Please be patient as the page loads
|
YYAP1_HUMAN
|
||||||
| SwissProt Accessions | Q9H869, Q5VYZ4, Q5VYZ7, Q7L4C3, Q7L5E2, Q8IXA6, Q8TEW5, Q8TF04, Q96HB6, Q9BQ64, Q9NV84 | Gene names | YY1AP1, HCCA2, YY1AP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YY1-associated protein 1 (Hepatocellular carcinoma susceptibility protein) (Hepatocellular carcinoma-associated protein 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
GON4L_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.884345 (rank : 2) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
GON4L_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.868108 (rank : 3) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
YYAP1_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9H869, Q5VYZ4, Q5VYZ7, Q7L4C3, Q7L5E2, Q8IXA6, Q8TEW5, Q8TF04, Q96HB6, Q9BQ64, Q9NV84 | Gene names | YY1AP1, HCCA2, YY1AP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YY1-associated protein 1 (Hepatocellular carcinoma susceptibility protein) (Hepatocellular carcinoma-associated protein 2). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 4) | NC score | 0.066966 (rank : 9) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
MUC2_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 5) | NC score | 0.067234 (rank : 8) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
PER1_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 6) | NC score | 0.085974 (rank : 6) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
SM1L2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 7) | NC score | 0.034976 (rank : 42) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q920F6 | Gene names | Smc1l2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 1-like 2 protein (SMC1beta protein). | |||||
|
ATF5_MOUSE
|
||||||
| θ value | 0.21417 (rank : 8) | NC score | 0.066637 (rank : 11) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O70191, Q58E51, Q99NH6 | Gene names | Atf5, Atfx, Nap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-5 (Activating transcription factor 5-alpha/beta) (Transcription factor ATFx) (Transcription factor-like protein ODA-10) (BZIP protein ATF7) (NRIF3- associated protein) (NAP1). | |||||
|
RTKN_HUMAN
|
||||||
| θ value | 0.365318 (rank : 9) | NC score | 0.057609 (rank : 18) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BST9, Q8WVN1, Q96PT6, Q9HB05 | Gene names | RTKN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rhotekin. | |||||
|
TTLL5_MOUSE
|
||||||
| θ value | 0.365318 (rank : 10) | NC score | 0.048602 (rank : 31) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CHB8 | Gene names | Ttll5, Kiaa0998 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 5. | |||||
|
PRDM2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 11) | NC score | 0.016510 (rank : 68) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 12) | NC score | 0.065519 (rank : 12) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
LCP2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.052397 (rank : 25) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q60787 | Gene names | Lcp2 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte cytosolic protein 2 (SH2 domain-containing leukocyte protein of 76 kDa) (SLP-76 tyrosine phosphoprotein) (SLP76). | |||||
|
MAP4_MOUSE
|
||||||
| θ value | 0.62314 (rank : 14) | NC score | 0.058459 (rank : 17) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
TCP10_HUMAN
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.043389 (rank : 34) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q12799 | Gene names | TCP10, TCP10A | |||
|
Domain Architecture |

|
|||||
| Description | T-complex protein 10A homolog. | |||||
|
FOXM1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 16) | NC score | 0.022467 (rank : 58) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q08050, O43258, O43259, O43260, Q4ZGG7, Q9BRL2 | Gene names | FOXM1, FKHL16, HFH11, MPP2, WIN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead-related protein FKHL16) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/fork-head homolog 11) (HFH-11) (Winged helix factor from INS-1 cells) (M-phase phosphoprotein 2) (MPM-2 reactive phosphoprotein 2) (Transcription factor Trident). | |||||
|
SMC1A_HUMAN
|
||||||
| θ value | 1.06291 (rank : 17) | NC score | 0.039213 (rank : 39) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
SMC1A_MOUSE
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.037778 (rank : 41) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
TTLL4_MOUSE
|
||||||
| θ value | 1.06291 (rank : 19) | NC score | 0.040953 (rank : 36) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80UG8, Q5U4C4 | Gene names | Ttll4, Kiaa0173 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 4. | |||||
|
WNK1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 20) | NC score | 0.012098 (rank : 74) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1158 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9H4A3, O15052, Q86WL5, Q8N673, Q9P1S9 | Gene names | WNK1, KDP, KIAA0344, PRKWNK1 | |||
