Please be patient as the page loads
|
HS70L_HUMAN
|
||||||
| SwissProt Accessions | P34931, O75634, Q2HXR3, Q8NE72, Q96QC9, Q9UQM1 | Gene names | HSPA1L | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 1L (Heat shock 70 kDa protein 1-like) (Heat shock 70 kDa protein 1-Hom) (HSP70-Hom). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
GRP78_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.982592 (rank : 13) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P11021, Q2EF78, Q9NPF1, Q9UK02 | Gene names | HSPA5, GRP78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP) (Endoplasmic reticulum lumenal Ca(2+)-binding protein grp78). | |||||
|
GRP78_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.982469 (rank : 14) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P20029, O35642, Q3UFF2, Q61630 | Gene names | Hspa5, Grp78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP). | |||||
|
HS70A_MOUSE
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.997308 (rank : 4) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61696, Q61697, Q7TQD8, Q9QWJ5 | Gene names | Hspa1a, Hsp70-3, Hsp70A1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock 70 kDa protein 1A (Heat shock 70 kDa protein 3) (HSP70.3) (Hsp68). | |||||
|
HS70B_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.997341 (rank : 3) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P17879, Q61689, Q925V6 | Gene names | Hspa1b, Hcp70.1, Hsp70-1, Hsp70a1, Hspa1 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 1B (HSP70.1). | |||||
|
HS70L_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P34931, O75634, Q2HXR3, Q8NE72, Q96QC9, Q9UQM1 | Gene names | HSPA1L | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 1L (Heat shock 70 kDa protein 1-like) (Heat shock 70 kDa protein 1-Hom) (HSP70-Hom). | |||||
|
HS70L_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.999079 (rank : 2) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P16627, O88686, Q61693 | Gene names | Hspa1l, Hsc70t | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 1L (Heat shock 70 kDa protein 1-like) (Heat shock 70 kDa-like protein 1) (Spermatid-specific heat shock protein 70). | |||||
|
HSP71_HUMAN
|
||||||
| θ value | 0 (rank : 7) | NC score | 0.996905 (rank : 5) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P08107, P19790, Q9UQL9, Q9UQM0 | Gene names | HSPA1A, HSPA1 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 1 (HSP70.1) (HSP70-1/HSP70-2). | |||||
|
HSP72_HUMAN
|
||||||
| θ value | 0 (rank : 8) | NC score | 0.996349 (rank : 7) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P54652, Q15508, Q9UE78 | Gene names | HSPA2 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock-related 70 kDa protein 2 (Heat shock 70 kDa protein 2). | |||||
|
HSP72_MOUSE
|
||||||
| θ value | 0 (rank : 9) | NC score | 0.996457 (rank : 6) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P17156 | Gene names | Hspa2, Hcp70.2, Hsp70-2 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock-related 70 kDa protein 2 (Heat shock protein 70.2). | |||||
|
HSP76_HUMAN
|
||||||
| θ value | 0 (rank : 10) | NC score | 0.994075 (rank : 8) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P17066, Q8IYK7, Q9BT95 | Gene names | HSPA6, HSP70B' | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 6 (Heat shock 70 kDa protein B'). | |||||
|
HSP7C_HUMAN
|
||||||
| θ value | 0 (rank : 11) | NC score | 0.992661 (rank : 9) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P11142, Q9H3R6 | Gene names | HSPA8, HSC70, HSP73, HSPA10 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock cognate 71 kDa protein (Heat shock 70 kDa protein 8). | |||||
|
HSP7C_MOUSE
|
||||||
| θ value | 0 (rank : 12) | NC score | 0.991959 (rank : 10) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P63017, P08109, P12225, Q62373, Q62374, Q62375 | Gene names | Hspa8, Hsc70, Hsc73 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock cognate 71 kDa protein (Heat shock 70 kDa protein 8). | |||||
|
GRP75_HUMAN
|
||||||
| θ value | 7.54289e-148 (rank : 13) | NC score | 0.982934 (rank : 12) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P38646, P30036, P31932, Q9BWB7 | Gene names | HSPA9B, GRP75, HSPA9 | |||
|
Domain Architecture |

