Please be patient as the page loads
|
TR19L_MOUSE
|
||||||
| SwissProt Accessions | Q8BX43, Q8BTV0 | Gene names | Tnfrsf19l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 19L precursor. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TR19L_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.947634 (rank : 2) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q969Z4, Q86V34, Q96JU1, Q9BUX7 | Gene names | TNFRSF19L, RELT | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 19L precursor (Receptor expressed in lymphoid tissues). | |||||
|
TR19L_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q8BX43, Q8BTV0 | Gene names | Tnfrsf19l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 19L precursor. | |||||
|
CE016_MOUSE
|
||||||
| θ value | 2.98157e-11 (rank : 3) | NC score | 0.472142 (rank : 3) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BRJ3 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C5orf16 homolog. | |||||
|
CE016_HUMAN
|
||||||
| θ value | 3.89403e-11 (rank : 4) | NC score | 0.461099 (rank : 4) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NC24, Q6P4E7, Q6UXY2 | Gene names | C5orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C5orf16. | |||||
|
TNR27_HUMAN
|
||||||
| θ value | 4.0297e-08 (rank : 5) | NC score | 0.456696 (rank : 5) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9HAV5, Q6UWM2, Q8IZA6 | Gene names | EDA2R, TNFRSF27, XEDAR | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 27 (X-linked ectodysplasin-A2 receptor) (EDA-A2 receptor). | |||||
|
TNR27_MOUSE
|
||||||
| θ value | 4.45536e-07 (rank : 6) | NC score | 0.440116 (rank : 6) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BX35, Q8BM50 | Gene names | Eda2r, Tnfrsf27, Xedar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 27 (X-linked ectodysplasin-A2 receptor) (EDA-A2 receptor). | |||||
|
TNR19_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 7) | NC score | 0.409465 (rank : 8) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NS68, Q9BXZ9, Q9BY00, Q9NZV2 | Gene names | TNFRSF19, TAJ, TROY | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 19 precursor (Toxicity and JNK inducer) (TRADE). | |||||
|
TNR19_MOUSE
|
||||||
| θ value | 2.88788e-06 (rank : 8) | NC score | 0.435297 (rank : 7) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JLL3, Q812G3, Q9JHF1, Q9JJH6, Q9JLL2, Q9QXW7 | Gene names | Tnfrsf19, Taj, Troy | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 19 precursor (Toxicity and JNK inducer) (TRADE). | |||||
|
TNR6B_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 9) | NC score | 0.393947 (rank : 9) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O95407 | Gene names | TNFRSF6B, DCR3, TR6 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 6B precursor (Decoy receptor for Fas ligand) (Decoy receptor 3) (DcR3) (M68). | |||||
|
TR11B_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 10) | NC score | 0.353333 (rank : 14) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O00300, O60236, Q53FX6, Q9UHP4 | Gene names | TNFRSF11B, OCIF, OPG | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 11B precursor (Osteoprotegerin) (Osteoclastogenesis inhibitory factor). | |||||
|
TNR4_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 11) | NC score | 0.372552 (rank : 10) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P43489, Q13663, Q5T7M0 | Gene names | TNFRSF4, TXGP1L | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 4 precursor (OX40L receptor) (ACT35 antigen) (TAX transcriptionally-activated glycoprotein 1 receptor) (CD134 antigen). | |||||
|
TR11B_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 12) | NC score | 0.359034 (rank : 13) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O08712, O70202 | Gene names | Tnfrsf11b, Ocif, Opg | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 11B precursor (Osteoprotegerin) (Osteoclastogenesis inhibitory factor). | |||||
|
TNR3_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 13) | NC score | 0.364168 (rank : 11) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P36941 | Gene names | LTBR, TNFCR, TNFRSF3 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 3 precursor (Lymphotoxin-beta receptor) (Tumor necrosis factor receptor 2-related protein) (Tumor necrosis factor C receptor). | |||||
|
TNR4_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 14) | NC score | 0.361862 (rank : 12) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P47741 | Gene names | Tnfrsf4, Ox40, Txgp1 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 4 precursor (OX40L receptor) (OX40 antigen). | |||||
|
TNR5_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 15) | NC score | 0.294008 (rank : 23) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P27512, Q99NE0, Q99NE1, Q99NE2, Q99NE3 | Gene names | Cd40, Tnfrsf5 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 5 precursor (CD40L receptor) (B-cell surface antigen CD40) (BP50) (CDw40). | |||||
|
TNR3_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 16) | NC score | 0.352417 (rank : 15) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P50284 | Gene names | Ltbr, Tnfcr, Tnfrsf3 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 3 precursor (Lymphotoxin-beta receptor). | |||||
|
TNR1A_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 17) | NC score | 0.300932 (rank : 19) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P25118 | Gene names | Tnfrsf1a, Tnfr-1, Tnfr1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 1A precursor (p60) (TNF-R1) (TNF-RI) (TNFR-I) (p55). | |||||
|
TNR9_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 18) | NC score | 0.332414 (rank : 17) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P20334 | Gene names | Tnfrsf9, Cd137, Cd157, Ila, Ly63 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 9 precursor (4-1BB ligand receptor) (T-cell antigen 4-1BB) (CD137 antigen). | |||||
|
LAMA5_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 19) | NC score | 0.126175 (rank : 44) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O15230, Q8WZA7, Q9H1P1 | Gene names | LAMA5, KIAA0533, KIAA1907 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
TNR14_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 20) | NC score | 0.342170 (rank : 16) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q92956, Q8WXR1, Q96J31, Q9UM65 | Gene names | TNFRSF14, HVEA, HVEM | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 14 precursor (Herpesvirus entry mediator A) (Tumor necrosis factor receptor-like 2) (TR2). | |||||
|
TNR1A_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 21) | NC score | 0.281882 (rank : 24) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P19438 | Gene names | TNFRSF1A, TNFAR, TNFR1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 1A precursor (p60) (TNF-R1) (TNF-RI) (TNFR-I) (p55) (CD120a antigen) [Contains: Tumor necrosis factor receptor superfamily member 1A, membrane form; Tumor necrosis factor-binding protein 1 (TBPI)]. | |||||
|
TNR1B_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 22) | NC score | 0.301421 (rank : 18) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P25119, O88734, P97893 | Gene names | Tnfrsf1b, Tnfr-2, Tnfr2 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 1B precursor (Tumor necrosis factor receptor 2) (TNF-R2) (Tumor necrosis factor receptor type II) (p75) (p80 TNF-alpha receptor). | |||||
|
TNR23_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 23) | NC score | 0.261328 (rank : 27) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9ER63, Q8VHC0 | Gene names | Tnfrsf23, Dctrailr1, Tnfrh1, Tnfrsf1al1 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 23 precursor (Tumor necrosis factor receptor p60 homolog 1) (TNF receptor family member SOB) (Decoy TRAIL receptor 1) (TNF receptor homolog 1). | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 24) | NC score | 0.032484 (rank : 131) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
LAMC2_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 25) | NC score | 0.106629 (rank : 52) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q13753, Q02536, Q02537, Q13752, Q14941, Q5VYE8 | Gene names | LAMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-2 chain precursor (Laminin 5 gamma 2 subunit) (Kalinin/nicein/epiligrin 100 kDa subunit) (Laminin B2t chain) (Cell- scattering factor 140 kDa subunit) (CSF 140 kDa subunit) (Large adhesive scatter factor 140 kDa subunit) (Ladsin 140 kDa subunit). | |||||
|
CS034_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 26) | NC score | 0.117008 (rank : 45) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NCQ2 | Gene names | C19orf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf34. | |||||
|
LAMA5_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 27) | NC score | 0.115380 (rank : 46) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q61001, Q9JHQ6 | Gene names | Lama5 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
STAB1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 28) | NC score | 0.100351 (rank : 58) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 546 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9NY15, Q8IUH0, Q8IUH1, Q93072 | Gene names | STAB1, FEEL1, KIAA0246 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein) (MS-1 antigen). | |||||
|
TNR25_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 29) | NC score | 0.265406 (rank : 25) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q93038, O00275, O00276, O00277, O00278, O00279, O00280, O14865, O14866, P78507, P78515, Q92983, Q93036, Q93037, Q99722, Q99830, Q99831, Q9BY86, Q9UME0, Q9UME1, Q9UME5 | Gene names | TNFRSF25, APO3, DDR3, DR3, TNFRSF12, WSL, WSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 25 precursor (WSL-1 protein) (Apoptosis-mediating receptor DR3) (Apoptosis-mediating receptor TRAMP) (Death domain receptor 3) (WSL protein) (Apoptosis- inducing receptor AIR) (Apo-3) (Lymphocyte-associated receptor of death) (LARD). | |||||
|
STAB2_MOUSE
|
||||||
| θ value | 0.125558 (rank : 30) | NC score | 0.098124 (rank : 61) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8R4U0, Q8BM87 | Gene names | Stab2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stabilin-2 precursor [Contains: Short form stabilin-2]. | |||||
|
TNR1B_HUMAN
|
||||||
| θ value | 0.125558 (rank : 31) | NC score | 0.299658 (rank : 20) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P20333, Q16042, Q6YI29, Q9UIH1 | Gene names | TNFRSF1B, TNFBR, TNFR2 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 1B precursor (Tumor necrosis factor receptor 2) (TNF-R2) (Tumor necrosis factor receptor type II) (p75) (p80 TNF-alpha receptor) (CD120b antigen) (Etanercept) [Contains: Tumor necrosis factor receptor superfamily member 1b, membrane form; Tumor necrosis factor-binding protein 2 (TBPII) (TBP- 2)]. | |||||
|
EPHB1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 32) | NC score | 0.030305 (rank : 133) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 988 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P54762, O43569, O95142, O95143 | Gene names | EPHB1, EPHT2, NET | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 1 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor EPH-2) (NET) (HEK6) (ELK). | |||||
|
TNR5_HUMAN
|
||||||
