Please be patient as the page loads
|
HES2_HUMAN
|
||||||
| SwissProt Accessions | Q9Y543, Q96EN4, Q9Y542 | Gene names | HES2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HES2_HUMAN
|
||||||
| θ value | 7.71989e-68 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Y543, Q96EN4, Q9Y542 | Gene names | HES2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HES2_MOUSE
|
||||||
| θ value | 3.72821e-46 (rank : 2) | NC score | 0.986247 (rank : 2) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O54792 | Gene names | Hes2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HES4_HUMAN
|
||||||
| θ value | 8.36355e-22 (rank : 3) | NC score | 0.916258 (rank : 3) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
HES1_HUMAN
|
||||||
| θ value | 1.86321e-21 (rank : 4) | NC score | 0.884145 (rank : 5) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
HES1_MOUSE
|
||||||
| θ value | 1.86321e-21 (rank : 5) | NC score | 0.884414 (rank : 4) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES6_MOUSE
|
||||||
| θ value | 1.47974e-10 (rank : 6) | NC score | 0.777908 (rank : 8) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9JHE6, Q3U2D7 | Gene names | Hes6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6). | |||||
|
HES6_HUMAN
|
||||||
| θ value | 1.9326e-10 (rank : 7) | NC score | 0.784409 (rank : 7) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96HZ4, Q8N2J2, Q96T93, Q9P2S3 | Gene names | HES6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6) (C- HAIRY1). | |||||
|
HEY2_MOUSE
|
||||||
| θ value | 4.30538e-10 (rank : 8) | NC score | 0.720168 (rank : 11) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HES7_HUMAN
|
||||||
| θ value | 1.63604e-09 (rank : 9) | NC score | 0.748039 (rank : 9) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES5_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 10) | NC score | 0.826413 (rank : 6) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
HES7_MOUSE
|
||||||
| θ value | 3.64472e-09 (rank : 11) | NC score | 0.739017 (rank : 10) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HEY2_HUMAN
|
||||||
| θ value | 3.64472e-09 (rank : 12) | NC score | 0.716751 (rank : 12) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY1_HUMAN
|
||||||
| θ value | 1.38499e-08 (rank : 13) | NC score | 0.704701 (rank : 14) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HEY1_MOUSE
|
||||||
| θ value | 1.80886e-08 (rank : 14) | NC score | 0.706993 (rank : 13) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
BHLH2_HUMAN
|
||||||
| θ value | 7.59969e-07 (rank : 15) | NC score | 0.535504 (rank : 16) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
HES3_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 16) | NC score | 0.674816 (rank : 15) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
BHLH2_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 17) | NC score | 0.508893 (rank : 17) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
BHLH3_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 18) | NC score | 0.500097 (rank : 19) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
BHLH3_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 19) | NC score | 0.501199 (rank : 18) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
PSL1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 20) | NC score | 0.041528 (rank : 37) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.029446 (rank : 41) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
SPR1A_MOUSE
|
||||||
| θ value | 1.06291 (rank : 22) | NC score | 0.034541 (rank : 39) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62266 | Gene names | Sprr1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cornifin A (Small proline-rich protein 1A) (SPR1A) (SPR1 A). | |||||
|
K1802_HUMAN
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.023081 (rank : 43) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
LRC46_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.025190 (rank : 42) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96FV0 | Gene names | LRRC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 46. | |||||
|
3MG_HUMAN
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.033500 (rank : 40) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P29372, Q13770, Q15275, Q15961, Q96BZ6, Q96S33, Q9NNX5 | Gene names | MPG, AAG, ANPG, MID1 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-3-methyladenine glycosylase (EC 3.2.2.21) (3-methyladenine DNA glycosidase) (ADPG) (3-alkyladenine DNA glycosylase) (N-methylpurine- DNA glycosylase). | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 26) | NC score | 0.093686 (rank : 21) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.095933 (rank : 20) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
SPR1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.036568 (rank : 38) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62267 | Gene names | Sprr1b | |||
|
Domain Architecture |

|
|||||
| Description | Cornifin B (Small proline-rich protein 1B) (SPR1B) (SPR1 B). | |||||
|
CAP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.015537 (rank : 47) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P40124 | Gene names | Cap1, Cap | |||
|
Domain Architecture |
|
|||||
| Description | Adenylyl cyclase-associated protein 1 (CAP 1). | |||||
|
DUET_HUMAN
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | -0.000775 (rank : 53) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 1220 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2A5, Q8TBQ5, Q9NSZ4 | Gene names | DUET, TRAD | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Duet (EC 2.7.11.1) (Serine/threonine kinase with Dbl- and pleckstrin homology domain). | |||||
|
HPS4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.022339 (rank : 44) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99KG7 | Gene names | Hps4, Le | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 4 protein homolog (Light-ear protein) (Le protein). | |||||
|
AMPB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.008063 (rank : 50) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VCT3 | Gene names | Rnpep | |||
|
Domain Architecture |