|
|||||
| Description | Stress-70 protein, mitochondrial precursor (75 kDa glucose-regulated protein) (GRP 75) (Peptide-binding protein 74) (PBP74) (Mortalin) (MOT). | |||||
|
GRP75_MOUSE
|
||||||
| θ value | 3.7435e-147 (rank : 14) | NC score | 0.984088 (rank : 11) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P38647, Q9CQ05 | Gene names | Hspa9a, Grp75, Hsp74, Hspa9 | |||
|
Domain Architecture |

|
|||||
| Description | Stress-70 protein, mitochondrial precursor (75 kDa glucose-regulated protein) (GRP 75) (Peptide-binding protein 74) (PBP74) (P66 MOT) (Mortalin). | |||||
|
HS74L_MOUSE
|
||||||
| θ value | 5.8972e-76 (rank : 15) | NC score | 0.903160 (rank : 19) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P48722, P97854, Q8BQD0, Q8CC45, Q91X29 | Gene names | Hspa4l, Apg1, Hsp4l, Osp94 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4L (Osmotic stress protein 94) (Heat shock 70-related protein APG-1). | |||||
|
HS74L_HUMAN
|
||||||
| θ value | 1.71583e-75 (rank : 16) | NC score | 0.900461 (rank : 20) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O95757, Q4W5M5, Q8IWA2 | Gene names | HSPA4L, APG1, OSP94 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4L (Osmotic stress protein 94) (Heat shock 70-related protein APG-1). | |||||
|
HSP74_MOUSE
|
||||||
| θ value | 2.24094e-75 (rank : 17) | NC score | 0.889591 (rank : 22) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61316 | Gene names | Hspa4, Hsp110 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4 (Heat shock 70-related protein APG-2). | |||||
|
HSP74_HUMAN
|
||||||
| θ value | 3.82246e-75 (rank : 18) | NC score | 0.903191 (rank : 18) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P34932, O95756 | Gene names | HSPA4 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4 (Heat shock 70-related protein APG-2) (HSP70RY). | |||||
|
STCH_HUMAN
|
||||||
| θ value | 4.99229e-75 (rank : 19) | NC score | 0.965484 (rank : 15) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P48723, Q8NE40 | Gene names | STCH | |||
|
Domain Architecture |
|
|||||
| Description | Stress 70 protein chaperone microsome-associated 60 kDa protein precursor (Microsomal stress 70 protein ATPase core). | |||||
|
STCH_MOUSE
|
||||||
| θ value | 7.20884e-74 (rank : 20) | NC score | 0.964665 (rank : 16) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BM72, Q3TII6, Q9D1X5 | Gene names | Stch | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stress 70 protein chaperone microsome-associated 60 kDa protein precursor (Microsomal stress 70 protein ATPase core). | |||||
|
HS105_HUMAN
|
||||||
| θ value | 1.60597e-73 (rank : 21) | NC score | 0.898856 (rank : 21) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q92598, O95739, Q5TBM6, Q5TBM8, Q9UPC4 | Gene names | HSPH1, HSP105, HSP110, KIAA0201 | |||
|
Domain Architecture |

|
|||||
| Description | Heat-shock protein 105 kDa (Heat shock 110 kDa protein) (Antigen NY- CO-25). | |||||
|
HS105_MOUSE
|
||||||
| θ value | 1.60597e-73 (rank : 22) | NC score | 0.906297 (rank : 17) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q61699, Q3TNS2, Q3UIY8, Q62578, Q62579, Q6A0A5, Q8C430, Q8VCW6 | Gene names | Hsph1, Hsp105, Hsp110, Kiaa0201 | |||
|
Domain Architecture |

|
|||||
| Description | Heat-shock protein 105 kDa (Heat shock-related 100 kDa protein E7I) (HSP-E7I) (Heat shock 110 kDa protein) (42 degrees C-HSP). | |||||
|
OXRP_HUMAN
|
||||||
| θ value | 1.19655e-52 (rank : 23) | NC score | 0.873258 (rank : 23) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 544 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y4L1 | Gene names | HYOU1, ORP150 | |||
|
Domain Architecture |