| θ value | 0.163984 (rank : 33) | NC score | 0.295618 (rank : 22) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P25942, Q9BYU0 | Gene names | CD40, TNFRSF5 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 5 precursor (CD40L receptor) (B-cell surface antigen CD40) (CDw40) (Bp50). | |||||
|
PHF8_MOUSE
|
||||||
| θ value | 0.21417 (rank : 34) | NC score | 0.045673 (rank : 125) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
TNR11_MOUSE
|
||||||
| θ value | 0.21417 (rank : 35) | NC score | 0.295698 (rank : 21) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O35305, Q8VCT7 | Gene names | Tnfrsf11a, Rank | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 11A precursor (Receptor activator of NF-KB) (Osteoclast differentiation factor receptor) (ODFR). | |||||
|
EPHB2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 36) | NC score | 0.029007 (rank : 135) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 983 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P29323, O43477, Q5T0U6, Q5T0U7, Q5T0U8 | Gene names | EPHB2, DRT, EPTH3, ERK, HEK5 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 2 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor EPH-3) (DRT) (Receptor protein-tyrosine kinase HEK5) (ERK) (NY-REN-47 antigen). | |||||
|
EPHB2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 37) | NC score | 0.028984 (rank : 136) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 982 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P54763, Q62213, Q9QVY4 | Gene names | Ephb2, Epth3, Nuk, Sek3 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 2 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor EPH-3) (Neural kinase) (Nuk receptor tyrosine kinase) (SEK-3). | |||||
|
TNR21_HUMAN
|
||||||
| θ value | 0.279714 (rank : 38) | NC score | 0.228401 (rank : 31) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O75509, Q96D86 | Gene names | TNFRSF21, DR6 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 21 precursor (TNFR- related death receptor 6) (Death receptor 6). | |||||
|
LAMB1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 39) | NC score | 0.094217 (rank : 67) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
EPHA1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 40) | NC score | 0.027878 (rank : 137) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P21709, Q15405 | Gene names | EPHA1, EPH, EPHT, EPHT1 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-A receptor 1 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor EPH). | |||||
|
LAMA2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 41) | NC score | 0.096986 (rank : 64) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
LAMB1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 42) | NC score | 0.097990 (rank : 62) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P02469 | Gene names | Lamb1-1, Lamb-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
STAB1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 43) | NC score | 0.098909 (rank : 60) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8R4Y4, Q8K0K6, Q8VC09 | Gene names | Stab1, Feel1 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein). | |||||
|
MEGF9_HUMAN
|
||||||
| θ value | 0.62314 (rank : 44) | NC score | 0.103252 (rank : 55) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9H1U4, O75098 | Gene names | MEGF9, EGFL5, KIAA0818 | |||
|
Domain Architecture |
|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
USH2A_MOUSE
|
||||||
| θ value | 0.62314 (rank : 45) | NC score | 0.096433 (rank : 65) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 626 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q2QI47, Q9JLP3 | Gene names | Ush2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein homolog) (Usher syndrome type IIa protein homolog). | |||||
|
SREC_HUMAN
|
||||||
| θ value | 0.813845 (rank : 46) | NC score | 0.093486 (rank : 69) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
GON4L_MOUSE
|
||||||
| θ value | 1.06291 (rank : 47) | NC score | 0.027308 (rank : 138) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
LAMC2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 48) | NC score | 0.108519 (rank : 51) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q61092 | Gene names | Lamc2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-2 chain precursor (Laminin 5 gamma 2 subunit) (Kalinin/nicein/epiligrin 100 kDa subunit) (Laminin B2t chain). | |||||
|
LPP_HUMAN
|
||||||
| θ value | 1.06291 (rank : 49) | NC score | 0.015623 (rank : 145) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
VINEX_HUMAN
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.020128 (rank : 141) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60504, Q9UQE4 | Gene names | SORBS3, SCAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Vinexin (Sorbin and SH3 domain-containing protein 3) (SH3-containing adapter molecule 1) (SCAM-1). | |||||
|
I11RA_HUMAN
|
||||||
| θ value | 1.38821 (rank : 51) | NC score | 0.023808 (rank : 139) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14626, Q16542, Q7KYJ7 | Gene names | IL11RA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-11 receptor alpha chain precursor (IL-11R-alpha) (IL- 11RA). | |||||
|
MEGF6_HUMAN
|
||||||
| θ value | 1.38821 (rank : 52) | NC score | 0.072258 (rank : 95) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
SCUB3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 53) | NC score | 0.071925 (rank : 96) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8IX30, Q5CZB3, Q86UZ9, Q8NAU9 | Gene names | SCUBE3, CEGF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal peptide, CUB and EGF-like domain-containing protein 3 precursor. | |||||
|
SCUB3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 54) | NC score | 0.071173 (rank : 97) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q66PY1, Q68FG9 | Gene names | Scube3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal peptide, CUB and EGF-like domain-containing protein 3 precursor. | |||||
|
TARSH_HUMAN
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.037597 (rank : 127) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
ZAR1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 56) | NC score | 0.036652 (rank : 129) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80SU3 | Gene names | Zar1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zygote arrest 1 (Oocyte-specific maternal effect factor). | |||||
|
SREC2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.091609 (rank : 71) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96GP6, Q8IXF3, Q9BW74 | Gene names | SCARF2, SREC2, SREPCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II) (SRECRP-1). | |||||
|
SREC2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.087480 (rank : 76) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 605 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P59222 | Gene names | Scarf2, Srec2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II). | |||||
|
CO9A1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 59) | NC score | 0.017714 (rank : 142) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P20849, Q13699, Q13700, Q5TF52, Q6P467, Q96BM8, Q99225, Q9H151, Q9H152, Q9Y6P2, Q9Y6P3 | Gene names | COL9A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
CO9A1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 60) | NC score | 0.015900 (rank : 144) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q05722, Q61269, Q61270, Q61433, Q61940 | Gene names | Col9a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
EPHB3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.030321 (rank : 132) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 996 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P54754, Q62214 | Gene names | Ephb3, Etk2, Mdk5, Sek4 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 3 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor MDK-5) (Developmental kinase 5) (SEK-4). | |||||
|
EPHB6_MOUSE
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.037269 (rank : 128) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O08644, Q8BN76, Q8K0A9 | Gene names | Ephb6, Cekl | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 6 precursor (Tyrosine-protein kinase-defective receptor EPH-6) (MEP). | |||||
|
LAMA2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.089783 (rank : 72) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
STAB2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.089444 (rank : 73) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8WWQ8, Q6ZMK2, Q7Z5N9, Q86UR4, Q8IUG9, Q8TES1, Q9H7H7, Q9NRY3 | Gene names | STAB2, FEEL2, FELL, FEX2, HARE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stabilin-2 precursor (FEEL-2 protein) (Fasciclin EGF-like laminin-type EGF-like and link domain-containing scavenger receptor 1) (FAS1 EGF- like and X-link domain-containing adhesion molecule 2) (Hyaluronan receptor for endocytosis) [Contains: 190 kDa form stabilin-2 (190 kDa hyaluronan receptor for endocytosis)]. | |||||
|
EPHB3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 65) | NC score | 0.033152 (rank : 130) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 996 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P54753, Q7Z740 | Gene names | EPHB3, ETK2, HEK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 3 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor HEK-2). | |||||
|
LAMA3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 66) | NC score | 0.095362 (rank : 66) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q61789, O08751, Q61788, Q61966, Q9JHQ7 | Gene names | Lama3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Nicein subunit alpha). | |||||
|
TNR22_MOUSE
|
||||||
| θ value | 3.0926 (rank : 67) | NC score | 0.247089 (rank : 28) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9ER62, Q8VHB9, Q9CZA4 | Gene names | Tnfrsf22, Dctrailr2, Tnfrh2, Tnfrsf1al2 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 22 (Tumor necrosis factor receptor p60 homolog 2) (TNF receptor family member SOBa) (Decoy TRAIL receptor 2) (TNF receptor homolog 2). | |||||
|
PHF8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.029790 (rank : 134) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
TNR16_HUMAN
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.217560 (rank : 32) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P08138 | Gene names | NGFR, TNFRSF16 | |||
|
Domain Architecture |
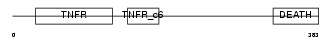
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 16 precursor (Low- affinity nerve growth factor receptor) (NGF receptor) (Gp80-LNGFR) (p75 ICD) (Low affinity neurotrophin receptor p75NTR) (CD271 antigen). | |||||
|
ZN687_HUMAN
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.002828 (rank : 152) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
MEGF8_HUMAN
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.086049 (rank : 80) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q7Z7M0, O75097 | Gene names | MEGF8, EGFL4, KIAA0817 | |||
|
Domain Architecture |
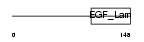
|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
