Please be patient as the page loads
|
FOXO4_HUMAN
|
||||||
| SwissProt Accessions | P98177, O43821, Q13720, Q3KPF1, Q8TDK9 | Gene names | MLLT7, AFX, AFX1, FOXO4 | |||
|
Domain Architecture |
|
|||||
| Description | Fork head domain transcription factor AFX1 (Forkhead box protein O4). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
FOXO4_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 197 | |
| SwissProt Accessions | P98177, O43821, Q13720, Q3KPF1, Q8TDK9 | Gene names | MLLT7, AFX, AFX1, FOXO4 | |||
|
Domain Architecture |

|
|||||
| Description | Fork head domain transcription factor AFX1 (Forkhead box protein O4). | |||||
|
FOXO4_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.988424 (rank : 2) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 111 | |
| SwissProt Accessions | Q9WVH3 | Gene names | Mllt7, Afx, Afx1, Fkhr3, Foxo4 | |||
|
Domain Architecture |
|
|||||
| Description | Fork head domain transcription factor AFX1 (Afxh) (Forkhead box protein O4). | |||||
|
FOXO3_HUMAN
|
||||||
| θ value | 4.99229e-75 (rank : 3) | NC score | 0.964746 (rank : 3) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | O43524, O15171, Q5T2I7, Q9BZ04 | Gene names | FOXO3A, FKHRL1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein O3A (Forkhead in rhabdomyosarcoma-like 1) (AF6q21 protein). | |||||
|
FOXO1_HUMAN
|
||||||
| θ value | 1.96316e-71 (rank : 4) | NC score | 0.954542 (rank : 5) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q12778, O43523, Q5VYC7, Q6NSK6 | Gene names | FOXO1A, FKHR | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein O1A (Forkhead in rhabdomyosarcoma). | |||||
|
FOXO1_MOUSE
|
||||||
| θ value | 4.37348e-71 (rank : 5) | NC score | 0.955560 (rank : 4) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q9R1E0 | Gene names | Foxo1a, Fkhr, Foxo1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein O1A (Forkhead in rhabdomyosarcoma). | |||||
|
FOXGB_HUMAN
|
||||||
| θ value | 7.32683e-18 (rank : 6) | NC score | 0.866174 (rank : 8) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | P55315 | Gene names | FOXG1B, FKHL1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1B (Forkhead-related protein FKHL1) (Transcription factor BF-1) (Brain factor 1) (BF1) (HFK1). | |||||
|
FOXGB_MOUSE
|
||||||
| θ value | 8.10077e-17 (rank : 7) | NC score | 0.866206 (rank : 7) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q60987, Q80VP3 | Gene names | Foxg1b, Fkhl1, Foxg1, Hfhbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1B (Forkhead-related protein FKHL1) (Transcription factor BF-1) (Brain factor 1) (BF1). | |||||
|
FOXJ3_HUMAN
|
||||||
| θ value | 8.10077e-17 (rank : 8) | NC score | 0.860605 (rank : 14) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q9UPW0, Q9NSS7 | Gene names | FOXJ3, KIAA1041 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein J3. | |||||
|
FOXJ3_MOUSE
|
||||||
| θ value | 8.10077e-17 (rank : 9) | NC score | 0.860570 (rank : 15) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q8BUR3, Q3UTD8 | Gene names | Foxj3, Kiaa1041 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein J3. | |||||
|
FOXC2_HUMAN
|
||||||
| θ value | 1.38178e-16 (rank : 10) | NC score | 0.845335 (rank : 46) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q99958 | Gene names | FOXC2, FKHL14, MFH1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
FOXI2_MOUSE
|
||||||
| θ value | 1.38178e-16 (rank : 11) | NC score | 0.869316 (rank : 6) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q3I5G5, Q8BIK9 | Gene names | Foxi2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein I2. | |||||
|
FOXD2_HUMAN
|
||||||
| θ value | 1.80466e-16 (rank : 12) | NC score | 0.858248 (rank : 17) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | O60548 | Gene names | FOXD2, FKHL17, FREAC9 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D2 (Forkhead-related protein FKHL17) (Forkhead- related transcription factor 9) (FREAC-9). | |||||
|
FOXI1_MOUSE
|
||||||
| θ value | 1.80466e-16 (rank : 13) | NC score | 0.861266 (rank : 12) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | Q922I5, Q5SRI5, Q9D299 | Gene names | Foxi1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein I1. | |||||
|
FOXD2_MOUSE
|
||||||
| θ value | 5.25075e-16 (rank : 14) | NC score | 0.856955 (rank : 23) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | O35392 | Gene names | Foxd2, Mf2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D2 (Mesoderm/mesenchyme forkhead 2) (MF-2). | |||||
|
FOXI1_HUMAN
|
||||||
| θ value | 8.95645e-16 (rank : 15) | NC score | 0.861119 (rank : 13) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q12951, Q14518, Q66SR7, Q8N6L8 | Gene names | FOXI1, FKHL10, FREAC6 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein I1 (Forkhead-related protein FKHL10) (Forkhead- related transcription factor 6) (FREAC-6) (Hepatocyte nuclear factor 3 forkhead homolog 3) (HNF-3/fork-head homolog 3) (HFH-3). | |||||
|
FOXGA_HUMAN
|
||||||
| θ value | 1.99529e-15 (rank : 16) | NC score | 0.862851 (rank : 11) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | P55316 | Gene names | FOXG1A, FKHL2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1A (Forkhead-related protein FKHL2) (Transcription factor BF-2) (Brain factor 2) (BF2) (HFK2). | |||||
|
FOXQ1_MOUSE
|
||||||
| θ value | 1.99529e-15 (rank : 17) | NC score | 0.865644 (rank : 9) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | O70220 | Gene names | Foxq1, Hfh1, Hfh1l | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein Q1 (Hepatocyte nuclear factor 3 forkhead homolog 1) (HNF-3/forkhead-like protein 1) (HFH-1) (HFH-1l). | |||||
|
FOXJ2_HUMAN
|
||||||
| θ value | 2.60593e-15 (rank : 18) | NC score | 0.853630 (rank : 26) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9P0K8, Q96PS9, Q9NSN5 | Gene names | FOXJ2, FHX | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein J2 (Fork head homologous X). | |||||
|
FOXJ2_MOUSE
|
||||||
| θ value | 2.60593e-15 (rank : 19) | NC score | 0.858673 (rank : 16) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | Q9ES18 | Gene names | Foxj2, Fhx | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein J2 (Fork head homologous X). | |||||
|
FOXQ1_HUMAN
|
||||||
| θ value | 2.60593e-15 (rank : 20) | NC score | 0.863482 (rank : 10) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q9C009, Q9NS06 | Gene names | FOXQ1, HFH1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein Q1 (Hepatocyte nuclear factor 3 forkhead homolog 1) (HNF-3/forkhead-like protein 1) (HFH-1). | |||||
|
FOXC1_HUMAN
|
||||||
| θ value | 7.58209e-15 (rank : 21) | NC score | 0.845183 (rank : 47) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q12948, Q86UP7, Q9BYM1, Q9NUE5, Q9UDD0, Q9UP06 | Gene names | FOXC1, FKHL7, FREAC3 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein C1 (Forkhead-related protein FKHL7) (Forkhead- related transcription factor 3) (FREAC-3). | |||||
|
FOXC1_MOUSE
|
||||||
| θ value | 7.58209e-15 (rank : 22) | NC score | 0.846344 (rank : 44) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 93 | |
| SwissProt Accessions | Q61572, O88409, Q61582 | Gene names | Foxc1, Fkh1, Fkhl7, Freac3, Mf1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein C1 (Forkhead-related protein FKHL7) (Forkhead- related transcription factor 3) (FREAC-3) (Transcription factor FKH-1) (Mesoderm/mesenchyme forkhead 1) (MF-1). | |||||
|
HNF3G_HUMAN
|
||||||
| θ value | 9.90251e-15 (rank : 23) | NC score | 0.853232 (rank : 27) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | P55318, Q9UMW9 | Gene names | FOXA3, HNF3G, TCF3G | |||
|
Domain Architecture |
|
|||||
| Description | Hepatocyte nuclear factor 3-gamma (HNF-3G) (Forkhead box protein A3) (Fork head-related protein FKH H3). | |||||
|
FOXE1_HUMAN
|
||||||
| θ value | 1.29331e-14 (rank : 24) | NC score | 0.850849 (rank : 32) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | O00358, O75765 | Gene names | FOXE1, FKHL15, TITF2, TTF2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein E1 (Thyroid transcription factor 2) (TTF-2) (Forkhead-related protein FKHL15). | |||||
|
FOXD1_HUMAN
|
||||||
| θ value | 1.68911e-14 (rank : 25) | NC score | 0.855192 (rank : 25) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q16676, Q12949 | Gene names | FOXD1, FKHL8, FREAC4 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D1 (Forkhead-related protein FKHL8) (Forkhead- related transcription factor 4) (FREAC-4). | |||||
|
FOXE1_MOUSE
|
||||||
| θ value | 1.68911e-14 (rank : 26) | NC score | 0.851860 (rank : 30) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q8R2I0 | Gene names | Foxe1, Titf2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein E1 (Thyroid transcription factor 2) (TTF-2). | |||||
|
FOXE3_MOUSE
|
||||||
| θ value | 2.20605e-14 (rank : 27) | NC score | 0.852012 (rank : 29) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q9QY14 | Gene names | Foxe3 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein E3. | |||||
|
FOXC2_MOUSE
|
||||||
| θ value | 2.88119e-14 (rank : 28) | NC score | 0.850154 (rank : 35) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q61850, P97948, Q63869, Q8C694 | Gene names | Foxc2, Fkh14, Fkhl14, Mfh1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
FOXD1_MOUSE
|
||||||
| θ value | 2.88119e-14 (rank : 29) | NC score | 0.857331 (rank : 21) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q61345 | Gene names | Foxd1, Fkhl8, Freac4, Hfhbf2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D1 (Forkhead-related protein FKHL8) (Forkhead- related transcription factor 4) (FREAC-4) (Brain factor 2) (HFH-BF-2). | |||||
|
FOXK2_HUMAN
|
||||||
| θ value | 1.86753e-13 (rank : 30) | NC score | 0.852741 (rank : 28) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | Q01167, Q13622, Q13623, Q13624 | Gene names | FOXK2, ILF, ILF1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein K2 (Interleukin enhancer-binding factor 1) (Cellular transcription factor ILF-1). | |||||
|
FOXL1_HUMAN
|
||||||
| θ value | 1.86753e-13 (rank : 31) | NC score | 0.857073 (rank : 22) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q12952, Q9H242 | Gene names | FOXL1, FKHL11, FREAC7 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein L1 (Forkhead-related protein FKHL11) (Forkhead- related transcription factor 7) (FREAC-7). | |||||
|
FOXF2_HUMAN
|
||||||
| θ value | 2.43908e-13 (rank : 32) | NC score | 0.858154 (rank : 18) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q12947, Q9UQ85 | Gene names | FOXF2, FKHL6, FREAC2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein F2 (Forkhead-related protein FKHL6) (Forkhead- related transcription factor 2) (FREAC-2) (Forkhead-related activator 2). | |||||
|
FOXL2_MOUSE
|
||||||
| θ value | 2.43908e-13 (rank : 33) | NC score | 0.857683 (rank : 19) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | O88470, Q8K3X9 | Gene names | Foxl2, Pfrk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein L2 (Pituitary forkhead factor) (P-Frk). | |||||
|
FOXN3_HUMAN
|
||||||
| θ value | 5.43371e-13 (rank : 34) | NC score | 0.846767 (rank : 43) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | O00409, Q96II7, Q9UIE7 | Gene names | CHES1, FOXN3 | |||
|
Domain Architecture |
|
|||||
| Description | Checkpoint suppressor 1 (Forkhead box protein N3). | |||||
|
FOXN3_MOUSE
|
||||||
| θ value | 5.43371e-13 (rank : 35) | NC score | 0.848898 (rank : 38) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q499D0, Q8C5N8, Q9CU83 | Gene names | Ches1, Foxn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Checkpoint suppressor 1 (Forkhead box protein N3). | |||||
|
HNF3A_MOUSE
|
||||||
| θ value | 5.43371e-13 (rank : 36) | NC score | 0.843790 (rank : 50) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | P35582, Q61108 | Gene names | Foxa1, Hnf3a, Tcf-3a, Tcf3a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-alpha (HNF-3A) (Forkhead box protein A1). | |||||
|
FOXB1_MOUSE
|
||||||
| θ value | 7.09661e-13 (rank : 37) | NC score | 0.844982 (rank : 48) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q64732, O08753 | Gene names | Foxb1, Fkh5, Foxb1a, Foxb1b, Mf3 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein B1 (Transcription factor FKH-5). | |||||
|
FOXD3_HUMAN
|
||||||
| θ value | 7.09661e-13 (rank : 38) | NC score | 0.849561 (rank : 36) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9UJU5, Q9BYM2, Q9UDD1 | Gene names | FOXD3 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D3 (HNF3/FH transcription factor genesis). | |||||
|
FOXD3_MOUSE
|
||||||
| θ value | 7.09661e-13 (rank : 39) | NC score | 0.846779 (rank : 42) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q61060 | Gene names | Foxd3, Hfh2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D3 (HNF3/FH transcription factor genesis) (Hepatocyte nuclear factor 3 forkhead homolog 2) (HFH-2). | |||||
|
HNF3A_HUMAN
|
||||||
| θ value | 7.09661e-13 (rank : 40) | NC score | 0.841621 (rank : 54) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | P55317, Q9H2A0 | Gene names | FOXA1, HNF3A, TCF3A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-alpha (HNF-3A) (Forkhead box protein A1). | |||||
|
FOXF1_HUMAN
|
||||||
| θ value | 9.26847e-13 (rank : 41) | NC score | 0.849354 (rank : 37) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q12946, Q5FWE5 | Gene names | FOXF1, FKHL5, FREAC1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein F1 (Forkhead-related protein FKHL5) (Forkhead- related transcription factor 1) (FREAC-1) (Forkhead-related activator 1). | |||||
|
FOXM1_HUMAN
|
||||||
| θ value | 9.26847e-13 (rank : 42) | NC score | 0.825369 (rank : 62) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q08050, O43258, O43259, O43260, Q4ZGG7, Q9BRL2 | Gene names | FOXM1, FKHL16, HFH11, MPP2, WIN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead-related protein FKHL16) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/fork-head homolog 11) (HFH-11) (Winged helix factor from INS-1 cells) (M-phase phosphoprotein 2) (MPM-2 reactive phosphoprotein 2) (Transcription factor Trident). | |||||
|
FOXM1_MOUSE
|
||||||
| θ value | 9.26847e-13 (rank : 43) | NC score | 0.832644 (rank : 57) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | O08696 | Gene names | Foxm1, Fkh16 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead homolog 16) (Winged-helix transcription factor Trident). | |||||
|
HNF3G_MOUSE
|
||||||
| θ value | 9.26847e-13 (rank : 44) | NC score | 0.850712 (rank : 33) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | P35584 | Gene names | Foxa3, Hnf3g, Tcf-3g, Tcf3g | |||
|
Domain Architecture |
|
|||||
| Description | Hepatocyte nuclear factor 3-gamma (HNF-3G) (Forkhead box protein A3). | |||||
|
FOXF1_MOUSE
|
||||||
| θ value | 1.2105e-12 (rank : 45) | NC score | 0.850438 (rank : 34) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q61080, Q61661 | Gene names | Foxf1, Fkhl5, Foxf1a, Freac1, Hfh8 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein F1 (Forkhead-related protein FKHL5) (Forkhead- related transcription factor 1) (FREAC-1) (Hepatocyte nuclear factor 3 forkhead homolog 8) (HFH-8). | |||||
|
FOXJ1_MOUSE
|
||||||
| θ value | 1.2105e-12 (rank : 46) | NC score | 0.846315 (rank : 45) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q61660 | Gene names | Foxj1, Fkhl13, Hfh4 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein J1 (Forkhead-related protein FKHL13) (Hepatocyte nuclear factor 3 forkhead homolog 4) (HFH-4). | |||||
|
FOXK1_MOUSE
|
||||||
| θ value | 1.2105e-12 (rank : 47) | NC score | 0.847232 (rank : 41) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | P42128, O35939 | Gene names | Foxk1, Mnf | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein K1 (Myocyte nuclear factor) (MNF). | |||||
|
FOXL1_MOUSE
|
||||||
| θ value | 1.58096e-12 (rank : 48) | NC score | 0.851658 (rank : 31) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q64731 | Gene names | Foxl1, Fkh6, Fkhl11 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein L1 (Forkhead-related protein FKHL11) (Transcription factor FKH-6). | |||||
|
FOXL2_HUMAN
|
||||||
| θ value | 1.58096e-12 (rank : 49) | NC score | 0.857590 (rank : 20) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P58012, Q4ZGJ3 | Gene names | FOXL2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein L2. | |||||
|
FOXP4_HUMAN
|
||||||
| θ value | 1.58096e-12 (rank : 50) | NC score | 0.788320 (rank : 73) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q8IVH2, Q96E19 | Gene names | FOXP4, FKHLA | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P4 (Fork head-related protein-like A). | |||||
|
HNF3B_MOUSE