|
|||||
| Description | 150 kDa oxygen-regulated protein precursor (Orp150) (Hypoxia up- regulated 1). | |||||
|
ANR45_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 24) | NC score | 0.252804 (rank : 24) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5TZF3, Q6ZST1 | Gene names | ANKRD45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 45. | |||||
|
HS12B_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 25) | NC score | 0.145050 (rank : 26) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9CZJ2 | Gene names | Hspa12b | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 12B. | |||||
|
E2AK2_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 26) | NC score | 0.008876 (rank : 40) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03963, Q61742, Q62026 | Gene names | Eif2ak2, Pkr, Prkr, Tik | |||
|
Domain Architecture |
|
|||||
| Description | Interferon-induced, double-stranded RNA-activated protein kinase (EC 2.7.11.1) (Interferon-inducible RNA-dependent protein kinase) (Protein kinase RNA-activated) (PKR) (p68 kinase) (P1/eIF-2A protein kinase) (Serine/threonine-protein kinase TIK). | |||||
|
ANR45_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 27) | NC score | 0.193476 (rank : 25) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q810N6 | Gene names | Ankrd45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 45. | |||||
|
HS12B_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 28) | NC score | 0.090858 (rank : 27) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96MM6, Q9BR52 | Gene names | HSPA12B, C20orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock 70 kDa protein 12B. | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 0.125558 (rank : 29) | NC score | 0.019571 (rank : 36) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 30) | NC score | 0.026059 (rank : 31) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
CTRO_MOUSE
|
||||||
| θ value | 1.38821 (rank : 31) | NC score | 0.000676 (rank : 49) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
FLNA_HUMAN
|
||||||
| θ value | 1.38821 (rank : 32) | NC score | 0.027873 (rank : 30) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P21333 | Gene names | FLNA, FLN, FLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Filamin-A (Alpha-filamin) (Filamin-1) (Endothelial actin-binding protein) (Actin-binding protein 280) (ABP-280) (Nonmuscle filamin). | |||||
|
KTN1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 33) | NC score | 0.005921 (rank : 44) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
JAK1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | -0.001714 (rank : 51) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 854 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P52332, Q62126 | Gene names | Jak1 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase JAK1 (EC 2.7.10.2) (Janus kinase 1) (JAK-1). | |||||
|
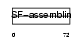
K2C3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 35) | NC score | 0.015367 (rank : 37) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P12035, Q701L8 | Gene names | KRT3 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin, type II cytoskeletal 3 (Cytokeratin-3) (CK-3) (Keratin-3) (K3) (65 kDa cytokeratin). | |||||
|
SYK_HUMAN
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.010211 (rank : 39) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15046, O14946, Q96J25 | Gene names | KARS, KIAA0070 | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl-tRNA synthetase (EC 6.1.1.6) (Lysine--tRNA ligase) (LysRS). | |||||
|
K2C1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.006906 (rank : 42) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P04264, Q14720, Q6GSJ0, Q9H298 | Gene names | KRT1, KRTA | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type II cytoskeletal 1 (Cytokeratin-1) (CK-1) (Keratin-1) (K1) (67 kDa cytokeratin) (Hair alpha protein). | |||||
|
PDP1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.020977 (rank : 34) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9P0J1, Q5U5K1 | Gene names | PDP1, PDP, PPM2C | |||
|
Domain Architecture |

|
|||||
| Description | [Pyruvate dehydrogenase [lipoamide]]-phosphatase 1, mitochondrial precursor (EC 3.1.3.43) (PDP 1) (Pyruvate dehydrogenase phosphatase, catalytic subunit 1) (PDPC 1) (Protein phosphatase 2C). | |||||
|
PDP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.020897 (rank : 35) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q3UV70 | Gene names | Pdp1, Ppm2c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | [Pyruvate dehydrogenase [lipoamide]]-phosphatase 1, mitochondrial precursor (EC 3.1.3.43) (PDP 1) (Pyruvate dehydrogenase phosphatase, catalytic subunit 1) (PDPC 1) (Protein phosphatase 2C). | |||||
|
SYWM_MOUSE
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.037563 (rank : 29) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CYK1 | Gene names | Wars2 | |||
|
Domain Architecture |
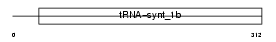
|
|||||
| Description | Tryptophanyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.2) (Tryptophan--tRNA ligase) (TrpRS) ((Mt)TrpRS). | |||||
|
TAF1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.005336 (rank : 45) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P21675, Q6IUZ1 | Gene names | TAF1, BA2R, CCG1, TAF2A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 1 (EC 2.7.11.1) (Transcription initiation factor TFIID 250 kDa subunit) (TAF(II)250) (TAFII-250) (TAFII250) (TBP-associated factor 250 kDa) (p250) (Cell cycle gene 1 protein). | |||||
|
DMD_MOUSE
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.003090 (rank : 47) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
FLNA_MOUSE
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.023301 (rank : 33) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BTM8, O54934, Q7TQI1, Q8BLK1, Q8BTN7, Q8VHX5, Q8VHX8, Q99KQ2 | Gene names | Flna, Fln, Fln1 | |||
|
Domain Architecture |

