Please be patient as the page loads
|
TFP11_MOUSE
|
||||||
| SwissProt Accessions | Q9ERA6, Q8VD06 | Gene names | Tfip11, Tip39 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11 (Tuftelin-interacting protein 39). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TFP11_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.990661 (rank : 2) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9UBB9, O95908, Q5H8V8, Q9UGV7, Q9Y2Q8 | Gene names | TFIP11 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11. | |||||
|
TFP11_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 95 | |
| SwissProt Accessions | Q9ERA6, Q8VD06 | Gene names | Tfip11, Tip39 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11 (Tuftelin-interacting protein 39). | |||||
|
ZGPAT_MOUSE
|
||||||
| θ value | 1.53129e-07 (rank : 3) | NC score | 0.383948 (rank : 12) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8VDM1, Q8BWW2 | Gene names | Zgpat | |||
|
Domain Architecture |
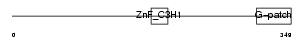
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein. | |||||
|
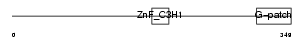
ZGPAT_HUMAN
|
||||||
| θ value | 2.61198e-07 (rank : 4) | NC score | 0.381523 (rank : 13) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8N5A5, Q5JWI9, Q8NC55, Q96JI0, Q96JU4, Q9H401 | Gene names | ZGPAT, KIAA1847, ZC3H9, ZC3HDC9 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein (Zinc finger CCCH domain-containing protein 9). | |||||
|
RBM10_HUMAN
|
||||||
| θ value | 7.59969e-07 (rank : 5) | NC score | 0.331201 (rank : 17) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
VPK7_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 6) | NC score | 0.398123 (rank : 3) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63123 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q23.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K18 Pro protein) (HERV-K110 Pro protein) (HERV-K(C1a) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
GPTC3_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 7) | NC score | 0.278256 (rank : 25) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96I76, Q5JYH2, Q8NDJ2, Q9H9Z3 | Gene names | GPATC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
GPTC3_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 8) | NC score | 0.288813 (rank : 23) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BIY1 | Gene names | Gpatc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
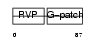
VPK10_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 9) | NC score | 0.392749 (rank : 7) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P10265, Q9UKH6 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | HERV-K_5q33.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K10 Pro protein) (HERV-K107 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK12_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 10) | NC score | 0.392652 (rank : 8) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P63131, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K102 Pro protein) (HERV-K(III) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK17_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 11) | NC score | 0.392142 (rank : 9) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P63125 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_11q22.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
VPK1_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 12) | NC score | 0.393400 (rank : 5) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P63119 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
VPK2_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 13) | NC score | 0.390435 (rank : 10) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y6I0, Q9UKH5 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K(HML-2.HOM) Pro protein) (HERV-K108 Pro protein) (HERV-K(C7) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK5_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 14) | NC score | 0.393073 (rank : 6) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P63121 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K113 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
VPK9_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 15) | NC score | 0.394163 (rank : 4) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P63124 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q13.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K104 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK11_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 16) | NC score | 0.384385 (rank : 11) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63129, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K101 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
RBM5_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 17) | NC score | 0.310886 (rank : 21) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P52756, Q93021, Q9BU14, Q9UKY8, Q9UL24 | Gene names | RBM5 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15) (Protein G15) (NY-REN-9 antigen). | |||||
|
RBM5_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 18) | NC score | 0.310143 (rank : 22) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q91YE7, Q3UZ19, Q99KV9 | Gene names | Rbm5, Luca15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15). | |||||
|
GPTC2_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 19) | NC score | 0.323593 (rank : 18) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NW75, Q86YE7 | Gene names | GPATC2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
GPTC2_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 20) | NC score | 0.319159 (rank : 19) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q7TQC7, Q8BNJ9, Q8BPM1, Q8CDH9 | Gene names | Gpatc2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
