Please be patient as the page loads
|
HBS1L_HUMAN
|
||||||
| SwissProt Accessions | Q9Y450, Q5T7G3, Q8NDW9, Q9UPW3 | Gene names | HBS1L, HBS1, KIAA1038 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HBS1-like protein (ERFS). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HBS1L_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9Y450, Q5T7G3, Q8NDW9, Q9UPW3 | Gene names | HBS1L, HBS1, KIAA1038 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HBS1-like protein (ERFS). | |||||
|
HBS1L_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.992304 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q69ZS7, Q91Z32, Q9CVT2, Q9CZ95, Q9WTY5 | Gene names | Hbs1l, Hbs1, Kiaa1038 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HBS1-like protein. | |||||
|
GSPT1_MOUSE
|
||||||
| θ value | 3.93732e-88 (rank : 3) | NC score | 0.950023 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8R050, O88179 | Gene names | Gspt1 | |||
|
Domain Architecture |

|
|||||
| Description | G1 to S phase transition protein 1 homolog. | |||||
|
GSPT1_HUMAN
|
||||||
| θ value | 5.14231e-88 (rank : 4) | NC score | 0.951068 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P15170 | Gene names | GSPT1 | |||
|
Domain Architecture |

|
|||||
| Description | G1 to S phase transition protein 1 homolog (GTP-binding protein GST1- HS). | |||||
|
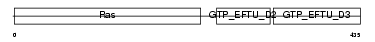
EF1A2_HUMAN
|
||||||
| θ value | 1.49618e-87 (rank : 5) | NC score | 0.942358 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q05639, P54266 | Gene names | EEF1A2, EEF1AL, STN | |||
|
Domain Architecture |
|
|||||
| Description | Elongation factor 1-alpha 2 (EF-1-alpha-2) (Elongation factor 1 A-2) (eEF1A-2) (Statin S1). | |||||
|
EF1A2_MOUSE
|
||||||
| θ value | 1.49618e-87 (rank : 6) | NC score | 0.942369 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P62631, P27706 | Gene names | Eef1a2, Eef1al, Stn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor 1-alpha 2 (EF-1-alpha-2) (Elongation factor 1 A-2) (eEF1A-2) (Statin S1). | |||||
|
EF1A1_HUMAN
|
||||||
| θ value | 1.40038e-85 (rank : 7) | NC score | 0.942232 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P68104, P04719, P04720 | Gene names | EEF1A1, EEF1A, EF1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor 1-alpha 1 (EF-1-alpha-1) (Elongation factor 1 A-1) (eEF1A-1) (Elongation factor Tu) (EF-Tu). | |||||
|
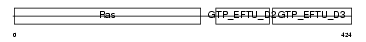
EF1A1_MOUSE
|
||||||
| θ value | 1.40038e-85 (rank : 8) | NC score | 0.942355 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10126, Q61511, Q6ZWN2, Q8BMB8, Q8BVS8, Q99KU5 | Gene names | Eef1a1, Eef1a | |||
|
Domain Architecture |
|
|||||
| Description | Elongation factor 1-alpha 1 (EF-1-alpha-1) (Elongation factor 1 A-1) (eEF1A-1) (Elongation factor Tu) (EF-Tu). | |||||
|
EFTU_MOUSE
|
||||||
| θ value | 4.89182e-30 (rank : 9) | NC score | 0.831674 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BFR5, Q6P919 | Gene names | Tufm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor Tu, mitochondrial precursor. | |||||
|
EFTU_HUMAN
|
||||||
| θ value | 1.4233e-29 (rank : 10) | NC score | 0.830649 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P49411, O15276 | Gene names | TUFM | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor Tu, mitochondrial precursor (EF-Tu) (P43). | |||||
|
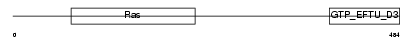
GTPB1_HUMAN
|
||||||
| θ value | 6.41864e-14 (rank : 11) | NC score | 0.603286 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O00178, Q6IC67 | Gene names | GTPBP1 | |||
|
Domain Architecture |
|
|||||
| Description | GTP-binding protein 1 (G-protein 1) (GP-1) (GP1). | |||||
|
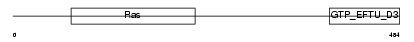
GTPB1_MOUSE
|
||||||
| θ value | 1.86753e-13 (rank : 12) | NC score | 0.599635 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O08582, Q3UD96, Q3UGW6, Q545R1, Q80ZY5 | Gene names | Gtpbp1, Gtpbp | |||
|
Domain Architecture |
|
|||||
| Description | GTP-binding protein 1 (G-protein 1) (GP-1) (GP1). | |||||
|
GTPB2_HUMAN
|
||||||
| θ value | 7.09661e-13 (rank : 13) | NC score | 0.554057 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BX10, Q5T7E8, Q8ND84, Q8TAH7, Q8WUA5, Q9HCS9, Q9NRU4, Q9NX60 | Gene names | GTPBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTP-binding protein 2. | |||||
|
GTPB2_MOUSE
|
||||||
| θ value | 7.09661e-13 (rank : 14) | NC score | 0.554038 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q3UJK4, Q9EST7, Q9JIX7 | Gene names | Gtpbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTP-binding protein 2 (GTP-binding-like protein 2). | |||||
|
SELB_MOUSE
|
||||||
| θ value | 2.0648e-12 (rank : 15) | NC score | 0.630301 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JHW4 | Gene names | Eefsec, Selb | |||
|
Domain Architecture |