|
||||||
| θ value | 1.58096e-12 (rank : 51) | NC score | 0.844699 (rank : 49) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | P35583, Q60602 | Gene names | Foxa2, Hnf3b, Tcf-3b, Tcf3b | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-beta (HNF-3B) (Forkhead box protein A2). | |||||
|
FOXE3_HUMAN
|
||||||
| θ value | 2.0648e-12 (rank : 52) | NC score | 0.848804 (rank : 39) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q13461, Q5SVY9, Q9NQV9 | Gene names | FOXE3, FKHL12, FREAC8 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein E3 (Forkhead-related protein FKHL12) (Forkhead- related transcription factor 8) (FREAC-8). | |||||
|
FOXJ1_HUMAN
|
||||||
| θ value | 2.0648e-12 (rank : 53) | NC score | 0.848717 (rank : 40) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q92949, O00630 | Gene names | FOXJ1, FKHL13, HFH4 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein J1 (Forkhead-related protein FKHL13) (Hepatocyte nuclear factor 3 forkhead homolog 4) (HFH-4). | |||||
|
FOXP1_MOUSE
|
||||||
| θ value | 2.69671e-12 (rank : 54) | NC score | 0.782322 (rank : 74) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | P58462 | Gene names | Foxp1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1 (Forkhead-related transcription factor 1). | |||||
|
FOXP1_HUMAN
|
||||||
| θ value | 4.59992e-12 (rank : 55) | NC score | 0.790664 (rank : 71) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9H334, Q9H332, Q9H333, Q9P0R1 | Gene names | FOXP1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1. | |||||
|
FOXB1_HUMAN
|
||||||
| θ value | 6.00763e-12 (rank : 56) | NC score | 0.842181 (rank : 53) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q99853, O60652, O75917 | Gene names | FOXB1, FKH5 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein B1 (Transcription factor FKH-5). | |||||
|
FOXN2_HUMAN
|
||||||
| θ value | 6.00763e-12 (rank : 57) | NC score | 0.856095 (rank : 24) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | P32314, Q15769 | Gene names | FOXN2, HTLF | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein N2 (Human T-cell leukemia virus enhancer factor). | |||||
|
HNF3B_HUMAN
|
||||||
| θ value | 6.00763e-12 (rank : 58) | NC score | 0.843664 (rank : 51) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9Y261, Q8WUW4, Q96DF7 | Gene names | FOXA2, HNF3B, TCF3B | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-beta (HNF-3B) (Forkhead box protein A2). | |||||
|
FOXP3_HUMAN
|
||||||
| θ value | 1.02475e-11 (rank : 59) | NC score | 0.817908 (rank : 63) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q9BZS1, O60827, Q4ZH51 | Gene names | FOXP3, IPEX | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P3 (Scurfin). | |||||
|
FOXP4_MOUSE
|
||||||
| θ value | 1.33837e-11 (rank : 60) | NC score | 0.788447 (rank : 72) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | Q9DBY0, Q80V92, Q8CG10, Q8CIS1 | Gene names | Foxp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein P4 (Fork head-related protein like A) (mFKHLA). | |||||
|
FREA_HUMAN
|
||||||
| θ value | 1.74796e-11 (rank : 61) | NC score | 0.838583 (rank : 56) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 93 | |
| SwissProt Accessions | O43638, Q96D28 | Gene names | FKHL18, FREAC10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead-related transcription factor 10 (FREAC-10) (Forkhead-like 18 protein). | |||||
|
FOXP2_HUMAN
|
||||||
| θ value | 3.89403e-11 (rank : 62) | NC score | 0.748795 (rank : 78) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | O15409, Q6ZND1, Q75MJ3, Q8IZE0, Q8N0W2, Q8N6B7, Q8N6B8, Q8NFQ1, Q8NFQ2, Q8NFQ3, Q8NFQ4, Q8TD74 | Gene names | FOXP2, CAGH44, TNRC10 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P2 (CAG repeat protein 44) (Trinucleotide repeat- containing gene 10 protein). | |||||
|
FOXP2_MOUSE
|
||||||
| θ value | 3.89403e-11 (rank : 63) | NC score | 0.746597 (rank : 79) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | P58463, Q6PD37, Q8C4F0, Q8R441 | Gene names | Foxp2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P2. | |||||
|
FOXN1_MOUSE
|
||||||
| θ value | 6.64225e-11 (rank : 64) | NC score | 0.791723 (rank : 70) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 100 | |
| SwissProt Accessions | Q61575 | Gene names | Foxn1, Fkh19, Hfh11, Whn | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N1 (Transcription factor winged-helix nude) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/forkhead homolog 11) (HFH-11). | |||||
|
FOXP3_MOUSE
|
||||||
| θ value | 6.64225e-11 (rank : 65) | NC score | 0.813948 (rank : 64) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q99JB6 | Gene names | Foxp3 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P3 (Scurfin). | |||||
|
FOXD4_HUMAN
|
||||||
| θ value | 1.47974e-10 (rank : 66) | NC score | 0.829479 (rank : 59) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q12950, Q5VVK1, Q8WXT6 | Gene names | FOXD4, FKHL9, FOXD4A, FREAC5 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D4 (Forkhead-related protein FKHL9) (Forkhead- related transcription factor 5) (FREAC-5) (Myeloid factor-alpha). | |||||
|
FREA_MOUSE
|
||||||
| θ value | 1.47974e-10 (rank : 67) | NC score | 0.840207 (rank : 55) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | Q61574, Q8C5N7 | Gene names | Fkhl18, Fkh3, Freac10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead-related transcription factor 10 (FREAC-10) (Forkhead-like 18 protein) (Transcription factor FKH-3). | |||||
|
FOXN1_HUMAN
|
||||||
| θ value | 2.52405e-10 (rank : 68) | NC score | 0.781881 (rank : 75) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 107 | |
| SwissProt Accessions | O15353, O15352 | Gene names | FOXN1, WHN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N1 (Transcription factor winged-helix nude). | |||||
|
FOXB2_MOUSE
|
||||||
| θ value | 3.29651e-10 (rank : 69) | NC score | 0.843258 (rank : 52) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q64733 | Gene names | Foxb2, Fkh4 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein B2 (Transcription factor FKH-4). | |||||
|
FX4L1_HUMAN
|
||||||
| θ value | 5.62301e-10 (rank : 70) | NC score | 0.829344 (rank : 61) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q9NU39 | Gene names | FOXD4L1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D4-like 1. | |||||
|
FX4L4_HUMAN
|
||||||
| θ value | 5.62301e-10 (rank : 71) | NC score | 0.829361 (rank : 60) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q8WXT5 | Gene names | FOXD4L4, FOXD4B | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D4-like 4 (Forkhead box protein D4B) (Myeloid factor-gamma). | |||||
|
FOXD4_MOUSE
|
||||||
| θ value | 7.34386e-10 (rank : 72) | NC score | 0.832370 (rank : 58) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q60688, Q61573 | Gene names | Foxd4, Fkh2, Fkhl9, Freac5 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D4 (Forkhead-related protein FKHL9) (Forkhead- related transcription factor 5) (FREAC-5) (Transcription factor FKH- 2). | |||||
|
FOXN4_HUMAN
|
||||||
| θ value | 2.13673e-09 (rank : 73) | NC score | 0.799028 (rank : 69) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 100 | |
| SwissProt Accessions | Q96NZ1, Q6ZMR4, Q96NZ0 | Gene names | FOXN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein N4. | |||||
|
FOXN4_MOUSE
|
||||||
| θ value | 6.21693e-09 (rank : 74) | NC score | 0.801945 (rank : 68) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q8K3Q3, Q920C0 | Gene names | Foxn4 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N4. | |||||
|
FOXH1_HUMAN
|
||||||
| θ value | 1.17247e-07 (rank : 75) | NC score | 0.806714 (rank : 66) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 99 | |
| SwissProt Accessions | O75593 | Gene names | FOXH1, FAST1, FAST2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein H1 (Forkhead activin signal transducer 1) (Fast- 1) (hFAST-1) (Forkhead activin signal transducer 2) (Fast-2). | |||||
|
FOXH1_MOUSE
|
||||||
| θ value | 1.17247e-07 (rank : 76) | NC score | 0.810278 (rank : 65) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | O88621, Q9QZL5, Q9R241 | Gene names | Foxh1, Fast1, Fast2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein H1 (Forkhead activin signal transducer 1) (Fast- 1) (Forkhead activin signal transducer 2) (Fast-2). | |||||
|
FOXE2_HUMAN
|
||||||
| θ value | 7.59969e-07 (rank : 77) | NC score | 0.802205 (rank : 67) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | Q99526 | Gene names | FOXE2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein E2 (HNF-3/fork head-like protein 5) (HFKL5) (HFKH4). | |||||
|
FOXR2_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 78) | NC score | 0.769572 (rank : 76) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q3UM89 | Gene names | Foxr2, Foxn6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein R2 (Forkhead box protein N6). | |||||
|
FOXR1_MOUSE
|
||||||
| θ value | 3.77169e-06 (rank : 79) | NC score | 0.762132 (rank : 77) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q3UTB7 | Gene names | Foxr1, Foxn5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein R1 (Forkhead box protein N5). | |||||
|
FOXR1_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 80) | NC score | 0.743744 (rank : 80) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q6PIV2, Q86UT9, Q8IXX2 | Gene names | FOXR1, DLNB13, FOXN5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein R1 (Forkhead box protein N5). | |||||
|
DHX37_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 81) | NC score | 0.023195 (rank : 121) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IY37, Q9BUI7, Q9P211 | Gene names | DHX37, DDX37, KIAA1517 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DHX37 (EC 3.6.1.-) (DEAH box protein 37). | |||||
|
FOXR2_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 82) | NC score | 0.630047 (rank : 81) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q6PJQ5 | Gene names | FOXR2, FOXN6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein R2 (Forkhead box protein N6). | |||||
|
DGKK_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 83) | NC score | 0.030732 (rank : 105) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
PRR12_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 84) | NC score | 0.052252 (rank : 86) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
CH013_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 85) | NC score | 0.079283 (rank : 84) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96KS9, Q8N3M3, Q9NSR0 | Gene names | C8orf13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C8orf13. | |||||
|
H15_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 86) | NC score | 0.031099 (rank : 103) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P43276, Q9CRM8 | Gene names | Hist1h1b, H1f5 | |||
|
Domain Architecture |
|
|||||
| Description | Histone H1.5 (H1 VAR.5) (H1b). | |||||
|
MMP9_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 87) | NC score | 0.019402 (rank : 130) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P41245, Q06788, Q9DC02 | Gene names | Mmp9, Clg4b | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-9 precursor (EC 3.4.24.35) (MMP-9) (92 kDa type IV collagenase) (92 kDa gelatinase) (Gelatinase B) (GELB). | |||||
|
H12_HUMAN
|
||||||
| θ value | 0.125558 (rank : 88) | NC score | 0.033254 (rank : 96) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P16403 | Gene names | HIST1H1C, H1F2 | |||
|
Domain Architecture |
|
|||||
| Description | Histone H1.2 (Histone H1d). | |||||
|
ZBT12_HUMAN
|
||||||
| θ value | 0.125558 (rank : 89) | NC score | -0.002828 (rank : 189) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y330, Q5JQ98 | Gene names | ZBTB12, C6orf46, G10, NG35 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 12 (Protein G10). | |||||
|
CS007_MOUSE
|
||||||
| θ value | 0.279714 (rank : 90) | NC score | 0.028949 (rank : 108) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
PTN23_MOUSE
|
||||||
| θ value | 0.279714 (rank : 91) | NC score | 0.024217 (rank : 118) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
LTBP4_MOUSE
|
||||||
| θ value | 0.365318 (rank : 92) | NC score | 0.004883 (rank : 172) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8K4G1, Q8K4G0 | Gene names | Ltbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 4 precursor (LTBP-4). | |||||
|
NOL3_MOUSE
|
||||||
| θ value | 0.365318 (rank : 93) | NC score | 0.031257 (rank : 102) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D1X0 | Gene names | Nol3 | |||
|
Domain Architecture |
|
|||||
| Description | Nucleolar protein 3. | |||||
|
PRP2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 94) | NC score | 0.049268 (rank : 87) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
K0460_MOUSE
|
||||||
| θ value | 0.47712 (rank : 95) | NC score | 0.029590 (rank : 106) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6NXI6, Q3U3L8, Q6ZQA7 | Gene names | Kiaa0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
BSN_HUMAN
|
||||||
| θ value | 0.62314 (rank : 96) | NC score | 0.026063 (rank : 112) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CO039_MOUSE
|
||||||
| θ value | 0.62314 (rank : 97) | NC score | 0.035886 (rank : 93) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
CS016_HUMAN
|
||||||
| θ value | 0.62314 (rank : 98) | NC score | 0.037208 (rank : 92) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8ND99 | Gene names | C19orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf16. | |||||
|
IF4G3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 99) | NC score | 0.013831 (rank : 146) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43432, Q15597, Q8NEN1 | Gene names | EIF4G3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
LRP5_MOUSE
|
||||||
| θ value | 0.62314 (rank : 100) | NC score | 0.018848 (rank : 134) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q91VN0, O88883, Q9R208 | Gene names | Lrp5, Lrp7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 5 precursor (LRP7) (Lr3). | |||||
|
BSPRY_MOUSE
|
||||||
| θ value | 0.813845 (rank : 101) | NC score | 0.010296 (rank : 155) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80YW5, Q3TU74, Q8BZF0, Q99KV7, Q9ER70 | Gene names | Bspry | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B box and SPRY domain-containing protein. | |||||
|
CABL2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 102) | NC score | 0.021289 (rank : 126) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BTV7, Q5JWL0, Q9BYK0 | Gene names | CABLES2, C20orf150 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 and ABL1 enzyme substrate 2 (Interactor with CDK3 2) (Ik3-2). | |||||
|
H11_MOUSE
|
||||||
| θ value | 0.813845 (rank : 103) | NC score | 0.027156 (rank : 110) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P43275 | Gene names | Hist1h1a, H1f1 | |||
|
Domain Architecture |
|
|||||
| Description | Histone H1.1 (H1 VAR.3) (H1a). | |||||
|
HNF1A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 104) | NC score | 0.033176 (rank : 97) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
SFR15_HUMAN
|
||||||
| θ value | 0.813845 (rank : 105) | NC score | 0.031906 (rank : 99) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O95104, Q6P1M5, Q8N3I8, Q9UFM1, Q9ULP8 | Gene names | SFRS15, KIAA1172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 15 (CTD-binding SR-like protein RA4). | |||||
|
PKHG4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 106) | NC score | 0.023756 (rank : 119) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q58EX7, Q4G0J8, Q4H485, Q56A69, Q9H7K4, Q9UFW0 | Gene names | PLEKHG4, PRTPHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Puratrophin-1 (Pleckstrin homology domain-containing family G member 4) (Purkinje cell atrophy-associated protein 1). | |||||
|
SELV_HUMAN
|
||||||
| θ value | 1.06291 (rank : 107) | NC score | 0.037958 (rank : 91) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P59797 | Gene names | SELV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Selenoprotein V. | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 108) | NC score | 0.042037 (rank : 88) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CNOT4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 109) | NC score | 0.028962 (rank : 107) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |
|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
ESPL1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 110) | NC score | 0.020786 (rank : 127) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14674 | Gene names | ESPL1, ESP1, KIAA0165 | |||
|
Domain Architecture |