|
Domain Architecture |
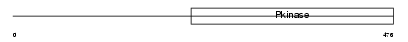
|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1) (Kinase deficient protein). | |||||
|
SRGP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 21) | NC score | 0.016652 (rank : 67) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z6B7, Q9H8A3, Q9P2P2 | Gene names | SRGAP1, ARHGAP13, KIAA1304 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 1 (srGAP1) (Rho-GTPase- activating protein 13). | |||||
|
FBX46_MOUSE
|
||||||
| θ value | 1.81305 (rank : 22) | NC score | 0.039921 (rank : 38) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BG80 | Gene names | Fbxo46, Fbx46, Fbxo34l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
LIPA1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.028291 (rank : 49) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
MAML1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.027003 (rank : 50) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92585, Q9NZ12 | Gene names | MAML1, KIAA0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.051814 (rank : 26) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
SP2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.005103 (rank : 82) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q02086 | Gene names | SP2, KIAA0048 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Sp2. | |||||
|
ZIMP7_HUMAN
|
||||||
| θ value | 1.81305 (rank : 27) | NC score | 0.028580 (rank : 48) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NF64, O94790, Q659A8, Q6JKL5, Q8WTX8, Q96Q01, Q9BQH7 | Gene names | ZIMP7, KIAA1886 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
CDN1C_HUMAN
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.066738 (rank : 10) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P49918 | Gene names | CDKN1C, KIP2 | |||
|
Domain Architecture |
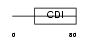
|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
NU214_HUMAN
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.053375 (rank : 20) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
GLE1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.031361 (rank : 43) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8R322, Q3TT10, Q3TU23, Q3UD65, Q8BT16, Q9D4A6 | Gene names | Gle1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin GLE1 (GLE1-like protein). | |||||
|
HCFC1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.046278 (rank : 33) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51610, Q6P4G5 | Gene names | HCFC1, HCF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) (VP16 accessory protein) (VCAF) (CFF) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N-terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C- terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
KTN1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.015724 (rank : 70) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.038583 (rank : 40) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
CLAP2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.015983 (rank : 69) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BRT1, Q8CHE3, Q99JI3, Q9DB80 | Gene names | Clasp2, Kiaa0627 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CLIP-associating protein 2 (Cytoplasmic linker-associated protein 2). | |||||
|
IF4G1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.019124 (rank : 62) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.024837 (rank : 55) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
PER1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.042448 (rank : 35) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PHC3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.025715 (rank : 52) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NDX5, Q5HYF0, Q6NSG2, Q8NFT7, Q8NFZ1, Q8TBM2, Q9H971, Q9H9I4 | Gene names | PHC3, EDR3, PH3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3 (hPH3) (Homolog of polyhomeotic 3) (Early development regulatory protein 3). | |||||
|
PLXB1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.019151 (rank : 61) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CJH3, Q6ZQC3, Q80ZZ1 | Gene names | Plxnb1, Kiaa0407 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor. | |||||
|
BSN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.050222 (rank : 28) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CEP57_MOUSE
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.016850 (rank : 64) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 580 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CEE0, Q6ZQJ3, Q7TN18, Q80X65, Q810F2, Q9D4J4, Q9D5S4, Q9D5W5 | Gene names | Cep57, Kiaa0092, Tsp57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin). | |||||
|
GCC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.016665 (rank : 66) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.014394 (rank : 72) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