VPK4_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 21) | NC score | 0.369453 (rank : 16) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63127, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K109 Pro protein) (HERV-K(C6) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK6_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 22) | NC score | 0.369632 (rank : 15) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63122 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K115 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK3_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 23) | NC score | 0.378373 (rank : 14) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63120 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19q12 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K(C19) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
SF04_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 24) | NC score | 0.251723 (rank : 30) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8IWZ8, O60378, Q6P3X9, Q8TCQ4, Q8WWT4, Q8WWT5, Q9NTG3 | Gene names | SF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4 (RNA-binding protein RBP). | |||||
|
K0082_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 25) | NC score | 0.260606 (rank : 27) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8N1G2, Q14670, Q96FJ9 | Gene names | KIAA0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
PINX1_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 26) | NC score | 0.316287 (rank : 20) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CZX5, Q91WZ9, Q9D0C2 | Gene names | Pinx1, Lpts | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (LPTS1) (67-11-3 protein). | |||||
|
SF04_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 27) | NC score | 0.242833 (rank : 34) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8CH02, Q8R094 | Gene names | Sf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4. | |||||
|
SFR14_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 28) | NC score | 0.244820 (rank : 33) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8IX01, O15071, O60369, Q5JPH7, Q8WUF7 | Gene names | SFRS14, KIAA0365 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
SFR14_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 29) | NC score | 0.235957 (rank : 35) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 30) | NC score | 0.103334 (rank : 42) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SON_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 31) | NC score | 0.150621 (rank : 37) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
K0082_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 32) | NC score | 0.250813 (rank : 31) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9DBC3, Q3U3G5, Q3U7Y9, Q6A0D5, Q8C7V0 | Gene names | Kiaa0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
TB182_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 33) | NC score | 0.103526 (rank : 41) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C0C2 | Gene names | TNKS1BP1, KIAA1741, TAB182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 182 kDa tankyrase 1-binding protein. | |||||
|
RAD50_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 34) | NC score | 0.060709 (rank : 45) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
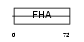
AGGF1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 35) | NC score | 0.261807 (rank : 26) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8N302, O00581, Q53YS3, Q9BU84, Q9NW66 | Gene names | AGGF1, VG5Q | |||
|
Domain Architecture |
|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (Vasculogenesis gene on 5q protein) (hVG5Q). | |||||
|
GOGA1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 36) | NC score | 0.051096 (rank : 47) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
NKRF_HUMAN
|
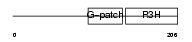
||||||
| θ value | 0.0252991 (rank : 37) | NC score | 0.252855 (rank : 29) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O15226, Q9UJ91 | Gene names | NKRF, ITBA4, NRF | |||
|
Domain Architecture |
|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF) (ITBA4 protein). | |||||
|
AGGF1_MOUSE
|
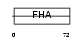
||||||
| θ value | 0.0330416 (rank : 38) | NC score | 0.259963 (rank : 28) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q7TN31, Q8R2S6, Q9CQR9, Q9CU87, Q9D768 | Gene names | Aggf1, Vg5q | |||
|
Domain Architecture |
|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (mVG5Q). | |||||
|
PINX1_HUMAN
|
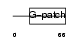
||||||
| θ value | 0.0330416 (rank : 39) | NC score | 0.282250 (rank : 24) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96BK5, Q7Z7J8, Q96QD7, Q9HBU7, Q9NWW2 | Gene names | PINX1, LPTS | |||
|
Domain Architecture |
|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (67-11-3 protein). | |||||
|
NKRF_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 40) | NC score | 0.250389 (rank : 32) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8BY02, Q8BJU5, Q9CRL2 | Gene names | Nkrf, Nrf | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF). | |||||
|
HXC10_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 41) | NC score | 0.045752 (rank : 49) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P31257, P31312, Q7TMT7 | Gene names | Hoxc10, Hox-3.6, Hoxc-10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein Hox-C10 (Hox-3.6). | |||||
|
ANR50_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 42) | NC score | 0.025720 (rank : 81) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULJ7, Q6N064, Q6ZSE6 | Gene names | ANKRD50, KIAA1223 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 50. | |||||
|
HXC10_HUMAN
|
||||||
| θ value | 0.163984 (rank : 43) | NC score | 0.043068 (rank : 51) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NYD6, O15219, O15220, Q9BVD5 | Gene names | HOXC10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-C10. | |||||
|
PEPL_MOUSE
|
||||||
| θ value | 0.21417 (rank : 44) | NC score | 0.038862 (rank : 52) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
RUFY2_MOUSE
|
||||||
| θ value | 0.21417 (rank : 45) | NC score | 0.044341 (rank : 50) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8R4C2, Q69ZH1 | Gene names | Rufy2, Kiaa1537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Leucine zipper FYVE-finger protein) (LZ-FYVE). | |||||
|
TSC1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 46) | NC score | 0.056721 (rank : 46) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q92574 | Gene names | TSC1, KIAA0243, TSC | |||
