Please be patient as the page loads
|
MIER1_MOUSE
|
||||||
| SwissProt Accessions | Q5UAK0, Q5UAK1, Q5UAK2, Q5UAK3, Q6ZPL6, Q9D402 | Gene names | Mier1, Kiaa1610 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mesoderm induction early response protein 1 (Mi-er1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MIER1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.991988 (rank : 2) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8N108, Q5T104, Q6AHY8, Q86TB4, Q8N156, Q8N161, Q8NC37, Q8NES4, Q8NES5, Q8NES6, Q8WWG2, Q9HCG2 | Gene names | MIER1, KIAA1610 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mesoderm induction early response protein 1 (Mi-er1) (hmi-er1) (Early response 1) (Er1). | |||||
|
MIER1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q5UAK0, Q5UAK1, Q5UAK2, Q5UAK3, Q6ZPL6, Q9D402 | Gene names | Mier1, Kiaa1610 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mesoderm induction early response protein 1 (Mi-er1). | |||||
|
MTA2_MOUSE
|
||||||
| θ value | 1.9326e-10 (rank : 3) | NC score | 0.514240 (rank : 3) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9R190, Q3TVT8 | Gene names | Mta2, Mta1l1 | |||
|
Domain Architecture |
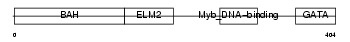
|
|||||
| Description | Metastasis-associated protein MTA2 (Metastasis-associated 1-like 1). | |||||
|
MTA2_HUMAN
|
||||||
| θ value | 5.62301e-10 (rank : 4) | NC score | 0.512251 (rank : 4) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O94776, Q9UQB5 | Gene names | MTA2, MTA1L1, PID | |||
|
Domain Architecture |
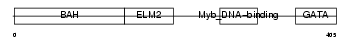
|
|||||
| Description | Metastasis-associated protein MTA2 (Metastasis-associated 1-like 1) (MTA1-L1 protein) (p53 target protein in deacetylase complex). | |||||
|
MTA3_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 5) | NC score | 0.506067 (rank : 5) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BTC8, Q9NSP2, Q9ULF4 | Gene names | MTA3, KIAA1266 | |||
|
Domain Architecture |
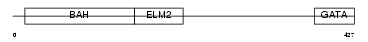
|
|||||
| Description | Metastasis-associated protein MTA3. | |||||
|
RERE_MOUSE
|
||||||
| θ value | 8.11959e-09 (rank : 6) | NC score | 0.381393 (rank : 16) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
RERE_HUMAN
|
||||||
| θ value | 4.0297e-08 (rank : 7) | NC score | 0.380345 (rank : 17) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
MTA3_MOUSE
|
||||||
| θ value | 1.53129e-07 (rank : 8) | NC score | 0.493271 (rank : 6) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q924K8, Q8VC18 | Gene names | Mta3 | |||
|
Domain Architecture |
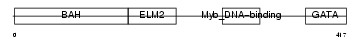
|
|||||
| Description | Metastasis-associated protein MTA3. | |||||
|
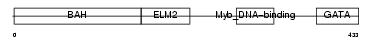
MTA1_HUMAN
|
||||||
| θ value | 5.81887e-07 (rank : 9) | NC score | 0.483346 (rank : 7) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q13330, Q8NFI8, Q96GI8 | Gene names | MTA1 | |||
|
Domain Architecture |
|
|||||
| Description | Metastasis-associated protein MTA1. | |||||
|
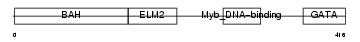
MTA1_MOUSE
|
||||||
| θ value | 5.81887e-07 (rank : 10) | NC score | 0.482264 (rank : 8) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K4B0, Q80UI1, Q8K4D4, Q924K9 | Gene names | Mta1 | |||
|
Domain Architecture |
|
|||||
| Description | Metastasis-associated protein MTA1. | |||||
|
RCOR3_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 11) | NC score | 0.454966 (rank : 11) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6PGA0 | Gene names | Rcor3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
RCOR3_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 12) | NC score | 0.454024 (rank : 12) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
ZN541_HUMAN
|
||||||
| θ value | 3.77169e-06 (rank : 13) | NC score | 0.425608 (rank : 15) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9H0D2, Q8NDK8 | Gene names | ZNF541 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 541. | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 14) | NC score | 0.305185 (rank : 19) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
RCOR1_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 15) | NC score | 0.465248 (rank : 9) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CFE3, Q3V092, Q8BK28, Q8CHI2 | Gene names | Rcor1, D12Wsu95e, Kiaa0071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 1 (Protein CoREST). | |||||
|
RCOR1_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 16) | NC score | 0.462567 (rank : 10) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UKL0, Q15044, Q6P2I9, Q86VG5 | Gene names | RCOR1, KIAA0071, RCOR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 1 (Protein CoREST). | |||||
|
TREF1_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 17) | NC score | 0.307190 (rank : 18) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96PN7, Q7Z6T2, Q7Z6T3, Q9NQ72, Q9NQ73, Q9NUN9 | Gene names | TRERF1, RAPA, TREP132 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional-regulating factor 1 (Transcriptional-regulating protein 132) (Zinc finger transcription factor TReP-132) (Zinc finger protein rapa). | |||||
|
NCOR1_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 18) | NC score | 0.298783 (rank : 21) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
NCOR1_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 19) | NC score | 0.302504 (rank : 20) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 20) | NC score | 0.287708 (rank : 23) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
TREF1_MOUSE
|
||||||
| θ value | 0.000121331 (rank : 21) | NC score | 0.297971 (rank : 22) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BXJ2, Q6PCQ4, Q80X25, Q810H8, Q8BY31 | Gene names | Trerf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional-regulating factor 1 (Transcriptional-regulating protein 132) (Zinc finger transcription factor TReP-132). | |||||
|
RCOR2_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 22) | NC score | 0.430956 (rank : 13) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IZ40, Q96FP3 | Gene names | RCOR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2. | |||||
|
RCOR2_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 23) | NC score | 0.430300 (rank : 14) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8C796, Q9JMK4 | Gene names | Rcor2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2 (M-CoREST). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 24) | NC score | 0.116576 (rank : 26) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
SEC63_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 25) | NC score | 0.140281 (rank : 24) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 26) | NC score | 0.133189 (rank : 25) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 27) | NC score | 0.083272 (rank : 35) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 28) | NC score | 0.105757 (rank : 30) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
SNUT1_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 29) | NC score | 0.115481 (rank : 27) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z315, Q8K155, Q9R1I9, Q9R270 | Gene names | Sart1, Haf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (Squamous cell carcinoma antigen recognized by T-cells 1) (SART-1) (mSART-1) (Hypoxia- associated factor). | |||||
|
EP400_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 30) | NC score | 0.050042 (rank : 54) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
RAG2_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 31) | NC score | 0.108910 (rank : 28) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P21784 | Gene names | Rag2, Rag-2 | |||
|
Domain Architecture |