MYH4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.007852 (rank : 80) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1476 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5SX39 | Gene names | Myh4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal). | |||||
|
NU153_HUMAN
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.024610 (rank : 56) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
PELP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.048426 (rank : 32) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
RBP24_HUMAN
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.016686 (rank : 65) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
RC3H1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.020770 (rank : 60) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
STA13_HUMAN
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.010666 (rank : 77) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y3M8, Q5HYH1, Q5TAE3, Q6UN61, Q86TP6, Q86WQ3, Q86XT1 | Gene names | STARD13, DLC2, GT650 | |||
|
Domain Architecture |

|
|||||
| Description | StAR-related lipid transfer protein 13 (StARD13) (START domain- containing protein 13) (46H23.2) (Deleted in liver cancer protein 2) (Rho GTPase-activating protein). | |||||
|
TOP2A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.021973 (rank : 59) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01320 | Gene names | Top2a, Top-2, Top2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
UN84B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.014781 (rank : 71) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BJS4 | Gene names | Unc84b, Sun2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sad1/unc-84-like protein 2. | |||||
|
BRD8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 52) | NC score | 0.024898 (rank : 54) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R3B7, Q8C049, Q8R583, Q8VDP0 | Gene names | Brd8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8. | |||||
|
HC127_HUMAN
|
||||||
| θ value | 6.88961 (rank : 53) | NC score | 0.031293 (rank : 44) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NQZ6, Q5JTF9, Q9H9C3, Q9H9H7, Q9ULQ4 | Gene names | HCA127, KIAA1166 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocellular carcinoma-associated antigen 127. | |||||
|
HC127_MOUSE
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.031293 (rank : 45) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q68FG0, Q6ZPW6 | Gene names | Hca127, Kiaa1166 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocellular carcinoma-associated antigen 127 homolog. | |||||
|
ITSN2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.010266 (rank : 78) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
LARP4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.024115 (rank : 57) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BWW4, Q3UR69 | Gene names | Larp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 4 (La ribonucleoprotein domain family member 4). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.030362 (rank : 46) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
SMG5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.013284 (rank : 73) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UPR3, Q5QJE7, Q8IXC0, Q8IY09, Q96IJ7 | Gene names | SMG5, EST1B, KIAA1089 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG5 (SMG-5 homolog) (EST1-like protein B) (LPTS-interacting protein) (LPTS-RP1). | |||||
|
TTBK1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.009326 (rank : 79) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1058 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5TCY1, Q2L6C6, Q8N444, Q96JH2 | Gene names | TTBK1, BDTK, KIAA1855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 1 (EC 2.7.11.1) (Brain-derived tau kinase). | |||||
|
2ACA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.011636 (rank : 76) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q06190, Q06189, Q9NPQ5 | Gene names | PPP2R3A, PPP2R3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein phosphatase 2A 72/130 kDa regulatory subunit B (PP2A, subunit B, B''-PR72/PR130) (PP2A, subunit B, B72/B130 isoforms) (PP2A, subunit B, PR72/PR130 isoforms) (PP2A, subunit B, R3 isoform). | |||||
|
BRPF3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.007774 (rank : 81) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||
|
HCFC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.026303 (rank : 51) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61191, Q684R1, Q7TSB0, Q8C2D0, Q9QWH2 | Gene names | Hcfc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N- terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C-terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
HXA10_MOUSE
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.002630 (rank : 83) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P31310, Q80Y22 | Gene names | Hoxa10, Hox-1.8, Hoxa-10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A10 (Hox-1.8). | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.029407 (rank : 47) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