PCSK5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.075195 (rank : 92) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 613 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q04592, Q62040 | Gene names | Pcsk5 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 5 precursor (EC 3.4.21.-) (Proprotein convertase PC5) (Subtilisin/kexin-like protease PC5) (PC6) (Subtilisin-like proprotein convertase 6) (SPC6). | |||||
|
TNR16_MOUSE
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.229069 (rank : 30) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Z0W1 | Gene names | Ngfr, Tnfrsf16 | |||
|
Domain Architecture |
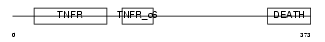
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 16 precursor (Low- affinity nerve growth factor receptor) (NGF receptor) (Low affinity neurotrophin receptor p75NTR). | |||||
|
C1QC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.014154 (rank : 146) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q02105 | Gene names | C1qc, C1qg | |||
|
Domain Architecture |
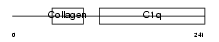
|
|||||
| Description | Complement C1q subcomponent subunit C precursor. | |||||
|
CO5A2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.012918 (rank : 147) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
FNDC3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.023807 (rank : 140) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BX90, Q2VEY7, Q6ZQ15, Q811D3, Q8BTM3 | Gene names | Fndc3a, D14Ertd453e, Fndc3, Kiaa0970 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibronectin type-III domain-containing protein 3a. | |||||
|
LAMA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.100120 (rank : 59) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 838 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P19137 | Gene names | Lama1, Lama, Lama-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
MEGF8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.082188 (rank : 81) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 520 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P60882, Q80TR3, Q80V41, Q8BMN9, Q8JZW7, Q8K0J3 | Gene names | Megf8, Egfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
NOTC3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.044327 (rank : 126) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61982 | Gene names | Notch3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
COBA2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.012167 (rank : 149) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
COFA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.015923 (rank : 143) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O35206, Q9EQD9 | Gene names | Col15a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV)]. | |||||
|
LAMC3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.102139 (rank : 56) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9R0B6, Q9WTW6 | Gene names | Lamc3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
NFAC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.007953 (rank : 150) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88942, O70345 | Gene names | Nfatc1, Nfat2, Nfatc | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 1 (NFAT transcription complex cytosolic component) (NF-ATc1) (NF-ATc). | |||||
|
SEPT9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.012704 (rank : 148) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UHD8, Q96QF3, Q96QF4, Q96QF5, Q9HA04, Q9UG40, Q9Y5W4 | Gene names | SEPT9, KIAA0991, MSF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (MLL septin-like fusion protein) (MLL septin-like fusion protein MSF-A) (Ovarian/Breast septin) (Ov/Br septin) (Septin D1). | |||||
|
TXLNB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.005773 (rank : 151) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
AGRIN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.054665 (rank : 112) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O00468, Q5SVA2, Q60FE1, Q7KYS8, Q8N4J5, Q96IC1, Q9BTD4 | Gene names | AGRIN, AGRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Agrin precursor. | |||||
|
ATRN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.051777 (rank : 118) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O75882, O60295, O95414, Q3MIT3, Q5TDA2, Q5TDA4, Q5VYW3, Q9NTQ3, Q9NTQ4, Q9NU01, Q9NZ57, Q9NZ58 | Gene names | ATRN, KIAA0548, MGCA | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany homolog) (DPPT-L). | |||||
|
ATRN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.050501 (rank : 123) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
CREL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.052694 (rank : 115) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96HD1, Q6I9X5, Q8NFT4, Q9Y409 | Gene names | CRELD1, CIRRIN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich with EGF-like domain protein 1 precursor. | |||||
|
CREL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.050147 (rank : 124) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91XD7, Q8BGJ8 | Gene names | Creld1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich with EGF-like domain protein 1 precursor. | |||||
|
EDAR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.187489 (rank : 39) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UNE0, Q9UND9 | Gene names | EDAR, DL | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member EDAR precursor (Anhidrotic ectodysplasin receptor 1) (Ectodysplasin-A receptor) (EDA- A1 receptor) (Ectodermal dysplasia receptor) (Downless homolog). | |||||
|
EDAR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.189763 (rank : 38) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9R187, Q9DC43 | Gene names | Edar, Dl | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member EDAR precursor (Anhidrotic ectodysplasin receptor 1) (Ectodysplasin-A receptor) (Ectodermal dysplasia receptor) (Downless). | |||||
|
ESM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.050913 (rank : 121) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NQ30, Q15330, Q96ES3 | Gene names | ESM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cell-specific molecule 1 precursor (ESM-1 secretory protein) (ESM-1). | |||||
|
ESM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.050552 (rank : 122) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QYY7 | Gene names | Esm1 | |||
|
Domain Architecture |
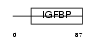
|
|||||
| Description | Endothelial cell-specific molecule 1 precursor (ESM-1 secretory protein) (ESM-1). | |||||
|
FRAS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.051044 (rank : 120) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q86XX4, Q86UZ4, Q8N3U9, Q8NAU7, Q96JW7, Q9H6N9, Q9P228 | Gene names | FRAS1, KIAA1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
FRAS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.052498 (rank : 116) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q80T14, Q80TC7, Q811H8, Q8BPZ4 | Gene names | Fras1, Kiaa1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
KR171_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.056636 (rank : 111) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BYP8 | Gene names | KRTAP17-1, KAP17.1, KRTAP16.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 17-1 (Keratin-associated protein 16.1). | |||||
|
KR510_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.073066 (rank : 94) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6L8G5 | Gene names | KRTAP5-10, KAP5.10, KRTAP5.10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-10 (Keratin-associated protein 5.10) (Ultrahigh sulfur keratin-associated protein 5.10). | |||||
|
KR511_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.066736 (rank : 108) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6L8G4 | Gene names | KRTAP5-11, KAP5.11, KRTAP5.11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-11 (Keratin-associated protein 5.11) (Ultrahigh sulfur keratin-associated protein 5.11). | |||||
|
KRA51_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.078254 (rank : 86) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6L8H4 | Gene names | KRTAP5-1, KAP5.1, KRTAP5.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-1 (Keratin-associated protein 5.1) (Ultrahigh sulfur keratin-associated protein 5.1). | |||||
|
KRA52_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.077153 (rank : 88) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q701N4 | Gene names | KRTAP5-2, KAP5-8, KAP5.2, KRTAP5-8, KRTAP5.2, KRTAP5.8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-2 (Keratin-associated protein 5.2) (Ultrahigh sulfur keratin-associated protein 5.2) (Keratin-associated protein 5-8) (Keratin-associated protein 5.8). | |||||
|
KRA53_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.068264 (rank : 101) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6L8H2, Q6PL44, Q701N3 | Gene names | KRTAP5-3, KAP5-9, KAP5.3, KRTAP5-9, KRTAP5.3, KRTAP5.9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-3 (Keratin-associated protein 5.3) (Ultrahigh sulfur keratin-associated protein 5.3) (Keratin-associated protein 5-9) (Keratin-associated protein 5.9) (UHS KerB-like). | |||||
|
KRA54_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.077618 (rank : 87) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6L8H1 | Gene names | KRTAP5-4, KAP5.4, KRTAP5.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-4 (Keratin-associated protein 5.4) (Ultrahigh sulfur keratin-associated protein 5.4). | |||||
|
KRA55_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.075482 (rank : 91) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q701N2 | Gene names | KRTAP5-5, KAP5-11, KAP5.5, KRTAP5-11, KRTAP5.11, KRTAP5.5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-5 (Keratin-associated protein 5.5) (Ultrahigh sulfur keratin-associated protein 5.5) (Keratin-associated protein 5-11) (Keratin-associated protein 5.11). | |||||
|
KRA56_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.071036 (rank : 98) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6L8G9 | Gene names | KRTAP5-6, KAP5.6, KRTAP5.6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-6 (Keratin-associated protein 5.6) (Ultrahigh sulfur keratin-associated protein 5.6). | |||||
|
KRA57_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.064821 (rank : 109) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6L8G8, Q701N5 | Gene names | KRTAP5-7, KAP5-7, KAP5.3, KRTAP5.3, KRTAP5.7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-7 (Keratin-associated protein 5.7) (Ultrahigh sulfur keratin-associated protein 5.7) (Keratin-associated protein 5-3) (Keratin-associated protein 5.3). | |||||
|
KRA58_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.063463 (rank : 110) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O75690, Q6L8G7, Q6UTX6 | Gene names | KRTAP5-8, KAP5.8, KRTAP5.8, UHSKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-8 (Keratin-associated protein 5.8) (Ultrahigh sulfur keratin-associated protein 5.8) (Keratin, ultra high-sulfur matrix protein B) (UHS keratin B) (UHS KerB). | |||||
|
KRA59_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.078383 (rank : 85) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