|
Domain Architecture |

|
|||||
| Description | Hamartin (Tuberous sclerosis 1 protein). | |||||
|
MYH10_HUMAN
|
||||||
| θ value | 0.279714 (rank : 47) | NC score | 0.032467 (rank : 65) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
VP13B_HUMAN
|
||||||
| θ value | 0.365318 (rank : 48) | NC score | 0.071812 (rank : 44) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z7G8, Q709C6, Q709C7, Q7Z7G4, Q7Z7G5, Q7Z7G6, Q7Z7G7, Q8NB77, Q9NWV1, Q9Y4E7 | Gene names | VPS13B, CHS1, COH1, KIAA0532 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vacuolar protein sorting-associated protein 13B (Cohen syndrome protein 1). | |||||
|
CP250_MOUSE
|
||||||
| θ value | 0.47712 (rank : 49) | NC score | 0.036738 (rank : 57) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
ARD1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 50) | NC score | 0.026987 (rank : 77) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 621 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P36406, Q9BZY4, Q9BZY5 | Gene names | TRIM23, ARD1, ARFD1, RNF46 | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein ARD-1 (ADP-ribosylation factor domain protein 1) (Tripartite motif-containing protein 23) (RING finger protein 46). | |||||
|
HXD10_HUMAN
|
||||||
| θ value | 0.62314 (rank : 51) | NC score | 0.036631 (rank : 59) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P28358 | Gene names | HOXD10, HOX4D | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D10 (Hox-4D) (Hox-4E). | |||||
|
SPF45_MOUSE
|
||||||
| θ value | 0.62314 (rank : 52) | NC score | 0.147471 (rank : 38) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8JZX4, O75939 | Gene names | Rbm17, Spf45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
KMHN1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 53) | NC score | 0.036873 (rank : 56) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q3V125, Q3TTQ8 | Gene names | Gm172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1 homolog. | |||||
|
MYH10_MOUSE
|
||||||
| θ value | 0.813845 (rank : 54) | NC score | 0.031408 (rank : 69) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 0.813845 (rank : 55) | NC score | 0.031965 (rank : 67) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
BAT4_HUMAN
|
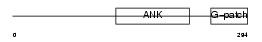
||||||
| θ value | 1.06291 (rank : 56) | NC score | 0.152328 (rank : 36) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O95872 | Gene names | BAT4, G5 | |||
|
Domain Architecture |
|
|||||
| Description | Protein BAT4 (HLA-B-associated transcript 4) (G5 protein). | |||||
|
MERL_HUMAN
|
||||||
| θ value | 1.06291 (rank : 57) | NC score | 0.028702 (rank : 73) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 532 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P35240, O95683, Q8WUJ2, Q969N0, Q969Q3, Q96T30, Q96T31, Q96T32, Q96T33, Q9BTW3, Q9UNG9, Q9UNH3, Q9UNH4 | Gene names | NF2, SCH | |||
|
Domain Architecture |

|
|||||
| Description | Merlin (Moesin-ezrin-radixin-like protein) (Neurofibromin-2) (Schwannomin) (Schwannomerlin). | |||||
|
MERL_MOUSE
|
||||||
| θ value | 1.06291 (rank : 58) | NC score | 0.028687 (rank : 75) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P46662, Q8BR03 | Gene names | Nf2, Nf-2 | |||
|
Domain Architecture |

|
|||||
| Description | Merlin (Moesin-ezrin-radixin-like protein) (Neurofibromin-2) (Schwannomin). | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 1.06291 (rank : 59) | NC score | 0.032362 (rank : 66) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
SPF45_HUMAN
|
||||||
| θ value | 1.06291 (rank : 60) | NC score | 0.142993 (rank : 40) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96I25, Q96GY6 | Gene names | RBM17, SPF45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
HXD10_MOUSE
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.034157 (rank : 63) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P28359, Q91WU9 | Gene names | Hoxd10, Hox-4.5, Hoxd-10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D10 (Hox-4.5) (Hox-5.3). | |||||
|
ARD1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.025266 (rank : 82) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGX0, Q8C2B6, Q8CDA4, Q8CDA7 | Gene names | Trim23, Arfd1 | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein ARD-1 (ADP-ribosylation factor domain protein 1) (Tripartite motif-containing protein 23). | |||||
|
BICD1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.036657 (rank : 58) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 837 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BR07, O55206, Q8BQ18, Q8BRR8, Q8C4D3, Q8CAK6, Q8R2J4, Q8R2J5, Q8R2J6, Q91YP7 | Gene names | Bicd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 1. | |||||
|
PKCB1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.035132 (rank : 62) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
SETX_HUMAN
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.029280 (rank : 71) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
HXB2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.014668 (rank : 86) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P0C1T1 | Gene names | Hoxb2, Hox-2.8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein Hox-B2 (Hox-2.8). | |||||
|
RABE2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.037299 (rank : 55) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 598 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q91WG2, Q99KN3 | Gene names | Rabep2, Rabpt5b | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 2 (Rabaptin-5beta). | |||||
|
BAT4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 68) | NC score | 0.145086 (rank : 39) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61858, Q9R065 | Gene names | Bat4, Bat-4, G5 | |||
|
Domain Architecture |
|
|||||
| Description | Protein BAT4 (HLA-B-associated transcript 4) (G5 protein). | |||||
|
CCD13_HUMAN
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.038709 (rank : 53) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8IYE1 | Gene names | CCDC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 13. | |||||
|