MNT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.018553 (rank : 63) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
PELP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.040397 (rank : 37) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
SCEL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.025655 (rank : 53) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95171 | Gene names | SCEL | |||
|
Domain Architecture |

|
|||||
| Description | Sciellin. | |||||
|
TXLNG_HUMAN
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.011817 (rank : 75) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NUQ3, Q9P0X1 | Gene names | TXLNG, CXorf15, LSR5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma-taxilin (Lipopolysaccharide-specific response protein 5). | |||||
|
VEZF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.000759 (rank : 84) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14119 | Gene names | VEZF1, DB1, ZNF161 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vascular endothelial zinc finger 1 (Zinc finger protein 161) (Putative transcription factor DB1). | |||||
|
ZN628_MOUSE
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.000146 (rank : 85) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 815 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CJ78, Q3U2L5, Q6P5B3 | Gene names | Znf628, Zec, Zfp628 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 628 (Zinc finger protein expressed in embryonal cells and certain adult organs). | |||||
|
AFF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.050197 (rank : 29) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.052796 (rank : 22) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
C8AP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.103887 (rank : 4) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
C8AP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.103622 (rank : 5) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
CC47_MOUSE
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.051071 (rank : 27) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
CN155_HUMAN
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.065476 (rank : 13) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
DCDC2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.054066 (rank : 19) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.059576 (rank : 15) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
MDC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.059295 (rank : 16) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.070382 (rank : 7) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
OXR1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.052751 (rank : 23) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
PO121_HUMAN
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.052601 (rank : 24) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
RAI2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.050075 (rank : 30) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5P3, Q8N6X7 | Gene names | RAI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2. | |||||
|
SLC2B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.052891 (rank : 21) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.064311 (rank : 14) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
YYAP1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9H869, Q5VYZ4, Q5VYZ7, Q7L4C3, Q7L5E2, Q8IXA6, Q8TEW5, Q8TF04, Q96HB6, Q9BQ64, Q9NV84 | Gene names | YY1AP1, HCCA2, YY1AP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YY1-associated protein 1 (Hepatocellular carcinoma susceptibility protein) (Hepatocellular carcinoma-associated protein 2). | |||||
|
GON4L_HUMAN
|
||||||
| NC score | 0.884345 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
GON4L_MOUSE
|
||||||
| NC score | 0.868108 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
C8AP2_HUMAN
|
||||||
| NC score | 0.103887 (rank : 4) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
C8AP2_MOUSE
|
||||||
| NC score | 0.103622 (rank : 5) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
PER1_MOUSE
|
||||||
| NC score | 0.085974 (rank : 6) | θ value | 0.0193708 (rank : 6) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.070382 (rank : 7) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
MUC2_HUMAN
|
||||||
| NC score | 0.067234 (rank : 8) | θ value | 0.0148317 (rank : 5) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.066966 (rank : 9) | θ value | 0.00869519 (rank : 4) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
CDN1C_HUMAN
|
||||||
| NC score | 0.066738 (rank : 10) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P49918 | Gene names | CDKN1C, KIP2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
ATF5_MOUSE
|
||||||
