Please be patient as the page loads
|
CDYL_MOUSE
|
||||||
| SwissProt Accessions | Q9WTK2 | Gene names | Cdyl | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain Y-like protein (CDY-like). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CDY1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.990155 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9Y6F8, O14600 | Gene names | CDY1 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific chromodomain protein Y 1. | |||||
|
CDY2_HUMAN
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.990115 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9Y6F7 | Gene names | CDY2A, CDY2 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific chromodomain protein Y 2. | |||||
|
CDYL1_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.992025 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9Y232 | Gene names | CDYL, CDYL1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain Y-like protein (CDY-like). | |||||
|
CDYL_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9WTK2 | Gene names | Cdyl | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain Y-like protein (CDY-like). | |||||
|
CDYL2_HUMAN
|
||||||
| θ value | 1.99421e-132 (rank : 5) | NC score | 0.979803 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8N8U2 | Gene names | CDYL2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain Y-like protein 2 (CDY-like 2). | |||||
|
PECI_HUMAN
|
||||||
| θ value | 1.97686e-47 (rank : 6) | NC score | 0.793250 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75521, Q5JYK5, Q8N0X0, Q9BUE9, Q9H0T9, Q9NQH1, Q9NYH7, Q9UN55 | Gene names | PECI, DRS1, HCA88 | |||
|
Domain Architecture |
|
|||||
| Description | Peroxisomal 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase) (DBI-related protein 1) (DRS-1) (Hepatocellular carcinoma- associated antigen 88) (NY-REN-1 antigen). | |||||
|
PECI_MOUSE
|
||||||
| θ value | 2.58187e-47 (rank : 7) | NC score | 0.806262 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9WUR2 | Gene names | Peci | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
ECH1_MOUSE
|
||||||
| θ value | 2.5182e-18 (rank : 8) | NC score | 0.690726 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35459, Q5M8P6 | Gene names | Ech1 | |||
|
Domain Architecture |
|
|||||
| Description | Delta3,5-delta2,4-dienoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.-). | |||||
|
ECH1_HUMAN
|
||||||
| θ value | 4.29542e-18 (rank : 9) | NC score | 0.685488 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13011, Q8WVX0, Q96EZ9 | Gene names | ECH1 | |||
|
Domain Architecture |
|
|||||
| Description | Delta3,5-delta2,4-dienoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.-). | |||||
|
ECHM_MOUSE
|
||||||
| θ value | 7.32683e-18 (rank : 10) | NC score | 0.675856 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BH95, Q3TJK2, Q6GQS2, Q6PEN1, Q99LX7 | Gene names | Echs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enoyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.17) (Short chain enoyl-CoA hydratase) (SCEH) (Enoyl-CoA hydratase 1). | |||||
|
ECHM_HUMAN
|
||||||
| θ value | 8.10077e-17 (rank : 11) | NC score | 0.671605 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P30084, O00739, Q5VWY1, Q96H54 | Gene names | ECHS1 | |||
|
Domain Architecture |
|
|||||
| Description | Enoyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.17) (Short chain enoyl-CoA hydratase) (SCEH) (Enoyl-CoA hydratase 1). | |||||
|
AUMH_HUMAN
|
||||||
| θ value | 3.52202e-12 (rank : 12) | NC score | 0.586173 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13825, Q8WUE4 | Gene names | AUH | |||
|
Domain Architecture |

|
|||||
| Description | Methylglutaconyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.18) (AU-specific RNA-binding enoyl-CoA hydratase) (AU-binding protein/enoyl-CoA hydratase). | |||||
|
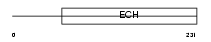
AUHM_MOUSE
|
||||||
| θ value | 4.59992e-12 (rank : 13) | NC score | 0.591455 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9JLZ3, Q80YD7, Q8QZS0, Q9CY78, Q9D155 | Gene names | Auh | |||
|
Domain Architecture |
|
|||||
| Description | Methylglutaconyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.18) (AU-specific RNA-binding enoyl-CoA hydratase) (AU-binding enoyl-CoA hydratase) (muAUH). | |||||
|
ECHA_HUMAN
|
||||||
| θ value | 2.52405e-10 (rank : 14) | NC score | 0.477684 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P40939, Q16679, Q96GT7 | Gene names | HADHA, HADH | |||
|
Domain Architecture |

