Please be patient as the page loads
|
CP053_HUMAN
|
||||||
| SwissProt Accessions | Q9BTK6 | Gene names | C16orf53 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf53. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CP053_HUMAN
|
||||||
| θ value | 1.39067e-109 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9BTK6 | Gene names | C16orf53 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf53. | |||||
|
CP053_MOUSE
|
||||||
| θ value | 1.3939e-101 (rank : 2) | NC score | 0.977074 (rank : 2) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99L02 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf53 homolog. | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 3) | NC score | 0.102712 (rank : 4) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 4) | NC score | 0.118047 (rank : 3) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 5) | NC score | 0.085855 (rank : 6) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 6) | NC score | 0.090089 (rank : 5) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
LAP4_HUMAN
|
||||||
| θ value | 0.163984 (rank : 7) | NC score | 0.024536 (rank : 35) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 608 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14160, Q6P496, Q7Z5D1, Q8WWV8, Q96C69, Q96GG1 | Gene names | SCRIB, CRIB1, KIAA0147, LAP4, SCRB1, VARTUL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog) (hScrib). | |||||
|
RIOK2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 8) | NC score | 0.056679 (rank : 17) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CQS5, Q91XF3 | Gene names | Riok2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase RIO2 (EC 2.7.11.1) (RIO kinase 2). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 0.813845 (rank : 9) | NC score | 0.055270 (rank : 19) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
F10A4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 10) | NC score | 0.073030 (rank : 9) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IZP2 | Gene names | FAM10A4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A4. | |||||
|
ZC3H6_MOUSE
|
||||||
| θ value | 1.06291 (rank : 11) | NC score | 0.040766 (rank : 26) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BYK8, Q80UK3, Q8C1H1, Q9D604 | Gene names | Zc3h6, Zc3hdc6 | |||
|
Domain Architecture |
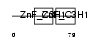
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
ABL1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 12) | NC score | 0.007664 (rank : 49) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1220 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P00519, Q13869, Q13870, Q16133, Q45F09 | Gene names | ABL1, ABL, JTK7 | |||
|
Domain Architecture |
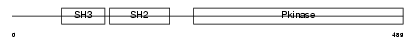
|
|||||
| Description | Proto-oncogene tyrosine-protein kinase ABL1 (EC 2.7.10.2) (p150) (c- ABL) (Abelson murine leukemia viral oncogene homolog 1). | |||||
|
F10A1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 13) | NC score | 0.072927 (rank : 10) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P50502, O14999 | Gene names | ST13, FAM10A1, HIP, SNC6 | |||
|
Domain Architecture |
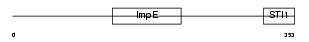
|
|||||
| Description | Hsc70-interacting protein (Hip) (Suppression of tumorigenicity protein 13) (Putative tumor suppressor ST13) (Protein FAM10A1) (Progesterone receptor-associated p48 protein) (NY-REN-33 antigen). | |||||
|
F10A5_HUMAN
|
||||||
| θ value | 2.36792 (rank : 14) | NC score | 0.070637 (rank : 12) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NFI4 | Gene names | FAM10A5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A5. | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 15) | NC score | 0.056069 (rank : 18) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
FA13C_HUMAN
|
||||||
| θ value | 3.0926 (rank : 16) | NC score | 0.040949 (rank : 25) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NE31, Q99787 | Gene names | FAM13C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
TSSK1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 17) | NC score | 0.009256 (rank : 45) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 840 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BXA7 | Gene names | TSSK1, SPOGA1, SPOGA4, STK22A, STK22D | |||
|
Domain Architecture |
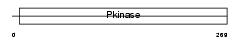
|
|||||
| Description | Testis-specific serine/threonine-protein kinase 1 (EC 2.7.11.1) (TSSK- 1) (Testis-specific kinase 1) (TSK-1) (Serine/threonine-protein kinase 22A). | |||||
|
DCC_MOUSE
|
||||||
| θ value | 4.03905 (rank : 18) | NC score | 0.008571 (rank : 47) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
FA13C_MOUSE
|
||||||
| θ value | 4.03905 (rank : 19) | NC score | 0.041690 (rank : 23) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DBR2 | Gene names | Fam13c1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
LTBP3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 20) | NC score | 0.007635 (rank : 50) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NS15, O15107, Q96HB9, Q9H7K2, Q9UFN4 | Gene names | LTBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 3 precursor (LTBP-3). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 21) | NC score | 0.017123 (rank : 41) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
WAPL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.041683 (rank : 24) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
ZFY19_HUMAN
