Please be patient as the page loads
|
SPF45_HUMAN
|
||||||
| SwissProt Accessions | Q96I25, Q96GY6 | Gene names | RBM17, SPF45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SPF45_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 94 | |
| SwissProt Accessions | Q96I25, Q96GY6 | Gene names | RBM17, SPF45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
SPF45_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.994041 (rank : 2) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q8JZX4, O75939 | Gene names | Rbm17, Spf45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
U2AF2_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 3) | NC score | 0.244929 (rank : 3) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P26368 | Gene names | U2AF2, U2AF65 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor U2AF 65 kDa subunit (U2 auxiliary factor 65 kDa subunit) (U2 snRNP auxiliary factor large subunit) (hU2AF(65)). | |||||
|
U2AF2_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 4) | NC score | 0.244049 (rank : 4) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P26369 | Gene names | U2af2, U2af65 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor U2AF 65 kDa subunit (U2 auxiliary factor 65 kDa subunit) (U2 snRNP auxiliary factor large subunit). | |||||
|
UHMK1_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 5) | NC score | 0.086742 (rank : 37) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TAS1, Q96C22 | Gene names | UHMK1, KIS, KIST | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Kist (EC 2.7.11.1) (Kinase interacting with stathmin) (U2AF homology motif kinase 1). | |||||
|
UHMK1_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 6) | NC score | 0.087215 (rank : 36) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 809 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97343, Q61775, Q9CYT1 | Gene names | Uhmk1, Kis, Kist | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Kist (EC 2.7.11.1) (Kinase interacting with stathmin) (U2AF homology motif kinase 1). | |||||
|
HTSF1_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 7) | NC score | 0.196335 (rank : 9) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
U2AF1_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 8) | NC score | 0.212708 (rank : 8) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q01081 | Gene names | U2AF1, U2AF35 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
U2AF1_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 9) | NC score | 0.213313 (rank : 7) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D883, Q564E4, Q99LX2, Q9CZ98 | Gene names | U2af1 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
K0082_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 10) | NC score | 0.228269 (rank : 5) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9DBC3, Q3U3G5, Q3U7Y9, Q6A0D5, Q8C7V0 | Gene names | Kiaa0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 11) | NC score | 0.101458 (rank : 31) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SON_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 12) | NC score | 0.146773 (rank : 25) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
HTSF1_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 13) | NC score | 0.188349 (rank : 10) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43719, Q59G06, Q99730 | Gene names | HTATSF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 (Tat-SF1). | |||||
|
K0082_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 14) | NC score | 0.228082 (rank : 6) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8N1G2, Q14670, Q96FJ9 | Gene names | KIAA0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
RBM5_HUMAN
|
||||||
| θ value | 0.163984 (rank : 15) | NC score | 0.143010 (rank : 27) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P52756, Q93021, Q9BU14, Q9UKY8, Q9UL24 | Gene names | RBM5 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15) (Protein G15) (NY-REN-9 antigen). | |||||
|
RBM5_MOUSE
|
||||||
| θ value | 0.163984 (rank : 16) | NC score | 0.142762 (rank : 29) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q91YE7, Q3UZ19, Q99KV9 | Gene names | Rbm5, Luca15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15). | |||||
|
PINX1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.175798 (rank : 11) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9CZX5, Q91WZ9, Q9D0C2 | Gene names | Pinx1, Lpts | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (LPTS1) (67-11-3 protein). | |||||
|
ITSN1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 18) | NC score | 0.021342 (rank : 91) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.066449 (rank : 43) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
RNPC2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.096232 (rank : 33) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q14498, Q14499 | Gene names | RNPC2, HCC1 | |||
|
Domain Architecture |
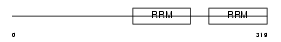
|
|||||
| Description | RNA-binding region-containing protein 2 (Hepatocellular carcinoma protein 1) (Splicing factor HCC1). | |||||
|
RNPC2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 21) | NC score | 0.096355 (rank : 32) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8VH51, Q99KV0 | Gene names | Rnpc2, Caper | |||
|
Domain Architecture |
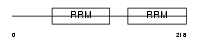
|
|||||
| Description | RNA-binding region-containing protein 2 (Coactivator of activating protein 1 and estrogen receptors) (Coactivator of AP-1 and ERs) (Transcription coactivator CAPER). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 22) | NC score | 0.049380 (rank : 73) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
VPK1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 23) | NC score | 0.172710 (rank : 13) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P63119 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
VPK9_HUMAN
|
||||||
| θ value | 0.365318 (rank : 24) | NC score | 0.173596 (rank : 12) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P63124 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q13.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K104 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK17_HUMAN
|
||||||
| θ value | 0.47712 (rank : 25) | NC score | 0.171198 (rank : 14) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63125 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_11q22.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
JHD1B_MOUSE
|
||||||
| θ value | 0.62314 (rank : 26) | NC score | 0.047697 (rank : 74) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
PABP5_HUMAN
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.074063 (rank : 38) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96DU9, Q6P529, Q9UFE5 | Gene names | PABPC5, PABP5 | |||
|
Domain Architecture |
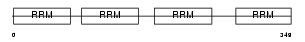
|
|||||
| Description | Polyadenylate-binding protein 5 (Poly(A)-binding protein 5) (PABP 5). | |||||
|