|
|||||
| Description | Trifunctional enzyme subunit alpha, mitochondrial precursor (TP-alpha) (78 kDa gastrin-binding protein) [Includes: Long-chain enoyl-CoA hydratase (EC 4.2.1.17); Long chain 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.211)]. | |||||
|
MPP8_HUMAN
|
||||||
| θ value | 7.34386e-10 (rank : 15) | NC score | 0.215944 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
ECHP_MOUSE
|
||||||
| θ value | 9.59137e-10 (rank : 16) | NC score | 0.447059 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9DBM2 | Gene names | Ehhadh | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal bifunctional enzyme (PBE) (PBFE) [Includes: Enoyl-CoA hydratase (EC 4.2.1.17); 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8); 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.35)]. | |||||
|
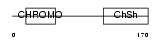
CBX5_HUMAN
|
||||||
| θ value | 1.25267e-09 (rank : 17) | NC score | 0.479876 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P45973 | Gene names | CBX5, HP1A | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha) (Antigen p25). | |||||
|
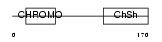
CBX5_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 18) | NC score | 0.481991 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q61686, Q9CS35, Q9CXD1 | Gene names | Cbx5, Hp1a | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha). | |||||
|
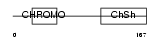
CBX1_HUMAN
|
||||||
| θ value | 8.97725e-08 (rank : 19) | NC score | 0.479153 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P83916, P23197 | Gene names | CBX1, CBX | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25) (HP1Hsbeta) (p25beta). | |||||
|
CBX1_MOUSE
|
||||||
| θ value | 8.97725e-08 (rank : 20) | NC score | 0.479153 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P83917, P23197 | Gene names | Cbx1, Cbx | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25). | |||||
|
CBX3_MOUSE
|
||||||
| θ value | 8.97725e-08 (rank : 21) | NC score | 0.489026 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P23198, Q921C4 | Gene names | Cbx3 | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (M32). | |||||
|
ECHP_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 22) | NC score | 0.401605 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q08426 | Gene names | EHHADH, ECHD | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal bifunctional enzyme (PBE) (PBFE) [Includes: Enoyl-CoA hydratase (EC 4.2.1.17); 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8); 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.35)]. | |||||
|
CBX4_MOUSE
|
||||||
| θ value | 5.81887e-07 (rank : 23) | NC score | 0.394049 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O55187 | Gene names | Cbx4, Pc2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 4 (Polycomb 2 homolog) (Pc2) (mPc2). | |||||
|
CBX3_HUMAN
|
||||||
| θ value | 7.59969e-07 (rank : 24) | NC score | 0.477001 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13185, Q96CD7, Q99409, Q9BVS3, Q9P0Z6 | Gene names | CBX3 | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (HECH). | |||||
|
CBX4_HUMAN
|
||||||
| θ value | 7.59969e-07 (rank : 25) | NC score | 0.395113 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O00257, Q96C04 | Gene names | CBX4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 4 (Polycomb 2 homolog) (Pc2) (hPc2). | |||||
|
CBX6_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 26) | NC score | 0.391521 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O95503, Q96EM5 | Gene names | CBX6 | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 6. | |||||
|
CBX6_MOUSE
|
||||||
| θ value | 6.43352e-06 (rank : 27) | NC score | 0.396176 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9DBY5, Q8BZH5 | Gene names | Cbx6 | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 6. | |||||
|
CBX7_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 28) | NC score | 0.439240 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O95931, Q86T17 | Gene names | CBX7 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 7. | |||||
|
CBX7_MOUSE
|
||||||
| θ value | 6.43352e-06 (rank : 29) | NC score | 0.457308 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8VDS3 | Gene names | Cbx7, D15Ertd417e | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 7. | |||||
|
CBX8_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 30) | NC score | 0.398789 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QXV1 | Gene names | Cbx8, Pc3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 8 (Polycomb 3 homolog) (Pc3) (mPc3). | |||||
|
CBX8_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 31) | NC score | 0.396292 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9HC52, Q96H39, Q9NR07 | Gene names | CBX8, PC3, RC1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 8 (Polycomb 3 homolog) (Pc3) (hPc3) (Rectachrome 1). | |||||
|
CBX2_MOUSE
|
||||||
| θ value | 5.44631e-05 (rank : 32) | NC score | 0.403814 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P30658, O35731, Q8CIA0 | Gene names | Cbx2, M33 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 2 (Modifier 3 protein) (M33). | |||||
|
CBX2_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 33) | NC score | 0.372706 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14781, Q9BTB1 | Gene names | CBX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromobox protein homolog 2. | |||||
|
SUV91_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 34) | NC score | 0.217622 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O54864, Q9JLC7, Q9JLP8 | Gene names | Suv39h1, Suv39h | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1) (Position-effect variegation 3-9 homolog). | |||||
|