|
|||||
| Description | Filamin-A (Alpha-filamin) (Filamin-1) (Endothelial actin-binding protein) (Actin-binding protein 280) (ABP-280) (Nonmuscle filamin). | |||||
|
NIN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.010566 (rank : 38) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
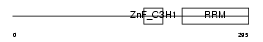
U2AFM_HUMAN
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.024094 (rank : 32) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15696 | Gene names | ZRSR2, U2AF1-RS2, U2AF1L2, U2AF1RS2 | |||
|
Domain Architecture |
|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2) (NY-REN-20 antigen). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.006740 (rank : 43) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
CELR1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | -0.001102 (rank : 50) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35161 | Gene names | Celsr1 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 1 precursor. | |||||
|
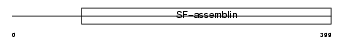
K2C7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.008309 (rank : 41) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P08729, Q92676, Q9BUD8, Q9Y3R7 | Gene names | KRT7, SCL | |||
|
Domain Architecture |
|
|||||
| Description | Keratin, type II cytoskeletal 7 (Cytokeratin-7) (CK-7) (Keratin-7) (K7) (Sarcolectin). | |||||
|
RRBP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.001504 (rank : 48) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
TEX9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.003221 (rank : 46) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D845, O54764 | Gene names | Tex9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 9 protein (Testis-specifically expressed protein 1) (Tsec-1). | |||||
|
SFR12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.059117 (rank : 28) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
HS70L_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P34931, O75634, Q2HXR3, Q8NE72, Q96QC9, Q9UQM1 | Gene names | HSPA1L | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 1L (Heat shock 70 kDa protein 1-like) (Heat shock 70 kDa protein 1-Hom) (HSP70-Hom). | |||||
|
HS70L_MOUSE
|
||||||
| NC score | 0.999079 (rank : 2) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P16627, O88686, Q61693 | Gene names | Hspa1l, Hsc70t | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 1L (Heat shock 70 kDa protein 1-like) (Heat shock 70 kDa-like protein 1) (Spermatid-specific heat shock protein 70). | |||||
|
HS70B_MOUSE
|
||||||
| NC score | 0.997341 (rank : 3) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P17879, Q61689, Q925V6 | Gene names | Hspa1b, Hcp70.1, Hsp70-1, Hsp70a1, Hspa1 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 1B (HSP70.1). | |||||
|
HS70A_MOUSE
|
||||||
| NC score | 0.997308 (rank : 4) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61696, Q61697, Q7TQD8, Q9QWJ5 | Gene names | Hspa1a, Hsp70-3, Hsp70A1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock 70 kDa protein 1A (Heat shock 70 kDa protein 3) (HSP70.3) (Hsp68). | |||||
|
HSP71_HUMAN
|
||||||
| NC score | 0.996905 (rank : 5) | θ value | 0 (rank : 7) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P08107, P19790, Q9UQL9, Q9UQM0 | Gene names | HSPA1A, HSPA1 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 1 (HSP70.1) (HSP70-1/HSP70-2). | |||||
|
HSP72_MOUSE
|
||||||
| NC score | 0.996457 (rank : 6) | θ value | 0 (rank : 9) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P17156 | Gene names | Hspa2, Hcp70.2, Hsp70-2 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock-related 70 kDa protein 2 (Heat shock protein 70.2). | |||||
|
HSP72_HUMAN
|
||||||
| NC score | 0.996349 (rank : 7) | θ value | 0 (rank : 8) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P54652, Q15508, Q9UE78 | Gene names | HSPA2 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock-related 70 kDa protein 2 (Heat shock 70 kDa protein 2). | |||||
|
HSP76_HUMAN
|
||||||
| NC score | 0.994075 (rank : 8) | θ value | 0 (rank : 10) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P17066, Q8IYK7, Q9BT95 | Gene names | HSPA6, HSP70B' | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 6 (Heat shock 70 kDa protein B'). | |||||
|
HSP7C_HUMAN
|
||||||
| NC score | 0.992661 (rank : 9) | θ value | 0 (rank : 11) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P11142, Q9H3R6 | Gene names | HSPA8, HSC70, HSP73, HSPA10 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock cognate 71 kDa protein (Heat shock 70 kDa protein 8). | |||||
|
HSP7C_MOUSE
|
||||||
| NC score | 0.991959 (rank : 10) | θ value | 0 (rank : 12) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P63017, P08109, P12225, Q62373, Q62374, Q62375 | Gene names | Hspa8, Hsc70, Hsc73 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock cognate 71 kDa protein (Heat shock 70 kDa protein 8). | |||||
|
GRP75_MOUSE
|
||||||
| NC score | 0.984088 (rank : 11) | θ value | 3.7435e-147 (rank : 14) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P38647, Q9CQ05 | Gene names | Hspa9a, Grp75, Hsp74, Hspa9 | |||
|
Domain Architecture |