NSD1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.023166 (rank : 83) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
RUFY2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.032988 (rank : 64) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
BICD1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.035838 (rank : 61) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96G01, O43892, O43893 | Gene names | BICD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 1. | |||||
|
IASPP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.014309 (rank : 87) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8WUF5, Q2PNZ9, Q5DU71, Q5I1X4, Q6P1R7, Q6PKF8, Q9Y290 | Gene names | PPP1R13L, IASPP, NKIP1, PPP1R13BL, RAI | |||
|
Domain Architecture |
|
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
MLKL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.006349 (rank : 95) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NB16, Q8N6V0 | Gene names | MLKL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mixed lineage kinase domain-like protein. | |||||
|
RAD50_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.050021 (rank : 48) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
REST_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.027442 (rank : 76) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
ANR18_HUMAN
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.026421 (rank : 79) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1106 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8IVF6, Q7Z468 | Gene names | ANKRD18A, KIAA2015 | |||
|
Domain Architecture |
|
|||||
| Description | Ankyrin repeat domain-containing protein 18A. | |||||
|
ANR26_HUMAN
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.026829 (rank : 78) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1614 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UPS8, Q2TAZ3, Q6ZR14, Q9H1Q1, Q9NSK9, Q9NTD5, Q9NW69 | Gene names | ANKRD26, KIAA1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
CEP35_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.029275 (rank : 72) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
CYTSA_MOUSE
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.026188 (rank : 80) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1104 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q2KN98, Q8CHF9 | Gene names | Cytsa, Kiaa0376 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytospin-A. | |||||
|
RNF17_MOUSE
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.019052 (rank : 84) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99MV7 | Gene names | Rnf17 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 17. | |||||
|
S6A16_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.006433 (rank : 94) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9GZN6, Q9Y5I9 | Gene names | SLC6A16, NTT5 | |||
|
Domain Architecture |

|
|||||
| Description | Orphan sodium- and chloride-dependent neurotransmitter transporter NTT5 (Solute carrier family 6 member 16). | |||||
|
SSNA1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | 0.038562 (rank : 54) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9JJ94 | Gene names | Ssna1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sjoegren syndrome nuclear autoantigen 1 homolog. | |||||
|
CTGE5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.031926 (rank : 68) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 928 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8R311, Q8CIE3 | Gene names | Ctage5, Mea6, Mgea6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cutaneous T-cell lymphoma-associated antigen 5 homolog (cTAGE-5 protein) (Meningioma-expressed antigen 6). | |||||
|
FCRLA_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.010022 (rank : 89) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q920A9, Q3T9Q2, Q8K3U9, Q8VHP5 | Gene names | Fcrla, Fcrl, Fcrl1, Fcrlm1, Fcrx, Freb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fc receptor-like and mucin-like 1 precursor (Fc receptor-like A) (Fc receptor homolog expressed in B-cells) (Fc receptor-related protein X) (FcRX) (Fc receptor-like protein). | |||||
|
OCRL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.009318 (rank : 90) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q01968, O60800, Q15684, Q15774, Q5JQF1, Q5JQF2, Q9UJG5, Q9UMA5 | Gene names | OCRL, INPP5F, OCRL1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol polyphosphate 5-phosphatase OCRL-1 (EC 3.1.3.36) (Lowe oculocerebrorenal syndrome protein). | |||||
|
SSNA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.036615 (rank : 60) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O43805, Q6FG70, Q9BVW8 | Gene names | SSNA1, NA14 | |||
|
Domain Architecture |
|
|||||
| Description | Sjoegren syndrome nuclear autoantigen 1 (Nuclear autoantigen of 14 kDa). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.029839 (rank : 70) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
DHX36_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.007731 (rank : 92) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H2U1, Q70JU3, Q8IYE5, Q9P240 | Gene names | DHX36, DDX36, KIAA1488, MLEL1, RHAU | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DHX36 (EC 3.6.1.-) (DEAH box protein 36) (MLE-like protein 1) (RNA helicase associated with AU-rich element ARE). | |||||
|
DHX36_MOUSE
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.007367 (rank : 93) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VHK9, Q6ZPP7, Q9CSE8 | Gene names | Dhx36, Ddx36, Kiaa1488, Mlel1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DHX36 (EC 3.6.1.-) (DEAH box protein 36) (MLE-like protein 1) (RNA helicase associated with AU-rich element ARE). | |||||
|
GOGA3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.028692 (rank : 74) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1437 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P55937, Q80VF5, Q8CCK4, Q9QYT2, Q9QYT3 | Gene names | Golga3, Mea2 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Male-enhanced antigen 2) (MEA-2). | |||||
|
NSD1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.017566 (rank : 85) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