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.031828 (rank : 28) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96K21, Q86WC2, Q8WU96 | Gene names | ZFYVE19, MPFYVE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 19 (MLL partner containing FYVE domain). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.072434 (rank : 11) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
SYTL1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.014848 (rank : 43) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99N80, Q99J26 | Gene names | Sytl1, Slp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.056742 (rank : 16) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
GAGE1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | 0.018870 (rank : 40) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13065 | Gene names | GAGE1 | |||
|
Domain Architecture |
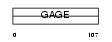
|
|||||
| Description | G antigen 1 (GAGE-1) (MZ2-F antigen). | |||||
|
LRRF1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 28) | NC score | 0.046417 (rank : 21) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3UZ39, O70323, Q3TS94 | Gene names | Lrrfip1, Flap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 1 (LRR FLII- interacting protein 1) (FLI-LRR-associated protein 1) (Flap-1) (H186 FLAP). | |||||
|
NALP5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 29) | NC score | 0.029789 (rank : 30) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
PHF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 30) | NC score | 0.023201 (rank : 36) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43189, O60929, Q96KM7 | Gene names | PHF1 | |||
|
Domain Architecture |
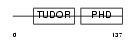
|
|||||
| Description | PHD finger protein 1 (Protein PHF1). | |||||
|
SDPR_MOUSE
|
||||||
| θ value | 6.88961 (rank : 31) | NC score | 0.029246 (rank : 31) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q63918, Q3V1P6, Q78EC3, Q8CBT4 | Gene names | Sdpr, Sdr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum deprivation-response protein (Phosphatidylserine-binding protein). | |||||
|
XRN1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.032412 (rank : 27) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P97789, O35651, P97790, Q3TE44, Q3TRE9, Q3UQL5 | Gene names | Xrn1, Dhm2, Exo, Sep1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5'-3' exoribonuclease 1 (EC 3.1.11.-) (mXRN1) (Strand-exchange protein 1 homolog) (Protein Dhm2). | |||||
|
CEP35_HUMAN
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.024607 (rank : 34) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
CGRE1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.027673 (rank : 32) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
DDX51_HUMAN
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.011083 (rank : 44) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
LRC50_HUMAN
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | 0.025917 (rank : 33) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NEP3, Q69YI8, Q69YJ0, Q96LP3 | Gene names | LRRC50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 50. | |||||
|
LRTM1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 37) | NC score | 0.007965 (rank : 48) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86UE6, Q96DN1 | Gene names | LRRTM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat transmembrane neuronal protein 1 precursor. | |||||
|
MINT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 38) | NC score | 0.020806 (rank : 39) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MTSS1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 39) | NC score | 0.021959 (rank : 38) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8R1S4, Q8BMM3, Q99LB3 | Gene names | Mtss1, Mim | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein). | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 40) | NC score | 0.003773 (rank : 51) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
OPTN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 41) | NC score | 0.008734 (rank : 46) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1120 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K3K8 | Gene names | Optn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Optineurin. | |||||
|
PRGC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.041864 (rank : 22) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
PRP40_MOUSE
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.031296 (rank : 29) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
RYR1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.022931 (rank : 37) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P21817, Q16314, Q16368, Q9NPK1, Q9P1U4 | Gene names | RYR1, RYDR | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 1 (Skeletal muscle-type ryanodine receptor) (RyR1) (RYR-1) (Skeletal muscle calcium release channel). | |||||
|
SNPC4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.015437 (rank : 42) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5SXM2, Q9Y6P7 | Gene names | SNAPC4, SNAP190 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit) (Proximal sequence element-binding transcription factor subunit alpha) (PSE-binding factor subunit alpha) (PTF subunit alpha). | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 46) | NC score | 0.052397 (rank : 20) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
NUCKS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.061343 (rank : 14) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