SUV91_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 35) | NC score | 0.207405 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O43463, Q53G60, Q6FHK6 | Gene names | SUV39H1, SUV39H | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1). | |||||
|
CHD1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 36) | NC score | 0.094633 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
SUV92_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 37) | NC score | 0.203797 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9EQQ0, Q9CUK3, Q9JLP7 | Gene names | Suv39h2 | |||
|
Domain Architecture |
|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
SUV92_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 38) | NC score | 0.201739 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9H5I1 | Gene names | SUV39H2 | |||
|
Domain Architecture |
|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
CHD1_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 39) | NC score | 0.096094 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CHD2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 40) | NC score | 0.091910 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O14647 | Gene names | CHD2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 2 (EC 3.6.1.-) (ATP- dependent helicase CHD2) (CHD-2). | |||||
|
CHD3_HUMAN
|
||||||
| θ value | 0.365318 (rank : 41) | NC score | 0.064857 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
C163B_HUMAN
|
||||||
| θ value | 0.47712 (rank : 42) | NC score | 0.014126 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NR16, Q6UWC2 | Gene names | CD163L1, CD163B, M160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor cysteine-rich type 1 protein M160 precursor (CD163 antigen-like 1) (CD163b antigen). | |||||
|
MS3L1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 43) | NC score | 0.107156 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WVG9 | Gene names | Msl3l1, Msl31 | |||
|
Domain Architecture |
|
|||||
| Description | Male-specific lethal 3-like 1 (MSL3-like 1) (Male-specific lethal-3 homolog 1). | |||||
|
DDX11_MOUSE
|
||||||
| θ value | 0.62314 (rank : 44) | NC score | 0.021077 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6AXC6 | Gene names | Ddx11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX11 (EC 3.6.1.-) (DEAD/H box protein 11). | |||||
|
SETX_HUMAN
|
||||||
| θ value | 0.813845 (rank : 45) | NC score | 0.023272 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
NFM_HUMAN
|
||||||
| θ value | 1.06291 (rank : 46) | NC score | 0.003369 (rank : 77) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
CC019_MOUSE
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.023539 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3U155, Q80XC8, Q8CA20 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C3orf19 homolog. | |||||
|
CCNF_MOUSE
|
||||||
| θ value | 1.81305 (rank : 48) | NC score | 0.015816 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P51944, Q60797, Q60799 | Gene names | Ccnf | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-F. | |||||
|
CO4A2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.003773 (rank : 76) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P08122, Q61375 | Gene names | Col4a2 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
NCOR1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.017452 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
PLST_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.014458 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P13797, Q86YI6 | Gene names | PLS3 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-3 (T-plastin). | |||||
|
PLST_MOUSE
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.014570 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99K51 | Gene names | Pls3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plastin-3 (T-plastin). | |||||
|
ZN433_HUMAN
|
||||||
| θ value | 2.36792 (rank : 53) | NC score | -0.002268 (rank : 78) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 782 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N7K0 | Gene names | ZNF433 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 433. | |||||
|
FA13C_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.013054 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NE31, Q99787 | Gene names | FAM13C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
PCCA_MOUSE
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.010456 (rank : 70) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91ZA3, Q80VU5, Q922N3 | Gene names | Pcca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Propionyl-CoA carboxylase alpha chain, mitochondrial precursor (EC 6.4.1.3) (PCCase subunit alpha) (Propanoyl-CoA:carbon dioxide ligase subunit alpha). | |||||
|
SG16_MOUSE
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.028139 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P02815, Q62263 | Gene names | Spt1, Spt-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 16.5 kDa submandibular gland glycoprotein precursor (Salivary protein 1). | |||||
|
ZDHC8_MOUSE
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.007423 (rank : 72) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5Y5T5, Q7TNF7 | Gene names | Zdhhc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable palmitoyltransferase ZDHHC8 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 8) (DHHC-8). | |||||
|
BCAS1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.020762 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80YN3, Q9CVA1 | Gene names | Bcas1, Nabc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 homolog (Novel amplified in breast cancer 1 homolog). | |||||
|
MS3L1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.058699 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N5Y2, Q9UG70, Q9Y5Z8 | Gene names | MSL3L1 | |||
|
Domain Architecture |