PRGC1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.068502 (rank : 40) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O70343 | Gene names | Ppargc1a, Pgc1, Pgc1a, Ppargc1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPAR gamma coactivator 1-alpha) (PPARGC-1-alpha) (PGC-1-alpha). | |||||
|
VPK11_HUMAN
|
||||||
| θ value | 0.62314 (rank : 29) | NC score | 0.167477 (rank : 16) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63129, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K101 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
VPK7_HUMAN
|
||||||
| θ value | 0.62314 (rank : 30) | NC score | 0.170452 (rank : 15) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63123 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q23.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K18 Pro protein) (HERV-K110 Pro protein) (HERV-K(C1a) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
PINX1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 31) | NC score | 0.154729 (rank : 24) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96BK5, Q7Z7J8, Q96QD7, Q9HBU7, Q9NWW2 | Gene names | PINX1, LPTS | |||
|
Domain Architecture |
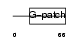
|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (67-11-3 protein). | |||||
|
TFP11_HUMAN
|
||||||
| θ value | 0.813845 (rank : 32) | NC score | 0.145504 (rank : 26) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UBB9, O95908, Q5H8V8, Q9UGV7, Q9Y2Q8 | Gene names | TFIP11 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11. | |||||
|
VPK3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 33) | NC score | 0.166610 (rank : 18) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63120 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19q12 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K(C19) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK4_HUMAN
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.161930 (rank : 23) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63127, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K109 Pro protein) (HERV-K(C6) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK6_HUMAN
|
||||||
| θ value | 0.813845 (rank : 35) | NC score | 0.162018 (rank : 22) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63122 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K115 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
JHD1B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 36) | NC score | 0.047195 (rank : 76) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
TFP11_MOUSE
|
||||||
| θ value | 1.06291 (rank : 37) | NC score | 0.142993 (rank : 28) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9ERA6, Q8VD06 | Gene names | Tfip11, Tip39 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11 (Tuftelin-interacting protein 39). | |||||
|
VPK10_HUMAN
|
||||||
| θ value | 1.06291 (rank : 38) | NC score | 0.166568 (rank : 19) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P10265, Q9UKH6 | Gene names | ||||
|
Domain Architecture |
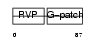
|
|||||
| Description | HERV-K_5q33.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K10 Pro protein) (HERV-K107 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK12_HUMAN
|
||||||
| θ value | 1.06291 (rank : 39) | NC score | 0.166528 (rank : 20) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P63131, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K102 Pro protein) (HERV-K(III) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK5_HUMAN
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.166686 (rank : 17) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P63121 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K113 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
WWC3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.051395 (rank : 71) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ULE0, Q659C1, Q9BTQ1 | Gene names | WWC3, KIAA1280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WWC family member 3. | |||||
|
DNJC7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 42) | NC score | 0.038155 (rank : 80) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99615 | Gene names | DNAJC7, TPR2, TTC2 | |||
|
Domain Architecture |
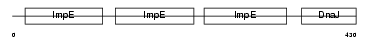
|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2). | |||||
|
DNJC7_MOUSE
|
||||||
| θ value | 1.38821 (rank : 43) | NC score | 0.038476 (rank : 79) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QYI3, Q3TKR1, Q8BPG3, Q8CIL2, Q9CT29, Q9D026 | Gene names | Dnajc7, Ttc2 | |||
|
Domain Architecture |
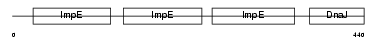
|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2) (MDj11). | |||||
|
MAMC1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 44) | NC score | 0.024257 (rank : 89) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P60755 | Gene names | Mamdc1, Mdga2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 1 precursor (MAM domain-containing glycosylphosphatidylinositol anchor protein 2). | |||||
|
MYH9_HUMAN
|
||||||
| θ value | 1.38821 (rank : 45) | NC score | 0.011871 (rank : 102) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
RB15B_HUMAN
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.065259 (rank : 44) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8NDT2, Q6QE19, Q9BV96 | Gene names | RBM15B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
RB15B_MOUSE
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.067696 (rank : 41) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6PHZ5, Q8C6G2 | Gene names | Rbm15b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
VPK2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 48) | NC score | 0.164098 (rank : 21) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y6I0, Q9UKH5 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K(HML-2.HOM) Pro protein) (HERV-K108 Pro protein) (HERV-K(C7) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
PABP4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.072111 (rank : 39) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13310, Q6P0N3 | Gene names | PABPC4, APP1, PABP4 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 4 (Poly(A)-binding protein 4) (PABP 4) (Inducible poly(A)-binding protein) (iPABP) (Activated-platelet protein 1) (APP-1). | |||||
|
CASP8_MOUSE
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.027723 (rank : 85) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O89110, O35669 | Gene names | Casp8 | |||
|
Domain Architecture |
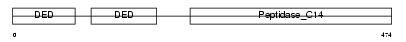
|
|||||
| Description | Caspase-8 precursor (EC 3.4.22.-) (CASP-8) [Contains: Caspase-8 subunit p18; Caspase-8 subunit p10]. | |||||
|
MYCB2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.027520 (rank : 86) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75592, Q5JSX8, Q5VZN6, Q6PIB6, Q9UQ11, Q9Y6E4 | Gene names | MYCBP2, KIAA0916, PAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
DNJC8_HUMAN