|
|||||
| Description | V(D)J recombination-activating protein 2 (RAG-2). | |||||
|
SNUT1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 32) | NC score | 0.108882 (rank : 29) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43290, Q53GB5 | Gene names | SART1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (U4/U6.U5 tri-snRNP-associated 110 kDa protein) (Squamous cell carcinoma antigen recognized by T cells 1) (SART-1) (hSART-1) (hSnu66). | |||||
|
SMRC2_MOUSE
|
||||||
| θ value | 0.21417 (rank : 33) | NC score | 0.078407 (rank : 39) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
SMRC2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 34) | NC score | 0.084767 (rank : 33) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
EDEM3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 35) | NC score | 0.048209 (rank : 55) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BZQ6, Q5TEZ0, Q9HCW1, Q9UFV7 | Gene names | EDEM3, C1orf22 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 3. | |||||
|
KI67_HUMAN
|
||||||
| θ value | 0.47712 (rank : 36) | NC score | 0.067804 (rank : 41) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
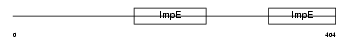
IFIT1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 37) | NC score | 0.050951 (rank : 51) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P09914, Q96QM5 | Gene names | IFIT1, G10P1, IFI56 | |||
|
Domain Architecture |
|
|||||
| Description | Interferon-induced protein with tetratricopeptide repeats 1 (IFIT-1) (Interferon-induced 56 kDa protein) (IFI-56K). | |||||
|
GNL3_MOUSE
|
||||||
| θ value | 0.813845 (rank : 38) | NC score | 0.057134 (rank : 47) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CI11, Q811R9, Q811S8, Q8BK21 | Gene names | Gnl3, Ns | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein-like 3 (Nucleolar GTP-binding protein 3) (Nucleostemin). | |||||
|
RAG2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 39) | NC score | 0.088200 (rank : 32) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P55895, Q8TBL4 | Gene names | RAG2 | |||
|
Domain Architecture |