| NC score | 0.066637 (rank : 11) | θ value | 0.21417 (rank : 8) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O70191, Q58E51, Q99NH6 | Gene names | Atf5, Atfx, Nap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-5 (Activating transcription factor 5-alpha/beta) (Transcription factor ATFx) (Transcription factor-like protein ODA-10) (BZIP protein ATF7) (NRIF3- associated protein) (NAP1). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.065519 (rank : 12) | θ value | 0.47712 (rank : 12) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
CN155_HUMAN
|
||||||
| NC score | 0.065476 (rank : 13) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.064311 (rank : 14) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.059576 (rank : 15) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.059295 (rank : 16) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MAP4_MOUSE
|
||||||
| NC score | 0.058459 (rank : 17) | θ value | 0.62314 (rank : 14) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
RTKN_HUMAN
|
||||||
| NC score | 0.057609 (rank : 18) | θ value | 0.365318 (rank : 9) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BST9, Q8WVN1, Q96PT6, Q9HB05 | Gene names | RTKN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rhotekin. | |||||
|
DCDC2_MOUSE
|
||||||
| NC score | 0.054066 (rank : 19) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
NU214_HUMAN
|
||||||
| NC score | 0.053375 (rank : 20) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
SLC2B_HUMAN
|
||||||
| NC score | 0.052891 (rank : 21) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.052796 (rank : 22) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
OXR1_MOUSE
|
||||||
| NC score | 0.052751 (rank : 23) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
PO121_HUMAN
|
||||||
| NC score | 0.052601 (rank : 24) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
LCP2_MOUSE
|
||||||
| NC score | 0.052397 (rank : 25) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q60787 | Gene names | Lcp2 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte cytosolic protein 2 (SH2 domain-containing leukocyte protein of 76 kDa) (SLP-76 tyrosine phosphoprotein) (SLP76). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.051814 (rank : 26) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
CC47_MOUSE
|
||||||
| NC score | 0.051071 (rank : 27) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.050222 (rank : 28) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
AFF1_HUMAN
|
||||||
| NC score | 0.050197 (rank : 29) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
RAI2_HUMAN
|
||||||
| NC score | 0.050075 (rank : 30) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5P3, Q8N6X7 | Gene names | RAI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2. | |||||
|
TTLL5_MOUSE
|
||||||
| NC score | 0.048602 (rank : 31) | θ value | 0.365318 (rank : 10) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CHB8 | Gene names | Ttll5, Kiaa0998 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 5. | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.048426 (rank : 32) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
HCFC1_HUMAN
|
||||||
| NC score | 0.046278 (rank : 33) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51610, Q6P4G5 | Gene names | HCFC1, HCF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) (VP16 accessory protein) (VCAF) (CFF) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N-terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C- terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
TCP10_HUMAN
|
||||||
| NC score | 0.043389 (rank : 34) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q12799 | Gene names | TCP10, TCP10A | |||
|
Domain Architecture |

|
|||||
| Description | T-complex protein 10A homolog. | |||||
|
PER1_HUMAN
|
||||||
| NC score | 0.042448 (rank : 35) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
TTLL4_MOUSE
|
||||||
| NC score | 0.040953 (rank : 36) | θ value | 1.06291 (rank : 19) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80UG8, Q5U4C4 | Gene names | Ttll4, Kiaa0173 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 4. | |||||
|
PELP1_HUMAN
|
||||||
| NC score | 0.040397 (rank : 37) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
FBX46_MOUSE
|
||||||
| NC score | 0.039921 (rank : 38) | θ value | 1.81305 (rank : 22) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BG80 | Gene names | Fbxo46, Fbx46, Fbxo34l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
SMC1A_HUMAN
|
||||||
| NC score | 0.039213 (rank : 39) | θ value | 1.06291 (rank : 17) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.038583 (rank : 40) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
SMC1A_MOUSE
|
||||||
| NC score | 0.037778 (rank : 41) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
SM1L2_MOUSE
|
||||||
| NC score | 0.034976 (rank : 42) | θ value | 0.163984 (rank : 7) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q920F6 | Gene names | Smc1l2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 1-like 2 protein (SMC1beta protein). | |||||