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.047272 (rank : 75) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O75937, Q6IBA4, Q8N4Z5, Q9P051, Q9P067 | Gene names | DNAJC8, SPF31 | |||
|
Domain Architecture |
|
|||||
| Description | DnaJ homolog subfamily C member 8 (Splicing protein spf31). | |||||
|
DNJC8_MOUSE
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.047020 (rank : 77) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6NZB0, Q6PG72, Q8C2M6 | Gene names | Dnajc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 8. | |||||
|
GDF2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.018302 (rank : 95) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UK05, Q9Y571 | Gene names | GDF2, BMP9 | |||
|
Domain Architecture |
|
|||||
| Description | Growth/differentiation factor 2 precursor (GDF-2) (Bone morphogenetic protein 9) (BMP-9). | |||||
|
GSCR2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.053419 (rank : 64) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NZM5, Q9BTC6, Q9HAX6, Q9NPP1, Q9NPR4, Q9UFI2 | Gene names | GLTSCR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 2 protein (p60). | |||||
|
LPIN1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.026174 (rank : 87) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14693 | Gene names | LPIN1, KIAA0188 | |||
|
Domain Architecture |
|
|||||
| Description | Lipin-1. | |||||
|
MAMC1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.020987 (rank : 92) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z553 | Gene names | MAMDC1, MDGA2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 1 precursor (MAM domain-containing glycosylphosphatidylinositol anchor protein 2). | |||||
|
MYH9_MOUSE
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.011699 (rank : 103) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
RBM10_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.112034 (rank : 30) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
RBM12_HUMAN
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.056705 (rank : 55) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NTZ6, O94865, Q8N3B1, Q9H196 | Gene names | RBM12, KIAA0765 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
RBM12_MOUSE
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.057239 (rank : 54) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8R4X3, Q8K373, Q8R302, Q9CS80 | Gene names | Rbm12 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
AGGF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.094655 (rank : 34) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7TN31, Q8R2S6, Q9CQR9, Q9CU87, Q9D768 | Gene names | Aggf1, Vg5q | |||
|
Domain Architecture |
|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (mVG5Q). | |||||
|
CTNA3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.014032 (rank : 98) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q65CL1, Q3UQ61, Q8C0N3 | Gene names | Ctnna3, Catna3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Catenin alpha-3 (Alpha T-catenin) (Cadherin-associated protein). | |||||
|
ENAM_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.034962 (rank : 81) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRM1, Q9H3D1 | Gene names | ENAM | |||
|
Domain Architecture |

|
|||||
| Description | Enamelin precursor. | |||||
|
MAGI2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.019933 (rank : 94) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
MYOC_HUMAN
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.017854 (rank : 96) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 489 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99972, O00620, Q7Z6Q9 | Gene names | MYOC, GLC1A, TIGR | |||
|
Domain Architecture |

|
|||||
| Description | Myocilin precursor (Trabecular meshwork-induced glucocorticoid response protein). | |||||
|
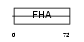
AGGF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.090672 (rank : 35) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8N302, O00581, Q53YS3, Q9BU84, Q9NW66 | Gene names | AGGF1, VG5Q | |||
|
Domain Architecture |
|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (Vasculogenesis gene on 5q protein) (hVG5Q). | |||||
|
MAGI2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.015450 (rank : 97) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WVQ1, Q3UH81, Q6GT88, Q8BYT1, Q8CA85 | Gene names | Magi2, Acvrinp1, Aip1, Arip1 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Activin receptor-interacting protein 1) (Acvrip1). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.025732 (rank : 88) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
ZN318_HUMAN
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.032517 (rank : 83) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5VUA4, O94796, Q4G0E4, Q8NEM6, Q9UNU8, Q9Y2W9 | Gene names | ZNF318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Endocrine regulatory protein). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.009255 (rank : 105) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
CCNK_MOUSE
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.033608 (rank : 82) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
CSTFT_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.056502 (rank : 56) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H0L4, O75174, Q53HK6, Q7LGE8, Q8N6T1 | Gene names | CSTF2T, KIAA0689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage stimulation factor 64 kDa subunit, tau variant (CSTF 64 kDa subunit, tau variant) (CF-1 64 kDa subunit, tau variant) (TauCstF-64). | |||||
|
CSTFT_MOUSE
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.059861 (rank : 49) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8C7E9, Q6A016, Q8BHH7, Q8C3W7, Q8R2Y1, Q9EPU3 | Gene names | Cstf2t, Kiaa0689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage stimulation factor 64 kDa subunit, tau variant (CSTF 64 kDa subunit, tau variant) (CF-1 64 kDa subunit, tau variant) (TauCstF-64). | |||||
|
ELOA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.021475 (rank : 90) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
GPTC3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.052779 (rank : 69) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BIY1 | Gene names | Gpatc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
RIMS2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.012211 (rank : 101) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UQ26, O43413, Q86XL9, Q8IWV9, Q8IWW1 | Gene names | RIMS2, KIAA0751, RIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2). | |||||
|
SF04_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.066586 (rank : 42) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8IWZ8, O60378, Q6P3X9, Q8TCQ4, Q8WWT4, Q8WWT5, Q9NTG3 | Gene names | SF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4 (RNA-binding protein RBP). | |||||
|
SFR14_MOUSE
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.059580 (rank : 50) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.031235 (rank : 84) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