|
|||||
| Description | Separin (EC 3.4.22.49) (Separase) (Caspase-like protein ESPL1) (Extra spindle poles-like 1 protein). | |||||
|
KTNB1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 111) | NC score | 0.000059 (rank : 183) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BVA0, O60620 | Gene names | KATNB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Katanin p80 WD40-containing subunit B1 (Katanin p80 subunit B1) (p80 katanin). | |||||
|
MUC2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 112) | NC score | 0.011428 (rank : 152) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
RAI2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 113) | NC score | 0.026651 (rank : 111) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QVY8 | Gene names | Rai2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2 (3f8). | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 114) | NC score | 0.016973 (rank : 141) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
SGIP1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 115) | NC score | 0.018774 (rank : 135) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8VD37, Q3UFU3, Q3UGA0, Q8BXX4, Q8C034, Q9CXT2 | Gene names | Sgip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3-containing GRB2-like protein 3-interacting protein 1 (Endophilin- 3-interacting protein). | |||||
|
ZN295_HUMAN
|
||||||
| θ value | 1.38821 (rank : 116) | NC score | -0.003196 (rank : 190) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULJ3, Q6P4R0 | Gene names | ZNF295, KIAA1227, ZBTB21 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 295 (Zinc finger and BTB domain-containing protein 21). | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 117) | NC score | 0.031664 (rank : 100) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
LPIN2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 118) | NC score | 0.009828 (rank : 159) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PI5, Q8C357, Q8C7I8, Q8CC85, Q8CHR7 | Gene names | Lpin2 | |||
|
Domain Architecture |
|
|||||
| Description | Lipin-2. | |||||
|
PTN23_HUMAN
|
||||||
| θ value | 1.81305 (rank : 119) | NC score | 0.015965 (rank : 143) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
SDC3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 120) | NC score | 0.017281 (rank : 139) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q64519, Q3UDD9, Q6ZQA4, Q7TQD4 | Gene names | Sdc3, Kiaa0468 | |||
|
Domain Architecture |
|
|||||
| Description | Syndecan-3 precursor (SYND3). | |||||
|
SON_HUMAN
|
||||||
| θ value | 1.81305 (rank : 121) | NC score | 0.033961 (rank : 95) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
CCD37_HUMAN
|
||||||
| θ value | 2.36792 (rank : 122) | NC score | 0.015806 (rank : 144) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q494V2, Q494V1, Q494V4, Q8N838 | Gene names | CCDC37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 37. | |||||
|
FIP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 123) | NC score | 0.027617 (rank : 109) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6UN15, Q499Y4, Q49AU3, Q7Z608, Q8WVN3, Q96F80, Q9H077 | Gene names | FIP1L1, FIP1, RHE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA 3'-end-processing factor FIP1 (FIP1-like 1) (Factor interacting with PAP) (hFip1) (Rearranged in hypereosinophilia). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 2.36792 (rank : 124) | NC score | 0.012172 (rank : 151) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 125) | NC score | 0.019070 (rank : 131) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
RAI2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 126) | NC score | 0.025777 (rank : 113) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y5P3, Q8N6X7 | Gene names | RAI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2. | |||||
|
RHG09_HUMAN
|
||||||
| θ value | 2.36792 (rank : 127) | NC score | 0.005951 (rank : 171) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BRR9, Q8TCJ3, Q8WYR0, Q96EZ2, Q96S74 | Gene names | ARHGAP9 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 9. | |||||
|
CCD72_HUMAN
|
||||||
| θ value | 3.0926 (rank : 128) | NC score | 0.141099 (rank : 82) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2S6, Q9P052 | Gene names | CCDC72 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 72. | |||||
|
CCD72_MOUSE
|
||||||
| θ value | 3.0926 (rank : 129) | NC score | 0.141099 (rank : 83) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K003 | Gene names | Ccdc72 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 72. | |||||
|
CDN1C_HUMAN
|
||||||
| θ value | 3.0926 (rank : 130) | NC score | 0.016944 (rank : 142) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P49918 | Gene names | CDKN1C, KIP2 | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
CSPG4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 131) | NC score | 0.007645 (rank : 165) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6UVK1, Q92675 | Gene names | CSPG4, MCSP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 4 precursor (Chondroitin sulfate proteoglycan NG2) (Melanoma chondroitin sulfate proteoglycan) (Melanoma-associated chondroitin sulfate proteoglycan). | |||||
|
DOT1L_HUMAN
|
||||||
| θ value | 3.0926 (rank : 132) | NC score | 0.022029 (rank : 124) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
FMN1A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 133) | NC score | 0.019056 (rank : 132) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q05860 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoforms I/II/III (Limb deformity protein). | |||||
|
FMN1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 134) | NC score | 0.019545 (rank : 129) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q05859 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoform IV (Limb deformity protein). | |||||
|
FRAT1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 135) | NC score | 0.013115 (rank : 148) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70339 | Gene names | Frat1 | |||
|
Domain Architecture |
|
|||||
| Description | Proto-oncogene FRAT1 (Frequently rearranged in advanced T-cell lymphomas) (Frat-1). | |||||
|
GGN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 136) | NC score | 0.022580 (rank : 122) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
GPR37_MOUSE
|
||||||
| θ value | 3.0926 (rank : 137) | NC score | -0.001748 (rank : 188) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QY42, Q8BKF8, Q8BXP9, Q8C022, Q9Z0G3 | Gene names | Gpr37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 37 precursor. | |||||
|
GSCR1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 138) | NC score | 0.035843 (rank : 94) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
HNF1A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 139) | NC score | 0.025482 (rank : 115) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P20823, Q99861 | Gene names | TCF1, HNF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1) (Transcription factor 1) (TCF-1). | |||||
|
KLF13_HUMAN
|
||||||
| θ value | 3.0926 (rank : 140) | NC score | -0.004201 (rank : 193) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y2Y9, Q9Y356 | Gene names | KLF13, BTEB3, NSLP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 13 (Transcription factor BTEB3) (Basic transcription element-binding protein 3) (BTE-binding protein 3) (RANTES factor of late activated T-lymphocytes 1) (RFLAT-1) (Transcription factor NSLP1) (Novel Sp1-like zinc finger transcription factor 1). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 3.0926 (rank : 141) | NC score | 0.057105 (rank : 85) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 102 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
ROBO3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 142) | NC score | 0.001083 (rank : 180) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z2I4 | Gene names | Robo3, Rbig1 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Retinoblastoma-inhibiting gene 1) (Rig-1). | |||||
|
BAG3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 143) | NC score | 0.018978 (rank : 133) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
BRD4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 144) | NC score | 0.006754 (rank : 168) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
CEP68_HUMAN
|
||||||
| θ value | 4.03905 (rank : 145) | NC score | 0.012250 (rank : 150) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q76N32, O60326, Q9BQ18, Q9UDM9 | Gene names | CEP68, KIAA0582 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 68 kDa (Cep68 protein). | |||||
|
CN092_HUMAN
|
||||||
| θ value | 4.03905 (rank : 146) | NC score | 0.008122 (rank : 162) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O94842 | Gene names | C14orf92, KIAA0737 | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal Langerhans cell protein LCP1. | |||||
|
FIP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 147) | NC score | 0.024328 (rank : 117) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D824, Q8BWX7, Q99LH0, Q9DBB2 | Gene names | Fip1l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA 3'-end-processing factor FIP1 (FIP1-like 1). | |||||
|
MYCB2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 148) | NC score | 0.006811 (rank : 167) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O75592, Q5JSX8, Q5VZN6, Q6PIB6, Q9UQ11, Q9Y6E4 | Gene names | MYCBP2, KIAA0916, PAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
SH22A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 149) | NC score | 0.011011 (rank : 153) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NP31, O43817, Q9UPA7 | Gene names | SH2D2A, SCAP, TSAD, VRAP | |||
|
Domain Architecture |
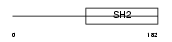
|
|||||
| Description | SH2 domain protein 2A (T cell-specific adapter protein) (TSAd) (VEGF receptor-associated protein) (SH2 domain-containing adapter protein). | |||||
|
WASIP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 150) | NC score | 0.033090 (rank : 98) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
XG_HUMAN
|
||||||
| θ value | 4.03905 (rank : 151) | NC score | 0.023720 (rank : 120) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55808, Q71BZ5 | Gene names | XG, PBDX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glycoprotein Xg precursor (Protein PBDX). | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 152) | NC score | 0.041666 (rank : 89) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CD2L6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 153) | NC score | -0.005739 (rank : 196) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BWU1, Q5JQZ7, Q5JR00, Q8TC78, Q9UPX2 | Gene names | CDC2L6, CDK11, KIAA1028 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 6 (EC 2.7.11.22) (CDC2- related protein kinase 6) (Death-preventing kinase) (Cyclin-dependent kinase 11). | |||||
|
CF182_MOUSE
|
||||||
| θ value | 5.27518 (rank : 154) | NC score | 0.001955 (rank : 178) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 551 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VDS7, Q9CZE0, Q9D5A1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf182 homolog. | |||||
|
CRUM2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 155) | NC score | -0.000559 (rank : 185) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5IJ48, Q5JS41, Q5JS43, Q6ZTA9, Q6ZWI6 | Gene names | CRB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crumbs homolog 2 precursor (Crumbs-like protein 2). | |||||
|
DIAP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 156) | NC score | 0.010090 (rank : 157) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O70566, Q8C2G8 | Gene names | Diaph2, Diap2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2) (mDia3). | |||||
|
GATA3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 157) | NC score | 0.006065 (rank : 170) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P23771, Q96J16 | Gene names | GATA3 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-acting T-cell-specific transcription factor GATA-3 (GATA-binding factor 3). | |||||
|
GCNL2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 158) | NC score | 0.010680 (rank : 154) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92830, Q9UCW1 | Gene names | GCN5L2, GCN5, HGCN5 | |||
|
Domain Architecture |

