank : 126) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q60722, Q60721, Q62211, Q64072, Q80UE8 | Gene names | Tcf4, Itf2, Sef2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (MITF-2) (SL3-3 enhancer factor 2) (SEF-2) (Class A helix-loop-helix transcription factor ME2). | |||||
|
JHD2A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.012121 (rank : 138) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6PCM1, Q2MJQ6, Q3TKW8, Q3UML3, Q6ZQ57, Q8K2J6, Q8K2K4, Q8R350 | Gene names | Jmjd1a, Jhdm2a, Kiaa0742 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2A (EC 1.14.11.-) (Jumonji domain-containing protein 1A). | |||||
|
K0408_HUMAN
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.027496 (rank : 107) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6ZU52, O43158, Q5TF20, Q7L2M2 | Gene names | KIAA0408 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0408. | |||||
|
KIF4A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.020045 (rank : 122) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
LASP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 127) | NC score | 0.006275 (rank : 152) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14847, Q96ED2, Q96IG0 | Gene names | LASP1, MLN50 | |||
|
Domain Architecture |
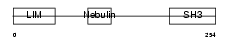
|
|||||
| Description | LIM and SH3 domain protein 1 (LASP-1) (MLN 50). | |||||
|
LMO7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 128) | NC score | 0.030347 (rank : 102) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
MYST4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 129) | NC score | 0.028726 (rank : 105) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
NT5D1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 130) | NC score | 0.014269 (rank : 135) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C5P5, Q3TAL3, Q3TPT8, Q3TTI3, Q3UL73, Q4G0D1 | Gene names | Nt5dc1, Nt5c2l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5'-nucleotidase domain-containing protein 1 (5'-nucleotidase cytosolic II-like protein 1). | |||||
|
NUD12_MOUSE
|
||||||
| θ value | 8.99809 (rank : 131) | NC score | 0.012762 (rank : 137) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DCN1, Q6PFA5 | Gene names | Nudt12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisomal NADH pyrophosphatase NUDT12 (EC 3.6.1.22) (Nucleoside diphosphate-linked moiety X motif 12) (Nudix motif 12). | |||||
|
PARD3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 132) | NC score | 0.010579 (rank : 146) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8TEW0, Q8TEW1, Q8TEW2, Q8TEW3, Q96K28, Q96RM6, Q96RM7, Q9BY57, Q9BY58, Q9HC48, Q9NWL4, Q9NYE6 | Gene names | PARD3, PAR3, PAR3A | |||
|
Domain Architecture |

|
|||||
| Description | Partitioning-defective 3 homolog (PARD-3) (PAR-3) (Atypical PKC isotype-specific-interacting protein) (ASIP) (CTCL tumor antigen se2- 5) (PAR3-alpha). | |||||
|
PRLPN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.008392 (rank : 148) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CGZ9, Q9DAX1 | Gene names | Prlpn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Placental prolactin-like protein N precursor (PRL-like protein N) (PLP-N). | |||||
|
SFR12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | 0.067491 (rank : 22) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
SFRS4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 135) | NC score | 0.052653 (rank : 53) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
SIA7A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.013772 (rank : 136) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9QZ39, Q9JJP5 | Gene names | St6galnac1, Siat7a | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase 1 (EC 2.4.99.3) (GalNAc alpha-2,6-sialyltransferase I) (ST6GalNAc I) (Sialyltransferase 7A). | |||||
|
SNCAP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.020261 (rank : 121) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y6H5 | Gene names | SNCAIP | |||
|
Domain Architecture |

|
|||||
| Description | Synphilin-1 (Alpha-synuclein-interacting protein). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.051945 (rank : 55) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TCOF_HUMAN
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.048950 (rank : 62) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
TENS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.011569 (rank : 140) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9HBL0 | Gene names | TNS1, TNS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-1. | |||||
|
THOC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.059151 (rank : 38) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.052361 (rank : 54) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
CEP35_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.050229 (rank : 61) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