|
|||||
| Description | Selenocysteine-specific elongation factor (Elongation factor sec) (Eukaryotic elongation factor, selenocysteine-tRNA-specific) (mSelB). | |||||
|
SELB_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 16) | NC score | 0.610842 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P57772 | Gene names | EEFSEC, SELB | |||
|
Domain Architecture |
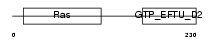
|
|||||
| Description | Selenocysteine-specific elongation factor (Elongation factor sec) (Eukaryotic elongation factor, selenocysteine-tRNA-specific). | |||||
|
IF2M_HUMAN
|
||||||
| θ value | 3.08544e-08 (rank : 17) | NC score | 0.444049 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P46199 | Gene names | MTIF2 | |||
|
Domain Architecture |
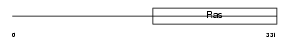
|
|||||
| Description | Translation initiation factor IF-2, mitochondrial precursor (IF-2Mt) (IF-2(Mt)). | |||||
|
EFG1_HUMAN
|
||||||
| θ value | 4.92598e-06 (rank : 18) | NC score | 0.382733 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96RP9, Q96T39 | Gene names | GFM1, EFG, EFG1, GFM | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 1, mitochondrial precursor (mEF-G 1) (Elongation factor G1). | |||||
|
EFG1_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 19) | NC score | 0.370854 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8K0D5, Q921D6, Q924I0 | Gene names | Gfm1, Efg, Efg1, Gfm | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 1, mitochondrial precursor (mEF-G 1) (Elongation factor G1). | |||||
|
EFG2_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 20) | NC score | 0.380663 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q969S9, Q8WYI0 | Gene names | GFM2, EFG2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 2, mitochondrial precursor (mEF-G 2) (Elongation factor G2). | |||||
|
EFG2_MOUSE
|
||||||
| θ value | 0.000121331 (rank : 21) | NC score | 0.368213 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8R2Q4 | Gene names | Gfm2, Efg2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 2, mitochondrial precursor (mEF-G 2) (Elongation factor G2). | |||||
|
IF2M_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 22) | NC score | 0.363672 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q91YJ5 | Gene names | Mtif2 | |||
|
Domain Architecture |
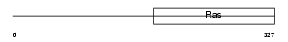
|
|||||
| Description | Translation initiation factor IF-2, mitochondrial precursor (IF-2Mt) (IF-2(Mt)). | |||||
|
IF2G_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 23) | NC score | 0.352699 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P41091 | Gene names | EIF2S3, EIF2G | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3 (Eukaryotic translation initiation factor 2 subunit gamma) (eIF-2-gamma). | |||||
|
IF2G_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 24) | NC score | 0.352690 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z0N1 | Gene names | Eif2s3x | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3, X-linked (Eukaryotic translation initiation factor 2 subunit gamma, X-linked) (eIF-2-gamma X). | |||||
|
IF2H_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 25) | NC score | 0.354572 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Z0N2 | Gene names | Eif2s3y | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3, Y-linked (Eukaryotic translation initiation factor 2 subunit gamma, Y-linked) (eIF-2-gamma Y). | |||||
|
U5S1_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 26) | NC score | 0.208099 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15029, Q9BUR0 | Gene names | EFTUD2, KIAA0031, SNRP116 | |||
|
Domain Architecture |

|
|||||
| Description | 116 kDa U5 small nuclear ribonucleoprotein component (U5 snRNP- specific protein, 116 kDa) (U5-116 kDa) (Elongation factor Tu GTP- binding domain protein 2) (hSNU114). | |||||
|
U5S1_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 27) | NC score | 0.208022 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O08810 | Gene names | Eftud2, Snrp116 | |||
|
Domain Architecture |