|
|||||
| Description | V(D)J recombination-activating protein 2 (RAG-2). | |||||
|
SMRC1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 40) | NC score | 0.080553 (rank : 37) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P97496, Q7TS80, Q7TT29 | Gene names | Smarcc1, Baf155, Srg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155) (SWI3-related protein). | |||||
|
UBP33_HUMAN
|
||||||
| θ value | 1.38821 (rank : 41) | NC score | 0.029289 (rank : 67) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TEY7, Q8TEY6, Q96AV6, Q9H9F0, Q9UPQ5 | Gene names | USP33, KIAA1097, VDU1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 33 (EC 3.1.2.15) (Ubiquitin thioesterase 33) (Ubiquitin-specific-processing protease 33) (Deubiquitinating enzyme 33) (VHL-interacting deubiquitinating enzyme 1). | |||||
|
JOSD3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 42) | NC score | 0.063424 (rank : 42) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H5J8, Q6I9Y6 | Gene names | JOSD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein JOSD3. | |||||
|
TGON2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 43) | NC score | 0.053678 (rank : 49) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
DIAP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.015237 (rank : 79) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60610, Q9UC76 | Gene names | DIAPH1, DIAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1). | |||||
|
K0146_MOUSE
|
||||||
| θ value | 2.36792 (rank : 45) | NC score | 0.029678 (rank : 66) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BGX7, Q3TCD0, Q5U457, Q6A0B6, Q6IS60, Q811E1, Q8BWD6, Q8R305 | Gene names | Kiaa0146 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0146. | |||||
|
UBP8_HUMAN
|
||||||
| θ value | 2.36792 (rank : 46) | NC score | 0.028661 (rank : 68) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
ARX_HUMAN
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.012830 (rank : 83) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96QS3 | Gene names | ARX | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein ARX (Aristaless-related homeobox). | |||||
|
CENPJ_HUMAN
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.036784 (rank : 57) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9HC77, Q5T6R5, Q96KS5, Q9C067 | Gene names | CENPJ, CPAP, LAP, LIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein J (CENP-J) (Centrosomal P4.1-associated protein) (LAG-3-associated protein) (LYST-interacting protein 1). | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 3.0926 (rank : 49) | NC score | 0.060234 (rank : 45) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
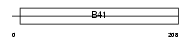
E41LA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 50) | NC score | 0.010541 (rank : 87) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HCS5 | Gene names | EPB41L4A, EPB41L4 | |||
|
Domain Architecture |
|
|||||
| Description | Band 4.1-like protein 4A (Protein NBL4). | |||||
|
GRIN1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 51) | NC score | 0.027444 (rank : 70) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
TNR6A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.033746 (rank : 59) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
ARX_MOUSE
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.012744 (rank : 84) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O35085, Q9QYT4 | Gene names | Arx | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein ARX (Aristaless-related homeobox). | |||||
|
C8AP2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.032904 (rank : 60) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
EP400_HUMAN
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.030092 (rank : 64) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
LUZP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.009759 (rank : 89) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
NOP14_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.035699 (rank : 58) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
RPGR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.022033 (rank : 72) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
WEE1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.002118 (rank : 95) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 794 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P47810, Q9EPL7 | Gene names | Wee1 | |||
|
Domain Architecture |