CIR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.064360 (rank : 27) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
CROP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.050606 (rank : 58) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5SUF2, Q3U9D5, Q8BUJ5, Q921Z3, Q9CRS7, Q9CTY4 | Gene names | Crop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein. | |||||
|
CYLC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.059512 (rank : 37) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.056068 (rank : 45) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
DMP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.053961 (rank : 48) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.053269 (rank : 52) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
ELOA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.063653 (rank : 28) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
ENL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.053545 (rank : 51) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.061941 (rank : 31) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.069376 (rank : 17) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.055401 (rank : 46) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.072707 (rank : 15) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
SFR11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.058463 (rank : 42) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
SPRL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.051648 (rank : 57) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.059930 (rank : 36) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
CJ006_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 141 | |
| SwissProt Accessions | Q6P9P0, Q8C041, Q8C0J5 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf6 homolog. | |||||
|
CJ006_HUMAN
|
||||||
| NC score | 0.937182 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IX21, Q5W0L8, Q9NPE8 | Gene names | C10orf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf6. | |||||
|
SFR17_HUMAN
|
||||||
| NC score | 0.136734 (rank : 3) | θ value | 0.00665767 (rank : 5) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.118565 (rank : 4) | θ value | 0.00020696 (rank : 3) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.104620 (rank : 5) | θ value | 0.0736092 (rank : 8) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.093697 (rank : 6) | θ value | 0.0193708 (rank : 6) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.091866 (rank : 7) | θ value | 0.365318 (rank : 24) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.089088 (rank : 8) | θ value | 0.279714 (rank : 21) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
SMCE1_HUMAN
|
||||||
| NC score | 0.088629 (rank : 9) | θ value | 0.125558 (rank : 12) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q969G3, O43539 | Gene names | SMARCE1, BAF57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
ZC313_HUMAN
|
||||||
| NC score | 0.086736 (rank : 10) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q5T200, O94936, Q5T1Z9, Q7Z7J3, Q8NDT6, Q9H0L6 | Gene names | ZC3H13, KIAA0853 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 13. | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.083833 (rank : 11) | θ value | 1.38821 (rank : 48) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
SMCE1_MOUSE
|
||||||
| NC score | 0.083103 (rank : 12) | θ value | 0.163984 (rank : 14) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O54941, Q8BPD9 | Gene names | Smarce1, Baf57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
SFR12_MOUSE
|
||||||
| NC score | 0.079491 (rank : 13) | θ value | 0.47712 (rank : 30) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.078354 (rank : 14) | θ value | 0.365318 (rank : 22) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.072707 (rank : 15) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
YTDC1_HUMAN
|
||||||
| NC score | 0.071380 (rank : 16) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.069376 (rank : 17) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
SEMG1_HUMAN
|
||||||
| NC score | 0.069362 (rank : 18) | θ value | 0.279714 (rank : 20) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P04279, Q6X4I9, Q6Y809, Q6Y822, Q6Y823, Q86U64, Q96QM3 | Gene names | SEMG1, SEMG | |||
|
Domain Architecture |

|
|||||
| Description | Semenogelin-1 precursor (Semenogelin I) (SGI) [Contains: Alpha- inhibin-92; Alpha-inhibin-31; Seminal basic protein]. | |||||
|
RB3GP_HUMAN
|
||||||
| NC score | 0.068823 (rank : 19) | θ value | 0.125558 (rank : 11) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15042, Q659F5, Q8TBB4 | Gene names | RAB3GAP1, KIAA0066, RAB3GAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab3 GTPase-activating protein catalytic subunit (RAB3 GTPase- activating protein 130 kDa subunit) (Rab3-GAP p130) (Rab3-GAP). | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.068593 (rank : 20) | θ value | 1.38821 (rank : 43) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
