Please be patient as the page loads
|
PIWL2_HUMAN
|
||||||
| SwissProt Accessions | Q8TC59, Q96SW6, Q9NW28 | Gene names | PIWIL2, HILI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 2. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PIWL1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.979444 (rank : 3) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96J94, O95404, Q8NA60, Q8TBY5, Q96JD5 | Gene names | PIWIL1, HIWI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 1. | |||||
|
PIWL1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.979173 (rank : 4) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JMB7 | Gene names | Piwil1, Miwi | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 1. | |||||
|
PIWL2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8TC59, Q96SW6, Q9NW28 | Gene names | PIWIL2, HILI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 2. | |||||
|
PIWL2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.998227 (rank : 2) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CDG1, Q3TQE8, Q99MV6, Q9JMB6 | Gene names | Piwil2, Mili | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 2. | |||||
|
PIWL4_HUMAN
|
||||||
| θ value | 1.91371e-159 (rank : 5) | NC score | 0.975385 (rank : 5) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7Z3Z4, Q68CZ4, Q8N8G9, Q8N9V8, Q8NEH2 | Gene names | PIWIL4, HILI2, PIWI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 4. | |||||
|
PIWL3_HUMAN
|
||||||
| θ value | 4.13891e-146 (rank : 6) | NC score | 0.974598 (rank : 6) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7Z3Z3 | Gene names | PIWIL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 3. | |||||
|
PIWL4_MOUSE
|
||||||
| θ value | 3.76091e-131 (rank : 7) | NC score | 0.967299 (rank : 7) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CGT6, Q8CC75 | Gene names | Piwil4, Miwi2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 4 (mAgo5). | |||||
|
I2C1_HUMAN
|
||||||
| θ value | 2.19075e-38 (rank : 8) | NC score | 0.755682 (rank : 8) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UL18, Q5TA57, Q6P4S0 | Gene names | EIF2C1, AGO1 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 1 (eIF2C 1) (eIF-2C 1) (Argonaute-1) (Putative RNA-binding protein Q99). | |||||
|
I2C1_MOUSE
|
||||||
| θ value | 1.42001e-37 (rank : 9) | NC score | 0.753091 (rank : 10) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CJG1 | Gene names | Eif2c1, Ago1 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 1 (eIF2C 1) (eIF-2C 1) (Argonaute-1) (Piwi/argonaute family protein meIF2C1). | |||||
|
I2C3_MOUSE
|
||||||
| θ value | 1.01765e-35 (rank : 10) | NC score | 0.751861 (rank : 11) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CJF9 | Gene names | Eif2c3, Ago3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 3 (eIF2C 3) (eIF-2C 3) (Argonaute-3) (Piwi/argonaute family protein meIF2C3). | |||||
|
I2C3_HUMAN
|
||||||
| θ value | 2.96089e-35 (rank : 11) | NC score | 0.751116 (rank : 12) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9H9G7, Q5TA55, Q9H1U6 | Gene names | EIF2C3, AGO3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 3 (eIF2C 3) (eIF-2C 3) (Argonaute-3). | |||||
|
I2C2_MOUSE
|
||||||
| θ value | 5.0505e-35 (rank : 12) | NC score | 0.754067 (rank : 9) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CJG0, Q4VAB3, Q571J6 | Gene names | Eif2c2, Ago2, Kiaa4215 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 2 (eIF2C 2) (eIF-2C 2) (Argonaute-2) (Slicer protein) (Piwi/argonaute family protein meIF2C2). | |||||
|
I2C2_HUMAN
|
||||||
| θ value | 1.46948e-34 (rank : 13) | NC score | 0.748918 (rank : 14) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UKV8, Q8TCZ5, Q8WV58, Q96ID1 | Gene names | EIF2C2, AGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 2 (eIF2C 2) (eIF-2C 2) (Argonaute-2) (Slicer protein) (PAZ Piwi domain protein) (PPD). | |||||
|
I2C4_MOUSE
|
||||||
| θ value | 4.27553e-34 (rank : 14) | NC score | 0.749073 (rank : 13) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CJF8 | Gene names | Eif2c4, Ago4 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 4 (eIF2C 4) (eIF-2C 4) (Argonaute-4) (Piwi/argonaute family protein meIF2C4). | |||||
|
I2C4_HUMAN
|
||||||
| θ value | 5.584e-34 (rank : 15) | NC score | 0.745118 (rank : 15) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9HCK5 | Gene names | EIF2C4, AGO4, KIAA1567 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 4 (eIF2C 4) (eIF-2C 4) (Argonaute-4). | |||||
|
DSRAD_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 16) | NC score | 0.034428 (rank : 18) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99MU3, O70375, Q80UZ6, Q8C222, Q99MU2, Q99MU4, Q99MU7 | Gene names | Adar | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (RNA adenosine deaminase 1). | |||||
|
RHG04_HUMAN
|
||||||
| θ value | 0.62314 (rank : 17) | NC score | 0.013488 (rank : 21) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98171, Q14144 | Gene names | ARHGAP4, KIAA0131, RGC1, RHOGAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 4 (Rho-GAP hematopoietic protein C1) (p115). | |||||
|
DICER_HUMAN
|
||||||
| θ value | 1.81305 (rank : 18) | NC score | 0.069867 (rank : 16) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UPY3, O95943, Q9UQ02 | Gene names | DICER1, DICER, HERNA, KIAA0928 | |||
|
Domain Architecture |