SRCA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.008700 (rank : 91) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7TQ48, Q8BS43 | Gene names | Srl, Sar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
SVIL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.013824 (rank : 88) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K4L3 | Gene names | Svil | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
TNIK_HUMAN
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.001873 (rank : 96) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
RBM6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.088104 (rank : 43) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P78332, O60549, O75524 | Gene names | RBM6, DEF3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 6 (RNA-binding motif protein 6) (RNA-binding protein DEF-3) (Lung cancer antigen NY-LU-12) (Protein G16). | |||||
|
TFP11_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 95 | |
| SwissProt Accessions | Q9ERA6, Q8VD06 | Gene names | Tfip11, Tip39 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11 (Tuftelin-interacting protein 39). | |||||
|
TFP11_HUMAN
|
||||||
| NC score | 0.990661 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9UBB9, O95908, Q5H8V8, Q9UGV7, Q9Y2Q8 | Gene names | TFIP11 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11. | |||||
|
VPK7_HUMAN
|
||||||
| NC score | 0.398123 (rank : 3) | θ value | 6.43352e-06 (rank : 6) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63123 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q23.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K18 Pro protein) (HERV-K110 Pro protein) (HERV-K(C1a) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK9_HUMAN
|
||||||
| NC score | 0.394163 (rank : 4) | θ value | 2.44474e-05 (rank : 15) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P63124 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q13.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K104 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK1_HUMAN
|
||||||
| NC score | 0.393400 (rank : 5) | θ value | 2.44474e-05 (rank : 12) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P63119 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
VPK5_HUMAN
|
||||||
| NC score | 0.393073 (rank : 6) | θ value | 2.44474e-05 (rank : 14) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P63121 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K113 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
VPK10_HUMAN
|
||||||
| NC score | 0.392749 (rank : 7) | θ value | 2.44474e-05 (rank : 9) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P10265, Q9UKH6 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-K_5q33.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K10 Pro protein) (HERV-K107 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK12_HUMAN
|
||||||
| NC score | 0.392652 (rank : 8) | θ value | 2.44474e-05 (rank : 10) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P63131, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K102 Pro protein) (HERV-K(III) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK17_HUMAN
|
||||||
| NC score | 0.392142 (rank : 9) | θ value | 2.44474e-05 (rank : 11) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P63125 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_11q22.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
VPK2_HUMAN
|
||||||
| NC score | 0.390435 (rank : 10) | θ value | 2.44474e-05 (rank : 13) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y6I0, Q9UKH5 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K(HML-2.HOM) Pro protein) (HERV-K108 Pro protein) (HERV-K(C7) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK11_HUMAN
|
||||||
| NC score | 0.384385 (rank : 11) | θ value | 5.44631e-05 (rank : 16) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63129, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K101 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
ZGPAT_MOUSE
|
||||||
| NC score | 0.383948 (rank : 12) | θ value | 1.53129e-07 (rank : 3) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8VDM1, Q8BWW2 | Gene names | Zgpat | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein. | |||||
|
ZGPAT_HUMAN
|
||||||
| NC score | 0.381523 (rank : 13) | θ value | 2.61198e-07 (rank : 4) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8N5A5, Q5JWI9, Q8NC55, Q96JI0, Q96JU4, Q9H401 | Gene names | ZGPAT, KIAA1847, ZC3H9, ZC3HDC9 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein (Zinc finger CCCH domain-containing protein 9). | |||||
|
VPK3_HUMAN
|
||||||
| NC score | 0.378373 (rank : 14) | θ value | 0.00035302 (rank : 23) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63120 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19q12 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K(C19) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK6_HUMAN
|
||||||
| NC score | 0.369632 (rank : 15) | θ value | 0.000270298 (rank : 22) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63122 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K115 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK4_HUMAN
|
||||||
| NC score | 0.369453 (rank : 16) | θ value | 0.000270298 (rank : 21) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63127, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K109 Pro protein) (HERV-K(C6) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
RBM10_HUMAN
|
||||||
| NC score | 0.331201 (rank : 17) | θ value | 7.59969e-07 (rank : 5) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
GPTC2_HUMAN
|
||||||
| NC score | 0.323593 (rank : 18) | θ value | 0.00020696 (rank : 19) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NW75, Q86YE7 | Gene names | GPATC2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
GPTC2_MOUSE
|
||||||
| NC score | 0.319159 (rank : 19) | θ value | 0.000270298 (rank : 20) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q7TQC7, Q8BNJ9, Q8BPM1, Q8CDH9 | Gene names | Gpatc2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
PINX1_MOUSE
|
||||||
| NC score | 0.316287 (rank : 20) | θ value | 0.00665767 (rank : 26) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CZX5, Q91WZ9, Q9D0C2 | Gene names | Pinx1, Lpts | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (LPTS1) (67-11-3 protein). | |||||