NUCKS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.062238 (rank : 13) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
PTMA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.084203 (rank : 7) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
PTMA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.080176 (rank : 8) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
PTMS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.057110 (rank : 15) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
CP053_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 1.39067e-109 (rank : 1) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9BTK6 | Gene names | C16orf53 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf53. | |||||
|
CP053_MOUSE
|
||||||
| NC score | 0.977074 (rank : 2) | θ value | 1.3939e-101 (rank : 2) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99L02 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf53 homolog. | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.118047 (rank : 3) | θ value | 0.0431538 (rank : 4) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.102712 (rank : 4) | θ value | 0.0148317 (rank : 3) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.090089 (rank : 5) | θ value | 0.0736092 (rank : 6) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.085855 (rank : 6) | θ value | 0.0563607 (rank : 5) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
PTMA_HUMAN
|
||||||
| NC score | 0.084203 (rank : 7) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
PTMA_MOUSE
|
||||||
| NC score | 0.080176 (rank : 8) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
F10A4_HUMAN
|
||||||
| NC score | 0.073030 (rank : 9) | θ value | 1.06291 (rank : 10) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IZP2 | Gene names | FAM10A4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A4. | |||||
|
F10A1_HUMAN
|
||||||
| NC score | 0.072927 (rank : 10) | θ value | 1.38821 (rank : 13) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P50502, O14999 | Gene names | ST13, FAM10A1, HIP, SNC6 | |||
|
Domain Architecture |

|
|||||
| Description | Hsc70-interacting protein (Hip) (Suppression of tumorigenicity protein 13) (Putative tumor suppressor ST13) (Protein FAM10A1) (Progesterone receptor-associated p48 protein) (NY-REN-33 antigen). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.072434 (rank : 11) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
F10A5_HUMAN
|
||||||
| NC score | 0.070637 (rank : 12) | θ value | 2.36792 (rank : 14) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NFI4 | Gene names | FAM10A5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A5. | |||||
|
NUCKS_MOUSE
|
||||||
| NC score | 0.062238 (rank : 13) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
NUCKS_HUMAN
|
||||||
| NC score | 0.061343 (rank : 14) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
PTMS_MOUSE
|
||||||
| NC score | 0.057110 (rank : 15) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.056742 (rank : 16) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
RIOK2_MOUSE
|
||||||
| NC score | 0.056679 (rank : 17) | θ value | 0.62314 (rank : 8) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CQS5, Q91XF3 | Gene names | Riok2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase RIO2 (EC 2.7.11.1) (RIO kinase 2). | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.056069 (rank : 18) | θ value | 3.0926 (rank : 15) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.055270 (rank : 19) | θ value | 0.813845 (rank : 9) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.052397 (rank : 20) | θ value | θ > 10 (rank : 46) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
LRRF1_MOUSE
|
||||||
| NC score | 0.046417 (rank : 21) | θ value | 6.88961 (rank : 28) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3UZ39, O70323, Q3TS94 | Gene names | Lrrfip1, Flap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 1 (LRR FLII- interacting protein 1) (FLI-LRR-associated protein 1) (Flap-1) (H186 FLAP). | |||||
|
PRGC2_MOUSE
|
||||||
| NC score | 0.041864 (rank : 22) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
FA13C_MOUSE
|
||||||
| NC score | 0.041690 (rank : 23) | θ value | 4.03905 (rank : 19) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DBR2 | Gene names | Fam13c1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
WAPL_MOUSE
|
||||||
| NC score | 0.041683 (rank : 24) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
FA13C_HUMAN
|
||||||
| NC score | 0.040949 (rank : 25) | θ value | 3.0926 (rank : 16) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NE31, Q99787 | Gene names | FAM13C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
ZC3H6_MOUSE
|
||||||
| NC score | 0.040766 (rank : 26) | θ value | 1.06291 (rank : 11) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BYK8, Q80UK3, Q8C1H1, Q9D604 | Gene names | Zc3h6, Zc3hdc6 | |||
|
Domain Architecture |
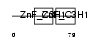
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
XRN1_MOUSE
|
||||||
| NC score | 0.032412 (rank : 27) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P97789, O35651, P97790, Q3TE44, Q3TRE9, Q3UQL5 | Gene names | Xrn1, Dhm2, Exo, Sep1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5'-3' exoribonuclease 1 (EC 3.1.11.-) (mXRN1) (Strand-exchange protein 1 homolog) (Protein Dhm2). | |||||
|
ZFY19_HUMAN
|
||||||
| NC score | 0.031828 (rank : 28) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96K21, Q86WC2, Q8WU96 | Gene names | ZFYVE19, MPFYVE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 19 (MLL partner containing FYVE domain). | |||||