|
|||||
| Description | Wee1-like protein kinase (EC 2.7.10.2) (Wee1A kinase). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.058083 (rank : 46) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
MYO10_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.009365 (rank : 90) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9HD67, Q9NYM7, Q9P110, Q9P111, Q9UHF6 | Gene names | MYO10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin X). | |||||
|
MYRIP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.016374 (rank : 77) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K3I4, Q8CFC0, Q8K4H5 | Gene names | Myrip, Slac2c | |||
|
Domain Architecture |
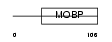
|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
NKTR_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.030260 (rank : 63) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
RBBP5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.018277 (rank : 75) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15291 | Gene names | RBBP5, RBQ3 | |||
|
Domain Architecture |
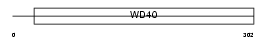
|
|||||
| Description | Retinoblastoma-binding protein 5 (RBBP-5) (Retinoblastoma-binding protein RBQ-3). | |||||
|
TIAM1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.014910 (rank : 80) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60610 | Gene names | Tiam1, Tiam-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
TXLNB_MOUSE
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.018752 (rank : 74) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
BPTF_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.023475 (rank : 71) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
DIAP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.012698 (rank : 85) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 839 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O08808 | Gene names | Diaph1, Diap1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1) (mDIA1) (p140mDIA). | |||||
|
MEFV_MOUSE
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.009017 (rank : 91) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JJ26 | Gene names | Mefv | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
MN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.031909 (rank : 61) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q10571 | Gene names | MN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable tumor suppressor protein MN1. | |||||
|
NEST_HUMAN
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.016399 (rank : 76) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
SMC4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.006112 (rank : 94) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 830 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CG47, Q8BTS7, Q8BTY9, Q99K21 | Gene names | Smc4l1, Capc, Smc4 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (XCAP-C homolog). | |||||
|
ATBF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.006341 (rank : 93) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
CCD47_HUMAN
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.019240 (rank : 73) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96A33, Q96D00, Q96JZ7, Q9H3E4, Q9NRG3 | Gene names | CCDC47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor. | |||||
|
CK5P2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.011221 (rank : 86) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
CLSPN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.030924 (rank : 62) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9HAW4, Q5VYG0, Q6P6H5, Q8IWI1 | Gene names | CLSPN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin (hClaspin) (Hu-Claspin). | |||||
|
CX039_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.014245 (rank : 81) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K2D0, Q3U3R2 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein CXorf39 homolog. | |||||
|
DNM3B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.008519 (rank : 92) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88509, O88510, O88511 | Gene names | Dnmt3b | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase MmuIIIB) (DNA MTase MmuIIIB) (M.MmuIIIB). | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.039332 (rank : 56) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
GP158_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.015811 (rank : 78) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5T848, Q6QR81, Q9ULT3 | Gene names | GPR158, KIAA1136 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 158 precursor. | |||||
|
IF2P_HUMAN
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.027446 (rank : 69) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
IF4B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.029790 (rank : 65) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P23588 | Gene names | EIF4B | |||
|
Domain Architecture |
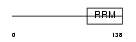
|
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
TIAM1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.013353 (rank : 82) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13009 | Gene names | TIAM1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
WRN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.010026 (rank : 88) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O09053, O09050, Q9JKD4, Q9Z241, Q9Z242 | Gene names | Wrn | |||
|
Domain Architecture |

|
|||||
| Description | Werner syndrome ATP-dependent helicase homolog (EC 3.6.1.-). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.095169 (rank : 31) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.055482 (rank : 48) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.050596 (rank : 53) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MYSM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.084143 (rank : 34) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5VVJ2, Q7Z3G8, Q96PX3 | Gene names | MYSM1, KIAA1915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MYSM1 (Myb-like, SWIRM and MPN domain-containing protein 1). | |||||
|
MYSM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.082798 (rank : 36) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q69Z66, Q3TPV7, Q8BRP5, Q8C4N1 | Gene names | Mysm1, Kiaa1915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MYSM1 (Myb-like, SWIRM and MPN domain-containing protein 1). | |||||
|
NUCKS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.050940 (rank : 52) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
PTMA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.078896 (rank : 38) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
PTMA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.072256 (rank : 40) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
PTMS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.052328 (rank : 50) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
PTMS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.061446 (rank : 43) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
SMRC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.060830 (rank : 44) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92922, Q6P172, Q8IWH2 | Gene names | SMARCC1, BAF155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155). | |||||
|
MIER1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q5UAK0, Q5UAK1, Q5UAK2, Q5UAK3, Q6ZPL6, Q9D402 | Gene names | Mier1, Kiaa1610 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mesoderm induction early response protein 1 (Mi-er1). | |||||
|
MIER1_HUMAN
|
||||||
| NC score | 0.991988 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8N108, Q5T104, Q6AHY8, Q86TB4, Q8N156, Q8N161, Q8NC37, Q8NES4, Q8NES5, Q8NES6, Q8WWG2, Q9HCG2 | Gene names | MIER1, KIAA1610 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mesoderm induction early response protein 1 (Mi-er1) (hmi-er1) (Early response 1) (Er1). | |||||
|
MTA2_MOUSE
|
||||||
| NC score | 0.514240 (rank : 3) | θ value | 1.9326e-10 (rank : 3) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9R190, Q3TVT8 | Gene names | Mta2, Mta1l1 | |||
|
Domain Architecture |