|
RBM5_HUMAN
|
||||||
| NC score | 0.310886 (rank : 21) | θ value | 7.1131e-05 (rank : 17) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P52756, Q93021, Q9BU14, Q9UKY8, Q9UL24 | Gene names | RBM5 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15) (Protein G15) (NY-REN-9 antigen). | |||||
|
RBM5_MOUSE
|
||||||
| NC score | 0.310143 (rank : 22) | θ value | 7.1131e-05 (rank : 18) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q91YE7, Q3UZ19, Q99KV9 | Gene names | Rbm5, Luca15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15). | |||||
|
GPTC3_MOUSE
|
||||||
| NC score | 0.288813 (rank : 23) | θ value | 2.44474e-05 (rank : 8) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BIY1 | Gene names | Gpatc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
PINX1_HUMAN
|
||||||
| NC score | 0.282250 (rank : 24) | θ value | 0.0330416 (rank : 39) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96BK5, Q7Z7J8, Q96QD7, Q9HBU7, Q9NWW2 | Gene names | PINX1, LPTS | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (67-11-3 protein). | |||||
|
GPTC3_HUMAN
|
||||||
| NC score | 0.278256 (rank : 25) | θ value | 8.40245e-06 (rank : 7) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96I76, Q5JYH2, Q8NDJ2, Q9H9Z3 | Gene names | GPATC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
AGGF1_HUMAN
|
||||||
| NC score | 0.261807 (rank : 26) | θ value | 0.0252991 (rank : 35) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8N302, O00581, Q53YS3, Q9BU84, Q9NW66 | Gene names | AGGF1, VG5Q | |||
|
Domain Architecture |

|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (Vasculogenesis gene on 5q protein) (hVG5Q). | |||||
|
K0082_HUMAN
|
||||||
| NC score | 0.260606 (rank : 27) | θ value | 0.00509761 (rank : 25) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8N1G2, Q14670, Q96FJ9 | Gene names | KIAA0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
AGGF1_MOUSE
|
||||||
| NC score | 0.259963 (rank : 28) | θ value | 0.0330416 (rank : 38) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q7TN31, Q8R2S6, Q9CQR9, Q9CU87, Q9D768 | Gene names | Aggf1, Vg5q | |||
|
Domain Architecture |

|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (mVG5Q). | |||||
|
NKRF_HUMAN
|
||||||
| NC score | 0.252855 (rank : 29) | θ value | 0.0252991 (rank : 37) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O15226, Q9UJ91 | Gene names | NKRF, ITBA4, NRF | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF) (ITBA4 protein). | |||||
|
SF04_HUMAN
|
||||||
| NC score | 0.251723 (rank : 30) | θ value | 0.00390308 (rank : 24) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8IWZ8, O60378, Q6P3X9, Q8TCQ4, Q8WWT4, Q8WWT5, Q9NTG3 | Gene names | SF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4 (RNA-binding protein RBP). | |||||
|
K0082_MOUSE
|
||||||
| NC score | 0.250813 (rank : 31) | θ value | 0.0148317 (rank : 32) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9DBC3, Q3U3G5, Q3U7Y9, Q6A0D5, Q8C7V0 | Gene names | Kiaa0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
NKRF_MOUSE
|
||||||
| NC score | 0.250389 (rank : 32) | θ value | 0.0431538 (rank : 40) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8BY02, Q8BJU5, Q9CRL2 | Gene names | Nkrf, Nrf | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF). | |||||
|
SFR14_HUMAN
|
||||||
| NC score | 0.244820 (rank : 33) | θ value | 0.00665767 (rank : 28) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8IX01, O15071, O60369, Q5JPH7, Q8WUF7 | Gene names | SFRS14, KIAA0365 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
SF04_MOUSE
|
||||||
| NC score | 0.242833 (rank : 34) | θ value | 0.00665767 (rank : 27) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8CH02, Q8R094 | Gene names | Sf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4. | |||||
|
SFR14_MOUSE
|
||||||
| NC score | 0.235957 (rank : 35) | θ value | 0.00665767 (rank : 29) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
BAT4_HUMAN
|
||||||
| NC score | 0.152328 (rank : 36) | θ value | 1.06291 (rank : 56) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O95872 | Gene names | BAT4, G5 | |||
|
Domain Architecture |

|
|||||
| Description | Protein BAT4 (HLA-B-associated transcript 4) (G5 protein). | |||||
|
SON_MOUSE
|
||||||
| NC score | 0.150621 (rank : 37) | θ value | 0.0113563 (rank : 31) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
SPF45_MOUSE
|
||||||
| NC score | 0.147471 (rank : 38) | θ value | 0.62314 (rank : 52) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8JZX4, O75939 | Gene names | Rbm17, Spf45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
BAT4_MOUSE
|
||||||
| NC score | 0.145086 (rank : 39) | θ value | 3.0926 (rank : 68) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61858, Q9R065 | Gene names | Bat4, Bat-4, G5 | |||
|
Domain Architecture |

|
|||||
| Description | Protein BAT4 (HLA-B-associated transcript 4) (G5 protein). | |||||
|
SPF45_HUMAN
|
||||||
| NC score | 0.142993 (rank : 40) | θ value | 1.06291 (rank : 60) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96I25, Q96GY6 | Gene names | RBM17, SPF45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
TB182_HUMAN
|
||||||
| NC score | 0.103526 (rank : 41) | θ value | 0.0148317 (rank : 33) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C0C2 | Gene names | TNKS1BP1, KIAA1741, TAB182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 182 kDa tankyrase 1-binding protein. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.103334 (rank : 42) | θ value | 0.0113563 (rank : 30) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
RBM6_HUMAN
|
||||||
| NC score | 0.088104 (rank : 43) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P78332, O60549, O75524 | Gene names | RBM6, DEF3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 6 (RNA-binding motif protein 6) (RNA-binding protein DEF-3) (Lung cancer antigen NY-LU-12) (Protein G16). | |||||
|
VP13B_HUMAN
|
||||||