SFRS4_HUMAN
|
||||||
| NC score | 0.067897 (rank : 21) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
SFR12_HUMAN
|
||||||
| NC score | 0.067491 (rank : 22) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
CIR_MOUSE
|
||||||
| NC score | 0.066096 (rank : 23) | θ value | 5.27518 (rank : 96) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.065994 (rank : 24) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
NBN_MOUSE
|
||||||
| NC score | 0.064571 (rank : 25) | θ value | 0.163984 (rank : 13) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9R207, O88981, Q3UY57, Q811I6, Q8CCY0, Q9R1X1 | Gene names | Nbn, Nbs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nibrin (Nijmegen breakage syndrome protein 1 homolog) (Cell cycle regulatory protein p95). | |||||
|
DEK_MOUSE
|
||||||
| NC score | 0.064397 (rank : 26) | θ value | 5.27518 (rank : 97) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
CIR_HUMAN
|
||||||
| NC score | 0.064360 (rank : 27) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
ELOA1_HUMAN
|
||||||
| NC score | 0.063653 (rank : 28) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
ATRX_HUMAN
|
||||||
| NC score | 0.063132 (rank : 29) | θ value | 0.365318 (rank : 23) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.062157 (rank : 30) | θ value | 0.62314 (rank : 33) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.061941 (rank : 31) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
RBMX2_HUMAN
|
||||||
| NC score | 0.061907 (rank : 32) | θ value | 0.365318 (rank : 26) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
PPIG_HUMAN
|
||||||
| NC score | 0.061840 (rank : 33) | θ value | 1.38821 (rank : 49) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
CROP_HUMAN
|
||||||
| NC score | 0.061600 (rank : 34) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O95232, Q6PHR9, Q9NUY0, Q9P2S7 | Gene names | CROP, CREAP1, O48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein (cAMP regulatory element-associated protein 1) (CRE-associated protein 1) (CREAP-1) (Luc7A) (Okadaic acid-inducible phosphoprotein OA48-18). | |||||
|
CENPC_HUMAN
|
||||||
| NC score | 0.060335 (rank : 35) | θ value | 2.36792 (rank : 59) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q03188, Q9P0M5 | Gene names | CENPC1, CENPC, ICEN7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C) (Interphase centromere complex protein 7). | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.059930 (rank : 36) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
CYLC2_HUMAN
|
||||||
| NC score | 0.059512 (rank : 37) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
THOC2_HUMAN
|
||||||
| NC score | 0.059151 (rank : 38) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.059024 (rank : 39) | θ value | 0.365318 (rank : 25) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.059019 (rank : 40) | θ value | 1.38821 (rank : 45) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
CG034_HUMAN
|
||||||
| NC score | 0.058535 (rank : 41) | θ value | 2.36792 (rank : 60) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 2 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96L11, Q8TDV6 | Gene names | C7orf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C7orf34 precursor (MSSP-binding protein CTM-1). | |||||
|
SFR11_HUMAN
|
||||||
| NC score | 0.058463 (rank : 42) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
NKTR_HUMAN
|
||||||
| NC score | 0.057567 (rank : 43) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
CENPJ_HUMAN
|
||||||
| NC score | 0.056674 (rank : 44) | θ value | 1.06291 (rank : 38) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9HC77, Q5T6R5, Q96KS5, Q9C067 | Gene names | CENPJ, CPAP, LAP, LIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein J (CENP-J) (Centrosomal P4.1-associated protein) (LAG-3-associated protein) (LYST-interacting protein 1). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.056068 (rank : 45) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.055401 (rank : 46) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
RENT2_HUMAN
|
||||||
| NC score | 0.054920 (rank : 47) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9HAU5, Q8N8U1, Q9H1J2, Q9NWL1, Q9P2D9, Q9Y4M9 | Gene names | UPF2, KIAA1408, RENT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 2 (Nonsense mRNA reducing factor 2) (Up-frameshift suppressor 2 homolog) (hUpf2). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.053961 (rank : 48) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
ARI4A_HUMAN
|
||||||
| NC score | 0.053951 (rank : 49) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
NIN_MOUSE
|
||||||
| NC score | 0.053849 (rank : 50) | θ value | 0.279714 (rank : 19) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||
|
ENL_HUMAN
|
||||||