TOR3A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.020442 (rank : 93) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H497, Q5M7Y7, Q8WVA7, Q8WWM2, Q9H495, Q9H6E7 | Gene names | TOR3A, ADIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Torsin-3A precursor (Torsin family 3 member A) (ATP-dependent interferon-responsive protein). | |||||
|
BRD8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.013038 (rank : 99) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H0E9, O43178, Q15355, Q59GN0, Q969M9 | Gene names | BRD8, SMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8 (p120) (Skeletal muscle abundant protein) (Thyroid hormone receptor coactivating protein 120kDa) (TrCP120). | |||||
|
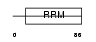
CSTF2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.058567 (rank : 52) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P33240, Q5H951, Q6LA74, Q8N502 | Gene names | CSTF2 | |||
|
Domain Architecture |
|
|||||
| Description | Cleavage stimulation factor 64 kDa subunit (CSTF 64 kDa subunit) (CF-1 64 kDa subunit) (CstF-64). | |||||
|
CSTF2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.059888 (rank : 48) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BIQ5, Q8K1Y6, Q9ERC2 | Gene names | Cstf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage stimulation factor 64 kDa subunit (CSTF 64 kDa subunit) (CF-1 64 kDa subunit) (CstF-64). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.042875 (rank : 78) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PABP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.060842 (rank : 46) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P29341 | Gene names | Pabpc1, Pabp1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 1 (Poly(A)-binding protein 1) (PABP 1). | |||||
|
PCD15_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.012213 (rank : 100) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99PJ1 | Gene names | Pcdh15 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin-15 precursor. | |||||
|
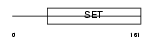
PRDM9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | -0.002113 (rank : 109) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NQV7 | Gene names | PRDM9, PFM6 | |||
|
Domain Architecture |
|
|||||
| Description | PR domain zinc finger protein 9 (PR domain-containing protein 9). | |||||
|
SEPT4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.008580 (rank : 107) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43236, Q9UM58 | Gene names | SEPT4, PNUTL2 | |||
|
Domain Architecture |

|
|||||
| Description | Septin-4 (Peanut-like protein 2) (Brain protein H5) (Cell division control-related protein 2) (hCDCREL-2) (Bradeion beta) (CE5B3 beta) (Cerebral protein 7). | |||||
|
TIA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.053116 (rank : 68) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P31483 | Gene names | TIA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolysin TIA-1 isoform p40 (RNA-binding protein TIA-1) (p40-TIA-1) [Contains: Nucleolysin TIA-1 isoform p15 (p15-TIA-1)]. | |||||
|
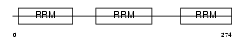
TIA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.052571 (rank : 70) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52912 | Gene names | Tia1, Tia | |||
|
Domain Architecture |
|
|||||
| Description | Nucleolysin TIA-1 (RNA-binding protein TIA-1). | |||||
|
TUFT1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.010329 (rank : 104) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 732 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O08970 | Gene names | Tuft1 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin. | |||||
|
ZFP64_MOUSE
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | -0.001684 (rank : 108) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 733 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99KE8, P97365, Q9CWR3 | Gene names | Zfp64 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 64 (Zfp-64). | |||||
|
ZYX_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.008774 (rank : 106) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62523, P70461 | Gene names | Zyx | |||
|
Domain Architecture |

|
|||||
| Description | Zyxin. | |||||
|
GPTC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.058066 (rank : 53) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9NW75, Q86YE7 | Gene names | GPATC2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
GPTC2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.053286 (rank : 66) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q7TQC7, Q8BNJ9, Q8BPM1, Q8CDH9 | Gene names | Gpatc2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
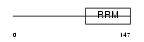
MAN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.054906 (rank : 59) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y2U8, Q9NT47, Q9NYA5 | Gene names | LEMD3, MAN1 | |||
|
Domain Architecture |
|
|||||
| Description | Inner nuclear membrane protein Man1 (LEM domain-containing protein 3). | |||||
|
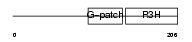
NKRF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.055050 (rank : 58) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15226, Q9UJ91 | Gene names | NKRF, ITBA4, NRF | |||
|
Domain Architecture |
|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF) (ITBA4 protein). | |||||
|
NKRF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.054450 (rank : 61) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BY02, Q8BJU5, Q9CRL2 | Gene names | Nkrf, Nrf | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF). | |||||
|
PABP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.053237 (rank : 67) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P11940, Q15097, Q93004 | Gene names | PABPC1, PAB1, PABP1, PABPC2 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 1 (Poly(A)-binding protein 1) (PABP 1). | |||||
|
PABP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.053396 (rank : 65) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9H361, Q8NHV0, Q9H086 | Gene names | PABPC3, PABP3, PABPL3 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 3 (Poly(A)-binding protein 3) (PABP 3) (Testis-specific poly(A)-binding protein). | |||||
|
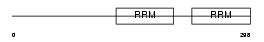
RBM23_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.055298 (rank : 57) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86U06, Q8ND16, Q8TB88, Q8WY40, Q9BUJ1, Q9NVV7 | Gene names | RBM23, RNPC4 | |||
|
Domain Architecture |
|
|||||
| Description | Probable RNA-binding protein 23 (RNA-binding motif protein 23) (RNA- binding region-containing protein 4) (Splicing factor SF2). | |||||
|
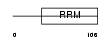
RBMX2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.054367 (rank : 62) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
RBMX2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.050890 (rank : 72) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8R0F5, Q3TI42 | Gene names | Rbmx2 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