|
|||||
| Description | Male-specific lethal 3-like 1 (MSL3-like 1) (Male-specific lethal-3 homolog 1). | |||||
|
LAP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.004238 (rank : 74) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96RT1, Q86W38, Q9NR18, Q9NW48, Q9ULJ5 | Gene names | ERBB2IP, ERBIN, KIAA1225, LAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
LAP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.004014 (rank : 75) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TH2, Q8BQ14, Q8CE41, Q8K171, Q99JU3, Q9JI47 | Gene names | Erbb2ip, Erbin, Kiaa1225, Lap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
SGOL1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.017432 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5FBB7, Q588H5, Q5FBB4, Q5FBB5, Q5FBB6, Q5FBB8, Q8N579, Q8WVL0, Q9BVA8, Q9H275 | Gene names | SGOL1, SGO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 1 (hSgo1) (Serologically defined breast cancer antigen NY-BR-85). | |||||
|
CHD4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.049480 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
CHD4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.049124 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
CHD5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.047856 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
LMD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.017644 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P29536 | Gene names | LMOD1 | |||
|
Domain Architecture |

|
|||||
| Description | Leiomodin-1 (Leiomodin, muscle form) (64 kDa autoantigen D1) (64 kDa autoantigen 1D) (64 kDa autoantigen 1D3) (Thyroid-associated ophthalmopathy autoantigen) (Smooth muscle leiomodin) (SM-Lmod). | |||||
|
OXR1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.015491 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
COFA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.005030 (rank : 73) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35206, Q9EQD9 | Gene names | Col15a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV)]. | |||||
|
LYRIC_MOUSE
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.009054 (rank : 71) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80WJ7, Q3U9F8, Q3UAQ8, Q8BN67, Q8CBT9, Q8CDL0, Q8CGI7, Q9D052 | Gene names | Mtdh, Lyric | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/LYRIC) (Metastasis adhesion protein) (Metadherin). | |||||
|
ACBD4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.105186 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NC06, Q8IUT1, Q9H8Q4 | Gene names | ACBD4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acyl-CoA-binding domain-containing protein 4. | |||||
|
ACBD7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.110315 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N6N7 | Gene names | ACBD7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acyl-CoA-binding domain-containing protein 7. | |||||
|
ACBP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.115569 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P07108, P08869, Q53SQ7, Q6IB48 | Gene names | DBI | |||
|
Domain Architecture |
|
|||||
| Description | Acyl-CoA-binding protein (ACBP) (Diazepam-binding inhibitor) (DBI) (Endozepine) (EP). | |||||
|
ACBP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.117284 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P31786 | Gene names | Dbi | |||
|
Domain Architecture |
|
|||||
| Description | Acyl-CoA-binding protein (ACBP) (Diazepam-binding inhibitor) (DBI) (Endozepine) (EP). | |||||
|
D3D2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.202488 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P42126, Q13290, Q7Z2L6, Q9BUB8, Q9BW05 | Gene names | DCI | |||
|
Domain Architecture |
|
|||||
| Description | 3,2-trans-enoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
D3D2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.183534 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P42125 | Gene names | Dci | |||
|
Domain Architecture |
|
|||||
| Description | 3,2-trans-enoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
DBIL5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.107985 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O09035 | Gene names | Dbil5 | |||
|
Domain Architecture |
|
|||||
| Description | Diazepam-binding inhibitor-like 5 (Endozepine-like peptide) (ELP). | |||||
|
HCDH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.078970 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q16836, O00324, O00397, O00753 | Gene names | HADHSC, HAD, SCHAD | |||
|
Domain Architecture |
|
|||||
| Description | Short chain 3-hydroxyacyl-CoA dehydrogenase, mitochondrial precursor (EC 1.1.1.35) (HCDH) (Medium and short chain L-3-hydroxyacyl-coenzyme A dehydrogenase). | |||||
|
HCDH_MOUSE
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.077699 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61425, Q3TF75, Q3THK8, Q3UFI0, Q8K149, Q925U9 | Gene names | Hadhsc, Hadh, Mschad, Schad | |||
|
Domain Architecture |
|
|||||
| Description | Short chain 3-hydroxyacyl-CoA dehydrogenase, mitochondrial precursor (EC 1.1.1.35) (HCDH) (Medium and short chain L-3-hydroxyacyl-coenzyme A dehydrogenase). | |||||
|
CDYL_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9WTK2 | Gene names | Cdyl | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain Y-like protein (CDY-like). | |||||
|
CDYL1_HUMAN
|
||||||
| NC score | 0.992025 (rank : 2) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9Y232 | Gene names | CDYL, CDYL1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain Y-like protein (CDY-like). | |||||
|
CDY1_HUMAN
|
||||||
| NC score | 0.990155 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9Y6F8, O14600 | Gene names | CDY1 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific chromodomain protein Y 1. | |||||
|
CDY2_HUMAN
|
||||||
| NC score | 0.990115 (rank : 4) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9Y6F7 | Gene names | CDY2A, CDY2 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific chromodomain protein Y 2. | |||||
|
CDYL2_HUMAN
|
||||||
| NC score | 0.979803 (rank : 5) | θ value | 1.99421e-132 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8N8U2 | Gene names | CDYL2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain Y-like protein 2 (CDY-like 2). | |||||
|
PECI_MOUSE
|
||||||
| NC score | 0.806262 (rank : 6) | θ value | 2.58187e-47 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9WUR2 | Gene names | Peci | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
PECI_HUMAN
|
||||||
| NC score | 0.793250 (rank : 7) | θ value | 1.97686e-47 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75521, Q5JYK5, Q8N0X0, Q9BUE9, Q9H0T9, Q9NQH1, Q9NYH7, Q9UN55 | Gene names | PECI, DRS1, HCA88 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase) (DBI-related protein 1) (DRS-1) (Hepatocellular carcinoma- associated antigen 88) (NY-REN-1 antigen). | |||||
|
ECH1_MOUSE
|
||||||
| NC score | 0.690726 (rank : 8) | θ value | 2.5182e-18 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35459, Q5M8P6 | Gene names | Ech1 | |||
|
Domain Architecture |