|
|||||
| Description | Metastasis-associated protein MTA2 (Metastasis-associated 1-like 1). | |||||
|
MTA2_HUMAN
|
||||||
| NC score | 0.512251 (rank : 4) | θ value | 5.62301e-10 (rank : 4) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O94776, Q9UQB5 | Gene names | MTA2, MTA1L1, PID | |||
|
Domain Architecture |

|
|||||
| Description | Metastasis-associated protein MTA2 (Metastasis-associated 1-like 1) (MTA1-L1 protein) (p53 target protein in deacetylase complex). | |||||
|
MTA3_HUMAN
|
||||||
| NC score | 0.506067 (rank : 5) | θ value | 2.79066e-09 (rank : 5) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BTC8, Q9NSP2, Q9ULF4 | Gene names | MTA3, KIAA1266 | |||
|
Domain Architecture |

|
|||||
| Description | Metastasis-associated protein MTA3. | |||||
|
MTA3_MOUSE
|
||||||
| NC score | 0.493271 (rank : 6) | θ value | 1.53129e-07 (rank : 8) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q924K8, Q8VC18 | Gene names | Mta3 | |||
|
Domain Architecture |

|
|||||
| Description | Metastasis-associated protein MTA3. | |||||
|
MTA1_HUMAN
|
||||||
| NC score | 0.483346 (rank : 7) | θ value | 5.81887e-07 (rank : 9) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q13330, Q8NFI8, Q96GI8 | Gene names | MTA1 | |||
|
Domain Architecture |

|
|||||
| Description | Metastasis-associated protein MTA1. | |||||
|
MTA1_MOUSE
|
||||||
| NC score | 0.482264 (rank : 8) | θ value | 5.81887e-07 (rank : 10) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K4B0, Q80UI1, Q8K4D4, Q924K9 | Gene names | Mta1 | |||
|
Domain Architecture |

|
|||||
| Description | Metastasis-associated protein MTA1. | |||||
|
RCOR1_MOUSE
|
||||||
| NC score | 0.465248 (rank : 9) | θ value | 1.09739e-05 (rank : 15) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CFE3, Q3V092, Q8BK28, Q8CHI2 | Gene names | Rcor1, D12Wsu95e, Kiaa0071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 1 (Protein CoREST). | |||||
|
RCOR1_HUMAN
|
||||||
| NC score | 0.462567 (rank : 10) | θ value | 2.44474e-05 (rank : 16) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UKL0, Q15044, Q6P2I9, Q86VG5 | Gene names | RCOR1, KIAA0071, RCOR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 1 (Protein CoREST). | |||||
|
RCOR3_MOUSE
|
||||||
| NC score | 0.454966 (rank : 11) | θ value | 2.21117e-06 (rank : 11) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6PGA0 | Gene names | Rcor3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
RCOR3_HUMAN
|
||||||
| NC score | 0.454024 (rank : 12) | θ value | 2.88788e-06 (rank : 12) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
RCOR2_HUMAN
|
||||||
| NC score | 0.430956 (rank : 13) | θ value | 0.00035302 (rank : 22) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IZ40, Q96FP3 | Gene names | RCOR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2. | |||||
|
RCOR2_MOUSE
|
||||||
| NC score | 0.430300 (rank : 14) | θ value | 0.00035302 (rank : 23) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8C796, Q9JMK4 | Gene names | Rcor2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2 (M-CoREST). | |||||
|
ZN541_HUMAN
|
||||||
| NC score | 0.425608 (rank : 15) | θ value | 3.77169e-06 (rank : 13) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9H0D2, Q8NDK8 | Gene names | ZNF541 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 541. | |||||
|
RERE_MOUSE
|
||||||
| NC score | 0.381393 (rank : 16) | θ value | 8.11959e-09 (rank : 6) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.380345 (rank : 17) | θ value | 4.0297e-08 (rank : 7) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
TREF1_HUMAN
|
||||||
| NC score | 0.307190 (rank : 18) | θ value | 4.1701e-05 (rank : 17) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96PN7, Q7Z6T2, Q7Z6T3, Q9NQ72, Q9NQ73, Q9NUN9 | Gene names | TRERF1, RAPA, TREP132 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional-regulating factor 1 (Transcriptional-regulating protein 132) (Zinc finger transcription factor TReP-132) (Zinc finger protein rapa). | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.305185 (rank : 19) | θ value | 8.40245e-06 (rank : 14) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
NCOR1_MOUSE
|
||||||
| NC score | 0.302504 (rank : 20) | θ value | 7.1131e-05 (rank : 19) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
NCOR1_HUMAN
|
||||||
| NC score | 0.298783 (rank : 21) | θ value | 7.1131e-05 (rank : 18) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
TREF1_MOUSE
|
||||||
| NC score | 0.297971 (rank : 22) | θ value | 0.000121331 (rank : 21) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BXJ2, Q6PCQ4, Q80X25, Q810H8, Q8BY31 | Gene names | Trerf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional-regulating factor 1 (Transcriptional-regulating protein 132) (Zinc finger transcription factor TReP-132). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.287708 (rank : 23) | θ value | 9.29e-05 (rank : 20) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.140281 (rank : 24) | θ value | 0.00175202 (rank : 25) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.133189 (rank : 25) | θ value | 0.00390308 (rank : 26) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.116576 (rank : 26) | θ value | 0.00134147 (rank : 24) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
SNUT1_MOUSE
|
||||||
| NC score | 0.115481 (rank : 27) | θ value | 0.0431538 (rank : 29) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z315, Q8K155, Q9R1I9, Q9R270 | Gene names | Sart1, Haf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (Squamous cell carcinoma antigen recognized by T-cells 1) (SART-1) (mSART-1) (Hypoxia- associated factor). | |||||
|
RAG2_MOUSE
|
||||||
| NC score | 0.108910 (rank : 28) | θ value | 0.0961366 (rank : 31) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P21784 | Gene names | Rag2, Rag-2 | |||
|
Domain Architecture |