| NC score | 0.053545 (rank : 51) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.053269 (rank : 52) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
SFRS4_MOUSE
|
||||||
| NC score | 0.052653 (rank : 53) | θ value | 8.99809 (rank : 135) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.052361 (rank : 54) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.051945 (rank : 55) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
NIN_HUMAN
|
||||||
| NC score | 0.051933 (rank : 56) | θ value | 0.47712 (rank : 28) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
SPRL1_MOUSE
|
||||||
| NC score | 0.051648 (rank : 57) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
CROP_MOUSE
|
||||||
| NC score | 0.050606 (rank : 58) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5SUF2, Q3U9D5, Q8BUJ5, Q921Z3, Q9CRS7, Q9CTY4 | Gene names | Crop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein. | |||||
|
RB3GP_MOUSE
|
||||||
| NC score | 0.050586 (rank : 59) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80UJ7, Q6A0D7, Q8C4Y0, Q8C679, Q8C6A5, Q8K324 | Gene names | Rab3gap1, Kiaa0066, Rab3gap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab3 GTPase-activating protein catalytic subunit (RAB3 GTPase- activating protein 130 kDa subunit) (Rab3-GAP p130) (Rab3-GAP). | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.050442 (rank : 60) | θ value | 0.00390308 (rank : 4) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
CEP35_HUMAN
|
||||||
| NC score | 0.050229 (rank : 61) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
TCOF_HUMAN
|
||||||
| NC score | 0.048950 (rank : 62) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
RBMX2_MOUSE
|
||||||
| NC score | 0.048777 (rank : 63) | θ value | 1.38821 (rank : 50) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8R0F5, Q3TI42 | Gene names | Rbmx2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
PCX1_HUMAN
|
||||||
| NC score | 0.048587 (rank : 64) | θ value | 1.06291 (rank : 39) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96RV3, O94897, Q96AI7, Q9Y2J9 | Gene names | PCNX, KIAA0805, KIAA0995, PCNXL1 | |||
|
Domain Architecture |

|
|||||
| Description | Pecanex-like protein 1 (Pecanex homolog). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.047049 (rank : 65) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
RNH2C_MOUSE
|
||||||
| NC score | 0.045726 (rank : 66) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CQ18, Q8R1N2 | Gene names | Rnaseh2c, Ayp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribonuclease H2 subunit C (RNase H2 subunit C) (Ribonuclease HI subunit C). | |||||
|
GRIN1_HUMAN
|
||||||
| NC score | 0.045666 (rank : 67) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
FA21C_HUMAN
|
||||||
| NC score | 0.044208 (rank : 68) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y4E1, Q5SQU4, Q5SQU5, Q7L521, Q9UG79 | Gene names | FAM21C, KIAA0592 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM21C. | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.043959 (rank : 69) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
RTP3_MOUSE
|
||||||
| NC score | 0.043238 (rank : 70) | θ value | 0.813845 (rank : 36) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5QGU6, Q8VDC2 | Gene names | Rtp3, Tmem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-transporting protein 3 (Transmembrane protein 7). | |||||
|
BRCA1_MOUSE
|
||||||
| NC score | 0.043076 (rank : 71) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P48754, Q60957, Q60983 | Gene names | Brca1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein homolog. | |||||
|
CT2NL_MOUSE
|
||||||
| NC score | 0.042741 (rank : 72) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
CDC37_MOUSE
|
||||||
| NC score | 0.042128 (rank : 73) | θ value | 4.03905 (rank : 79) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61081, Q3TGP0 | Gene names | Cdc37 | |||
|
Domain Architecture |

|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
CROCC_MOUSE
|
||||||
| NC score | 0.039161 (rank : 74) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.039022 (rank : 75) | θ value | 1.81305 (rank : 53) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
DHX37_HUMAN
|
||||||
| NC score | 0.037819 (rank : 76) | θ value | 0.0961366 (rank : 10) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IY37, Q9BUI7, Q9P211 | Gene names | DHX37, DDX37, KIAA1517 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DHX37 (EC 3.6.1.-) (DEAH box protein 37). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.037739 (rank : 77) | θ value | 5.27518 (rank : 104) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
CCNB3_HUMAN
|
||||||
| NC score | 0.037588 (rank : 78) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WWL7, Q96SB5, Q96SB6, Q96SB7, Q9NT38 | Gene names | CCNB3, CYCB3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
MYH8_HUMAN
|
||||||
| NC score | 0.037450 (rank : 79) | θ value | 0.21417 (rank : 16) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