|
|||||
| Description | Delta3,5-delta2,4-dienoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.-). | |||||
|
ECH1_HUMAN
|
||||||
| NC score | 0.685488 (rank : 9) | θ value | 4.29542e-18 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13011, Q8WVX0, Q96EZ9 | Gene names | ECH1 | |||
|
Domain Architecture |

|
|||||
| Description | Delta3,5-delta2,4-dienoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.-). | |||||
|
ECHM_MOUSE
|
||||||
| NC score | 0.675856 (rank : 10) | θ value | 7.32683e-18 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BH95, Q3TJK2, Q6GQS2, Q6PEN1, Q99LX7 | Gene names | Echs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enoyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.17) (Short chain enoyl-CoA hydratase) (SCEH) (Enoyl-CoA hydratase 1). | |||||
|
ECHM_HUMAN
|
||||||
| NC score | 0.671605 (rank : 11) | θ value | 8.10077e-17 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P30084, O00739, Q5VWY1, Q96H54 | Gene names | ECHS1 | |||
|
Domain Architecture |

|
|||||
| Description | Enoyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.17) (Short chain enoyl-CoA hydratase) (SCEH) (Enoyl-CoA hydratase 1). | |||||
|
AUHM_MOUSE
|
||||||
| NC score | 0.591455 (rank : 12) | θ value | 4.59992e-12 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9JLZ3, Q80YD7, Q8QZS0, Q9CY78, Q9D155 | Gene names | Auh | |||
|
Domain Architecture |

|
|||||
| Description | Methylglutaconyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.18) (AU-specific RNA-binding enoyl-CoA hydratase) (AU-binding enoyl-CoA hydratase) (muAUH). | |||||
|
AUMH_HUMAN
|
||||||
| NC score | 0.586173 (rank : 13) | θ value | 3.52202e-12 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13825, Q8WUE4 | Gene names | AUH | |||
|
Domain Architecture |

|
|||||
| Description | Methylglutaconyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.18) (AU-specific RNA-binding enoyl-CoA hydratase) (AU-binding protein/enoyl-CoA hydratase). | |||||
|
CBX3_MOUSE
|
||||||
| NC score | 0.489026 (rank : 14) | θ value | 8.97725e-08 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P23198, Q921C4 | Gene names | Cbx3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (M32). | |||||
|
CBX5_MOUSE
|
||||||
| NC score | 0.481991 (rank : 15) | θ value | 2.79066e-09 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q61686, Q9CS35, Q9CXD1 | Gene names | Cbx5, Hp1a | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha). | |||||
|
CBX5_HUMAN
|
||||||
| NC score | 0.479876 (rank : 16) | θ value | 1.25267e-09 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P45973 | Gene names | CBX5, HP1A | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha) (Antigen p25). | |||||
|
CBX1_HUMAN
|
||||||
| NC score | 0.479153 (rank : 17) | θ value | 8.97725e-08 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P83916, P23197 | Gene names | CBX1, CBX | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25) (HP1Hsbeta) (p25beta). | |||||
|
CBX1_MOUSE
|
||||||
| NC score | 0.479153 (rank : 18) | θ value | 8.97725e-08 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P83917, P23197 | Gene names | Cbx1, Cbx | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25). | |||||
|
ECHA_HUMAN
|
||||||
| NC score | 0.477684 (rank : 19) | θ value | 2.52405e-10 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P40939, Q16679, Q96GT7 | Gene names | HADHA, HADH | |||
|
Domain Architecture |