|
|||||
| Description | V(D)J recombination-activating protein 2 (RAG-2). | |||||
|
SNUT1_HUMAN
|
||||||
| NC score | 0.108882 (rank : 29) | θ value | 0.0961366 (rank : 32) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43290, Q53GB5 | Gene names | SART1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (U4/U6.U5 tri-snRNP-associated 110 kDa protein) (Squamous cell carcinoma antigen recognized by T cells 1) (SART-1) (hSART-1) (hSnu66). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.105757 (rank : 30) | θ value | 0.0148317 (rank : 28) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.095169 (rank : 31) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
RAG2_HUMAN
|
||||||
| NC score | 0.088200 (rank : 32) | θ value | 0.813845 (rank : 39) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P55895, Q8TBL4 | Gene names | RAG2 | |||
|
Domain Architecture |

|
|||||
| Description | V(D)J recombination-activating protein 2 (RAG-2). | |||||
|
SMRC2_HUMAN
|
||||||
| NC score | 0.084767 (rank : 33) | θ value | 0.279714 (rank : 34) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
MYSM1_HUMAN
|
||||||
| NC score | 0.084143 (rank : 34) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5VVJ2, Q7Z3G8, Q96PX3 | Gene names | MYSM1, KIAA1915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MYSM1 (Myb-like, SWIRM and MPN domain-containing protein 1). | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.083272 (rank : 35) | θ value | 0.0113563 (rank : 27) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
MYSM1_MOUSE
|
||||||
| NC score | 0.082798 (rank : 36) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q69Z66, Q3TPV7, Q8BRP5, Q8C4N1 | Gene names | Mysm1, Kiaa1915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MYSM1 (Myb-like, SWIRM and MPN domain-containing protein 1). | |||||
|
SMRC1_MOUSE
|
||||||
| NC score | 0.080553 (rank : 37) | θ value | 1.38821 (rank : 40) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P97496, Q7TS80, Q7TT29 | Gene names | Smarcc1, Baf155, Srg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155) (SWI3-related protein). | |||||
|
PTMA_HUMAN
|
||||||
| NC score | 0.078896 (rank : 38) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
SMRC2_MOUSE
|
||||||
| NC score | 0.078407 (rank : 39) | θ value | 0.21417 (rank : 33) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
PTMA_MOUSE
|
||||||
| NC score | 0.072256 (rank : 40) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.067804 (rank : 41) | θ value | 0.47712 (rank : 36) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
JOSD3_HUMAN
|
||||||
| NC score | 0.063424 (rank : 42) | θ value | 1.81305 (rank : 42) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H5J8, Q6I9Y6 | Gene names | JOSD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein JOSD3. | |||||
|
PTMS_MOUSE
|
||||||
| NC score | 0.061446 (rank : 43) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
SMRC1_HUMAN
|
||||||
| NC score | 0.060830 (rank : 44) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92922, Q6P172, Q8IWH2 | Gene names | SMARCC1, BAF155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155). | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.060234 (rank : 45) | θ value | 3.0926 (rank : 49) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.058083 (rank : 46) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
GNL3_MOUSE
|
||||||
| NC score | 0.057134 (rank : 47) | θ value | 0.813845 (rank : 38) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CI11, Q811R9, Q811S8, Q8BK21 | Gene names | Gnl3, Ns | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein-like 3 (Nucleolar GTP-binding protein 3) (Nucleostemin). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.055482 (rank : 48) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.053678 (rank : 49) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
PTMS_HUMAN
|
||||||
| NC score | 0.052328 (rank : 50) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
IFIT1_HUMAN
|
||||||
| NC score | 0.050951 (rank : 51) | θ value | 0.62314 (rank : 37) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P09914, Q96QM5 | Gene names | IFIT1, G10P1, IFI56 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced protein with tetratricopeptide repeats 1 (IFIT-1) (Interferon-induced 56 kDa protein) (IFI-56K). | |||||
|
NUCKS_MOUSE
|
||||||