|
GLE1_MOUSE
|
||||||
| NC score | 0.031361 (rank : 43) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8R322, Q3TT10, Q3TU23, Q3UD65, Q8BT16, Q9D4A6 | Gene names | Gle1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin GLE1 (GLE1-like protein). | |||||
|
HC127_HUMAN
|
||||||
| NC score | 0.031293 (rank : 44) | θ value | 6.88961 (rank : 53) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NQZ6, Q5JTF9, Q9H9C3, Q9H9H7, Q9ULQ4 | Gene names | HCA127, KIAA1166 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocellular carcinoma-associated antigen 127. | |||||
|
HC127_MOUSE
|
||||||
| NC score | 0.031293 (rank : 45) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q68FG0, Q6ZPW6 | Gene names | Hca127, Kiaa1166 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocellular carcinoma-associated antigen 127 homolog. | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.030362 (rank : 46) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.029407 (rank : 47) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
ZIMP7_HUMAN
|
||||||
| NC score | 0.028580 (rank : 48) | θ value | 1.81305 (rank : 27) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NF64, O94790, Q659A8, Q6JKL5, Q8WTX8, Q96Q01, Q9BQH7 | Gene names | ZIMP7, KIAA1886 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
LIPA1_HUMAN
|
||||||
| NC score | 0.028291 (rank : 49) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
MAML1_HUMAN
|
||||||
| NC score | 0.027003 (rank : 50) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92585, Q9NZ12 | Gene names | MAML1, KIAA0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
HCFC1_MOUSE
|
||||||
| NC score | 0.026303 (rank : 51) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61191, Q684R1, Q7TSB0, Q8C2D0, Q9QWH2 | Gene names | Hcfc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N- terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C-terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
PHC3_HUMAN
|
||||||
| NC score | 0.025715 (rank : 52) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NDX5, Q5HYF0, Q6NSG2, Q8NFT7, Q8NFZ1, Q8TBM2, Q9H971, Q9H9I4 | Gene names | PHC3, EDR3, PH3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3 (hPH3) (Homolog of polyhomeotic 3) (Early development regulatory protein 3). | |||||
|
SCEL_HUMAN
|
||||||
| NC score | 0.025655 (rank : 53) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95171 | Gene names | SCEL | |||
|
Domain Architecture |

|
|||||
| Description | Sciellin. | |||||
|
BRD8_MOUSE
|
||||||
| NC score | 0.024898 (rank : 54) | θ value | 6.88961 (rank : 52) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R3B7, Q8C049, Q8R583, Q8VDP0 | Gene names | Brd8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8. | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.024837 (rank : 55) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
NU153_HUMAN
|
||||||
| NC score | 0.024610 (rank : 56) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
LARP4_MOUSE
|
||||||
| NC score | 0.024115 (rank : 57) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BWW4, Q3UR69 | Gene names | Larp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 4 (La ribonucleoprotein domain family member 4). | |||||
|
FOXM1_HUMAN
|
||||||
| NC score | 0.022467 (rank : 58) | θ value | 0.813845 (rank : 16) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q08050, O43258, O43259, O43260, Q4ZGG7, Q9BRL2 | Gene names | FOXM1, FKHL16, HFH11, MPP2, WIN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead-related protein FKHL16) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/fork-head homolog 11) (HFH-11) (Winged helix factor from INS-1 cells) (M-phase phosphoprotein 2) (MPM-2 reactive phosphoprotein 2) (Transcription factor Trident). | |||||
|
TOP2A_MOUSE
|
||||||
| NC score | 0.021973 (rank : 59) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01320 | Gene names | Top2a, Top-2, Top2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
RC3H1_HUMAN
|
||||||
| NC score | 0.020770 (rank : 60) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
PLXB1_MOUSE
|
||||||
| NC score | 0.019151 (rank : 61) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CJH3, Q6ZQC3, Q80ZZ1 | Gene names | Plxnb1, Kiaa0407 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor. | |||||
|
IF4G1_MOUSE
|
||||||
| NC score | 0.019124 (rank : 62) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
MNT_HUMAN
|
||||||
| NC score | 0.018553 (rank : 63) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
CEP57_MOUSE
|
||||||
| NC score | 0.016850 (rank : 64) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 580 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CEE0, Q6ZQJ3, Q7TN18, Q80X65, Q810F2, Q9D4J4, Q9D5S4, Q9D5W5 | Gene names | Cep57, Kiaa0092, Tsp57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin). | |||||
|
RBP24_HUMAN
|
||||||
| NC score | 0.016686 (rank : 65) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
GCC2_HUMAN
|
||||||