|
|||||
| Description | General control of amino acid synthesis protein 5-like 2 (EC 2.3.1.48) (Histone acetyltransferase GCN5) (hsGCN5) (STAF97). | |||||
|
MK07_MOUSE
|
||||||
| θ value | 5.27518 (rank : 159) | NC score | -0.004270 (rank : 194) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1054 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9WVS8 | Gene names | Mapk7, Erk5 | |||
|
Domain Architecture |
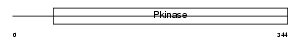
|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (BMK1 kinase). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 160) | NC score | 0.013819 (rank : 147) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
MOT8_MOUSE
|
||||||
| θ value | 5.27518 (rank : 161) | NC score | 0.007820 (rank : 164) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O70324, Q8K3S9 | Gene names | Slc16a2, Mct8, Xpct | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Monocarboxylate transporter 8 (MCT 8) (Solute carrier family 16 member 2) (X-linked PEST-containing transporter). | |||||
|
SAPS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 162) | NC score | 0.018337 (rank : 136) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 163) | NC score | 0.024990 (rank : 116) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 164) | NC score | 0.030894 (rank : 104) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
ADA19_HUMAN
|
||||||
| θ value | 6.88961 (rank : 165) | NC score | 0.000383 (rank : 181) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
ARTN_HUMAN
|
||||||
| θ value | 6.88961 (rank : 166) | NC score | 0.008130 (rank : 161) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5T4W7, O95441, O96030, Q6P6A3 | Gene names | ARTN, EVN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Artemin precursor (Enovin) (Neublastin). | |||||
|
ATBF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 167) | NC score | -0.000419 (rank : 184) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
ATBF1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 168) | NC score | 0.000229 (rank : 182) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
CD2L5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 169) | NC score | -0.006748 (rank : 197) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14004, Q6DKQ9, Q75MH4, Q75MH5, Q96JN4, Q9H4A0, Q9H4A1, Q9UDR4 | Gene names | CDC2L5, CDC2L, CHED, KIAA1791 | |||
|
Domain Architecture |
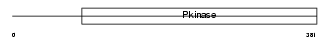
|
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5) (Cholinesterase-related cell division controller). | |||||
|
CS007_HUMAN
|
||||||
| θ value | 6.88961 (rank : 170) | NC score | 0.022330 (rank : 123) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
GP1BA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 171) | NC score | -0.000986 (rank : 186) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P07359, Q14441, Q16469, Q8N1F3, Q8NG39, Q9HDC7, Q9UEK1, Q9UQS4 | Gene names | GP1BA | |||
|
Domain Architecture |

|
|||||
| Description | Platelet glycoprotein Ib alpha chain precursor (Glycoprotein Ibalpha) (GP-Ib alpha) (GPIbA) (GPIb-alpha) (Antigen CD42b-alpha) (CD42b antigen) [Contains: Glycocalicin]. | |||||
|
H10_HUMAN
|
||||||
| θ value | 6.88961 (rank : 172) | NC score | 0.020419 (rank : 128) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P07305 | Gene names | H1F0, H1FV | |||
|
Domain Architecture |
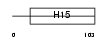
|
|||||
| Description | Histone H1.0 (H1(0)) (Histone H1'). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 173) | NC score | 0.012385 (rank : 149) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 174) | NC score | 0.025605 (rank : 114) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
2DMA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 175) | NC score | 0.002243 (rank : 177) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28078 | Gene names | H2-DMa, H2-Ma, Ma | |||
|
Domain Architecture |
|
|||||
| Description | Class II histocompatibility antigen, M alpha chain precursor. | |||||
|
ASPP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 176) | NC score | 0.001154 (rank : 179) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q13625, Q12892, Q86X75, Q96KQ3 | Gene names | TP53BP2, ASPP2, BBP | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 2 (Tumor suppressor p53-binding protein 2) (p53-binding protein 2) (p53BP2) (53BP2) (Bcl2-binding protein) (Bbp) (NY-REN-51 antigen). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 177) | NC score | 0.031455 (rank : 101) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
DIAP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 178) | NC score | 0.007883 (rank : 163) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O60610, Q9UC76 | Gene names | DIAPH1, DIAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1). | |||||
|
FMNL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 179) | NC score | 0.018257 (rank : 138) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
Domain Architecture |
|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
GASP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 180) | NC score | 0.006854 (rank : 166) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5U4C1, Q8BKR8, Q8BUN4, Q8BYK9, Q8CHF4, Q8R095 | Gene names | Gprasp1, Kiaa0443 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 1 (GASP-1). | |||||
|
GGA3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 181) | NC score | 0.003602 (rank : 175) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZ52, Q15017, Q6IS16, Q9UJY3 | Gene names | GGA3, KIAA0154 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-binding protein GGA3 (Golgi-localized, gamma ear-containing, ARF-binding protein 3). | |||||
|
K0460_HUMAN
|
||||||
| θ value | 8.99809 (rank : 182) | NC score | 0.018276 (rank : 137) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5VT52, O75048, Q5VT53, Q6MZL4, Q86XD2 | Gene names | KIAA0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
LRP5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 183) | NC score | 0.009896 (rank : 158) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75197, Q96TD6, Q9UP66 | Gene names | LRP5, LRP7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 5 precursor. | |||||
|
MEF2B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 184) | NC score | 0.009687 (rank : 160) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q02080 | Gene names | MEF2B, XMEF2 | |||
|
Domain Architecture |
|
|||||
| Description | Myocyte-specific enhancer factor 2B (Serum response factor-like protein 2) (XMEF2) (RSRFR2). | |||||
|
MK07_HUMAN
|
||||||
| θ value | 8.99809 (rank : 185) | NC score | -0.005012 (rank : 195) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1087 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q13164, Q16634 | Gene names | MAPK7, ERK4, ERK5, PRKM7 | |||
|
Domain Architecture |
|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (ERK4) (BMK1 kinase). | |||||
|
MMP9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 186) | NC score | 0.010141 (rank : 156) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P14780, Q3LR70, Q8N725, Q9H4Z1 | Gene names | MMP9, CLG4B | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-9 precursor (EC 3.4.24.35) (MMP-9) (92 kDa type IV collagenase) (92 kDa gelatinase) (Gelatinase B) (GELB) [Contains: 67 kDa matrix metalloproteinase-9; 82 kDa matrix metalloproteinase-9]. | |||||
|
PHLA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 187) | NC score | 0.021418 (rank : 125) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8WV24, Q15184, Q2TAN2, Q9NZ17 | Gene names | PHLDA1, PHRIP, TDAG51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Apoptosis-associated nuclear protein) (Proline- and histidine-rich protein) (Proline- and glutamine-rich protein) (PQ-rich protein). | |||||
|
PHLA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 188) | NC score | 0.015024 (rank : 145) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q62392, Q3TY48, Q3UG87 | Gene names | Phlda1, Tdag51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Proline- and glutamine-rich protein) (PQR protein). | |||||
|
PININ_MOUSE
|
||||||
| θ value | 8.99809 (rank : 189) | NC score | 0.002305 (rank : 176) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 190) | NC score | 0.039924 (rank : 90) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
ROBO2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 191) | NC score | -0.001726 (rank : 187) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TPD3, Q8BJ59 | Gene names | Robo2, Kiaa1568 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 2 precursor. | |||||
|
SEM6D_HUMAN
|
||||||
| θ value | 8.99809 (rank : 192) | NC score | 0.003615 (rank : 174) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NFY4, Q8NFY3, Q8NFY5, Q8NFY6, Q8NFY7, Q9P249 | Gene names | SEMA6D, KIAA1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
SEM6D_MOUSE
|
||||||
| θ value | 8.99809 (rank : 193) | NC score | 0.003773 (rank : 173) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q76KF0, Q76KF1, Q76KF2, Q76KF3, Q76KF4, Q80TD0 | Gene names | Sema6d, Kiaa1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
SFPQ_HUMAN
|
||||||
| θ value | 8.99809 (rank : 194) | NC score | 0.017018 (rank : 140) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
SMG7_MOUSE
|
||||||
| θ value | 8.99809 (rank : 195) | NC score | 0.006417 (rank : 169) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5RJH6, Q63ZW5, Q6ZQF3 | Gene names | Smg7, Est1c, Kiaa0250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
SPEG_HUMAN
|
||||||
| θ value | 8.99809 (rank : 196) | NC score | -0.003583 (rank : 191) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 197) | NC score | -0.003808 (rank : 192) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
FOXO4_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 197 | |
| SwissProt Accessions | P98177, O43821, Q13720, Q3KPF1, Q8TDK9 | Gene names | MLLT7, AFX, AFX1, FOXO4 | |||
|
Domain Architecture |

|
|||||
| Description | Fork head domain transcription factor AFX1 (Forkhead box protein O4). | |||||
|
FOXO4_MOUSE
|
||||||
| NC score | 0.988424 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 111 | |
| SwissProt Accessions | Q9WVH3 | Gene names | Mllt7, Afx, Afx1, Fkhr3, Foxo4 | |||
|
Domain Architecture |