| NC score | 0.050940 (rank : 52) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.050596 (rank : 53) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
EP400_MOUSE
|
||||||
| NC score | 0.050042 (rank : 54) | θ value | 0.0563607 (rank : 30) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
EDEM3_HUMAN
|
||||||
| NC score | 0.048209 (rank : 55) | θ value | 0.47712 (rank : 35) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BZQ6, Q5TEZ0, Q9HCW1, Q9UFV7 | Gene names | EDEM3, C1orf22 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 3. | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.039332 (rank : 56) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
CENPJ_HUMAN
|
||||||
| NC score | 0.036784 (rank : 57) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9HC77, Q5T6R5, Q96KS5, Q9C067 | Gene names | CENPJ, CPAP, LAP, LIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein J (CENP-J) (Centrosomal P4.1-associated protein) (LAG-3-associated protein) (LYST-interacting protein 1). | |||||
|
NOP14_HUMAN
|
||||||
| NC score | 0.035699 (rank : 58) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
TNR6A_HUMAN
|
||||||
| NC score | 0.033746 (rank : 59) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
C8AP2_HUMAN
|
||||||
| NC score | 0.032904 (rank : 60) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
MN1_HUMAN
|
||||||
| NC score | 0.031909 (rank : 61) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q10571 | Gene names | MN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable tumor suppressor protein MN1. | |||||
|
CLSPN_HUMAN
|
||||||
| NC score | 0.030924 (rank : 62) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9HAW4, Q5VYG0, Q6P6H5, Q8IWI1 | Gene names | CLSPN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin (hClaspin) (Hu-Claspin). | |||||
|
NKTR_HUMAN
|
||||||
| NC score | 0.030260 (rank : 63) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
EP400_HUMAN
|
||||||
| NC score | 0.030092 (rank : 64) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
IF4B_HUMAN
|
||||||
| NC score | 0.029790 (rank : 65) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P23588 | Gene names | EIF4B | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
K0146_MOUSE
|
||||||
| NC score | 0.029678 (rank : 66) | θ value | 2.36792 (rank : 45) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BGX7, Q3TCD0, Q5U457, Q6A0B6, Q6IS60, Q811E1, Q8BWD6, Q8R305 | Gene names | Kiaa0146 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0146. | |||||
|
UBP33_HUMAN
|
||||||
| NC score | 0.029289 (rank : 67) | θ value | 1.38821 (rank : 41) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TEY7, Q8TEY6, Q96AV6, Q9H9F0, Q9UPQ5 | Gene names | USP33, KIAA1097, VDU1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 33 (EC 3.1.2.15) (Ubiquitin thioesterase 33) (Ubiquitin-specific-processing protease 33) (Deubiquitinating enzyme 33) (VHL-interacting deubiquitinating enzyme 1). | |||||
|
UBP8_HUMAN
|
||||||
| NC score | 0.028661 (rank : 68) | θ value | 2.36792 (rank : 46) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
IF2P_HUMAN
|
||||||
| NC score | 0.027446 (rank : 69) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
GRIN1_HUMAN
|
||||||
| NC score | 0.027444 (rank : 70) | θ value | 3.0926 (rank : 51) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
BPTF_HUMAN
|
||||||
| NC score | 0.023475 (rank : 71) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
RPGR_HUMAN
|
||||||
| NC score | 0.022033 (rank : 72) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
CCD47_HUMAN
|
||||||
| NC score | 0.019240 (rank : 73) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96A33, Q96D00, Q96JZ7, Q9H3E4, Q9NRG3 | Gene names | CCDC47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor. | |||||
|
TXLNB_MOUSE
|
||||||
| NC score | 0.018752 (rank : 74) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
RBBP5_HUMAN
|
||||||
| NC score | 0.018277 (rank : 75) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15291 | Gene names | RBBP5, RBQ3 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-binding protein 5 (RBBP-5) (Retinoblastoma-binding protein RBQ-3). | |||||
|
NEST_HUMAN
|
||||||
| NC score | 0.016399 (rank : 76) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
MYRIP_MOUSE
|
||||||
| NC score | 0.016374 (rank : 77) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K3I4, Q8CFC0, Q8K4H5 | Gene names | Myrip, Slac2c | |||
|
Domain Architecture |