|
|||||
| Description | 116 kDa U5 small nuclear ribonucleoprotein component (U5 snRNP- specific protein, 116 kDa) (U5-116 kDa) (Elongation factor Tu GTP- binding domain protein 2). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 28) | NC score | 0.038426 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
SYBU_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 29) | NC score | 0.056484 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NX95, Q5R1T1, Q5R1T2, Q5R1T3, Q5Y2M6, Q8ND49, Q8TCR6, Q9P256 | Gene names | SYBU, GOLSYN, KIAA1472 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntabulin (Syntaxin-1-binding protein) (Golgi-localized syntaphilin- related protein). | |||||
|
IF2P_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 30) | NC score | 0.145802 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
MUC13_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 31) | NC score | 0.037862 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
BRE1B_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 32) | NC score | 0.015812 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75150, Q6AHZ6, Q6N005, Q7L3T6, Q8N615, Q96T18, Q9BSV9, Q9HC82 | Gene names | RNF40, BRE1B, KIAA0661 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1B (EC 6.3.2.-) (BRE1-B) (RING finger protein 40) (95 kDa retinoblastoma-associated protein) (RBP95). | |||||
|
NNP1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 33) | NC score | 0.042247 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56183, O35712, Q9ERE1, Q9JI07, Q9JK67, Q9JKU2 | Gene names | Nnp1 | |||
|
Domain Architecture |
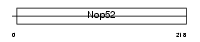
|
|||||
| Description | NNP-1 protein (Novel nuclear protein 1) (Nop52). | |||||
|
EF2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 34) | NC score | 0.249246 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P13639 | Gene names | EEF2, EF2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 2 (EF-2). | |||||
|
EF2_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 35) | NC score | 0.251460 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P58252, Q544E4 | Gene names | Eef2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 2 (EF-2). | |||||
|
SYBU_MOUSE
|
||||||
| θ value | 0.279714 (rank : 36) | NC score | 0.048394 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BHS8, Q3TQN7, Q401F3, Q401F4, Q571C1, Q6P1J2, Q80XH0, Q8BHS7, Q8BI27 | Gene names | Sybu, Golsyn, Kiaa1472 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntabulin (Syntaxin-1-binding protein) (Golgi-localized syntaphilin- related protein) (m-Golsyn). | |||||
|
BAIP2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 37) | NC score | 0.043601 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BKX1, Q91V97, Q923V9, Q923W0 | Gene names | Baiap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain-specific angiogenesis inhibitor 1-associated protein 2 (BAI1- associated protein 2) (BAI-associated protein 2) (Insulin receptor substrate p53) (IRSp53) (Insulin receptor substrate protein of 53 kDa) (Insulin receptor tyrosine kinase 53 kDa substrate). | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 0.62314 (rank : 38) | NC score | 0.035282 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
TCOF_HUMAN
|
||||||
| θ value | 0.62314 (rank : 39) | NC score | 0.028421 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
BAIP2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 40) | NC score | 0.046124 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UQB8, O43858, Q53HB1, Q86WC1, Q8N5C0, Q96CR7, Q9UBR3, Q9UQ43 | Gene names | BAIAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain-specific angiogenesis inhibitor 1-associated protein 2 (BAI1- associated protein 2) (BAI-associated protein 2) (Protein BAP2) (Insulin receptor substrate p53) (IRSp53) (Insulin receptor substrate protein of 53 kDa) (Insulin receptor substrate p53/p58) (IRSp53/58) (IRS-58) (Fas ligand-associated factor 3) (FLAF3). | |||||
|
MLF1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 41) | NC score | 0.027205 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P58340 | Gene names | MLF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myeloid leukemia factor 1 (Myelodysplasia-myeloid leukemia factor 1). | |||||
|
P66A_MOUSE
|
||||||
| θ value | 1.38821 (rank : 42) | NC score | 0.021471 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CHY6, Q8BTQ2, Q8VEC9 | Gene names | Gatad2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional repressor p66 alpha (GATA zinc finger domain- containing protein 2A). | |||||
|
BCL7A_MOUSE
|
||||||
| θ value | 2.36792 (rank : 43) | NC score | 0.017514 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CXE2, Q8C361, Q8C8M8, Q8VD15 | Gene names | Bcl7a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell CLL/lymphoma 7 protein family member A. | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.010713 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
TM131_MOUSE
|
||||||
| θ value | 2.36792 (rank : 45) | NC score | 0.022611 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O70472 | Gene names | Tmem131, D1Bwg0491e, Rw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 131 (Protein RW1). | |||||
|
CCD82_MOUSE
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.017950 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6PG04, Q3V462 | Gene names | Ccdc82 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 82. | |||||
|
LIMA1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.011899 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9ERG0, Q9ERG1 | Gene names | Lima1, D15Ertd366e, Eplin | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm) (mEPLIN). | |||||
|