SF04_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.054161 (rank : 63) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CH02, Q8R094 | Gene names | Sf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4. | |||||
|
U2AFL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.060592 (rank : 47) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15695, Q13570 | Gene names | ZRSR1, U2AF1-RS1, U2AF1L1, U2AF1RS1, U2AFBPL | |||
|
Domain Architecture |
|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1). | |||||
|
U2AFL_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.061533 (rank : 45) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q64707 | Gene names | Zrsr1, Sp2, Sp2-7, U2af1-rs1, U2af1l1 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1) (SP2). | |||||
|
U2AFM_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.059167 (rank : 51) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q15696 | Gene names | ZRSR2, U2AF1-RS2, U2AF1L2, U2AF1RS2 | |||
|
Domain Architecture |
|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2) (NY-REN-20 antigen). | |||||
|
U2AFM_MOUSE
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.054742 (rank : 60) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q62377 | Gene names | Zrsr2, U2af1-rs2, U2af1l2, U2af1rs2 | |||
|
Domain Architecture |
|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2). | |||||
|
SPF45_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 94 | |
| SwissProt Accessions | Q96I25, Q96GY6 | Gene names | RBM17, SPF45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
SPF45_MOUSE
|
||||||
| NC score | 0.994041 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q8JZX4, O75939 | Gene names | Rbm17, Spf45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
U2AF2_HUMAN
|
||||||
| NC score | 0.244929 (rank : 3) | θ value | 1.43324e-05 (rank : 3) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P26368 | Gene names | U2AF2, U2AF65 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor U2AF 65 kDa subunit (U2 auxiliary factor 65 kDa subunit) (U2 snRNP auxiliary factor large subunit) (hU2AF(65)). | |||||
|
U2AF2_MOUSE
|
||||||
| NC score | 0.244049 (rank : 4) | θ value | 1.43324e-05 (rank : 4) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P26369 | Gene names | U2af2, U2af65 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor U2AF 65 kDa subunit (U2 auxiliary factor 65 kDa subunit) (U2 snRNP auxiliary factor large subunit). | |||||
|
K0082_MOUSE
|
||||||
| NC score | 0.228269 (rank : 5) | θ value | 0.00390308 (rank : 10) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9DBC3, Q3U3G5, Q3U7Y9, Q6A0D5, Q8C7V0 | Gene names | Kiaa0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
K0082_HUMAN
|
||||||
| NC score | 0.228082 (rank : 6) | θ value | 0.00509761 (rank : 14) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8N1G2, Q14670, Q96FJ9 | Gene names | KIAA0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
U2AF1_MOUSE
|
||||||
| NC score | 0.213313 (rank : 7) | θ value | 0.00134147 (rank : 9) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D883, Q564E4, Q99LX2, Q9CZ98 | Gene names | U2af1 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
U2AF1_HUMAN
|
||||||
| NC score | 0.212708 (rank : 8) | θ value | 0.00134147 (rank : 8) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q01081 | Gene names | U2AF1, U2AF35 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
HTSF1_MOUSE
|
||||||
| NC score | 0.196335 (rank : 9) | θ value | 0.00102713 (rank : 7) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
HTSF1_HUMAN
|
||||||
| NC score | 0.188349 (rank : 10) | θ value | 0.00509761 (rank : 13) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43719, Q59G06, Q99730 | Gene names | HTATSF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 (Tat-SF1). | |||||
|
PINX1_MOUSE
|
||||||
| NC score | 0.175798 (rank : 11) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9CZX5, Q91WZ9, Q9D0C2 | Gene names | Pinx1, Lpts | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (LPTS1) (67-11-3 protein). | |||||
|
VPK9_HUMAN
|
||||||
| NC score | 0.173596 (rank : 12) | θ value | 0.365318 (rank : 24) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P63124 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q13.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K104 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK1_HUMAN
|
||||||
| NC score | 0.172710 (rank : 13) | θ value | 0.365318 (rank : 23) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P63119 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
VPK17_HUMAN
|
||||||
| NC score | 0.171198 (rank : 14) | θ value | 0.47712 (rank : 25) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63125 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_11q22.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
VPK7_HUMAN
|
||||||
| NC score | 0.170452 (rank : 15) | θ value | 0.62314 (rank : 30) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63123 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q23.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K18 Pro protein) (HERV-K110 Pro protein) (HERV-K(C1a) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK11_HUMAN
|
||||||
| NC score | 0.167477 (rank : 16) | θ value | 0.62314 (rank : 29) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63129, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K101 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
VPK5_HUMAN
|
||||||
| NC score | 0.166686 (rank : 17) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P63121 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K113 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
VPK3_HUMAN
|
||||||
| NC score | 0.166610 (rank : 18) | θ value | 0.813845 (rank : 33) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63120 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19q12 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K(C19) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK10_HUMAN
|
||||||
| NC score | 0.166568 (rank : 19) | θ value | 1.06291 (rank : 38) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P10265, Q9UKH6 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-K_5q33.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K10 Pro protein) (HERV-K107 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK12_HUMAN
|
||||||
| NC score | 0.166528 (rank : 20) | θ value | 1.06291 (rank : 39) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P63131, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K102 Pro protein) (HERV-K(III) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK2_HUMAN
|
||||||
| NC score | 0.164098 (rank : 21) | θ value | 1.38821 (rank : 48) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y6I0, Q9UKH5 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K(HML-2.HOM) Pro protein) (HERV-K108 Pro protein) (HERV-K(C7) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK6_HUMAN