| NC score | 0.016665 (rank : 66) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
SRGP1_HUMAN
|
||||||
| NC score | 0.016652 (rank : 67) | θ value | 1.38821 (rank : 21) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z6B7, Q9H8A3, Q9P2P2 | Gene names | SRGAP1, ARHGAP13, KIAA1304 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 1 (srGAP1) (Rho-GTPase- activating protein 13). | |||||
|
PRDM2_HUMAN
|
||||||
| NC score | 0.016510 (rank : 68) | θ value | 0.47712 (rank : 11) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
CLAP2_MOUSE
|
||||||
| NC score | 0.015983 (rank : 69) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BRT1, Q8CHE3, Q99JI3, Q9DB80 | Gene names | Clasp2, Kiaa0627 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CLIP-associating protein 2 (Cytoplasmic linker-associated protein 2). | |||||
|
KTN1_MOUSE
|
||||||
| NC score | 0.015724 (rank : 70) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
UN84B_MOUSE
|
||||||
| NC score | 0.014781 (rank : 71) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BJS4 | Gene names | Unc84b, Sun2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sad1/unc-84-like protein 2. | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.014394 (rank : 72) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
SMG5_HUMAN
|
||||||
| NC score | 0.013284 (rank : 73) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UPR3, Q5QJE7, Q8IXC0, Q8IY09, Q96IJ7 | Gene names | SMG5, EST1B, KIAA1089 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG5 (SMG-5 homolog) (EST1-like protein B) (LPTS-interacting protein) (LPTS-RP1). | |||||
|
WNK1_HUMAN
|
||||||
| NC score | 0.012098 (rank : 74) | θ value | 1.06291 (rank : 20) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1158 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9H4A3, O15052, Q86WL5, Q8N673, Q9P1S9 | Gene names | WNK1, KDP, KIAA0344, PRKWNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1) (Kinase deficient protein). | |||||
|
TXLNG_HUMAN
|
||||||
| NC score | 0.011817 (rank : 75) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NUQ3, Q9P0X1 | Gene names | TXLNG, CXorf15, LSR5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma-taxilin (Lipopolysaccharide-specific response protein 5). | |||||
|
2ACA_HUMAN
|
||||||
| NC score | 0.011636 (rank : 76) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q06190, Q06189, Q9NPQ5 | Gene names | PPP2R3A, PPP2R3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein phosphatase 2A 72/130 kDa regulatory subunit B (PP2A, subunit B, B''-PR72/PR130) (PP2A, subunit B, B72/B130 isoforms) (PP2A, subunit B, PR72/PR130 isoforms) (PP2A, subunit B, R3 isoform). | |||||
|
STA13_HUMAN
|
||||||
| NC score | 0.010666 (rank : 77) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y3M8, Q5HYH1, Q5TAE3, Q6UN61, Q86TP6, Q86WQ3, Q86XT1 | Gene names | STARD13, DLC2, GT650 | |||
|
Domain Architecture |

|
|||||
| Description | StAR-related lipid transfer protein 13 (StARD13) (START domain- containing protein 13) (46H23.2) (Deleted in liver cancer protein 2) (Rho GTPase-activating protein). | |||||
|
ITSN2_MOUSE
|
||||||
| NC score | 0.010266 (rank : 78) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
TTBK1_HUMAN
|
||||||
| NC score | 0.009326 (rank : 79) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1058 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5TCY1, Q2L6C6, Q8N444, Q96JH2 | Gene names | TTBK1, BDTK, KIAA1855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 1 (EC 2.7.11.1) (Brain-derived tau kinase). | |||||
|
MYH4_MOUSE
|
||||||
| NC score | 0.007852 (rank : 80) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1476 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5SX39 | Gene names | Myh4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal). | |||||
|
BRPF3_HUMAN
|
||||||
| NC score | 0.007774 (rank : 81) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||
|
SP2_HUMAN
|
||||||
| NC score | 0.005103 (rank : 82) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q02086 | Gene names | SP2, KIAA0048 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Sp2. | |||||
|
HXA10_MOUSE
|
||||||
| NC score | 0.002630 (rank : 83) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P31310, Q80Y22 | Gene names | Hoxa10, Hox-1.8, Hoxa-10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A10 (Hox-1.8). | |||||
|
VEZF1_HUMAN
|
||||||
| NC score | 0.000759 (rank : 84) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14119 | Gene names | VEZF1, DB1, ZNF161 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vascular endothelial zinc finger 1 (Zinc finger protein 161) (Putative transcription factor DB1). | |||||
|
ZN628_MOUSE
|
||||||
| NC score | 0.000146 (rank : 85) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 815 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CJ78, Q3U2L5, Q6P5B3 | Gene names | Znf628, Zec, Zfp628 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 628 (Zinc finger protein expressed in embryonal cells and certain adult organs). | |||||