BRCA2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.012844 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51587, O00183, O15008, Q13879 | Gene names | BRCA2, FANCD1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein (Fanconi anemia group D1 protein). | |||||
|
ELL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.012093 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O08856 | Gene names | Ell | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL (Eleven-nineteen lysine-rich leukemia protein). | |||||
|
H10_HUMAN
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.016638 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P07305 | Gene names | H1F0, H1FV | |||
|
Domain Architecture |
|
|||||
| Description | Histone H1.0 (H1(0)) (Histone H1'). | |||||
|
LRRK1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.000051 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q38SD2, Q6NVH5, Q6NYC0, Q6ZNL9, Q6ZNM9, Q96JN5, Q9H5S3 | Gene names | LRRK1, KIAA1790 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 1 (EC 2.7.11.1). | |||||
|
SFRS4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.005576 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
BRD7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.009435 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NPI1, Q4VC09, Q8N2L9, Q96KA4, Q9BV48, Q9UH59 | Gene names | BRD7, BP75, CELTIX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 7 (75 kDa bromodomain protein) (Protein CELTIX-1). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.007229 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
HRBL_HUMAN
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.014511 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95081, O75429, Q96AB9, Q96GL4 | Gene names | HRBL, RABR | |||
|
Domain Architecture |
|
|||||
| Description | HIV-1 Rev-binding protein-like protein (Rev/Rex activation domain- binding protein related) (RAB-R). | |||||
|
HRBL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.013194 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80WC7, Q8BKS5, Q99J67 | Gene names | Hrbl, Rabr | |||
|
Domain Architecture |
|
|||||
| Description | HIV-1 Rev-binding protein-like protein (Rev/Rex activation domain- binding protein related) (RAB-R). | |||||
|
LRRK1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | -0.000119 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1267 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UHC2, Q3U476, Q66JQ4, Q6GQR9, Q6NZF5, Q6ZPI4, Q8BKP3, Q8BU93, Q8BUY0, Q8BVV2, Q8R085 | Gene names | Lrrk1, Kiaa1790 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 1 (EC 2.7.11.1). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.005243 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
REST_HUMAN
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.002730 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
RRBP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.004103 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
TCOF_MOUSE
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.020920 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
EVI2B_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.012620 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VD58, Q3U1A3 | Gene names | Evi2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EVI2B protein precursor (Ecotropic viral integration site 2B protein). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.006446 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
TNF10_MOUSE
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.010485 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50592 | Gene names | Tnfsf10, Trail | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor ligand superfamily member 10 (TNF-related apoptosis-inducing ligand) (Protein TRAIL) (CD253 antigen). | |||||
|
WAPL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.010051 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
MLF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.018378 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QWV4, O70217 | Gene names | Mlf1, Hls7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myeloid leukemia factor 1 (Myelodysplasia-myeloid leukemia factor 1) (Hematopoietic lineage switch 7). | |||||
|
RABE2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.004589 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 545 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H5N1 | Gene names | RABEP2, RABPT5B | |||
|
Domain Architecture |
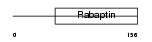
|
|||||
| Description | Rab GTPase-binding effector protein 2 (Rabaptin-5beta). | |||||
|
SERA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.015881 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43175, Q5SZU3, Q9BQ01 | Gene names | PHGDH, PGDH3 | |||
|
Domain Architecture |
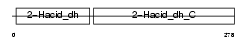
|
|||||
| Description | D-3-phosphoglycerate dehydrogenase (EC 1.1.1.95) (3-PGDH). | |||||
|
ZAN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.006481 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y493, O00218, Q96L85, Q96L86, Q96L87, Q96L88, Q96L89, Q96L90, Q9BXN9, Q9BZ83, Q9BZ84, Q9BZ85, Q9BZ86, Q9BZ87, Q9BZ88 | Gene names | ZAN | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
HBS1L_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9Y450, Q5T7G3, Q8NDW9, Q9UPW3 | Gene names | HBS1L, HBS1, KIAA1038 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HBS1-like protein (ERFS). | |||||
|
HBS1L_MOUSE
|
||||||
| NC score | 0.992304 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q69ZS7, Q91Z32, Q9CVT2, Q9CZ95, Q9WTY5 | Gene names | Hbs1l, Hbs1, Kiaa1038 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HBS1-like protein. | |||||
|
GSPT1_HUMAN
|
||||||
| NC score | 0.951068 (rank : 3) | θ value | 5.14231e-88 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P15170 | Gene names | GSPT1 | |||
|
Domain Architecture |