|
|||||
| Description | Trifunctional enzyme subunit alpha, mitochondrial precursor (TP-alpha) (78 kDa gastrin-binding protein) [Includes: Long-chain enoyl-CoA hydratase (EC 4.2.1.17); Long chain 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.211)]. | |||||
|
CBX3_HUMAN
|
||||||
| NC score | 0.477001 (rank : 20) | θ value | 7.59969e-07 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13185, Q96CD7, Q99409, Q9BVS3, Q9P0Z6 | Gene names | CBX3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (HECH). | |||||
|
CBX7_MOUSE
|
||||||
| NC score | 0.457308 (rank : 21) | θ value | 6.43352e-06 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8VDS3 | Gene names | Cbx7, D15Ertd417e | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 7. | |||||
|
ECHP_MOUSE
|
||||||
| NC score | 0.447059 (rank : 22) | θ value | 9.59137e-10 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9DBM2 | Gene names | Ehhadh | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal bifunctional enzyme (PBE) (PBFE) [Includes: Enoyl-CoA hydratase (EC 4.2.1.17); 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8); 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.35)]. | |||||
|
CBX7_HUMAN
|
||||||
| NC score | 0.439240 (rank : 23) | θ value | 6.43352e-06 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O95931, Q86T17 | Gene names | CBX7 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 7. | |||||
|
CBX2_MOUSE
|
||||||
| NC score | 0.403814 (rank : 24) | θ value | 5.44631e-05 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P30658, O35731, Q8CIA0 | Gene names | Cbx2, M33 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 2 (Modifier 3 protein) (M33). | |||||
|
ECHP_HUMAN
|
||||||
| NC score | 0.401605 (rank : 25) | θ value | 1.99992e-07 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q08426 | Gene names | EHHADH, ECHD | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal bifunctional enzyme (PBE) (PBFE) [Includes: Enoyl-CoA hydratase (EC 4.2.1.17); 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8); 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.35)]. | |||||
|
CBX8_MOUSE
|
||||||
| NC score | 0.398789 (rank : 26) | θ value | 1.09739e-05 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QXV1 | Gene names | Cbx8, Pc3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 8 (Polycomb 3 homolog) (Pc3) (mPc3). | |||||
|
CBX8_HUMAN
|
||||||
| NC score | 0.396292 (rank : 27) | θ value | 4.1701e-05 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9HC52, Q96H39, Q9NR07 | Gene names | CBX8, PC3, RC1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 8 (Polycomb 3 homolog) (Pc3) (hPc3) (Rectachrome 1). | |||||
|
CBX6_MOUSE
|
||||||
| NC score | 0.396176 (rank : 28) | θ value | 6.43352e-06 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9DBY5, Q8BZH5 | Gene names | Cbx6 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 6. | |||||
|
CBX4_HUMAN
|
||||||
| NC score | 0.395113 (rank : 29) | θ value | 7.59969e-07 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O00257, Q96C04 | Gene names | CBX4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 4 (Polycomb 2 homolog) (Pc2) (hPc2). | |||||
|
CBX4_MOUSE
|
||||||
| NC score | 0.394049 (rank : 30) | θ value | 5.81887e-07 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O55187 | Gene names | Cbx4, Pc2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 4 (Polycomb 2 homolog) (Pc2) (mPc2). | |||||
|
CBX6_HUMAN
|
||||||
| NC score | 0.391521 (rank : 31) | θ value | 6.43352e-06 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O95503, Q96EM5 | Gene names | CBX6 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 6. | |||||
|
CBX2_HUMAN
|
||||||
| NC score | 0.372706 (rank : 32) | θ value | 7.1131e-05 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14781, Q9BTB1 | Gene names | CBX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromobox protein homolog 2. | |||||
|
SUV91_MOUSE
|
||||||
| NC score | 0.217622 (rank : 33) | θ value | 0.00298849 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O54864, Q9JLC7, Q9JLP8 | Gene names | Suv39h1, Suv39h | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1) (Position-effect variegation 3-9 homolog). | |||||
|
MPP8_HUMAN
|
||||||
| NC score | 0.215944 (rank : 34) | θ value | 7.34386e-10 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
SUV91_HUMAN
|
||||||
| NC score | 0.207405 (rank : 35) | θ value | 0.00509761 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O43463, Q53G60, Q6FHK6 | Gene names | SUV39H1, SUV39H | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1). | |||||
|
SUV92_MOUSE
|
||||||
| NC score | 0.203797 (rank : 36) | θ value | 0.0330416 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9EQQ0, Q9CUK3, Q9JLP7 | Gene names | Suv39h2 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
D3D2_HUMAN
|
||||||
| NC score | 0.202488 (rank : 37) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P42126, Q13290, Q7Z2L6, Q9BUB8, Q9BW05 | Gene names | DCI | |||
|
Domain Architecture |

|
|||||
| Description | 3,2-trans-enoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
SUV92_HUMAN
|
||||||
| NC score | 0.201739 (rank : 38) | θ value | 0.0431538 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9H5I1 | Gene names | SUV39H2 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
D3D2_MOUSE
|
||||||
| NC score | 0.183534 (rank : 39) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P42125 | Gene names | Dci | |||
|
Domain Architecture |