LAMA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.093251 (rank : 70) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 804 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P25391 | Gene names | LAMA1, LAMA | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
LAMA3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.089129 (rank : 74) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q16787, Q13679, Q13680, Q96TG0 | Gene names | LAMA3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Epiligrin 170 kDa subunit) (E170) (Nicein subunit alpha). | |||||
|
LAMA4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.078567 (rank : 84) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q16363, Q14735, Q15335, Q4LE44, Q5SZG8, Q9UE18, Q9UJN9 | Gene names | LAMA4 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-4 chain precursor. | |||||
|
LAMA4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.081901 (rank : 82) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P97927, O88785, P70409 | Gene names | Lama4 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-4 chain precursor. | |||||
|
LAMB2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.093845 (rank : 68) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P55268, Q16321 | Gene names | LAMB2, LAMS | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (Laminin B1s chain). | |||||
|
LAMB2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.101628 (rank : 57) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q61292, Q62182 | Gene names | Lamb2, Lams | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (S-LAM). | |||||
|
LAMB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.080327 (rank : 83) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q13751, O14947, Q14733, Q9UJK4, Q9UJL1 | Gene names | LAMB3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-3 chain precursor (Laminin 5 beta 3) (Laminin B1k chain) (Kalinin B1 chain). | |||||
|
LAMB3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.076273 (rank : 89) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61087 | Gene names | Lamb3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-3 chain precursor (Laminin 5 beta 3) (Kalinin B1 chain). | |||||
|
LAMC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.086438 (rank : 79) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P11047 | Gene names | LAMC1, LAMB2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
LAMC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.087739 (rank : 75) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P02468 | Gene names | Lamc1, Lamb-2, Lamc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
LAMC3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.105076 (rank : 54) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9Y6N6 | Gene names | LAMC3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
MEGF9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.087035 (rank : 78) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
NAGPA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.053363 (rank : 114) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UK23, Q96EJ8 | Gene names | NAGPA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
NAGPA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.052441 (rank : 117) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8BJ48, Q3UUT5, Q8CHQ8, Q9QZE6 | Gene names | Nagpa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
NET1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.066748 (rank : 107) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O95631 | Gene names | NTN1, NTN1L | |||
|
Domain Architecture |

|
|||||
| Description | Netrin-1 precursor. | |||||
|
NET1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.067151 (rank : 105) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O09118, Q60832, Q9QY50 | Gene names | Ntn1 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin-1 precursor. | |||||
|
NET2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.069248 (rank : 100) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00634 | Gene names | NTN2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-2-like protein precursor. | |||||
|
NET2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.069330 (rank : 99) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9R1A3, Q9QY49, Q9WVA6 | Gene names | Ntn2l, Ntn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-2-like protein precursor (Netrin-3). | |||||
|
NET4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.076238 (rank : 90) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9HB63, Q658K9, Q7L3F1, Q7L9D6, Q7Z5B6, Q9BZP1, Q9NT44, Q9P133 | Gene names | NTN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-4 precursor (Beta-netrin) (Hepar-derived netrin-like protein). | |||||
|
NET4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.075108 (rank : 93) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JI33 | Gene names | Ntn4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-4 precursor (Beta-netrin). | |||||
|
NTNG1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.067991 (rank : 103) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y2I2, Q5VU86, Q5VU87, Q5VU89, Q5VU90, Q5VU91, Q7Z2Y3, Q8N633 | Gene names | NTNG1, KIAA0976, LMNT1 | |||
|
Domain Architecture |
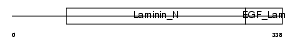
|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
NTNG1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.068025 (rank : 102) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8R4G0, Q68FE5, Q69ZU3, Q8R4F3, Q8R4F4, Q8R4F5, Q8R4F6, Q8R4F7, Q8R4F8, Q8R4F9, Q9ESR3, Q9ESR4, Q9ESR5, Q9ESR6, Q9ESR7, Q9ESR8 | Gene names | Ntng1, Kiaa0976, Lmnt1 | |||
|
Domain Architecture |
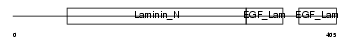
|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
NTNG2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.066960 (rank : 106) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96CW9, Q6UXY0, Q96JH0 | Gene names | NTNG2, KIAA1857, LMNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G2 precursor (Laminet-2). | |||||
|
NTNG2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.067499 (rank : 104) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8R4F1, Q8R4F2, Q8VIP6, Q8VIP7, Q8VIP8 | Gene names | Ntng2, Lmnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G2 precursor (Laminet-2). | |||||
|
PGBM_MOUSE
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.054106 (rank : 113) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q05793 | Gene names | Hspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
TNR11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.262684 (rank : 26) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y6Q6 | Gene names | TNFRSF11A, RANK | |||
|
Domain Architecture |
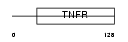
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 11A precursor (Receptor activator of NF-KB) (Osteoclast differentiation factor receptor) (ODFR) (CD265 antigen). | |||||
|
TNR18_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.145636 (rank : 42) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y5U5, O95851, Q9NYJ9 | Gene names | TNFRSF18, AITR, GITR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 18 precursor (Glucocorticoid-induced TNFR-related protein) (Activation-inducible TNFR family receptor). | |||||
|
TNR18_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.110443 (rank : 48) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O35714, Q9JKR1, Q9JKR2, Q9JKR3 | Gene names | Tnfrsf18, Gitr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 18 precursor (Glucocorticoid-induced TNFR-related protein). | |||||
|
TNR21_MOUSE
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.206123 (rank : 34) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9EPU5, Q91W77, Q91XH9 | Gene names | Tnfrsf21, Dr6 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 21 precursor (TNFR- related death receptor 6) (Death receptor 6). | |||||
|
TNR26_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.213730 (rank : 33) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P83626 | Gene names | Tnfrsf26, Tnfrh3 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 26 precursor (TNF receptor homolog 3). | |||||
|
TNR6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.165848 (rank : 40) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P25445, Q14292, Q14293, Q14294, Q14295, Q16652, Q6SSE9 | Gene names | FAS, APT1, FAS1, TNFRSF6 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 6 precursor (FASLG receptor) (Apoptosis-mediating surface antigen FAS) (Apo-1 antigen) (CD95 antigen). | |||||
|
TNR6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.190127 (rank : 37) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P25446, Q9DCQ1 | Gene names | Fas, Apt1, Tnfrsf6 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 6 precursor (FASLG receptor) (Apoptosis-mediating surface antigen FAS) (Apo-1 antigen) (CD95 antigen). | |||||
|
TNR7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.203128 (rank : 35) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P26842 | Gene names | TNFRSF7, CD27 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 7 precursor (CD27L receptor) (T-cell activation antigen CD27) (T14). | |||||
|
TNR7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.192762 (rank : 36) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P41272 | Gene names | Tnfrsf7, Cd27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 7 precursor (CD27L receptor) (T-cell activation antigen CD27). | |||||
|
TNR8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.163476 (rank : 41) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P28908 | Gene names | TNFRSF8, CD30 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 8 precursor (CD30L receptor) (Lymphocyte activation antigen CD30) (KI-1 antigen). | |||||
|
TNR8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.126491 (rank : 43) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q60846 | Gene names | Tnfrsf8 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 8 precursor (CD30L receptor) (Lymphocyte activation antigen CD30). | |||||
|
TNR9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.241576 (rank : 29) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q07011 | Gene names | TNFRSF9, CD137, ILA | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 9 precursor (4-1BB ligand receptor) (T-cell antigen 4-1BB homolog) (T-cell antigen ILA) (CD137 antigen). | |||||
|
TR10A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.106028 (rank : 53) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O00220, Q96E62 | Gene names | TNFRSF10A, APO2, DR4, TRAILR1 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 10A precursor (Death receptor 4) (TNF-related apoptosis-inducing ligand receptor 1) (TRAIL receptor 1) (TRAIL-R1) (CD261 antigen). | |||||
|
TR10B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.109256 (rank : 50) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O14763, O14720, O15508, O15517, O15531, Q9BVE0 | Gene names | TNFRSF10B, DR5, KILLER, TRAILR2, TRICK2, ZTNFR9 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 10B precursor (Death receptor 5) (TNF-related apoptosis-inducing ligand receptor 2) (TRAIL receptor 2) (TRAIL-R2) (CD262 antigen). | |||||
|