|
|||||
| Description | Endoribonuclease Dicer (EC 3.1.26.-) (Helicase with RNase motif) (Helicase-MOI). | |||||
|
DICER_MOUSE
|
||||||
| θ value | 1.81305 (rank : 19) | NC score | 0.069318 (rank : 17) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R418 | Gene names | Dicer1, Dicer, Mdcr | |||
|
Domain Architecture |

|
|||||
| Description | Endoribonuclease Dicer (EC 3.1.26.-) (Double-strand-specific ribonuclease mDCR-1). | |||||
|
TRI62_HUMAN
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.006438 (rank : 25) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BVG3, Q9NVG0 | Gene names | TRIM62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 62. | |||||
|
TXTP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 21) | NC score | 0.008952 (rank : 23) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P53007, Q9BSK6 | Gene names | SLC25A1, SLC20A3 | |||
|
Domain Architecture |
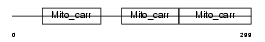
|
|||||
| Description | Tricarboxylate transport protein, mitochondrial precursor (Citrate transport protein) (CTP) (Tricarboxylate carrier protein) (Solute carrier family 25 member 1). | |||||
|
CXXC6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 22) | NC score | 0.017175 (rank : 19) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
CEP68_HUMAN
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.013561 (rank : 20) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q76N32, O60326, Q9BQ18, Q9UDM9 | Gene names | CEP68, KIAA0582 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 68 kDa (Cep68 protein). | |||||
|
TRI62_MOUSE
|
||||||
| θ value | 4.03905 (rank : 24) | NC score | 0.005956 (rank : 26) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80V85 | Gene names | Trim62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 62. | |||||
|
F120A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.009958 (rank : 22) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NZB2, O60649, Q14688, Q4VXF4, Q4VXF5, Q4VXG2, Q96I21, Q9NZB1 | Gene names | FAM120A, C9orf10, KIAA0183 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120A. | |||||
|
CECR2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.005009 (rank : 28) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BXF3, Q658Z4, Q96P58, Q9C0C3 | Gene names | CECR2, KIAA1740 | |||
|
Domain Architecture |

|
|||||
| Description | Cat eye syndrome critical region protein 2. | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | -0.000770 (rank : 30) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
DPOE2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.008263 (rank : 24) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O54956, Q6P3Y7, Q8BP72 | Gene names | Pole2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase epsilon subunit 2 (EC 2.7.7.7) (DNA polymerase II subunit 2) (DNA polymerase epsilon subunit B). | |||||
|
EFS_MOUSE
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.005214 (rank : 27) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64355 | Gene names | Efs, Sin | |||
|
Domain Architecture |
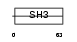
|
|||||
| Description | Embryonal Fyn-associated substrate (SRC-interacting protein) (Signal- integrating protein). | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | -0.001057 (rank : 31) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
SMOO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.004041 (rank : 29) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53814, O00569, O95769, O95937, Q9P1S8, Q9UIT1, Q9UIT2 | Gene names | SMTN, SMSMO | |||
|
Domain Architecture |