|
|||||
| Description | Fork head domain transcription factor AFX1 (Afxh) (Forkhead box protein O4). | |||||
|
FOXO3_HUMAN
|
||||||
| NC score | 0.964746 (rank : 3) | θ value | 4.99229e-75 (rank : 3) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | O43524, O15171, Q5T2I7, Q9BZ04 | Gene names | FOXO3A, FKHRL1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein O3A (Forkhead in rhabdomyosarcoma-like 1) (AF6q21 protein). | |||||
|
FOXO1_MOUSE
|
||||||
| NC score | 0.955560 (rank : 4) | θ value | 4.37348e-71 (rank : 5) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q9R1E0 | Gene names | Foxo1a, Fkhr, Foxo1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein O1A (Forkhead in rhabdomyosarcoma). | |||||
|
FOXO1_HUMAN
|
||||||
| NC score | 0.954542 (rank : 5) | θ value | 1.96316e-71 (rank : 4) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q12778, O43523, Q5VYC7, Q6NSK6 | Gene names | FOXO1A, FKHR | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein O1A (Forkhead in rhabdomyosarcoma). | |||||
|
FOXI2_MOUSE
|
||||||
| NC score | 0.869316 (rank : 6) | θ value | 1.38178e-16 (rank : 11) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q3I5G5, Q8BIK9 | Gene names | Foxi2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein I2. | |||||
|
FOXGB_MOUSE
|
||||||
| NC score | 0.866206 (rank : 7) | θ value | 8.10077e-17 (rank : 7) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q60987, Q80VP3 | Gene names | Foxg1b, Fkhl1, Foxg1, Hfhbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1B (Forkhead-related protein FKHL1) (Transcription factor BF-1) (Brain factor 1) (BF1). | |||||
|
FOXGB_HUMAN
|
||||||
| NC score | 0.866174 (rank : 8) | θ value | 7.32683e-18 (rank : 6) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | P55315 | Gene names | FOXG1B, FKHL1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1B (Forkhead-related protein FKHL1) (Transcription factor BF-1) (Brain factor 1) (BF1) (HFK1). | |||||
|
FOXQ1_MOUSE
|
||||||
| NC score | 0.865644 (rank : 9) | θ value | 1.99529e-15 (rank : 17) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | O70220 | Gene names | Foxq1, Hfh1, Hfh1l | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein Q1 (Hepatocyte nuclear factor 3 forkhead homolog 1) (HNF-3/forkhead-like protein 1) (HFH-1) (HFH-1l). | |||||
|
FOXQ1_HUMAN
|
||||||
| NC score | 0.863482 (rank : 10) | θ value | 2.60593e-15 (rank : 20) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q9C009, Q9NS06 | Gene names | FOXQ1, HFH1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein Q1 (Hepatocyte nuclear factor 3 forkhead homolog 1) (HNF-3/forkhead-like protein 1) (HFH-1). | |||||
|
FOXGA_HUMAN
|
||||||
| NC score | 0.862851 (rank : 11) | θ value | 1.99529e-15 (rank : 16) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | P55316 | Gene names | FOXG1A, FKHL2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1A (Forkhead-related protein FKHL2) (Transcription factor BF-2) (Brain factor 2) (BF2) (HFK2). | |||||
|
FOXI1_MOUSE
|
||||||
| NC score | 0.861266 (rank : 12) | θ value | 1.80466e-16 (rank : 13) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | Q922I5, Q5SRI5, Q9D299 | Gene names | Foxi1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein I1. | |||||
|
FOXI1_HUMAN
|
||||||
| NC score | 0.861119 (rank : 13) | θ value | 8.95645e-16 (rank : 15) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q12951, Q14518, Q66SR7, Q8N6L8 | Gene names | FOXI1, FKHL10, FREAC6 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein I1 (Forkhead-related protein FKHL10) (Forkhead- related transcription factor 6) (FREAC-6) (Hepatocyte nuclear factor 3 forkhead homolog 3) (HNF-3/fork-head homolog 3) (HFH-3). | |||||
|
FOXJ3_HUMAN
|
||||||
| NC score | 0.860605 (rank : 14) | θ value | 8.10077e-17 (rank : 8) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q9UPW0, Q9NSS7 | Gene names | FOXJ3, KIAA1041 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J3. | |||||
|
FOXJ3_MOUSE
|
||||||
| NC score | 0.860570 (rank : 15) | θ value | 8.10077e-17 (rank : 9) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q8BUR3, Q3UTD8 | Gene names | Foxj3, Kiaa1041 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J3. | |||||
|
FOXJ2_MOUSE
|
||||||
| NC score | 0.858673 (rank : 16) | θ value | 2.60593e-15 (rank : 19) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | Q9ES18 | Gene names | Foxj2, Fhx | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J2 (Fork head homologous X). | |||||
|
FOXD2_HUMAN
|
||||||
| NC score | 0.858248 (rank : 17) | θ value | 1.80466e-16 (rank : 12) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | O60548 | Gene names | FOXD2, FKHL17, FREAC9 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D2 (Forkhead-related protein FKHL17) (Forkhead- related transcription factor 9) (FREAC-9). | |||||
|
FOXF2_HUMAN
|
||||||
| NC score | 0.858154 (rank : 18) | θ value | 2.43908e-13 (rank : 32) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q12947, Q9UQ85 | Gene names | FOXF2, FKHL6, FREAC2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein F2 (Forkhead-related protein FKHL6) (Forkhead- related transcription factor 2) (FREAC-2) (Forkhead-related activator 2). | |||||
|
FOXL2_MOUSE
|
||||||
| NC score | 0.857683 (rank : 19) | θ value | 2.43908e-13 (rank : 33) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | O88470, Q8K3X9 | Gene names | Foxl2, Pfrk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein L2 (Pituitary forkhead factor) (P-Frk). | |||||
|
FOXL2_HUMAN
|
||||||
| NC score | 0.857590 (rank : 20) | θ value | 1.58096e-12 (rank : 49) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P58012, Q4ZGJ3 | Gene names | FOXL2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein L2. | |||||
|
FOXD1_MOUSE
|
||||||
| NC score | 0.857331 (rank : 21) | θ value | 2.88119e-14 (rank : 29) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q61345 | Gene names | Foxd1, Fkhl8, Freac4, Hfhbf2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D1 (Forkhead-related protein FKHL8) (Forkhead- related transcription factor 4) (FREAC-4) (Brain factor 2) (HFH-BF-2). | |||||
|
FOXL1_HUMAN
|
||||||
| NC score | 0.857073 (rank : 22) | θ value | 1.86753e-13 (rank : 31) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q12952, Q9H242 | Gene names | FOXL1, FKHL11, FREAC7 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein L1 (Forkhead-related protein FKHL11) (Forkhead- related transcription factor 7) (FREAC-7). | |||||
|
FOXD2_MOUSE
|
||||||
| NC score | 0.856955 (rank : 23) | θ value | 5.25075e-16 (rank : 14) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | O35392 | Gene names | Foxd2, Mf2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D2 (Mesoderm/mesenchyme forkhead 2) (MF-2). | |||||
|
FOXN2_HUMAN
|
||||||
| NC score | 0.856095 (rank : 24) | θ value | 6.00763e-12 (rank : 57) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | P32314, Q15769 | Gene names | FOXN2, HTLF | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N2 (Human T-cell leukemia virus enhancer factor). | |||||
|
FOXD1_HUMAN
|
||||||
| NC score | 0.855192 (rank : 25) | θ value | 1.68911e-14 (rank : 25) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q16676, Q12949 | Gene names | FOXD1, FKHL8, FREAC4 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D1 (Forkhead-related protein FKHL8) (Forkhead- related transcription factor 4) (FREAC-4). | |||||
|
FOXJ2_HUMAN
|
||||||
| NC score | 0.853630 (rank : 26) | θ value | 2.60593e-15 (rank : 18) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9P0K8, Q96PS9, Q9NSN5 | Gene names | FOXJ2, FHX | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J2 (Fork head homologous X). | |||||
|
HNF3G_HUMAN
|
||||||
| NC score | 0.853232 (rank : 27) | θ value | 9.90251e-15 (rank : 23) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | P55318, Q9UMW9 | Gene names | FOXA3, HNF3G, TCF3G | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-gamma (HNF-3G) (Forkhead box protein A3) (Fork head-related protein FKH H3). | |||||
|
FOXK2_HUMAN
|
||||||
| NC score | 0.852741 (rank : 28) | θ value | 1.86753e-13 (rank : 30) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | Q01167, Q13622, Q13623, Q13624 | Gene names | FOXK2, ILF, ILF1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein K2 (Interleukin enhancer-binding factor 1) (Cellular transcription factor ILF-1). | |||||
|
FOXE3_MOUSE
|
||||||
| NC score | 0.852012 (rank : 29) | θ value | 2.20605e-14 (rank : 27) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q9QY14 | Gene names | Foxe3 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein E3. | |||||
|
FOXE1_MOUSE
|
||||||
| NC score | 0.851860 (rank : 30) | θ value | 1.68911e-14 (rank : 26) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q8R2I0 | Gene names | Foxe1, Titf2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein E1 (Thyroid transcription factor 2) (TTF-2). | |||||
|
FOXL1_MOUSE
|
||||||
| NC score | 0.851658 (rank : 31) | θ value | 1.58096e-12 (rank : 48) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q64731 | Gene names | Foxl1, Fkh6, Fkhl11 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein L1 (Forkhead-related protein FKHL11) (Transcription factor FKH-6). | |||||
|
FOXE1_HUMAN
|
||||||
| NC score | 0.850849 (rank : 32) | θ value | 1.29331e-14 (rank : 24) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | O00358, O75765 | Gene names | FOXE1, FKHL15, TITF2, TTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein E1 (Thyroid transcription factor 2) (TTF-2) (Forkhead-related protein FKHL15). | |||||
|
HNF3G_MOUSE
|
||||||
| NC score | 0.850712 (rank : 33) | θ value | 9.26847e-13 (rank : 44) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | P35584 | Gene names | Foxa3, Hnf3g, Tcf-3g, Tcf3g | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-gamma (HNF-3G) (Forkhead box protein A3). | |||||
|
FOXF1_MOUSE
|
||||||
| NC score | 0.850438 (rank : 34) | θ value | 1.2105e-12 (rank : 45) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q61080, Q61661 | Gene names | Foxf1, Fkhl5, Foxf1a, Freac1, Hfh8 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein F1 (Forkhead-related protein FKHL5) (Forkhead- related transcription factor 1) (FREAC-1) (Hepatocyte nuclear factor 3 forkhead homolog 8) (HFH-8). | |||||
|
FOXC2_MOUSE
|
||||||
| NC score | 0.850154 (rank : 35) | θ value | 2.88119e-14 (rank : 28) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q61850, P97948, Q63869, Q8C694 | Gene names | Foxc2, Fkh14, Fkhl14, Mfh1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
FOXD3_HUMAN
|
||||||
| NC score | 0.849561 (rank : 36) | θ value | 7.09661e-13 (rank : 38) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9UJU5, Q9BYM2, Q9UDD1 | Gene names | FOXD3 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D3 (HNF3/FH transcription factor genesis). | |||||
|
FOXF1_HUMAN
|
||||||
| NC score | 0.849354 (rank : 37) | θ value | 9.26847e-13 (rank : 41) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q12946, Q5FWE5 | Gene names | FOXF1, FKHL5, FREAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein F1 (Forkhead-related protein FKHL5) (Forkhead- related transcription factor 1) (FREAC-1) (Forkhead-related activator 1). | |||||
|
FOXN3_MOUSE
|
||||||
| NC score | 0.848898 (rank : 38) | θ value | 5.43371e-13 (rank : 35) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q499D0, Q8C5N8, Q9CU83 | Gene names | Ches1, Foxn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Checkpoint suppressor 1 (Forkhead box protein N3). | |||||
|
FOXE3_HUMAN
|
||||||
| NC score | 0.848804 (rank : 39) | θ value | 2.0648e-12 (rank : 52) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q13461, Q5SVY9, Q9NQV9 | Gene names | FOXE3, FKHL12, FREAC8 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein E3 (Forkhead-related protein FKHL12) (Forkhead- related transcription factor 8) (FREAC-8). | |||||
|
FOXJ1_HUMAN
|
||||||
| NC score | 0.848717 (rank : 40) | θ value | 2.0648e-12 (rank : 53) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q92949, O00630 | Gene names | FOXJ1, FKHL13, HFH4 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J1 (Forkhead-related protein FKHL13) (Hepatocyte nuclear factor 3 forkhead homolog 4) (HFH-4). | |||||
|
FOXK1_MOUSE
|
||||||
| NC score | 0.847232 (rank : 41) | θ value | 1.2105e-12 (rank : 47) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | P42128, O35939 | Gene names | Foxk1, Mnf | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein K1 (Myocyte nuclear factor) (MNF). | |||||
|
FOXD3_MOUSE
|
||||||
| NC score | 0.846779 (rank : 42) | θ value | 7.09661e-13 (rank : 39) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q61060 | Gene names | Foxd3, Hfh2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D3 (HNF3/FH transcription factor genesis) (Hepatocyte nuclear factor 3 forkhead homolog 2) (HFH-2). | |||||
|
FOXN3_HUMAN
|
||||||
| NC score | 0.846767 (rank : 43) | θ value | 5.43371e-13 (rank : 34) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | O00409, Q96II7, Q9UIE7 | Gene names | CHES1, FOXN3 | |||
|
Domain Architecture |