TR10B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.109919 (rank : 49) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9QZM4, Q9JJL5, Q9JJL6 | Gene names | Tnfrsf10b, Dr5, Killer | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 10B precursor (Death receptor 5) (MK). | |||||
|
TR10C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.114928 (rank : 47) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O14798, O14755, Q6UXM5 | Gene names | TNFRSF10C, DCR1, LIT, TRAILR3, TRID | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 10C precursor (Decoy receptor 1) (DcR1) (Decoy TRAIL receptor without death domain) (TNF- related apoptosis-inducing ligand receptor 3) (TRAIL receptor 3) (TRAIL-R3) (Trail receptor without an intracellular domain) (Lymphocyte inhibitor of TRAIL) (Antagonist decoy receptor for TRAIL/Apo-2L) (CD263 antigen). | |||||
|
TR10D_HUMAN
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.097256 (rank : 63) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UBN6, Q9Y6Q4 | Gene names | TNFRSF10D, DCR2, TRAILR4, TRUNDD | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 10D precursor (Decoy receptor 2) (DcR2) (TNF-related apoptosis-inducing ligand receptor 4) (TRAIL receptor 4) (TRAIL-R4) (TRAIL receptor with a truncated death domain) (CD264 antigen). | |||||
|
USH2A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.087184 (rank : 77) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O75445, Q5VVM9, Q6S362, Q9NS27 | Gene names | USH2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein) (Usher syndrome type IIa protein). | |||||
|
ZAN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.051544 (rank : 119) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
TR19L_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q8BX43, Q8BTV0 | Gene names | Tnfrsf19l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 19L precursor. | |||||
|
TR19L_HUMAN
|
||||||
| NC score | 0.947634 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q969Z4, Q86V34, Q96JU1, Q9BUX7 | Gene names | TNFRSF19L, RELT | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 19L precursor (Receptor expressed in lymphoid tissues). | |||||
|
CE016_MOUSE
|
||||||
| NC score | 0.472142 (rank : 3) | θ value | 2.98157e-11 (rank : 3) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BRJ3 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C5orf16 homolog. | |||||
|
CE016_HUMAN
|
||||||
| NC score | 0.461099 (rank : 4) | θ value | 3.89403e-11 (rank : 4) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NC24, Q6P4E7, Q6UXY2 | Gene names | C5orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C5orf16. | |||||
|
TNR27_HUMAN
|
||||||
| NC score | 0.456696 (rank : 5) | θ value | 4.0297e-08 (rank : 5) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9HAV5, Q6UWM2, Q8IZA6 | Gene names | EDA2R, TNFRSF27, XEDAR | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 27 (X-linked ectodysplasin-A2 receptor) (EDA-A2 receptor). | |||||
|
TNR27_MOUSE
|
||||||
| NC score | 0.440116 (rank : 6) | θ value | 4.45536e-07 (rank : 6) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BX35, Q8BM50 | Gene names | Eda2r, Tnfrsf27, Xedar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 27 (X-linked ectodysplasin-A2 receptor) (EDA-A2 receptor). | |||||
|
TNR19_MOUSE
|
||||||
| NC score | 0.435297 (rank : 7) | θ value | 2.88788e-06 (rank : 8) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JLL3, Q812G3, Q9JHF1, Q9JJH6, Q9JLL2, Q9QXW7 | Gene names | Tnfrsf19, Taj, Troy | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 19 precursor (Toxicity and JNK inducer) (TRADE). | |||||
|
TNR19_HUMAN
|
||||||
| NC score | 0.409465 (rank : 8) | θ value | 2.88788e-06 (rank : 7) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NS68, Q9BXZ9, Q9BY00, Q9NZV2 | Gene names | TNFRSF19, TAJ, TROY | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 19 precursor (Toxicity and JNK inducer) (TRADE). | |||||
|
TNR6B_HUMAN
|
||||||
| NC score | 0.393947 (rank : 9) | θ value | 6.43352e-06 (rank : 9) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O95407 | Gene names | TNFRSF6B, DCR3, TR6 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 6B precursor (Decoy receptor for Fas ligand) (Decoy receptor 3) (DcR3) (M68). | |||||
|
TNR4_HUMAN
|
||||||
| NC score | 0.372552 (rank : 10) | θ value | 7.1131e-05 (rank : 11) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P43489, Q13663, Q5T7M0 | Gene names | TNFRSF4, TXGP1L | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 4 precursor (OX40L receptor) (ACT35 antigen) (TAX transcriptionally-activated glycoprotein 1 receptor) (CD134 antigen). | |||||
|
TNR3_HUMAN
|
||||||
| NC score | 0.364168 (rank : 11) | θ value | 0.00134147 (rank : 13) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P36941 | Gene names | LTBR, TNFCR, TNFRSF3 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 3 precursor (Lymphotoxin-beta receptor) (Tumor necrosis factor receptor 2-related protein) (Tumor necrosis factor C receptor). | |||||
|
TNR4_MOUSE
|
||||||
| NC score | 0.361862 (rank : 12) | θ value | 0.00175202 (rank : 14) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P47741 | Gene names | Tnfrsf4, Ox40, Txgp1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 4 precursor (OX40L receptor) (OX40 antigen). | |||||
|
TR11B_MOUSE
|
||||||
| NC score | 0.359034 (rank : 13) | θ value | 0.000158464 (rank : 12) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O08712, O70202 | Gene names | Tnfrsf11b, Ocif, Opg | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 11B precursor (Osteoprotegerin) (Osteoclastogenesis inhibitory factor). | |||||
|
TR11B_HUMAN
|
||||||
| NC score | 0.353333 (rank : 14) | θ value | 4.1701e-05 (rank : 10) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O00300, O60236, Q53FX6, Q9UHP4 | Gene names | TNFRSF11B, OCIF, OPG | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 11B precursor (Osteoprotegerin) (Osteoclastogenesis inhibitory factor). | |||||
|
TNR3_MOUSE
|
||||||
| NC score | 0.352417 (rank : 15) | θ value | 0.00298849 (rank : 16) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P50284 | Gene names | Ltbr, Tnfcr, Tnfrsf3 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 3 precursor (Lymphotoxin-beta receptor). | |||||
|
TNR14_HUMAN
|
||||||
| NC score | 0.342170 (rank : 16) | θ value | 0.0252991 (rank : 20) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q92956, Q8WXR1, Q96J31, Q9UM65 | Gene names | TNFRSF14, HVEA, HVEM | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 14 precursor (Herpesvirus entry mediator A) (Tumor necrosis factor receptor-like 2) (TR2). | |||||
|
TNR9_MOUSE
|
||||||
| NC score | 0.332414 (rank : 17) | θ value | 0.00390308 (rank : 18) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P20334 | Gene names | Tnfrsf9, Cd137, Cd157, Ila, Ly63 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 9 precursor (4-1BB ligand receptor) (T-cell antigen 4-1BB) (CD137 antigen). | |||||
|
TNR1B_MOUSE
|
||||||
| NC score | 0.301421 (rank : 18) | θ value | 0.0330416 (rank : 22) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P25119, O88734, P97893 | Gene names | Tnfrsf1b, Tnfr-2, Tnfr2 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 1B precursor (Tumor necrosis factor receptor 2) (TNF-R2) (Tumor necrosis factor receptor type II) (p75) (p80 TNF-alpha receptor). | |||||
|
TNR1A_MOUSE
|
||||||
| NC score | 0.300932 (rank : 19) | θ value | 0.00390308 (rank : 17) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P25118 | Gene names | Tnfrsf1a, Tnfr-1, Tnfr1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 1A precursor (p60) (TNF-R1) (TNF-RI) (TNFR-I) (p55). | |||||
|
TNR1B_HUMAN
|
||||||
| NC score | 0.299658 (rank : 20) | θ value | 0.125558 (rank : 31) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P20333, Q16042, Q6YI29, Q9UIH1 | Gene names | TNFRSF1B, TNFBR, TNFR2 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 1B precursor (Tumor necrosis factor receptor 2) (TNF-R2) (Tumor necrosis factor receptor type II) (p75) (p80 TNF-alpha receptor) (CD120b antigen) (Etanercept) [Contains: Tumor necrosis factor receptor superfamily member 1b, membrane form; Tumor necrosis factor-binding protein 2 (TBPII) (TBP- 2)]. | |||||
|
TNR11_MOUSE
|
||||||
| NC score | 0.295698 (rank : 21) | θ value | 0.21417 (rank : 35) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O35305, Q8VCT7 | Gene names | Tnfrsf11a, Rank | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 11A precursor (Receptor activator of NF-KB) (Osteoclast differentiation factor receptor) (ODFR). | |||||
|
TNR5_HUMAN
|
||||||
| NC score | 0.295618 (rank : 22) | θ value | 0.163984 (rank : 33) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P25942, Q9BYU0 | Gene names | CD40, TNFRSF5 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 5 precursor (CD40L receptor) (B-cell surface antigen CD40) (CDw40) (Bp50). | |||||
|
TNR5_MOUSE
|
||||||
| NC score | 0.294008 (rank : 23) | θ value | 0.00228821 (rank : 15) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P27512, Q99NE0, Q99NE1, Q99NE2, Q99NE3 | Gene names | Cd40, Tnfrsf5 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 5 precursor (CD40L receptor) (B-cell surface antigen CD40) (BP50) (CDw40). | |||||
|
TNR1A_HUMAN
|
||||||
| NC score | 0.281882 (rank : 24) | θ value | 0.0330416 (rank : 21) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P19438 | Gene names | TNFRSF1A, TNFAR, TNFR1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 1A precursor (p60) (TNF-R1) (TNF-RI) (TNFR-I) (p55) (CD120a antigen) [Contains: Tumor necrosis factor receptor superfamily member 1A, membrane form; Tumor necrosis factor-binding protein 1 (TBPI)]. | |||||
|
TNR25_HUMAN
|
||||||
| NC score | 0.265406 (rank : 25) | θ value | 0.0961366 (rank : 29) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q93038, O00275, O00276, O00277, O00278, O00279, O00280, O14865, O14866, P78507, P78515, Q92983, Q93036, Q93037, Q99722, Q99830, Q99831, Q9BY86, Q9UME0, Q9UME1, Q9UME5 | Gene names | TNFRSF25, APO3, DDR3, DR3, TNFRSF12, WSL, WSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 25 precursor (WSL-1 protein) (Apoptosis-mediating receptor DR3) (Apoptosis-mediating receptor TRAMP) (Death domain receptor 3) (WSL protein) (Apoptosis- inducing receptor AIR) (Apo-3) (Lymphocyte-associated receptor of death) (LARD). | |||||
|
TNR11_HUMAN
|
||||||
| NC score | 0.262684 (rank : 26) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y6Q6 | Gene names | TNFRSF11A, RANK | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 11A precursor (Receptor activator of NF-KB) (Osteoclast differentiation factor receptor) (ODFR) (CD265 antigen). | |||||
|
TNR23_MOUSE
|
||||||
| NC score | 0.261328 (rank : 27) | θ value | 0.0431538 (rank : 23) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9ER63, Q8VHC0 | Gene names | Tnfrsf23, Dctrailr1, Tnfrh1, Tnfrsf1al1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 23 precursor (Tumor necrosis factor receptor p60 homolog 1) (TNF receptor family member SOB) (Decoy TRAIL receptor 1) (TNF receptor homolog 1). | |||||
|
TNR22_MOUSE
|
||||||
| NC score | 0.247089 (rank : 28) | θ value | 3.0926 (rank : 67) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9ER62, Q8VHB9, Q9CZA4 | Gene names | Tnfrsf22, Dctrailr2, Tnfrh2, Tnfrsf1al2 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 22 (Tumor necrosis factor receptor p60 homolog 2) (TNF receptor family member SOBa) (Decoy TRAIL receptor 2) (TNF receptor homolog 2). | |||||