|
|||||
| Description | G1 to S phase transition protein 1 homolog (GTP-binding protein GST1- HS). | |||||
|
GSPT1_MOUSE
|
||||||
| NC score | 0.950023 (rank : 4) | θ value | 3.93732e-88 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8R050, O88179 | Gene names | Gspt1 | |||
|
Domain Architecture |

|
|||||
| Description | G1 to S phase transition protein 1 homolog. | |||||
|
EF1A2_MOUSE
|
||||||
| NC score | 0.942369 (rank : 5) | θ value | 1.49618e-87 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P62631, P27706 | Gene names | Eef1a2, Eef1al, Stn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor 1-alpha 2 (EF-1-alpha-2) (Elongation factor 1 A-2) (eEF1A-2) (Statin S1). | |||||
|
EF1A2_HUMAN
|
||||||
| NC score | 0.942358 (rank : 6) | θ value | 1.49618e-87 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q05639, P54266 | Gene names | EEF1A2, EEF1AL, STN | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 1-alpha 2 (EF-1-alpha-2) (Elongation factor 1 A-2) (eEF1A-2) (Statin S1). | |||||
|
EF1A1_MOUSE
|
||||||
| NC score | 0.942355 (rank : 7) | θ value | 1.40038e-85 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10126, Q61511, Q6ZWN2, Q8BMB8, Q8BVS8, Q99KU5 | Gene names | Eef1a1, Eef1a | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 1-alpha 1 (EF-1-alpha-1) (Elongation factor 1 A-1) (eEF1A-1) (Elongation factor Tu) (EF-Tu). | |||||
|
EF1A1_HUMAN
|
||||||
| NC score | 0.942232 (rank : 8) | θ value | 1.40038e-85 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P68104, P04719, P04720 | Gene names | EEF1A1, EEF1A, EF1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor 1-alpha 1 (EF-1-alpha-1) (Elongation factor 1 A-1) (eEF1A-1) (Elongation factor Tu) (EF-Tu). | |||||
|
EFTU_MOUSE
|
||||||
| NC score | 0.831674 (rank : 9) | θ value | 4.89182e-30 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BFR5, Q6P919 | Gene names | Tufm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor Tu, mitochondrial precursor. | |||||
|
EFTU_HUMAN
|
||||||
| NC score | 0.830649 (rank : 10) | θ value | 1.4233e-29 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P49411, O15276 | Gene names | TUFM | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor Tu, mitochondrial precursor (EF-Tu) (P43). | |||||
|
SELB_MOUSE
|
||||||
| NC score | 0.630301 (rank : 11) | θ value | 2.0648e-12 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JHW4 | Gene names | Eefsec, Selb | |||
|
Domain Architecture |

|
|||||
| Description | Selenocysteine-specific elongation factor (Elongation factor sec) (Eukaryotic elongation factor, selenocysteine-tRNA-specific) (mSelB). | |||||
|
SELB_HUMAN
|
||||||
| NC score | 0.610842 (rank : 12) | θ value | 1.133e-10 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P57772 | Gene names | EEFSEC, SELB | |||
|
Domain Architecture |