|
|||||
| Description | Checkpoint suppressor 1 (Forkhead box protein N3). | |||||
|
FOXC1_MOUSE
|
||||||
| NC score | 0.846344 (rank : 44) | θ value | 7.58209e-15 (rank : 22) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 93 | |
| SwissProt Accessions | Q61572, O88409, Q61582 | Gene names | Foxc1, Fkh1, Fkhl7, Freac3, Mf1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein C1 (Forkhead-related protein FKHL7) (Forkhead- related transcription factor 3) (FREAC-3) (Transcription factor FKH-1) (Mesoderm/mesenchyme forkhead 1) (MF-1). | |||||
|
FOXJ1_MOUSE
|
||||||
| NC score | 0.846315 (rank : 45) | θ value | 1.2105e-12 (rank : 46) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q61660 | Gene names | Foxj1, Fkhl13, Hfh4 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J1 (Forkhead-related protein FKHL13) (Hepatocyte nuclear factor 3 forkhead homolog 4) (HFH-4). | |||||
|
FOXC2_HUMAN
|
||||||
| NC score | 0.845335 (rank : 46) | θ value | 1.38178e-16 (rank : 10) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q99958 | Gene names | FOXC2, FKHL14, MFH1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
FOXC1_HUMAN
|
||||||
| NC score | 0.845183 (rank : 47) | θ value | 7.58209e-15 (rank : 21) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q12948, Q86UP7, Q9BYM1, Q9NUE5, Q9UDD0, Q9UP06 | Gene names | FOXC1, FKHL7, FREAC3 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein C1 (Forkhead-related protein FKHL7) (Forkhead- related transcription factor 3) (FREAC-3). | |||||
|
FOXB1_MOUSE
|
||||||
| NC score | 0.844982 (rank : 48) | θ value | 7.09661e-13 (rank : 37) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q64732, O08753 | Gene names | Foxb1, Fkh5, Foxb1a, Foxb1b, Mf3 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein B1 (Transcription factor FKH-5). | |||||
|
HNF3B_MOUSE
|
||||||
| NC score | 0.844699 (rank : 49) | θ value | 1.58096e-12 (rank : 51) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | P35583, Q60602 | Gene names | Foxa2, Hnf3b, Tcf-3b, Tcf3b | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-beta (HNF-3B) (Forkhead box protein A2). | |||||
|
HNF3A_MOUSE
|
||||||
| NC score | 0.843790 (rank : 50) | θ value | 5.43371e-13 (rank : 36) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | P35582, Q61108 | Gene names | Foxa1, Hnf3a, Tcf-3a, Tcf3a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-alpha (HNF-3A) (Forkhead box protein A1). | |||||
|
HNF3B_HUMAN
|
||||||
| NC score | 0.843664 (rank : 51) | θ value | 6.00763e-12 (rank : 58) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9Y261, Q8WUW4, Q96DF7 | Gene names | FOXA2, HNF3B, TCF3B | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-beta (HNF-3B) (Forkhead box protein A2). | |||||
|
FOXB2_MOUSE
|
||||||
| NC score | 0.843258 (rank : 52) | θ value | 3.29651e-10 (rank : 69) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q64733 | Gene names | Foxb2, Fkh4 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein B2 (Transcription factor FKH-4). | |||||
|
FOXB1_HUMAN
|
||||||
| NC score | 0.842181 (rank : 53) | θ value | 6.00763e-12 (rank : 56) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q99853, O60652, O75917 | Gene names | FOXB1, FKH5 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein B1 (Transcription factor FKH-5). | |||||
|
HNF3A_HUMAN
|
||||||
| NC score | 0.841621 (rank : 54) | θ value | 7.09661e-13 (rank : 40) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | P55317, Q9H2A0 | Gene names | FOXA1, HNF3A, TCF3A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-alpha (HNF-3A) (Forkhead box protein A1). | |||||
|
FREA_MOUSE
|
||||||
| NC score | 0.840207 (rank : 55) | θ value | 1.47974e-10 (rank : 67) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | Q61574, Q8C5N7 | Gene names | Fkhl18, Fkh3, Freac10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead-related transcription factor 10 (FREAC-10) (Forkhead-like 18 protein) (Transcription factor FKH-3). | |||||
|
FREA_HUMAN
|
||||||
| NC score | 0.838583 (rank : 56) | θ value | 1.74796e-11 (rank : 61) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 93 | |
| SwissProt Accessions | O43638, Q96D28 | Gene names | FKHL18, FREAC10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead-related transcription factor 10 (FREAC-10) (Forkhead-like 18 protein). | |||||
|
FOXM1_MOUSE
|
||||||
| NC score | 0.832644 (rank : 57) | θ value | 9.26847e-13 (rank : 43) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | O08696 | Gene names | Foxm1, Fkh16 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead homolog 16) (Winged-helix transcription factor Trident). | |||||
|
FOXD4_MOUSE
|
||||||
| NC score | 0.832370 (rank : 58) | θ value | 7.34386e-10 (rank : 72) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q60688, Q61573 | Gene names | Foxd4, Fkh2, Fkhl9, Freac5 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D4 (Forkhead-related protein FKHL9) (Forkhead- related transcription factor 5) (FREAC-5) (Transcription factor FKH- 2). | |||||
|
FOXD4_HUMAN
|
||||||
| NC score | 0.829479 (rank : 59) | θ value | 1.47974e-10 (rank : 66) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q12950, Q5VVK1, Q8WXT6 | Gene names | FOXD4, FKHL9, FOXD4A, FREAC5 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D4 (Forkhead-related protein FKHL9) (Forkhead- related transcription factor 5) (FREAC-5) (Myeloid factor-alpha). | |||||
|
FX4L4_HUMAN
|
||||||
| NC score | 0.829361 (rank : 60) | θ value | 5.62301e-10 (rank : 71) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q8WXT5 | Gene names | FOXD4L4, FOXD4B | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D4-like 4 (Forkhead box protein D4B) (Myeloid factor-gamma). | |||||
|
FX4L1_HUMAN
|
||||||
| NC score | 0.829344 (rank : 61) | θ value | 5.62301e-10 (rank : 70) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q9NU39 | Gene names | FOXD4L1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D4-like 1. | |||||
|
FOXM1_HUMAN
|
||||||
| NC score | 0.825369 (rank : 62) | θ value | 9.26847e-13 (rank : 42) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q08050, O43258, O43259, O43260, Q4ZGG7, Q9BRL2 | Gene names | FOXM1, FKHL16, HFH11, MPP2, WIN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead-related protein FKHL16) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/fork-head homolog 11) (HFH-11) (Winged helix factor from INS-1 cells) (M-phase phosphoprotein 2) (MPM-2 reactive phosphoprotein 2) (Transcription factor Trident). | |||||
|
FOXP3_HUMAN
|
||||||
| NC score | 0.817908 (rank : 63) | θ value | 1.02475e-11 (rank : 59) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q9BZS1, O60827, Q4ZH51 | Gene names | FOXP3, IPEX | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P3 (Scurfin). | |||||
|
FOXP3_MOUSE
|
||||||
| NC score | 0.813948 (rank : 64) | θ value | 6.64225e-11 (rank : 65) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q99JB6 | Gene names | Foxp3 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P3 (Scurfin). | |||||
|
FOXH1_MOUSE
|
||||||
| NC score | 0.810278 (rank : 65) | θ value | 1.17247e-07 (rank : 76) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | O88621, Q9QZL5, Q9R241 | Gene names | Foxh1, Fast1, Fast2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein H1 (Forkhead activin signal transducer 1) (Fast- 1) (Forkhead activin signal transducer 2) (Fast-2). | |||||
|
FOXH1_HUMAN
|
||||||
| NC score | 0.806714 (rank : 66) | θ value | 1.17247e-07 (rank : 75) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 99 | |
| SwissProt Accessions | O75593 | Gene names | FOXH1, FAST1, FAST2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein H1 (Forkhead activin signal transducer 1) (Fast- 1) (hFAST-1) (Forkhead activin signal transducer 2) (Fast-2). | |||||
|
FOXE2_HUMAN
|
||||||
| NC score | 0.802205 (rank : 67) | θ value | 7.59969e-07 (rank : 77) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | Q99526 | Gene names | FOXE2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein E2 (HNF-3/fork head-like protein 5) (HFKL5) (HFKH4). | |||||
|
FOXN4_MOUSE
|
||||||
| NC score | 0.801945 (rank : 68) | θ value | 6.21693e-09 (rank : 74) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q8K3Q3, Q920C0 | Gene names | Foxn4 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N4. | |||||
|
FOXN4_HUMAN
|
||||||
| NC score | 0.799028 (rank : 69) | θ value | 2.13673e-09 (rank : 73) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 100 | |
| SwissProt Accessions | Q96NZ1, Q6ZMR4, Q96NZ0 | Gene names | FOXN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein N4. | |||||
|
FOXN1_MOUSE
|
||||||
| NC score | 0.791723 (rank : 70) | θ value | 6.64225e-11 (rank : 64) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 100 | |
| SwissProt Accessions | Q61575 | Gene names | Foxn1, Fkh19, Hfh11, Whn | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N1 (Transcription factor winged-helix nude) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/forkhead homolog 11) (HFH-11). | |||||
|
FOXP1_HUMAN
|
||||||
| NC score | 0.790664 (rank : 71) | θ value | 4.59992e-12 (rank : 55) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9H334, Q9H332, Q9H333, Q9P0R1 | Gene names | FOXP1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1. | |||||
|
FOXP4_MOUSE
|
||||||
| NC score | 0.788447 (rank : 72) | θ value | 1.33837e-11 (rank : 60) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | Q9DBY0, Q80V92, Q8CG10, Q8CIS1 | Gene names | Foxp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein P4 (Fork head-related protein like A) (mFKHLA). | |||||
|
FOXP4_HUMAN
|
||||||
| NC score | 0.788320 (rank : 73) | θ value | 1.58096e-12 (rank : 50) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q8IVH2, Q96E19 | Gene names | FOXP4, FKHLA | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P4 (Fork head-related protein-like A). | |||||
|
FOXP1_MOUSE
|
||||||
| NC score | 0.782322 (rank : 74) | θ value | 2.69671e-12 (rank : 54) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | P58462 | Gene names | Foxp1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1 (Forkhead-related transcription factor 1). | |||||
|
FOXN1_HUMAN
|
||||||
| NC score | 0.781881 (rank : 75) | θ value | 2.52405e-10 (rank : 68) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 107 | |
| SwissProt Accessions | O15353, O15352 | Gene names | FOXN1, WHN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N1 (Transcription factor winged-helix nude). | |||||
|
FOXR2_MOUSE
|
||||||
| NC score | 0.769572 (rank : 76) | θ value | 2.21117e-06 (rank : 78) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q3UM89 | Gene names | Foxr2, Foxn6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein R2 (Forkhead box protein N6). | |||||
|
FOXR1_MOUSE
|
||||||
| NC score | 0.762132 (rank : 77) | θ value | 3.77169e-06 (rank : 79) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q3UTB7 | Gene names | Foxr1, Foxn5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein R1 (Forkhead box protein N5). | |||||
|
FOXP2_HUMAN
|
||||||
| NC score | 0.748795 (rank : 78) | θ value | 3.89403e-11 (rank : 62) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | O15409, Q6ZND1, Q75MJ3, Q8IZE0, Q8N0W2, Q8N6B7, Q8N6B8, Q8NFQ1, Q8NFQ2, Q8NFQ3, Q8NFQ4, Q8TD74 | Gene names | FOXP2, CAGH44, TNRC10 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P2 (CAG repeat protein 44) (Trinucleotide repeat- containing gene 10 protein). | |||||
|
FOXP2_MOUSE
|
||||||
| NC score | 0.746597 (rank : 79) | θ value | 3.89403e-11 (rank : 63) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | P58463, Q6PD37, Q8C4F0, Q8R441 | Gene names | Foxp2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P2. | |||||
|
FOXR1_HUMAN
|
||||||
| NC score | 0.743744 (rank : 80) | θ value | 7.1131e-05 (rank : 80) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q6PIV2, Q86UT9, Q8IXX2 | Gene names | FOXR1, DLNB13, FOXN5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein R1 (Forkhead box protein N5). | |||||
|
FOXR2_HUMAN
|
||||||
| NC score | 0.630047 (rank : 81) | θ value | 0.0252991 (rank : 82) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q6PJQ5 | Gene names | FOXR2, FOXN6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein R2 (Forkhead box protein N6). | |||||
|
CCD72_HUMAN
|
||||||
| NC score | 0.141099 (rank : 82) | θ value | 3.0926 (rank : 128) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2S6, Q9P052 | Gene names | CCDC72 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 72. | |||||
|
CCD72_MOUSE
|
||||||
| NC score | 0.141099 (rank : 83) | θ value | 3.0926 (rank : 129) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K003 | Gene names | Ccdc72 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 72. | |||||
|
CH013_HUMAN
|
||||||
| NC score | 0.079283 (rank : 84) | θ value | 0.0736092 (rank : 85) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96KS9, Q8N3M3, Q9NSR0 | Gene names | C8orf13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C8orf13. | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.057105 (rank : 85) | θ value | 3.0926 (rank : 141) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 102 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.052252 (rank : 86) | θ value | 0.0330416 (rank : 84) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.049268 (rank : 87) | θ value | 0.365318 (rank : 94) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.042037 (rank : 88) | θ value | 1.38821 (rank : 108) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.041666 (rank : 89) | θ value | 5.27518 (rank : 152) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.039924 (rank : 90) | θ value | 8.99809 (rank : 190) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
SELV_HUMAN
|
||||||
| NC score | 0.037958 (rank : 91) | θ value | 1.06291 (rank : 107) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P59797 | Gene names | SELV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Selenoprotein V. | |||||
|
CS016_HUMAN
|
||||||
| NC score | 0.037208 (rank : 92) | θ value | 0.62314 (rank : 98) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8ND99 | Gene names | C19orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf16. | |||||
|
CO039_MOUSE
|
||||||
| NC score | 0.035886 (rank : 93) | θ value | 0.62314 (rank : 97) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
GSCR1_HUMAN
|
||||||
| NC score | 0.035843 (rank : 94) | θ value | 3.0926 (rank : 138) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.033961 (rank : 95) | θ value | 1.81305 (rank : 121) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
H12_HUMAN
|
||||||
| NC score | 0.033254 (rank : 96) | θ value | 0.125558 (rank : 88) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P16403 | Gene names | HIST1H1C, H1F2 | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.2 (Histone H1d). | |||||
|
HNF1A_MOUSE
|
||||||
| NC score | 0.033176 (rank : 97) | θ value | 0.813845 (rank : 104) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
WASIP_MOUSE
|
||||||
| NC score | 0.033090 (rank : 98) | θ value | 4.03905 (rank : 150) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
SFR15_HUMAN
|
||||||
| NC score | 0.031906 (rank : 99) | θ value | 0.813845 (rank : 105) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O95104, Q6P1M5, Q8N3I8, Q9UFM1, Q9ULP8 | Gene names | SFRS15, KIAA1172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 15 (CTD-binding SR-like protein RA4). | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.031664 (rank : 100) | θ value | 1.81305 (rank : 117) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.031455 (rank : 101) | θ value | 8.99809 (rank : 177) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
NOL3_MOUSE
|
||||||
| NC score | 0.031257 (rank : 102) | θ value | 0.365318 (rank : 93) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D1X0 | Gene names | Nol3 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar protein 3. | |||||
|
H15_MOUSE
|
||||||
| NC score | 0.031099 (rank : 103) | θ value | 0.0961366 (rank : 86) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P43276, Q9CRM8 | Gene names | Hist1h1b, H1f5 | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.5 (H1 VAR.5) (H1b). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.030894 (rank : 104) | θ value | 5.27518 (rank : 164) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
DGKK_HUMAN
|
||||||
| NC score | 0.030732 (rank : 105) | θ value | 0.0330416 (rank : 83) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
K0460_MOUSE
|
||||||
| NC score | 0.029590 (rank : 106) | θ value | 0.47712 (rank : 95) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6NXI6, Q3U3L8, Q6ZQA7 | Gene names | Kiaa0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
CNOT4_MOUSE
|
||||||
| NC score | 0.028962 (rank : 107) | θ value | 1.38821 (rank : 109) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
CS007_MOUSE
|
||||||
| NC score | 0.028949 (rank : 108) | θ value | 0.279714 (rank : 90) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
FIP1_HUMAN
|
||||||
| NC score | 0.027617 (rank : 109) | θ value | 2.36792 (rank : 123) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6UN15, Q499Y4, Q49AU3, Q7Z608, Q8WVN3, Q96F80, Q9H077 | Gene names | FIP1L1, FIP1, RHE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA 3'-end-processing factor FIP1 (FIP1-like 1) (Factor interacting with PAP) (hFip1) (Rearranged in hypereosinophilia). | |||||
|
H11_MOUSE
|
||||||
| NC score | 0.027156 (rank : 110) | θ value | 0.813845 (rank : 103) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P43275 | Gene names | Hist1h1a, H1f1 | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.1 (H1 VAR.3) (H1a). | |||||
|
RAI2_MOUSE
|
||||||
| NC score | 0.026651 (rank : 111) | θ value | 1.38821 (rank : 113) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QVY8 | Gene names | Rai2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2 (3f8). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.026063 (rank : 112) | θ value | 0.62314 (rank : 96) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
RAI2_HUMAN
|
||||||
| NC score | 0.025777 (rank : 113) | θ value | 2.36792 (rank : 126) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y5P3, Q8N6X7 | Gene names | RAI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2. | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.025605 (rank : 114) | θ value | 6.88961 (rank : 174) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
HNF1A_HUMAN
|
||||||
| NC score | 0.025482 (rank : 115) | θ value | 3.0926 (rank : 139) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P20823, Q99861 | Gene names | TCF1, HNF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1) (Transcription factor 1) (TCF-1). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.024990 (rank : 116) | θ value | 5.27518 (rank : 163) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
FIP1_MOUSE
|
||||||
| NC score | 0.024328 (rank : 117) | θ value | 4.03905 (rank : 147) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D824, Q8BWX7, Q99LH0, Q9DBB2 | Gene names | Fip1l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA 3'-end-processing factor FIP1 (FIP1-like 1). | |||||
|
PTN23_MOUSE
|
||||||
| NC score | 0.024217 (rank : 118) | θ value | 0.279714 (rank : 91) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
PKHG4_HUMAN
|
||||||
| NC score | 0.023756 (rank : 119) | θ value | 1.06291 (rank : 106) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q58EX7, Q4G0J8, Q4H485, Q56A69, Q9H7K4, Q9UFW0 | Gene names | PLEKHG4, PRTPHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Puratrophin-1 (Pleckstrin homology domain-containing family G member 4) (Purkinje cell atrophy-associated protein 1). | |||||
|
XG_HUMAN
|
||||||
| NC score | 0.023720 (rank : 120) | θ value | 4.03905 (rank : 151) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55808, Q71BZ5 | Gene names | XG, PBDX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glycoprotein Xg precursor (Protein PBDX). | |||||
|
DHX37_HUMAN
|
||||||
| NC score | 0.023195 (rank : 121) | θ value | 0.0148317 (rank : 81) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IY37, Q9BUI7, Q9P211 | Gene names | DHX37, DDX37, KIAA1517 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DHX37 (EC 3.6.1.-) (DEAH box protein 37). | |||||
|
GGN_MOUSE
|
||||||
| NC score | 0.022580 (rank : 122) | θ value | 3.0926 (rank : 136) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
CS007_HUMAN
|
||||||
| NC score | 0.022330 (rank : 123) | θ value | 6.88961 (rank : 170) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
DOT1L_HUMAN
|
||||||
| NC score | 0.022029 (rank : 124) | θ value | 3.0926 (rank : 132) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
PHLA1_HUMAN
|
||||||
| NC score | 0.021418 (rank : 125) | θ value | 8.99809 (rank : 187) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8WV24, Q15184, Q2TAN2, Q9NZ17 | Gene names | PHLDA1, PHRIP, TDAG51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Apoptosis-associated nuclear protein) (Proline- and histidine-rich protein) (Proline- and glutamine-rich protein) (PQ-rich protein). | |||||
|
CABL2_HUMAN
|
||||||
| NC score | 0.021289 (rank : 126) | θ value | 0.813845 (rank : 102) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BTV7, Q5JWL0, Q9BYK0 | Gene names | CABLES2, C20orf150 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 and ABL1 enzyme substrate 2 (Interactor with CDK3 2) (Ik3-2). | |||||
|
ESPL1_HUMAN
|
||||||
| NC score | 0.020786 (rank : 127) | θ value | 1.38821 (rank : 110) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14674 | Gene names | ESPL1, ESP1, KIAA0165 | |||
|
Domain Architecture |