|
|||||
| Description | Smoothelin. | |||||
|
PIWL2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8TC59, Q96SW6, Q9NW28 | Gene names | PIWIL2, HILI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 2. | |||||
|
PIWL2_MOUSE
|
||||||
| NC score | 0.998227 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CDG1, Q3TQE8, Q99MV6, Q9JMB6 | Gene names | Piwil2, Mili | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 2. | |||||
|
PIWL1_HUMAN
|
||||||
| NC score | 0.979444 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96J94, O95404, Q8NA60, Q8TBY5, Q96JD5 | Gene names | PIWIL1, HIWI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 1. | |||||
|
PIWL1_MOUSE
|
||||||
| NC score | 0.979173 (rank : 4) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JMB7 | Gene names | Piwil1, Miwi | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 1. | |||||
|
PIWL4_HUMAN
|
||||||
| NC score | 0.975385 (rank : 5) | θ value | 1.91371e-159 (rank : 5) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7Z3Z4, Q68CZ4, Q8N8G9, Q8N9V8, Q8NEH2 | Gene names | PIWIL4, HILI2, PIWI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 4. | |||||
|
PIWL3_HUMAN
|
||||||
| NC score | 0.974598 (rank : 6) | θ value | 4.13891e-146 (rank : 6) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7Z3Z3 | Gene names | PIWIL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 3. | |||||
|
PIWL4_MOUSE
|
||||||
| NC score | 0.967299 (rank : 7) | θ value | 3.76091e-131 (rank : 7) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CGT6, Q8CC75 | Gene names | Piwil4, Miwi2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 4 (mAgo5). | |||||
|
I2C1_HUMAN
|
||||||
| NC score | 0.755682 (rank : 8) | θ value | 2.19075e-38 (rank : 8) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UL18, Q5TA57, Q6P4S0 | Gene names | EIF2C1, AGO1 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 1 (eIF2C 1) (eIF-2C 1) (Argonaute-1) (Putative RNA-binding protein Q99). | |||||
|
I2C2_MOUSE
|
||||||
| NC score | 0.754067 (rank : 9) | θ value | 5.0505e-35 (rank : 12) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CJG0, Q4VAB3, Q571J6 | Gene names | Eif2c2, Ago2, Kiaa4215 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 2 (eIF2C 2) (eIF-2C 2) (Argonaute-2) (Slicer protein) (Piwi/argonaute family protein meIF2C2). | |||||
|
I2C1_MOUSE
|
||||||
| NC score | 0.753091 (rank : 10) | θ value | 1.42001e-37 (rank : 9) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CJG1 | Gene names | Eif2c1, Ago1 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 1 (eIF2C 1) (eIF-2C 1) (Argonaute-1) (Piwi/argonaute family protein meIF2C1). | |||||
|
I2C3_MOUSE
|
||||||
| NC score | 0.751861 (rank : 11) | θ value | 1.01765e-35 (rank : 10) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CJF9 | Gene names | Eif2c3, Ago3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 3 (eIF2C 3) (eIF-2C 3) (Argonaute-3) (Piwi/argonaute family protein meIF2C3). | |||||
|
I2C3_HUMAN
|
||||||
| NC score | 0.751116 (rank : 12) | θ value | 2.96089e-35 (rank : 11) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9H9G7, Q5TA55, Q9H1U6 | Gene names | EIF2C3, AGO3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 3 (eIF2C 3) (eIF-2C 3) (Argonaute-3). | |||||
|
I2C4_MOUSE
|
||||||
| NC score | 0.749073 (rank : 13) | θ value | 4.27553e-34 (rank : 14) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CJF8 | Gene names | Eif2c4, Ago4 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 4 (eIF2C 4) (eIF-2C 4) (Argonaute-4) (Piwi/argonaute family protein meIF2C4). | |||||
|
I2C2_HUMAN
|
||||||
| NC score | 0.748918 (rank : 14) | θ value | 1.46948e-34 (rank : 13) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UKV8, Q8TCZ5, Q8WV58, Q96ID1 | Gene names | EIF2C2, AGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 2 (eIF2C 2) (eIF-2C 2) (Argonaute-2) (Slicer protein) (PAZ Piwi domain protein) (PPD). | |||||
|
I2C4_HUMAN
|
||||||
| NC score | 0.745118 (rank : 15) | θ value | 5.584e-34 (rank : 15) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9HCK5 | Gene names | EIF2C4, AGO4, KIAA1567 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2C 4 (eIF2C 4) (eIF-2C 4) (Argonaute-4). | |||||
|
DICER_HUMAN
|
||||||
| NC score | 0.069867 (rank : 16) | θ value | 1.81305 (rank : 18) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UPY3, O95943, Q9UQ02 | Gene names | DICER1, DICER, HERNA, KIAA0928 | |||
|
Domain Architecture |