LC7L2_MOUSE
|
||||||
| NC score | 0.037360 (rank : 80) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7TNC4, Q99LM5, Q99PC3 | Gene names | Luc7l2 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein Luc7-like 2 (CGI-74 homolog). | |||||
|
LSP1_MOUSE
|
||||||
| NC score | 0.037281 (rank : 81) | θ value | 3.0926 (rank : 72) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19973, P97339, Q04950, Q62022, Q62023, Q62024, Q8CD28, Q99L65 | Gene names | Lsp1, Pp52, S37, Wp34 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte-specific protein 1 (Protein pp52) (52 kDa phosphoprotein) (Lymphocyte-specific antigen WP34) (S37 protein). | |||||
|
REST_MOUSE
|
||||||
| NC score | 0.036907 (rank : 82) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
MYH3_HUMAN
|
||||||
| NC score | 0.035939 (rank : 83) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
GP179_HUMAN
|
||||||
| NC score | 0.035508 (rank : 84) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
TOP2A_MOUSE
|
||||||
| NC score | 0.035486 (rank : 85) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q01320 | Gene names | Top2a, Top-2, Top2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
ANR12_HUMAN
|
||||||
| NC score | 0.035058 (rank : 86) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1081 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q6UB98, O94951, Q658K1, Q6QMF7, Q9H231, Q9H784 | Gene names | ANKRD12, ANCO2, KIAA0874 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 12 (Ankyrin repeat-containing cofactor 2) (GAC-1 protein). | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.034991 (rank : 87) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
MYH13_HUMAN
|
||||||
| NC score | 0.034692 (rank : 88) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
MYH6_MOUSE
|
||||||
| NC score | 0.034470 (rank : 89) | θ value | 1.81305 (rank : 52) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
A4_HUMAN
|
||||||
| NC score | 0.034108 (rank : 90) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P05067, P09000, P78438, Q13764, Q13778, Q13793, Q16011, Q16014, Q16019, Q16020, Q9BT38, Q9UCA9, Q9UCB6, Q9UCC8, Q9UCD1, Q9UQ58 | Gene names | APP, A4, AD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein) (Cerebral vascular amyloid peptide) (CVAP) (Protease nexin-II) (PN-II) (APPI) (PreA4) [Contains: Soluble APP-alpha (S-APP- alpha); Soluble APP-beta (S-APP-beta); C99; Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)); Gamma-CTF(57) (Gamma- secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
MYH6_HUMAN
|
||||||
| NC score | 0.033786 (rank : 91) | θ value | 4.03905 (rank : 85) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
LIPA1_HUMAN
|
||||||
| NC score | 0.033409 (rank : 92) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
CDC37_HUMAN
|
||||||
| NC score | 0.033295 (rank : 93) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q16543 | Gene names | CDC37 | |||
|
Domain Architecture |

|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.033275 (rank : 94) | θ value | 0.0431538 (rank : 7) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
MECP2_MOUSE
|
||||||
| NC score | 0.033257 (rank : 95) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z2D6 | Gene names | Mecp2 | |||
|
Domain Architecture |

|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
MYH8_MOUSE
|
||||||
| NC score | 0.033214 (rank : 96) | θ value | 4.03905 (rank : 86) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1600 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P13542, Q5SX36 | Gene names | Myh8, Myhsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
ANR26_MOUSE
|
||||||
| NC score | 0.032435 (rank : 97) | θ value | 2.36792 (rank : 58) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
AP3B1_HUMAN
|
||||||
| NC score | 0.031434 (rank : 98) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
RED_MOUSE
|
||||||
| NC score | 0.031041 (rank : 99) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z1M8 | Gene names | Ik, Red, Rer | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
RED_HUMAN
|
||||||
| NC score | 0.030793 (rank : 100) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13123 | Gene names | IK, RED, RER | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
CP135_HUMAN
|
||||||
| NC score | 0.030544 (rank : 101) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
LMO7_HUMAN
|
||||||
| NC score | 0.030347 (rank : 102) | θ value | 8.99809 (rank : 128) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
GNL3_MOUSE
|
||||||
| NC score | 0.029040 (rank : 103) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CI11, Q811R9, Q811S8, Q8BK21 | Gene names | Gnl3, Ns | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein-like 3 (Nucleolar GTP-binding protein 3) (Nucleostemin). | |||||
|
P52K_HUMAN
|
||||||