|
|||||
| Description | Separin (EC 3.4.22.49) (Separase) (Caspase-like protein ESPL1) (Extra spindle poles-like 1 protein). | |||||
|
H10_HUMAN
|
||||||
| NC score | 0.020419 (rank : 128) | θ value | 6.88961 (rank : 172) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P07305 | Gene names | H1F0, H1FV | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.0 (H1(0)) (Histone H1'). | |||||
|
FMN1B_MOUSE
|
||||||
| NC score | 0.019545 (rank : 129) | θ value | 3.0926 (rank : 134) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q05859 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoform IV (Limb deformity protein). | |||||
|
MMP9_MOUSE
|
||||||
| NC score | 0.019402 (rank : 130) | θ value | 0.0961366 (rank : 87) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P41245, Q06788, Q9DC02 | Gene names | Mmp9, Clg4b | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-9 precursor (EC 3.4.24.35) (MMP-9) (92 kDa type IV collagenase) (92 kDa gelatinase) (Gelatinase B) (GELB). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.019070 (rank : 131) | θ value | 2.36792 (rank : 125) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
FMN1A_MOUSE
|
||||||
| NC score | 0.019056 (rank : 132) | θ value | 3.0926 (rank : 133) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q05860 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoforms I/II/III (Limb deformity protein). | |||||
|
BAG3_MOUSE
|
||||||
| NC score | 0.018978 (rank : 133) | θ value | 4.03905 (rank : 143) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
LRP5_MOUSE
|
||||||
| NC score | 0.018848 (rank : 134) | θ value | 0.62314 (rank : 100) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q91VN0, O88883, Q9R208 | Gene names | Lrp5, Lrp7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 5 precursor (LRP7) (Lr3). | |||||
|
SGIP1_MOUSE
|
||||||
| NC score | 0.018774 (rank : 135) | θ value | 1.38821 (rank : 115) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8VD37, Q3UFU3, Q3UGA0, Q8BXX4, Q8C034, Q9CXT2 | Gene names | Sgip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3-containing GRB2-like protein 3-interacting protein 1 (Endophilin- 3-interacting protein). | |||||
|
SAPS1_HUMAN
|
||||||
| NC score | 0.018337 (rank : 136) | θ value | 5.27518 (rank : 162) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
K0460_HUMAN
|
||||||
| NC score | 0.018276 (rank : 137) | θ value | 8.99809 (rank : 182) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5VT52, O75048, Q5VT53, Q6MZL4, Q86XD2 | Gene names | KIAA0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
FMNL_HUMAN
|
||||||
| NC score | 0.018257 (rank : 138) | θ value | 8.99809 (rank : 179) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
Domain Architecture |

|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
SDC3_MOUSE
|
||||||
| NC score | 0.017281 (rank : 139) | θ value | 1.81305 (rank : 120) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q64519, Q3UDD9, Q6ZQA4, Q7TQD4 | Gene names | Sdc3, Kiaa0468 | |||
|
Domain Architecture |

|
|||||
| Description | Syndecan-3 precursor (SYND3). | |||||
|
SFPQ_HUMAN
|
||||||
| NC score | 0.017018 (rank : 140) | θ value | 8.99809 (rank : 194) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.016973 (rank : 141) | θ value | 1.38821 (rank : 114) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
CDN1C_HUMAN
|
||||||
| NC score | 0.016944 (rank : 142) | θ value | 3.0926 (rank : 130) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P49918 | Gene names | CDKN1C, KIP2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
PTN23_HUMAN
|
||||||
| NC score | 0.015965 (rank : 143) | θ value | 1.81305 (rank : 119) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
CCD37_HUMAN
|
||||||
| NC score | 0.015806 (rank : 144) | θ value | 2.36792 (rank : 122) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q494V2, Q494V1, Q494V4, Q8N838 | Gene names | CCDC37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 37. | |||||
|
PHLA1_MOUSE
|
||||||
| NC score | 0.015024 (rank : 145) | θ value | 8.99809 (rank : 188) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q62392, Q3TY48, Q3UG87 | Gene names | Phlda1, Tdag51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Proline- and glutamine-rich protein) (PQR protein). | |||||
|
IF4G3_HUMAN
|
||||||
| NC score | 0.013831 (rank : 146) | θ value | 0.62314 (rank : 99) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43432, Q15597, Q8NEN1 | Gene names | EIF4G3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.013819 (rank : 147) | θ value | 5.27518 (rank : 160) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
FRAT1_MOUSE
|
||||||
| NC score | 0.013115 (rank : 148) | θ value | 3.0926 (rank : 135) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70339 | Gene names | Frat1 | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene FRAT1 (Frequently rearranged in advanced T-cell lymphomas) (Frat-1). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.012385 (rank : 149) | θ value | 6.88961 (rank : 173) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
CEP68_HUMAN
|
||||||
| NC score | 0.012250 (rank : 150) | θ value | 4.03905 (rank : 145) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q76N32, O60326, Q9BQ18, Q9UDM9 | Gene names | CEP68, KIAA0582 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 68 kDa (Cep68 protein). | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.012172 (rank : 151) | θ value | 2.36792 (rank : 124) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MUC2_HUMAN
|
||||||
| NC score | 0.011428 (rank : 152) | θ value | 1.38821 (rank : 112) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
SH22A_HUMAN
|
||||||
| NC score | 0.011011 (rank : 153) | θ value | 4.03905 (rank : 149) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NP31, O43817, Q9UPA7 | Gene names | SH2D2A, SCAP, TSAD, VRAP | |||
|
Domain Architecture |