|
|||||
| Description | Aminopeptidase B (EC 3.4.11.6) (Ap-B) (Arginyl aminopeptidase) (Arginine aminopeptidase) (Cytosol aminopeptidase IV). | |||||
|
AP180_HUMAN
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.016853 (rank : 46) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60641, Q5JX13, Q68DL9, Q6P9D3, Q9NTY7 | Gene names | SNAP91, KIAA0656 | |||
|
Domain Architecture |
|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein). | |||||
|
AP180_MOUSE
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.017981 (rank : 45) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61548, Q61547 | Gene names | Snap91 | |||
|
Domain Architecture |
|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein) (Phosphoprotein F1- 20). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.008628 (rank : 49) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PDLI7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | 0.005047 (rank : 52) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 547 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NR12, Q14250, Q5XG82, Q6NVZ5, Q96C91, Q9BXB8, Q9BXB9 | Gene names | PDLIM7, ENIGMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ and LIM domain protein 7 (LIM mineralization protein) (LMP) (Protein enigma). | |||||
|
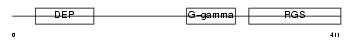
RGS11_HUMAN
|
||||||
| θ value | 8.99809 (rank : 37) | NC score | 0.006318 (rank : 51) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O94810, O75883 | Gene names | RGS11 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 11 (RGS11). | |||||
|
SNX19_HUMAN
|
||||||
| θ value | 8.99809 (rank : 38) | NC score | 0.015320 (rank : 48) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92543 | Gene names | SNX19, KIAA0254 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-19. | |||||
|
MAD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 39) | NC score | 0.065570 (rank : 33) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MITF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 40) | NC score | 0.083810 (rank : 25) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 41) | NC score | 0.079457 (rank : 29) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MLX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 42) | NC score | 0.084460 (rank : 24) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 43) | NC score | 0.084665 (rank : 23) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
SRBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 44) | NC score | 0.075137 (rank : 30) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
SRBP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 45) | NC score | 0.064454 (rank : 34) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
SRBP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 46) | NC score | 0.081172 (rank : 26) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
TFE3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.084834 (rank : 22) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
TFEB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.061524 (rank : 36) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.062755 (rank : 35) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
USF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.080483 (rank : 27) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.080448 (rank : 28) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.070692 (rank : 32) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
USF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.074802 (rank : 31) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
HES2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 7.71989e-68 (rank : 1) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Y543, Q96EN4, Q9Y542 | Gene names | HES2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HES2_MOUSE
|
||||||
| NC score | 0.986247 (rank : 2) | θ value | 3.72821e-46 (rank : 2) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O54792 | Gene names | Hes2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HES4_HUMAN
|
||||||
| NC score | 0.916258 (rank : 3) | θ value | 8.36355e-22 (rank : 3) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
HES1_MOUSE
|
||||||
| NC score | 0.884414 (rank : 4) | θ value | 1.86321e-21 (rank : 5) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES1_HUMAN
|
||||||
| NC score | 0.884145 (rank : 5) | θ value | 1.86321e-21 (rank : 4) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
HES5_MOUSE
|
||||||
| NC score | 0.826413 (rank : 6) | θ value | 2.79066e-09 (rank : 10) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
HES6_HUMAN
|
||||||
| NC score | 0.784409 (rank : 7) | θ value | 1.9326e-10 (rank : 7) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96HZ4, Q8N2J2, Q96T93, Q9P2S3 | Gene names | HES6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6) (C- HAIRY1). | |||||
|
HES6_MOUSE
|
||||||
| NC score | 0.777908 (rank : 8) | θ value | 1.47974e-10 (rank : 6) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9JHE6, Q3U2D7 | Gene names | Hes6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6). | |||||
|
HES7_HUMAN
|
||||||
| NC score | 0.748039 (rank : 9) | θ value | 1.63604e-09 (rank : 9) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES7_MOUSE
|
||||||
| NC score | 0.739017 (rank : 10) | θ value | 3.64472e-09 (rank : 11) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.720168 (rank : 11) | θ value | 4.30538e-10 (rank : 8) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HEY2_HUMAN
|
||||||
| NC score | 0.716751 (rank : 12) | θ value | 3.64472e-09 (rank : 12) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY1_MOUSE
|
||||||
| NC score | 0.706993 (rank : 13) | θ value | 1.80886e-08 (rank : 14) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HEY1_HUMAN
|
||||||
| NC score | 0.704701 (rank : 14) | θ value | 1.38499e-08 (rank : 13) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HES3_MOUSE
|
||||||
| NC score | 0.674816 (rank : 15) | θ value | 7.1131e-05 (rank : 16) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
BHLH2_HUMAN
|
||||||
| NC score | 0.535504 (rank : 16) | θ value | 7.59969e-07 (rank : 15) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
BHLH2_MOUSE
|
||||||
| NC score | 0.508893 (rank : 17) | θ value | 0.00035302 (rank : 17) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
BHLH3_HUMAN
|
||||||
| NC score | 0.501199 (rank : 18) | θ value | 0.00390308 (rank : 19) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BHLH3_MOUSE
|
||||||
| NC score | 0.500097 (rank : 19) | θ value | 0.00175202 (rank : 18) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.095933 (rank : 20) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.093686 (rank : 21) | θ value | 3.0926 (rank : 26) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.084834 (rank : 22) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.084665 (rank : 23) | θ value | θ > 10 (rank : 43) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.084460 (rank : 24) | θ value | θ > 10 (rank : 42) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.083810 (rank : 25) | θ value | θ > 10 (rank : 40) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
SRBP2_HUMAN
|
||||||
| NC score | 0.081172 (rank : 26) | θ value | θ > 10 (rank : 46) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.080483 (rank : 27) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.080448 (rank : 28) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.079457 (rank : 29) | θ value | θ > 10 (rank : 41) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
SRBP1_HUMAN
|
||||||
| NC score | 0.075137 (rank : 30) | θ value | θ > 10 (rank : 44) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.074802 (rank : 31) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.070692 (rank : 32) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
MAD1_MOUSE
|
||||||
| NC score | 0.065570 (rank : 33) | θ value | θ > 10 (rank : 39) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
SRBP1_MOUSE
|
||||||
| NC score | 0.064454 (rank : 34) | θ value | θ > 10 (rank : 45) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.062755 (rank : 35) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.061524 (rank : 36) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
PSL1_HUMAN
|
||||||
| NC score | 0.041528 (rank : 37) | θ value | 0.125558 (rank : 20) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
SPR1B_MOUSE
|
||||||
| NC score | 0.036568 (rank : 38) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62267 | Gene names | Sprr1b | |||
|
Domain Architecture |