|
||||||
| NC score | 0.162018 (rank : 22) | θ value | 0.813845 (rank : 35) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63122 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K115 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK4_HUMAN
|
||||||
| NC score | 0.161930 (rank : 23) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P63127, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K109 Pro protein) (HERV-K(C6) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
PINX1_HUMAN
|
||||||
| NC score | 0.154729 (rank : 24) | θ value | 0.813845 (rank : 31) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96BK5, Q7Z7J8, Q96QD7, Q9HBU7, Q9NWW2 | Gene names | PINX1, LPTS | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (67-11-3 protein). | |||||
|
SON_MOUSE
|
||||||
| NC score | 0.146773 (rank : 25) | θ value | 0.00390308 (rank : 12) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
TFP11_HUMAN
|
||||||
| NC score | 0.145504 (rank : 26) | θ value | 0.813845 (rank : 32) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UBB9, O95908, Q5H8V8, Q9UGV7, Q9Y2Q8 | Gene names | TFIP11 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11. | |||||
|
RBM5_HUMAN
|
||||||
| NC score | 0.143010 (rank : 27) | θ value | 0.163984 (rank : 15) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P52756, Q93021, Q9BU14, Q9UKY8, Q9UL24 | Gene names | RBM5 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15) (Protein G15) (NY-REN-9 antigen). | |||||
|
TFP11_MOUSE
|
||||||
| NC score | 0.142993 (rank : 28) | θ value | 1.06291 (rank : 37) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9ERA6, Q8VD06 | Gene names | Tfip11, Tip39 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11 (Tuftelin-interacting protein 39). | |||||
|
RBM5_MOUSE
|
||||||
| NC score | 0.142762 (rank : 29) | θ value | 0.163984 (rank : 16) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q91YE7, Q3UZ19, Q99KV9 | Gene names | Rbm5, Luca15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15). | |||||
|
RBM10_HUMAN
|
||||||
| NC score | 0.112034 (rank : 30) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.101458 (rank : 31) | θ value | 0.00390308 (rank : 11) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
RNPC2_MOUSE
|
||||||
| NC score | 0.096355 (rank : 32) | θ value | 0.365318 (rank : 21) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8VH51, Q99KV0 | Gene names | Rnpc2, Caper | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding region-containing protein 2 (Coactivator of activating protein 1 and estrogen receptors) (Coactivator of AP-1 and ERs) (Transcription coactivator CAPER). | |||||
|
RNPC2_HUMAN
|
||||||
| NC score | 0.096232 (rank : 33) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q14498, Q14499 | Gene names | RNPC2, HCC1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding region-containing protein 2 (Hepatocellular carcinoma protein 1) (Splicing factor HCC1). | |||||
|
AGGF1_MOUSE
|
||||||
| NC score | 0.094655 (rank : 34) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7TN31, Q8R2S6, Q9CQR9, Q9CU87, Q9D768 | Gene names | Aggf1, Vg5q | |||
|
Domain Architecture |

|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (mVG5Q). | |||||
|
AGGF1_HUMAN
|
||||||
| NC score | 0.090672 (rank : 35) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8N302, O00581, Q53YS3, Q9BU84, Q9NW66 | Gene names | AGGF1, VG5Q | |||
|
Domain Architecture |

|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (Vasculogenesis gene on 5q protein) (hVG5Q). | |||||
|
UHMK1_MOUSE
|
||||||
| NC score | 0.087215 (rank : 36) | θ value | 0.000158464 (rank : 6) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 809 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97343, Q61775, Q9CYT1 | Gene names | Uhmk1, Kis, Kist | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Kist (EC 2.7.11.1) (Kinase interacting with stathmin) (U2AF homology motif kinase 1). | |||||
|
UHMK1_HUMAN
|
||||||
| NC score | 0.086742 (rank : 37) | θ value | 0.000158464 (rank : 5) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TAS1, Q96C22 | Gene names | UHMK1, KIS, KIST | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Kist (EC 2.7.11.1) (Kinase interacting with stathmin) (U2AF homology motif kinase 1). | |||||
|
PABP5_HUMAN
|
||||||
| NC score | 0.074063 (rank : 38) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96DU9, Q6P529, Q9UFE5 | Gene names | PABPC5, PABP5 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 5 (Poly(A)-binding protein 5) (PABP 5). | |||||
|
PABP4_HUMAN
|
||||||
| NC score | 0.072111 (rank : 39) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13310, Q6P0N3 | Gene names | PABPC4, APP1, PABP4 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 4 (Poly(A)-binding protein 4) (PABP 4) (Inducible poly(A)-binding protein) (iPABP) (Activated-platelet protein 1) (APP-1). | |||||
|
PRGC1_MOUSE
|
||||||
| NC score | 0.068502 (rank : 40) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O70343 | Gene names | Ppargc1a, Pgc1, Pgc1a, Ppargc1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPAR gamma coactivator 1-alpha) (PPARGC-1-alpha) (PGC-1-alpha). | |||||
|
RB15B_MOUSE
|
||||||
| NC score | 0.067696 (rank : 41) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6PHZ5, Q8C6G2 | Gene names | Rbm15b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
SF04_HUMAN
|
||||||
| NC score | 0.066586 (rank : 42) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8IWZ8, O60378, Q6P3X9, Q8TCQ4, Q8WWT4, Q8WWT5, Q9NTG3 | Gene names | SF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4 (RNA-binding protein RBP). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.066449 (rank : 43) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
RB15B_HUMAN
|
||||||
| NC score | 0.065259 (rank : 44) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8NDT2, Q6QE19, Q9BV96 | Gene names | RBM15B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
U2AFL_MOUSE
|
||||||
| NC score | 0.061533 (rank : 45) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q64707 | Gene names | Zrsr1, Sp2, Sp2-7, U2af1-rs1, U2af1l1 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1) (SP2). | |||||
|
PABP1_MOUSE
|
||||||
| NC score | 0.060842 (rank : 46) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P29341 | Gene names | Pabpc1, Pabp1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 1 (Poly(A)-binding protein 1) (PABP 1). | |||||
|
U2AFL_HUMAN
|
||||||
| NC score | 0.060592 (rank : 47) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15695, Q13570 | Gene names | ZRSR1, U2AF1-RS1, U2AF1L1, U2AF1RS1, U2AFBPL | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1). | |||||
|
CSTF2_MOUSE
|
||||||
| NC score | 0.059888 (rank : 48) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BIQ5, Q8K1Y6, Q9ERC2 | Gene names | Cstf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage stimulation factor 64 kDa subunit (CSTF 64 kDa subunit) (CF-1 64 kDa subunit) (CstF-64). | |||||
|
CSTFT_MOUSE
|
||||||