|
|||||
| Description | Stress-70 protein, mitochondrial precursor (75 kDa glucose-regulated protein) (GRP 75) (Peptide-binding protein 74) (PBP74) (P66 MOT) (Mortalin). | |||||
|
GRP75_HUMAN
|
||||||
| NC score | 0.982934 (rank : 12) | θ value | 7.54289e-148 (rank : 13) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P38646, P30036, P31932, Q9BWB7 | Gene names | HSPA9B, GRP75, HSPA9 | |||
|
Domain Architecture |

|
|||||
| Description | Stress-70 protein, mitochondrial precursor (75 kDa glucose-regulated protein) (GRP 75) (Peptide-binding protein 74) (PBP74) (Mortalin) (MOT). | |||||
|
GRP78_HUMAN
|
||||||
| NC score | 0.982592 (rank : 13) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P11021, Q2EF78, Q9NPF1, Q9UK02 | Gene names | HSPA5, GRP78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP) (Endoplasmic reticulum lumenal Ca(2+)-binding protein grp78). | |||||
|
GRP78_MOUSE
|
||||||
| NC score | 0.982469 (rank : 14) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P20029, O35642, Q3UFF2, Q61630 | Gene names | Hspa5, Grp78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP). | |||||
|
STCH_HUMAN
|
||||||
| NC score | 0.965484 (rank : 15) | θ value | 4.99229e-75 (rank : 19) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P48723, Q8NE40 | Gene names | STCH | |||
|
Domain Architecture |

|
|||||
| Description | Stress 70 protein chaperone microsome-associated 60 kDa protein precursor (Microsomal stress 70 protein ATPase core). | |||||
|
STCH_MOUSE
|
||||||
| NC score | 0.964665 (rank : 16) | θ value | 7.20884e-74 (rank : 20) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BM72, Q3TII6, Q9D1X5 | Gene names | Stch | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stress 70 protein chaperone microsome-associated 60 kDa protein precursor (Microsomal stress 70 protein ATPase core). | |||||
|
HS105_MOUSE
|
||||||
| NC score | 0.906297 (rank : 17) | θ value | 1.60597e-73 (rank : 22) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q61699, Q3TNS2, Q3UIY8, Q62578, Q62579, Q6A0A5, Q8C430, Q8VCW6 | Gene names | Hsph1, Hsp105, Hsp110, Kiaa0201 | |||
|
Domain Architecture |

|
|||||
| Description | Heat-shock protein 105 kDa (Heat shock-related 100 kDa protein E7I) (HSP-E7I) (Heat shock 110 kDa protein) (42 degrees C-HSP). | |||||
|
HSP74_HUMAN
|
||||||
| NC score | 0.903191 (rank : 18) | θ value | 3.82246e-75 (rank : 18) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P34932, O95756 | Gene names | HSPA4 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4 (Heat shock 70-related protein APG-2) (HSP70RY). | |||||
|
HS74L_MOUSE
|
||||||
| NC score | 0.903160 (rank : 19) | θ value | 5.8972e-76 (rank : 15) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P48722, P97854, Q8BQD0, Q8CC45, Q91X29 | Gene names | Hspa4l, Apg1, Hsp4l, Osp94 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4L (Osmotic stress protein 94) (Heat shock 70-related protein APG-1). | |||||
|
HS74L_HUMAN
|
||||||
| NC score | 0.900461 (rank : 20) | θ value | 1.71583e-75 (rank : 16) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O95757, Q4W5M5, Q8IWA2 | Gene names | HSPA4L, APG1, OSP94 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4L (Osmotic stress protein 94) (Heat shock 70-related protein APG-1). | |||||
|
HS105_HUMAN
|
||||||
| NC score | 0.898856 (rank : 21) | θ value | 1.60597e-73 (rank : 21) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q92598, O95739, Q5TBM6, Q5TBM8, Q9UPC4 | Gene names | HSPH1, HSP105, HSP110, KIAA0201 | |||
|
Domain Architecture |