|
PRP40_MOUSE
|
||||||
| NC score | 0.031296 (rank : 29) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
NALP5_MOUSE
|
||||||
| NC score | 0.029789 (rank : 30) | θ value | 6.88961 (rank : 29) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
SDPR_MOUSE
|
||||||
| NC score | 0.029246 (rank : 31) | θ value | 6.88961 (rank : 31) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q63918, Q3V1P6, Q78EC3, Q8CBT4 | Gene names | Sdpr, Sdr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum deprivation-response protein (Phosphatidylserine-binding protein). | |||||
|
CGRE1_HUMAN
|
||||||
| NC score | 0.027673 (rank : 32) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
LRC50_HUMAN
|
||||||
| NC score | 0.025917 (rank : 33) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NEP3, Q69YI8, Q69YJ0, Q96LP3 | Gene names | LRRC50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 50. | |||||
|
CEP35_HUMAN
|
||||||
| NC score | 0.024607 (rank : 34) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
LAP4_HUMAN
|
||||||
| NC score | 0.024536 (rank : 35) | θ value | 0.163984 (rank : 7) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 608 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14160, Q6P496, Q7Z5D1, Q8WWV8, Q96C69, Q96GG1 | Gene names | SCRIB, CRIB1, KIAA0147, LAP4, SCRB1, VARTUL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog) (hScrib). | |||||
|
PHF1_HUMAN
|
||||||
| NC score | 0.023201 (rank : 36) | θ value | 6.88961 (rank : 30) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43189, O60929, Q96KM7 | Gene names | PHF1 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 1 (Protein PHF1). | |||||
|
RYR1_HUMAN
|
||||||
| NC score | 0.022931 (rank : 37) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P21817, Q16314, Q16368, Q9NPK1, Q9P1U4 | Gene names | RYR1, RYDR | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 1 (Skeletal muscle-type ryanodine receptor) (RyR1) (RYR-1) (Skeletal muscle calcium release channel). | |||||
|
MTSS1_MOUSE
|
||||||
| NC score | 0.021959 (rank : 38) | θ value | 8.99809 (rank : 39) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8R1S4, Q8BMM3, Q99LB3 | Gene names | Mtss1, Mim | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.020806 (rank : 39) | θ value | 8.99809 (rank : 38) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
GAGE1_HUMAN
|
||||||
| NC score | 0.018870 (rank : 40) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13065 | Gene names | GAGE1 | |||
|
Domain Architecture |

|
|||||
| Description | G antigen 1 (GAGE-1) (MZ2-F antigen). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.017123 (rank : 41) | θ value | 4.03905 (rank : 21) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SNPC4_HUMAN
|
||||||
| NC score | 0.015437 (rank : 42) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5SXM2, Q9Y6P7 | Gene names | SNAPC4, SNAP190 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit) (Proximal sequence element-binding transcription factor subunit alpha) (PSE-binding factor subunit alpha) (PTF subunit alpha). | |||||
|
SYTL1_MOUSE
|
||||||
| NC score | 0.014848 (rank : 43) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99N80, Q99J26 | Gene names | Sytl1, Slp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7). | |||||
|
DDX51_HUMAN
|
||||||
| NC score | 0.011083 (rank : 44) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
TSSK1_HUMAN
|
||||||
| NC score | 0.009256 (rank : 45) | θ value | 3.0926 (rank : 17) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 840 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BXA7 | Gene names | TSSK1, SPOGA1, SPOGA4, STK22A, STK22D | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific serine/threonine-protein kinase 1 (EC 2.7.11.1) (TSSK- 1) (Testis-specific kinase 1) (TSK-1) (Serine/threonine-protein kinase 22A). | |||||
|
OPTN_MOUSE
|
||||||
| NC score | 0.008734 (rank : 46) | θ value | 8.99809 (rank : 41) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1120 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K3K8 | Gene names | Optn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Optineurin. | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.008571 (rank : 47) | θ value | 4.03905 (rank : 18) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
LRTM1_HUMAN
|
||||||
| NC score | 0.007965 (rank : 48) | θ value | 8.99809 (rank : 37) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86UE6, Q96DN1 | Gene names | LRRTM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat transmembrane neuronal protein 1 precursor. | |||||
|
ABL1_HUMAN
|
||||||
| NC score | 0.007664 (rank : 49) | θ value | 1.38821 (rank : 12) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1220 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P00519, Q13869, Q13870, Q16133, Q45F09 | Gene names | ABL1, ABL, JTK7 | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene tyrosine-protein kinase ABL1 (EC 2.7.10.2) (p150) (c- ABL) (Abelson murine leukemia viral oncogene homolog 1). | |||||
|
LTBP3_HUMAN
|
||||||
| NC score | 0.007635 (rank : 50) | θ value | 4.03905 (rank : 20) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NS15, O15107, Q96HB9, Q9H7K2, Q9UFN4 | Gene names | LTBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 3 precursor (LTBP-3). | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | 0.003773 (rank : 51) | θ value | 8.99809 (rank : 40) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||