| NC score | 0.071812 (rank : 44) | θ value | 0.365318 (rank : 48) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z7G8, Q709C6, Q709C7, Q7Z7G4, Q7Z7G5, Q7Z7G6, Q7Z7G7, Q8NB77, Q9NWV1, Q9Y4E7 | Gene names | VPS13B, CHS1, COH1, KIAA0532 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vacuolar protein sorting-associated protein 13B (Cohen syndrome protein 1). | |||||
|
RAD50_MOUSE
|
||||||
| NC score | 0.060709 (rank : 45) | θ value | 0.0193708 (rank : 34) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
TSC1_HUMAN
|
||||||
| NC score | 0.056721 (rank : 46) | θ value | 0.21417 (rank : 46) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q92574 | Gene names | TSC1, KIAA0243, TSC | |||
|
Domain Architecture |

|
|||||
| Description | Hamartin (Tuberous sclerosis 1 protein). | |||||
|
GOGA1_HUMAN
|
||||||
| NC score | 0.051096 (rank : 47) | θ value | 0.0252991 (rank : 36) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
RAD50_HUMAN
|
||||||
| NC score | 0.050021 (rank : 48) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
HXC10_MOUSE
|
||||||
| NC score | 0.045752 (rank : 49) | θ value | 0.0736092 (rank : 41) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P31257, P31312, Q7TMT7 | Gene names | Hoxc10, Hox-3.6, Hoxc-10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein Hox-C10 (Hox-3.6). | |||||
|
RUFY2_MOUSE
|
||||||
| NC score | 0.044341 (rank : 50) | θ value | 0.21417 (rank : 45) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8R4C2, Q69ZH1 | Gene names | Rufy2, Kiaa1537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Leucine zipper FYVE-finger protein) (LZ-FYVE). | |||||
|
HXC10_HUMAN
|
||||||
| NC score | 0.043068 (rank : 51) | θ value | 0.163984 (rank : 43) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NYD6, O15219, O15220, Q9BVD5 | Gene names | HOXC10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-C10. | |||||
|
PEPL_MOUSE
|
||||||
| NC score | 0.038862 (rank : 52) | θ value | 0.21417 (rank : 44) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
CCD13_HUMAN
|
||||||
| NC score | 0.038709 (rank : 53) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8IYE1 | Gene names | CCDC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 13. | |||||
|
SSNA1_MOUSE
|
||||||
| NC score | 0.038562 (rank : 54) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9JJ94 | Gene names | Ssna1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sjoegren syndrome nuclear autoantigen 1 homolog. | |||||
|
RABE2_MOUSE
|
||||||
| NC score | 0.037299 (rank : 55) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 598 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q91WG2, Q99KN3 | Gene names | Rabep2, Rabpt5b | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 2 (Rabaptin-5beta). | |||||
|
KMHN1_MOUSE
|
||||||
| NC score | 0.036873 (rank : 56) | θ value | 0.813845 (rank : 53) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q3V125, Q3TTQ8 | Gene names | Gm172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1 homolog. | |||||
|
CP250_MOUSE
|
||||||
| NC score | 0.036738 (rank : 57) | θ value | 0.47712 (rank : 49) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
BICD1_MOUSE
|
||||||
| NC score | 0.036657 (rank : 58) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 837 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BR07, O55206, Q8BQ18, Q8BRR8, Q8C4D3, Q8CAK6, Q8R2J4, Q8R2J5, Q8R2J6, Q91YP7 | Gene names | Bicd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 1. | |||||
|
HXD10_HUMAN
|
||||||
| NC score | 0.036631 (rank : 59) | θ value | 0.62314 (rank : 51) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P28358 | Gene names | HOXD10, HOX4D | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D10 (Hox-4D) (Hox-4E). | |||||
|
SSNA1_HUMAN
|
||||||
| NC score | 0.036615 (rank : 60) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O43805, Q6FG70, Q9BVW8 | Gene names | SSNA1, NA14 | |||
|
Domain Architecture |

|
|||||
| Description | Sjoegren syndrome nuclear autoantigen 1 (Nuclear autoantigen of 14 kDa). | |||||
|
BICD1_HUMAN
|
||||||
| NC score | 0.035838 (rank : 61) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96G01, O43892, O43893 | Gene names | BICD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 1. | |||||
|
PKCB1_HUMAN
|
||||||
| NC score | 0.035132 (rank : 62) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
HXD10_MOUSE
|
||||||
| NC score | 0.034157 (rank : 63) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P28359, Q91WU9 | Gene names | Hoxd10, Hox-4.5, Hoxd-10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D10 (Hox-4.5) (Hox-5.3). | |||||
|
RUFY2_HUMAN
|
||||||
| NC score | 0.032988 (rank : 64) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
MYH10_HUMAN
|
||||||
| NC score | 0.032467 (rank : 65) | θ value | 0.279714 (rank : 47) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.032362 (rank : 66) | θ value | 1.06291 (rank : 59) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | 0.031965 (rank : 67) | θ value | 0.813845 (rank : 55) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
CTGE5_MOUSE
|
||||||
| NC score | 0.031926 (rank : 68) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 928 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8R311, Q8CIE3 | Gene names | Ctage5, Mea6, Mgea6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cutaneous T-cell lymphoma-associated antigen 5 homolog (cTAGE-5 protein) (Meningioma-expressed antigen 6). | |||||
|
MYH10_MOUSE
|
||||||
| NC score | 0.031408 (rank : 69) | θ value | 0.813845 (rank : 54) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.029839 (rank : 70) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
SETX_HUMAN
|
||||||
| NC score | 0.029280 (rank : 71) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
CEP35_HUMAN
|
||||||
| NC score | 0.029275 (rank : 72) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
MERL_HUMAN
|
||||||
| NC score | 0.028702 (rank : 73) | θ value | 1.06291 (rank : 57) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 532 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P35240, O95683, Q8WUJ2, Q969N0, Q969Q3, Q96T30, Q96T31, Q96T32, Q96T33, Q9BTW3, Q9UNG9, Q9UNH3, Q9UNH4 | Gene names | NF2, SCH | |||
|
Domain Architecture |

|
|||||
| Description | Merlin (Moesin-ezrin-radixin-like protein) (Neurofibromin-2) (Schwannomin) (Schwannomerlin). | |||||
|
GOGA3_MOUSE
|
||||||