|
|||||
| Description | Heat-shock protein 105 kDa (Heat shock 110 kDa protein) (Antigen NY- CO-25). | |||||
|
HSP74_MOUSE
|
||||||
| NC score | 0.889591 (rank : 22) | θ value | 2.24094e-75 (rank : 17) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61316 | Gene names | Hspa4, Hsp110 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4 (Heat shock 70-related protein APG-2). | |||||
|
OXRP_HUMAN
|
||||||
| NC score | 0.873258 (rank : 23) | θ value | 1.19655e-52 (rank : 23) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 544 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y4L1 | Gene names | HYOU1, ORP150 | |||
|
Domain Architecture |

|
|||||
| Description | 150 kDa oxygen-regulated protein precursor (Orp150) (Hypoxia up- regulated 1). | |||||
|
ANR45_HUMAN
|
||||||
| NC score | 0.252804 (rank : 24) | θ value | 7.1131e-05 (rank : 24) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5TZF3, Q6ZST1 | Gene names | ANKRD45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 45. | |||||
|
ANR45_MOUSE
|
||||||
| NC score | 0.193476 (rank : 25) | θ value | 0.0330416 (rank : 27) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q810N6 | Gene names | Ankrd45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 45. | |||||
|
HS12B_MOUSE
|
||||||
| NC score | 0.145050 (rank : 26) | θ value | 0.00175202 (rank : 25) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9CZJ2 | Gene names | Hspa12b | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock 70 kDa protein 12B. | |||||
|
HS12B_HUMAN
|
||||||
| NC score | 0.090858 (rank : 27) | θ value | 0.0961366 (rank : 28) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96MM6, Q9BR52 | Gene names | HSPA12B, C20orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock 70 kDa protein 12B. | |||||
|
SFR12_HUMAN
|
||||||
| NC score | 0.059117 (rank : 28) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
SYWM_MOUSE
|
||||||
| NC score | 0.037563 (rank : 29) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CYK1 | Gene names | Wars2 | |||
|
Domain Architecture |
|
|||||
| Description | Tryptophanyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.2) (Tryptophan--tRNA ligase) (TrpRS) ((Mt)TrpRS). | |||||
|
FLNA_HUMAN
|
||||||
| NC score | 0.027873 (rank : 30) | θ value | 1.38821 (rank : 32) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P21333 | Gene names | FLNA, FLN, FLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Filamin-A (Alpha-filamin) (Filamin-1) (Endothelial actin-binding protein) (Actin-binding protein 280) (ABP-280) (Nonmuscle filamin). | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.026059 (rank : 31) | θ value | 0.163984 (rank : 30) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
U2AFM_HUMAN
|
||||||
| NC score | 0.024094 (rank : 32) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15696 | Gene names | ZRSR2, U2AF1-RS2, U2AF1L2, U2AF1RS2 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2) (NY-REN-20 antigen). | |||||
|
FLNA_MOUSE
|
||||||
| NC score | 0.023301 (rank : 33) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BTM8, O54934, Q7TQI1, Q8BLK1, Q8BTN7, Q8VHX5, Q8VHX8, Q99KQ2 | Gene names | Flna, Fln, Fln1 | |||
|
Domain Architecture |

|
|||||
| Description | Filamin-A (Alpha-filamin) (Filamin-1) (Endothelial actin-binding protein) (Actin-binding protein 280) (ABP-280) (Nonmuscle filamin). | |||||
|
PDP1_HUMAN
|
||||||
| NC score | 0.020977 (rank : 34) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9P0J1, Q5U5K1 | Gene names | PDP1, PDP, PPM2C | |||
|
Domain Architecture |