|
|||||
| Description | Selenocysteine-specific elongation factor (Elongation factor sec) (Eukaryotic elongation factor, selenocysteine-tRNA-specific). | |||||
|
GTPB1_HUMAN
|
||||||
| NC score | 0.603286 (rank : 13) | θ value | 6.41864e-14 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O00178, Q6IC67 | Gene names | GTPBP1 | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein 1 (G-protein 1) (GP-1) (GP1). | |||||
|
GTPB1_MOUSE
|
||||||
| NC score | 0.599635 (rank : 14) | θ value | 1.86753e-13 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O08582, Q3UD96, Q3UGW6, Q545R1, Q80ZY5 | Gene names | Gtpbp1, Gtpbp | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein 1 (G-protein 1) (GP-1) (GP1). | |||||
|
GTPB2_HUMAN
|
||||||
| NC score | 0.554057 (rank : 15) | θ value | 7.09661e-13 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BX10, Q5T7E8, Q8ND84, Q8TAH7, Q8WUA5, Q9HCS9, Q9NRU4, Q9NX60 | Gene names | GTPBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTP-binding protein 2. | |||||
|
GTPB2_MOUSE
|
||||||
| NC score | 0.554038 (rank : 16) | θ value | 7.09661e-13 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q3UJK4, Q9EST7, Q9JIX7 | Gene names | Gtpbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTP-binding protein 2 (GTP-binding-like protein 2). | |||||
|
IF2M_HUMAN
|
||||||
| NC score | 0.444049 (rank : 17) | θ value | 3.08544e-08 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P46199 | Gene names | MTIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Translation initiation factor IF-2, mitochondrial precursor (IF-2Mt) (IF-2(Mt)). | |||||
|
EFG1_HUMAN
|
||||||
| NC score | 0.382733 (rank : 18) | θ value | 4.92598e-06 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96RP9, Q96T39 | Gene names | GFM1, EFG, EFG1, GFM | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 1, mitochondrial precursor (mEF-G 1) (Elongation factor G1). | |||||
|
EFG2_HUMAN
|
||||||
| NC score | 0.380663 (rank : 19) | θ value | 2.44474e-05 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q969S9, Q8WYI0 | Gene names | GFM2, EFG2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 2, mitochondrial precursor (mEF-G 2) (Elongation factor G2). | |||||
|
EFG1_MOUSE
|
||||||
| NC score | 0.370854 (rank : 20) | θ value | 1.43324e-05 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8K0D5, Q921D6, Q924I0 | Gene names | Gfm1, Efg, Efg1, Gfm | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 1, mitochondrial precursor (mEF-G 1) (Elongation factor G1). | |||||
|
EFG2_MOUSE
|
||||||
| NC score | 0.368213 (rank : 21) | θ value | 0.000121331 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8R2Q4 | Gene names | Gfm2, Efg2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 2, mitochondrial precursor (mEF-G 2) (Elongation factor G2). | |||||
|
IF2M_MOUSE
|
||||||
| NC score | 0.363672 (rank : 22) | θ value | 0.00298849 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q91YJ5 | Gene names | Mtif2 | |||
|
Domain Architecture |

|
|||||
| Description | Translation initiation factor IF-2, mitochondrial precursor (IF-2Mt) (IF-2(Mt)). | |||||
|
IF2H_MOUSE
|
||||||
| NC score | 0.354572 (rank : 23) | θ value | 0.00390308 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Z0N2 | Gene names | Eif2s3y | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3, Y-linked (Eukaryotic translation initiation factor 2 subunit gamma, Y-linked) (eIF-2-gamma Y). | |||||
|
IF2G_HUMAN
|
||||||
| NC score | 0.352699 (rank : 24) | θ value | 0.00390308 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P41091 | Gene names | EIF2S3, EIF2G | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3 (Eukaryotic translation initiation factor 2 subunit gamma) (eIF-2-gamma). | |||||
|
IF2G_MOUSE
|
||||||
| NC score | 0.352690 (rank : 25) | θ value | 0.00390308 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z0N1 | Gene names | Eif2s3x | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3, X-linked (Eukaryotic translation initiation factor 2 subunit gamma, X-linked) (eIF-2-gamma X). | |||||
|
EF2_MOUSE
|
||||||
| NC score | 0.251460 (rank : 26) | θ value | 0.0961366 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P58252, Q544E4 | Gene names | Eef2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 2 (EF-2). | |||||
|
EF2_HUMAN
|
||||||
| NC score | 0.249246 (rank : 27) | θ value | 0.0961366 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P13639 | Gene names | EEF2, EF2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 2 (EF-2). | |||||
|
U5S1_HUMAN
|
||||||
| NC score | 0.208099 (rank : 28) | θ value | 0.00665767 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15029, Q9BUR0 | Gene names | EFTUD2, KIAA0031, SNRP116 | |||
|
Domain Architecture |

|
|||||
| Description | 116 kDa U5 small nuclear ribonucleoprotein component (U5 snRNP- specific protein, 116 kDa) (U5-116 kDa) (Elongation factor Tu GTP- binding domain protein 2) (hSNU114). | |||||
|
U5S1_MOUSE
|
||||||
| NC score | 0.208022 (rank : 29) | θ value | 0.00665767 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O08810 | Gene names | Eftud2, Snrp116 | |||
|
Domain Architecture |