| NC score | 0.028967 (rank : 104) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43422, Q8WTW1, Q9Y3Z4 | Gene names | PRKRIR, DAP4, P52RIPK, THAP0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (Death-associated protein 4) (THAP domain-containing protein 0). | |||||
|
MYST4_HUMAN
|
||||||
| NC score | 0.028726 (rank : 105) | θ value | 8.99809 (rank : 129) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
CD248_MOUSE
|
||||||
| NC score | 0.028016 (rank : 106) | θ value | 0.0961366 (rank : 9) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91V98, Q3UMV6, Q91ZV1 | Gene names | Cd248, Tem1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
K0408_HUMAN
|
||||||
| NC score | 0.027496 (rank : 107) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6ZU52, O43158, Q5TF20, Q7L2M2 | Gene names | KIAA0408 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0408. | |||||
|
DACT1_HUMAN
|
||||||
| NC score | 0.026912 (rank : 108) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NYF0, Q86TY0 | Gene names | DACT1, DPR1, HNG3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dapper homolog 1 (hDPR1) (Heptacellular carcinoma novel gene 3 protein). | |||||
|
TOPRS_HUMAN
|
||||||
| NC score | 0.026478 (rank : 109) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NS56, O43273, Q6P987, Q9NS55, Q9UNR9 | Gene names | TOPORS, LUN, TP53BPL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
CTGE2_HUMAN
|
||||||
| NC score | 0.026469 (rank : 110) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96RT6 | Gene names | CTAGE1, CTAGE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein cTAGE-2. | |||||
|
FA76B_HUMAN
|
||||||
| NC score | 0.026318 (rank : 111) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5HYJ3, Q6PIU3, Q8TC53 | Gene names | FAM76B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM76B. | |||||
|
ZAN_HUMAN
|
||||||
| NC score | 0.026315 (rank : 112) | θ value | 1.38821 (rank : 51) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y493, O00218, Q96L85, Q96L86, Q96L87, Q96L88, Q96L89, Q96L90, Q9BXN9, Q9BZ83, Q9BZ84, Q9BZ85, Q9BZ86, Q9BZ87, Q9BZ88 | Gene names | ZAN | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
PYGO1_HUMAN
|
||||||
| NC score | 0.024579 (rank : 113) | θ value | 4.03905 (rank : 88) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y3Y4 | Gene names | PYGO1 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 1. | |||||
|
SPTA1_HUMAN
|
||||||
| NC score | 0.023894 (rank : 114) | θ value | 5.27518 (rank : 103) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P02549, Q15514, Q6LDY5 | Gene names | SPTA1, SPTA | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin alpha chain, erythrocyte (Erythroid alpha-spectrin). | |||||
|
EP15_HUMAN
|
||||||
| NC score | 0.023713 (rank : 115) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1021 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P42566 | Gene names | EPS15, AF1P | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15 (Protein Eps15) (AF-1p protein). | |||||
|
CE110_HUMAN
|
||||||
| NC score | 0.023284 (rank : 116) | θ value | 5.27518 (rank : 95) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O43303, O43335, Q68DV9, Q8NE13 | Gene names | CEP110, CP110, KIAA0419 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 110 kDa (Cep110 protein). | |||||
|
PER2_MOUSE
|
||||||
| NC score | 0.022644 (rank : 117) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
FGD6_HUMAN
|
||||||
| NC score | 0.022523 (rank : 118) | θ value | 0.47712 (rank : 27) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZV73, Q6ZR53, Q7Z2Z7, Q96D44, Q9NUR8, Q9P2I5 | Gene names | FGD6, KIAA1362, ZFYVE24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6 (Zinc finger FYVE domain-containing protein 24). | |||||
|
TAGAP_HUMAN
|
||||||
| NC score | 0.021905 (rank : 119) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N103, Q2NKM8, Q8NI40, Q96KZ2, Q96QA2 | Gene names | TAGAP, TAGAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell activation Rho GTPase-activating protein (T-cell activation GTPase-activating protein). | |||||
|
ELL_MOUSE
|
||||||
| NC score | 0.020385 (rank : 120) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O08856 | Gene names | Ell | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL (Eleven-nineteen lysine-rich leukemia protein). | |||||
|
SNCAP_HUMAN
|
||||||
| NC score | 0.020261 (rank : 121) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y6H5 | Gene names | SNCAIP | |||
|
Domain Architecture |

|
|||||
| Description | Synphilin-1 (Alpha-synuclein-interacting protein). | |||||
|
KIF4A_MOUSE
|
||||||
| NC score | 0.020045 (rank : 122) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
CA026_HUMAN
|
||||||
| NC score | 0.019042 (rank : 123) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5T5J6, Q8NEK9, Q9BZQ7, Q9NXQ0 | Gene names | C1orf26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf26. | |||||
|
PHC1_HUMAN
|
||||||