|
|||||
| Description | Cornifin B (Small proline-rich protein 1B) (SPR1B) (SPR1 B). | |||||
|
SPR1A_MOUSE
|
||||||
| NC score | 0.034541 (rank : 39) | θ value | 1.06291 (rank : 22) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62266 | Gene names | Sprr1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cornifin A (Small proline-rich protein 1A) (SPR1A) (SPR1 A). | |||||
|
3MG_HUMAN
|
||||||
| NC score | 0.033500 (rank : 40) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P29372, Q13770, Q15275, Q15961, Q96BZ6, Q96S33, Q9NNX5 | Gene names | MPG, AAG, ANPG, MID1 | |||
|
Domain Architecture |
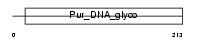
|
|||||
| Description | DNA-3-methyladenine glycosylase (EC 3.2.2.21) (3-methyladenine DNA glycosidase) (ADPG) (3-alkyladenine DNA glycosylase) (N-methylpurine- DNA glycosylase). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.029446 (rank : 41) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
LRC46_HUMAN
|
||||||
| NC score | 0.025190 (rank : 42) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96FV0 | Gene names | LRRC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 46. | |||||
|
K1802_HUMAN
|
||||||
| NC score | 0.023081 (rank : 43) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
HPS4_MOUSE
|
||||||
| NC score | 0.022339 (rank : 44) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99KG7 | Gene names | Hps4, Le | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 4 protein homolog (Light-ear protein) (Le protein). | |||||
|
AP180_MOUSE
|
||||||
| NC score | 0.017981 (rank : 45) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61548, Q61547 | Gene names | Snap91 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein) (Phosphoprotein F1- 20). | |||||
|
AP180_HUMAN
|
||||||
| NC score | 0.016853 (rank : 46) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60641, Q5JX13, Q68DL9, Q6P9D3, Q9NTY7 | Gene names | SNAP91, KIAA0656 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein). | |||||
|
CAP1_MOUSE
|
||||||
| NC score | 0.015537 (rank : 47) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P40124 | Gene names | Cap1, Cap | |||
|
Domain Architecture |

|
|||||
| Description | Adenylyl cyclase-associated protein 1 (CAP 1). | |||||
|
SNX19_HUMAN
|
||||||
| NC score | 0.015320 (rank : 48) | θ value | 8.99809 (rank : 38) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92543 | Gene names | SNX19, KIAA0254 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-19. | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.008628 (rank : 49) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
AMPB_MOUSE
|
||||||
| NC score | 0.008063 (rank : 50) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VCT3 | Gene names | Rnpep | |||
|
Domain Architecture |

|
|||||
| Description | Aminopeptidase B (EC 3.4.11.6) (Ap-B) (Arginyl aminopeptidase) (Arginine aminopeptidase) (Cytosol aminopeptidase IV). | |||||
|
RGS11_HUMAN
|
||||||
| NC score | 0.006318 (rank : 51) | θ value | 8.99809 (rank : 37) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O94810, O75883 | Gene names | RGS11 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 11 (RGS11). | |||||
|
PDLI7_HUMAN
|
||||||
| NC score | 0.005047 (rank : 52) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 547 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NR12, Q14250, Q5XG82, Q6NVZ5, Q96C91, Q9BXB8, Q9BXB9 | Gene names | PDLIM7, ENIGMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ and LIM domain protein 7 (LIM mineralization protein) (LMP) (Protein enigma). | |||||
|
DUET_HUMAN
|
||||||
| NC score | -0.000775 (rank : 53) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 1220 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2A5, Q8TBQ5, Q9NSZ4 | Gene names | DUET, TRAD | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Duet (EC 2.7.11.1) (Serine/threonine kinase with Dbl- and pleckstrin homology domain). | |||||