| NC score | 0.028692 (rank : 74) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1437 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P55937, Q80VF5, Q8CCK4, Q9QYT2, Q9QYT3 | Gene names | Golga3, Mea2 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Male-enhanced antigen 2) (MEA-2). | |||||
|
MERL_MOUSE
|
||||||
| NC score | 0.028687 (rank : 75) | θ value | 1.06291 (rank : 58) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P46662, Q8BR03 | Gene names | Nf2, Nf-2 | |||
|
Domain Architecture |

|
|||||
| Description | Merlin (Moesin-ezrin-radixin-like protein) (Neurofibromin-2) (Schwannomin). | |||||
|
REST_HUMAN
|
||||||
| NC score | 0.027442 (rank : 76) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
ARD1_HUMAN
|
||||||
| NC score | 0.026987 (rank : 77) | θ value | 0.62314 (rank : 50) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 621 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P36406, Q9BZY4, Q9BZY5 | Gene names | TRIM23, ARD1, ARFD1, RNF46 | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein ARD-1 (ADP-ribosylation factor domain protein 1) (Tripartite motif-containing protein 23) (RING finger protein 46). | |||||
|
ANR26_HUMAN
|
||||||
| NC score | 0.026829 (rank : 78) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1614 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UPS8, Q2TAZ3, Q6ZR14, Q9H1Q1, Q9NSK9, Q9NTD5, Q9NW69 | Gene names | ANKRD26, KIAA1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
ANR18_HUMAN
|
||||||
| NC score | 0.026421 (rank : 79) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1106 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8IVF6, Q7Z468 | Gene names | ANKRD18A, KIAA2015 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat domain-containing protein 18A. | |||||
|
CYTSA_MOUSE
|
||||||
| NC score | 0.026188 (rank : 80) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1104 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q2KN98, Q8CHF9 | Gene names | Cytsa, Kiaa0376 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytospin-A. | |||||
|
ANR50_HUMAN
|
||||||
| NC score | 0.025720 (rank : 81) | θ value | 0.0961366 (rank : 42) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULJ7, Q6N064, Q6ZSE6 | Gene names | ANKRD50, KIAA1223 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 50. | |||||
|
ARD1_MOUSE
|
||||||
| NC score | 0.025266 (rank : 82) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGX0, Q8C2B6, Q8CDA4, Q8CDA7 | Gene names | Trim23, Arfd1 | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein ARD-1 (ADP-ribosylation factor domain protein 1) (Tripartite motif-containing protein 23). | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.023166 (rank : 83) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
RNF17_MOUSE
|
||||||
| NC score | 0.019052 (rank : 84) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99MV7 | Gene names | Rnf17 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 17. | |||||
|
NSD1_MOUSE
|
||||||
| NC score | 0.017566 (rank : 85) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
HXB2_MOUSE
|
||||||
| NC score | 0.014668 (rank : 86) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P0C1T1 | Gene names | Hoxb2, Hox-2.8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein Hox-B2 (Hox-2.8). | |||||
|
IASPP_HUMAN
|
||||||
| NC score | 0.014309 (rank : 87) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8WUF5, Q2PNZ9, Q5DU71, Q5I1X4, Q6P1R7, Q6PKF8, Q9Y290 | Gene names | PPP1R13L, IASPP, NKIP1, PPP1R13BL, RAI | |||
|
Domain Architecture |

|
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
SVIL_MOUSE
|
||||||
| NC score | 0.013824 (rank : 88) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K4L3 | Gene names | Svil | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
FCRLA_MOUSE
|
||||||
| NC score | 0.010022 (rank : 89) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q920A9, Q3T9Q2, Q8K3U9, Q8VHP5 | Gene names | Fcrla, Fcrl, Fcrl1, Fcrlm1, Fcrx, Freb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fc receptor-like and mucin-like 1 precursor (Fc receptor-like A) (Fc receptor homolog expressed in B-cells) (Fc receptor-related protein X) (FcRX) (Fc receptor-like protein). | |||||
|
OCRL_HUMAN
|
||||||
| NC score | 0.009318 (rank : 90) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q01968, O60800, Q15684, Q15774, Q5JQF1, Q5JQF2, Q9UJG5, Q9UMA5 | Gene names | OCRL, INPP5F, OCRL1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol polyphosphate 5-phosphatase OCRL-1 (EC 3.1.3.36) (Lowe oculocerebrorenal syndrome protein). | |||||
|
SRCA_MOUSE
|
||||||
| NC score | 0.008700 (rank : 91) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7TQ48, Q8BS43 | Gene names | Srl, Sar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
DHX36_HUMAN
|
||||||
| NC score | 0.007731 (rank : 92) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H2U1, Q70JU3, Q8IYE5, Q9P240 | Gene names | DHX36, DDX36, KIAA1488, MLEL1, RHAU | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DHX36 (EC 3.6.1.-) (DEAH box protein 36) (MLE-like protein 1) (RNA helicase associated with AU-rich element ARE). | |||||
|
DHX36_MOUSE
|
||||||
| NC score | 0.007367 (rank : 93) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VHK9, Q6ZPP7, Q9CSE8 | Gene names | Dhx36, Ddx36, Kiaa1488, Mlel1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DHX36 (EC 3.6.1.-) (DEAH box protein 36) (MLE-like protein 1) (RNA helicase associated with AU-rich element ARE). | |||||
|
S6A16_HUMAN
|
||||||
| NC score | 0.006433 (rank : 94) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9GZN6, Q9Y5I9 | Gene names | SLC6A16, NTT5 | |||
|
Domain Architecture |

|
|||||
| Description | Orphan sodium- and chloride-dependent neurotransmitter transporter NTT5 (Solute carrier family 6 member 16). | |||||
|
MLKL_HUMAN
|
||||||
| NC score | 0.006349 (rank : 95) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NB16, Q8N6V0 | Gene names | MLKL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mixed lineage kinase domain-like protein. | |||||
|
TNIK_HUMAN
|
||||||
| NC score | 0.001873 (rank : 96) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 95 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||