| NC score | 0.018638 (rank : 124) | θ value | 4.03905 (rank : 87) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P78364, Q8WVM3, Q9BU63 | Gene names | PHC1, EDR1, PH1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (hPH1) (Early development regulatory protein 1). | |||||
|
TSH1_MOUSE
|
||||||
| NC score | 0.017947 (rank : 125) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5DTH5, Q6PE60, Q8VD19, Q9JLD5 | Gene names | Tshz1, Kiaa4206, Sdccag33, Tsh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 1 (Serologically defined colon cancer antigen 3 homolog). | |||||
|
ITF2_MOUSE
|
||||||
| NC score | 0.017120 (rank : 126) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q60722, Q60721, Q62211, Q64072, Q80UE8 | Gene names | Tcf4, Itf2, Sef2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (MITF-2) (SL3-3 enhancer factor 2) (SEF-2) (Class A helix-loop-helix transcription factor ME2). | |||||
|
FSBP_HUMAN
|
||||||
| NC score | 0.016737 (rank : 127) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95073, Q8N4S5 | Gene names | FSBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibrinogen silencer-binding protein. | |||||
|
PIWL4_MOUSE
|
||||||
| NC score | 0.016649 (rank : 128) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CGT6, Q8CC75 | Gene names | Piwil4, Miwi2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 4 (mAgo5). | |||||
|
RAB20_HUMAN
|
||||||
| NC score | 0.016008 (rank : 129) | θ value | 0.47712 (rank : 29) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NX57, Q9NX49 | Gene names | RAB20 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-related protein Rab-20. | |||||
|
WDHD1_HUMAN
|
||||||
| NC score | 0.015960 (rank : 130) | θ value | 3.0926 (rank : 78) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75717 | Gene names | WDHD1, AND1 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
EST31_MOUSE
|
||||||
| NC score | 0.015927 (rank : 131) | θ value | 1.38821 (rank : 44) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q63880 | Gene names | Es31 | |||
|
Domain Architecture |

|
|||||
| Description | Liver carboxylesterase 31 precursor (EC 3.1.1.1) (ES-Male) (Esterase- 31). | |||||
|
AF10_HUMAN
|
||||||
| NC score | 0.015757 (rank : 132) | θ value | 5.27518 (rank : 94) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P55197 | Gene names | MLLT10, AF10 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-10. | |||||
|
ITF2_HUMAN
|
||||||
| NC score | 0.015698 (rank : 133) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P15884, Q15439, Q15440 | Gene names | TCF4, ITF2, SEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (SL3-3 enhancer factor 2) (SEF-2). | |||||
|
MO4L2_HUMAN
|
||||||
| NC score | 0.015643 (rank : 134) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15014, Q567V0, Q8J026 | Gene names | MORF4L2, KIAA0026, MRGX | |||
|
Domain Architecture |

|
|||||
| Description | Mortality factor 4-like protein 2 (MORF-related gene X protein) (Transcription factor-like protein MRGX) (MSL3-2 protein). | |||||
|
NT5D1_MOUSE
|
||||||
| NC score | 0.014269 (rank : 135) | θ value | 8.99809 (rank : 130) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C5P5, Q3TAL3, Q3TPT8, Q3TTI3, Q3UL73, Q4G0D1 | Gene names | Nt5dc1, Nt5c2l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5'-nucleotidase domain-containing protein 1 (5'-nucleotidase cytosolic II-like protein 1). | |||||
|
SIA7A_MOUSE
|
||||||
| NC score | 0.013772 (rank : 136) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9QZ39, Q9JJP5 | Gene names | St6galnac1, Siat7a | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase 1 (EC 2.4.99.3) (GalNAc alpha-2,6-sialyltransferase I) (ST6GalNAc I) (Sialyltransferase 7A). | |||||
|
NUD12_MOUSE
|
||||||
| NC score | 0.012762 (rank : 137) | θ value | 8.99809 (rank : 131) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DCN1, Q6PFA5 | Gene names | Nudt12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisomal NADH pyrophosphatase NUDT12 (EC 3.6.1.22) (Nucleoside diphosphate-linked moiety X motif 12) (Nudix motif 12). | |||||
|
JHD2A_MOUSE
|
||||||
| NC score | 0.012121 (rank : 138) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6PCM1, Q2MJQ6, Q3TKW8, Q3UML3, Q6ZQ57, Q8K2J6, Q8K2K4, Q8R350 | Gene names | Jmjd1a, Jhdm2a, Kiaa0742 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2A (EC 1.14.11.-) (Jumonji domain-containing protein 1A). | |||||
|
ZEP1_HUMAN
|
||||||
| NC score | 0.012053 (rank : 139) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
TENS1_HUMAN
|
||||||
| NC score | 0.011569 (rank : 140) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9HBL0 | Gene names | TNS1, TNS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-1. | |||||
|
SNTA1_HUMAN
|
||||||
| NC score | 0.011228 (rank : 141) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13424, Q16438 | Gene names | SNTA1, SNT1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-1-syntrophin (59 kDa dystrophin-associated protein A1 acidic component 1) (Pro-TGF-alpha cytoplasmic domain-interacting protein 1) (TACIP1) (Syntrophin 1). | |||||