|
TNR9_HUMAN
|
||||||
| NC score | 0.241576 (rank : 29) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q07011 | Gene names | TNFRSF9, CD137, ILA | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 9 precursor (4-1BB ligand receptor) (T-cell antigen 4-1BB homolog) (T-cell antigen ILA) (CD137 antigen). | |||||
|
TNR16_MOUSE
|
||||||
| NC score | 0.229069 (rank : 30) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Z0W1 | Gene names | Ngfr, Tnfrsf16 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 16 precursor (Low- affinity nerve growth factor receptor) (NGF receptor) (Low affinity neurotrophin receptor p75NTR). | |||||
|
TNR21_HUMAN
|
||||||
| NC score | 0.228401 (rank : 31) | θ value | 0.279714 (rank : 38) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O75509, Q96D86 | Gene names | TNFRSF21, DR6 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 21 precursor (TNFR- related death receptor 6) (Death receptor 6). | |||||
|
TNR16_HUMAN
|
||||||
| NC score | 0.217560 (rank : 32) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P08138 | Gene names | NGFR, TNFRSF16 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 16 precursor (Low- affinity nerve growth factor receptor) (NGF receptor) (Gp80-LNGFR) (p75 ICD) (Low affinity neurotrophin receptor p75NTR) (CD271 antigen). | |||||
|
TNR26_MOUSE
|
||||||
| NC score | 0.213730 (rank : 33) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P83626 | Gene names | Tnfrsf26, Tnfrh3 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 26 precursor (TNF receptor homolog 3). | |||||
|
TNR21_MOUSE
|
||||||
| NC score | 0.206123 (rank : 34) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9EPU5, Q91W77, Q91XH9 | Gene names | Tnfrsf21, Dr6 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 21 precursor (TNFR- related death receptor 6) (Death receptor 6). | |||||
|
TNR7_HUMAN
|
||||||
| NC score | 0.203128 (rank : 35) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P26842 | Gene names | TNFRSF7, CD27 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 7 precursor (CD27L receptor) (T-cell activation antigen CD27) (T14). | |||||
|
TNR7_MOUSE
|
||||||
| NC score | 0.192762 (rank : 36) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P41272 | Gene names | Tnfrsf7, Cd27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 7 precursor (CD27L receptor) (T-cell activation antigen CD27). | |||||
|
TNR6_MOUSE
|
||||||
| NC score | 0.190127 (rank : 37) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P25446, Q9DCQ1 | Gene names | Fas, Apt1, Tnfrsf6 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 6 precursor (FASLG receptor) (Apoptosis-mediating surface antigen FAS) (Apo-1 antigen) (CD95 antigen). | |||||
|
EDAR_MOUSE
|
||||||
| NC score | 0.189763 (rank : 38) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9R187, Q9DC43 | Gene names | Edar, Dl | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member EDAR precursor (Anhidrotic ectodysplasin receptor 1) (Ectodysplasin-A receptor) (Ectodermal dysplasia receptor) (Downless). | |||||
|
EDAR_HUMAN
|
||||||
| NC score | 0.187489 (rank : 39) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UNE0, Q9UND9 | Gene names | EDAR, DL | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member EDAR precursor (Anhidrotic ectodysplasin receptor 1) (Ectodysplasin-A receptor) (EDA- A1 receptor) (Ectodermal dysplasia receptor) (Downless homolog). | |||||
|
TNR6_HUMAN
|
||||||
| NC score | 0.165848 (rank : 40) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P25445, Q14292, Q14293, Q14294, Q14295, Q16652, Q6SSE9 | Gene names | FAS, APT1, FAS1, TNFRSF6 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 6 precursor (FASLG receptor) (Apoptosis-mediating surface antigen FAS) (Apo-1 antigen) (CD95 antigen). | |||||
|
TNR8_HUMAN
|
||||||
| NC score | 0.163476 (rank : 41) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P28908 | Gene names | TNFRSF8, CD30 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 8 precursor (CD30L receptor) (Lymphocyte activation antigen CD30) (KI-1 antigen). | |||||
|
TNR18_HUMAN
|
||||||
| NC score | 0.145636 (rank : 42) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y5U5, O95851, Q9NYJ9 | Gene names | TNFRSF18, AITR, GITR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 18 precursor (Glucocorticoid-induced TNFR-related protein) (Activation-inducible TNFR family receptor). | |||||
|
TNR8_MOUSE
|
||||||
| NC score | 0.126491 (rank : 43) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q60846 | Gene names | Tnfrsf8 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 8 precursor (CD30L receptor) (Lymphocyte activation antigen CD30). | |||||
|
LAMA5_HUMAN
|
||||||
| NC score | 0.126175 (rank : 44) | θ value | 0.0252991 (rank : 19) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O15230, Q8WZA7, Q9H1P1 | Gene names | LAMA5, KIAA0533, KIAA1907 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
CS034_HUMAN
|
||||||
| NC score | 0.117008 (rank : 45) | θ value | 0.0961366 (rank : 26) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NCQ2 | Gene names | C19orf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf34. | |||||
|
LAMA5_MOUSE
|
||||||
| NC score | 0.115380 (rank : 46) | θ value | 0.0961366 (rank : 27) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q61001, Q9JHQ6 | Gene names | Lama5 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
TR10C_HUMAN
|
||||||
| NC score | 0.114928 (rank : 47) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O14798, O14755, Q6UXM5 | Gene names | TNFRSF10C, DCR1, LIT, TRAILR3, TRID | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 10C precursor (Decoy receptor 1) (DcR1) (Decoy TRAIL receptor without death domain) (TNF- related apoptosis-inducing ligand receptor 3) (TRAIL receptor 3) (TRAIL-R3) (Trail receptor without an intracellular domain) (Lymphocyte inhibitor of TRAIL) (Antagonist decoy receptor for TRAIL/Apo-2L) (CD263 antigen). | |||||
|
TNR18_MOUSE
|
||||||
| NC score | 0.110443 (rank : 48) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O35714, Q9JKR1, Q9JKR2, Q9JKR3 | Gene names | Tnfrsf18, Gitr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 18 precursor (Glucocorticoid-induced TNFR-related protein). | |||||
|
TR10B_MOUSE
|
||||||
| NC score | 0.109919 (rank : 49) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9QZM4, Q9JJL5, Q9JJL6 | Gene names | Tnfrsf10b, Dr5, Killer | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 10B precursor (Death receptor 5) (MK). | |||||
|
TR10B_HUMAN
|
||||||
| NC score | 0.109256 (rank : 50) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O14763, O14720, O15508, O15517, O15531, Q9BVE0 | Gene names | TNFRSF10B, DR5, KILLER, TRAILR2, TRICK2, ZTNFR9 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 10B precursor (Death receptor 5) (TNF-related apoptosis-inducing ligand receptor 2) (TRAIL receptor 2) (TRAIL-R2) (CD262 antigen). | |||||
|
LAMC2_MOUSE
|
||||||
| NC score | 0.108519 (rank : 51) | θ value | 1.06291 (rank : 48) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q61092 | Gene names | Lamc2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-2 chain precursor (Laminin 5 gamma 2 subunit) (Kalinin/nicein/epiligrin 100 kDa subunit) (Laminin B2t chain). | |||||
|
LAMC2_HUMAN
|
||||||
| NC score | 0.106629 (rank : 52) | θ value | 0.0736092 (rank : 25) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q13753, Q02536, Q02537, Q13752, Q14941, Q5VYE8 | Gene names | LAMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-2 chain precursor (Laminin 5 gamma 2 subunit) (Kalinin/nicein/epiligrin 100 kDa subunit) (Laminin B2t chain) (Cell- scattering factor 140 kDa subunit) (CSF 140 kDa subunit) (Large adhesive scatter factor 140 kDa subunit) (Ladsin 140 kDa subunit). | |||||
|
TR10A_HUMAN
|
||||||
| NC score | 0.106028 (rank : 53) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O00220, Q96E62 | Gene names | TNFRSF10A, APO2, DR4, TRAILR1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 10A precursor (Death receptor 4) (TNF-related apoptosis-inducing ligand receptor 1) (TRAIL receptor 1) (TRAIL-R1) (CD261 antigen). | |||||
|
LAMC3_HUMAN
|
||||||
| NC score | 0.105076 (rank : 54) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9Y6N6 | Gene names | LAMC3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
MEGF9_HUMAN
|
||||||
| NC score | 0.103252 (rank : 55) | θ value | 0.62314 (rank : 44) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9H1U4, O75098 | Gene names | MEGF9, EGFL5, KIAA0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
LAMC3_MOUSE
|
||||||
| NC score | 0.102139 (rank : 56) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9R0B6, Q9WTW6 | Gene names | Lamc3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
LAMB2_MOUSE
|
||||||
| NC score | 0.101628 (rank : 57) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q61292, Q62182 | Gene names | Lamb2, Lams | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (S-LAM). | |||||
|
STAB1_HUMAN
|
||||||
| NC score | 0.100351 (rank : 58) | θ value | 0.0961366 (rank : 28) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 546 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9NY15, Q8IUH0, Q8IUH1, Q93072 | Gene names | STAB1, FEEL1, KIAA0246 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein) (MS-1 antigen). | |||||
|
LAMA1_MOUSE
|
||||||
| NC score | 0.100120 (rank : 59) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 838 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P19137 | Gene names | Lama1, Lama, Lama-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
STAB1_MOUSE
|
||||||
| NC score | 0.098909 (rank : 60) | θ value | 0.47712 (rank : 43) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8R4Y4, Q8K0K6, Q8VC09 | Gene names | Stab1, Feel1 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein). | |||||
|
STAB2_MOUSE
|
||||||
| NC score | 0.098124 (rank : 61) | θ value | 0.125558 (rank : 30) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8R4U0, Q8BM87 | Gene names | Stab2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stabilin-2 precursor [Contains: Short form stabilin-2]. | |||||
|
LAMB1_MOUSE
|
||||||
| NC score | 0.097990 (rank : 62) | θ value | 0.47712 (rank : 42) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P02469 | Gene names | Lamb1-1, Lamb-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
TR10D_HUMAN
|
||||||
| NC score | 0.097256 (rank : 63) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UBN6, Q9Y6Q4 | Gene names | TNFRSF10D, DCR2, TRAILR4, TRUNDD | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 10D precursor (Decoy receptor 2) (DcR2) (TNF-related apoptosis-inducing ligand receptor 4) (TRAIL receptor 4) (TRAIL-R4) (TRAIL receptor with a truncated death domain) (CD264 antigen). | |||||
|
LAMA2_HUMAN
|
||||||
| NC score | 0.096986 (rank : 64) | θ value | 0.47712 (rank : 41) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
USH2A_MOUSE
|
||||||
| NC score | 0.096433 (rank : 65) | θ value | 0.62314 (rank : 45) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 626 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q2QI47, Q9JLP3 | Gene names | Ush2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein homolog) (Usher syndrome type IIa protein homolog). | |||||