| NC score | 0.059861 (rank : 49) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8C7E9, Q6A016, Q8BHH7, Q8C3W7, Q8R2Y1, Q9EPU3 | Gene names | Cstf2t, Kiaa0689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage stimulation factor 64 kDa subunit, tau variant (CSTF 64 kDa subunit, tau variant) (CF-1 64 kDa subunit, tau variant) (TauCstF-64). | |||||
|
SFR14_MOUSE
|
||||||
| NC score | 0.059580 (rank : 50) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
U2AFM_HUMAN
|
||||||
| NC score | 0.059167 (rank : 51) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q15696 | Gene names | ZRSR2, U2AF1-RS2, U2AF1L2, U2AF1RS2 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2) (NY-REN-20 antigen). | |||||
|
CSTF2_HUMAN
|
||||||
| NC score | 0.058567 (rank : 52) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P33240, Q5H951, Q6LA74, Q8N502 | Gene names | CSTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Cleavage stimulation factor 64 kDa subunit (CSTF 64 kDa subunit) (CF-1 64 kDa subunit) (CstF-64). | |||||
|
GPTC2_HUMAN
|
||||||
| NC score | 0.058066 (rank : 53) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9NW75, Q86YE7 | Gene names | GPATC2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
RBM12_MOUSE
|
||||||
| NC score | 0.057239 (rank : 54) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8R4X3, Q8K373, Q8R302, Q9CS80 | Gene names | Rbm12 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
RBM12_HUMAN
|
||||||
| NC score | 0.056705 (rank : 55) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NTZ6, O94865, Q8N3B1, Q9H196 | Gene names | RBM12, KIAA0765 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
CSTFT_HUMAN
|
||||||
| NC score | 0.056502 (rank : 56) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H0L4, O75174, Q53HK6, Q7LGE8, Q8N6T1 | Gene names | CSTF2T, KIAA0689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage stimulation factor 64 kDa subunit, tau variant (CSTF 64 kDa subunit, tau variant) (CF-1 64 kDa subunit, tau variant) (TauCstF-64). | |||||
|
RBM23_HUMAN
|
||||||
| NC score | 0.055298 (rank : 57) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86U06, Q8ND16, Q8TB88, Q8WY40, Q9BUJ1, Q9NVV7 | Gene names | RBM23, RNPC4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable RNA-binding protein 23 (RNA-binding motif protein 23) (RNA- binding region-containing protein 4) (Splicing factor SF2). | |||||
|
NKRF_HUMAN
|
||||||
| NC score | 0.055050 (rank : 58) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15226, Q9UJ91 | Gene names | NKRF, ITBA4, NRF | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF) (ITBA4 protein). | |||||
|
MAN1_HUMAN
|
||||||
| NC score | 0.054906 (rank : 59) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y2U8, Q9NT47, Q9NYA5 | Gene names | LEMD3, MAN1 | |||
|
Domain Architecture |

|
|||||
| Description | Inner nuclear membrane protein Man1 (LEM domain-containing protein 3). | |||||
|
U2AFM_MOUSE
|
||||||
| NC score | 0.054742 (rank : 60) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q62377 | Gene names | Zrsr2, U2af1-rs2, U2af1l2, U2af1rs2 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2). | |||||
|
NKRF_MOUSE
|
||||||
| NC score | 0.054450 (rank : 61) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BY02, Q8BJU5, Q9CRL2 | Gene names | Nkrf, Nrf | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF). | |||||
|
RBMX2_HUMAN
|
||||||
| NC score | 0.054367 (rank : 62) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
SF04_MOUSE
|
||||||
| NC score | 0.054161 (rank : 63) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CH02, Q8R094 | Gene names | Sf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4. | |||||
|
GSCR2_HUMAN
|
||||||
| NC score | 0.053419 (rank : 64) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NZM5, Q9BTC6, Q9HAX6, Q9NPP1, Q9NPR4, Q9UFI2 | Gene names | GLTSCR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 2 protein (p60). | |||||
|
PABP3_HUMAN
|
||||||
| NC score | 0.053396 (rank : 65) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9H361, Q8NHV0, Q9H086 | Gene names | PABPC3, PABP3, PABPL3 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 3 (Poly(A)-binding protein 3) (PABP 3) (Testis-specific poly(A)-binding protein). | |||||
|
GPTC2_MOUSE
|
||||||
| NC score | 0.053286 (rank : 66) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q7TQC7, Q8BNJ9, Q8BPM1, Q8CDH9 | Gene names | Gpatc2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
PABP1_HUMAN
|
||||||
| NC score | 0.053237 (rank : 67) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P11940, Q15097, Q93004 | Gene names | PABPC1, PAB1, PABP1, PABPC2 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 1 (Poly(A)-binding protein 1) (PABP 1). | |||||
|
TIA1_HUMAN
|
||||||
| NC score | 0.053116 (rank : 68) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P31483 | Gene names | TIA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolysin TIA-1 isoform p40 (RNA-binding protein TIA-1) (p40-TIA-1) [Contains: Nucleolysin TIA-1 isoform p15 (p15-TIA-1)]. | |||||
|
GPTC3_MOUSE
|
||||||
| NC score | 0.052779 (rank : 69) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BIY1 | Gene names | Gpatc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
TIA1_MOUSE
|
||||||
| NC score | 0.052571 (rank : 70) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52912 | Gene names | Tia1, Tia | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolysin TIA-1 (RNA-binding protein TIA-1). | |||||
|
WWC3_HUMAN
|
||||||
| NC score | 0.051395 (rank : 71) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ULE0, Q659C1, Q9BTQ1 | Gene names | WWC3, KIAA1280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WWC family member 3. | |||||
|
RBMX2_MOUSE
|
||||||
| NC score | 0.050890 (rank : 72) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8R0F5, Q3TI42 | Gene names | Rbmx2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.049380 (rank : 73) | θ value | 0.365318 (rank : 22) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
JHD1B_MOUSE
|
||||||
| NC score | 0.047697 (rank : 74) | θ value | 0.62314 (rank : 26) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
DNJC8_HUMAN
|
||||||
| NC score | 0.047272 (rank : 75) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O75937, Q6IBA4, Q8N4Z5, Q9P051, Q9P067 | Gene names | DNAJC8, SPF31 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 8 (Splicing protein spf31). | |||||
|
JHD1B_HUMAN
|
||||||
| NC score | 0.047195 (rank : 76) | θ value | 1.06291 (rank : 36) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
DNJC8_MOUSE
|
||||||
| NC score | 0.047020 (rank : 77) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6NZB0, Q6PG72, Q8C2M6 | Gene names | Dnajc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 8. | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.042875 (rank : 78) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
DNJC7_MOUSE
|
||||||
| NC score | 0.038476 (rank : 79) | θ value | 1.38821 (rank : 43) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QYI3, Q3TKR1, Q8BPG3, Q8CIL2, Q9CT29, Q9D026 | Gene names | Dnajc7, Ttc2 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2) (MDj11). | |||||