|
|||||
| Description | Endoribonuclease Dicer (EC 3.1.26.-) (Helicase with RNase motif) (Helicase-MOI). | |||||
|
DICER_MOUSE
|
||||||
| NC score | 0.069318 (rank : 17) | θ value | 1.81305 (rank : 19) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R418 | Gene names | Dicer1, Dicer, Mdcr | |||
|
Domain Architecture |

|
|||||
| Description | Endoribonuclease Dicer (EC 3.1.26.-) (Double-strand-specific ribonuclease mDCR-1). | |||||
|
DSRAD_MOUSE
|
||||||
| NC score | 0.034428 (rank : 18) | θ value | 0.0193708 (rank : 16) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99MU3, O70375, Q80UZ6, Q8C222, Q99MU2, Q99MU4, Q99MU7 | Gene names | Adar | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (RNA adenosine deaminase 1). | |||||
|
CXXC6_HUMAN
|
||||||
| NC score | 0.017175 (rank : 19) | θ value | 3.0926 (rank : 22) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
CEP68_HUMAN
|
||||||
| NC score | 0.013561 (rank : 20) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q76N32, O60326, Q9BQ18, Q9UDM9 | Gene names | CEP68, KIAA0582 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 68 kDa (Cep68 protein). | |||||
|
RHG04_HUMAN
|
||||||
| NC score | 0.013488 (rank : 21) | θ value | 0.62314 (rank : 17) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98171, Q14144 | Gene names | ARHGAP4, KIAA0131, RGC1, RHOGAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 4 (Rho-GAP hematopoietic protein C1) (p115). | |||||
|
F120A_HUMAN
|
||||||
| NC score | 0.009958 (rank : 22) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NZB2, O60649, Q14688, Q4VXF4, Q4VXF5, Q4VXG2, Q96I21, Q9NZB1 | Gene names | FAM120A, C9orf10, KIAA0183 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120A. | |||||
|
TXTP_HUMAN
|
||||||
| NC score | 0.008952 (rank : 23) | θ value | 2.36792 (rank : 21) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P53007, Q9BSK6 | Gene names | SLC25A1, SLC20A3 | |||
|
Domain Architecture |

|
|||||
| Description | Tricarboxylate transport protein, mitochondrial precursor (Citrate transport protein) (CTP) (Tricarboxylate carrier protein) (Solute carrier family 25 member 1). | |||||
|
DPOE2_MOUSE
|
||||||
| NC score | 0.008263 (rank : 24) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O54956, Q6P3Y7, Q8BP72 | Gene names | Pole2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase epsilon subunit 2 (EC 2.7.7.7) (DNA polymerase II subunit 2) (DNA polymerase epsilon subunit B). | |||||
|
TRI62_HUMAN
|
||||||
| NC score | 0.006438 (rank : 25) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BVG3, Q9NVG0 | Gene names | TRIM62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 62. | |||||
|
TRI62_MOUSE
|
||||||
| NC score | 0.005956 (rank : 26) | θ value | 4.03905 (rank : 24) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80V85 | Gene names | Trim62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 62. | |||||
|
EFS_MOUSE
|
||||||
| NC score | 0.005214 (rank : 27) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64355 | Gene names | Efs, Sin | |||
|
Domain Architecture |

|
|||||
| Description | Embryonal Fyn-associated substrate (SRC-interacting protein) (Signal- integrating protein). | |||||
|
CECR2_HUMAN
|
||||||
| NC score | 0.005009 (rank : 28) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BXF3, Q658Z4, Q96P58, Q9C0C3 | Gene names | CECR2, KIAA1740 | |||
|
Domain Architecture |

|
|||||
| Description | Cat eye syndrome critical region protein 2. | |||||
|
SMOO_HUMAN
|
||||||
| NC score | 0.004041 (rank : 29) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53814, O00569, O95769, O95937, Q9P1S8, Q9UIT1, Q9UIT2 | Gene names | SMTN, SMSMO | |||
|
Domain Architecture |

|
|||||
| Description | Smoothelin. | |||||
|
MYH11_MOUSE
|
||||||
| NC score | -0.000770 (rank : 30) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | -0.001057 (rank : 31) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||