|
|||||
| Description | 116 kDa U5 small nuclear ribonucleoprotein component (U5 snRNP- specific protein, 116 kDa) (U5-116 kDa) (Elongation factor Tu GTP- binding domain protein 2). | |||||
|
IF2P_HUMAN
|
||||||
| NC score | 0.145802 (rank : 30) | θ value | 0.0330416 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
SYBU_HUMAN
|
||||||
| NC score | 0.056484 (rank : 31) | θ value | 0.0252991 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NX95, Q5R1T1, Q5R1T2, Q5R1T3, Q5Y2M6, Q8ND49, Q8TCR6, Q9P256 | Gene names | SYBU, GOLSYN, KIAA1472 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntabulin (Syntaxin-1-binding protein) (Golgi-localized syntaphilin- related protein). | |||||
|
SYBU_MOUSE
|
||||||
| NC score | 0.048394 (rank : 32) | θ value | 0.279714 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BHS8, Q3TQN7, Q401F3, Q401F4, Q571C1, Q6P1J2, Q80XH0, Q8BHS7, Q8BI27 | Gene names | Sybu, Golsyn, Kiaa1472 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntabulin (Syntaxin-1-binding protein) (Golgi-localized syntaphilin- related protein) (m-Golsyn). | |||||
|
BAIP2_HUMAN
|
||||||
| NC score | 0.046124 (rank : 33) | θ value | 0.813845 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UQB8, O43858, Q53HB1, Q86WC1, Q8N5C0, Q96CR7, Q9UBR3, Q9UQ43 | Gene names | BAIAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain-specific angiogenesis inhibitor 1-associated protein 2 (BAI1- associated protein 2) (BAI-associated protein 2) (Protein BAP2) (Insulin receptor substrate p53) (IRSp53) (Insulin receptor substrate protein of 53 kDa) (Insulin receptor substrate p53/p58) (IRSp53/58) (IRS-58) (Fas ligand-associated factor 3) (FLAF3). | |||||
|
BAIP2_MOUSE
|
||||||
| NC score | 0.043601 (rank : 34) | θ value | 0.62314 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BKX1, Q91V97, Q923V9, Q923W0 | Gene names | Baiap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain-specific angiogenesis inhibitor 1-associated protein 2 (BAI1- associated protein 2) (BAI-associated protein 2) (Insulin receptor substrate p53) (IRSp53) (Insulin receptor substrate protein of 53 kDa) (Insulin receptor tyrosine kinase 53 kDa substrate). | |||||
|
NNP1_MOUSE
|
||||||
| NC score | 0.042247 (rank : 35) | θ value | 0.0736092 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56183, O35712, Q9ERE1, Q9JI07, Q9JK67, Q9JKU2 | Gene names | Nnp1 | |||
|
Domain Architecture |

|
|||||
| Description | NNP-1 protein (Novel nuclear protein 1) (Nop52). | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.038426 (rank : 36) | θ value | 0.0113563 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
MUC13_MOUSE
|
||||||
| NC score | 0.037862 (rank : 37) | θ value | 0.0330416 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.035282 (rank : 38) | θ value | 0.62314 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
TCOF_HUMAN
|
||||||
| NC score | 0.028421 (rank : 39) | θ value | 0.62314 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
MLF1_HUMAN
|
||||||
| NC score | 0.027205 (rank : 40) | θ value | 1.38821 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P58340 | Gene names | MLF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myeloid leukemia factor 1 (Myelodysplasia-myeloid leukemia factor 1). | |||||
|
TM131_MOUSE
|
||||||
| NC score | 0.022611 (rank : 41) | θ value | 2.36792 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O70472 | Gene names | Tmem131, D1Bwg0491e, Rw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 131 (Protein RW1). | |||||
|
P66A_MOUSE
|
||||||
| NC score | 0.021471 (rank : 42) | θ value | 1.38821 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CHY6, Q8BTQ2, Q8VEC9 | Gene names | Gatad2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional repressor p66 alpha (GATA zinc finger domain- containing protein 2A). | |||||
|
TCOF_MOUSE
|
||||||
| NC score | 0.020920 (rank : 43) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
MLF1_MOUSE
|
||||||
| NC score | 0.018378 (rank : 44) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QWV4, O70217 | Gene names | Mlf1, Hls7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myeloid leukemia factor 1 (Myelodysplasia-myeloid leukemia factor 1) (Hematopoietic lineage switch 7). | |||||
|
CCD82_MOUSE
|
||||||
| NC score | 0.017950 (rank : 45) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6PG04, Q3V462 | Gene names | Ccdc82 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 82. | |||||
|
BCL7A_MOUSE
|
||||||
| NC score | 0.017514 (rank : 46) | θ value | 2.36792 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CXE2, Q8C361, Q8C8M8, Q8VD15 | Gene names | Bcl7a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell CLL/lymphoma 7 protein family member A. | |||||
|
H10_HUMAN
|
||||||
| NC score | 0.016638 (rank : 47) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P07305 | Gene names | H1F0, H1FV | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.0 (H1(0)) (Histone H1'). | |||||
|
SERA_HUMAN
|
||||||
| NC score | 0.015881 (rank : 48) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43175, Q5SZU3, Q9BQ01 | Gene names | PHGDH, PGDH3 | |||
|
Domain Architecture |