|
|||||
| Description | [Pyruvate dehydrogenase [lipoamide]]-phosphatase 1, mitochondrial precursor (EC 3.1.3.43) (PDP 1) (Pyruvate dehydrogenase phosphatase, catalytic subunit 1) (PDPC 1) (Protein phosphatase 2C). | |||||
|
PDP1_MOUSE
|
||||||
| NC score | 0.020897 (rank : 35) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q3UV70 | Gene names | Pdp1, Ppm2c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | [Pyruvate dehydrogenase [lipoamide]]-phosphatase 1, mitochondrial precursor (EC 3.1.3.43) (PDP 1) (Pyruvate dehydrogenase phosphatase, catalytic subunit 1) (PDPC 1) (Protein phosphatase 2C). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.019571 (rank : 36) | θ value | 0.125558 (rank : 29) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
K2C3_HUMAN
|
||||||
| NC score | 0.015367 (rank : 37) | θ value | 1.81305 (rank : 35) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P12035, Q701L8 | Gene names | KRT3 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type II cytoskeletal 3 (Cytokeratin-3) (CK-3) (Keratin-3) (K3) (65 kDa cytokeratin). | |||||
|
NIN_HUMAN
|
||||||
| NC score | 0.010566 (rank : 38) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
SYK_HUMAN
|
||||||
| NC score | 0.010211 (rank : 39) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15046, O14946, Q96J25 | Gene names | KARS, KIAA0070 | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl-tRNA synthetase (EC 6.1.1.6) (Lysine--tRNA ligase) (LysRS). | |||||
|
E2AK2_MOUSE
|
||||||
| NC score | 0.008876 (rank : 40) | θ value | 0.0193708 (rank : 26) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03963, Q61742, Q62026 | Gene names | Eif2ak2, Pkr, Prkr, Tik | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced, double-stranded RNA-activated protein kinase (EC 2.7.11.1) (Interferon-inducible RNA-dependent protein kinase) (Protein kinase RNA-activated) (PKR) (p68 kinase) (P1/eIF-2A protein kinase) (Serine/threonine-protein kinase TIK). | |||||
|
K2C7_HUMAN
|
||||||
| NC score | 0.008309 (rank : 41) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P08729, Q92676, Q9BUD8, Q9Y3R7 | Gene names | KRT7, SCL | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type II cytoskeletal 7 (Cytokeratin-7) (CK-7) (Keratin-7) (K7) (Sarcolectin). | |||||
|
K2C1_HUMAN
|
||||||
| NC score | 0.006906 (rank : 42) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P04264, Q14720, Q6GSJ0, Q9H298 | Gene names | KRT1, KRTA | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type II cytoskeletal 1 (Cytokeratin-1) (CK-1) (Keratin-1) (K1) (67 kDa cytokeratin) (Hair alpha protein). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.006740 (rank : 43) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
KTN1_MOUSE
|
||||||
| NC score | 0.005921 (rank : 44) | θ value | 1.38821 (rank : 33) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
TAF1_HUMAN
|
||||||
| NC score | 0.005336 (rank : 45) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P21675, Q6IUZ1 | Gene names | TAF1, BA2R, CCG1, TAF2A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 1 (EC 2.7.11.1) (Transcription initiation factor TFIID 250 kDa subunit) (TAF(II)250) (TAFII-250) (TAFII250) (TBP-associated factor 250 kDa) (p250) (Cell cycle gene 1 protein). | |||||
|
TEX9_MOUSE
|
||||||
| NC score | 0.003221 (rank : 46) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D845, O54764 | Gene names | Tex9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 9 protein (Testis-specifically expressed protein 1) (Tsec-1). | |||||
|
DMD_MOUSE
|
||||||
| NC score | 0.003090 (rank : 47) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
RRBP1_HUMAN
|
||||||
| NC score | 0.001504 (rank : 48) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
CTRO_MOUSE
|
||||||
| NC score | 0.000676 (rank : 49) | θ value | 1.38821 (rank : 31) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
CELR1_MOUSE
|
||||||
| NC score | -0.001102 (rank : 50) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35161 | Gene names | Celsr1 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 1 precursor. | |||||
|
JAK1_MOUSE
|
||||||
| NC score | -0.001714 (rank : 51) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 854 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P52332, Q62126 | Gene names | Jak1 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase JAK1 (EC 2.7.10.2) (Janus kinase 1) (JAK-1). | |||||