|
|||||
| Description | 3,2-trans-enoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
ACBP_MOUSE
|
||||||
| NC score | 0.117284 (rank : 40) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P31786 | Gene names | Dbi | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-CoA-binding protein (ACBP) (Diazepam-binding inhibitor) (DBI) (Endozepine) (EP). | |||||
|
ACBP_HUMAN
|
||||||
| NC score | 0.115569 (rank : 41) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P07108, P08869, Q53SQ7, Q6IB48 | Gene names | DBI | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-CoA-binding protein (ACBP) (Diazepam-binding inhibitor) (DBI) (Endozepine) (EP). | |||||
|
ACBD7_HUMAN
|
||||||
| NC score | 0.110315 (rank : 42) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N6N7 | Gene names | ACBD7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acyl-CoA-binding domain-containing protein 7. | |||||
|
DBIL5_MOUSE
|
||||||
| NC score | 0.107985 (rank : 43) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O09035 | Gene names | Dbil5 | |||
|
Domain Architecture |

|
|||||
| Description | Diazepam-binding inhibitor-like 5 (Endozepine-like peptide) (ELP). | |||||
|
MS3L1_MOUSE
|
||||||
| NC score | 0.107156 (rank : 44) | θ value | 0.47712 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WVG9 | Gene names | Msl3l1, Msl31 | |||
|
Domain Architecture |

|
|||||
| Description | Male-specific lethal 3-like 1 (MSL3-like 1) (Male-specific lethal-3 homolog 1). | |||||
|
ACBD4_HUMAN
|
||||||
| NC score | 0.105186 (rank : 45) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NC06, Q8IUT1, Q9H8Q4 | Gene names | ACBD4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acyl-CoA-binding domain-containing protein 4. | |||||
|
CHD1_MOUSE
|
||||||
| NC score | 0.096094 (rank : 46) | θ value | 0.0563607 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CHD1_HUMAN
|
||||||
| NC score | 0.094633 (rank : 47) | θ value | 0.0330416 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CHD2_HUMAN
|
||||||
| NC score | 0.091910 (rank : 48) | θ value | 0.163984 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O14647 | Gene names | CHD2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 2 (EC 3.6.1.-) (ATP- dependent helicase CHD2) (CHD-2). | |||||
|
HCDH_HUMAN
|
||||||
| NC score | 0.078970 (rank : 49) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q16836, O00324, O00397, O00753 | Gene names | HADHSC, HAD, SCHAD | |||
|
Domain Architecture |

|
|||||
| Description | Short chain 3-hydroxyacyl-CoA dehydrogenase, mitochondrial precursor (EC 1.1.1.35) (HCDH) (Medium and short chain L-3-hydroxyacyl-coenzyme A dehydrogenase). | |||||
|
HCDH_MOUSE
|
||||||
| NC score | 0.077699 (rank : 50) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61425, Q3TF75, Q3THK8, Q3UFI0, Q8K149, Q925U9 | Gene names | Hadhsc, Hadh, Mschad, Schad | |||
|
Domain Architecture |

|
|||||
| Description | Short chain 3-hydroxyacyl-CoA dehydrogenase, mitochondrial precursor (EC 1.1.1.35) (HCDH) (Medium and short chain L-3-hydroxyacyl-coenzyme A dehydrogenase). | |||||
|
CHD3_HUMAN
|
||||||
| NC score | 0.064857 (rank : 51) | θ value | 0.365318 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
MS3L1_HUMAN
|
||||||
| NC score | 0.058699 (rank : 52) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N5Y2, Q9UG70, Q9Y5Z8 | Gene names | MSL3L1 | |||
|
Domain Architecture |

|
|||||
| Description | Male-specific lethal 3-like 1 (MSL3-like 1) (Male-specific lethal-3 homolog 1). | |||||
|
CHD4_HUMAN
|
||||||
| NC score | 0.049480 (rank : 53) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
CHD4_MOUSE
|
||||||
| NC score | 0.049124 (rank : 54) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
CHD5_HUMAN
|
||||||
| NC score | 0.047856 (rank : 55) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
SG16_MOUSE
|
||||||
| NC score | 0.028139 (rank : 56) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P02815, Q62263 | Gene names | Spt1, Spt-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 16.5 kDa submandibular gland glycoprotein precursor (Salivary protein 1). | |||||
|
CC019_MOUSE
|
||||||
| NC score | 0.023539 (rank : 57) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3U155, Q80XC8, Q8CA20 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C3orf19 homolog. | |||||
|
SETX_HUMAN
|
||||||
| NC score | 0.023272 (rank : 58) | θ value | 0.813845 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
DDX11_MOUSE
|
||||||
| NC score | 0.021077 (rank : 59) | θ value | 0.62314 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6AXC6 | Gene names | Ddx11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX11 (EC 3.6.1.-) (DEAD/H box protein 11). | |||||
|
BCAS1_MOUSE
|
||||||
| NC score | 0.020762 (rank : 60) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80YN3, Q9CVA1 | Gene names | Bcas1, Nabc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 homolog (Novel amplified in breast cancer 1 homolog). | |||||
|
LMD1_HUMAN
|
||||||
| NC score | 0.017644 (rank : 61) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P29536 | Gene names | LMOD1 | |||
|
Domain Architecture |