|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
GP158_HUMAN
|
||||||
| NC score | 0.015811 (rank : 78) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5T848, Q6QR81, Q9ULT3 | Gene names | GPR158, KIAA1136 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 158 precursor. | |||||
|
DIAP1_HUMAN
|
||||||
| NC score | 0.015237 (rank : 79) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60610, Q9UC76 | Gene names | DIAPH1, DIAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1). | |||||
|
TIAM1_MOUSE
|
||||||
| NC score | 0.014910 (rank : 80) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60610 | Gene names | Tiam1, Tiam-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
CX039_MOUSE
|
||||||
| NC score | 0.014245 (rank : 81) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K2D0, Q3U3R2 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein CXorf39 homolog. | |||||
|
TIAM1_HUMAN
|
||||||
| NC score | 0.013353 (rank : 82) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13009 | Gene names | TIAM1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
ARX_HUMAN
|
||||||
| NC score | 0.012830 (rank : 83) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96QS3 | Gene names | ARX | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein ARX (Aristaless-related homeobox). | |||||
|
ARX_MOUSE
|
||||||
| NC score | 0.012744 (rank : 84) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O35085, Q9QYT4 | Gene names | Arx | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein ARX (Aristaless-related homeobox). | |||||
|
DIAP1_MOUSE
|
||||||
| NC score | 0.012698 (rank : 85) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 839 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O08808 | Gene names | Diaph1, Diap1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1) (mDIA1) (p140mDIA). | |||||
|
CK5P2_HUMAN
|
||||||
| NC score | 0.011221 (rank : 86) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
E41LA_HUMAN
|
||||||
| NC score | 0.010541 (rank : 87) | θ value | 3.0926 (rank : 50) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HCS5 | Gene names | EPB41L4A, EPB41L4 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 4A (Protein NBL4). | |||||
|
WRN_MOUSE
|
||||||
| NC score | 0.010026 (rank : 88) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O09053, O09050, Q9JKD4, Q9Z241, Q9Z242 | Gene names | Wrn | |||
|
Domain Architecture |

|
|||||
| Description | Werner syndrome ATP-dependent helicase homolog (EC 3.6.1.-). | |||||
|
LUZP1_MOUSE
|
||||||
| NC score | 0.009759 (rank : 89) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
MYO10_HUMAN
|
||||||
| NC score | 0.009365 (rank : 90) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9HD67, Q9NYM7, Q9P110, Q9P111, Q9UHF6 | Gene names | MYO10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin X). | |||||
|
MEFV_MOUSE
|
||||||
| NC score | 0.009017 (rank : 91) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JJ26 | Gene names | Mefv | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
DNM3B_MOUSE
|
||||||
| NC score | 0.008519 (rank : 92) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88509, O88510, O88511 | Gene names | Dnmt3b | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase MmuIIIB) (DNA MTase MmuIIIB) (M.MmuIIIB). | |||||
|
ATBF1_HUMAN
|
||||||
| NC score | 0.006341 (rank : 93) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
SMC4_MOUSE
|
||||||
| NC score | 0.006112 (rank : 94) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 830 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CG47, Q8BTS7, Q8BTY9, Q99K21 | Gene names | Smc4l1, Capc, Smc4 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (XCAP-C homolog). | |||||
|
WEE1_MOUSE
|
||||||
| NC score | 0.002118 (rank : 95) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 794 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P47810, Q9EPL7 | Gene names | Wee1 | |||
|
Domain Architecture |

|
|||||
| Description | Wee1-like protein kinase (EC 2.7.10.2) (Wee1A kinase). | |||||