|
|||||
| Description | SH2 domain protein 2A (T cell-specific adapter protein) (TSAd) (VEGF receptor-associated protein) (SH2 domain-containing adapter protein). | |||||
|
GCNL2_HUMAN
|
||||||
| NC score | 0.010680 (rank : 154) | θ value | 5.27518 (rank : 158) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92830, Q9UCW1 | Gene names | GCN5L2, GCN5, HGCN5 | |||
|
Domain Architecture |

|
|||||
| Description | General control of amino acid synthesis protein 5-like 2 (EC 2.3.1.48) (Histone acetyltransferase GCN5) (hsGCN5) (STAF97). | |||||
|
BSPRY_MOUSE
|
||||||
| NC score | 0.010296 (rank : 155) | θ value | 0.813845 (rank : 101) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80YW5, Q3TU74, Q8BZF0, Q99KV7, Q9ER70 | Gene names | Bspry | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B box and SPRY domain-containing protein. | |||||
|
MMP9_HUMAN
|
||||||
| NC score | 0.010141 (rank : 156) | θ value | 8.99809 (rank : 186) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P14780, Q3LR70, Q8N725, Q9H4Z1 | Gene names | MMP9, CLG4B | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-9 precursor (EC 3.4.24.35) (MMP-9) (92 kDa type IV collagenase) (92 kDa gelatinase) (Gelatinase B) (GELB) [Contains: 67 kDa matrix metalloproteinase-9; 82 kDa matrix metalloproteinase-9]. | |||||
|
DIAP2_MOUSE
|
||||||
| NC score | 0.010090 (rank : 157) | θ value | 5.27518 (rank : 156) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O70566, Q8C2G8 | Gene names | Diaph2, Diap2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2) (mDia3). | |||||
|
LRP5_HUMAN
|
||||||
| NC score | 0.009896 (rank : 158) | θ value | 8.99809 (rank : 183) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75197, Q96TD6, Q9UP66 | Gene names | LRP5, LRP7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 5 precursor. | |||||
|
LPIN2_MOUSE
|
||||||
| NC score | 0.009828 (rank : 159) | θ value | 1.81305 (rank : 118) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PI5, Q8C357, Q8C7I8, Q8CC85, Q8CHR7 | Gene names | Lpin2 | |||
|
Domain Architecture |

|
|||||
| Description | Lipin-2. | |||||
|
MEF2B_HUMAN
|
||||||
| NC score | 0.009687 (rank : 160) | θ value | 8.99809 (rank : 184) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q02080 | Gene names | MEF2B, XMEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2B (Serum response factor-like protein 2) (XMEF2) (RSRFR2). | |||||
|
ARTN_HUMAN
|
||||||
| NC score | 0.008130 (rank : 161) | θ value | 6.88961 (rank : 166) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5T4W7, O95441, O96030, Q6P6A3 | Gene names | ARTN, EVN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Artemin precursor (Enovin) (Neublastin). | |||||
|
CN092_HUMAN
|
||||||
| NC score | 0.008122 (rank : 162) | θ value | 4.03905 (rank : 146) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O94842 | Gene names | C14orf92, KIAA0737 | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal Langerhans cell protein LCP1. | |||||
|
DIAP1_HUMAN
|
||||||
| NC score | 0.007883 (rank : 163) | θ value | 8.99809 (rank : 178) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O60610, Q9UC76 | Gene names | DIAPH1, DIAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1). | |||||
|
MOT8_MOUSE
|
||||||
| NC score | 0.007820 (rank : 164) | θ value | 5.27518 (rank : 161) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O70324, Q8K3S9 | Gene names | Slc16a2, Mct8, Xpct | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Monocarboxylate transporter 8 (MCT 8) (Solute carrier family 16 member 2) (X-linked PEST-containing transporter). | |||||
|
CSPG4_HUMAN
|
||||||
| NC score | 0.007645 (rank : 165) | θ value | 3.0926 (rank : 131) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6UVK1, Q92675 | Gene names | CSPG4, MCSP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 4 precursor (Chondroitin sulfate proteoglycan NG2) (Melanoma chondroitin sulfate proteoglycan) (Melanoma-associated chondroitin sulfate proteoglycan). | |||||
|
GASP1_MOUSE
|
||||||
| NC score | 0.006854 (rank : 166) | θ value | 8.99809 (rank : 180) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5U4C1, Q8BKR8, Q8BUN4, Q8BYK9, Q8CHF4, Q8R095 | Gene names | Gprasp1, Kiaa0443 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 1 (GASP-1). | |||||
|
MYCB2_HUMAN
|
||||||
| NC score | 0.006811 (rank : 167) | θ value | 4.03905 (rank : 148) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O75592, Q5JSX8, Q5VZN6, Q6PIB6, Q9UQ11, Q9Y6E4 | Gene names | MYCBP2, KIAA0916, PAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
BRD4_MOUSE
|
||||||
| NC score | 0.006754 (rank : 168) | θ value | 4.03905 (rank : 144) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
SMG7_MOUSE
|
||||||
| NC score | 0.006417 (rank : 169) | θ value | 8.99809 (rank : 195) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5RJH6, Q63ZW5, Q6ZQF3 | Gene names | Smg7, Est1c, Kiaa0250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
GATA3_HUMAN
|
||||||
| NC score | 0.006065 (rank : 170) | θ value | 5.27518 (rank : 157) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P23771, Q96J16 | Gene names | GATA3 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-acting T-cell-specific transcription factor GATA-3 (GATA-binding factor 3). | |||||
|
RHG09_HUMAN
|
||||||
| NC score | 0.005951 (rank : 171) | θ value | 2.36792 (rank : 127) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BRR9, Q8TCJ3, Q8WYR0, Q96EZ2, Q96S74 | Gene names | ARHGAP9 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 9. | |||||
|
LTBP4_MOUSE
|
||||||
| NC score | 0.004883 (rank : 172) | θ value | 0.365318 (rank : 92) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8K4G1, Q8K4G0 | Gene names | Ltbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 4 precursor (LTBP-4). | |||||
|
SEM6D_MOUSE
|
||||||
| NC score | 0.003773 (rank : 173) | θ value | 8.99809 (rank : 193) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q76KF0, Q76KF1, Q76KF2, Q76KF3, Q76KF4, Q80TD0 | Gene names | Sema6d, Kiaa1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
SEM6D_HUMAN
|
||||||
| NC score | 0.003615 (rank : 174) | θ value | 8.99809 (rank : 192) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NFY4, Q8NFY3, Q8NFY5, Q8NFY6, Q8NFY7, Q9P249 | Gene names | SEMA6D, KIAA1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
GGA3_HUMAN
|
||||||
| NC score | 0.003602 (rank : 175) | θ value | 8.99809 (rank : 181) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZ52, Q15017, Q6IS16, Q9UJY3 | Gene names | GGA3, KIAA0154 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-binding protein GGA3 (Golgi-localized, gamma ear-containing, ARF-binding protein 3). | |||||
|
PININ_MOUSE
|
||||||
| NC score | 0.002305 (rank : 176) | θ value | 8.99809 (rank : 189) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
2DMA_MOUSE
|
||||||
| NC score | 0.002243 (rank : 177) | θ value | 8.99809 (rank : 175) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28078 | Gene names | H2-DMa, H2-Ma, Ma | |||
|
Domain Architecture |

|
|||||
| Description | Class II histocompatibility antigen, M alpha chain precursor. | |||||
|
CF182_MOUSE
|
||||||
| NC score | 0.001955 (rank : 178) | θ value | 5.27518 (rank : 154) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 551 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VDS7, Q9CZE0, Q9D5A1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf182 homolog. | |||||
|
ASPP2_HUMAN
|
||||||
| NC score | 0.001154 (rank : 179) | θ value | 8.99809 (rank : 176) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q13625, Q12892, Q86X75, Q96KQ3 | Gene names | TP53BP2, ASPP2, BBP | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 2 (Tumor suppressor p53-binding protein 2) (p53-binding protein 2) (p53BP2) (53BP2) (Bcl2-binding protein) (Bbp) (NY-REN-51 antigen). | |||||
|
ROBO3_MOUSE
|
||||||
| NC score | 0.001083 (rank : 180) | θ value | 3.0926 (rank : 142) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z2I4 | Gene names | Robo3, Rbig1 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Retinoblastoma-inhibiting gene 1) (Rig-1). | |||||
|
ADA19_HUMAN
|
||||||
| NC score | 0.000383 (rank : 181) | θ value | 6.88961 (rank : 165) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
ATBF1_MOUSE
|
||||||
| NC score | 0.000229 (rank : 182) | θ value | 6.88961 (rank : 168) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
KTNB1_HUMAN
|
||||||
| NC score | 0.000059 (rank : 183) | θ value | 1.38821 (rank : 111) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BVA0, O60620 | Gene names | KATNB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Katanin p80 WD40-containing subunit B1 (Katanin p80 subunit B1) (p80 katanin). | |||||
|
ATBF1_HUMAN
|
||||||
| NC score | -0.000419 (rank : 184) | θ value | 6.88961 (rank : 167) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
CRUM2_HUMAN
|
||||||
| NC score | -0.000559 (rank : 185) | θ value | 5.27518 (rank : 155) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5IJ48, Q5JS41, Q5JS43, Q6ZTA9, Q6ZWI6 | Gene names | CRB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crumbs homolog 2 precursor (Crumbs-like protein 2). | |||||
|
GP1BA_HUMAN
|
||||||
| NC score | -0.000986 (rank : 186) | θ value | 6.88961 (rank : 171) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P07359, Q14441, Q16469, Q8N1F3, Q8NG39, Q9HDC7, Q9UEK1, Q9UQS4 | Gene names | GP1BA | |||
|
Domain Architecture |

|
|||||
| Description | Platelet glycoprotein Ib alpha chain precursor (Glycoprotein Ibalpha) (GP-Ib alpha) (GPIbA) (GPIb-alpha) (Antigen CD42b-alpha) (CD42b antigen) [Contains: Glycocalicin]. | |||||
|
ROBO2_MOUSE
|
||||||
| NC score | -0.001726 (rank : 187) | θ value | 8.99809 (rank : 191) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TPD3, Q8BJ59 | Gene names | Robo2, Kiaa1568 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 2 precursor. | |||||
|
GPR37_MOUSE
|
||||||
| NC score | -0.001748 (rank : 188) | θ value | 3.0926 (rank : 137) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QY42, Q8BKF8, Q8BXP9, Q8C022, Q9Z0G3 | Gene names | Gpr37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 37 precursor. | |||||
|
ZBT12_HUMAN
|
||||||
| NC score | -0.002828 (rank : 189) | θ value | 0.125558 (rank : 89) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y330, Q5JQ98 | Gene names | ZBTB12, C6orf46, G10, NG35 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 12 (Protein G10). | |||||
|
ZN295_HUMAN
|
||||||
| NC score | -0.003196 (rank : 190) | θ value | 1.38821 (rank : 116) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULJ3, Q6P4R0 | Gene names | ZNF295, KIAA1227, ZBTB21 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 295 (Zinc finger and BTB domain-containing protein 21). | |||||
|
SPEG_HUMAN
|
||||||
| NC score | -0.003583 (rank : 191) | θ value | 8.99809 (rank : 196) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
WNK2_HUMAN
|
||||||
| NC score | -0.003808 (rank : 192) | θ value | 8.99809 (rank : 197) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
KLF13_HUMAN
|
||||||
| NC score | -0.004201 (rank : 193) | θ value | 3.0926 (rank : 140) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y2Y9, Q9Y356 | Gene names | KLF13, BTEB3, NSLP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 13 (Transcription factor BTEB3) (Basic transcription element-binding protein 3) (BTE-binding protein 3) (RANTES factor of late activated T-lymphocytes 1) (RFLAT-1) (Transcription factor NSLP1) (Novel Sp1-like zinc finger transcription factor 1). | |||||
|
MK07_MOUSE
|
||||||
| NC score | -0.004270 (rank : 194) | θ value | 5.27518 (rank : 159) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1054 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9WVS8 | Gene names | Mapk7, Erk5 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (BMK1 kinase). | |||||
|
MK07_HUMAN
|
||||||
| NC score | -0.005012 (rank : 195) | θ value | 8.99809 (rank : 185) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 1087 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q13164, Q16634 | Gene names | MAPK7, ERK4, ERK5, PRKM7 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (ERK4) (BMK1 kinase). | |||||
|
CD2L6_HUMAN
|
||||||
| NC score | -0.005739 (rank : 196) | θ value | 5.27518 (rank : 153) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BWU1, Q5JQZ7, Q5JR00, Q8TC78, Q9UPX2 | Gene names | CDC2L6, CDK11, KIAA1028 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 6 (EC 2.7.11.22) (CDC2- related protein kinase 6) (Death-preventing kinase) (Cyclin-dependent kinase 11). | |||||
|
CD2L5_HUMAN
|
||||||
| NC score | -0.006748 (rank : 197) | θ value | 6.88961 (rank : 169) | |||
| Query Neighborhood Hits | 197 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14004, Q6DKQ9, Q75MH4, Q75MH5, Q96JN4, Q9H4A0, Q9H4A1, Q9UDR4 | Gene names | CDC2L5, CDC2L, CHED, KIAA1791 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5) (Cholinesterase-related cell division controller). | |||||