|
DNJC7_HUMAN
|
||||||
| NC score | 0.038155 (rank : 80) | θ value | 1.38821 (rank : 42) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99615 | Gene names | DNAJC7, TPR2, TTC2 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2). | |||||
|
ENAM_HUMAN
|
||||||
| NC score | 0.034962 (rank : 81) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRM1, Q9H3D1 | Gene names | ENAM | |||
|
Domain Architecture |

|
|||||
| Description | Enamelin precursor. | |||||
|
CCNK_MOUSE
|
||||||
| NC score | 0.033608 (rank : 82) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
ZN318_HUMAN
|
||||||
| NC score | 0.032517 (rank : 83) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5VUA4, O94796, Q4G0E4, Q8NEM6, Q9UNU8, Q9Y2W9 | Gene names | ZNF318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Endocrine regulatory protein). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.031235 (rank : 84) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
CASP8_MOUSE
|
||||||
| NC score | 0.027723 (rank : 85) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O89110, O35669 | Gene names | Casp8 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase-8 precursor (EC 3.4.22.-) (CASP-8) [Contains: Caspase-8 subunit p18; Caspase-8 subunit p10]. | |||||
|
MYCB2_HUMAN
|
||||||
| NC score | 0.027520 (rank : 86) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75592, Q5JSX8, Q5VZN6, Q6PIB6, Q9UQ11, Q9Y6E4 | Gene names | MYCBP2, KIAA0916, PAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
LPIN1_HUMAN
|
||||||
| NC score | 0.026174 (rank : 87) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14693 | Gene names | LPIN1, KIAA0188 | |||
|
Domain Architecture |

|
|||||
| Description | Lipin-1. | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.025732 (rank : 88) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
MAMC1_MOUSE
|
||||||
| NC score | 0.024257 (rank : 89) | θ value | 1.38821 (rank : 44) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P60755 | Gene names | Mamdc1, Mdga2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 1 precursor (MAM domain-containing glycosylphosphatidylinositol anchor protein 2). | |||||
|
ELOA1_HUMAN
|
||||||
| NC score | 0.021475 (rank : 90) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
ITSN1_MOUSE
|
||||||
| NC score | 0.021342 (rank : 91) | θ value | 0.365318 (rank : 18) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
MAMC1_HUMAN
|
||||||
| NC score | 0.020987 (rank : 92) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z553 | Gene names | MAMDC1, MDGA2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 1 precursor (MAM domain-containing glycosylphosphatidylinositol anchor protein 2). | |||||
|
TOR3A_HUMAN
|
||||||
| NC score | 0.020442 (rank : 93) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H497, Q5M7Y7, Q8WVA7, Q8WWM2, Q9H495, Q9H6E7 | Gene names | TOR3A, ADIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Torsin-3A precursor (Torsin family 3 member A) (ATP-dependent interferon-responsive protein). | |||||
|
MAGI2_HUMAN
|
||||||
| NC score | 0.019933 (rank : 94) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
GDF2_HUMAN
|
||||||
| NC score | 0.018302 (rank : 95) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UK05, Q9Y571 | Gene names | GDF2, BMP9 | |||
|
Domain Architecture |

|
|||||
| Description | Growth/differentiation factor 2 precursor (GDF-2) (Bone morphogenetic protein 9) (BMP-9). | |||||
|
MYOC_HUMAN
|
||||||
| NC score | 0.017854 (rank : 96) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 489 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99972, O00620, Q7Z6Q9 | Gene names | MYOC, GLC1A, TIGR | |||
|
Domain Architecture |

|
|||||
| Description | Myocilin precursor (Trabecular meshwork-induced glucocorticoid response protein). | |||||
|
MAGI2_MOUSE
|
||||||
| NC score | 0.015450 (rank : 97) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WVQ1, Q3UH81, Q6GT88, Q8BYT1, Q8CA85 | Gene names | Magi2, Acvrinp1, Aip1, Arip1 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Activin receptor-interacting protein 1) (Acvrip1). | |||||
|
CTNA3_MOUSE
|
||||||
| NC score | 0.014032 (rank : 98) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q65CL1, Q3UQ61, Q8C0N3 | Gene names | Ctnna3, Catna3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Catenin alpha-3 (Alpha T-catenin) (Cadherin-associated protein). | |||||
|
BRD8_HUMAN
|
||||||
| NC score | 0.013038 (rank : 99) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H0E9, O43178, Q15355, Q59GN0, Q969M9 | Gene names | BRD8, SMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8 (p120) (Skeletal muscle abundant protein) (Thyroid hormone receptor coactivating protein 120kDa) (TrCP120). | |||||
|
PCD15_MOUSE
|
||||||
| NC score | 0.012213 (rank : 100) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99PJ1 | Gene names | Pcdh15 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin-15 precursor. | |||||
|
RIMS2_HUMAN
|
||||||
| NC score | 0.012211 (rank : 101) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UQ26, O43413, Q86XL9, Q8IWV9, Q8IWW1 | Gene names | RIMS2, KIAA0751, RIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2). | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.011871 (rank : 102) | θ value | 1.38821 (rank : 45) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH9_MOUSE
|
||||||
| NC score | 0.011699 (rank : 103) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
TUFT1_MOUSE
|
||||||
| NC score | 0.010329 (rank : 104) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 732 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O08970 | Gene names | Tuft1 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin. | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.009255 (rank : 105) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
ZYX_MOUSE
|
||||||
| NC score | 0.008774 (rank : 106) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62523, P70461 | Gene names | Zyx | |||
|
Domain Architecture |

|
|||||
| Description | Zyxin. | |||||
|
SEPT4_HUMAN
|
||||||
| NC score | 0.008580 (rank : 107) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43236, Q9UM58 | Gene names | SEPT4, PNUTL2 | |||
|
Domain Architecture |

|
|||||
| Description | Septin-4 (Peanut-like protein 2) (Brain protein H5) (Cell division control-related protein 2) (hCDCREL-2) (Bradeion beta) (CE5B3 beta) (Cerebral protein 7). | |||||
|
ZFP64_MOUSE
|
||||||
| NC score | -0.001684 (rank : 108) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 733 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99KE8, P97365, Q9CWR3 | Gene names | Zfp64 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 64 (Zfp-64). | |||||
|
PRDM9_HUMAN
|
||||||
| NC score | -0.002113 (rank : 109) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NQV7 | Gene names | PRDM9, PFM6 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 9 (PR domain-containing protein 9). | |||||