|
|||||
| Description | D-3-phosphoglycerate dehydrogenase (EC 1.1.1.95) (3-PGDH). | |||||
|
BRE1B_HUMAN
|
||||||
| NC score | 0.015812 (rank : 49) | θ value | 0.0736092 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75150, Q6AHZ6, Q6N005, Q7L3T6, Q8N615, Q96T18, Q9BSV9, Q9HC82 | Gene names | RNF40, BRE1B, KIAA0661 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1B (EC 6.3.2.-) (BRE1-B) (RING finger protein 40) (95 kDa retinoblastoma-associated protein) (RBP95). | |||||
|
HRBL_HUMAN
|
||||||
| NC score | 0.014511 (rank : 50) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95081, O75429, Q96AB9, Q96GL4 | Gene names | HRBL, RABR | |||
|
Domain Architecture |

|
|||||
| Description | HIV-1 Rev-binding protein-like protein (Rev/Rex activation domain- binding protein related) (RAB-R). | |||||
|
HRBL_MOUSE
|
||||||
| NC score | 0.013194 (rank : 51) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80WC7, Q8BKS5, Q99J67 | Gene names | Hrbl, Rabr | |||
|
Domain Architecture |

|
|||||
| Description | HIV-1 Rev-binding protein-like protein (Rev/Rex activation domain- binding protein related) (RAB-R). | |||||
|
BRCA2_HUMAN
|
||||||
| NC score | 0.012844 (rank : 52) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51587, O00183, O15008, Q13879 | Gene names | BRCA2, FANCD1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein (Fanconi anemia group D1 protein). | |||||
|
EVI2B_MOUSE
|
||||||
| NC score | 0.012620 (rank : 53) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VD58, Q3U1A3 | Gene names | Evi2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EVI2B protein precursor (Ecotropic viral integration site 2B protein). | |||||
|
ELL_MOUSE
|
||||||
| NC score | 0.012093 (rank : 54) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O08856 | Gene names | Ell | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL (Eleven-nineteen lysine-rich leukemia protein). | |||||
|
LIMA1_MOUSE
|
||||||
| NC score | 0.011899 (rank : 55) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9ERG0, Q9ERG1 | Gene names | Lima1, D15Ertd366e, Eplin | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm) (mEPLIN). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.010713 (rank : 56) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
TNF10_MOUSE
|
||||||
| NC score | 0.010485 (rank : 57) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50592 | Gene names | Tnfsf10, Trail | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor ligand superfamily member 10 (TNF-related apoptosis-inducing ligand) (Protein TRAIL) (CD253 antigen). | |||||
|
WAPL_MOUSE
|
||||||
| NC score | 0.010051 (rank : 58) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
BRD7_HUMAN
|
||||||
| NC score | 0.009435 (rank : 59) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NPI1, Q4VC09, Q8N2L9, Q96KA4, Q9BV48, Q9UH59 | Gene names | BRD7, BP75, CELTIX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 7 (75 kDa bromodomain protein) (Protein CELTIX-1). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.007229 (rank : 60) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
ZAN_HUMAN
|
||||||
| NC score | 0.006481 (rank : 61) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y493, O00218, Q96L85, Q96L86, Q96L87, Q96L88, Q96L89, Q96L90, Q9BXN9, Q9BZ83, Q9BZ84, Q9BZ85, Q9BZ86, Q9BZ87, Q9BZ88 | Gene names | ZAN | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.006446 (rank : 62) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
SFRS4_MOUSE
|
||||||
| NC score | 0.005576 (rank : 63) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.005243 (rank : 64) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
RABE2_HUMAN
|
||||||
| NC score | 0.004589 (rank : 65) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 545 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H5N1 | Gene names | RABEP2, RABPT5B | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 2 (Rabaptin-5beta). | |||||
|
RRBP1_HUMAN
|
||||||
| NC score | 0.004103 (rank : 66) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
REST_HUMAN
|
||||||
| NC score | 0.002730 (rank : 67) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
LRRK1_HUMAN
|
||||||
| NC score | 0.000051 (rank : 68) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q38SD2, Q6NVH5, Q6NYC0, Q6ZNL9, Q6ZNM9, Q96JN5, Q9H5S3 | Gene names | LRRK1, KIAA1790 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 1 (EC 2.7.11.1). | |||||
|
LRRK1_MOUSE
|
||||||
| NC score | -0.000119 (rank : 69) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1267 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UHC2, Q3U476, Q66JQ4, Q6GQR9, Q6NZF5, Q6ZPI4, Q8BKP3, Q8BU93, Q8BUY0, Q8BVV2, Q8R085 | Gene names | Lrrk1, Kiaa1790 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 1 (EC 2.7.11.1). | |||||