|
SIX2_MOUSE
|
||||||
| NC score | 0.011211 (rank : 142) | θ value | 3.0926 (rank : 75) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62232, P70179 | Gene names | Six2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein SIX2 (Sine oculis homeobox homolog 2). | |||||
|
FGD3_HUMAN
|
||||||
| NC score | 0.011100 (rank : 143) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
CDKL5_HUMAN
|
||||||
| NC score | 0.011084 (rank : 144) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O76039, Q14198, Q5H985, Q9UJL6 | Gene names | CDKL5, STK9 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase-like 5 (EC 2.7.11.22) (Serine/threonine- protein kinase 9). | |||||
|
WWP2_HUMAN
|
||||||
| NC score | 0.010707 (rank : 145) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
PARD3_HUMAN
|
||||||
| NC score | 0.010579 (rank : 146) | θ value | 8.99809 (rank : 132) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8TEW0, Q8TEW1, Q8TEW2, Q8TEW3, Q96K28, Q96RM6, Q96RM7, Q9BY57, Q9BY58, Q9HC48, Q9NWL4, Q9NYE6 | Gene names | PARD3, PAR3, PAR3A | |||
|
Domain Architecture |

|
|||||
| Description | Partitioning-defective 3 homolog (PARD-3) (PAR-3) (Atypical PKC isotype-specific-interacting protein) (ASIP) (CTCL tumor antigen se2- 5) (PAR3-alpha). | |||||
|
CNGA1_HUMAN
|
||||||
| NC score | 0.008726 (rank : 147) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P29973, Q16279, Q16485 | Gene names | CNGA1, CNCG, CNCG1 | |||
|
Domain Architecture |
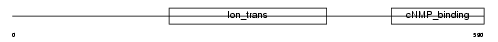
|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
PRLPN_MOUSE
|
||||||
| NC score | 0.008392 (rank : 148) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CGZ9, Q9DAX1 | Gene names | Prlpn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Placental prolactin-like protein N precursor (PRL-like protein N) (PLP-N). | |||||
|
SCN2A_HUMAN
|
||||||
| NC score | 0.007851 (rank : 149) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99250, Q14472, Q9BZC9, Q9BZD0 | Gene names | SCN2A, NAC2, SCN2A2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 2 subunit alpha (Sodium channel protein type II subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.2) (Sodium channel protein, brain II subunit alpha) (HBSC II). | |||||
|
IFNW1_HUMAN
|
||||||
| NC score | 0.007499 (rank : 150) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P05000 | Gene names | IFNW1 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon omega-1 precursor (Interferon alpha-II-1). | |||||
|
NEK1_MOUSE
|
||||||
| NC score | 0.006993 (rank : 151) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P51954 | Gene names | Nek1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek1 (EC 2.7.11.1) (NimA-related protein kinase 1). | |||||
|
LASP1_HUMAN
|
||||||
| NC score | 0.006275 (rank : 152) | θ value | 8.99809 (rank : 127) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14847, Q96ED2, Q96IG0 | Gene names | LASP1, MLN50 | |||
|
Domain Architecture |

|
|||||
| Description | LIM and SH3 domain protein 1 (LASP-1) (MLN 50). | |||||
|
CASR_HUMAN
|
||||||
| NC score | 0.006066 (rank : 153) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P41180, Q13912, Q16108, Q16109, Q16110, Q16379, Q4PJ19 | Gene names | CASR, GPRC2A, PCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Extracellular calcium-sensing receptor precursor (CaSR) (Parathyroid Cell calcium-sensing receptor). | |||||
|
ZFP37_MOUSE
|
||||||
| NC score | 0.005943 (rank : 154) | θ value | 0.47712 (rank : 32) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P17141, Q62514 | Gene names | Zfp37, Zfp-37 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 37 (Zfp-37) (Male germ cell-specific zinc finger protein). | |||||
|
MARK2_HUMAN
|
||||||
| NC score | 0.005856 (rank : 155) | θ value | 0.813845 (rank : 35) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||
|
MAK_HUMAN
|
||||||
| NC score | 0.004342 (rank : 156) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 857 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20794, Q9NUH7 | Gene names | MAK | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MAK (EC 2.7.11.22) (Male germ cell- associated kinase). | |||||
|
EVI1_MOUSE
|
||||||
| NC score | 0.003118 (rank : 157) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 826 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P14404 | Gene names | Evi1, Evi-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ecotropic virus integration 1 site protein. | |||||
|
ZN467_MOUSE
|
||||||
| NC score | 0.003014 (rank : 158) | θ value | 0.813845 (rank : 37) | |||
| Query Neighborhood Hits | 141 | Target Neighborhood Hits | 794 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8JZL0, Q9JJ98 | Gene names | Znf467, Ezi, Zfp467 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 467 (Endothelial cell-derived zinc finger protein) (EZI). | |||||