|
|||||
| Description | Leiomodin-1 (Leiomodin, muscle form) (64 kDa autoantigen D1) (64 kDa autoantigen 1D) (64 kDa autoantigen 1D3) (Thyroid-associated ophthalmopathy autoantigen) (Smooth muscle leiomodin) (SM-Lmod). | |||||
|
NCOR1_MOUSE
|
||||||
| NC score | 0.017452 (rank : 62) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
SGOL1_HUMAN
|
||||||
| NC score | 0.017432 (rank : 63) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5FBB7, Q588H5, Q5FBB4, Q5FBB5, Q5FBB6, Q5FBB8, Q8N579, Q8WVL0, Q9BVA8, Q9H275 | Gene names | SGOL1, SGO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 1 (hSgo1) (Serologically defined breast cancer antigen NY-BR-85). | |||||
|
CCNF_MOUSE
|
||||||
| NC score | 0.015816 (rank : 64) | θ value | 1.81305 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P51944, Q60797, Q60799 | Gene names | Ccnf | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-F. | |||||
|
OXR1_MOUSE
|
||||||
| NC score | 0.015491 (rank : 65) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
PLST_MOUSE
|
||||||
| NC score | 0.014570 (rank : 66) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99K51 | Gene names | Pls3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plastin-3 (T-plastin). | |||||
|
PLST_HUMAN
|
||||||
| NC score | 0.014458 (rank : 67) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P13797, Q86YI6 | Gene names | PLS3 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-3 (T-plastin). | |||||
|
C163B_HUMAN
|
||||||
| NC score | 0.014126 (rank : 68) | θ value | 0.47712 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NR16, Q6UWC2 | Gene names | CD163L1, CD163B, M160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor cysteine-rich type 1 protein M160 precursor (CD163 antigen-like 1) (CD163b antigen). | |||||
|
FA13C_HUMAN
|
||||||
| NC score | 0.013054 (rank : 69) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NE31, Q99787 | Gene names | FAM13C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
PCCA_MOUSE
|
||||||
| NC score | 0.010456 (rank : 70) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91ZA3, Q80VU5, Q922N3 | Gene names | Pcca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Propionyl-CoA carboxylase alpha chain, mitochondrial precursor (EC 6.4.1.3) (PCCase subunit alpha) (Propanoyl-CoA:carbon dioxide ligase subunit alpha). | |||||
|
LYRIC_MOUSE
|
||||||
| NC score | 0.009054 (rank : 71) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80WJ7, Q3U9F8, Q3UAQ8, Q8BN67, Q8CBT9, Q8CDL0, Q8CGI7, Q9D052 | Gene names | Mtdh, Lyric | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/LYRIC) (Metastasis adhesion protein) (Metadherin). | |||||
|
ZDHC8_MOUSE
|
||||||
| NC score | 0.007423 (rank : 72) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5Y5T5, Q7TNF7 | Gene names | Zdhhc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable palmitoyltransferase ZDHHC8 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 8) (DHHC-8). | |||||
|
COFA1_MOUSE
|
||||||
| NC score | 0.005030 (rank : 73) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35206, Q9EQD9 | Gene names | Col15a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV)]. | |||||
|
LAP2_HUMAN
|
||||||
| NC score | 0.004238 (rank : 74) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96RT1, Q86W38, Q9NR18, Q9NW48, Q9ULJ5 | Gene names | ERBB2IP, ERBIN, KIAA1225, LAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
LAP2_MOUSE
|
||||||
| NC score | 0.004014 (rank : 75) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TH2, Q8BQ14, Q8CE41, Q8K171, Q99JU3, Q9JI47 | Gene names | Erbb2ip, Erbin, Kiaa1225, Lap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
CO4A2_MOUSE
|
||||||
| NC score | 0.003773 (rank : 76) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P08122, Q61375 | Gene names | Col4a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
NFM_HUMAN
|
||||||
| NC score | 0.003369 (rank : 77) | θ value | 1.06291 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
ZN433_HUMAN
|
||||||
| NC score | -0.002268 (rank : 78) | θ value | 2.36792 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 782 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N7K0 | Gene names | ZNF433 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 433. | |||||