|
LAMA3_MOUSE
|
||||||
| NC score | 0.095362 (rank : 66) | θ value | 3.0926 (rank : 66) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q61789, O08751, Q61788, Q61966, Q9JHQ7 | Gene names | Lama3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Nicein subunit alpha). | |||||
|
LAMB1_HUMAN
|
||||||
| NC score | 0.094217 (rank : 67) | θ value | 0.365318 (rank : 39) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
LAMB2_HUMAN
|
||||||
| NC score | 0.093845 (rank : 68) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P55268, Q16321 | Gene names | LAMB2, LAMS | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (Laminin B1s chain). | |||||
|
SREC_HUMAN
|
||||||
| NC score | 0.093486 (rank : 69) | θ value | 0.813845 (rank : 46) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
LAMA1_HUMAN
|
||||||
| NC score | 0.093251 (rank : 70) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 804 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P25391 | Gene names | LAMA1, LAMA | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
SREC2_HUMAN
|
||||||
| NC score | 0.091609 (rank : 71) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96GP6, Q8IXF3, Q9BW74 | Gene names | SCARF2, SREC2, SREPCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II) (SRECRP-1). | |||||
|
LAMA2_MOUSE
|
||||||
| NC score | 0.089783 (rank : 72) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
STAB2_HUMAN
|
||||||
| NC score | 0.089444 (rank : 73) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8WWQ8, Q6ZMK2, Q7Z5N9, Q86UR4, Q8IUG9, Q8TES1, Q9H7H7, Q9NRY3 | Gene names | STAB2, FEEL2, FELL, FEX2, HARE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stabilin-2 precursor (FEEL-2 protein) (Fasciclin EGF-like laminin-type EGF-like and link domain-containing scavenger receptor 1) (FAS1 EGF- like and X-link domain-containing adhesion molecule 2) (Hyaluronan receptor for endocytosis) [Contains: 190 kDa form stabilin-2 (190 kDa hyaluronan receptor for endocytosis)]. | |||||
|
LAMA3_HUMAN
|
||||||
| NC score | 0.089129 (rank : 74) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q16787, Q13679, Q13680, Q96TG0 | Gene names | LAMA3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Epiligrin 170 kDa subunit) (E170) (Nicein subunit alpha). | |||||
|
LAMC1_MOUSE
|
||||||
| NC score | 0.087739 (rank : 75) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P02468 | Gene names | Lamc1, Lamb-2, Lamc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
SREC2_MOUSE
|
||||||
| NC score | 0.087480 (rank : 76) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 605 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P59222 | Gene names | Scarf2, Srec2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II). | |||||
|
USH2A_HUMAN
|
||||||
| NC score | 0.087184 (rank : 77) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O75445, Q5VVM9, Q6S362, Q9NS27 | Gene names | USH2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein) (Usher syndrome type IIa protein). | |||||
|
MEGF9_MOUSE
|
||||||
| NC score | 0.087035 (rank : 78) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
LAMC1_HUMAN
|
||||||
| NC score | 0.086438 (rank : 79) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P11047 | Gene names | LAMC1, LAMB2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
MEGF8_HUMAN
|
||||||
| NC score | 0.086049 (rank : 80) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q7Z7M0, O75097 | Gene names | MEGF8, EGFL4, KIAA0817 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
MEGF8_MOUSE
|
||||||
| NC score | 0.082188 (rank : 81) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 520 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P60882, Q80TR3, Q80V41, Q8BMN9, Q8JZW7, Q8K0J3 | Gene names | Megf8, Egfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
LAMA4_MOUSE
|
||||||
| NC score | 0.081901 (rank : 82) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P97927, O88785, P70409 | Gene names | Lama4 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-4 chain precursor. | |||||
|
LAMB3_HUMAN
|
||||||
| NC score | 0.080327 (rank : 83) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q13751, O14947, Q14733, Q9UJK4, Q9UJL1 | Gene names | LAMB3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-3 chain precursor (Laminin 5 beta 3) (Laminin B1k chain) (Kalinin B1 chain). | |||||
|
LAMA4_HUMAN
|
||||||
| NC score | 0.078567 (rank : 84) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q16363, Q14735, Q15335, Q4LE44, Q5SZG8, Q9UE18, Q9UJN9 | Gene names | LAMA4 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-4 chain precursor. | |||||
|
KRA59_HUMAN
|
||||||
| NC score | 0.078383 (rank : 85) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
KRA51_HUMAN
|
||||||
| NC score | 0.078254 (rank : 86) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6L8H4 | Gene names | KRTAP5-1, KAP5.1, KRTAP5.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-1 (Keratin-associated protein 5.1) (Ultrahigh sulfur keratin-associated protein 5.1). | |||||
|
KRA54_HUMAN
|
||||||
| NC score | 0.077618 (rank : 87) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6L8H1 | Gene names | KRTAP5-4, KAP5.4, KRTAP5.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-4 (Keratin-associated protein 5.4) (Ultrahigh sulfur keratin-associated protein 5.4). | |||||
|
KRA52_HUMAN
|
||||||
| NC score | 0.077153 (rank : 88) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q701N4 | Gene names | KRTAP5-2, KAP5-8, KAP5.2, KRTAP5-8, KRTAP5.2, KRTAP5.8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-2 (Keratin-associated protein 5.2) (Ultrahigh sulfur keratin-associated protein 5.2) (Keratin-associated protein 5-8) (Keratin-associated protein 5.8). | |||||
|
LAMB3_MOUSE
|
||||||
| NC score | 0.076273 (rank : 89) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61087 | Gene names | Lamb3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-3 chain precursor (Laminin 5 beta 3) (Kalinin B1 chain). | |||||
|
NET4_HUMAN
|
||||||
| NC score | 0.076238 (rank : 90) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9HB63, Q658K9, Q7L3F1, Q7L9D6, Q7Z5B6, Q9BZP1, Q9NT44, Q9P133 | Gene names | NTN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-4 precursor (Beta-netrin) (Hepar-derived netrin-like protein). | |||||
|
KRA55_HUMAN
|
||||||
| NC score | 0.075482 (rank : 91) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q701N2 | Gene names | KRTAP5-5, KAP5-11, KAP5.5, KRTAP5-11, KRTAP5.11, KRTAP5.5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-5 (Keratin-associated protein 5.5) (Ultrahigh sulfur keratin-associated protein 5.5) (Keratin-associated protein 5-11) (Keratin-associated protein 5.11). | |||||
|
PCSK5_MOUSE
|
||||||
| NC score | 0.075195 (rank : 92) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 613 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q04592, Q62040 | Gene names | Pcsk5 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 5 precursor (EC 3.4.21.-) (Proprotein convertase PC5) (Subtilisin/kexin-like protease PC5) (PC6) (Subtilisin-like proprotein convertase 6) (SPC6). | |||||
|
NET4_MOUSE
|
||||||
| NC score | 0.075108 (rank : 93) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JI33 | Gene names | Ntn4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-4 precursor (Beta-netrin). | |||||
|
KR510_HUMAN
|
||||||
| NC score | 0.073066 (rank : 94) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6L8G5 | Gene names | KRTAP5-10, KAP5.10, KRTAP5.10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-10 (Keratin-associated protein 5.10) (Ultrahigh sulfur keratin-associated protein 5.10). | |||||
|
MEGF6_HUMAN
|
||||||
| NC score | 0.072258 (rank : 95) | θ value | 1.38821 (rank : 52) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
SCUB3_HUMAN
|
||||||
| NC score | 0.071925 (rank : 96) | θ value | 1.38821 (rank : 53) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8IX30, Q5CZB3, Q86UZ9, Q8NAU9 | Gene names | SCUBE3, CEGF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal peptide, CUB and EGF-like domain-containing protein 3 precursor. | |||||
|
SCUB3_MOUSE
|
||||||
| NC score | 0.071173 (rank : 97) | θ value | 1.38821 (rank : 54) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q66PY1, Q68FG9 | Gene names | Scube3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal peptide, CUB and EGF-like domain-containing protein 3 precursor. | |||||
|
KRA56_HUMAN
|
||||||
| NC score | 0.071036 (rank : 98) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6L8G9 | Gene names | KRTAP5-6, KAP5.6, KRTAP5.6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-6 (Keratin-associated protein 5.6) (Ultrahigh sulfur keratin-associated protein 5.6). | |||||
|
NET2_MOUSE
|
||||||
| NC score | 0.069330 (rank : 99) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9R1A3, Q9QY49, Q9WVA6 | Gene names | Ntn2l, Ntn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-2-like protein precursor (Netrin-3). | |||||
|
NET2_HUMAN
|
||||||
| NC score | 0.069248 (rank : 100) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00634 | Gene names | NTN2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-2-like protein precursor. | |||||
|
KRA53_HUMAN
|
||||||
| NC score | 0.068264 (rank : 101) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6L8H2, Q6PL44, Q701N3 | Gene names | KRTAP5-3, KAP5-9, KAP5.3, KRTAP5-9, KRTAP5.3, KRTAP5.9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-3 (Keratin-associated protein 5.3) (Ultrahigh sulfur keratin-associated protein 5.3) (Keratin-associated protein 5-9) (Keratin-associated protein 5.9) (UHS KerB-like). | |||||
|
NTNG1_MOUSE
|
||||||
| NC score | 0.068025 (rank : 102) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8R4G0, Q68FE5, Q69ZU3, Q8R4F3, Q8R4F4, Q8R4F5, Q8R4F6, Q8R4F7, Q8R4F8, Q8R4F9, Q9ESR3, Q9ESR4, Q9ESR5, Q9ESR6, Q9ESR7, Q9ESR8 | Gene names | Ntng1, Kiaa0976, Lmnt1 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
NTNG1_HUMAN
|
||||||
| NC score | 0.067991 (rank : 103) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y2I2, Q5VU86, Q5VU87, Q5VU89, Q5VU90, Q5VU91, Q7Z2Y3, Q8N633 | Gene names | NTNG1, KIAA0976, LMNT1 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
NTNG2_MOUSE
|
||||||
| NC score | 0.067499 (rank : 104) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8R4F1, Q8R4F2, Q8VIP6, Q8VIP7, Q8VIP8 | Gene names | Ntng2, Lmnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G2 precursor (Laminet-2). | |||||
|
NET1_MOUSE
|
||||||
| NC score | 0.067151 (rank : 105) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O09118, Q60832, Q9QY50 | Gene names | Ntn1 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin-1 precursor. | |||||
|
NTNG2_HUMAN
|
||||||
| NC score | 0.066960 (rank : 106) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96CW9, Q6UXY0, Q96JH0 | Gene names | NTNG2, KIAA1857, LMNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G2 precursor (Laminet-2). | |||||
|
NET1_HUMAN
|
||||||
| NC score | 0.066748 (rank : 107) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O95631 | Gene names | NTN1, NTN1L | |||
|
Domain Architecture |

|
|||||
| Description | Netrin-1 precursor. | |||||
|
KR511_HUMAN
|
||||||
| NC score | 0.066736 (rank : 108) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6L8G4 | Gene names | KRTAP5-11, KAP5.11, KRTAP5.11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-11 (Keratin-associated protein 5.11) (Ultrahigh sulfur keratin-associated protein 5.11). | |||||
|
KRA57_HUMAN
|
||||||
| NC score | 0.064821 (rank : 109) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6L8G8, Q701N5 | Gene names | KRTAP5-7, KAP5-7, KAP5.3, KRTAP5.3, KRTAP5.7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-7 (Keratin-associated protein 5.7) (Ultrahigh sulfur keratin-associated protein 5.7) (Keratin-associated protein 5-3) (Keratin-associated protein 5.3). | |||||
|
KRA58_HUMAN
|
||||||
| NC score | 0.063463 (rank : 110) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O75690, Q6L8G7, Q6UTX6 | Gene names | KRTAP5-8, KAP5.8, KRTAP5.8, UHSKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-8 (Keratin-associated protein 5.8) (Ultrahigh sulfur keratin-associated protein 5.8) (Keratin, ultra high-sulfur matrix protein B) (UHS keratin B) (UHS KerB). | |||||
|
KR171_HUMAN
|
||||||
| NC score | 0.056636 (rank : 111) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BYP8 | Gene names | KRTAP17-1, KAP17.1, KRTAP16.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 17-1 (Keratin-associated protein 16.1). | |||||
|
AGRIN_HUMAN
|
||||||
| NC score | 0.054665 (rank : 112) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O00468, Q5SVA2, Q60FE1, Q7KYS8, Q8N4J5, Q96IC1, Q9BTD4 | Gene names | AGRIN, AGRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Agrin precursor. | |||||
|
PGBM_MOUSE
|
||||||
| NC score | 0.054106 (rank : 113) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q05793 | Gene names | Hspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
NAGPA_HUMAN
|
||||||
| NC score | 0.053363 (rank : 114) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UK23, Q96EJ8 | Gene names | NAGPA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
CREL1_HUMAN
|
||||||
| NC score | 0.052694 (rank : 115) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96HD1, Q6I9X5, Q8NFT4, Q9Y409 | Gene names | CRELD1, CIRRIN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich with EGF-like domain protein 1 precursor. | |||||
|
FRAS1_MOUSE
|
||||||
| NC score | 0.052498 (rank : 116) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q80T14, Q80TC7, Q811H8, Q8BPZ4 | Gene names | Fras1, Kiaa1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
NAGPA_MOUSE
|
||||||
| NC score | 0.052441 (rank : 117) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8BJ48, Q3UUT5, Q8CHQ8, Q9QZE6 | Gene names | Nagpa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
ATRN_HUMAN
|
||||||
| NC score | 0.051777 (rank : 118) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O75882, O60295, O95414, Q3MIT3, Q5TDA2, Q5TDA4, Q5VYW3, Q9NTQ3, Q9NTQ4, Q9NU01, Q9NZ57, Q9NZ58 | Gene names | ATRN, KIAA0548, MGCA | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany homolog) (DPPT-L). | |||||
|
ZAN_MOUSE
|
||||||
| NC score | 0.051544 (rank : 119) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
FRAS1_HUMAN
|
||||||
| NC score | 0.051044 (rank : 120) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q86XX4, Q86UZ4, Q8N3U9, Q8NAU7, Q96JW7, Q9H6N9, Q9P228 | Gene names | FRAS1, KIAA1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
ESM1_HUMAN
|
||||||
| NC score | 0.050913 (rank : 121) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NQ30, Q15330, Q96ES3 | Gene names | ESM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cell-specific molecule 1 precursor (ESM-1 secretory protein) (ESM-1). | |||||
|
ESM1_MOUSE
|
||||||
| NC score | 0.050552 (rank : 122) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QYY7 | Gene names | Esm1 | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial cell-specific molecule 1 precursor (ESM-1 secretory protein) (ESM-1). | |||||
|
ATRN_MOUSE
|
||||||
| NC score | 0.050501 (rank : 123) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
CREL1_MOUSE
|
||||||
| NC score | 0.050147 (rank : 124) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91XD7, Q8BGJ8 | Gene names | Creld1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich with EGF-like domain protein 1 precursor. | |||||
|
PHF8_MOUSE
|
||||||
| NC score | 0.045673 (rank : 125) | θ value | 0.21417 (rank : 34) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
NOTC3_MOUSE
|
||||||
| NC score | 0.044327 (rank : 126) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61982 | Gene names | Notch3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
TARSH_HUMAN
|
||||||
| NC score | 0.037597 (rank : 127) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
EPHB6_MOUSE
|
||||||
| NC score | 0.037269 (rank : 128) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O08644, Q8BN76, Q8K0A9 | Gene names | Ephb6, Cekl | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 6 precursor (Tyrosine-protein kinase-defective receptor EPH-6) (MEP). | |||||
|
ZAR1_MOUSE
|
||||||
| NC score | 0.036652 (rank : 129) | θ value | 1.38821 (rank : 56) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80SU3 | Gene names | Zar1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zygote arrest 1 (Oocyte-specific maternal effect factor). | |||||
|
EPHB3_HUMAN
|
||||||
| NC score | 0.033152 (rank : 130) | θ value | 3.0926 (rank : 65) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 996 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P54753, Q7Z740 | Gene names | EPHB3, ETK2, HEK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 3 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor HEK-2). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.032484 (rank : 131) | θ value | 0.0563607 (rank : 24) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
EPHB3_MOUSE
|
||||||
| NC score | 0.030321 (rank : 132) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 996 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P54754, Q62214 | Gene names | Ephb3, Etk2, Mdk5, Sek4 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 3 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor MDK-5) (Developmental kinase 5) (SEK-4). | |||||
|
EPHB1_HUMAN
|
||||||
| NC score | 0.030305 (rank : 133) | θ value | 0.163984 (rank : 32) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 988 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P54762, O43569, O95142, O95143 | Gene names | EPHB1, EPHT2, NET | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 1 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor EPH-2) (NET) (HEK6) (ELK). | |||||
|
PHF8_HUMAN
|
||||||
| NC score | 0.029790 (rank : 134) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
EPHB2_HUMAN
|
||||||
| NC score | 0.029007 (rank : 135) | θ value | 0.279714 (rank : 36) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 983 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P29323, O43477, Q5T0U6, Q5T0U7, Q5T0U8 | Gene names | EPHB2, DRT, EPTH3, ERK, HEK5 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 2 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor EPH-3) (DRT) (Receptor protein-tyrosine kinase HEK5) (ERK) (NY-REN-47 antigen). | |||||
|
EPHB2_MOUSE
|
||||||
| NC score | 0.028984 (rank : 136) | θ value | 0.279714 (rank : 37) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 982 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P54763, Q62213, Q9QVY4 | Gene names | Ephb2, Epth3, Nuk, Sek3 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 2 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor EPH-3) (Neural kinase) (Nuk receptor tyrosine kinase) (SEK-3). | |||||
|
EPHA1_HUMAN
|
||||||
| NC score | 0.027878 (rank : 137) | θ value | 0.47712 (rank : 40) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P21709, Q15405 | Gene names | EPHA1, EPH, EPHT, EPHT1 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-A receptor 1 precursor (EC 2.7.10.1) (Tyrosine-protein kinase receptor EPH). | |||||
|
GON4L_MOUSE
|
||||||
| NC score | 0.027308 (rank : 138) | θ value | 1.06291 (rank : 47) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
I11RA_HUMAN
|
||||||
| NC score | 0.023808 (rank : 139) | θ value | 1.38821 (rank : 51) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14626, Q16542, Q7KYJ7 | Gene names | IL11RA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-11 receptor alpha chain precursor (IL-11R-alpha) (IL- 11RA). | |||||
|
FNDC3_MOUSE
|
||||||
| NC score | 0.023807 (rank : 140) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BX90, Q2VEY7, Q6ZQ15, Q811D3, Q8BTM3 | Gene names | Fndc3a, D14Ertd453e, Fndc3, Kiaa0970 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibronectin type-III domain-containing protein 3a. | |||||
|
VINEX_HUMAN
|
||||||
| NC score | 0.020128 (rank : 141) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60504, Q9UQE4 | Gene names | SORBS3, SCAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Vinexin (Sorbin and SH3 domain-containing protein 3) (SH3-containing adapter molecule 1) (SCAM-1). | |||||
|
CO9A1_HUMAN
|
||||||
| NC score | 0.017714 (rank : 142) | θ value | 2.36792 (rank : 59) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P20849, Q13699, Q13700, Q5TF52, Q6P467, Q96BM8, Q99225, Q9H151, Q9H152, Q9Y6P2, Q9Y6P3 | Gene names | COL9A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
COFA1_MOUSE
|
||||||
| NC score | 0.015923 (rank : 143) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O35206, Q9EQD9 | Gene names | Col15a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV)]. | |||||
|
CO9A1_MOUSE
|
||||||
| NC score | 0.015900 (rank : 144) | θ value | 2.36792 (rank : 60) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q05722, Q61269, Q61270, Q61433, Q61940 | Gene names | Col9a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
LPP_HUMAN
|
||||||
| NC score | 0.015623 (rank : 145) | θ value | 1.06291 (rank : 49) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
C1QC_MOUSE
|
||||||
| NC score | 0.014154 (rank : 146) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q02105 | Gene names | C1qc, C1qg | |||
|
Domain Architecture |

|
|||||
| Description | Complement C1q subcomponent subunit C precursor. | |||||
|
CO5A2_HUMAN
|
||||||
| NC score | 0.012918 (rank : 147) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
SEPT9_HUMAN
|
||||||
| NC score | 0.012704 (rank : 148) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UHD8, Q96QF3, Q96QF4, Q96QF5, Q9HA04, Q9UG40, Q9Y5W4 | Gene names | SEPT9, KIAA0991, MSF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (MLL septin-like fusion protein) (MLL septin-like fusion protein MSF-A) (Ovarian/Breast septin) (Ov/Br septin) (Septin D1). | |||||
|
COBA2_HUMAN
|
||||||
| NC score | 0.012167 (rank : 149) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
NFAC1_MOUSE
|
||||||
| NC score | 0.007953 (rank : 150) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88942, O70345 | Gene names | Nfatc1, Nfat2, Nfatc | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 1 (NFAT transcription complex cytosolic component) (NF-ATc1) (NF-ATc). | |||||
|
TXLNB_MOUSE
|
||||||
| NC score | 0.005773 (rank : 151) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
ZN687_HUMAN
|
||||||
| NC score | 0.002828 (rank : 152) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||